Abstract
Exposure to mixtures of toxic metals is known to cause adverse health effects through epigenetic alterations. Here we aimed to examine the unexplored area of aberrant DNA methylation in the H19/IGF2 domain following combined toxic metal exposure. An in vitro epigenotoxicity assay using the human normal liver epithelial cell line THLE-3 was conducted. When THLE-3 cells were exposed to specific concentrations of either organic arsenic or MeHgCl, an increase in the H19 lncRNA levels and a marked reduction in the IGF2 mRNA levels were observed. In contrast, combined exposures coupled with CdCl2 resulted in the transcriptional repression of H19 and transcriptional activation of IGF2. It should be noted that the correlation between the dysregulated expression of H19/IGF2 and the hypermethylated CpG sites within the H19 differentially methylated region (DMR) was statistically significant. Furthermore, we performed transcriptomic analysis of the hepatocytes exposed to toxic metal combinations indicating enrichment of pro-inflammatory and anti-proliferative pathways compared to the unexposed cells. Our results suggest that hazardous metal mixtures may trigger epigenetic aberrations at the H19/IGF2 locus. We propose that altered CpG methylation in the H19 DMR could be a candidate biomarker for hepatic epigenotoxicity, in part, due to environmental exposure.
Similar content being viewed by others
Introduction
With the advancement of industrialization and urbanization, toxic metals have permeated our environment, appearing in air, soil, water, food, and tobacco smoke1,2. Consequently, this widespread contamination presents a significant risk of exposure to the general population through various means such as drinking water, food consumption, air inhalation, and skin contact3. Combined exposures to multiple toxic metals, such as arsenic, cadmium, and mercury, have been observed in populations due to environmental and occupational factors4. Such exposures can lead to additive and synergistic effects, exacerbating health risks5,6. Recent studies, including data from large-scale health surveys such as the National Health and Nutrition Examination Survey (NHANES) in the United States and the Korean National Environmental Health Survey (KoNEHS), have indicated that individuals worldwide are exposed to multiple toxic metals simultaneously7,8. The adverse health effects of toxic metal exposure have led to global ecological and health concerns, and several metals (e.g., arsenic, cadmium and methylmercury) have the potential to cause epigenetic changes, such as DNA methylation, and lead to disease9. Toxic metals can be classified as epigenotoxic substances, which are defined as contaminants that cause epigenetic alterations and adverse health outcomes by triggering signaling pathways10. However, the complexity and variability of multiple exposures have historically posed challenges to comprehensive studies, contributing to the limited research in this area. Consequently, these epigenotoxic events are valuable as biomarkers for early disease diagnosis and prediction, as well as for identifying potential therapeutic targets.
Chronic liver inflammation, often resulting in liver cirrhosis, is a major cause of death worldwide, including in the United States11. This condition encompasses various chronic pathologies, with the main causes including viral infections (such as hepatitis B and C), alcoholic liver disease, nonalcoholic fatty liver disease (NAFLD), and autoimmune disorders12. While these factors are the primary drivers of chronic hepatitis, environmental toxicants, including toxic metals, are increasingly recognized as significant contributors to liver inflammation and dysfunction13. Toxic metals, specifically, are known to induce liver inflammation through pathways involving TNF-α, pro-inflammatory cytokines, MAPK, and ERK. In particular, arsenic, cadmium, and mercury, in particular, have been associated with various hepatotoxic effects, including oxidative stress, mitochondrial dysfunction, and alterations in cell signaling pathways14. However, while the pathogenic effects of individual toxic metal exposure on the liver have been studied, the inflammatory consequences of metal mixture exposure remain less understood. Over time, these metals can accumulate in various human tissues, organs, and systems15, with the liver being a primary site of accumulation during prolonged exposure16.
The importance of studying combined toxic metal exposures cannot be overstated, as real-world scenarios often involve simultaneous exposure to multiple contaminants. Recent population studies have highlighted the prevalence of co-exposures to multiple toxic metals. For instance, one study by found metal accumulation in the sediments following dam construction including arsenic, lead, and mercury17. Similarly, neurodevelopmental impairments have been reported in children with prenatal exposure to relatively high levels of both mercury and arsenic18. These findings underscore the need for more comprehensive research on the health effects of combined metal exposures, especially in vulnerable populations.
Epigenetic alterations are increasingly recognized as a crucial mechanism by which toxic metals exert their detrimental effects on human health. Several studies have demonstrated that exposure to individual toxic metals can induce epigenetic changes, including altered DNA methylation, histone modifications and non-coding RNA expression9. For instance, arsenic exposure has been associated with hepatic global DNA hypomethylation and gene-specific hypermethylation19, while cadmium has been shown to alter histone modifications and microRNA expression20,21. However, the epigenetic and/or epigenomic effects of metal mixture exposures, which more closely reflect real-world scenarios, remain largely unexplored22. Given that individuals are often simultaneously exposed to multiple toxic metals, there is an urgent need for research monitoring the epigenotoxic impacts of metal mixtures, particularly on target organs such as the liver. While previous research has explored lead-mediated epigenetic alterations of H19/IGF223, studies on the effects of other toxic metals, such as arsenic, cadmium, and mercury, remain scarce. Due to the high toxicity and widespread environmental presence of these metals, research on their impact on H19/IGF2 dysregulation is urgently needed.
The H19/IGF2 imprinted domain is of particular interest in the context of metal-induced hepatotoxicity. This domain, located on human chromosome 11, is epigenetically regulated and plays crucial roles in growth regulation and liver function23. Abnormal DNA methylation in the H19/IGF2 imprinted domain has been observed in hepatocellular carcinoma (HCC) tissues and cell lines, suggesting a potential role for epigenetic dysregulation of these genes as biomarkers of liver malignancy24. Moreover, the H19 long non-coding RNA (lncRNA) and IGF2 mRNA have been implicated in various liver pathologies, including NAFLD and liver fibrosis25. Despite these important functions, the epigenotoxic influences of metal exposure, particularly mixture exposures, on the epigenetic regulation of the H19/IGF2 domain in liver cells remain poorly understood.
In this study, we aimed to investigate the epigenetic effects of combined toxic metal exposure on the H19/IGF2 domain in normal human liver epithelial cells, addressing the critical knowledge gap in understanding real-world, multi-metal exposure scenarios. Specifically, our objectives were threefold: (1) to develop an in vitro epigenotoxicity assay for assessing the impacts of individual and combined metal exposures on gene expression and DNA methylation of the H19/IGF2 locus; (2) to examine the methylation profiles at key regulatory regions of the H19/IGF2 domain in response to metal mixtures; and (3) to conduct transcriptome profiling to elucidate potential links between H19/IGF2 dysregulation and altered biological pathways indicative of hepatic malfunction. By integrating these approaches, we sought to identify novel epigenetic biomarkers for metal-induced hepatotoxicity and provide mechanistic insights into how metal mixture exposures may contribute to liver pathogenesis. This research aims to advance our understanding of environmental epigenotoxicity and inform risk assessment strategies for complex metal exposures.
