Abstract
Micropapillary adenocarcinoma (MPC) is an aggressive histological subtype of lung adenocarcinoma (LUAD). MPC is composed of small clusters of cancer cells exhibiting inverted polarity. However, the mechanism underlying its formation is poorly understood. Here we show that hypoxia is involved in MPC formation. Hypoxia induced the formation of MPC-like structures (MLSs) in a three-dimensional culture system using A549 human LUAD cells, and HIF-1α was indispensable for MLS formation. RNA sequencing analysis demonstrated that A549 cells forming MLSs exhibited a gene expression signature similar to that of lung MPC. Moreover, MLS formation enhanced the resistance of A549 cells to natural killer cell cytotoxicity. Our findings suggest that hypoxia drives lung MPC formation through HIF-1α and that immune escape from natural killer cells might underlie the aggressiveness of MPC.
Similar content being viewed by others
Introduction
Lung cancer is the leading cause of cancer-related death worldwide1, and lung adenocarcinoma (LUAD) is the most prevalent histological type of lung cancer2. Most LUADs are histologically heterogeneous and display a mixed growth pattern. In 2015, the World Health Organization proposed definitions of five LUAD subtypes (lepidic, acinar, papillary, micropapillary, and solid) to address this histological heterogeneity2. Among them, micropapillary adenocarcinoma (MPC) has been considered a high-grade subtype with a poor prognosis because of frequent lymphatic invasion and lymph node metastases2,3,4,5,6. Even a small component of MPC within a tumor can accelerate disease progression and recurrence6. MPC is also recognized as an aggressive variant in organs other than the lungs (including the breast, bladder, salivary glands, and colon)3,7. MPC is characterized by small, tightly cohesive clusters of tumor cells freely dispersed within alveolar spaces or clear empty spaces in the invasive portion of the lesion3,4,5,6. These clusters of tumor cells in MPC exhibit an inverted-polarity configuration in which their apical surfaces face the stroma8,9,10. This is in striking contrast to the non-MPC pattern, in which the apical membrane lines the inside of the lumen on the opposite side of the stroma. Although the higher metastatic potential of MPC can be attributed to its characteristic structure, the molecular mechanisms underlying the formation of lung MPC are almost completely unknown because of the lack of in vitro models.
An intratumoral hypoxic environment is a critical hallmark of cancer and is widely associated with aggressive cancer progression, therapeutic resistance, and a poor prognosis11,12,13. In patients with stage I LUAD, hypoxia is a dominant risk factor for poor overall survival14. A central component of hypoxic adaptation is hypoxia-inducible factor (HIF)-1, which exists as a heterodimer composed of HIF-1α and HIF-1β subunits and activates the expression of numerous genes involved in cell survival, invasion/metastasis, angiogenesis, glycolysis, and immune escape15,16,17. HIF-1α is rapidly degraded in cells under normoxic conditions and remains stable under hypoxic conditions15. Thus, it is an oxygen sensor and the critical factor mediating the adaptive response to hypoxia.
In this study, we developed a three-dimensional (3D) culture model with A549 cells. This model demonstrated that hypoxia exposure induced the formation of MPC-like structures (MLSs), which exhibit inverted polarity and have a gene expression signature similar to that of lung MPC. We found that HIF-1α and RhoA/Rho-kinase signaling pathways were involved in the hypoxia-induced formation of MLSs. Furthermore, MLS formation enhanced the resistance of A549 cells to natural killer (NK) cell cytotoxicity. Our results suggest that HIF-1α is required for the formation of lung MPC and may represent a new molecular target for lung MPC therapy.
Results
Hypoxia induces the formation of MLSs in a 3D culture system
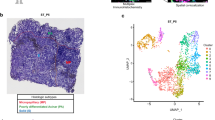
First, we attempted to generate a 3D cell culture system to construct MLSs, which are cell clusters exhibiting inverted polarity similar to the clusters in lung MPC. We cultured various LUAD cell lines on thick Matrigel and established evaluation criteria for whether the cell clusters on Matrigel were MLSs (Fig. 1a). When the apical marker phospho-ezrin/radixin/moesin (pERM) was positive along at least half of the periphery of cell clusters by immunohistochemical staining, the clusters were classified as MLSs (cell clusters with inverted polarity). Conversely, if the cell clusters did not meet this criterion, they were classified as non-MLSs (cell clusters with non-inverted polarity). We combined hypoxia exposure or recombinant TGF-β (rTGF-β) stimulation with the 3D culture system. This was done in light of substantial evidence indicating that such types of stimulation play a role in the morphological changes and invasive potential of tumor cells. Strikingly, when A549 cells were used, the number of MLSs significantly increased to 70.0% ± 3.3% under hypoxia (1% oxygen concentration), whereas the other LUAD cell lines (H1650, H1792, H1975, and HCC827) showed no such changes (Fig. 1b). Although stimulation of A549 cells with rTGF-β also increased the number of MLSs to 32.2% ± 8.4%, the proportion of increased MLSs was lower than in cells exposed to hypoxia (Fig. 1c, Supplementary Fig. S1). In addition, hypoxia increased the number of MLSs in an oxygen concentration-dependent and time-dependent manner (Fig. 1d, e). Therefore, we utilized hypoxia exposure to induce MLSs in the 3D culture system.
Hypoxia induced the formation of MLSs of A549 human LUAD cells in a 3D culture system. (a) Scheme representing a 3D cell culture system. Cells were cultured in the 3D system, then fixed and immunostained with pERM antibody. MLSs (clusters with inverted polarity) were defined as the cell clusters with pERM immunostaining on at least half of their periphery, and the remaining cell clusters were defined as non-MLSs (those with non-inverted polarity). MLSs in 30 to 35 cell clusters were counted, and the percentage of MLSs was calculated. (b) Representative images of each LUAD cell line cultured in the 3D system for 5 days under normoxia (20% oxygen) or hypoxia (1% oxygen). These cells were fixed and immunostained for pERM. F-actin and the nucleus were stained with Alex Flour 594 phalloidin and DAPI, respectively. N, normoxia; H, hypoxia. Scale bar: 20 μm. The percentage of MLSs was quantified as shown in Fig. 1a. Data are presented as mean + SD of three different areas for each condition and are representative of three independent experiments. c, Each LUAD cell line was cultured in the 3D system for 5 days with 10 ng/mL rTGF-β protein. The percentage of MLSs was quantified as shown in Fig. 1b. Data are presented as mean + SD of three different areas for each condition and are representative of three independent experiments. (d, e) A549 cells were cultured in the 3D system for 5 days under the indicated oxygen concentrations (d) or for the indicated time periods under normoxia (20% oxygen) or hypoxia (1% oxygen) (e). The percentage of MLSs was quantified as shown in Fig. 1b. Data are presented as mean ± SD of three different areas for each condition and are representative of three independent experiments. (f) Representative images of A549 cells cultured in the 3D system for 5 days under normoxia (20% oxygen) or hypoxia (1% oxygen). The cells were fixed and immunostained for MUC1, ZO-1, β-catenin, β1-integrin, and pERM. F-actin and the nucleus were stained with Alex Flour 594 phalloidin and DAPI, respectively. Z-stack images were generated from image stacks. T, top; B, bottom. Scale bar, 20 μm. (g, h) Immunoblot analysis for MUC1 in A549 cells cultured in the 2D or 3D system under the indicated concentrations of oxygen for 5 days (g) or for the indicated time periods under hypoxia (1% oxygen) (h). Data are representative of two biologically independent experiments. β-actin was used as a loading control. (i, j) After 5 days of culture in the 3D system under hypoxia (1% oxygen), A549 cells were cultured under normoxia (20% oxygen) or hypoxia (1% oxygen) for the indicated time periods. The percentage of MLSs was quantified as shown in Fig. 1b (i), and MUC1 expression was analyzed via immunoblot (j). Data are presented as mean ± SD of three different areas for each condition and are representative of two independent experiments. P values were calculated using the two-tailed Student’s t-test (***P < 0.001; **P < 0.01).
