Abstract
The human-brain is a vital and complicated organ within the body. Identifying brain-related diseases can be challenging. Typically, Magnetic Resonance Imaging (MRI) scanning methods are used to gain insights of the protected regions in the body. Brain segmentation can result in identifying region boundaries as a set of contours. However, segmenting brain images poses several challenges, including noise, bias field, and partial volume effect (PVE). Removing noise, accurately segmenting tissues and tumors are crucial for effective evaluation. To enhance tissue and tumor segmentation, a new machine learning-based method called as Gaussian-Kernelized Enhanced Intuitionistic Fuzzy-C-Means (GKEIFCM) has been proposed. Approach enhances Improved Intuitionistic Fuzzy-C-Means Algorithm (IIFCM) by utilizing Gaussian kernelized distance between pixels, resulting in uncomplicated segmentation with reduced computational times and improved efficiency. This proposed novel method proved to be expertise in tissue and tumor classification and identification respectively. The results demonstrate the effectiveness of GKEIFCM interms of Dice, Jaccard-similarity-index, Accuracy and Execution time.
Similar content being viewed by others
Introduction
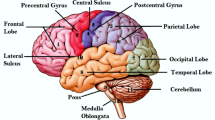
The MR image of a human-brain can be primarily segmented into three tissues. They are Cerebrospinal-Fluid(CSF), Gray-Matter(GM) and White-Matter(WM). As mentioned before, MRI segmentation helps radiologist or doctor to accurately identify various disease tumors, brain related disorders and cause of a particular headache etc. As it is well-known, it is difficult to find definite boundaries between various tissues in human brain. Segmentation of an image is to extract several objects. Segmentation plays a vibrant role in a huge number of biomedical applications like detection of disease, surgery and treatment planning1. This task is more challenging in processing the medical images due to different shapes, locations, and intensities of the tissues.
In medical imaging, segmentation is the prerequisite and treated as the initial step in quantifying the analysis. These days physicians consider using MRI image segmentation techniques to correctly diagnose many dreadful disorders like tumors and diseases like Alzheimer’s, edema, dementia and schizophrenia etc. to save time and life of humans. MRI image segmentation is a challenging-task attributed to various noises present in MRI image acquisition process. Many segmentation algorithms were developed for this purpose. The extraction of quantitative and qualitative information about tissues is comprised of noise in the images during the process of acquisition. In the literature, various methods are introduced to overcome the difficulties associated with the MRI segmentation process. Among them thresholding, region, edge, deformable model based, and classification methods are classical segmentation methods.
Explicitly, classification approaches are divided into supervised and unsupervised. Artificial-Neural-Network (ANN), Support-Vector-Machine (SVM), Bayes-Classifiers1 are the examples of supervised methods whereas clustering algorithms (CA) such as Markov-Random-Fields (MRF), Expectation-Maximization (EM method), fuzzy-clustering, K-Means, and atlas-based segmentations2 are the examples of unsupervised methods. Clustering-based algorithms are most popularly used and they are the best example of unsupervised learning. In this, data is grouped with the existing data by finding similarities. Clustering is very famous in the fields: data science, Artificial-Intelligence3, pattern recognition4 etc.
This paper organizes as follows: Firstly, Section-2 briefs literature review associated to our work, Next, Section-3 deals with description of algorithm and other segmentation algorithms used for comparison. Later, Section-4 demonstrates results. Finally, Section-5 concludes paper.
Related works
In the literature, researchers suggested denoising methods to suppress the noise of an image without the degradation of the attributes of the original image. To reduce the noise from MR images, numerous denoising filters like bilateral, PCA, non-local means and bilateral can be utilized. Denoising filter analysis is carried out using various denoising techniques and revealed that the Spatially Adaptive Non-Local Means filter gives finest results than existing ones5. In general, CA are classified into two types: hard and fuzzy6. The goal of hard clustering is to subdivide the dataset for instance, Y into non-overlapping non-empty subsets 1 … n. Whereas, the goal of the fuzzy clustering is to subdivide the dataset Y into divisions, in order to allow the data object to fit into a specific unit (probably null) for each and every single fuzzy cluster. The robust FCM algorithm7,8 is proposed with the modification in objective function of the conventional FCM, including the local-spatial term allowing the computation of the smooth membership-grade. It improves the segmentation process and it’s in-sensitive to noise only till a certain level. By considering an additional term in the objective function, the fuzzy clustering with spatial constraints9 is proposed to allow smoothing of pixels by its neighboring pixels to overcome the difficulty of the intensity in-homogeneity. This process is not noise sensitive, however it consumes more run time as it encompasses the computation of neighboring pixels in complete iterations.