Results
Individual exposure to toxic metals induced the changes in gene expression of both H19 and IGF2
Our investigation into how single toxic metal exposure affects gene expression in the normal human liver cell line THLE-3 showed that exposure to organic arsenic (oAs) at concentrations ranging from 1 nM to 10 µM approximately doubled the expression levels of the H19 lncRNA. In contrast, cadmium chloride (CdCl2) or methylmercury chloride (MeHgCl) exposure did not alter the H19 lncRNA levels in THLE-3, except for a significant increase observed at 100 pM MeHgCl exposure (p < 0.05) (Fig. 1a). Similarly, no significant changes were observed in the mRNA levels of IGF2 across other concentration ranges of oAs or CdCl2. Of note, a significant reduction (p < 0.001) in the IGF2 mRNA levels was found after exposure to 10 and 100 nM MeHgCl, respectively (Fig. 1b).
Gene expression changes in the H19/IGF2 domain after individual exposures to increasing doses of each toxic metal. RT-qPCR analysis showed the expression levels of (a) H19 lncRNA and (b) IGF2 mRNA in THLE-3 cells after individual exposure to toxic metals for 72 h. The relative expression levels were normalized by the mean Ct values of ACTB and GAPDH. Shown are the mean ± standard error of independent experiments in triplicate for each dose of toxic metals. One-way ANOVA followed by post hoc Tukey’s test was performed using the unexposed control as the reference group for all comparisons (*p < 0.05, **p < 0.01, ***p < 0.001).
Combined exposure to toxic metals provoked the opposite pattern of gene expressions compared to the individual exposures
Next, the treatments with toxic metal mixtures in THLE-3 cells aimed to detect any additive or synergistic effects on the H19/IGF2 locus. Interestingly, we found the distinct pattern of gene expression changes in response to the combined exposures compared to individual metal treatments. Specifically, we observed significant downregulation of H19 lncRNA and an upregulation of IGF2 mRNA at certain metal combinations: 10 µM of oAs + 1 µM of CdCl2, and 1 µM of CdCl2 + 10 nM of MeHgCl. These changes were not observed when cells were exposed to these metal concentrations individually, suggesting a unique response to metal mixtures. Notably, the combined exposure to oAs and MeHgCl did not induce marked transcriptional dysregulation, except for a slight decrease in the H19 expression at lower concentration ranges (Fig. 2). This differential response to various metal combinations underscores the complexity of metal mixture effects and highlights the importance of studying diverse combinations of environmental contaminants.
Combined exposure to toxic metals coupled with CdCl2 impacted on the expression of the H19/IGF2 domain in THLE-3 cells. The expression of (a) H19 and (b) IGF2 was assessed in THLE-3 cells after exposure to toxic metals in combination for 72 h. The relative expression levels were normalized by the mean Ct values of ACTB and GAPDH. Shown are the mean ± standard error of independent experiments in triplicate for each dose of toxic metals. One-way ANOVA followed by post hoc Tukey’s test was performed using the unexposed control as the reference group for all comparisons (*p < 0.05, **p < 0.01, ***p < 0.001).
Epigenotoxic effect of toxic metals mixture on DNA methylation status of the H19/IGF2 domain
To investigate whether the mutually exclusive alterations in gene expression were linked to toxic metal-induced epigenetic modifications within the H19/IGF2 domain, we conducted a comprehensive quantitative analysis of methylation levels at the H19 differentially methylated region (DMR) and imprinting control region (ICR). Bisulfite amplicon sequencing (BSAS) was employed to analyze the genomic regions that are crucial for the establishment of imprinting regulation. Specifically, we focused on the H19 DMR and ICR within the H19/IGF2 domain, which are functional elements directing the allelic transcription of the H19 and IGF2 genes; a genomic region (chr11:1,998,273-1,998,659) harboring 21 CpGs within the H19 DMR and another genomic region (chr11:2,001,023 − 2,001,370) encompassing 17 CpGs within the ICR, respectively.
We compared the altered DNA methylation levels of the 21 CpG sites within the H19 DMR between the samples exposed to individual toxic metals and those exposed to a combination of toxic metals, e.g., the H19-downregulating and IGF2-upregulating samples. While no significant CpG methylation changes were observed in the H19 DMR after exposure to individual toxic metals, exposure to 10 µM of oAs showed an overall, statistically insignificant hypermethylation in the H19 DMR. Conversely, significant CpG hypermethylation in the H19 DMR was exhibited in the THLE-3 cells exposed to toxic metal mixtures, related to dysregulated expression of the H19 and IGF2 genes (Fig. 3a). This correlation between hypermethylation of the H19 DMR and the downregulation of H19 expression, along with the upregulation of IGF2, suggests a potential mechanistic link between metal-induced epigenetic changes and gene expression alterations in this imprinted domain. Notably, the combined exposure to 10 µM of oAs + 1 µM of CdCl2 led to a significantly higher DNA methylation level of CpG loci (p < 0.01) compared to 1 µM of CdCl2 + 10 nM of MeHgCl exposure (p < 0.05). Taken together, the results of our quantitative analysis suggest that the H19 DMR examined could be considered a responsive genomic region of exposure to toxic metal mixtures.
Epigenetic dysfunction of CpG loci within the H19 DMR and the ICR in the presence of exposure to toxic metals. (a) BSAS analysis results in the H19 DMR following exposure to toxic metals, either alone or in combination. (b) BSAS analysis results in ICR following exposure to toxic metals, either alone or in combination. Shown are the mean ± standard error of independent experiments in triplicate for each dose of toxic metals. One-way ANOVA followed by post hoc Tukey’s test was performed using the unexposed control as the reference group for all comparisons (*p < 0.05, **p < 0.01, ***p < 0.001). DMR, differentially methylated region; ICR, Imprinting control region; BSAS, Bisulfite amplicon sequencing; oAs, 10 µM of oAs; CdCl2, 1 µM of CdCl2; MeHgCl, 10 nM of MeHgCl; oAs + CdCl2, 10 µM of oAs + 1 µM of CdCl2; CdCl2 + MeHgCl, 1 µM of CdCl2 + 10 nM of MeHgCl.
Subsequently, we assessed the DNA methylation levels of the 17 CpG sites within the ICR using the equivalent exposure concentrations. The significant changes in the CpG methylation levels were not detected in the ICR, compared to the H19 DMR (Fig. 3b). Intriguingly, individual exposure to 10 µM of oAs or 1 µM of CdCl2 resulted in a significant reduction in the DNA methylation levels at the site-specific CpGs. In contrast to the H19 DMR, these findings imply that combined exposure to toxic metals might not impact on the ICR examined to elicit epigenetic dysregulation in the H19/IGF2 locus.