The MLSs generated in the system exhibited various features that have been described in previous studies of MPC8,18,19. Compared to the cell clusters cultured under normoxia, the apical markers MUC1 and ZO-1 as well as prominent filamentous actin (F-actin) bundles were enriched at the periphery of MLSs in hypoxia (Fig. 1f). By contrast, irrespective of the oxygen concentration, the adherens junction protein β-catenin was expressed at the contact sites between the cells, and the basolateral proteinβ1-integrin was expressed at both the periphery of the cell clusters and at the contact sites between the cells. Z-stack images showed that both pERM and F-actin bundles were highly localized even on the bottom of the cell clusters, where they contacted the Matrigel. Further, we confirmed that hypoxia induced the formation of MLSs even when the entire periphery of the cell cluster was surrounded by Matrigel (Supplementary Fig. S2).
MUC1 is a molecule that characterizes MPC8,9,10, and hypoxia upregulates its expression in lung and renal carcinoma20,21. Indeed, MUC1 expression was augmented in an oxygen concentration-dependent and hypoxia exposure time-dependent manner in A549 cells cultured in the 3D culture system (Fig. 1g, h). Furthermore, the expression level of hypoxia-induced MUC1 was significantly higher in the 3D culture system than in the two-dimensional (2D) culture system.
When MLSs induced by hypoxia were cultured under normoxia, both the number of MLSs and the expression level of MUC1 significantly decreased in a time-dependent manner (Fig. 1i, j). These results indicate that both MLS formation and hypoxia-induced MUC1 expression are reversibly altered depending on the oxygen concentration.
A549 cells forming MLSs exhibit a gene expression signature similar to that of lung MPC
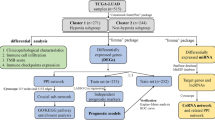
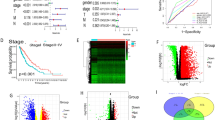
To characterize the hypoxia-induced MLSs in detail, we next performed RNA sequencing (RNA-seq) analysis of A549 cells cultured in the 2D or 3D culture system under normoxia or hypoxia (Fig. 2a). We examined whether there was a difference in the expression of hypoxia signature genes. The expression of hypoxia signature genes obtained from the Hallmark gene set was increased in both the 2D- and 3D-Hypoxia group, and these gene sets tended to be slightly higher in 3D (Fig. 2b). We then investigated how well the hypoxia-induced MLSs mimicked lung MPC with the data from another research group. A microarray dataset obtained from the National Center for Biotechnology Information Gene Expression Omnibus (NCBI GEO) (accession number GSE58772) was used to define the signature genes for lung MPC. The data set included normalized microarray data from the five LUAD subtypes (acinar, solid, lepidic, MPC, and PC) upon laser capture microdissection of fresh frozen LUAD tissue samples from 48 patients22 (Fig. 2c). The top 50 genes upregulated in MPC compared to other LUAD subtypes were defined as the MPC signature genes (Supplementary Table S1). In the microarray dataset, we confirmed that the MPC gene signature score for each MPC case tended to be higher than that for the other LUAD subtype cases. Then we analyzed the expression of the MPC signature genes in A549 cells cultured under each condition shown in Fig. 2a (Fig. 2d, Supplementary Table S2). Notably, the heatmap in Fig. 2d shows that the expression of the MPC signature genes was increased only in the 3D-Hypoxia group, which is the condition under which the MLSs were induced, and the calculated MPC gene signature score for each condition was consistent with the heatmap. These results suggest that MLSs formed by A549 cells truly mimic lung MPC and that both hypoxic conditions and a 3D cell culture system are required for their formation.
A549 cells forming MLSs exhibited a gene expression signature similar to that of lung MPC. (a) Schematic representation of the preparation of RNA-seq samples from A549 cells cultured in the 2D or 3D system normoxia (20% oxygen) or hypoxia (1% oxygen). (b) The left panel indicates a heatmap showing the expression values for the hypoxia signature genes obtained from the Hallmark gene set (HALLMARK_HYPOXIA) in each culture condition of A549 cells as indicated in Fig. 2a. The right panel shows the hypoxia gene signature scores calculated using RNA-seq data from each culture condition of A549 cells as indicated in Fig. 2a. (c) The upper panel shows a schematic representation of the analysis of a public microarray dataset from five different adenocarcinoma architectures upon laser capture microdissection of 48 LUAD cases (acinar from 10 cases, solid from 10 cases, lepidic from 10 cases, MPC from 9 cases, and PC from 9 cases). Fifty genes upregulated in MPC compared with other LUAD subtypes were identified using the dataset, and they were defined as the MPC signature genes. The lower panel shows the MPC gene signature scores for each LUAD subtype. Each dot in the graph represents a single case. (d) The left panel shows a heatmap exhibiting the expression values for the MPC signature genes in each culture condition of A549 cells as indicated in Fig. 2a. The right panel shows the MPC gene signature scores calculated using RNA-seq data from each culture condition of A549 cells as indicated in Fig. 2a.
HIF-1α is indispensable for hypoxia-induced formation of MLSs
To elucidate the molecular mechanism underlying hypoxia-induced MLS formation, we examined the changes in the expression of HIF-1α in A549 cell clusters cultured in the 3D system. Hypoxia exposure increased the expression of HIF-1α in an oxygen concentration-dependent manner (Supplementary Fig. S3a), and this effect was correlated with the hypoxia-induced increase in the number of MLSs (Fig. 1d). The expression of HIF-1α peaked 4 h after hypoxia exposure and then gradually decreased (Supplementary Fig. S3b). Next, we examined the effect of the HIF-1α inhibitor YC-1 on hypoxia-induced MLS formation in A549 cells. YC-1 inhibited the formation in a dose-dependent manner (Fig. 3a). In addition, YC-1 treatment for a 5-day period after induction of MLSs by hypoxia reduced the number of MLSs, suggesting that HIF-1α is required to maintain the structure of MLSs (Fig. 3b). To further elucidate the involvement of HIF-1α in MLS formation, we disrupted the expression of HIF-1α in A549 cells using a CRISPR/Cas9 system (Fig. 3c). HIF-1α knockout (KO) significantly reduced the number of hypoxia-induced MLSs at each oxygen concentration (Fig. 3d). However, HIF-1α-KO clone number 17 could not be cultured for 5 days in 1% oxygen. Moreover, HIF-1α KO reduced the hypoxia-induced MUC1 expression in 2.5% oxygen (Fig. 3e). These results indicate that HIF-1α is required for both hypoxia-induced formation of MLSs and enhancement of MUC1 expression in A549 cells. However, MUC1 KO in A549 cells using the CRISPR/Cas9 system did not affect hypoxia-induced MLS formation (Fig. 3f,g).
