Abstract
Symbiotic microorganisms living in the digestive tracts of invertebrates can be crucial in host-symbiont interactions, as they play fundamental roles in important biological processes. Urbanization-related habitat alteration and disturbance, however, considerably affect the environment of host insects, from which their gut microbiota is derived. Still, relatively few studies, all on flying insects, have assessed the impact of urbanization on the gut microbiota of insects. Here, we compared the gut bacterial microbiota in rural and urban individuals of a flightless ground beetle, Carabus convexus, using next generation sequencing. Across the 48 gut samples we identified 1163 different bacterial operational taxonomic units (OTUs), forming significantly different gut bacterial communities in rural versus urban beetles. The taxonomic diversity of the gut bacterial microbiota expressed by the Hill numbers was significantly higher in rural than urban individuals, as well as in rural males vs. females. Smaller differences were found in functional diversity, assessed by the Rao’s quadratic entropy which was marginally significantly higher in urban than rural beetles.
Similar content being viewed by others
Introduction
Global anthropogenic activities, including habitat conversion, exploitation, and urbanization significantly threaten biodiversity as well as ecosystem functions and services1,2,3,4. Nowadays, urbanization —characterised by a global increase in urban land cover and human population5—is the most intensively increasing anthropogenic process6.
Urbanization heavily modifies the abiotic components of the environment, from microclimate7, to pollutants8 and nutrients9. The most frequent overall effect on urban-living organisms is a reduction in their diversity both taxonomically10 and functionally11, as well as homogenisation12. Most of these effects have been documented in multicellular organisms, but microorganisms are expected to react similarly13. Indeed, there is evidence of environmental changes directly affecting microbes14.
In recent decades, we have gradually recognised the significant roles played by symbiotic microorganisms15, especially the gut microbiota16. Furthermore, novel methodologies (improved culturing techniques and very cheap, extremely thorough, sequencing methods) have greatly advanced the field of microbiology. The intestinal microbiota play crucial roles in important biological processes, such as the digestion of poorly degradable materials, the synthesis of essential vitamins and nutrients16, the degradation of toxins17, the regulation of host’s immune system and cellular homeostasis18, and the protection against pathogens and parasites16,19.
There are examples of vertical transmission of gut microbes, but it is assumed that the gut bacterial microbiota are largely acquired from microorganisms found in food items and other sources within the environment16,20. Thus, habitat alteration and changes in food can influence the composition and diversity of the gut microbial community21. Urbanization profoundly modifies several environmental conditions, including the quality and quantity of available food22, therefore it is reasonable to assume that urbanization also affects the composition and diversity of the gut microbiota. Such an impact has been documented for several flying insects23,24. Studies on terrestrial insects with limited dispersal power, however, are much rarer. Recently, Magura et al.25 described the bacterial microbiota of the flightless ground beetle, Carabus convexus F. 1775 collected from rural and urban habitats. Using next generation sequencing and analysing amplicon sequence variants (ASVs), they identified 11 bacterial phyla and 59 families. However, no study to date has investigated the functional diversity of the insect gut microbiota, even though a functional reduction can have detrimental effects on host survival and performance26. We aimed to test how urbanization acts on the functional diversity of the microbiota in this flightless carabid.
Specifically, we hypothesised that (1) due to urbanization-driven habitat alteration, biodiversity impoverishment and food spectrum reduction, the diversity of gut bacterial microbiota would be both taxonomically and functionally reduced in urban beetles compared to their rural conspecifics. Further, (2) given that carabid males are more mobile than females27, we hypothesised that they acquire a more diverse set of symbionts from their environment.
We found that urbanization had a significant effect on the taxonomic but not on the functional diversity of the gut bacterial microbiota. Furthermore, we found sex-specific differences: rural males had significantly more diverse bacterial microbiota than conspecific females.
Materials and methods
Study area and study design
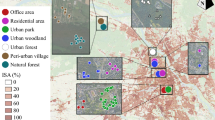
The research was conducted in eastern Hungary, in and around the city of Debrecen (47° 32′ N; 21° 38′ E). Debrecen is the second-largest city in Hungary (199,858 inhabitants in 2022), with extensive lowland forests in and around town. We used the research infrastructure established according to the international GLOBENET protocol28, and described in detail by Magura et al.25.
Studied model organism and sampling design
Our model organism, C. convexus is a widespread Eurasian species of medium size (14–23 mm), spring-breeding, wingless (brachypterous), with limited dispersal power. It is predominantly nocturnal and predatory with extraintestinal digestion29. In the studied lowland region (Great Hungarian Plain) C. convexus is a forest specialist that also occurs in urban forest fragments though in significantly lower numbers than in continuous rural stands30.
Adults of C. convexus were captured using unbaited pitfall traps during the breeding period, from late March to mid-May in 2021 as described by Magura et al.25.
We collected 106 C. convexus individuals (49 females and 40 males from the rural, while 8 females and 9 males from the urban sites), all with unworn mandibles, indicating that they were in their first breeding season. Given the very uneven numbers collected, we processed all urban-collected adults but only 15 randomly selected rural females and 16 males, ensuring that roughly similar numbers were obtained from each of the four rural sites. Selected beetles were starved for one day to reduce the influence of microbiota from freshly consumed prey, after which they were euthanized and stored in 96% ethanol.