Toxic metals exposure-specific correlation between the gene expression changes and the altered CpG methylation levels at the H19/IGF2 domain
To pinpoint potential biomarkers for toxic metal exposures within the H19/IGF2 domain, we performed correlation analysis between gene expression levels and the CpG methylation levels in the H19 DMR and ICR regulatory elements. We found that hypermethylation of CpG loci in the H19 DMR was inversely correlated with the H19 lncRNA expression and positively correlated with the IGF2 mRNA expression, regardless of the type of toxic metal exposure. In the THLE-3 cells exposed to 10 µM of oAs + 1 µM of CdCl2, the identical 16 out of 21 CpG units displayed significant correlations (p < 0.05) for the H19 repression and the IGF2 activation (Fig. 4a). In addition, the 5 CpGs for the IGF2 activation and the overlapping 3 CpGs for the H19 repression exhibited a more significant correlation (p < 0.01) in the cells exposed to 1 µM of CdCl2 + 10 nM of MeHgCl (Fig. 4b). In contrast, any significant correlations between the H19/IGF2 gene expression changes and the altered CpG methylation at the ICR examined were not detected, compared to the results from the H19 DMR (Fig. S1).
Correlation between DNA methylation changes of H19 DMR and expression of H19/IGF2 domain in the presence of combined exposure to toxic metals. Pairwise Spearman rank correlation between DNA methylation level of the H19 DMR and expression of target genes (H19 and IGF2). Toxic metal-specific correlations are shown for exposure to (a) 10 µM of oAs + 1 µM of CdCl2 and (b) 1 µM of CdCl2 + 10 nM of MeHgCl, respectively. Pairwise Spearman rank correlation, *p < 0.05, **p < 0.01; DMR, differentially methylated region; oAs + CdCl2, 10 µM of oAs + 1 µM of CdCl2; CdCl2 + MeHgCl, 1 µM of CdCl2 + 10 nM of MeHgCl.
Mixture exposure to oAs + CdCl2 or CdCl2 + MeHgCl triggered transcriptomic responses related to the hepatic pro-inflammation and anti-proliferation
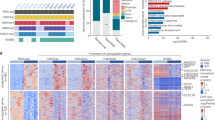
To investigate the epigenetic changes in the H19/IGF2 domain induced by two different exposure combinations, 10 µM of oAs + 1 µM of CdCl2 and 1 µM of CdCl2 + 10 nM of MeHgCl, a transcriptome analysis was performed in THLE-3 cells. In the mixture exposure of 10 µM of oAs + 1 µM of CdCl2, two more enriched pathway with an FDR q-value less than 0.05 was presented in a positive manner compared to the CdCl2 + oAs (6 upregulated & 5 downregulated pathways in Fig. 5a). In contrast, in the combined exposure of 1 µM CdCl2 + 10 nM MeHgCl, six additional enriched pathways were negatively affected compared to the oAs + CdCl2 group (4 upregulated & 10 downregulated pathways in Fig. 5b). Additionally, the significant changes with an FDR q-value less than 0.01 (red bars in Fig. 5) in the enriched pathways were found to be greater in the mixture exposure of 1 µM of CdCl2 + 10 nM of MeHgCl. Among them, the TNF-α signaling via NF-κB-related gene set was upregulated under both metal mixture exposures, while MYC targets V1, Mitotic Spindle, and E2F Targets-related gene sets were downregulated under both combined metal exposures (Fig. 5). It should be noted that the oxidative phosphorylation pathway, was upregulated in the presence of the oAs + CdCl2 mixture, but downregulated following the mixture exposure to the CdCl2 + MeHgCl mixture (Fig. S2b).
Hepatic transcriptome profiling of combined exposure to toxic metals in the THLE-3 cells using RNA-seq analysis. GSEA exhibiting the enriched gene sets in the THLE-3 cells exposed to (a) 10 µM of oAs + 1 µM of CdCl2 and (b) 1 µM of CdCl2 + 10 nM of MeHgCl compared to the unexposed control. Bars in red represent significant enrichment at FDR q-value < 0.01, whereas bars in black indicate gene sets with an FDR q-value between 0.01 and 0.05. Control, unexposed control; oAs + CdCl2, 10 µM of oAs + 1 µM of CdCl2; CdCl2 + MeHgCl, 1 µM of CdCl2 + 10 nM of MeHgCl.
Discussion
Environmental pollutants can have a considerable impact on gene expression, which can lead to various health risks26. In this study, we investigated the epigenotoxic effects of metal mixtures, including oAs, CdCl2, and MeHgCl, on the H19/IGF2 domain in the THLE-3 cell line. Through the use of an in vitro epigenotoxicity assay, we observed significant transcriptional alterations of the biallelically expressed imprinted genes H19 and IGF2, showing the opposite transcriptional pattern between individual and combined exposure. Next, quantitative analysis of the DNA methylation levels revealed that these alterations were closely linked to hypermethylation at the site-specific CpG units within the H19 DMR. After the correlation analysis between the H19/IGF2 gene expression changes and the altered CpG methylation levels, we identified the distinct CpG dinucleotide loci in the H19 DMR that could be affected by different combinations of toxic metals. Furthermore, hepatic transcriptome profiling indicated a potential link between the metal mixture-mediated epigenetic dysregulation and compelling pathophysiological responses, such as pro-inflammation and anti-proliferation. Consequently, our findings provided valuable insights into the complex epigenetic dynamics derived by exposure to toxic metal mixtures, in part, reflecting its potential impact on hepatic gene regulation and health implications.
Supporting our hypothesis, we observed the modulation of gene expression in human hepatocytes at low concentrations of toxic metals, which is consistent with the effects of endocrine-disrupting chemicals (EDCs) that produce non-monotonic dose-response curves (NMDRCs) and disrupt hormone functions at low doses27,28. These NMDRCs occur when a chemical’s effect does not increase proportionally with higher doses, and instead, shows varying responses at different concentrations. Such responses may involve mechanisms like complex feedback loops or indirect effects, which are not seen with traditional monotonic dose-response models29. Importantly, epigenetic changes at low concentrations have been documented in various studies, supporting the hypothesis that indirect mechanisms, such as those influencing hormonal and cellular pathways, can trigger these alterations without a clear linear pattern30.
Intriguingly, exposure to lower concentrations of lead decreased IGF2 expression in a non-monotonic dose-response manner in the human embryonic kidney cell line (HEK-293), but did not affect H19 expression31. However, our study is the first to investigate these epigenotoxic effects in the context of combined metal exposures, which more closely mimics real-world scenarios. We discovered a significant downregulation of H19 lncRNA expression following exposure to specific combinations of toxic metals. Conversely, the IGF2 mRNA expression was upregulated after co-exposure to 10 µM oAs + 1 µM of CdCl2 and 1 µM of CdCl2 + 10 nM of MeHgCl, with p-values less than 0.001 in both cases (Fig. 2). These findings indicate that when toxic metals are combined, their influence on gene expression can be either antagonistic or synergistic, depending on the specific doses of each metal, highlighting the complex nature of mixture toxicity.
Our results suggest that toxic metal exposure may disrupt the regulation of the H19/IGF2 expression, which could have significant implications for liver health. Increased IGF2 expression has been observed in patients with liver inflammation32,33, a precursor to more severe liver diseases such as NAFLD and non-alcoholic steatohepatitis (NASH)34,35. While the role of the H19 lncRNA in these conditions is still under investigation36, our findings provide new insights into how environmental exposures might influence its expression. The H19 transcript might function in various capacities by affecting miRNAs, RNA-binding proteins, and signaling pathways, as indicated by studies in cultured hepatocyte cell lines, animal models, and human samples25,37,38.
