Abstract
Metabolite exchange mediates crucial interactions in microbial communities, significantly impacting global carbon and nitrogen cycling. Understanding these chemically-mediated interactions is essential for elucidating natural community functions and developing engineered synthetic communities. This study investigated membrane-separated bioreactors (mBRs) as a novel tool to identify transient metabolites and their producers/consumers in mixed microbial communities. We compared three co-culture methods (direct mixed, 2-chamber mBR, and 3-chamber mBR) to grow a synthetic binary community of the cyanobacterium Synechococcus elongatus PCC 7942 and the fungus Rhodotorula toruloides NBRC 0880, as well as axenic S. elongatus. Despite not being natural lichen constituents, these organisms exhibited interactions resembling those in cyanolichens. S. elongatus fixed CO2 into sugars as the primary shared metabolite, while R. toruloides secreted various biochemicals, predominantly sugar alcohols, mirroring the metabolite exchange observed in natural lichens. The mBR systems successfully captured metabolite gradients and revealed rapidly consumed compounds, including TCA cycle intermediates and amino acids. Our approach demonstrated that the 2-chamber mBR optimally balanced metabolite exchange and growth dynamics. This study provides insights into cross-species metabolic interactions and presents a valuable tool for investigating and engineering synthetic microbial communities with potential applications in biotechnology and environmental science.
Similar content being viewed by others
Introduction
Cyanobacteria play a crucial role in global carbon cycling, both in marine environments and terrestrial ecosystems. The cyanobacterium Synechococcusis one of the two most abundant phytoplankton genera dominating photosynthesis in the marine biological carbon pump1, contributing nearly one-fifth of global primary production of organic matter2,3. A substantial portion of this fixed carbon (estimated between 5% and 50%, with an average of 10–20%4,5,6,7,8,9) is directly released as dissolved organic matter (DOM), representing the major repository of organic carbon in the ocean10,11,12, and regulating the biosphere through carbon sequestration13,14.
In addition, cyanobacteria are essential components of lichens, another globally significant carbon-fixing system15,16. In cyanolichens, cyanobacteria fix CO2and share reduced carbon as energy-rich sugars with their fungal partners (mycobionts)17. Both marine and lichen systems demonstrate the critical role of cyanobacteria in supporting diverse heterotrophic communities through the release of organic carbon metabolites. These metabolites serve a role of primary carbon substrate for the associated heterotrophic microbiome but also include nutrients, exoproteins, allelochemicals, and other signaling molecules18,19,20,21, which drive the overall community structure and phenotype22,23,24,25,26.
In marine ecosystems, heterotrophic bacteria provide crucial feedback to cyanobacteria through cross-feeding of by-product metabolites. These metabolites can promote or inhibit phytoplankton growth27,28,29, enhance survival under nutrient starvation30,31or light deprivation32,33, protect against oxidative34,35and stationary-phase stresses29, and alleviate auxotrophy bottlenecks35. Similarly, in lichens, mycobionts transform cyanobacteria-provided sugars and secrete sugar alcohols (e.g., mannitol, arabitol, glycerol, erythritol)17,36,37,38,39,40,41as well as hundreds of other secondary metabolites42,43,44,45. Diverse by-product compounds from fungi improve desiccation tolerance, provide radiation screening, aid in thermal energy dissipation, trigger antioxidant defenses, and restore macromolecules and membranes46,47,48,49. Additionally, they facilitate communication50, quorum-sensing51, and signaling52,53, enabling lichens to thrive in extreme environments54,55.
The symbiotic relationships with heterotrophs have driven the evolution of mixotrophic and even heterotrophic capabilities in cyanobacteria like Synechococcus56,57,58. These adaptations and partnership with heterotrophic bacteria allow them to survive in challenging conditions, such as the extended darkness of deep euphotic zones. Cyanobacteria have developed an array of transporters59,60to assimilate various organic metabolites, including amino acids56,61,62, organosulfur compounds63,64,65, carboxylic acids, and nucleobases66,67, expanding their metabolic flexibility beyond autotrophy.
Indeed, Synecchococcusand other marine phytoplankton species evolve mixotrophy and even heterotrophy capabilities as a strategy to source carbon, nitrogen, sulfur, phosphorus and energy for survival in the extended darkness of deep euphotic zones, which hascontributed to their ecological success56,57,58. In this way, Synecchococcusis equipped with an array of transporters59,60to assimilate a variety of organic metabolites including amino-acids56,61,62, organosulfur compounds63,64,65, carboxylic acids and nucleobases66,67, which demonstrates that microbial interactions extend beyond the flux of reduced carbon from the photoautotroph to heterotrophic microbiome.
The robustness and metabolic capabilities of natural microbial communities have inspired the development of engineered multi-microbe systems, including “synthetic lichen” partnerships between cyanobacteria and fungi or heterotrophic bacteria. These systems show potential as bioplatforms for bioremediation68 or the generation of carbon-neutral bioproducts from CO269,70,71,72. Recent progress in genome-scale modeling has further elucidated cross-feeding and metabolic exchanges in these synthetic communities73,74. Synthetic lichens not only have significant translational potential across the fields of carbon sequestration, biological materials, and bioremediation, but also serve as valuable tools for understanding the biology of symbiosis. By constructing these synthetic systems, researchers can generate insights into the complex interactions and metabolic processes occurring in natural lichens, demonstrating the utility of synthetic biology in advancing fundamental science. Moreover, these engineered communities have the potential to enable sustainable biotechnologies that address major global challenges while simultaneously deepening our understanding of biological systems.
Despite these advances, we have yet to fully uncover the intricate metabolic interactions in both natural and synthetic communities. These interactions form the basis of mutualistic, commensal, parasitic, or competitive relationships among species, which can dynamically shift depending on nutrient availability and coadaptation30,75,76,77. A more comprehensive understanding of these interactions is crucial for harnessing and controlling microbial community functions in ecological, industrial, and biotechnological applications70,78,79.
Collecting comprehensive microbial community exometabolomics presents several challenges, including distinguishing between metabolites that are secreted but not consumed (accumulating in the growth medium) and those that are both secreted and consumed by partner organisms (exchanged metabolites), assigning metabolites to specific partners, and detecting rapidly consumed compounds. To address these issues, we developed and tested membrane-separated bioreactor systems (mBRs) as a tool to characterize metabolite exchange in a synthetic lichen community. Crucially, the membrane separation allows for the detection of rapidly consumed compounds by creating a diffusion barrier. In a well-mixed system, these compounds might be consumed as quickly as they are produced, leaving no detectable trace. However, in the mBR system, the membrane slows the diffusion of metabolites between compartments, allowing for the accumulation and detection of these otherwise elusive compounds in the producer’s compartment before they are fully consumed by the partner organism. This novel approach enables the establishment of concentration gradients for secreted metabolites, revealing diffusion flux and directional exchange between species in co-culture, which is often challenging to discern in well-mixed systems.
In this study, we established a synthetic binary community consisting of a sucrose-secreting Synechococcus elongatus strain PCC 7942 CscB/SPS strain and the basidiomycetous fungus Rhodotorula toruloides strain NBRC 0880. This unique pairing allows us to explore the potential for establishing synthetic symbioses between organisms from different ecosystems but with potentially complementing metabolic capabilities. By elucidating complex interactions and metabolic exchanges in our synthetic system, we aim to gain insights into the mechanisms driving community dynamics and ecosystem functioning in natural environments. Furthermore, this study contributes to the design and optimization of synthetic ecology bioproduction systems for converting CO₂ into non-fossil fuel chemicals, potentially revolutionizing our approach to carbon-neutral chemical synthesis.
Results
Growth of axenic S. elongatus and different S. elongatus – R. Toruloides co-culture spatial configurations
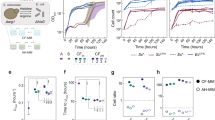
To investigate the interactions between Synechococcus elongatus and Rhodotorula toruloides, their effects on microbial growth, and the role of spatial separation, we compared the growth of the metabolically engineered sucrose-producing cyanobacterium S. elongatus under four conditions: (1) axenic culture, (2) same-volume mixed co-culture with R. toruloides, (3) spatially separated co-culture in a 2-unit membrane bioreactor (2-mBR; Fig. 1A), and (4) spatially separated co-culture in a 3-unit membrane bioreactor (3-mBR; Fig. 1B). In these co-cultures, S. elongatus fixes CO2 and converts it to sucrose, which is secreted and provided to R. toruloides as the principal source of organic carbon. Additional information on the cultivation systems used, including a manufacturing schematic of the custom-made membrane bioreactor, is provided in Supplementary Figures S1-S3.