Gut dissection, DNA extraction, PCR amplification and high-throughput sequencing
Beetles stored in 96% ethanol at + 5 °C were dissected within 3 days after collection to avoid changes in the gut microbiota community31. The intestinal tracts (foregut and midgut) were dissected as described by Magura et al.25. To avoid contamination, before the dissection, the surface of the beetles, the forceps and needles used for dissection were cleaned and sterilized with 96% ethanol. The study was performed in accordance with guidelines and regulations of the University of Debrecen.
Bacterial DNA was extracted using the QIAamp PowerFecal Pro DNA Kit, following the manufacturer’s protocol. The hypervariable V3 and V4 regions of the bacterial 16S ribosomal RNA (rRNA) genes were amplified by polymerase chain reactions (PCR) using the universal primers: 5′-CCTACGGGNGGCWGCAG-3′ and 5′-GACTACHVGGGTATCTAATCC-3′32. To identify samples after sequencing, both forward and reverse primers were labeled at the 5′ end with Illumina overhang adapter sequences following the Illumina 16S Metagenomic Library Preparation Guide (15044223-B). Further procedures were as described in the relevant manufacturers’ instructions, also detailed by Magura et al.25.
The normalized, pooled and denatured library samples were sequenced on the Illumina MiSeq platform (Illumina, Inc., San Diego, CA, United States; MiSeq Reagent Kit v3, 300 bp paired-end V3 chemistry, 600 cycle). All laboratory works were performed under sterile conditions at the UD-GenoMed Medical Genomic Technologies Ltd., University of Debrecen, Hungary. We followed the recommendations of Eisenhofer et al.33 to ensure the replicability and reliability of our study. Positive controls in microbiota studies of ground beetles is not widespread34, and in harmony with this practice, we did not used positive controls. Two types of negative controls were used: DNA extraction negative control (containing sterile water without a gut sample) and PCR amplification negative control (containing sterile water without template DNA). Negative controls had no microbial reads, validating the robustness and reliability of our results.
Bioinformatic and statistical analyses
The processing of raw MiSeq forward and reverse paired-end reads was based on the FROGS pipeline35. Paired-end reads were merged using the FLASH tool36. Dereplicated sequences were clustered first with aggregation parameter d = 1, then with d = 3 with Swarm37. Chimera detection and removal was by following the UCHIME method38,39, and the usual 0.005% abundance noise threshold40. Quality-controlled sequences were clustered into operational taxonomic units (OTUs) using the identity threshold of 97%39. Sequence taxonomic assignment of the operational taxonomic units (OTUs) was by the BLAST procedure41 on the SILVA 16S database for bacterial sequences (release 138.1)42.
The predicted functional roles (functional traits) of the OTUs assigned to bacterial genera was determined using the Functional Annotation of Prokaryotic Taxa (FAPROTAX) database43. Based on this, OTUs assigned to bacterial genera were classified into groups according to their assumed function (metabolism of carbon-based, hydrogen-based, nitrogen-based, sulphur-based, carbon–nitrogen-based, and aromatic compounds, cellulose, chitin, hydrocarbon, methanol, nitrate, and xylan metabolism, acetogenesis, methanogenesis, nitrogen fixation, pathogenicity) and their respiration types (anaerobic, aerobic, or variable).
All statistical analyses were performed in R (version 4.3.144). Before analysis, the bioinformatically processed dataset was median normalized45 with the phyloseq package46 to deal with differences in sequencing depth47. The composition of the gut bacterial communities was analysed by non-metric multidimensional scaling ordination using the Bray–Curtis index of dissimilarity in the vegan package48. Convex hulls around the samples of the groups were visualised using the standard errors of the averages (centroids) of samples multiplied with the 95% confidence value from the Chi-squared distribution (d.f. = 2 ) utilising the vegan48 and MASS49 packages. To test for significant differences in centroids by PERMANOVA we used the vegan48 and RVAideMemoire50 packages.
Common taxonomic diversity indices vary in sensitivities to the relative abundance of species (or any other taxonomic level considered) and rank communities in different ways51. The scalable diversity approach offers a solution to this problem52. The Hill numbers are appropriate candidates for scalable diversity comparisons, as the parameter value, q, determines the sensitivity of the Hill number to the relative abundances of taxa53,54. At q = 0, the Hill number (0D) corresponds to taxon richness and is very sensitive to the rare taxa, for q → 1 the Hill number (1D) is related to the Shannon diversity , while at q = 2, 2D corresponds to the Simpson diversity and is more influenced by abundant than rare taxa53,55. We calculated the Hill diversity using the hillR package56.
The predicted functional diversity of the gut bacterial communities was expressed using the Rao’s quadratic entropy (\(Q\)57,58) based on the traits related to the earlier-mentioned functional roles:
where \(\text{S}\) is the number of OTUs in a gut sample; \({d}_{ij}\) is the difference between the i-th and j-th OTU based on functional traits (\({d}_{ij}={d}_{ji}\) and \({d}_{ii}=0\)); \({p}_{i}\) and \({p}_{j}\) are the relative median normalized read numbers of the i-th and j-th OTU.