H19 and IGF2 are the most widely studied mammalian elements of a cluster of imprinted genes; the H19 gene is transcribed from the maternal allele, while the IGF2 gene is expressed from the paternal allele39. The parent-of-origin-specific expression pattern of the H19/IGF2 locus is controlled by the allele-specific physical interactions between the DMRs40. On the maternal allele, the unmethylated H19 DMR, which is bound by CCCTC-binding factor (CTCF), and the IGF2 DMR1 interact, resulting in an active chromatin domain with H19 in close vicinity to its enhancers and IGF2 in an inactive domain away from the enhancers. On the paternal allele, the methylated H19 DMR associates with the methylated IGF2 DMR2, moving IGF2 into the active chromatin domain41,42. The H19/IGF2 domain is imprinted in the fetal liver, where only one parental allele is expressed, but it is regulated by the P1 promoter in the adult human liver, resulting in biallelic expression43. We provided the evidence that the altered expression of H19 and IGF2 by combined toxic metal exposures was mainly due to CpG hypermethylation at the H19 DMR (Fig. 3a). In contrast, no significant changes in the CpG methylation level were observed in the examined ICR (Fig. 3b). Our findings may contradict the previous studies demonstrating the ICR plays a pivotal role in directing the allelic expression of the H19/IGF2 domain. Therefore, it is necessary to perform regional methylation analysis for the remaining CpG loci in the ICR to trace the any possible epigenetic events linked to mixture exposure to toxic metals.
Followed by the quantitative analysis of gene expression and DNA methylation, we performed a correlation analysis between the altered H19/IGF2 expression levels and the altered CpG methylation levels, to determine the core CpG units of the regulatory elements responsible for significant epigenetic dysregulation at the H19/IGF2 domain. It was revealed that the CpG loci within the H19 DMR were more highly correlated with metal-mixtures-mediated H19/IGF2 modulation than the CpGs within the examined ICR (Fig. 4 and S1). Interestingly, the correlation between the CpG methylation levels and the H19/IGF2 expression varied with different metal combinations, indicating metal-specific outcomes. These findings suggest that a variety of adverse environmental exposures including toxic metals may exert epigenetic effects on the gene modulation in a chemical-specific manner. In mammalian tissues, the parental imprints at the H19/IGF2 locus are maintained through the methylation status of the ICR, which dominates the reciprocally exclusive expression of these imprinted genes44. Our results, however, demonstrated that the methylation status at the H19 DMR may play a crucial role in regulating the mutually exclusive pattern of H19/IGF2 expression in the human hepatocytes in response to toxic metal exposures. Furthermore, our findings suggested that exposure to a mixture of toxic metals might influence on the methylation levels of the site-specific CpGs in the H19 DMR and ICR, along with the modulation of H19 and IGF2 expression.
In our transcriptome analysis, we aimed to explore the impact of combined toxic metal exposures on global gene expression profiles in hepatocytes and to assess whether these transcriptional changes are associated with epigenetic modifications at the H19/IGF2 locus, which could serve as potential biomarkers of metal exposure. Upon exposure to 10 µM of oAs + 1 µM of CdCl2 and 1 µM of CdCl2 + 10 nM of MeHgCl, we identified a total of 180 and 177 differentially expressed genes (DEGs), respectively. Among them, 51 DEGs were found to be common to both conditions (Fig. S2a). Our study revealed that the enriched pathways with an FDR q-value below 0.01, particularly the upregulation of TNF-α signaling via the NF-κB pathway (Fig. 5), were similar to those observed in studies of various stages of liver disease progression. Previous research has demonstrated that toxic metal exposure, such as cadmium and methylmercury, enhances NF-κB activity, leading to increased production of pro-inflammatory cytokines such as TNF-α, which play a critical role in liver inflammation and hepatic steatosis45,46. Additionally, the observed downregulation of MYC targets V1, Mitotic Spindle, and E2F Targets observed in our study corroborates findings from previous research that highlight the disruption of cell cycle regulation and consequently, anti-proliferation47,48. This downregulation has been associated with impaired hepatocyte proliferation, which can contribute to the progression of liver diseases such as NAFLD and NASH, where cellular proliferation has been found to be reduced. We have also focused on the evidence of mitochondrial-epigenetic crosstalk in patients with imprinting disorders characterized by overexpression of IGF2 that could potentially suppress oxidative phosphorylation (OXPHOS)49. Distinct enrichment of OXPHOS pathways were identified between the two combined exposures: OXPHOS was upregulated in cells exposed to the oAs + CdCl2 mixture, but this pathway was downregulated in cells exposed to the CdCl2 + MeHgCl mixture. The contrasting impacts on OXPHOS may imply the mutually exclusive patterns of mitochondrial dysfunction, attributed to the specific metal combinations. Given that toxic metals such as arsenic, cadmium, and mercury can damage mitochondria in various ways50, these findings highlight the intricate nature of their combined impact on mitochondrial pathways. Although OXPHOS metabolic pathway has been documented to be universally downregulated in human cancer51,52, metabolic alterations including upregulated OXPHOS have been widely accepted as a hallmark of cancer cells53,54. This bioenergetic propensity is also frequently associated with development of HCC. Further in-depth studies are needed to elucidate the underlying mechanisms of this differential OXPHOS regulation and its implications for hepatocarcinogenesis. By establishing a link between these modified pathways and the observed alterations in methylation patterns at the H19/IGF2 locus, our study provides further evidence that epigenetic modifications resulting from toxic metal exposure play a pivotal role in the pathogenesis of hepatic disease. The disruption of these pathways underscores the complex role of epigenetic events in liver function and disease progression, highlighting the necessity for further studies to elucidate the underlying mechanisms. Moreover, future studies should employ multi-omics approaches and functional assays to validate H19/IGF2 dysregulation and its role in altered pathways. Correlation analyses between H19/IGF2 expression and other DEGs will further clarify the broader impact of metal exposure on cellular processes.
While our study provides valuable insights into the epigenetic effects of metal mixtures, we acknowledge certain limitations. Firstly, the in vitro nature of the research, including the use of only one cell line, may limit the generalizability of our results. Although the THLE-3 cell lines are useful for understanding specific mechanisms in the liver pathogenesis, they do not fully recapitulate the complex pathophysiological responses indicative of diverse human health outcomes. Also, the controlled conditions of in vitro studies may not accurately represent the intricate interactions that occur in vivo or across different populations. Future studies should consider using primary human hepatocytes or 3D liver organoids to further validate our findings. Additionally, while our research focused on three specific metals (organic arsenic, cadmium chloride, and methylmercury chloride) and their binary combinations, this study is part of a larger research plan. We initiated our research with these metals due to their significant health impacts, prevalence in environmental exposure, and the existing data gaps regarding their combined effects. Future studies will include other heavy metals and more complex metal mixtures to provide a more comprehensive understanding of various metal interactions. Lastly, investigating the long-term consequences of these epigenetic changes and their potential reversibility could provide important information for therapeutic interventions. These aspects highlight the need for caution in extrapolating our results to broader environmental and health contexts.