During the initial 75 h of incubation, all experimental configurations showed only minor differences in S. elongatus growth. However, S. elongatus cultivated with R. toruloides in the spatially separated 2-mBR reached the highest cell density of 1.5 × 109 cells per mL by the end of experiment (Fig. 1C), which was 7% higher than the spatially separated 3-mBR and 13% higher than axenic S. elongatus culture with the mixed co-culture the lowest of all 4 conditions. These results suggest that the 2-mBR configuration allows for beneficial interactions between the two species while minimizing potential negative effects. The enhanced growth in the 2-mBR relative to the mixed culture implies that spatial separation may facilitate positive metabolic exchanges while reducing direct competition or inhibition.
The growth of S. elongatus in the 3-mBR, while better than in axenic and mixed cultures, was lower than in the 2-mBR. This suggests that the additional diffusion barrier in the 3-mBR may impede some beneficial interactions. This hypothesis is further supported by the significantly delayed growth of R. toruloides in the 3-mBR setup compared to both the mixed co-culture and the 2-mBR co-culture (Fig. 1D). A minor reduction in growth of S. elongatus in the mixed co-culture compared to spatially separated conditions indicates that direct contact between the species may lead to minor antagonistic interactions. These could include competition for resources, alterations in the microenvironment, or contact-dependent inhibition.
To further elucidate the nature of these interactions and to validate that the membrane bioreactor configurations allow us to capture concentration gradients for metabolites produced by one partner and consumed by another, we analyzed sucrose content profiles in the different experimental setups (Fig. 1E and F). As expected, sucrose steadily accumulated in axenic S. elongatus cultures but was nearly completely consumed by R. toruloides in mixed co-culture after 75 h of cultivation. Both mBR configurations resulted in the establishment of sucrose concentration gradients across the membrane, with S. elongatus as the producer and a gradual reduction moving towards the R. toruloides compartment. This gradient formation demonstrates the occurrence of metabolite exchange between the two species in the spatially separated systems.
These findings highlight the complex nature of interactions between S. elongatus and R. toruloides, and demonstrate that spatial separation can significantly modulate these interactions, affecting growth patterns and metabolite exchange. To further characterize these interactions at the metabolic level, we conducted a comprehensive analysis of the exometabolomes from all cultures.
Spatially separated and mixed co-cultivation of S. elongatus and R. toruloides, with comparison to axenic S. elongatus culture. (A) 2-unit membrane bioreactor (2-mBR) and (B) 3-unit membrane bioreactor (3-mBR) setups. Detailed schematics for mBR and mixed bioreactor are provided in Supplementary Information Figures S1-S3. (C, D) Cell density comparisons: ‘Se 2-mBR’ and ‘Rt 2-mBR’ – S. elongatus and R. toruloides in 2-mBR, respectively; ‘Se 3-mBR’ and ‘Rt 3-mBR’ – S. elongatus and R. toruloides in 3-mBR, respectively; ‘Se mixed’ and ‘Rt mixed’ – S. elongatus and R. toruloides in same-volume mixed co-culture; ‘Se axenic’ – S. elongatus grown alone. (E, F) Extracellular sucrose content in different mBR units (bars), mixed co-culture (‘Se-Rt mixed’, red diamonds), and axenic S. elongatus culture (‘Se axenic’, green circles). In mixed co-culture, cells interact directly in a single volume. Note: Axenic S. elongatus culture confirms sucrose production and serves as a reference for potential production levels. In ‘Rt 2-mBR’, ‘Rt 3-mBR’ and late ‘Se-Rt mixed’, sucrose levels remain low due to rapid consumption by R. toruloides. Sucrose gradients in mBRs illustrate the production-consumption balance. Data points represent mean ± standard deviation of three biological replicates.
Profiling of exchange metabolites in the middle growth phase of S. elongatus – R. toruloides co-culture
To characterize the metabolic interactions between S. elongatus – R. toruloides and evaluate the impact of spatial separation on these interactions, we analyzed the exometabolomes of the various culture configurations during the middle growth phase of the co-culture (101 h of cultivation).
High-resolution Q-TOF-MS was employed to identify extracellular metabolites. The resulting metabolomics data underwent pre-processing, including sample normalization by sum, Log10 transformation, and Pareto scaling (see Supplementary Figures S4-S7). Principal component analysis (PCA) of the resulting data revealed distinct groupings of samples based on their metabolite profiles (Fig. 2). Exometabolomes from S. elongatus units of both 2- and 3-unit mBRs clustered tightly with the axenic S. elongatus samples in the lower right quadrant of the PCA plot, indicating similar metabolite profiles. Likewise, exometabolomes from R. toruloides units of both mBR configurations showed high similarity. As expected, samples from the middle unit of the 3-unit mBR were positioned between the S. elongatus and R. toruloides clusters. Notably, the exometabolome of the S. elongatus – R. toruloides same-volume mixed co-culture was distinct from all other samples, potentially due to rapid metabolite utilization or the production of specific metabolites resulting from direct cell-to-cell contact.
Principal Component Analysis (PCA) of exometabolome profiles at 101 h for cultures of axenic S. elongatus (Se axenic), S. elongatus units of the membrane separated bioreactors (Se 2-mBR and Se 3-mBR, respectively), R. toruloides inits of the mBRs (Rt 2-mBR and Rt 3-mBR, respectively), middle unit of the 3-unit mBR (mddl 3-mBR), and S. elongatus – R. toruloides same-volume mixed co-culture (Se-Rt mixed). Each circle represents a biological replicate (n = 3 for each condition).
To elucidate relationships and co-variation between extracellular metabolites, we calculated a pairwise Pearson correlation matrix and visualized the associations among 99 identified metabolites using a hierarchically clustered heatmap (Supplementary Figure S8). We further analyzed significant correlations (|R| > 0.7, p< 0.05) using network analysis in Cytoscape80with the Girvan-Newman fast greedy clustering algorithm81. The resulting metabolite correlation network comprised 83 nodes with 259 edges, grouped into five clusters of more than 3 nodes (Fig. 3). This network topology analysis can reveal non-intuitive metabolic relationships and identify metabolites of central importance82,83.
Metabolites with high centrality that position as “bottlenecks” bridging large groups of metabolites or as “hubs” directly connected to a large number of other metabolites are often those of principle importance. Top 20 metabolites with highest betweenness centrality (“bottleneck”) and degree (“hub”) metabolites are shown in Table 1. As anticipated, sucrose, the primary carbon source exchanged between S. elongatus and R. toruloides, was among the top metabolites in both centrality measures. Other important metabolites included sugar derivatives (myo-inositol, D-ribose 5-phosphate, methyl-β-D-galactoside, and iso-erythritol), carboxylic acids (3-hydroxy-isobutanoate and putative oxalate), and the amino acid valine.
Network analysis of extracellular metabolite correlations at 101 h. The network, visualized using Cytoscape, summarizes significant pairwise correlations (Pearson |R| > 0.7, p < 0.05) among extracellular metabolites. Nodes represent individual metabolites, with colors indicating betweenness centrality. Edges represent correlations, with blue for positive and red for negative correlations. Edge width corresponds to correlation strength. Dashed edges indicate correlations between metabolites in different clusters. The network is divided into five major clusters (> 3 nodes each) using Cytoscape’s GLay plugin. Metabolite identification confidence: No symbol: high (score > 0.75); *: moderate (score < 0.75); **: putative (best spectral match).
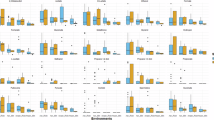
Hierarchical clustering analysis revealed three principal clusters of extracellular metabolites based on their relative abundance across different samples (top 40 based on T-test are shown in Fig. 4 and full list can be found in Supplementary Figure S9-S10). Metabolites in the Cluster I included sucrose as well as other sugars and glycosides (e.g. arabinose, maltose, glucosyl glycerol, xylitol, arbutin, N-acetyl-D-mannosamine). They were found in high abundance in all S. elongatus cultures including S. elongatus – R. toruloides mixed co-cultures, and lowest abundance in the R. toruloides unit of the 3-unit mBR. These metabolites corresponded to Cluster 2 in the correlation network (Fig. 3) and were likely produced and excreted by the cyanobacterium. Contrastingly, Cluster III on the heatmap (Fig. 4) included diverse list of metabolites with higher abundance in the R. toruloides units of the mBRs, but low abundance in the presence of S. elongatus including S. elongatus – R. toruloides mixed co-culture. This pattern suggests production by the yeast and potential consumption by the cyanobacterium. These metabolites were predominantly found in Cluster 5 of the correlation network. Finally, metabolites in the Cluster II had highest abundance in the S. elongatus – R. toruloides same-volume mixed co-culture implying that they can be either produced by both species or their production can be induced by physical cell-to-cell contact in the mixed co-culture.