Distances between OTUs based on predicted functional traits (\({d}_{ij}\)) were calculated using Gower’s distance metric, considering both functional classes as categorical variables. Rao’s quadratic entropy was computed with the SYNCSA package59 while Gower’s distance was calculated using the StatMatch package60.
The effects of urbanization level (or habitat type: rural vs. urban) and sex (female vs. male) on the diversity of gut bacterial microbiota was analysed by linear mixed models (LMM) using the lme4 package61. Before modelling, the best-fitting probability distribution for the Hill numbers and the Rao’s quadratic entropy were identified using the the car62 and MASS packages49. Accordingly, both the Hill numbers and the Rao’s quadratic entropy values were modelled with normal error distributions. In the models with nested design (sampling sites nested within sampling areas) the habitat type, sex and their interaction were considered as fixed effects. When the model indicated significant difference between means, the Tukey test for multiple comparison among means63 was performed.
Results
Composition of the gut bacterial microbiota
The sequenced 48 gut samples yielded 1163 different OTUs with a total read count of 8,202,927 (mean = 170,894.3 and SD = 59,956.48). From the 31 rural gut bacterial samples, 1138 OTUs with 5,140,946 reads (mean = 165,837 and SD = 69,654.33), while from the 17 urban samples 1059 OTUs with 3,061,981 reads (mean = 180,116.5 and SD = 36,369.76) were obtained. Metadata and raw sequence reads were deposited in the NCBI SRA database (BioProject PRJNA1051584, https://www.ncbi.nlm.nih.gov/bioproject/PRJNA1051584).
Overall, 21 bacterial phyla were identified. Based on the average relative abundance, the most common phyla were: Firmicutes (average 48.56% in all individuals, 42.95% in rural and 58.79% in urban beetles), Bacteroidetes (overall average 28.51%, 33.31% in rural and 19.76% in urban individuals), an unclassified phylum (average 7.90% in all individuals, 8.36% in rural and 7.06% in urban beetles), and Proteobacteria (overall mean 5.85%, 5.16% in rural vs. 7.11% in urban individuals). These four phyla represent 90.82% (89.78% in rural, 92.72% in urban) of the whole bacterial microbiota.
At a family level, 104 bacterial families were identified (Table S1). The most abundant families were: Enterococcaceae (average relative abundance in all samples: 14.49%), Prevotellaceae (13.10%), Ruminococcaceae (12.03%), an unclassified family (7.90%), and Rikenellaceae (6.09%). Ninety-seven families were present in both habitat types, while 6 families were recorded only in beetles from rural and 1 family only from urban habitats. Thirty-four families were present in more than 90% of the samples, so they can be considered common members of the gut bacterial microbiota. In contrast, 12 families were recorded in less than 10% of the samples, indicating that they were likely to be facultative members (Table S1). We observed few differences in the average abundance of the common families between the rural and urban samples, but the Nocardiaceae were more abundant (but not significantly; one-way ANOVA, F1, 46 = 0.5553, p = 0.460) in rural, while Enterococcaceae were significantly (one-way ANOVA, F1, 46 = 11.6330, p = 0.001) more abundant in urban beetles (Fig. 1).
Mean abundance per gut sample of the common (> 1% average abundance in all samples) bacterial families (OTUs) in rural and urban C. convexus individuals.
The ordination plot centroids revealed significantly different gut bacterial microbiota between habitat types (Fig. 2A; R2 = 0.1193, p = 0.003, number of permutations: 999). Centroids were also significantly different between females and males (Fig. 2B; R2 = 0.1836, p = 0.007, number of permutations: 999). However, the pairwise comparisons between group centroids showed that only the rural male—urban male, as well as the rural male—urban female group pairs were significantly different (p = 0.003 for both comparisons).
Non-metric multidimensional scaling ordination using the Bray–Curtis index of dissimilarity of the gut bacterial samples (OTUs) from C. convexus individuals (A) at the habitat-level (open green circles represent gut bacterial samples from rural, while open red ones represent samples from urban beetles), and (B) at the level of habitat type × sex interaction (green symbols represent rural, red ones urban beetles, while open squares represent females, open down triangles males). Ellipses (convex hulls) were based on the standard errors of the averages (centroids) of points multiplied with the 95% confidence value from the Chi-squared distribution (number of permutations: 999). Final stress value: 17.0069.
Taxonomic diversity of the gut bacterial microbiota
The taxonomic diversity of the gut bacterial microbiota expressed by the Hill numbers was significantly higher in the rural than urban beetles for all parameter values, except q = 0, indicating that, apart from the number of OTUs, their bacterial gut microbiota was more diverse (Table 1, Fig. 3). Biologically relevant sex-specific difference in the Hill number at q = 2 of the gut bacterial microbiota indicated that abundant bacterial taxa were slightly more diverse in males than females. While there was no significant habitat × sex interaction (Fig. 3), LMMs performed separately for the rural and urban habitats (considering sex of the studied beetles as a fixed, and site as a random factor) showed that the gut bacterial microbiota was significantly more diverse in rural males than females (Table S2), while there was no such difference in the urban habitat (Table S2).