Despite these limitations, our study represents significant methodological advances and insights. We utilized physiologically based pharmacokinetic (PBPK) models to estimate concentrations corresponding to toxicity benchmarks and applied these estimates to liver cell lines, thereby reducing experimental error (Table S1). This approach enhances the relevance and accuracy of our findings, particularly in understanding the cellular impacts of toxic metal exposures at realistic doses. Our investigation of the effects of predicted exposure combinations and their link to epigenetic modulation offers a novel perspective on the mechanisms underlying exposure-induced health outcomes. In conclusion, the hypermethylated CpG loci within the H19 DMR as potential biomarkers for metal mixtures can open new avenues for early detection of liver damage due to environmental exposures. Furthermore, these epigenotoxic response-sensing candidates may contribute to the development of novel preventive and/or therapeutic strategies for metal-induced adverse outcomes. These advancements also have potential for future use in exposure monitoring and health impact assessment, contributing to the field of environmental health and safety.
Materials and methods
Cell culture and reagents
THLE-3, a cell line derived from normal adult hepatic epithelial cells, was obtained from the American Type Culture Collection (ATCC, Manassas, VA, USA). THLE-3 cells were cultured in BEGM Bronchial Epithelial Cell Growth Basal Medium (Lonza, Walkersville, MD, USA) supplemented with 10% FBS (ATCC, Manassas, VA, USA), 1% antibiotic antimycotic solution (WELGENE Inc., Daegu, South Korea), 5 ng/mL human recombinant EGF (Corning Inc., Corning, NY, USA), 70 ng/mL phosphoethanolamine (Sigma Aldrich, St Louis, MO, USA), and the additives from the BEGM Bullet Kit (without gentamycin/amphotericin (GA) and epinephrine). Culture plates were maintained at 37 °C in humidified 5% CO2 atmosphere, and the passage number for exposure assessment was four to five.
Monomethylarsonous acid (MMA III, CAS No. 25400-23-1) and dimethylarsinous acid (DMA III, CAS No. 55094-22-9) were procured from Chem Service Inc. (West Chester, PA, USA) and Sigma Aldrich (St Louis, MO, USA), respectively. Cadmium chloride (CdCl2, CAS No. 10108-64-2) and methylmercury (II) chloride (MeHgCl, CAS No. 115-09-3) were sourced from Sigma Aldrich (St Louis, MO, USA) and Alfa Aesar (Ward Hill, MA, USA), respectively. The cells were seeded at a density of approximately 4–6 × 104 cells/cm2 in 90-mm plates one day prior to metal exposure. Subsequent treatments involved either individual or combined exposure to oAs, CdCl2, and MeHgCl, or a control condition using sterile distilled water for 72 h. This exposure duration was chosen based on preliminary studies showing stable epigenetic changes while minimizing confounding effects from prolonged cell culture55.
In vitro epigenotoxicity assay of toxic metals based on physiologically based pharmacokinetic (PBPK) models
In this study, we selected organic forms of arsenic, cadmium chloride, and methylmercury as the research chemicals for several reasons. Firstly, these metals are commonly found in combination in contaminated environments and have been consistently detected in general populations, including those in Korea56,57. Secondly, recent reports indicate that the general Korean population is predominantly exposed to oAs variants58, making it a relevant choice for our study. To simulate the in vivo metabolized form of inorganic arsenic, we used trivalent organic arsenic species (MMA III and DMA III), which have been reported to exhibit higher toxicity compared to inorganic arsenic59,60. These two metabolites were mixed in a 1:9 ratio based on their proportions observed in human studies to more accurately reflect real-world exposure scenarios61. The decision to focus on these three metals was based on their known high toxicity, environmental prevalence, and their potential to act as potent epigenetic disruptors. While the effects of lead exposure on H19/IGF2 methylation have been previously reported, the combined impact of arsenic, cadmium, and mercury on this locus remains unexplored. Therefore, this study serves as a preliminary investigation, forming the basis for future research that will include additional heavy metals and more complex mixture exposures to comprehensively evaluate the epigenetic and toxicological effects of various metal combinations.
To determine environmentally relevant exposure concentrations, we utilized PBPK models. Reference dose (RfD) values set by the US Environmental Protection Agency (EPA) were converted into biomonitoring equivalents (BEs) in the liver using PBPK models61,62,63. This conversion was instrumental in determining the specific concentrations of toxic metals for cell exposure (refer to Table S1 for details). We applied the transformed BEs at dilutions of 1/10 and 1/100 to gauge low dose effects, and at 10 and 100 times the BEs to assess substance toxicity at higher concentrations to capture a range of potential effects. For combined exposures, we focused on binary mixtures to systematically examine potential interactions while maintaining experimental feasibility.
Reverse transcription-quantitative PCR analysis
Total RNA was isolated using the RNeasy Mini Kit (QIAGEN, Hilden, Germany) according to the manufacturer’s protocol. cDNA synthesis was carried out on 1 µg of total RNA using the High-Capacity cDNA Reverse Transcription Kit (Applied Biosystems, Foster City, CA, USA) for reverse transcription-quantitative PCR (RT-qPCR) following the manufacturer’s instructions. The QuantStudio™ 6 Flex Real-Time PCR system (Applied Biosystems, Foster City, CA, USA) was used to amplify and quantify the cDNA samples in MicroAmp Fast 96well Rx Plate (Applied Biosystems, Foster City, CA, USA). The QuantStudio™ Real-Time PCR Software v1.3 (Applied Biosystems, Foster City, CA, USA) was used to perform thermal cycling using the following standard conditions: 95 °C for 10 min, followed by 40 cycles of 95 °C for 15 s and 60 °C for 1 min. All experiments were performed in triplicate. ACTB and GAPDH were utilized as endogenous controls for amplification, and their mean Ct values were employed for normalization. The 2−ΔΔCt method was then used to calculate fold changes relative to the unexposed control. The oligonucleotide sequences of the qPCR primers are listed in the Table S2.
Bisulfite amplicon sequencing (BSAS) analysis
The gDNA was extracted using the QIAamp DNA Mini Kit (QIAGEN, Hilden, Germany) following the manufacturer’s protocol. Bisulfite modification of the 2 µg of gDNA was performed using the EZ DNA methylation-lightning kit (Zymo Research, Irvine, CA, USA). The bisulfite-converted DNA was amplified to a 387-bp sequence element harboring 21 CpG sites located at the H19 DMR and a 348-bp element involving 17 CpG sites at the ICR using TaKaRa EX Taq DNA polymerase (Takara Bio Inc., Japan). The library was prepared according to Illumina’s recommendations and sequenced using the 300-cycle MiSeq reagent kit V3 (paired end, 2 × 300 bp) on an Illumina MiSeq instrument (Illumina Inc., San Diego, CA, USA) for BSAS analysis. Instrument preparation and setup were performed following the manufacturer’s instructions. To eliminate reads with high sequencing errors, the data for each sample were pre-processed using FastQC (version 0.11.5). Potentially present sequencing adapters and low-quality bases present in the low reads were trimmed using Skewer (version 0.2.2)64. After removing the low-quality bases and sequencing adapters, the cleaned high-quality reads were mapped to the reference genome using BS-seeker2 software (version 2.1.2)65. A 10% mis-mapping rate was allowed, as the alignment is performed with reference to certain specified genic regions of the genome sequence. BS-seeker2 was used to determine the methylation levels from the mapping result. The methylation percentages were calculated at single base resolution. The oligonucleotide sequences of the BSAS PCR primers used are listed in the Table S3.