Hierarchical clustering of top extracellular metabolites at 101 h. The heatmap shows hierarchical clustering analysis of selected untargeted extracellular metabolites (top 40 based on T-test). Clustering was performed using Euclidean distance measure and Ward’s algorithm. Color intensity corresponds to the normalized relative abundance of each metabolite across different experimental conditions. Metabolite identification confidence: No symbol: high (score > 0.75); *: moderate (score < 0.75); **: putative (best spectral match).
We further analyzed the effect of spatial separation of S. elongatus and R. toruloides on the number and identity of differentially abundant exometabolites (log2 FC > 1, padjusted < 0.05) after 101 h of incubation (Fig. 5). Comparisons were made between axenic S. elongatus culture and mixed co-culture (Fig. 5A), and between S. elongatus and R. toruloides units in both 2-unit (Fig. 5B) and 3-unit (Fig. 5C) mBRs. Interestingly, while S. elongatus was the primary organic carbon supplier, a larger number of differentially abundant exometabolites were associated with R. toruloides, particularly in mixed co-culture and 2-unit mBR configurations.
Effect of spatial separation of the S. elongatus and R. toruloides in co-cultures on a number and specifics of the differentially abundant exometabolites (log2 FC > 1, padjusted < 0.05) after 101 h of incubation is shown for in Fig. 5 for axenic S. elongatus culture vs. S. elongatus – R. toruloides same-volume mixed co-culture (Fig. 5A), S. elongatus unit vs. R. toruloides unit of a 2-unit mBR (Fig. 5B), S. elongatus unit vs. R. toruloides unit of a 3-unit mBR (Fig. 5C). While cyanobacterium was the primary organic carbon supplier in this binary synthetic community, larger number of differentially abundant exometabolites were observed in the broth with R. toruloides especially for mixed co-culture and 2-unit mBR.
The complete set of statistically significant differentially abundant metabolites is presented in three-way Edwards–Venn diagrams (Fig. 5D-E). Functional enrichment analysis using the hypergeometric test revealed that these metabolites significantly overrepresented several metabolic pathways in both organisms, including the TCA cycle, Glyoxylate and dicarboxylate metabolism, Pentose phosphate pathways (PPP), metabolism of various amino-acids, Fructose metabolism (Fruc) in yeast, Butanoate metabolism and Carbon fixation in cyanobacteria.
These results highlight the complex metabolic interactions between S. elongatus and R. toruloides and demonstrate that spatial separation can significantly influence the production and exchange of metabolites in this synthetic community.
Differential exometabolite analysis at 101 h across cultivation conditions. (A-C) Volcano plots showing differential content of exometabolites for: (A) axenic S. elongatus culture vs. S. elongatus – R. toruloides same-volume mixed co-culture; (B) S. elongatus unit vs. R. toruloides unit of a 2-unit mBR; (C) S. elongatus unit vs. R. toruloides unit of a 3-unit mBR. Red dots indicate metabolites with statistically significant differential abundance (|log2 FC| > 1 and padj < 0.1); Negative and positive log2 FC likely correspond to metabolites associated with production by S. elongatus and R. toruloides, respectively. (D and E) Three-way Edwards–Venn diagrams showing metabolites with (D) higher (likely associated with production by S. elongatus) and (E) lower (likely associated with production by R. toruloides) content (log2 FC > 1, padj < 0.1) in axenic S. elongatus and S. elongatus mBR units compared to mixed co-culture and R. toruloides mBR units, respectively. (F) Functional enrichment (Hypergeometric Test) and pathway topology analysis (relative betweenness centrality based on KEGG metabolic pathways) of differential metabolites. Metabolite identification confidence: No symbol: high (score > 0.75); *: moderate (score < 0.75); **: putative (best spectral match).
Profiling of exchange metabolites in established mature co-culture of S. elongatus – R. toruloides
To investigate the progression of metabolic interactions between S. elongatus and R. toruloides over time, we analyzed exometabolome profiles at 173 h of cultivation, representing the established mature phase of the co-culture.
Principal component analysis (PCA) of the 173-hour exometabolome profiles (Fig. 6) revealed patterns similar to those observed at 101 h, with substantial differences between S. elongatus and R. toruloides units in both 2- and 3-unit mBRs. Profiles from S. elongatus units across mBR configurations clustered together, as did those from R. toruloides units, indicating high similarity of the exometabolome profiles. Notably, profilies from axenic S. elongatus cultures and S. elongatus – R. toruloides mixed co-cultures positioned discretely, reflecting their distinct metabolic profiles.
Network analysis of significant metabolite associations (Pearson |R| > 0.73, p < 0.05) yielded a correlation network of 60 nodes and 247 edges, grouped into three connected clusters (49 nodes) and one stand-alone cluster (5 nodes) (Fig. 7). Full matrix of hierarchically clustered correlation coefficients is visualized as a heatmap in Supplementary Figure S11. Clusters 1 and 2 dominated the network, representing > 85% of nodes and > 75% of edges, with exclusively positive correlations. In contrast, cluster 3 exhibited negative intra-connections. Valine emerged as the top “bottleneck” (highest betweenness centrality) and one of the top “hub” (highest degree) metabolites (Table 2). Other key metabolites included sugar alcohols (erythritol, ribitol, xylitol, galactitol, and acetol/hydroxyacetone) and carboxylic acids (fumarate, succinate, oxalate, and glycolate).
Principal Component Analysis (PCA) of exometabolome profiles at 173 h for cultures of axenic S. elongatus (Se axenic), S. elongatus units of the membrane separated bioreactors (Se 2-mBR and Se 3-mBR, respectively), R. toruloides inits of the mBRs (Rt 2-mBR and Rt 3-mBR, respectively), middle unit of the 3-unit mBR (mddl 3-mBR), and S. elongatus – R. toruloides same-volume mixed co-culture (Se-Rt mixed). Each circle represents a biological replicate (n = 3 for each condition).
Network analysis of extracellular metabolite correlations at 173 h. The network, visualized using Cytoscape, summarizes significant pairwise correlations (Pearson |R| > 0.7, p < 0.05) among extracellular metabolites. Nodes represent individual metabolites, with colors indicating betweenness centrality. Edges represent correlations, with blue for positive and red for negative correlations. Edge width corresponds to correlation strength. Dashed edges indicate correlations between metabolites in different clusters. The network is divided into five major clusters (> 3 nodes each) using Cytoscape’s GLay plugin. Metabolite identification confidence: No symbol: high (score > 0.75); *: moderate (score < 0.75); **: putative (best spectral match).
Hierarchical clustering analysis revealed six distinct groups of extracellular metabolites based on their relative abundance across samples (top 40 based on T-test are shown in Fig. 8 and full list can be found in Supplementary Figure S9-S10). Clusters I, II, and III contained metabolites with highest abundance in all cultures with R. toruloides and lowest in axenic S. elongatus, suggesting yeast origin. However, there were some vatiations potentally resulting from differences in utilization of certain metabolites by cyanobacteria. Cluster I, peaking in mixed co-cultures, included sugar alcohols (arabinitol, mannitol, galactitol, ribitol and erythritol), TCA intermediates (succinate, fumarate, and malate), and aromatic metabolites (phenylalanine and 2-coumarate or 2-hydroxycinnamic acid), indicating potential stimulation of their production by close proximity of S. elongatus Accumulation of these metabolites in the mixed co-culture suggests their unlikely utilization by the cyanobacteria or augmented secretion by R. toruloides in the presence of S. elongatus. In contrast, Clusters II and III included metabolites not peaking in mixed co-cultures, including other TCA metabolites (oxalate and ketoglutarate), amino acids (valine and threonine), D-ribose-5-phosphate, heptadecanoic acid and few other. Cluster II metabolites showed lower abundance in mixed cultures compared to R. toruloides units of the mBRs, suggesting utilization by S. elongatus. Cluster III metabolites had relatively even abondance across all cultures with R. toruloides including S. elongatus – R. toruloides co-culture implying either reduced or absence of consumption by cyanobacteria. Cluster IV included sugars secreted by S. elongatus and robustly utilized by yeast, including sucrose, maltose, and arbutin. Finally, Cluster VI metabolites were in highest abundance in mixed co-cultures followed by axenic S. elongatus, suggesting cyanobacterial production stimulated by yeast proximity.
Hierarchical clustering of top extracellular metabolites at 173 h. The heatmap shows hierarchical clustering analysis of selected untargeted extracellular metabolites (top 40 based on T-test). Clustering was performed using Euclidean distance measure and Ward’s algorithm. Color intensity corresponds to the normalized relative abundance of each metabolite across different experimental conditions. Metabolite identification confidence: No symbol: high (score > 0.75); *: moderate (score < 0.75); **: putative (best spectral match).