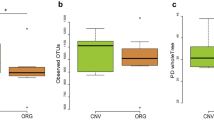
Boxplot of taxonomic diversity of the gut bacterial microbiota (OTUs) of C. convexus females and males from rural and urban habitats using the Hill numbers (A) when the parameter value, q = 0 (0D), (B) when the parameter value, q → 1 (1D), and (C) when the parameter value, q = 2 (2D). In boxplots horizontal lines represent median values, boxes show the interquartile ranges, whiskers represent the minimum and maximum values, while points outside the whiskers are outliers.
Functional diversity of the gut bacterial microbiota
Of the 1,163 identified OTUs 62.6% (69.4% of the total reads) could be assigned traits from the FAPROTAX database. The predicted functional diversity of these 728 OTUs was marginally significantly (p = 0.0700) higher in urban than rural beetles (Table 1, Fig. 4). LMM models revealed no significant sex-specific difference whether combined (Table 1) or analysed separately (Table S2). Similarly, the habitat × sex interaction was not a significant predictor for the predicted functional diversity of the gut bacterial microbiota (Table 1).
Boxplot of functional diversity of the gut bacterial microbiota (OTUs) of C. convexus females and males from rural and urban habitats expressed by the Rao’s quadratic entropy using the traits related to functional roles and respiratory types of the OTUs. In boxplots horizontal lines represent median values, boxes show the interquartile ranges, whiskers represent the minimum and maximum values, while points outside the whiskers are outliers.
Discussion
Methodological considerations
Recent studies of microbiota are increasingly use ASVs because these are generally considered more reliable64. Analysing microbiota using OTUs versus ASVs can give slightly different results. Even in C. convexus higher bacterial “species richness” was indicated using ASVs25 than in our OTU-based analysis. However, by the ASV-based analysis the most common group of the C. convexus microbiota was an unidentified phylum25. Much of our knowledge concerning insect microbiota originates from OTU-based studies. Consequently, a functional analysis—at least currently—ought to rely on OTU basis.
An additional hurdle is that the target 16s RNA genes do not code proteins, so functions can only be attributed by attaching known functions to the identified bacteria which is inevitably indirect, uncertain and imprecise. The resulting analysis should therefore be interpreted cautiously. However, the differences found in functional diversity do not merely reflect differences in taxonomic composition, because there were differences in their reaction to urbanization. The gut microbiota was more diverse in rural individuals, while urban beetles had a higher functional diversity of the gut microbiota.
Composition of the gut bacterial microbiota
The identified OTU richness was similar to previous findings, such as the 1,245 bacterial OTUs in the gut of the carabid Chlaenius pallipes (Gebler, 1823)65.
Similarly, the 21 identified bacterial phyla are not much different from the 19 bacterial phyla found in 218 insect species from 21 taxonomic orders66. The predominant bacterial phyla identified in the intestinal tract of C. convexus individuals sampled from wild populations (Firmicutes, Bacteroidetes, and Proteobacteria, representing more than 80% of the gut-associated bacteria) are generally frequent and thus also common in insects66,67, including ground beetles65,68.
The most abundant families in the gut of C. convexus were Enterococcaceae, Prevotellaceae, and Ruminococcaceae. Similarly, Enterococcaceae was among the most common families in the digestive tracts of ground beetles with various trophic habits31,34,65,69. In our study the average abundance of the Enterococcaceae, however, was significantly higher in the guts of urban than rural adults, suggesting some urbanization-related disturbance on the gut microbial composition. Similarly, another carabid, Pseudoophonus rufipes (DeGeer, 1774) had higher Enterococcaceae abundance when collected from conventionally managed fields with regular disturbance by tillage70.
Gut microbiota can significantly differ between sexes66. However, a previous study on the riparian bombardier beetle, Brachinus elongatulus Chaudoir, 187671, found no significant sex-specific differences in these bacterial communities. In contrast, we found significant differences, both by sex and by habitat. Assuming that microbial taxa in the host gut could be derived from the surrounding environment72, this habitat-related compositional difference can be explained by an impoverishment in the diversity of soil bacteria due to urbanization73. Sex-specific gut bacterial compositional difference, however, may also be triggered by the different mobility of sexes. Males with greater mobility, especially during the breeding period, are likely to pick up a wider range of bacterial symbionts from their environment than the less mobile females, especially in rural sites with a more diverse bacterial landscape. Higher between-individual variability of the gut microbiota in both rural and urban females suggest that diet-mediated stochastic acquisition could be another driver of the composition of gut bacterial microbiota. To cover the energy demand to produce eggs, females may consume a wider range of food items they encounter27, contributing to the between-individual variance in the gut microbiota.
Effects of urbanization on composition and diversity of the gut bacterial microbiota
Urbanization-triggered changes in the various environmental conditions can be reflected in behavioural, physiological, morphological, genetic and life-history parameters, and population size in ground beetles22.