RNA sequencing (RNA-seq)
Transcriptome analysis was conducted to elucidate global gene expression changes and to identify key biological pathways modulated under varying toxic metal exposure conditions. For all samples, the RNA integrity number (RIN) was ≥ 7, as determined using an Agilent 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA). Paired-end sequencing (2 × 101 bp) was performed on the NovaSeq6000 platform (Illumina Inc., San Diego, CA, USA) in accordance with the manufacturer’s instructions. For this, cDNA libraries were constructed from 1 µg of total RNA using the TruSeq Stranded mRNA Library Prep Kit (Illumina Inc., San Diego, CA, USA). On average, each library produced 30–33 million read pairs. The quality of the read data was assessed using FastQC. Subsequently, raw reads were processed and trimmed with Trim Galore (version 0.6.6) to eliminate potential adapter sequences and low-quality reads. Reads were then aligned to the human genome version GRCh38 using the STAR aligner (version 2.7.9a)66. Gene expression levels were normalized by calculating the number of fragments per kilobase of transcript per million mapped reads (FPKM) with StringTie (version 2.1.7b)67. To identify DEGs, we utilized the edgeR package (v3.38.4) on Bioconductor (v3.14)68. DEGs were defined as genes with absolute log2 fold change (LFC) values ≥ 1 and FDR-corrected p-values ≤ 0.05.
Pathway enrichment analysis was conducted as described previously69. To gain deeper insight into the gene expression changes resulting from toxic metal exposure, ranked gene list analyses were performed using the gene set enrichment analysis (GSEA) software (version 4.3.2) with the following parameters: ranked gene list = (-1) * log(p-value) * log2FC, gene sets database = Hallmark gene sets (h.all.v2024.1.Hs.symbols.gmt), number of permutations = 1000, permutation type = gene_set, metric for ranking genes = Signal2Noise, and normalization mode = meandiv70.
Statistical analysis
All data were presented as mean ± standard error of the mean. Differences in sample group means were analyzed using one-way analysis of variance (ANOVA) followed by Tukey’s post hoc test, using the unexposed control as the reference group for all comparisons. The p-value < 0.05 was considered statistically significant. We calculated the pairwise correlation of gene expression with methylation level of various methylation sites using Spearman’s test. RStudio (version 4.1.3) was used for statistical and graphical analyses.
Data availability
All relevant data are available within the paper. The datasets generated and/or analyzed during this study are available in the Gene Expression Omnibus (GEO) repository under the accession numbers GSE280025 (BSAS) and GSE280026 (RNA-seq).
References
Maqsood, Q., Hussain, N., Mumtaz, M., Bilal, M. & Iqbal, H. M. N. Novel strategies and advancement in reducing heavy metals from the contaminated environment. Arch. Microbiol. 204, 478. https://doi.org/10.1007/s00203-022-03087-2 (2022).
Clemens, S. & Ma, J. F. Toxic heavy metal and metalloid accumulation in crop plants and foods. Annu. Rev. Plant. Biol. 67, 489–512. https://doi.org/10.1146/annurev-arplant-043015-112301 (2016).
Witkowska, D., Slowik, J. & Chilicka, K. Heavy Metals and Human Health: Possible exposure pathways and the competition for protein binding sites. Molecules 26 https://doi.org/10.3390/molecules26196060 (2021).
Sanders, A. P. et al. Combined exposure to lead, cadmium, mercury, and arsenic and kidney health in adolescents age 12–19 in NHANES 2009–2014. Environ. Int. 131, 104993. https://doi.org/10.1016/j.envint.2019.104993 (2019).
Silins, I. & Hogberg, J. Combined toxic exposures and human health: Biomarkers of exposure and effect. Int. J. Environ. Res. Public. Health 8, 629–647. https://doi.org/10.3390/ijerph8030629 (2011).
Singh, N., Gupta, V. K., Kumar, A. & Sharma, B. Synergistic effects of heavy metals and pesticides in living systems. Front. Chem. 5, 70. https://doi.org/10.3389/fchem.2017.00070 (2017).
Kim, D. W., Ock, J., Moon, K. W. & Park, C. H. Association between heavy metal exposure and dyslipidemia among Korean adults: From the Korean National Environmental Health Survey, 2015–2017. Int. J. Environ. Res. Public Health 19, 3181. https://doi.org/10.3390/ijerph19063181 (2022).
Huang, Q. et al. Association between manganese exposure in heavy metals mixtures and the prevalence of Sarcopenia in US adults from NHANES 2011–2018. J. Hazard. Mater. 464, 133005. https://doi.org/10.1016/j.jhazmat.2023.133005 (2024).
Kefayati, F., Babaahmadi, K., Mousavi, A., Hodjat, T., Abdollahi, M. & M. & Epigenotoxicity: A danger to the future life. J. Environ. Sci. Health Tox Hazard. Subst. Environ. Eng. 58, 382–411. https://doi.org/10.1080/10934529.2023.2190713 (2023).
Hu, J. & Yu, Y. Epigenetic response profiles into environmental epigenotoxicant screening and health risk assessment: A critical review. Chemosphere 226, 259–272. https://doi.org/10.1016/j.chemosphere.2019.03.096 (2019).
Mokdad, A. A. et al. Liver cirrhosis mortality in 187 countries between 1980 and 2010: A systematic analysis. BMC Med. 12, 145. https://doi.org/10.1186/s12916-014-0145-y (2014).
Koyama, Y. & Brenner, D. A. Liver inflammation and fibrosis. J. Clin. Invest. 127, 55–64. https://doi.org/10.1172/JCI88881 (2017).
Wang, X. et al. Systemic inflammation mediates the association of heavy metal exposures with liver injury: A study in general Chinese urban adults. J. Hazard. Mater. 419, 126497. https://doi.org/10.1016/j.jhazmat.2021.126497 (2021).
Renu, K. et al. Molecular mechanism of heavy metals (lead, Chromium, Arsenic, Mercury, Nickel and Cadmium) - induced hepatotoxicity—A review. Chemosphere 271, 129735. https://doi.org/10.1016/j.chemosphere.2021.129735 (2021).
Seenivasan, S., Manikandan, N., Muraleedharan, N. N. & Selvasundaram, R. Heavy metal content of black teas from south India. Food Control 19, 746–749. https://doi.org/10.1016/j.foodcont.2007.07.012 (2008).
Vinodhini, R. & Narayanan, M. Bioaccumulation of heavy metals in organs of fresh water fish Cyprinus Earpio (Common carp). Int. J. Environ. Sci. Te 5, 179–182 (2008). doi:Doi 10.1007/Bf03326011.