The FCM-S1 and FCM-S210,11 are suggested to reduce the execution time of the FCM-S algorithm. FCM-S1 and FCM-S2 methods are used to compute mean and median filtered images to replace neighborhood pixels of the FCM-S algorithm. An improved fuzzy c-means (Im-FCM) proposed12, in acquaintance with the neighborhood desirability term with its distance-measure, depending on two factors: (i) pixel intensities, (ii) spatial location of the adjacent pixels. The Im-FCM uses two parameters, whose finest values are acquired using ANN to amend the degree of two factors which needs more run time to acquire the parameter. The fast-generalized fuzzy-c-means (FGFCM)13 combines the grey level as well as local spatial information using a similarity measure factor. The fuzzy local information c-means (FLICM) algorithm14 stated that by adding a new fuzzy local neighborhood factor in the objective function, the intensity-level and neighborhood relationship in the spatial domain can be found and parameter setting can be avoided. However, images are treated as fuzzy because of ambiguity resulting in classification, with regard to areas, boundaries and imperfect grey-levels.
The fuzzy clustering is most often considered in-case of partial membership clusters of an element. In image segmentation, clustering considers image pixel as the data object, especially every pixel is assigned to a cluster depending on its resemblance of prominent features15. While discussing the ambiguity, indecision arises in the image while defining the membership function in the hybrid algorithms16,17. Ever since the membership grades are inaccurate and varies on individual’s options, there are several kinds of uncertainties to an extent, arising due to absence of well-defined information in outlining the membership function. Leads to a definition of higher order fuzzy-sets, referred as Intuitionistic-Fuzzy-Set (IFS) theory proposed18, which considers the membership and the non-membership grades also. In IFS, because of hesitation degree19, the non-complement of the membership grade is greater than the membership grade. Many methodologies were introduced to mitigate the disadvantages of FCM. The basics of optimal sets is presented in20 where the number of sets are not defined, by using the Shannon’s entropy a standard function is hosted to capitalize the good points in the class.
The type 2 fuzzy clustering is proposed21, with the ambiguity in a fuzzy set Type 2 membership by giving triangular membership grades for Type 1 fuzzy. The clustering technique22,23,24,25 is defined in which an intuitionistic fuzzy similarity matrix is transfigured to interval valued fuzzy equivalence matrix, depending on the λ-cutting matrix of the intuitionistic fuzzy equivalence matrix. Clustering is performed using the λ-cutting matrix of the related to association equivalence matrix. Hence, there is a need to develop an algorithm which solves this issue.
Proposed methodology
The primary problem in brain MR image segmentation is to control the uncertainty which arises at boundaries among various brain and non-brain tissues along with the tumors. Highlights of the proposed algorithm are as follows:
-
(I)
Gaussian kernelized distance metric is used between pixels, which reduces the number of iterations to segment the tissues thus improves the dice and Jaccard similarity values and execution times are less.
-
(II)
Employed IIFCM algorithm with Gaussian kernelized distance improves the accuracy in detecting the tumors.
-
(III)
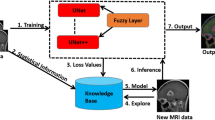
The spatial distance between the pixels is reduced with the proposed novel GKEIFCM algorithm, thus leads to minimization of the objective function. Figure 1 shows the block diagram of the proposed work.
Block diagram of the proposed method.
First, the MRI image is considered as input and skull stripping is performed on slice number 90 as a pre-processing step as shown in Fig. 2. On the skull stripped image, the new proposed Gaussian Kernelized Enhanced intuitionistic FCM (GKEIFCM) algorithm is applied for segmentation. The resultant clustering outputs are contrasted with the original images by using quantitative metrics to estimate the performance of the proposed method. The GKEIFCM method is proposed from IIFCM, which in turn developed from the IFCM. By the hybridization of IFCM, IIFCM methods, a novel GKEIFCM algorithm is proposed.
(a) Input MRI image slice 90, (b) Skull stripped image.