To our knowledge, our OTU-based study is the first to assess the effect of urbanization on the gut bacterial microbiota of ground beetles. Our results show that the diversity of the gut bacterial microbiota was significantly reduced in urban habitats. Similarly reduced microbiota diversity has been observed in urban black-tailed prairie dogs (Cynomys ludovicianus (Ord, 1815)74), and in eastern grey squirrels (Sciurus carolinensis Gmelin, 178875). Urban house sparrows (Passer domesticus (Linnaeus, 1758)) also have less diverse gut microbiota than rural conspecifics76. However, urban environments can also allow a more diverse microbiota, as registered in coyotes (Canis latrans Say, 182377), white-crowned sparrow (Zonotrichia leucophrys (Forster, 1772)78), and eastern water dragons (Intellagama lesueurii Gray, 183179).
In flying insects, the gut bacterial microbiota diversity can also be higher (e.g., bumblebees, Bombus terrestris (L., 1758)23), or lower (e.g., honey bee, Apis mellifera L, 175880) in rural than urban individuals, while no such difference exists in adult dragonflies24. These inconsistent results indicate that the gut bacterial communities in rural and urban individuals may differ even in closely related species. This is exacerbated if a single diversity measure (richness or a single taxonomic diversity index (e.g. Shannon or the Simpson diversity) is used when evaluating diversity. Therefore, using a scalable diversity approach (such as Hill numbers) is recommended to get a more detailed understanding of the diversity of gut bacterial microbiota.
Bacterial microbiota are acquired from the surrounding environment16,72 and/or from food items81. The lower diversity of gut bacterial microbiota in urban C. convexus could reflect the impoverished microbial diversity and reduced food spectrum in urban habitats. However, microbial diversity in urban soils can be lower82, higher83,84,85, or non-different86 in urban than in rural habitats. Although anthropogenic activities may cause serious decrease in arthropod biomass4, estimating the range of potential food items for a carnivorous ground beetle remains a serious challenge.
Previous work in these habitats indicates that the abundance of potential prey items for several groups (other beetles, chilopods, diplopods, gastropods, isopods) was significantly lower in the studied urban than rural areas30,87. Thus, urban C. convexus can experience a less diverse bacterial landscape, leading to microbiota of higher variability with some new functions but overall, a reduced diversity.
The size of the convex hull on the ordination plot, a measure of β-diversity, of the gut samples from C. convexus, however, was bigger in urban than rural habitat, irrespective of whether the sexes were combined or treated separately, indicating higher between-individual variability in urban than rural beetles. Similarly, there is higher β-diversity of the gut bacterial microbiota in urban than rural habitats in mammals74, and birds88. Furthermore, higher β-diversity (between-sites variability) of soil bacterial communities in urban than rural habitats was also reported84. Thus, we assume that soil bacterial diversity was lower in our urban habitat but their composition varied more, resulting in higher soil bacterial β-diversity. This can also be manifested in the gut microbiota through stochastic acquisition of bacteria from soils31. In urban habitats the occasional consumption of human-supplied food items (e.g. pet food, garbage) could also substantially affect the hosts’ gut microbiota composition89. Thus, consumption of anthropogenic food may contribute to the higher β-diversity of gut bacterial communities in urban C. convexus.
These new types of resources may promote the growth of specific bacteria with various functional roles, possibly assisting in more efficient digestion of new food items. This could expand the diet breath, allowing beetles to exploit various food resources81. Indeed, a broader food spectrum is associated with richer, and probably functionally more diverse gut microbiota66,81.
Conclusions
Using next generation sequencing of the bacterial 16S rRNA gene, we showed that the composition of the gut bacterial microbiota in C. convexus was significantly different between rural and urban individuals, while microbiota diversity was significantly reduced in urban habitats. However, their predicted functional diversity was not significantly reduced. As a substantial part of gut bacteria in beetles is derived from the surrounding environment by horizontal transmission31, the observed changes were plausibly triggered by urbanization.
Biodiversity in urban habitats has several advantages for humans. To make urban areas healthier for their inhabitants, maintaining existing urban green habitats, restoring degraded ones, creating green roofs, green walls, urban and community gardens are suggested90. These measures would probably favour microbes as well, with hopefully positive effects on higher organisms via more diverse microbial landscapes.
Data availability
Metadata and raw sequence reads are deposited in the NCBI SRA database (BioProject PRJNA1051584, https://www.ncbi.nlm.nih.gov/bioproject/PRJNA1051584).
References
Forester, D. J. & Machlist, G. E. Modeling human factors that affect the loss of biodiversity. Conserv. Biol.10, 1253–1263 (1996).
Norris, K. Agriculture and biodiversity conservation: Opportunity knocks. Conserv. Lett.1, 2–11 (2008).
Paillet, Y. et al. Biodiversity differences between managed and unmanaged forests: Meta-analysis of species richness in Europe. Conserv. Biol.24, 101–112 (2010).
Hahs, A. K. et al. Urbanisation generates multiple trait syndromes for terrestrial animal taxa worldwide. Nat. Commun.14, 4751 (2023).
McIntyre, N. E. Urban ecology—Definitions and goals. In The Routledge Handbook of Urban Ecology (eds. Douglas, I., Goode, D., Houck, M. & Wang, R.) 7–16 (Routledge, 2011).
Parris, K. M. Ecology of Urban Environments (Wiley-Blackwell, 2016).