HIF-1α was indispensable for the formation of MLSs. (a) A549 cells were treated with the indicated concentrations of YC-1, an HIF-1α inhibitor, in the 3D system for 5 days under hypoxia (1% oxygen). The percentage of MLSs was quantified as shown in Fig. 1b. Data are presented as mean + SD of three different areas for each condition and are representative of three independent experiments. (b) After 5 days of culture in the 3D system under hypoxia (1% oxygen), A549 cells were treated with the indicated concentrations of YC-1 under hypoxia (1% oxygen) for 5 days. The percentage of MLSs was quantified as shown in Fig. 1b. Data are presented as mean ± SD of three different areas for each condition and are representative of three independent experiments. (c) Immunoblot analysis for HIF-1α in parent cells and HIF-1α-KO A549 cells cultured for 4 h under normoxia (20% oxygen) or hypoxia (1% oxygen). Data are representative of two biologically independent experiments. N, normoxia; H, hypoxia. (d) Parent cells and HIF-1α-KO A549 cells were cultured in the 3D system for 5 days under the indicated oxygen concentrations. The percentage of MLSs was quantified as shown in Fig. 1b. Data are presented as mean + SD of three different areas for each condition and are representative of three independent experiments. ND, not determined. F-actin and the nucleus were stained with Alex Flour 594 phalloidin and DAPI, respectively. (e), Immunoblot analysis for MUC1 in parent cells and HIF-1α-KO A549 cells cultured in the 3D system for 5 days under normoxia (20% oxygen) or hypoxia (1% oxygen). Data are representative of two biologically independent experiments. N, normoxia; H, hypoxia. GAPDH was used as a loading control. (f) Immunoblot analysis for MUC1 in control (Ctrl) and MUC1-KO A549 cells cultured in the 3D system for 5 days under normoxia (20% oxygen) or hypoxia (1% oxygen), repeated in two biologically independent experiments. (g) Control and MUC1-KO A549 cells were cultured in the 3D system for 5 days under normoxia (20% oxygen) or hypoxia (1% oxygen). The percentage of MLSs was quantified as shown in Fig. 1b. Data are presented as mean + SD of three different areas for each condition and are representative of three independent experiments. P values were calculated using the two-tailed Student’s t-test (***P < 0.001; **P < 0.01; *P < 0.05; NS, not significant).
RhoA/Rho-kinase signaling is involved in hypoxia-induced formation of MLSs
Because the prominent F-actin bundles were localized at the periphery of MLSs (Fig. 1f), we hypothesized that increased actomyosin contractility is important for MLS formation. In line with this hypothesis, immunostaining revealed that the phosphorylated myosin light chain (phospho-MLC) at Ser19, which activates myosin II and regulates the actomyosin assembly, was localized at the peripheral apical cortex of MLSs (Fig. 4a). Next, we examined the effect of inhibitors of myosin II or Rho-kinase, which phosphorylate the MLC23 and play a role in regulating apicobasal polarity in Madin–Darby canine kidney (MDCK) cells24. The Rho-kinase inhibitors Y27632 and fasudil and the myosin II inhibitor blebbistatin significantly abolished the hypoxia-induced MLS formation (Fig. 4b). We confirmed that Y27632 and fasudil reduced the phosphorylation of MLC at Ser19 (Fig. 4c). To elucidate the involvement of RhoA (the upstream molecule of Rho-kinase), we generated A549 cells that stably expressed RhoA T19N, a dominant-negative form of RhoA (Fig. 4d). Notably, RhoA T19N expression significantly reduced the number of hypoxia-induced MLSs (Fig. 4e). These results suggest that activation of RhoA/Rho-kinase is required for hypoxia-induced MLS formation.
RhoA/Rho-kinase signaling was involved in the formation of MLSs. (a) Representative images of A549 cells cultured in the 3D system for 5 days under normoxia (20% oxygen) or hypoxia (1% oxygen). The cells were fixed and immunostained for phosphor-MLC (Ser19). F-actin and the nucleus were stained with Alex Flour 594 phalloidin and DAPI, respectively. (b) Representative images of A549 cells in the control condition (DMSO) or treated with 5 μM Y27632, fasudil, or blebbistatin in the 3D system for 5 days under hypoxia (1% oxygen) and immunostained for pERM. F-actin and the nucleus were stained with Alex Flour 594 phalloidin and DAPI, respectively. The percentage of MLSs was quantified as shown in Fig. 1b. Data are presented as mean + SD of three different areas for each condition and are representative of three independent experiments. Scale bars, 20 μm. (c) Immunoblot analysis for phospho-MLC in A549 cells treated with 5 μM Y27632, fasudil, blebbistatin, or DMSO as a negative control in the 3D system for 48 h under hypoxia (1% oxygen). β-actin was used as a loading control. (d) Immunoblot analysis for HA in A549 cells stably expressing HA-tagged RhoA T19N. Data are representative of two biologically independent experiments. (e) Control and A549 cells expressing HA-tagged RhoA T19N were cultured in the 3D system for 5 days under normoxia (20% oxygen) or hypoxia (1% oxygen). The percentage of MLSs was quantified as shown in Fig. 1b. Data are presented as mean + SD of three different areas for each condition and are representative of three independent experiments. P values were calculated using the two-tailed Student’s t-test (***P < 0.001; **P < 0.01).
Immunohistological analysis of the molecules associated with the A549 3D cell culture model
We evaluated the expression levels of molecules associated with the A549 3D cell culture model in LUAD specimens containing both MPC and PC components. Immunostaining revealed that both MPC and PC components exhibited robust expression of MUC1 at the plasma membrane. However, there was a discrepancy in the ratio of positive margins. To evaluate this, we devised a distinctive scoring method, the tumor cell (TC)-score, to ensure accurate and consistent results (Fig. 5a). Notably, the TC-score showed that MUC1 expression levels were higher in MPC cells than in PC cells (Fig. 5b,c). With regard to the expression of HIF1A, the immunostaining did not show significant statistical differences between MPC and PC (Fig. 5d,e). Immunostaining for pERM showed no statistically significant difference between MPC and PC for its expression level (Fig. 5f,g).