Segmentation of intuitionistic fuzzy set
In segmentation, intuitionistic fuzzy data will be used for intuitionistic fuzzification and IFS is the representation for the image. Let us consider A is the image with \(\:MxN\), L is the number of grey levels with \(\:0\:to\:L-1\) varying range. Image \(\:Y\:\)with regard to IFS13 is represented as \(\:A=\left\{\left(sj\:,\mu\:Y\left(sj\right),\:vY\left(sj\right)\right)\right\},\:j=\text{1,2},\dots\:,P\) where \(\:\mu\:Y\left(sj\right),\:vY\left(sj\right)\) are membership \(\:{(s}_{j})\:\)and non-membership degrees of a pixel \(\:{s}_{j}.\:\) This \(\:{s}_{j}\) is given with regard to normalised intensity-level (L) as:
where, \(\:j=\text{1,2},\dots\:,N,\)\(\:{s}_{j}Є\{0,\dots\:.L-1\},\:{\left({s}_{j}\right)}_{max}\:\:and\:{\left({s}_{j}\right)}_{min}\) are maximum and minimum grey-levels. Sugeno’s-negation function is also used to authenticate the non-membership-degree which is referred to as:
Therefore, IFS is given as:
Where, \(\:\mu\:Y\) and \(\:vY\) are the membership and non-membership degrees of a pixel \(\:s\).
Proposed methodology: Gaussian kernelized enhanced intuitionistic Fuzzy-C-means clustering technique (GKEIFCM)
The Euclidean distance metric used by the IFCM and IIFCM algorithms26, is usually applicable to the hypothesis of un-correlated segments as well as segments with spherical shape. Here, the Gaussian kernelized metric is used as a replacement to the Euclidian distance in the IIFCM algorithm. To increase the segmentation performance and the robustness w.r.to outliers, the Euclidean-distance is replaced with the kernel \(\:{∥\phi\:\left({x}_{i}\right)-\phi\:\left({v}_{j}\right)∥}^{2}\). The kernelized distance is simplified as:
where,
Hence by replacing the Euclidean-distance with Gaussian kernel-distance, we will have the new objective function as:
The kernelized distance, which minizines the objective-function of the intuitionistic FCM algorithm. This leads to a smaller number of iteration and faster in computations and lesser execution times.
The centroids and objective functions are changed accordingly as shown in Eq. (7). where, U is the membership matrix and \(\:{u}_{ij}\) denotes membership degree. Lagrangian method is used in literature to minimize the objective function. On simplification, we get:
To find \(\:{K}_{ij}\), similarity measure among two elements \(\:xi,\:xj\in\:S\) in IFS \(\:{D}_{k}\) is expressed as:
where,
The intuitionistic fuzzy factor is also changed as:
The flow chart of GKIIFCM algorithm is shown in Fig. 3, which explains the step-by-step procedure.
Flow chart of proposed GKIIFCM algorithm.
Simulation results
The Proposed method exhibits both qualitative and quantitative analysis verified on real world tumor and non- tumor images. In addition, the case studies are also verified at Vijaya Diagnostic Centre, Hyderabad, Telangana, India. To assess the efficacy of our method, the experiments are conducted on MRI brain images containing 3%, 5% and 7% noises [http://www.bic.mni.mcgill.ca/brainweb/], tumor images [https://www.med.harvard.edu/aanlib/] and Ground Truth (GT) images are obtained with the help of a radiologist. MRI slices are chosen based on the mean slice, Slice selection is attained by applying a 1-Dimensional, linear magnetic field gradient during the period that the RF pulse is applied. System specifications are mentioned in results section. The algorithm is implemented in MATLAB 2022a and run on Intel core i5 processor, 2.50 GHz with 32G RAM and Windows8 operating system. The obtained segmented outputs of MR images are evaluated against the GT images consolidated by the radiologists. The similarities between the GT and resultant segmented MRI brain images are calculated using performance metrics such as Dice-Similarity-Index (DSI) and Jaccard-Similarity-Index (JSI). The DSI and JSI27 between A and B images computed with the help of the formulae as in Eq. 12.
Here, A is GT image and B is the segmented output image obtained by the proposed and existing methods. Better segmented outputs will have better dice similarity and Jaccard indices values. The values range from 0 to 1. Verification of all the algorithms for 55 iterations is considered in addition to inputs with different noise percentages. Segmentation results can be obtained in terms of similarity indices, execution times and accuracies.