The presence of a semipermeable barrier (0.2 μm membrane) significantly affected the number of differentially abundant exometabolites (log2 FC > 1, padjusted < 0.1) at 173 h. Notably, sucrose was the only differentially abundant exometabolite in axenic S. elongatus versus mixed co-culture (Fig. 9A and D). The mBR systems revealed additional differentially abundant metabolites in S. elongatus units, including glucose, maltose, arbutin, and other sugar derivatives (putative lactose, glucosamine phosphate, acetylmannosamine and gluconolactone) (Fig. 9B and C).
R. toruloides cultures exhibited a significantly larger number of differentially abundant exometabolites compared to S. elongatus cultures. We identified 27 exometabolites consistently present in all three types of cultures with R. toruloides cells and an additional 21 in at least two (Fig. 9D and E). These metabolites primarily comprised TCA intermediates (succinate, fumarate, oxalate and malate), sugar alcohols (mannitol, erythritol, arabitol, acetol, and glycerol), carboxylic (glycolate, hydroxypropanoate, lactate, hydroxyisobutyrate, pimelate) and amino acids (glycine, valine, threonine).
In contrast, a relatively large number of exometabolites with statistically significant increased abundance were identified in cultures with R. toruloides cells compared to corresponding S. elongatus cultures. Specifically, 27 exometabolites were consistently identified in all three cultures with R. toruloides and 21 more were identified in at least two of the yeast cultures. All differentially abundant metabolites are shown for all three cultivation configurations on the three-way Edwards–Venn diagrams in Fig. 5D (higher abundance in S. elongatus units) and Fig. 5D (higher abundance in R. toruloides units). Major groups of metabolites were represented by the TCA intermediates (succinate, fumarate, oxalate and malate), sugar alcohols (mannitol, erythritol, arabitol, acetol, and glycerol), carboxylic (glycolate, hydroxypropanoate, lactate, hydroxyisobutyrate, pimelate) and amino acids (glycine, valine, threonine). In addition to metabolites COG’s TCA, Glyoxylate and dicarboxylate, PPP, amino-acids, Butanoate and Carbon fixation pathways identified in 101 h samples, the exometabolites in 173 h sample overrepresented COG’s Aminoacyl-tRNA biosynthesis, Starch and sucrose, Glycerolipid, and Pyruvate metabolism pathways (Fig. 9F).
Pathway enrichment analysis of differentially abundant metabolites at 173 h revealed overrepresentation of several metabolic pathways beyond those identified at 101 h, including Aminoacyl-tRNA biosynthesis, Starch and sucrose metabolism, Glycerolipid metabolism, and Pyruvate metabolism (Fig. 9F).
These results demonstrate the dynamic nature of metabolic interactions between S. elongatus and R. toruloides over time, and highlight the impact of spatial separation on metabolite exchange in this synthetic community.
Differential exometabolite analysis at 173 h across cultivation conditions. (A-C) Volcano plots showing differential content of exometabolites for: (A) axenic S. elongatus culture vs. S. elongatus – R. toruloides same-volume mixed co-culture; (B) S. elongatus unit vs. R. toruloides unit of a 2-unit mBR; (C) S. elongatus unit vs. R. toruloides unit of a 3-unit mBR. Red dots indicate metabolites with statistically significant differential abundance (|log2 FC| > 1 and padj < 0.1); Negative and positive log2 FC likely correspond to metabolites associated with production by S. elongatus and R. toruloides, respectively. (D-E) Three-way Edwards–Venn diagrams showing metabolites with (D) higher (likely associated with production by S. elongatus) and (E) lower (likely associated with production by R. toruloides) content (log2 FC > 1, padj < 0.1) in axenic S. elongatus and S. elongatus mBR units compared to mixed co-culture and R. toruloides mBR units, respectively. (F) Functional enrichment (Hypergeometric Test) and pathway topology analysis (relative betweenness centrality based on KEGG metabolic pathways) of differential metabolites. Metabolite identification confidence: No symbol: high (score > 0.75); *: moderate (score < 0.75); **: putative (best spectral match).
Discussion
Microbial communities, both natural and engineered, represent complex and powerful biological systems that play crucial roles in global biogeochemical cycles, waste management, and even human health. While advances in genomics have enabled the identification and quantification of species within these communities, understanding the intricate metabolic interactions that drive community dynamics remains challenging. This is particularly true for extracellular metabolites, which are often difficult to attribute to specific community members due to the shared metabolic capabilities of many microorganisms.
In this study, we deployed 2-unit and 3-unit membrane bioreactor (mBR) systems to investigate cross-feeding and interactions in a synthetic lichen partnership between the photoautotrophic cyanobacterium Synechococcus elongatus and the heterotrophic yeast Rhodotorula toruloides. This novel approach allowed us to capture the directionality of metabolite flow between community members as diffusion-limited gradients and to detect transient extracellular metabolites that might otherwise go unnoticed in well-mixed co-cultures.
While similar approaches using microplate-based devices like SubTap and BioMe84,85 have been developed for high-throughput screening of microbial interactions, our mBR system offers distinct advantages for certain types of studies. Unlike microplate-based systems, which are limited by small sample volumes, our mBRs provide larger volumes that allow for time-course studies and multi-omics analyses requiring substantial sample quantities. Additionally, the larger volume in our mBRs mitigates issues related to evaporation that can affect microplate-based systems during extended experiments.
One of the most striking observations in our study was the emergence of carbon exchange patterns in the mature synthetic co-culture that resembled those found in natural cyanolichens. S. elongatus fixed CO2 and shared reduced carbon with its fungal partner primarily in the form of sugars, including not only the engineered sucrose secretion but also glucose, arabinose, maltose, and other sugar derivatives. In return, R. toruloidessecreted a variety of secondary metabolites, with sugar alcohols being particularly prominent. Indeed, sugar alcohols like mannitol, arabitol, erythritol, and ribitol were reported to be the main secreted carbon metabolites secreted by fungi in cyanolichens17,36,37,38,39,40,41. Therefore, the observed exchange of sugars for sugar alcohols mirrors the fundamental cross-transfer observed in natural cyanolichens, suggesting that this may be a conserved pattern in cyanobacteria-fungi interactions. It is important to note that while natural lichens function under dynamic conditions including diurnal cycles, fluctuating light intensities, alternating periods of desiccation and hydration, and seasonal changes, our current study employed constant light conditions86,87,88. This simplification allowed us to establish a baseline for metabolic exchange in our synthetic system. Future studies incorporating light-dark cycles would be valuable to more closely mimic natural conditions and potentially reveal additional insights into the temporal dynamics of metabolite exchange in cyanobacteria-fungi partnerships.
The use of mBR systems proved particularly valuable in identifying rapidly consumed metabolic intermediates that mediate carbon flow through the community. During the early growth phase of the heterotroph, we detected higher concentrations of organic acids (such as glycolic acid, pimelic acid, lactic acid, citric acid, and oxalic acid) in the R. toruloides chambers of the mBRs. This suggests that these compounds are produced by the yeast and rapidly consumed by S. elongatus, potentially serving as additional carbon sources beyond CO2. Conversely, during late-stage growth, sugars produced by S. elongatus (including sucrose, maltose, and trehalose) were found at significantly lower concentrations in the R. toruloides chambers compared to the S. elongatus chambers, suggesting that sugar production by the cyanobacteria may become a limiting factor for yeast growth in the later stages of co-culture.
In general, comparing the exometabolomes at 101 and 173 h revealed dynamic shifts in metabolite profiles over time. While sugar exchange remained prominent throughout, we observed an increase in the diversity and abundance of sugar alcohols and TCA cycle intermediates in the later time point, particularly in R. toruloides cultures. This shift suggests a maturation of the metabolic relationship between the two organisms, with R. toruloides potentially adapting its metabolism to more efficiently process the increasing flux of carbon sources provided by S. elongatus as the culture density increased. The greater abundance and diversity of yeast-produced exometabolites at 173 h likely reflects both this increased sugar availability and the yeast’s metabolic adaptation to the co-culture environment over time.
The mBR approach also allowed us to establish concentration gradients for various metabolites, providing insights into the directionality of metabolite flows without the need for costly labeled substrates. We found that approximately 33% (69 out of 107) of the detected metabolites exhibited significant concentration differences (log2 FC > 1, p < 0.01) between the R. toruloides and S. elongatus chambers in at least one experimental setup. Of these, 13 metabolites exhibited significant differential gradients in both the 2- and 3-mBR systems with 100% agreement in directionality of the gradient between systems. For metabolites where only one system identified a significant directional gradient, the other mBR was in soft agreement (same direction but non-significant) in 85% (48/56) of the observations, which underscores the reproducibility and reliability of this approach. Across the middle and late time-points, seven metabolites exhibited high-confidence significant gradients across experimental systems including sucrose, maltose, and proline produced by S. elongatus and a variety of modified sugars produced by R. toruloides (ethyl-glucuronide, methyl-beta-D-galactopyranoside, and ribose-5-phosphate) demonstrating carbon flow in both directions. Notably, both sucrose and maltose were specifically lower in concentration during mixed co-cultivation during late-stage growth of the yeast partner. These results demonstrate that membrane separated bioreactors aid identification of metabolites representing major forms of organic carbon exchanged by microbial partners and detection of the primary producer from concentration gradients.