Shim, M. J., Yang, Y. M., Oh, D. Y., Lee, S. H. & Yoon, Y. Y. Spatial distribution of heavy metal accumulation in the sediments after dam construction. Environ. Monit. Assess. 187, 733. https://doi.org/10.1007/s10661-015-4967-7 (2015).
Nyanza, E. C. et al. Effects of prenatal exposure and co-exposure to metallic or metalloid elements on early infant neurodevelopmental outcomes in areas with small-scale gold mining activities in Northern Tanzania. Environ. Int. 149, 106104. https://doi.org/10.1016/j.envint.2020.106104 (2021).
Chen, H. et al. Chronic inorganic arsenic exposure induces hepatic global and individual gene hypomethylation: Implications for arsenic hepatocarcinogenesis. Carcinogenesis 25, 1779–1786. https://doi.org/10.1093/carcin/bgh161 (2004).
Sun, Y. et al. C-myc promotes miR-92a-2-5p transcription in rat ovarian granulosa cells after cadmium exposure. Toxicol. Appl. Pharmacol. 421, 115536. https://doi.org/10.1016/j.taap.2021.115536 (2021).
Cartularo, L. et al. Gene expression and pathway analysis of human hepatocellular carcinoma cells treated with cadmium. Toxicol. Appl. Pharmacol. 288, 399–408. https://doi.org/10.1016/j.taap.2015.08.011 (2015).
Hernandez, A. F. & Tsatsakis, A. M. Human exposure to chemical mixtures: Challenges for the integration of toxicology with epidemiology data in risk assessment. Food Chem. Toxicol. 103, 188–193. https://doi.org/10.1016/j.fct.2017.03.012 (2017).
Han, L., Lee, D. H. & Szabo, P. E. CTCF is the master organizer of domain-wide allele-specific chromatin at the H19/Igf2 imprinted region. Mol. Cell. Biol. 28, 1124–1135. https://doi.org/10.1128/MCB.01361-07 (2008).
Vernucci, M. et al. The H19 endodermal enhancer is required for Igf2 activation and tumor formation in experimental liver carcinogenesis. Oncogene 19, 6376–6385. https://doi.org/10.1038/sj.onc.1204024 (2000).
Zhang, J. et al. A transforming growth factor-beta and H19 Signaling Axis in Tumor-Initiating hepatocytes that regulates hepatic carcinogenesis. Hepatology 69, 1549–1563. https://doi.org/10.1002/hep.30153 (2019).
Wu, H., Eckhardt, C. M. & Baccarelli, A. A. Molecular mechanisms of environmental exposures and human disease. Nat. Rev. Genet. 24, 332–344. https://doi.org/10.1038/s41576-022-00569-3 (2023).
Schug, T. T., Janesick, A., Blumberg, B. & Heindel, J. J. Endocrine disrupting chemicals and disease susceptibility. J. Steroid Biochem. Mol. Biol. 127, 204–215. https://doi.org/10.1016/j.jsbmb.2011.08.007 (2011).
Myers, J. P., Zoeller, R. T. & vom Saal, F. A clash of old and new scientific concepts in toxicity, with important implications for public health. Environ. Health Perspect. 117, 1652–1655. https://doi.org/10.1289/ehp.0900887 (2009).
Lagarde, F. et al. Non-monotonic dose-response relationships and endocrine disruptors: A qualitative method of assessment. Environ. Health 14, 13. https://doi.org/10.1186/1476-069X-14-13 (2015).
Zoeller, R. T. & Vandenberg, L. N. assessing dose–response relationships for endocrine disrupting chemicals (EDCs): A focus on non-monotonicity. Environ. Health 14, 42. https://doi.org/10.1186/s12940-015-0029-4 (2015).
Nye, M. D., Hoyo, C. & Murphy, S. K. In vitro lead exposure changes DNA methylation and expression of IGF2 and PEG1/MEST. Toxicol. Vitro 29, 544–550. https://doi.org/10.1016/j.tiv.2015.01.002 (2015).
Adamek, A. & Kasprzak, A. Insulin-like growth factor (IGF) system in Liver diseases. Int. J. Mol. Sci. 19 https://doi.org/10.3390/ijms19051308 (2018).
Belfiore, A. et al. IGF2: A role in Metastasis and Tumor Evasion from Immune Surveillance? Biomedicines 11 https://doi.org/10.3390/biomedicines11010229 (2023).
Pope, C., Mishra, S., Russell, J., Zhou, Q. & Zhong, X. B. Targeting H19, an imprinted long non-coding RNA, in hepatic functions and Liver diseases. Diseases 5 https://doi.org/10.3390/diseases5010011 (2017).
Schwartz, B. E. et al. Discovery and Targeting of the signaling controls of PNPLA3 to effectively reduce transcription, expression, and function in pre-clinical NAFLD/NASH settings. Cells 9 https://doi.org/10.3390/cells9102247 (2020).
Tietze, L. & Kessler, S. M. The Good, the bad, the Question-H19 in Hepatocellular Carcinoma. Cancers (Basel) 12 https://doi.org/10.3390/cancers12051261 (2020).
Sun, Z. et al. Aberrant NSUN2-mediated m(5)C modification of H19 lncRNA is associated with poor differentiation of hepatocellular carcinoma. Oncogene 39, 6906–6919. https://doi.org/10.1038/s41388-020-01475-w (2020).
Ye, Y. et al. Macrophages-induced long noncoding RNA H19 up-regulation triggers and activates the miR-193b/MAPK1 axis and promotes cell aggressiveness in hepatocellular carcinoma. Cancer Lett. 469, 310–322. https://doi.org/10.1016/j.canlet.2019.11.001 (2020).
Ferguson-Smith, A. C., Sasaki, H., Cattanach, B. M. & Surani, M. A. Parental-origin-specific epigenetic modification of the mouse H19 gene. Nature 362, 751–755. https://doi.org/10.1038/362751a0 (1993).
Murrell, A., Heeson, S. & Reik, W. Interaction between differentially methylated regions partitions the imprinted genes Igf2 and H19 into parent-specific chromatin loops. Nat. Genet. 36, 889–893. https://doi.org/10.1038/ng1402 (2004).
Reik, W. et al. Chromosome loops, insulators, and histone methylation: New insights into regulation of imprinting in clusters. Cold Spring Harb Symp. Quant. Biol. 69, 29–37. https://doi.org/10.1101/sqb.2004.69.29 (2004).
Lopes, S. et al. Epigenetic modifications in an imprinting cluster are controlled by a hierarchy of DMRs suggesting long-range chromatin interactions. Hum. Mol. Genet. 12, 295–305. https://doi.org/10.1093/hmg/ddg022 (2003).
Netchine, I. et al. 11p15 imprinting center region 1 loss of methylation is a common and specific cause of typical Russell-Silver syndrome: Clinical scoring system and epigenetic-phenotypic correlations. J. Clin. Endocrinol. Metab. 92, 3148–3154. https://doi.org/10.1210/jc.2007-0354 (2007).