Segmentation of tissues in MRI brain images
The obtained results are verified for various MRI slice numbers 60, 65, 70, 75, 80, 85, 90, 95, 100, 105, 110,115, 120,125 respectively. The input and skull stripped images are shown in Fig. 2 for the slice number 90. This data-set includes a variety of noise levels (0–9%), T1-weighted images with various noise levels are used. The volume 1-mm slice thickness, 1 × 1 × 1 mm3 as voxel-size and voxel-resolution of 1-mm3, images are of dimension 181 × 217 × 181 voxels are used. The results are shown for the segmented images with 3% noise in MRI images for slice number 90 in Figs. 4 and 5 for grey matter and white matter respectively.
Segmentation results of GM tissue.
The figures show the comparison of obtained results with existing methods and proposed method. The existing methods for comparison with the proposed GKEIFCM unsupervised machine learning algorithm considered are FCM, FCM-S1 and S2, FLICM, FGFCM, IFCM and IIFCM.
Segmentation results of WM tissue.
Grey and white matters of the images as shown in Figs. 4 and 5 are segmented using different algorithms and the performance metrics are presented in the Tables 1, 2, 3, 4, 5, 6 and 7. Table 1 displays the dice similarity and Jaccard similarity indices of input slice 90 with 3% noise. From the Table 1, it is clearly indicated that the dice similarity and Jaccard coefficients of the grey matter tissue are 98.43% and 92.72% respectively for the proposed method, whereas for the conventional FCM method, they are recorded as 81.64% and 68.97% respectively. From the outcomes, we can clearly state that proposed method provides better tissue segmentation results with an improvement of 20.10% and 34.4%.
Table 2 displays DSI and JSI of input slice 90 with 5% noise and Table 3 displays DSI and JSI of input slice 90 with 7% noise respectively from the analysis of the tabular columns the proposed GKEIFCM gives the best performance values in-terms of Dice and Jaccard indices measures.
Dice similarity index comparison grey matter for slice 90 with 3%, 5% and 7% of noises.
The graphical representation of DSI for noises 3%, 5% and 7% are shown in Fig. 6. Indicates the improvement of results for DSI metric is 20.04% when compared to the FCM_S1 and the proposed GKEIFCM method for 3% noise. The kernel distance metric used in proposed method leads to better results when compared to conventional methods with Euclidean distance metric utilization.
The graphical representation of JSI for noises 3%, 5% and 7% are shown in the Fig. 7. Indicates that the proposed method shows 35.02% improvement on results of dice similarity index when compared to conventional FLICM method and the proposed GKEIFCM method for 7% noise. The kernel-distance metric is used in proposed method to minimize objective function which leads to decrease in number of iteration comparability with conventional methods.
Jaccard index comparison grey matter for slice 90 with 3%, 5% and 7% of noises.
From the results, the GKEIFCM algorithm provides the highest dice and Jaccard indices compared to other algorithms and the indices keep on decreasing as the noise level in the MRI increases. Tables 4, 5 and 6 show the similarity indices for slice no. 60 with noise percentages 3%, 5%, and 7% respectively. For the 60th slice as well, it is observed that the GKEIFCM algorithm is giving the improved segmentation results.
Table 4 displays the dice similarity and Jaccard similarity indices of input slice 60 with 3% noise. From the tabulations, it is clearly indicated that the dice similarity coefficients of the grey matter and white matter tissue are 98.13% and 98.34% respectively for the proposed method, whereas for the conventional FCM method, they are recorded as 89.64% and 92.96% respectively. We can clearly state from the results that proposed GKEIFCM algorithm provides better tissue segmentation results.
From the Table 5, it is clearly indicated that the dice similarity and Jaccard coefficients of the white matter tissue are 97.34% and 93.29% respectively for the proposed method, whereas for the FLICM method, they are recorded as 88.90% and 80.02% respectively. Obtained results clearly state that the proposed GKEIFCM algorithm provides better tissue segmentation results for both grey matter and white matter for the performance measures of Dice and Jaccard indices.