Our findings have implications for understanding both natural and synthetic microbial communities. In natural systems, such as aquatic environments and cyanolichens, the identified metabolite exchange patterns may help explain the stability and resilience of these communities. For instance, the ability of cyanobacteria to utilize TCA intermediayes and amino-acids produced by heterotrophs may contribute to their survival during periods of limited light availability in deep euphotic zones56,57,58,61,62,66. Receiving TCA intermediates and amino-acids in return for sugars could be another cross-feeding pattern between phototrophs and heterotrophs developed through adaptive evolution.
From a synthetic biology perspective, our results demonstrate the potential for creating artificial symbioses between diverse organisms that do not naturally form lichens. This opens up possibilities for designing novel synthetic communities with tailored metabolic capabilities. The mBR approach we developed could be a valuable tool for optimizing these synthetic systems, allowing for fine-tuning of metabolite exchange and community composition.
Moreover, our study contributes to the broader field of synthetic ecology and its potential applications in biotechnology. By elucidating the complex interactions and metabolic exchanges in our synthetic system, we provide insights that could inform the design of more efficient and robust microbial consortia for sustainable bioproduction. For example, the observed patterns of sugar and organic acid exchange could be exploited to develop systems for converting CO2 or waste organic carbon into value-added chemicals or biofuels .
However, it’s important to note that while our 3-mBR system allowed for the detection of additional metabolites due to increased diffusion time, it also further limited metabolite exchange and impacted partner species growth. This highlights the need for careful consideration of experimental design when studying time-dynamic metabolic fluxes in microbial communities.
In conclusion, our study demonstrates the power of membrane-separated bioreactor systems in unraveling the complex metabolic interactions within microbial communities. By providing a means to capture transient metabolites and establish concentration gradients, this approach offers new possibilities for studying and engineering microbial consortia. Future work could extend this method to more complex communities or focus on engineering specific metabolic exchanges based on the patterns observed here. Ultimately, these insights could lead to the development of more sophisticated synthetic microbial communities with applications ranging from bioremediation to the production of novel biomaterials.
Conclusions
This study demonstrates the efficacy of membrane-separated bioreactor systems in elucidating complex metabolic interactions within synthetic microbial communities. By employing 2-unit and 3-unit mBRs, we successfully captured the directionality of metabolite flow and detected transient extracellular metabolites in a synthetic lichen partnership between Synechococcus elongatus and Rhodotorula toruloides. Our findings reveal striking similarities between the carbon exchange patterns in this synthetic system and those observed in natural cyanolichens, particularly the trade of sugars for sugar alcohols.
The mBR approach proved invaluable in identifying rapidly consumed metabolites and establishing concentration gradients, offering insights into metabolite flows without the need for labeled substrates. This method’s reproducibility and ability to capture time-dynamic metabolic fluxes underscore its potential as a powerful tool for studying and engineering microbial consortia.
Our results have significant implications for understanding both natural microbial communities and designing synthetic ones. The observed metabolite exchange patterns may explain the stability of natural systems and inform the development of more efficient synthetic communities for biotechnological applications. Future research could extend this approach to more complex communities or focus on engineering specific metabolic exchanges, potentially leading to novel solutions in bioremediation, sustainable bioproduction, and the creation of new biomaterials.
Materials and methods
Culture media and maintaining
Sucrose secreting Synechococcus elongatus PCC 7942 cscB/SPS strain (S. elongatus) engineered to overexpress sucrose phosphate synthase (SPS) and sucrose permease (CscB) was generously provided by Dr. Daniel Ducat89. Rhodotorula toruloides (formerly Rhodosporidium toruloides) strain NBRC 0880 (otherwise known as IFO0880) was used as a yeast partner to establish a synthetic binary co-culture. Axenic S. elongatus and its co-culture with R. toruloides were grown at 30 ± 2 °C temperature and supplemented with 2% CO2 in nitrogen under approximately 200 µmol photons m−2 s−1 of light provided by 15 W LED light. Cultures were mixed by PFTE Octagon Spinbar Magnetic Stirring Bars (9.5 mm x 25.4 mm) at ~ 200 r.p.m. in BG11 medium supplemented with 0.09 g/L of yeast nitrogen base without amino acids and ammonium sulfate, 0.264 g L−1 ammonium sulfate, 0.174 g L−1 potassium phosphate dibasic, and 3 g L−1 HEPES to a final pH of 8.3 with NaOH. Axenic R. toruloides was grown in same medium supplemented with 10 g L−1 of sucrose. Where appropriate, 1 mM IPTG was added to induce cscB and sps expression. During routine maintenance of cyanobacterial cultures, flasks were back-diluted to approximately 0.2 OD750 for ≥ 3 days prior starting target experiment, growth medium was supplemented with experiments 25 mg L−1 of chloramphenicol and 25 mg L−1 of kanamycin. Antibiotics were removed prior to conducting any of the reported experiments.
Two- and three-unit membrane separated bioreactors for co-cultivation
A custom-made membrane separated bioreactor (mBR) setup was implemented to identify exometabolite profile, metabolite producer and relative gradient for exchanging metabolites in the binary communities. A 2-unit mBR consisted of modified commercially available glass 500 mL and 250 mL heavy duty GL45 round bottles. The bottles were customized into coupled cultivation units by adding a side opening and a 60 mm neck with NW63 glass flat flanges connected with knuckle clamps and teflon centered o-rings to ensure a leak proof sealing. The flanges house 0.22 μm hydrophilic polyvinylidene fluoride (PVDF) membrane filter (Durapore, Millipore, Billerica, MA, USA) to establish a separation barrier for microorganisms but allowing diffusion of the metabolites during co-culture. A 3-unit mBR included additional third chamber between two cultivation units. All units were equipped with standard 20 mm septum fittings to allow aseptic collection of samples.
Sucrose and cell number quantification
Sucrose production and export was activated by addition of 1 mM IPTG inducing expression of sps and cscB. Cells were separated by centrifugation at 15,000 g, the supernatant was recovered and analyzed using the Sucrose/d-Glucose Assay Kit (Megazyme: K-SUCGL) and Genesys 20 Spectrophotometer (Thermo Scientific).
Cell counting was performed using a NovoCyte 3000 VYB Flow Cytometer System (Agilent Technologies Inc., Santa Clara, CA) equipped with violet (405 nm), blue (488 nm), and yellow (561 nm) lasers. S. elongatus and R. toruloides were distinguished and counted based on their distinct forward scatter (FSC) properties and the high red autofluorescence of photosynthetic pigments in S. elongatus. All samples were analyzed fresh (within an hour after collection), briefly vortexed, and loaded along with negative controls (medium and autoclaved distilled water) on a 24-tube loader. Analysis was performed using a flow rate of 50 µL/min and run to achieve at least 100,000 S. elongatus cell events. A series of 3 rinses was performed between each sample, with a homogenization of 1 cycle per sample at 1,000 rpm for 10 s. Excitation was performed with two lasers at 405 nm and 488 nm. Pigment autofluorescence was measured using a red filter (660/20 nm). Data was analyzed with NovoExpress Software (Agilent Technologies Inc., Santa Clara, CA).
Extracellular metabolomics
Comprehensive analysis of extracellular metabolites was conducted using a modified protocol based on previous studies90. Briefly, culture supernatants were dehydrated and subjected to chemical derivatization by methoxyamination and trimethylsilyation. The resulting samples were then examined using gas chromatography-mass spectrometry (GC-MS). The GC-MS analysis was performed on an Agilent GC 7890 A system, utilizing a HP-5MS column (30 m × 0.25 mm × 0.25 μm; Agilent Technologies, Santa Clara, CA, United States) coupled with a single quadrupole MSD 5975 C (Agilent Technologies). To ensure accurate retention time calibration, a fatty acid methyl ester standard mixture was analyzed alongside the samples. Sample (1 mL) was injected into a splitless port at constant temperature of 250 °C. The GC temperature gradient started at 60 °C and hold for 1 min after injection. Then, the temperature was increased to 325 °C at a rate of 10 °C/min and hold at this temperature for 5-min. Fatty acid methyl ester standard mix (C8-28) (Sigma-Aldrich) was analyzed longside the samples as standard for retention time calibration. Raw GC-MS data processing was carried out in Metabolite Detector software91.