The expression of MUC1, HIF-1α, and pERM in LUAD specimens by immunohistochemistry. (a) TC (tumor cell)-score schematic and formula. In this scoring system, one tumor cell was randomly selected from the MPC or PC component. MUC1 expression level in the tumor cell was classified as either high intensity (score 2) or low intensity (score 1). The score was multiplied by the positivity rate of the tumor cell periphery and calculated using the following formula: [1 × (% of the circumference with score 1)] or [2 × (% of the circumference with score 2)]. (b) Representative immunohistochemistry images showing MUC1 expression. Scale bars, 100 μm. (c) Box plot of MUC1 TC-scores calculated for 100 PC and 100 MPC cells in 10 cases of LUAD specimens. (d) Representative immunohistochemistry images showing HIF-1α expression. Scale bars, 100 μm. (e) Box plot of HIF-1α H-scores calculated for PC and MPC components. (f) Representative immunohistochemistry images showing pERM expression. Scale bars, 100 μm. (g) Boxplot of pERM H-scores calculated for PC and MPC components. P values were calculated using the two-tailed Student’s t-test (***P < 0.001; **P < 0.01; *P < 0.05; NS, not significant).
A549 cells forming MLSs showed enhanced resistance to NK cell cytotoxicity
HIF-1α is involved in tumor immune escape25,26, including escape from immune surveillance by NK cells27, leading to the promotion of tumorigenesis. Therefore, we hypothesized that the malignant behaviors of lung MPC may be caused by its immune escape from NK cells. Indeed, immunohistochemical staining showed a reduction in CD56 + NK cell infiltration in MPC compared to PC (Supplementary Fig. S4).
To verify this hypothesis, we examined the sensitivity of A549 cells forming MLSs to NK cell cytotoxic activity using a lactate dehydrogenase (LDH) assay (Fig. 6a). When A549 cells were cultured in the 2D culture system, the human NK cell line KHYG-1 showed cytotoxicity against the cells in a target/effector cell ratio-dependent manner, and the cytotoxic activity was attenuated in the cells cultured under hypoxia (Fig. 6b). Notably, when A549 cells were cultured in the 3D culture system under hypoxic conditions and formed MLSs, the cytotoxic activity of KHYG-1 cells against these cells was further attenuated compared to the activity against the cells cultured in the 2D system (Fig. 6b). In addition, the cytotoxic activity of NK-92 MI, another human NK cell line, against A549 cells was attenuated under hypoxic conditions when the cells were cultured in the 3D system but not in the 2D system (Fig. 6c). These results suggest that the structure of MLSs contributes to the attenuation of NK cell cytotoxic activity. KO of HIF-1α in A549 cells almost completely restored their sensitivity to the cytotoxic activity of both KHYG-1 and NK-92 MI cells (Fig. 6d, e). However, KO of MUC1 in A549 cells did not affect their sensitivity to the cytotoxic activity of KHYG-1 cells (Fig. 6f). To further investigate the involvement of the structure of MLSs in the escape from NK cell cytotoxicity, we disrupted the actin cytoskeletal structure of MLSs by Y27632. Treatment of MLSs with Y27632 for 3 h significantly reduced the number of MLSs in a dose-dependent manner, and the effect was sustained for 24 h under normoxia even after the removal of Y27632 (Fig. 6g). The Y27632 treatment partially restored the sensitivity of A549 cells to NK cell cytotoxicity, suggesting that the actin cytoskeletal structure of MLSs (as well as other factors induced by HIF-1α) is responsible for the enhanced resistance to NK cell cytotoxicity (Fig. 6h).
A549 cells forming MLSs showed low sensitivity to NK cell cytotoxicity, and KO of HIF-1α restored the sensitivity to NK cells. (a) Flow chart showing the experimental scheme to evaluate NK cell-mediated cytotoxicity for A549 cells using the LDH assay. (b) LDH-release-based NK cell cytotoxicity test with KHYG-1 cells. A549 cells were cultured in the 2D or 3D culture system for 5 days under normoxia (20% oxygen) or hypoxia (1% oxygen), and then KHYG-1 cells were added at the indicated ratios. Medium (50 μL) was collected after 24 h coculture, and LDH release was measured according to the manufacturer’s instructions. Data are presented as mean + SD of three biologically independent samples and are representative of three independent experiments. (c) LDH-release-based NK cell cytotoxicity test with NK-92 MI cells. A549 cells were cultured in the 3D culture system for 5 days under normoxia (20% oxygen) or hypoxia (1% oxygen), and then NK-92 MI cells were added at a 1:20 ratio. Medium (50 μL) was collected after 6 h coculture, and LDH release was measured as shown in Fig. 6b. The results are shown as relative cytotoxicity to normoxia culture. Data are presented as mean + SD of three biologically independent samples and are representative of three independent experiments. (d, e) LDH-release-based NK cell cytotoxicity test with KHYG-1 cells (d) or NK-92 MI cells (e) for parent cells and HIF-1α-KO A549 cells. Cells were cultured in the 3D system for 5 days under normoxia (20% oxygen) or hypoxia (2.5% oxygen), and then KHYG-1 cells or NK-92 MI cells were added at a 1:20 ratio. Medium (50 μL) was collected after 24 h (KHYG-1 cells) or 6 h (NK-92 MI cells) of coculture, and LDH release was measured as shown in Fig. 6b. The results are shown as relative cytotoxicity to normoxia culture. Data are presented as mean + SD of three biologically independent samples and are representative of three independent experiments. (f) LDH-release-based NK cell cytotoxicity test with KHYG-1 cells for control cells and MUC1-KO A549 cells. Cells were cultured in the 3D culture system for 5 days under normoxia (20% oxygen) or hypoxia (1% oxygen), and then KHYG-1 cells were added at a 1:20 ratio. Medium (50 μL) was collected after 24 h coculture, and LDH release was measured as shown in Fig. 6b. The results are shown as relative cytotoxicity to normoxia culture. Data are presented as mean + SD of three biologically independent samples and are representative of three independent experiments. (g) After 5 days of culture in the 3D system under hypoxia (1% oxygen), A549 cells were treated with the indicated concentrations of Y27632 for 3 h under normoxia. Then Y27632 was removed by media replacement, and A549 cells were cultured under normoxia for an additional 21 h. The percentage of MLSs was quantified as shown in Fig. 1b. Data are presented as mean ± SD of three different areas for each condition and are representative of two biologically independent experiments. (h) LDH-release-based NK cell cytotoxicity test with KHYG-1 cells for A549 cells treated with Y27632 as shown in Fig. 6g. After treatment of A549 cells with Y27632 for 3 h, KHYG-1 cells were added at a 1:20 ratio. Medium (50 μL) was collected after 24 h coculture, and LDH release was measured as shown in Fig. 6b. Data are presented as mean + SD of three biologically independent samples and are representative of three independent experiments. P values were determined by two-tailed Student’s t-test.