Kalnay, E. & Cai, M. Impact of urbanization and land-use change on climate. Nature423, 528–531 (2003).
Grimm, N. B. et al. Global change and the ecology of cities. Science.319, 756–760 (2008).
McDonnell, M. J. et al. Ecosystem processes along an urban-to-rural gradient. Urban Ecosyst.1, 21–36 (1997).
McKinney, M. L. Effects of urbanization on species richness: A review of plants and animals. Urban Ecosyst.11, 161–176 (2008).
Sol, D. et al. The worldwide impact of urbanisation on avian functional diversity. Ecol. Lett.23, 962–972 (2020).
McKinney, M. L. Urbanization as a major cause of biotic homogenization. Biol. Conserv.127, 247–260 (2006).
Zhang, X., Chen, X., Liu, M., Xu, Z. & Wei, H. Coupled changes in soil organic carbon fractions and microbial community composition in urban and suburban forests. Sci. Rep.10, 15933 (2020).
Mukhtar, H. et al. Soil microbiome feedback to climate change and options for mitigation. Sci. Total Environ.882, 163412 (2023).
Ottman, N., Smidt, H., de Vos, W. & Belzer, C. The function of our microbiota: who is out there and what do they do?. Front. Cell. Infect. Microbiol.2, 104 (2012).
Engel, P. & Moran, N. A. The gut microbiota of insects—Diversity in structure and function. FEMS Microbiol. Rev.37, 699–735 (2013).
Kikuchi, Y. et al. Symbiont-mediated insecticide resistance. Proc. Natl. Acad. Sci.109, 8618–8622 (2012).
Bai, S., Yao, Z., Raza, M. F., Cai, Z. & Zhang, H. Regulatory mechanisms of microbial homeostasis in insect gut. Insect Sci.28, 286–301 (2021).
Douglas, A. E. Multiorganismal insects: Diversity and function of resident microorganisms. Annu. Rev. Entomol.60, 17–34 (2015).
Borer, E. T., Kinkel, L. L., May, G. & Seabloom, E. W. The world within: Quantifying the determinants and outcomes of a host’s microbiome. Basic Appl. Ecol.14, 533–539 (2013).
Wang, H., Jin, L. & Zhang, H. Comparison of the diversity of the bacterial communities in the intestinal tract of adult Bactrocera dorsalis from three different populations. J. Appl. Microbiol.110, 1390–1401 (2011).
Magura, T. & Lövei, G. L. Consequences of urban living: Urbanization and ground beetles. Curr. Landsc. Ecol. Reports6, 9–21 (2021).
Bosmans, L. et al. Habitat-specific variation in gut microbial communities and pathogen prevalence in bumblebee queens (Bombus terrestris). PLoS One13, 1–19 (2018).
Nobles, S. & Jackson, C. R. Effects of life stage, site, and species on the dragonfly gut microbiome. Microorganisms8, 183 (2020).
Magura, T. et al. Urbanization reduces gut bacterial microbiome diversity in a specialist ground beetle. Carabus convexus. Mol. Ecol.33, e17265 (2024).
Brooks, A. W., Kohl, K. D., Brucker, R. M., van Opstal, E. J. & Bordenstein, S. R. Phylosymbiosis: Relationships and functional effects of microbial communities across host evolutionary history. PLoS Biol.14, e2000225 (2016).
Lövei, G. L. & Sunderland, K. D. Ecology and behavior of ground beetles (Coleoptera: Carabidae). Annu. Rev. Entomol.41, 231–256 (1996).
Niemelä, J. et al. The search for common anthropogenic impacts on biodiversity: A global network. J. Insect Conserv.4, 3–9 (2000).
Turin, H., Penev, L. & Casale, A. The Genus Carabus in Europe—A Synthesis (Pensoft, 2003).
Magura, T., Lövei, G. L. & Tóthmérész, B. Time-consistent rearrangement of carabid beetle assemblages by an urbanisation gradient in Hungary. Acta Oecol.34, 233–243 (2008).
Kudo, R., Masuya, H., Endoh, R., Kikuchi, T. & Ikeda, H. Gut bacterial and fungal communities in ground-dwelling beetles are associated with host food habit and habitat. ISME J.13, 676–685 (2019).
Klindworth, A. et al. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucl. Acids Res.41, e1 (2013).
Eisenhofer, R. et al. Contamination in low microbial biomass microbiome studies: Issues and recommendations. Trends Microbiol.27, 105–117 (2019).
Giglio, A., Vommaro, M. L., Gionechetti, F. & Pallavicini, A. Gut microbial community response to herbicide exposure in a ground beetle. J. Appl. Entomol.145, 986–1000 (2021).
Escudié, F. et al. FROGS: Find, rapidly, OTUs with galaxy solution. Bioinformatics34, 1287–1294 (2018).
Magoč, T. & Salzberg, S. L. FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics27, 2957–2963 (2011).
Mahé, F., Rognes, T., Quince, C., de Vargas, C. & Dunthorn, M. Swarm: Robust and fast clustering method for amplicon-based studies. PeerJ2, e593 (2014).
Edgar, R. C., Haas, B. J., Clemente, J. C., Quince, C. & Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics27, 2194–2200 (2011).