This involved calculating retention indices for the detected metabolites based on the analysis of a FAMEs mixture, and chromatographic aligning of them across all samples after deconvolution. Metabolite identification was accomplished through a two-step process: first, by comparing experimental spectra with an extensive, custom-augmented version of “Agilent GC-MS Metabolomics Library” library, and second, by matching unknown peaks against a standard GC-MS library with spectra and validated retention indices for over 850 metabolites. Next, the unknown peaks were additionally matched with the NIST17/Wiley11 GC-MS library. To minimize errors and false identifications, all metabolite identifications and quantification ions underwent rigorous validation. Identified metabolites were categorized into three groups: (1) a main group without any indices, corresponding to metabolites with a high matching score > 0.75; (2) a second group marked with a single asterisk (*), corresponding to metabolites with a matching score < 0.75; and (3) a third group denoted with two asterisks (**), corresponding to metabolites with no direct but best spectral match. This careful approach not only ensured the reliability and accuracy of the metabolomic data obtained from the extracellular samples, but also allowed us to include and display a broader range of metabolites, providing a more comprehensive view of the metabolic landscape. Supplementary Table S1 provides a complete list of identified metabolites, including abbreviated names used in figures and tables, full names, and relevant database IDs (KEGG, BioCyc, PubChem, ChEBI). Abbreviated names include a unique identifier number to facilitate cross-referencing throughout the paper and in functional enrichment analysis.
Metabolomics data analysis and visualization
The acquired metabolite peak intensity data was pre-processed using MetaboAnalyst v5.0 platform92,93 to adjust for systematic differences among samples and variation in metabolite peak scales through sample normalization by sum, Log10 transformation and Pareto scaling (mean-centered and divided by the square root of the standard deviation of each variable) (see Supplementary Figures S4-S7). Pre-processed data was subjected to principal component analysis (PCA) using the R Statspackage (version 4.2.2)94, which applies singular value decomposition method, and visualized using Factoextrapackage (version 1.0.7)95. Pearson pairwise correlation coefficients among metabolite intensities (concentrations), hierarchical clustering based on Euclidean distance for metabolite intensities and samples (see Supplementary Figures S5-S10). Then, the metabolites with statistically significant differential intensities between cultures (Log2 FC of > 1 and adjusted p-value < 0.05) were determined using MetaboAnalyst v5.092,93 (see Supplementary Tables S2-S3). Most significant associations among metabolites, representing top ~ 250 correlation coefficients with R > 0.8 for 101 h and R > 0.83 for 173 h, were visualized as a correlation-based network using Cytoscapesoftware80. Metabolites were clustered through community clustering app (GLAY) based on the Girvan-Newman fast greedy algorithm implemented in the clusterMakerplugin81.
Results of the metabolite differential intensity and functional enrichment analysis were visualized on Volcano plots, Edwards–Venn and pathway topology diagrams made by using R EnhancedVolcanopackage (version 1.16.0)96, EVennweb tool97 and Pathway Analysistool of the MetaboAnalyst v5.092,93. Pathway over-representation analysis was performed using the hypergeometric test (FDR ≤ 0.05 was considered significant) using metabolite sets based on normal metabolic pathways and the reference pathway libraries for S. elongatus PCC 7942 and S. cerevisiaeavailable through the Kyoto Encyclopedia of Genes and Genomes (KEGG)98.
Data availability
Data is provided within the manuscript or supplementary information files.
References
Flombaum, P. et al. Present and future global distributions of the marine Cyanobacteria Prochlorococcus and Synechococcus. Proceedings of the National Academy of Sciences, 110(24): pp. 9824–9829. (2013).
Field, C. B., Behrenfeld, M. J., Randerson, J. T. & Falkowski, P. Primary production of the Biosphere: integrating Terrestrial and Oceanic Components. Science. 281 (5374), 237–240 (1998).
Jardillier, L., Zubkov, M. V., Pearman, J. & Scanlan, D. J. Significant CO2 fixation by small prymnesiophytes in the subtropical and tropical northeast Atlantic Ocean. ISME J. 4 (9), 1180–1192 (2010).
Teiser, M. L. O. Extracellular low Molecular Weight Organic Compounds Produced by Synechococcus sp. and Their Roles in the food web of Alkaline hot Spring Microbial mat Communities, in Department of Biology (University of Oregon, 1993).
Biddanda, B. & Benner, R. Carbon, nitrogen, and carbohydrate fluxes during the production of particulate and dissolved organic matter by marine phytoplankton. Limnol. Oceanogr. 42 (3), 506–518 (1997).
Bertilsson, S., Berglund, O., Pullin, M. & Chisholm, S. Release of Dissolved Organic Matter by Prochlorococcus55p. 225–231 (Vie et Milieu/Life & Environment, 2005).
Morán, X. A. G., Gasol, J. M., Pedrós-Alió, C. & Estrada, M. Partitioning of phytoplanktonic organic carbon production and bacterial production along a coastal-offshore gradient in the NE Atlantic during different hydrographic regimes. Aquat. Microb. Ecol. 29, 239–252 (2002).
Marañón, E., Cermeño, P., Fernández, E., Rodríguez, J. & Zabala, L. Significance and mechanisms of photosynthetic production of dissolved organic carbon in a coastal eutrophic ecosystem. Limnol. Oceanogr. 49 (5), 1652–1666 (2004).
Teira, E., José Pazó, M., Serret, P. & Fernández, E. Dissolved organic carbon production by microbial populations in the Atlantic Ocean. Limnol. Oceanogr. 46 (6), 1370–1377 (2001).
Jiao, N. et al. Microbial production of recalcitrant dissolved organic matter: long-term carbon storage in the global ocean. Nat. Rev. Microbiol. 8 (8), 593–599 (2010).
Amon, R. M. W. Ocean dissolved organics matter. Nat. Geosci. 9 (12), 864–865 (2016).
Moran, M. A. et al. Deciphering ocean carbon in a changing world. Proceedings of the National Academy of Sciences, 113(12): pp. 3143–3151. (2016).
Sexton, P. F. et al. Eocene global warming events driven by ventilation of oceanic dissolved organic carbon. Nature. 471 (7338), 349–352 (2011).
Rothman, D. H., Hayes, J. M. & Summons, R. E. Dynamics of the Neoproterozoic carbon cycle. Proceedings of the National Academy of Sciences, 100(14): pp. 8124–8129. (2003).
Porada, P., Weber, B., Elbert, W., Pöschl, U. & Kleidon, A. Estimating impacts of lichens and bryophytes on global biogeochemical cycles. Glob. Biogeochem. Cycles. 28 (2), 71–85 (2014).
Elbert, W. et al. Contribution of cryptogamic covers to the global cycles of carbon and nitrogen. Nat. Geosci. 5 (7), 459–462 (2012).
Honegger, R. Functional aspects of the Lichen Symbiosis. Annu. Rev. Plant Physiol. Plant Mol. Biol. 42 (1), 553–578 (1991).
Morán, X. A. G., Ducklow, H. W. & Erickson, M. Carbon fluxes through estuarine bacteria reflect coupling with phytoplankton. Mar. Ecol. Prog. Ser. 489, 75–85 (2013).
Luo, Y. W., Friedrichs, M. A. M., Doney, S. C., Church, M. J. & Ducklow, H. W. Oceanic heterotrophic bacterial nutrition by semilabile DOM as revealed by data assimilative modeling. Aquat. Microb. Ecol. 60 (3), 273–287 (2010).
Li, B. et al. Catalytic promiscuity in the biosynthesis of cyclic peptide secondary metabolites in planktonic marine cyanobacteria. Proceedings of the National Academy of Sciences, 107(23): pp. 10430–10435. (2010).
Christie-Oleza, J. A., Armengaud, J., Guerin, P. & Scanlan, D. J. Functional distinctness in the exoproteomes of marineSynechococcus. Environ. Microbiol. 17 (10), 3781–3794 (2015).
Landa, M. et al. Phylogenetic and structural response of heterotrophic bacteria to dissolved organic matter of different chemical composition in a continuous culture study. Environ. Microbiol. 16 (6), 1668–1681 (2014).
Becker, J. W. et al. Closely Relat. Phytoplankton Species Produce Similar Suites Dissolved Org. Matter Front. Microbiol., 5. (2014).
Teeling, H. et al. Substrate-controlled succession of Marine Bacterioplankton populations Induced by a Phytoplankton Bloom. Science. 336 (6081), 608–611 (2012).
Pontiller, B., Martínez-García, S., Lundin, D. & Pinhassi, J. Labile dissolved Organic Matter compound characteristics select for divergence in Marine bacterial activity and transcription. Front. Microbiol., 11. (2020).