Collectively, these findings support the idea that lung MPC can be induced by hypoxia via HIF-1α and that its structure can be reversibly altered depending on the oxygen concentration in the tumor microenvironment. Furthermore, immune escape of lung MPC from NK cells may be brought about by not only hypoxic conditions but also the actin cytoskeletal structure of MLSs, leading to the malignancy of MPC (Fig. 7).
Schematic diagram of the hypoxia-induced lung MPC structure through HIF-1α. Hypoxia induces activation of HIF-1α in lung adenocarcinoma cells, which induces MPC structure through RhoA-dependent formation of inverted polarity and high expression of MUC1. This results in immune escape from NK cells.
Discussion
The recent development of 3D cell culture models has facilitated the ability to elucidate the structure, organization, and functionality of tumors in vivo28,29. In this study, combining a 3D cell culture system using Matrigel with hypoxic conditions, we constructed MLSs of A549 cells, which are cell clusters that have inverted polarity and exhibit a gene expression signature similar to that of lung MPC.
The most remarkable feature of MPC is its inverted polarity8,9,10. The mechanisms of polarity formation have been studied using multiple models, particularly MDCK cells30. When aggregates of MDCK cells are embedded in an extracellular matrix (ECM) such as collagen or Matrigel, they form a hollow cyst lined by a single cell layer, and the apical membrane forms inside the lumen31,32. By contrast, defects in ECM signaling by either suspension culture33 or treatment with a function-blocking antibody of β1-integrin31,32 lead to inverted polarity, forming an apical membrane on the outside of the cell cluster. Inhibition of RhoA/Rho-kinase, which is downregulated by ECM signals through β1-integrin at the ECM interface, eliminates the inverted polarity24,32. Although similarity between the inverted polarity of MDCK cells and the MPC phenotype has been suggested34, the cell clusters in MPC and MLSs in this study differed from MDCK cells in that they exhibited inverted polarity despite being surrounded by ECM and were in a state of abnormal polarity. Notably, the development of inverted polarity in MLSs was suppressed by inhibition of RhoA/Rho-kinase signaling as well as that in MDCK (Fig. 4b, e). These results suggest that in MLSs, RhoA/Rho-kinase is activated despite the presence of an ECM such as Matrigel. These findings combined with the finding that HIF-1α is required for the formation of MLSs (Fig. 3d) indicate that HIF-1α might be involved in the activation of RhoA/Rho-kinase. However, further studies are needed to elucidate the precise molecular mechanisms underlying how hypoxia induces the formation of MLSs.
The tumor microenvironment, which is composed of tumor cells, stromal cells, immune cells, blood vessels, and ECM, plays multiple central roles in cancer progression35. Hypoxia is one of the most important features of the tumor microenvironment36. In this study, we showed that hypoxia and HIF-1α activation are key factors that induce the inverted polarity characteristic of lung MPC. Interestingly, hypoxia increased the frequency of MLSs in A549 cell clusters in an oxygen concentration-dependent manner (Fig. 1d), and MLSs once induced by hypoxia returned to non-MLSs under normoxia (Fig. 1i). These results might explain why pure MPC is rarely observed in human LUAD tissues and why MPC is a component of LUAD with mixed subtypes6,37,38. Specifically, the differences in the oxygen concentration within the lung tumor microenvironment may be responsible for differences in the forces that drive the formation or maintenance of MPC.
The immune microenvironment is one of the most important factors in the tumor microenvironment35. Research in the field of tumor immunobiology has identified immune escape as a classic hallmark of cancer and has characterized different escape strategies utilized by cancer cells39,40. Importantly, hypoxia is involved in tumor immune escape25,26, including escape from immune surveillance by NK cells27, and promotes tumorigenesis. In this study, A549 cells forming MLSs exhibited enhanced resistance to the cytotoxic activity of NK cells, and the actin cytoskeletal structure of MLSs was at least partially responsible for the enhanced resistance (Fig. 6g, h). In a previous study, breast cancer cells responded to NK cell attack via surprisingly rapid and massive accumulation of F-actin near the immunologic synapse, and this actin response contributed to tumor cell resistance to NK cell-mediated cell death41. In our study, abundant localization of F-actin was observed on the outer surface of MLSs (Fig. 1f), and the data indicated that this F-actin might be involved in the hypoxia-induced enhancement of resistance to the cytotoxic activity of NK cells. Further studies are needed to elucidate the detailed mechanisms.
In summary, we explored the mechanism underlying lung MPC formation by combining a 3D culture system and hypoxic conditions. We discovered that hypoxia drives the formation of the inverted polarity characteristic of lung MPC via HIF-1α. Our results suggest that both the morphology and malignancy of LUAD can be altered by the oxygen concentration in the lung tumor microenvironment and that HIF-1α is a potential therapeutic target for lung MPC.
Methods
Antibodies and reagents
Rabbit monoclonal antibody to HIF-1α (ab51608) and MUC1 (ab109185), rabbit polyclonal antibody to β-catenin (ab16051), and mouse monoclonal antibody to β1-integrin (ab24693) were purchased from Abcam (Cambridge, UK). Mouse monoclonal antibody to MUC1 (ACR319B) was purchased from Biocare Medical (Pacheco, CA, USA). Rabbit polyclonal antibody to phospho-MLC (Ser19) (no. 3671S) and β-actin (conjugated to horseradish peroxidase) (no. 5125) and rabbit monoclonal antibody to phospho-ezrin (Thr567)/radixin (Thr564)/moesin (Thr558) (no. 3726S) and GAPDH (no. 2118S) were purchased from Cell Signaling Technology (Danvers, MA, USA). Rabbit polyclonal antibody to ZO-1 (no. 617300) was purchased from Thermo Fisher Scientific (Waltham, MA, USA). Mouse monoclonal antibody to HA-tag (M180–3) was purchased from Medical & Biological Laboratories (Tokyo, Japan). Other reagents were obtained from the following sources: Alexa Fluor 594 phalloidin was obtained from Thermo Fisher Scientific. rTGF-β1 (human cell-expressed) protein was obtained from R&D Systems (Minneapolis, MN, USA). YC-1 was obtained from Sigma-Aldrich (St. Louis, MO, USA). Puromycin was obtained from InvivoGen (San Diego, CA, USA). Y27632 was obtained from Enzo Life Sciences (Farmingdale, NY, USA). Blebbistatin was obtained from Wako (Tokyo, Japan). Fasudil hydrochloride was obtained from Tokyo Chemical Industry (Tokyo, Japan).
Surgical specimens from patients with LUAD
LUAD tissues were obtained from patients who underwent surgical resection at Osaka University Hospital. These tissue specimens were fixed in 10% formalin and routinely processed for paraffin embedding. Informed consent was obtained from all patients.
Ethics
All experimental protocols involving human specimens were approved by the Ethics Review Board of the Graduate School of Medicine, Osaka University (approval no. 15234) and carried out in accordance with the committee’s guidelines and regulations. The study was performed in accordance with the Declaration of Helsinki.