Rognes, T., Flouri, T., Nichols, B., Quince, C. & Mahé, F. VSEARCH: A versatile open source tool for metagenomics. PeerJ4, e2584 (2016).
Bokulich, N. A. et al. Quality-filtering vastly improves diversity estimates from Illumina amplicon sequencing. Nat. Methods10, 57–59 (2013).
Camacho, C. et al. BLAST+: Architecture and applications. BMC Bioinform.10, 421 (2009).
Quast, C. et al. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucl. Acids Res.41, D590–D596 (2013).
Louca, S., Parfrey, L. W. & Doebeli, M. Decoupling function and taxonomy in the global ocean microbiome. Science353, 1272–1277 (2016).
R Core Team. R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing (2023).
Dillies, M.-A. et al. A comprehensive evaluation of normalization methods for Illumina high-throughput RNA sequencing data analysis. Brief. Bioinform.14, 671–683 (2012).
McMurdie, P. J. & Holmes, S. phyloseq: An R Package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE8, 1–11 (2013).
Evans, C., Hardin, J. & Stoebel, D. M. Selecting between-sample RNA-Seq normalization methods from the perspective of their assumptions. Brief. Bioinform.19, 776–792 (2018).
Oksanen, J. et al. vegan: Community ecology package. (2022).
Venables, W. & Ripley, B. Modern Applied Statistics with S. (Springer, Berlin, 2002).
Hervé, M. RVAideMemoire: Testing and plotting procedures for biostatistics. (2021).
Hurlbert, S. H. The nonconcept of species diversity: A critique and alternative parameters. Ecology52, 577–586 (1971).
Maurer, B. A. & McGill, B. J. Measurement of species diversity. In Biological Diversity: Frontiers in Measurement and Assessment (eds. Magurran, A. E. & McGill, B. J.) 55–65 (Oxford University Press, 2011).
Hill, M. O. Diversity and evenness: A unifying notation and its consequences. Ecology54, 427–432 (1973).
Ricotta, C. & Feoli, E. Hill numbers everywhere. Does it make ecological sense?. Ecol. Indic.161, 111971 (2024).
Chao, A. et al. Rarefaction and extrapolation with Hill numbers: A framework for sampling and estimation in species diversity studies. Ecol. Monogr.84, 45–67 (2014).
Li, D. hillR: taxonomic, functional, and phylogenetic diversity and similarity through Hill numbers. J. Open Source Softw.3, 1041 (2018).
Botta-Dukát, Z. Rao’s quadratic entropy as a measure of functional diversity based on multiple traits. J. Veg. Sci.16, 533–540 (2005).
Rao, C. R. Diversity and dissimilarity coefficients: A unified approach. Theor. Popul. Biol.21, 24–43 (1982).
Debastiani, V. J. & Pillar, V. D. SYNCSA - R tool for analysis of metacommunities based on functional traits and phylogeny of the community components. Bioinformatics28, 2067–2068 (2012).
D’Orazio, M., Di Zio, M. & Scanu, M. Statistical Matching: Theory and Practice (Wiley, 2006). https://doi.org/10.1002/0470023554.
Bates, D., Mächler, M., Bolker, B. & Walker, S. Fitting linear mixed-effects models using lme4.. J. Stat. Softw. Artic.67, 1–48 (2015).
Fox, J. & Weisberg, S. An R Companion to Applied Regression (SAGE Publications, 2018).
Zuur, A., Ieno, E. N., Walker, N., Saveliev, A. A. & Smith, G. M. Mixed Effects Models and Extensions in Ecology with R (Springer, 2009). https://doi.org/10.1007/978-0-387-87458-6.
Chiarello, M., McCauley, M., Villéger, S. & Jackson, C. R. Ranking the biases: The choice of OTUs versus ASVs in 16S rRNA amplicon data analysis has stronger effects on diversity measures than rarefaction and OTU identity threshold. PLoS ONE17, 1–19 (2022).
Do, Y., Park, J.-K., Park, W.-B. & Kim, M.-S. The gut bacterial community of Chlaenius pallipes (Coleoptera: Carabidae) associates with their habitat and morphology. Insects13, 1099 (2022).
Yun, J.-H. et al. Insect gut bacterial diversity determined by environmental habitat, diet, developmental stage, and phylogeny of host. Appl. Environ. Microbiol.80, 5254–5264 (2014).
Colman, D. R., Toolson, E. C. & Takacs-Vesbach, C. D. Do diet and taxonomy influence insect gut bacterial communities?. Mol. Ecol.21, 5124–5137 (2012).
Do, Y. et al. Host and environmental influences on the gut bacterial community of carabid beetles in distinct paddy fields. Entomol. Res.53, 509–517 (2023).
Kolasa, M. et al. How hosts taxonomy, trophy, and endosymbionts shape microbiome diversity in beetles. Microb. Ecol.78, 995–1013 (2019).
Magagnoli, S. et al. The ground beetle Pseudoophonus rufipes gut microbiome is influenced by the farm management system. Sci. Rep.12, 22638 (2022).