Xu, J. et al. You Exude What You Eat: How Carbon-, Nitrogen-, and Sulfur-Rich Organic Substrates Shape Microbial Community Composition and the Dissolved Organic Matter Pool (Applied and Environmental Microbiology, 2022). 88(23).
Sher, D., Thompson, J. W., Kashtan, N., Croal, L. & Chisholm, S. W. Response of Prochlorococcus ecotypes to co-culture with diverse marine bacteria. ISME J. 5 (7), 1125–1132 (2011).
Morris, J. J., Kirkegaard, R., Szul, M. J., Johnson, Z. I. & Zinser, E. R. Facilitation of Robust Growth of Prochlorococcus colonies and dilute liquid cultures by helper heterotrophic Bacteria. Appl. Environ. Microbiol. 74 (14), 4530–4534 (2008).
Weissberg, O., Aharonovich, D. & Sher, D. Phototroph-heterotroph interactions during growth and long-term starvation across Prochlorococcus and Alteromonas diversity. ISME J. 17 (2), 227–237 (2022).
Christie-Oleza, J. A., Sousoni, D., Lloyd, M., Armengaud, J. & Scanlan, D. J. Nutrient recycling facilitates long-term stability of marine microbial phototroph–heterotroph interactions. Nat. Microbiol., (2017). 2(9).
Roth-Rosenberg, D. et al. Prochlorococcus cells rely on microbial interactions rather than on chlorotic resting stages to survive long-term nutrient starvation. mBio, 11(4). (2020).
Coe, A. et al. Survival of Prochlorococcus in extended darkness. Limnol. Oceanogr. 61 (4), 1375–1388 (2016).
Biller, S. J., Coe, A., Roggensack, S. E., Chisholm, S. W. & Mason, O. Heterotroph Interact. Alter. Prochlorococcus Transcriptome Dynamics Dur. Ext. Periods Darkn. mSystems, 3(3). (2018).
Morris, J. J., Johnson, Z. I., Szul, M. J., Keller, M. & Zinser, E. R. Dependence of the Cyanobacterium Prochlorococcus on Hydrogen Peroxide Scavenging Microbes for Growth at the Ocean’s Surface. PLoS ONE, 6(2). (2011).
Bohutskyi, P. et al. Metabolic effects of vitamin B12 on physiology, stress resistance, growth rate and biomass productivity of cyanobacterium stanieri planktonic and biofilm cultures. Algal Res., 42. (2019).
Smith, D., Muscatine, L. & Lewis, D. Carbohydrate movement from autotrophs to heterotrophs in parasitic and mutualistic symbiosis. Biol. Rev. 44 (1), 17–85 (1969).
Lines, C. E. M., Ratcliffe, R. G., Rees, T. A. V. & Southon, T. E. A 13 C NMR study of photosynthate transport and metabolism in the lichen Xanthoria Calcicola Oxner. New Phytol. 111 (3), 447–456 (1989).
Honegger, R., Kutasi, V. & Ruffner, H. P. Polyol patterns in eleven species of aposymbiotically cultured lichen mycobionts. Mycol. Res. 97 (1), 35–39 (1993).
Fahselt, D. Carbon Metabolism in Lichens. Symbiosis. 17, 127–182 (1994).
Armstrong, R. A. & Smith, S. N. The Levels of Ribitol, Arabitol and Mannitol in Individual Lobes of the Lichen Parmelia Conspersa (Ehrh. ex Ach.) ACH34p. 253–260 (Environmental and Experimental Botany, 1994). 3.
Aubert, S., Juge, C., Boisson, A. M., Gout, E. & Bligny, R. Metabolic processes sustaining the reviviscence of lichen Xanthoria elegans (Link) in high mountain environments. Planta. 226 (5), 1287–1297 (2007).
Honegger, R. Metabolic Interactions at the Mycobiont-Photobiont Interface in Lichens, in Plant Relationships. pp. 209–221. (1997).
Huneck, S. & Yoshimura, I. Identification of Lichen Substances, in Identification of Lichen Substances. pp. 11–123. (1996).
Stocker-Wörgötter, E. Metabolic diversity of lichen-forming ascomycetous fungi: culturing, polyketide and shikimatemetabolite production, and PKS genes. Nat. Prod. Rep. 25 (1), 188–200 (2008).
Calcott, M. J., Ackerley, D. F., Knight, A., Keyzers, R. A. & Owen, J. G. Secondary metabolism in the lichen symbiosis. Chem. Soc. Rev. 47 (5), 1730–1760 (2018).
Williamson, J. D., Jennings, D. B., Guo, W. W., Pharr, D. M. & Ehrenshaft, M. Sugar Alcohols, Salt stress, and Fungal Resistance: polyols—multifunctional Plant Protection? J. Am. Soc. Hortic. Sci. 127 (4), 467–473 (2002).
Kosugi, M. et al. Arabitol provided by Lichenous Fungi Enhances Ability To Dissipate Excess Light Energy in a Symbiotic Green Alga under Desiccation. Plant Cell Physiol. 54 (8), 1316–1325 (2013).
Kranner, I. et al. Antioxidants and photoprotection in a lichen as compared with its isolated symbiotic partners. Proceedings of the National Academy of Sciences, 102(8): pp. 3141–3146. (2005).
Millot, M., Di Meo, F., Tomasi, S., Boustie, J. & Trouillas, P. Photoprotective capacities of lichen metabolites: a joint theoretical and experimental study. J. Photochem. Photobiol., B. 111, 17–26 (2012).
Straight, P. D. & Kolter, R. Interspecies Chemical Communication in Bacterial Development. Annu. Rev. Microbiol. 63 (1), 99–118 (2009).
Gökalsın, B., Berber, D., Özyiğitoğlu, G. Ç., Yeşilada, E. & Sesal, N. C. Quorum sensing attenuation properties of ethnobotanically valuable lichens againstPseudomonas aeruginosa. Plant. Biosystems - Int. J. Dealing all Aspects Plant. Biology. 154 (6), 792–799 (2019).
Nazem-Bokaee, H., Hom, E. F. Y., Warden, A. C., Mathews, S. & Gueidan, C. Towards a Systems Biology Approach to Understanding the Lichen Symbiosis: Opportunities and Challenges of Implementing Network Modelling12 (Frontiers in Microbiology, 2021).
Amin, S. A. et al. Interaction and signalling between a cosmopolitan phytoplankton and associated bacteria. Nature. 522 (7554), 98–101 (2015).
Friedmann, E. I., Hua, M. & Ocampo-Friedmann, R. 3.6 cryptoendolithic lichen and cyanobacterial communities of the Ross Desert, Antarctica. Polarforschung. 58 (2/3), 251–259 (1988).
Li, H. & Wei, J. C. Functional analysis of thioredoxin from the desert lichen-forming fungus, endocarpon pusillum Hedwig, reveals its role in stress tolerance. Sci. Rep. 6, 27184 (2016).
Paoli, A., Celussi, M., Del Negro, P., Fonda Umani, S. & Talarico, L. Ecological advantages from light adaptation and heterotrophic-like behavior in Synechococcus harvested from the Gulf of Trieste (Northern Adriatic Sea). FEMS Microbiol. Ecol. 64 (2), 219–229 (2008).
Muñoz-Marín, M. C. Mixotrophy in depth. Nat. Microbiol. 7 (12), 1949–1950 (2022).
Wu, Z. et al. Single-cell Measurements Modelling Reveal. Substantial Org. Carbon Acquisition Prochlorococcus Nat. Microbiol., 7(12): 2068–2077. (2022).
Yelton, A. P. et al. Global genetic capacity for mixotrophy in marine picocyanobacteria. ISME J. 10 (12), 2946–2957 (2016).
Muñoz-Marín, M. C. et al. Mixotrophy in Marine picocyanobacteria: use of organic compounds by Prochlorococcus and Synechococcus. ISME J. 14 (5), 1065–1073 (2020).
Mary, I. et al. Diel rhythmicity in amino acid uptake byProchlorococcus. Environ. Microbiol. 10 (8), 2124–2131 (2008).
Michelou, V. K., Cottrell, M. T. & Kirchman, D. L. Light-stimulated bacterial production and amino acid assimilation by Cyanobacteria and other microbes in the North Atlantic Ocean. Appl. Environ. Microbiol. 73 (17), 5539–5546 (2007).
Malmstrom, R. R., Kiene, R. P., Vila, M. & Kirchman, D. L. Dimethylsulfoniopropionate (DMSP) assimilation by Synechococcus in the Gulf of Mexico and northwest Atlantic Ocean. Limnol. Oceanogr. 50 (6), 1924–1931 (2005).