Cell lines and cell culture (2D cell culture system)
LUAD cell lines (A549, H1650, H1792, H1975, and HCC827) were obtained from the American Type Culture Collection (Manassas, VA, USA). These cells were cultured in Dulbecco’s modified Eagle medium (DMEM) supplemented with 10% fetal bovine serum (FBS), penicillin (100 IU/mL), and streptomycin (100 μg/mL); they were cultured as a monolayer in a flat petri dish attached to a plastic surface at 37 °C in a 5% carbon dioxide atmosphere (2D cell culture system). The human NK cell line KHYG-1, obtained from JCRB Cell Bank (Osaka, Japan), was cultured in Roswell Park Memorial Institute (RPMI) 1640 medium containing 10% FBS and 200 units/mL recombinant interleukin-2 (Peprotech, Rocky Hill, NJ, USA). Human NK-92 MI cells were purchased from the American Type Culture Collection and cultured in MyeloCult™ H5100 (STEMCELL Technologies, Vancouver, Canada).
Cell culture on or in Matrigel (3D cell culture system)
For cell culture on Matrigel, we spread 60 μL growth factor-reduced Matrigel (Corning, Corning, NY, USA) on a round coverslip (15 mm diameter) and incubated it at 37 °C for 15 min to solidify the gel. LUAD cell lines were trypsinized to a single-cell suspension at 5 × 104 cells/mL in DMEM supplemented with 5% FBS, seeded on the solidified thick gel, and then incubated for the indicated time periods. For cell culture in Matrigel, the cell pellet (4 × 104 cells) was resuspended in 30 mL Matrigel and pipetted into the center of a well of a 24-well plate to form a dome. The plate was placed in a humidified incubator to polymerize the Matrigel (37 °C, 10–15 min), and then DMEM supplemented with 5% FBS was added on the top of the solidified dome (1 mL/well in a 24-well plate). The cells were cultured at 37 °C for 5 days.
Immunofluorescence
Samples were washed in PBS and fixed in 4% paraformaldehyde for 15 min. Permeabilization and blocking were performed in PBS containing 0.25% Triton X-100 and 2% bovine serum albumin. Then the cells were incubated with each primary antibody (anti-phospho-ERM [1:500 dilution], anti-MUC1 [1:250 dilution; Biocare Medical], anti-ZO-1 [1:250 dilution], anti-β-catenin [1:250 dilution], anti-β1-integrin [1:250 dilution], and anti-phospho-MLC [Ser19; 1:250 dilution]) overnight at 4 °C in PBS containing 2% bovine serum albumin. Secondary antibody (anti-rabbit or anti-mouse antibody conjugated with Alexa Fluor 488) and Alexa Fluor 594 phalloidin were incubated for 1.5 h at room temperature. Finally, the slide was mounted using Fluoro-KEEPER Antifade Reagent, Non-Hardening Type with DAPI (Nacalai Tesque, Kyoto, Japan). Fluorescent images were obtained using an LSM880 or LSM710 confocal laser scanning microscope (Carl Zeiss, Oberkochen, Germany) and analyzed with Zen software (Carl Zeiss).
Immunocytochemistry of the cell line
A549 cells cultured in Matrigel under normoxia or hypoxia (1% oxygen) were washed in phosphate-buffered saline (PBS) and fixed in 4% paraformaldehyde for 15 min. After centrifugation, the fixed cell pellets were pre-embedded in agarose gel and then embedded in paraffin using the same method used for tissue sample processing. The paraffin-embedded cells were sectioned, processed, and subjected to immunohistochemistry with an anti-phospho-ERM antibody (1:400 dilution) and an anti-MUC1 antibody (1:800 dilution; Abcam, Cambridge, UK) in accordance with the manufacturer’s instructions. 3,3'-Diaminobenzidine (DAB) (Dako/Agilent, Santa Clara, CA, USA) was used as the chromogen.
Immunohistochemistry of the sectioned tissues
Paraffin-embedded tissues were sectioned, processed, and subjected to immunohistochemistry using Dako Autostainer Link 48 (Dako/Agilent) with an anti-MUC1 (1:800 dilution; Abcam), anti-HIF-1α (1:400 dilution), anti-pERM (1:400 dilution), and anti-CD56 antibody (1:200 dilution). DAB (Dako/Agilent) was used as the chromogen. The expression was assessed using a visual grading system based on the intensity of staining under a light microscope. High intensity (score 2) and low intensity (score 1) were defined as strong and weak staining, respectively. H-scores were assigned using the following formula: (1 × (% cells of score 1) + 2 × (% cells of score 2)). In the TC (tumor cell)-scoring system, one tumor cell was randomly selected from the MPC or PC component. MUC1 expression level in the tumor cell was classified as either high intensity (score 2) or low intensity (score 1). The score was multiplied by the positivity rate of the tumor cell periphery and calculated using the following formula: [1 × (% of the circumference with score 1)] or [2 × (% of the circumference with score 2)].
Generation of HIF-1α or MUC1-KO A549 cells
For HIF-1α-KO, we used the CRISPR/Cas9 system manufactured by Integrated DNA Technologies (Coralville, IA, USA). Guide RNA Hs.Cas9.HIF1A.1.AC was first duplexed with ATTO 550 Alt-R CRISPR/Cas9 trans-activating CRISPR RNA, then complexed to Alt-R S.p. HiFi Cas9 Nuclease V3 according to the manufacturer’s instructions. The complex was delivered into A549 cells using Lipofectamine RNAiMAX Transfection Reagent (Thermo Fisher Scientific). A549 cells positive for ATTO 550 were sorted using an SH800 cell sorter (Sony, Tokyo, Japan) into single cells 48 h after transfection. The gene-edited clone that had been selected using a T7 endonuclease I cleavage assay (Integrated DNA Technologies) was transfected with the above Cas9 ribonucleoprotein complex and sorted again in the same way. Among the sorted clones, HIF-1α-KO clones were screened by immunoblotting. For MUC1-KO, we used the CRISPR/Cas9 system manufactured by Santa Cruz Biotechnology (Dallas, TX, USA). MUC1 CRISPR/Cas9 KO plasmid (cat# sc-416876) and MUC1 HDR plasmid (cat# sc-416876-HDR) were mixed and delivered into A549 cells using Lipofectamine 3000 Transfection Reagent (Thermo Fisher Scientific). Transfected cells were selected and cloned in DMEM containing 1 μg/mL puromycin. Among the puromycin-resistant clones, MUC1-KO clones were screened by immunoblotting. Similarly, we transfected the control CRISPR/Cas9 plasmid (cat# sc-418922) and the MUC1 HDR plasmid into A549 cells and constructed a puromycin-resistant clone, which was used as a control.