McManus, R., Ravenscraft, A. & Moore, W. Bacterial associates of a gregarious riparian beetle with explosive defensive chemistry. Front. Microbiol.9, 2361 (2018).
Dillon, R. J. & Dillon, V. M. The gut bacteria of insects: Nonpathogenic interactions. Annu. Rev. Entomol.49, 71–92 (2004).
Qu, Y. et al. Characters and environmental driving factors of bacterial community in soil of Beijing urban parks. Environ. Res.215, 114178 (2022).
Neha, S. A. & Salazar-Bravo, J. Fine-scale spatial variation shape fecal microbiome diversity and composition in black-tailed prairie dogs (Cynomys ludovicianus). BMC Microbiol.23, 51 (2023).
Stothart, M. R. & Newman, A. E. M. Shades of grey: host phenotype dependent effect of urbanization on the bacterial microbiome of a wild mammal. Anim. Microbiome3, 46 (2021).
Teyssier, A. et al. Diet contributes to urban-induced alterations in gut microbiota: Experimental evidence from a wild passerine. Proc. R. Soc. B Biol. Sci.287, 20192182 (2020).
Sugden, S., Sanderson, D., Ford, K., Stein, L. Y. & St. Clair, C. C. An altered microbiome in urban coyotes mediates relationships between anthropogenic diet and poor health. Sci. Rep.10, 22207 (2020).
Berlow, M., Phillips, J. N. & Derryberry, E. P. Effects of urbanization and landscape on gut microbiomes in white-crowned sparrows. Microb. Ecol.81, 253–266 (2021).
Littleford-Colquhoun, B. L., Weyrich, L. S., Jackson, N. & Frere, C. H. City life alters the gut microbiome and stable isotope profiling of the eastern water dragon (Intellagama lesueurii). Mol. Ecol.28, 4592–4607 (2019).
Damico, M. E., Rueppell, O., Shaffer, Z., Han, B. & Raymann, K. High royal jelly production does not impact the gut microbiome of honey bees. Anim. Microbiome3, 60 (2021).
Brunetti, M., Magoga, G., Gionechetti, F., De Biase, A. & Montagna, M. Does diet breadth affect the complexity of the phytophagous insect microbiota? The case study of Chrysomelidae. Environ. Microbiol.24, 3565–3579 (2022).
Wang, H. et al. Changes in land use driven by urbanization impact nitrogen cycling and the microbial community composition in soils. Sci. Rep.7, 44049 (2017).
Gao, D. et al. Urbanization imprint on soil bacterial communities in forests and grasslands. Forests14, 38 (2023).
Scholier, T. et al. Urban forest soils harbour distinct and more diverse communities of bacteria and fungi compared to less disturbed forest soils. Mol. Ecol.32, 504–517 (2023).
Li, M. et al. Effects of urban–rural environmental gradient on soil microbial community in rapidly urbanizing area. Ecosyst. Heal. Sustain.9, 0118 (2024).
Epp Schmidt, D. J. et al. Urbanization erodes ectomycorrhizal fungal diversity and may cause microbial communities to converge. Nat. Ecol. Evol.1, 123 (2017).
Magura, T., Tóthmérész, B. & Molnár, T. A species-level comparison of occurrence patterns in carabids along an urbanisation gradient. Landsc. Urban Plan.86, 134–140 (2008).
Phillips, J. N., Berlow, M. & Derryberry, E. P. The effects of landscape urbanization on the gut microbiome: An exploration into the gut of urban and rural white-crowned sparrows. Front. Ecol. Evol.6, 148 (2018).
Knutie, S. A., Chaves, J. A. & Gotanda, K. M. Human activity can influence the gut microbiota of Darwin’s finches in the Galapagos Islands. Mol. Ecol.28, 2441–2450 (2019).
Fenoglio, M. S., Calviño, A., González, E., Salvo, A. & Videla, M. Urbanisation drivers and underlying mechanisms of terrestrial insect diversity loss in cities. Ecol. Entomol.46, 757–771 (2021).
Acknowledgements
We thank I. Nagy (Aarhus University, Denmark) for advice on analysing functional diversity. This research was funded by the National Research, Development and Innovation Fund, grant numbers OTKA K-131459 and OTKA K-146628. Sampling of beetles was by permission of the Department of Green Infrastructure of the Mayor’s Office of Debrecen (no.: ZÖLD-82312-2/2021). Authorship is by the “first-and-last-author-emphasis” (FLAE) principle.
Author information
Authors and Affiliations
Contributions
T.M.: conceptualization, data curation, formal analysis, funding acquisition, investigation, methodology, supervision, project administration, resources, validation, visualization, writing — original draft, writing—review and editing. S.M.: investigation, methodology, writing—review and editing. R.H.: investigation, methodology, writing—review and editing. M.T.: investigation, methodology, writing—review and editing. G.L.L.: conceptualization, methodology, supervision, validation, writing — original draft, writing—review and editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Magura, T., Mizser, S., Horváth, R. et al. Urbanization impoverishes taxonomic but not functional diversity of the gut microbiota in a forest specialist ground beetle, Carabus convexus. Sci Rep 14, 25546 (2024). https://doi.org/10.1038/s41598-024-75864-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-75864-6