Vila-Costa, M. et al. Dimethylsulfoniopropionate Uptake Mar. Phytoplankton Sci., 314(5799): 652–654. (2006).
Ruiz-González, C., Simó, R., Vila-Costa, M., Sommaruga, R. & Gasol, J. M. Sunlight modulates the relative importance of heterotrophic bacteria and picophytoplankton in DMSP-sulphur uptake. ISME J. 6 (3), 650–659 (2011).
Cuhel, R. L. & Waterbury, J. B. Biochemical composition and short term nutrient incorporation patterns in a unicellular marine cyanobacterium, Synechococcus (WH7803)1. Limnology and Oceanography, 29(2): pp. 370–374. (1984).
Martinez, J., Riera, M., Lalucat, J. & Vives-Rego, J. Thymidine incorporation into algal DNA from axenic cultures of Synechococcus, Chlorella and Tetraselmis. Lett. Appl. Microbiol. 8 (4), 135–138 (1989).
Wang, J. et al. Construction of fungi-microalgae Symbiotic System and Adsorption Study of Heavy Metal ions268 (Separation and Purification Technology, 2021).
Li, T. et al. Creating a synthetic lichen: Mutualistic co-culture of fungi and extracellular polysaccharide-secreting cyanobacterium Nostoc PCC 7413. Algal Res., 45. (2020).
Li, T. et al. Mimicking lichens: incorporation of yeast strains together with sucrose-secreting cyanobacteria improves survival, growth, ROS removal, and lipid production in a stable mutualistic co-culture production platform. Biotechnol. Biofuels, 10(1). (2017).
Jiang, L. et al. Evidence for a mutualistic relationship between the cyanobacteria Nostoc and fungi Aspergilli in different environments. Appl. Microbiol. Biotechnol. 104 (14), 6413–6426 (2020).
Weiss, T. L., Young, E. J. & Ducat, D. C. A synthetic, light-driven consortium of cyanobacteria and heterotrophic bacteria enables stable polyhydroxybutyrate production. Metab. Eng. 44, 236–245 (2017).
Zuñiga, C. et al. Synthetic microbial communities of heterotrophs and phototrophs facilitate sustainable growth. Nature Communications, 11(1). (2020).
DiMucci, D., Kon, M., Segrè, D. & Typas, N. Machine learning reveals Missing edges and Putative Interaction mechanisms in Microbial Ecosystem Networks. mSystems, 3(5). (2018).
Zhang, Z. et al. Long-term survival of Synechococcus and heterotrophic Bacteria without external nutrient supply after changes in their relationship from antagonism to Mutualism. mBio, 12(4). (2021).
Nair, S. et al. Inherent tendency of Synechococcus and heterotrophic bacteria for mutualism on long-term coexistence despite environmental interference. Sci. Adv., 8(39). (2022).
Zuñiga, C. et al. Environmental stimuli drive a transition from cooperation to competition in synthetic phototrophic communities. Nat. Microbiol. 4 (12), 2184–2191 (2019).
Bohutskyi, P. et al. Production of lipid-containing algal-bacterial polyculture in wastewater and biomethanation of lipid extracted residues: enhancing methane yield through hydrothermal pretreatment and relieving solvent toxicity through co-digestion. Sci. Total Environ. 653, 1377–1394 (2019).
Bohutskyi, P. et al. Conversion of stranded waste-stream carbon and nutrients into value-added products via metabolically coupled binary heterotroph-photoautotroph system. Bioresour. Technol. 260, 68–75 (2018).
Shannon, P. et al. Cytoscape: a Software Environment for Integrated Models of Biomolecular Interaction Networks. Genome Res. 13 (11), 2498–2504 (2003).
Morris, J. H. et al. clusterMaker: a multi-algorithm clustering plugin for Cytoscape. BMC Bioinform., 12(1). (2011).
De Perez, L., Alseekh, S., Brotman, Y. & Fernie, A. R. Network-based strategies in metabolomics data analysis and interpretation: from molecular networking to biological interpretation. Expert Rev. Proteomics. 17 (4), 243–255 (2020).
Jahagirdar, S. & Saccenti, E. On the use of correlation and MI as a measure of Metabolite—Metabolite Association for Network Differential Connectivity Analysis. Metabolites, 10(4). (2020).
Birer-Williams, C. M. C., Chu, R. K., Anderton, C. R., Wright, E. S. & Bernstein, H. C. SubTap, a versatile 3D printed platform for eavesdropping on extracellular interactions. mSystems, 6(4). (2021).
Jo, C. et al. Construction and modeling of a coculture microplate for real-time measurement of Microbial interactions. mSystems, 8(2). (2023).
Lange, O. L., Green, T. G. A. & Heber, U. Hydration-dependent photosynthetic production of lichens: what do laboratory studies tell us about field performance? J. Exp. Bot. 52 (363), 2033–2042 (2001).
Osyczka, P. & Myśliwa-Kurdziel, B. The pattern of photosynthetic response and adaptation to changing light conditions in lichens is linked to their ecological range. Photosynth. Res. 157 (1), 21–35 (2023).
Cowan, D. A., Green, T. G. A. & Wilson, A. T. Lichen Metabolism. 2. Aspects of light and dark physiology. New Phytol. 83 (3), 761–769 (2006).
Abramson, B. W., Kachel, B., Kramer, D. M. & Ducat, D. C. Increased photochemical efficiency in Cyanobacteria via an Engineered sucrose Sink. Plant Cell Physiol. 57 (12), 2451–2460 (2016).
Pomraning, K. R. et al. Integration of Proteomics and Metabolomics Into the Design, Build, Test, Learn Cycle to Improve 3-Hydroxypropionic Acid Production in Aspergillus pseudoterreus. Frontiers in Bioengineering and Biotechnology, 9. (2021).
Hiller, K. et al. MetaboliteDetector: Comprehensive Analysis Tool for targeted and nontargeted GC/MS based Metabolome Analysis. Anal. Chem. 81 (9), 3429–3439 (2009).
Pang, Z. et al. Using MetaboAnalyst 5.0 for LC–HRMS spectra processing, multi-omics integration and covariate adjustment of global metabolomics data. Nat. Protoc. 17 (8), 1735–1761 (2022).
Xia, J., Psychogios, N., Young, N. & Wishart, D. MetaboAnalyst: a web server for metabolomic data analysis and interpretation. Nucleic Acids Res. 37 (Web Server), W652–W660 (2009).
Team, R. C. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. URL (2022). https://www.R-project.org/
Kassambara, A. & Mundt, F. Factoextra: Extract and Visualize the Results of Multivariate Data Analyses. R Package Version 1.0.7. (2020). https://CRAN.R-project.org/package=factoextra
Blighe, K., Rana, S. & Lewis, M. EnhancedVolcano: Publication-ready volcano plots with enhanced colouring and labeling. R package version 1.16.0. (2022). https://github.com/kevinblighe/EnhancedVolcano.
Chen, T., Zhang, H., Liu, Y., Liu, Y. X. & Huang, L. EVenn: Easy to create repeatable and editable Venn diagrams and Venn networks online. J. Genet. Genomics. 48 (9), 863–866 (2021).
Kanehisa, M., Goto, S., Sato, Y., Furumichi, M. & Tanabe, M. KEGG for integration and interpretation of large-scale molecular data sets. Nucleic Acids Res. 40 (D1), D109–D114 (2011).
Acknowledgements
This research was supported by the Predictive Phenomics Initiative Laboratory Directed Research and Development Program at Pacific Northwest National Laboratory (PNNL), and U.S. Department of Energy Genomic Science Program under award DE-SC0019388. PNNL is operated by Battelle for the DOE under Contract DE-AC05-76RL01830.
Author information
Authors and Affiliations
Contributions
P.B., J.M., and M.B. funding acquisition and project administration; P.B., J.M., and M.B. conception and design; P.B. collection, assembly and analysis of cultivation experimental data; Y-M. K. nd B.C.P. analysis of metabolics samples; P.B. and Y-M. K. processing and interpretation of metabolomics data; P.B. writing – original draft; P.B., K.R.P. and M.B writing – review & editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Conflict of interest
The authors wish to confirm that there are no known conflicts of interest associated with this publication.
Authors agreement to authorship and submission
All persons designated as author agree to submit the manuscript for peer review.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Bohutskyi, P., Pomraning, K.R., Jenkins, J. . et al. Mixed and membrane-separated culturing of synthetic cyanobacteria-yeast consortia reveals metabolic cross-talk mimicking natural cyanolichens. Sci Rep 14, 25303 (2024). https://doi.org/10.1038/s41598-024-74743-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-74743-4
Keywords
This article is cited by
-
Advances in organic acid production using cyanobacteria: strategies and applications
Blue Biotechnology (2025)