Generation of A549 cells stably expressing the dominant-negative mutant RhoA T19N
pEF-BOS-HA-RhoA T19N containing the HA-tagged dominant-negative mutant RhoA T19N was kindly supplied by Kozo Kaibuchi (Fujita Health University, Toyoake, Japan). A549 cells were cotransfected with the plasmid and a Linear Puromycin Marker (Takara Bio, Shiga, Japan) using Lipofectamine 3000 (Thermo Fisher Scientific). Two days after transfection, the cells were trypsinized and diluted. One day later, the medium was replaced with fresh medium containing 1 μg/mL puromycin to select and establish stably transfected cells. Similarly, we transfected the Linear Puromycin Marker into A549 cells and constructed a puromycin-resistant clone, which was used as a control.
Cytotoxicity analysis
To evaluate the cytotoxic activity of human NK cell lines (KHYG-1 and NK-92 MI) to A549 cells, we performed an LDH assay using a CyQUANT LDH cytotoxicity assay kit (Thermo Fisher Scientific) in accordance with the manufacturer’s protocol. The release of LDH into the cell culture medium is an indicator of cytotoxicity and cytolysis. In brief, A549 cells (5 × 103 cells/well) were cultured for 5 days in 96-well plates coated with Matrigel (30 µL/well) under normoxia or hypoxia and then cocultured with the KHYG-1 and NK-92 MI cells under normoxia for 24 h and 6 h, respectively. Next, 50 µL coculture medium was transferred to a new plate and added to 50 µL LDH reaction mixture. After incubation at room temperature in the dark, the reaction was stopped by adding 50 µL LDH stop solution. LDH activity was determined at 490 nm and 680 nm using a microplate reader. NK cytotoxicity was calculated as follows: % cytotoxicity = (experimental value – target cell spontaneous control) / (target cell maximum control – target cell spontaneous control) × 100.
Immunoblotting
Cells were rinsed with ice-cold PBS and lysed in ice-cold lysis buffer (1% NP-40, 25 mM Tris–HCl, 150 mM NaCl, 1 mM EDTA, 10 mM MgCl2, and 2% glycerol) containing EDTA-free complete protease inhibitor cocktail (Roche Diagnostics, Basel, Switzerland). The cells were immediately scraped off the plate and sonicated for 10 s. The soluble fractions from the cell lysates were isolated by centrifugation at 13,000 rpm for 5 min in a microcentrifuge. Next, 2 × SDS buffer (4% SDS, 125 mM Tris–HCl [pH 6.8], 20% glycerol, 0.01% bromophenol blue, and 10% 2-mercaptoethanol) was added to the cell lysates and boiled for 5 min at 98 °C. Proteins were analyzed by SDS–polyacrylamide gel electrophoresis and immunoblotting following standard protocols using the following antibodies: anti-HIF-1α (1:1,000 dilution), anti-MUC1 (1:1,000 dilution; Abcam), anti-phospho-MLC (Ser19) (1:1,000 dilution), anti-HA (1:1,000 dilution), anti-GAPDH (1:1,000 dilution), and anti-β-actin (horseradish peroxidase-conjugated) (1:1,000 dilution).
RNA-seq and data analysis
Library preparation was prepared using a TruSeq stranded mRNA sample prep kit (Illumina, San Diego, CA), following the manufacturer’s instructions. Subsequent sequencing was performed on an Illumina NovaSeq 6000 sequencer (Illumina) in the 101-base single-end mode. The sequenced reads were preprocessed by Trim Galore v0.6.7 (RRID:SCR_011847) and quantified by Salmon v1.4.0 (RRID:SCR_017036) with the flags gcBias and validateMappings. The GENCODE v37 (GRCh38.p13, RRID:SCR_014966) annotation was used as the transcript reference. All procedures were implemented using the RNA-seq pipeline ikra v2.0.1 (http://doi.org/https://doi.org/10.5281/zenodo.4718200) with default parameters. The quantified transcript-level TPM (transcripts per million) was summarized into a gene-level TPM using tximport v1.22.0 (RRID:SCR_016752). Low-expression genes that had < 1 TPM were filtered. In order to define gene signature for micropapillary subtype, microarray dataset obtained from the Gene Expression Omnibus (GEO, RRID:SCR_005012) with accession number GSE58772 were used. The data set includes normalized microarray data at the probe set level from different adenocarcinoma architectures upon laser capture microdissection of fresh frozen lung adenocarcinoma samples. Multiple probe sets for the same gene were collapsed using avereps function by LIMMA package (RRID:SCR_010943). For filtering out probes that were not significantly expressed, only probes that were expressed in at least ten arrays according to a detection p-value of 0.05 were kept. Differential expression analysis was performed using LIMMA and the results were sorted by the p-value. Top 50 upregulated genes in micropapillary group which were commonly detected in in-house RNA-seq data were defined as micropapillary gene signature.
Statistical analyses
Data are presented as mean ± standard deviation (SD). Significance was tested with the unpaired two-tailed Student’s t-test using GraphPad Prism according to the manufacturer’s instructions. Statistical significance was set at P < 0.05.
Data availability
The RNA-sequencing data obtained in this study are publicly available in Gene Expression Omnibus (GEO) at GSE240551. The data in this study are available from the corresponding authors (S Nojima and E Morii) upon reasonable request.
References
Sung, H. et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71, 209–249 (2021).
Nicholson, A. G. et al. The 2021 WHO Classification of lung tumors: Impact of advances since 2015. J. Thorac. Oncol. 17, 362–387 (2022).
Nassar, H. Carcinomas with micropapillary morphology: Clinical significance and current concepts. Adv. Anat. Pathol. 11, 297–303 (2004).
Nagano, T. et al. Structural and biological properties of a papillary component generating a micropapillary component in lung adenocarcinoma. Lung Cancer 67, 282–289 (2010).
Monroig-Bosque, P. D. C. et al. Micropapillary adenocarcinoma of lung: Morphological criteria and diagnostic reproducibility among pulmonary pathologists. Ann. Diagn. Pathol. 41, 43–50 (2019).
Lee, G. et al. Clinical impact of minimal micropapillary pattern in invasive lung adenocarcinoma: Prognostic significance and survival outcomes. Am. J. Surg. Pathol. 39, 660–666 (2015).
Sakamoto, K. et al. Primary invasive micropapillary carcinoma of the colon. Histopathology 47, 479–484 (2005).
Nassar, H. et al. Pathogenesis of invasive micropapillary carcinoma: Role of MUC1 glycoprotein. Mod. Pathol. 17, 1045–1050 (2004).
Kawakami, T. et al. Micropapillary pattern and grade of stromal invasion in pT1 adenocarcinoma of the lung: Usefulness as prognostic factors. Mod. Pathol. 20, 514–521 (2007).
Tsutsumida, H. et al. A micropapillary pattern is predictive of a poor prognosis in lung adenocarcinoma, and reduced surfactant apoprotein A expression in the micropapillary pattern is an excellent indicator of a poor prognosis. Mod. Pathol. 20, 638–647 (2007).
Brahimi-Horn, M. C., Chiche, J. & Pouysségur, J. Hypoxia and cancer. J. Mol. Med. (Berl) 85, 1301–1307 (2007).