Table 6 displays the dice similarity and Jaccard similarity indices of input slice 60 with 7% noise. From the tabulations, it is clearly indicated that the dice similarity coefficients of the grey matter and white matter tissue are 88.95% and 97.63% respectively for the proposed method, whereas for the FGFCM method, provides 83.26% and FLICM gives 87.90% respectively. With these results, we can clearly state that the proposed GKEIFCM algorithm provides better tissue segmentation results in both dice and Jaccard similarity indices.
Segmentation of tumor using GKEIFCM algorithm
In this section, the results are shown for the segmented tumor from tumor images using the proposed GKEIFCM algorithm are shown in Fig. 8 and their accuracies are calculated.
Segmentation of tumors with accuracies in MRI brain images for the proposed GKEIFCM method.
Table 7 shows the results of accuracy for the segmented tumors. It can be seen that FCM_S1 method produces 92.83% and 92.54% accuracies noted as least value for 3%, 7% of noises respectively, and FLICM method results least for 5% of noise with 91.77 when compared to the proposed GKEIFCM algorithm.
Comparison of execution Times
The execution time of the implemented algorithms and the proposed GKEIFCM are presented in Table 8. It can be clearly observed that the GKEIFCM algorithm takes less computational time for segmentation of the tissues (GM and WM) and tumors with an average of 9.86 s, when compared to the FCM-S1 algorithm on-average of 91.88 s execution time and other algorithms.
The conventional FCM method takes less execution time but lacks in providing the good dice similarity and Jaccard indices in tissue segmentation and accuracy values in tumor extraction.
The proposed method assist radiologists and oncologists by reducing the manual workload involved in tumor delineation, which is time-intensive and prone to variability across practitioners. By automating this process with a high degree of accuracy, our approach may increase diagnostic efficiency, allowing clinicians to focus more on complex decision-making and patient care.
Conclusion
The effectiveness of MRI brain tissue and tumor segmentation is crucial for the exact diagnosis and treatment of neurological diseases. In this study, a novel machine learning method called Gaussian kernelized improved intuitionistic FCM algorithm (GKEIFCM) was proposed to improve the accuracy of segmentation. This algorithm uses a Gaussian kernelized distance metric to decrease the spatial distance among the pixels and enhance the accuracy of segmentation but lacks to achieve the prominent results for large datasets. Results showed that GKEIFCM outperformed other commonly used techniques, such as FCM, FCM with spatial functions S1 and S2, FLICM, IFCM, and IIFCM, in terms of accuracy, DSI, JSI metrics. Specifically, GKEIFCM achieved an accuracy of 98.54%, a DSI of 98.37%, and a JSI of 96.68% with only 3% noise. The proposed hybrid GKEIFCM segmentation algorithm was found to be robust to noise artefacts and can accurately locate tumors in affected regions. These findings suggest that GKEIFCM is a promising technique for segmentation. It can be furtherly combined with advanced deep learning techniques for better results for even large datasets.
Data availability
The datasets used and/or analysed during the current study available from the corresponding author on reasonable request.
Abbreviations
- MRI:
-
Magnetic-resonance-imaging
- PVE:
-
Partial-volume-effect
- GKEIFCM:
-
Gaussian-kernelized enhanced intuitionistic fuzzy-C-means
- FCM:
-
Fuzzy-C-means
- CSF:
-
Cerebrospinal-fluid
- GM:
-
Gray-matter
- WM:
-
White-matter
- ANN:
-
Artificial-neural-network
- SVM:
-
Support-vector-machine
- MRF:
-
Markov-random-fields
- EM method:
-
Expectation-maximization
- Im-FCM:
-
Improved fuzzy c-means
- FGFCM:
-
Fast-generalized fuzzy-c-means
- FLICM:
-
Fuzzy local information c-means
- IFS:
-
Intuitionistic fuzzy set theory
References
Schmarje, L., Santarossa, M., Schroder, S. M. & Koch, R. A survey on semi-, self-and unsupervised learning for image classification. IEEE Access. 9, 82146–82168 (2021).
Lavanya, K. G., Dhanalakshmi, P. & Nandhini, M. Computerized segmentation of MR brain tumor: An integrated approach of multi-modal fusion and unsupervised clustering. Int. J. Inform. Technol. 1–15. (2023).
Gunasekara, S., Ramesh, H. N. T. K., Kaldera & Maheshi, B. Dissanayake. A systematic approach for MRI brain tumor localization and segmentation using deep learning and active contouring. J. Healthcare Eng. (2021).