Banerjee, S., Smallwood, A., Lamond, S., Campbell, S. & Nargund, G. Igf2/H19 imprinting control region (ICR): An insulator or a position-dependent silencer? ScientificWorldJournal 1, 218–224. https://doi.org/10.1100/tsw.2001.50 (2001).
Qu, F. & Zheng, W. Cadmium exposure: Mechanisms and pathways of toxicity and Implications for Human Health. Toxics 12, 388. https://doi.org/10.3390/toxics12060388 (2024).
Alshehri, A. S. et al. Kaempferol prevents cadmium chloride-induced liver damage by upregulating Nrf2 and suppressing NF-κB and keap1. Environ. Sci. Pollut R 29, 13917–13929. https://doi.org/10.1007/s11356-021-16711-3 (2022).
Ren, B. et al. E2F integrates cell cycle progression with DNA repair, replication, and G(2)/M checkpoints. Genes Dev. 16, 245–256. https://doi.org/10.1101/gad.949802 (2002).
Daniel, C. J. et al. T-cell dysfunction upon expression of MYC with altered phosphorylation at Threonine 58 and serine 62. Mol. Cancer Res. 20, 1151–1165. https://doi.org/10.1158/1541-7786.MCR-21-0560 (2022).
Wallace, D. C. & Fan, W. Energetics, epigenetics, mitochondrial genetics. Mitochondrion 10, 12–31. https://doi.org/10.1016/j.mito.2009.09.006 (2010).
Reddam, A., McLarnan, S. & Kupsco, A. Environmental Chemical exposures and mitochondrial dysfunction: A review of recent literature. Curr. Environ. Health Rep. 9, 631–649. https://doi.org/10.1007/s40572-022-00371-7 (2022).
Tian, H., Zhu, X., Lv, Y., Jiao, Y. & Wang, G. Glucometabolic reprogramming in the Hepatocellular Carcinoma Microenvironment: Cause and Effect. Cancer Manag Res. 12, 5957–5974. https://doi.org/10.2147/cmar.S258196 (2020).
Wang, H. et al. Inhibition of hepatocellular carcinoma by metabolic normalization. PLoS One 14, e0218186. https://doi.org/10.1371/journal.pone.0218186 (2019).
Hanahan, D. & Weinberg, R. A. Hallmarks of Cancer: The Next Generation. Cell 144, 646–674. https://doi.org/10.1016/j.cell.2011.02.013 (2011).
Rohatgi, N., Ghoshdastider, U., Baruah, P., Kulshrestha, T. & Skanderup, A. J. A pan-cancer metabolic atlas of the tumor microenvironment. Cell. Rep. 39, 110800. https://doi.org/10.1016/j.celrep.2022.110800 (2022).
Tsai, H. C. et al. Transient low doses of DNA-demethylating agents exert durable antitumor effects on hematological and epithelial tumor cells. Cancer Cell. 21, 430–446. https://doi.org/10.1016/j.ccr.2011.12.029 (2012).
Kim, J. H. et al. Lead and mercury levels in repeatedly collected urine samples of young children: A longitudinal biomonitoring study. Environ. Res. 189, 109901. https://doi.org/10.1016/j.envres.2020.109901 (2020).
Kim, Y. & Lee, B. K. Associations of blood lead, cadmium, and mercury with estimated glomerular filtration rate in the Korean general population: Analysis of 2008–2010 Korean National Health and Nutrition Examination Survey data. Environ. Res. 118, 124–129. https://doi.org/10.1016/j.envres.2012.06.003 (2012).
Choi, J. W. et al. Concentrations of blood and urinary arsenic species and their characteristics in general Korean population. Environ. Res. 214, 113846. https://doi.org/10.1016/j.envres.2022.113846 (2022).
Kobayashi, Y. & Agusa, T. in Arsenic Contamination in Asia: Biological Effects and Preventive Measures (eds Hiroshi Yamauchi & Guifan Sun) 13–28Springer Singapore, (2019).
Raessler, M. The Arsenic Contamination of drinking and groundwaters in Bangladesh: Featuring Biogeochemical aspects and implications on Public Health. Arch. Environ. Contam. Toxicol. 75, 1–7. https://doi.org/10.1007/s00244-018-0511-4 (2018).
El-Masri, H. A. & Kenyon, E. M. Development of a human physiologically based pharmacokinetic (PBPK) model for inorganic arsenic and its mono- and di-methylated metabolites. J. Pharmacokinet. Pharmacodyn 35, 31–68. https://doi.org/10.1007/s10928-007-9075-z (2008).
Carrier, G., Bouchard, M., Brunet, R. C. & Caza, M. A toxicokinetic model for predicting the tissue distribution and elimination of organic and inorganic mercury following exposure to methyl mercury in animals and humans. II. Application and validation of the model in humans. Toxicol. Appl. Pharmacol. 171, 50–60. https://doi.org/10.1006/taap.2000.9113 (2001).
Kjellstrom, T. & Nordberg, G. F. A kinetic model of cadmium metabolism in the human being. Environ. Res. 16, 248–269. https://doi.org/10.1016/0013-9351(78)90160-3 (1978).
Jiang, H., Lei, R., Ding, S. W. & Zhu, S. Skewer: A fast and accurate adapter trimmer for next-generation sequencing paired-end reads. BMC Bioinform. 15, 182. https://doi.org/10.1186/1471-2105-15-182 (2014).
Guo, W. et al. BS-Seeker2: A versatile aligning pipeline for bisulfite sequencing data. BMC Genom. 14, 774. https://doi.org/10.1186/1471-2164-14-774 (2013).
Dobin, A. et al. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21. https://doi.org/10.1093/bioinformatics/bts635 (2013).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295. https://doi.org/10.1038/nbt.3122 (2015).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140. https://doi.org/10.1093/bioinformatics/btp616 (2010).
Reimand, J. et al. Pathway enrichment analysis and visualization of omics data using g:Profiler, GSEA, Cytoscape and EnrichmentMap. Nat. Protoc. 14, 482–517. https://doi.org/10.1038/s41596-018-0103-9 (2019).
Subramanian, A. et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. U S A 102, 15545–15550. https://doi.org/10.1073/pnas.0506580102 (2005).
Acknowledgements
This study was supported by the Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education [2020R1A2C2006763] and National Research Foundation of Korea (BK21 PLUS).
Author information
Authors and Affiliations
Contributions
Yehoon Jo: Writing - Original draft, Software, Investigation, Visualization; Eugene Lim: Methodology, Data curation, Formal analysis; Jihye Park: Formal analysis, Visualization, Software; Keunsoo Kang: Formal analysis, Visualization, Supervision; Mi-Yeon Shin: Investigation, Software; Jeong Weon Choi: Investigation, Software; Sungkyoon Kim: Conceptualization, Resources, Writing – Review & Editing, Investigation, Funding acquisition; Jaehyouk Lee: Conceptualization, Methodology, Writing – Review & Editing, Investigation, Data Curation, Supervision.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Jo, Y., Lim, E., Park, J. et al. Epigenetic dysregulation of H19/IGF2 in hepatic cells exposed to toxic metal mixtures in vitro. Sci Rep 14, 29413 (2024). https://doi.org/10.1038/s41598-024-80142-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-80142-6