Gilkes, D. M., Semenza, G. L. & Wirtz, D. Hypoxia and the extracellular matrix: Drivers of tumour metastasis. Nat. Rev. Cancer 14, 430–439 (2014).
Muz, B., de la Puente, P., Azab, F. & Azab, A. K. The role of hypoxia in cancer progression, angiogenesis, metastasis, and resistance to therapy. Hypoxia (Auckl) 3, 83–92 (2015).
Shi, R. et al. Identification and validation of hypoxia-derived gene signatures to predict clinical outcomes and therapeutic responses in stage I lung adenocarcinoma patients. Theranostics 11, 5061–5076 (2021).
Semenza, G. L. Targeting HIF-1 for cancer therapy. Nat. Rev. Cancer 3, 721–732 (2003).
Lee, P., Chandel, N. S. & Simon, M. C. Cellular adaptation to hypoxia through hypoxia inducible factors and beyond. Nat. Rev. Mol. Cell Biol. 21, 268–283 (2020).
Bui, B. P., Nguyen, P. L., Lee, K. & Cho, J. Hypoxia-inducible factor-1: A novel therapeutic target for the management of cancer, drug Resistance, and cancer-related pain. Cancers (Basel) 14, 6054 (2022).
Kamiya, K. et al. Histopathological features and prognostic significance of the micropapillary pattern in lung adenocarcinoma. Mod. Pathol. 21, 992–1001 (2008).
Liu, B. et al. Overexpression of β1 integrin contributes to polarity reversal and a poor prognosis of breast invasive micropapillary carcinoma. Oncotarget 9, 4338–4353 (2017).
Mikami, Y. et al. Hypoxia enhances MUC1 expression in a lung adenocarcinoma cell line. Biochem. Biophys. Res. Commun. 379, 1060–1065 (2009).
Aubert, S. et al. MUC1, a new hypoxia inducible factor target gene, is an actor in clear renal cell carcinoma tumor progression. Cancer Res. 69, 5707–5715 (2009).
Zabeck, H. et al. Molecular signatures in IASLC/ATS/ERS classified growth patterns of lung adenocarcinoma. PLoS One 13, e0206132 (2018).
Somlyo, A. P. & Somlyo, A. V. Ca2+ sensitivity of smooth muscle and nonmuscle myosin II: Modulated by G proteins, kinases, and myosin phosphatase. Physiol. Rev. 83, 1325–1358 (2003).
Yu, W. et al. Involvement of RhoA, ROCK I and myosin II in inverted orientation of epithelial polarity. EMBO Rep. 9, 923–929 (2008).
Vito, A., El-Sayes, N. & Mossman, K. Hypoxia-driven immune escape in the tumor microenvironment. Cells 9, 992 (2020).
Wu, Q. et al. Hypoxia-inducible factors: Master regulators of hypoxic tumor immune escape. J. Hematol. Oncol. 15, 77 (2022).
Vitale, M. & Parodi, M. Blocking HIF to enhance NK cells: Hints for new anti-tumor therapeutic strategies?. Vaccines (Basel) 9, 1144 (2021).
Salinas-Vera, Y. M. et al. Three-dimensional 3D culture models in gynecological and breast cancer research. Front. Oncol. 12, 826113 (2022).
Jubelin, C. et al. Three-dimensional in vitro culture models in oncology research. Cell Biosci. 12, 155 (2022).
Bryant, D. M. & Mostov, K. E. From cells to organs: Building polarized tissue. Nat. Rev. Mol. Cell Biol. 9, 887–901 (2008).
Yu, W. et al. β1-integrin orients epithelial polarity via Rac1 and laminin. Mol. Biol. Cell 16, 433–445 (2005).
Bryant, D. M. et al. A molecular switch for the orientation of epithelial cell polarization. Dev. Cell 31, 171–187 (2014).
Liu, K. D. et al. Rac1 is required for reorientation of polarity and lumen formation through a PI 3-kinase-dependent pathway. Am. J. Physiol. Renal Physiol. 293, F1633–F1640 (2007).
Campbell, F. C. et al. Mechanistic insights into colorectal cancer phenomics from fundamental and organotypic model studies. Am. J. Pathol. 188, 1936–1948 (2018).
Jin, M. Z. & Jin, W. L. The updated landscape of tumor microenvironment and drug repurposing. Signal Transduct. Target Ther. 5, 166 (2020).
Petrova, V., Annicchiarico-Petruzzelli, M., Melino, G. & Amelio, I. The hypoxic tumour microenvironment. Oncogenesis 7, 10 (2018).
Solis, L. M. et al. Histologic patterns and molecular characteristics of lung adenocarcinoma associated with clinical outcome. Cancer 118, 2889–2899 (2012).
Cadioli, A. et al. Lung cancer histologic and immunohistochemical heterogeneity in the era of molecular therapies: analysis of 172 consecutive surgically resected, entirely sampled pulmonary carcinomas. Am. J. Surg. Pathol. 38, 502–509 (2014).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: The next generation. Cell 144, 646–674 (2011).
Gonzalez, H., Hagerling, C. & Werb, Z. Roles of the immune system in cancer: From tumor initiation to metastatic progression. Genes Dev. 32, 1267–1284 (2018).
Absi, A. A. et al. Actin cytoskeleton remodeling drives breast cancer cell escape from natural killer-mediated cytotoxicity. Cancer Res. 78, 5631–5643 (2018).
Acknowledgements
We thank Mr. Masaharu Kohara, Ms. Takako Sawamura, Ms. Megumi Nihei, and Ms. Etsuko Maeno-Fujinami from the Department of Pathology, Osaka University Graduate School of Medicine, for their technical assistance. This study was supported by Mr. Eiji Oiki, Center for Medical Research and Education, Graduate School of Medicine, Osaka University.
Funding
This research was supported by by JSPS KAKENHI (grant numbers A19H034520, T22K194330, A23H027000, T21K069030, T21K070980, and T24K102850) and AMED (JP21ae0121049).
Author information
Authors and Affiliations
Contributions
D.U. conducted most of the experiments and data analyses. D.U., S.N., and A.H. wrote the manuscript. D.M. and D.O. also performed RNA-seq. A.H. analyzed the RNA-seq data. Y.S. contributed to collection of clinical samples. S.T., M.K., K.K., T.T., H.K., K.U., T.M., S.M., Y.S., and A.K. provided advice regarding project planning, experimental techniques, and data interpretation. D.U., S.N., and E.M. designed the experiments. E.M. supervised the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
All experimental protocols involving human specimens were approved by the Ethics Review Board of the Graduate School of Medicine, Osaka University (approval no. 15234) and carried out in accordance with the committee’s guidelines and regulations. The study was performed in accordance with the Declaration of Helsinki.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Umeda, D., Harada, A., Motooka, D. et al. Hypoxia drives the formation of lung micropapillary adenocarcinoma-like structure through hypoxia-inducible factor-1α. Sci Rep 14, 31642 (2024). https://doi.org/10.1038/s41598-024-80280-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-80280-x