Jain, A. K., Narasimha Murty, M. & Patrick, J. Flynn. Data clustering: A review. ACM Comput. Surveys (CSUR) 31(3), 264–323. (1999).
Jabbar, H., Hatem, R. M., Muttasher & Ali Fattah, D. Segmentation of brain tissue using improved kernelized rough-fuzzy c-means technique. Indonesian J. Electr. Eng. Comput. Sci. 32 (1), 216–226 (2023).
Ouchicha, C., Ammor, O. & Meknassi, M. A new approach based on exponential entropy with modified kernel fuzzy c-means clustering for MRI brain segmentation. Evol. Intel. 16 (2), 651–665 (2023).
Yang, R. & Li, D. Adaptive wavelet transform based on artificial fish swarm optimization and fuzzy C-means method for noisy image segmentation. Comput. Sci. Inform. Syst. 00, 39–39 (2022).
Pham, D. L. Spatial models for fuzzy clustering. Computer vision and image understanding 84(2), 285–297. (2001).
Ahmed, M. N. et al. A modified fuzzy c-means algorithm for bias field estimation and segmentation of MRI data. IEEE Trans. Med. Imag. 21(3), 193–199 (2002).
Wu, C., Huang, C. & Zhang, J. Local Information-Driven Intuitionistic Fuzzy C-Means Algorithm Integrating Total Bregman Divergence and Kernel Metric for Noisy Image Segmentation Circuits Syst. Signal. Process. 42(3), 1522–1572 (2023).
Chen, S. & Zhang, D. Robust image segmentation using FCM with spatial constraints based on new kernel-induced distance measure. IEEE Trans. Syst. Man. Cybernetics Part. B (Cybernetics). 34 (4), 1907–1916 (2004).
Shen, S., Sandham, W., Granat, M. & Sterr, A. MRI fuzzy segmentation of brain tissue using neighborhood attraction with neural-network optimization. IEEE Trans. Inf Technol. Biomed. 9 (3), 459–467 (2005).
Cai, W., Chen, S. & Zhang, D. Fast and robust fuzzy c-means clustering algorithms incorporating local information for image segmentation. Pattern Recog. 40(3), 825–838. (2007).
Krinidis, S. & Chatzis, V. A robust fuzzy local information C-means clustering algorithm. IEEE Trans. Image Process. 19(5), 1328–1337. (2010).
Ouchicha, C., Ammor, O. & Meknassi, M. A new approach based on exponential entropy with modified kernel fuzzy c-means clustering for MRI brain segmentation. Evol. Intel. 1–15. (2022).
Arulselvarani, S. & Manimekalai, S. Brain tumor segmentation & detection of Mr images using intuitionistic fuzzy clustering mean (IFCM). Ann. Roman. Soc. Cell Biol. 25(3), 7669–7680 (2021).
Ray, M., Mahata, N. & Jamuna Kanta Sing. Uncertainty parameter weighted entropy-based fuzzy c-means algorithm using complemented membership functions for noisy volumetric brain MR image segmentation. Biomed. Signal Process. Control. 85, 104925 (2023).
Atanassov, K. Intuitionistic fuzzy sets. Int. J. Bioautomation. 20, 1 (2016).
Aruna Kumar, S. V., Yaghoubi, E. & Proença, H. A fuzzy Consensus Clustering Algorithm for MRI Brain tissue segmentation. Appl. Sci. 12, 15 (2022).
Ferahta, N. New fuzzy clustering algorithm applied to RMN segmentation. IEEE Trans. Eng. Comput. Technol. 12, 9–13 (2006).
Rhee, F. C. H. & Hwang, C. A type-2 fuzzy C-means clustering algorithm. In Proceedings joint 9th IFSA world congress and 20th NAFIPS international conference (Cat. No. 01TH8569), vol. 4, pp. 1926–1929. IEEE, (2001).
Zhang, H., Xu, Z. & Chen, Q. Clustering Approach Intuitionistic Fuzzy sets Control Decis. 22, 8 : 882. (2007).
Kala, R. & Deepa, P. Spatial rough intuitionistic fuzzy C-means clustering for MRI segmentation. Neural Process. Lett. 53 (2), 1305–1353 (2021).
Dahiya, S. & Gosain, A. A novel type-II intuitionistic fuzzy clustering algorithm for mammograms segmentation. J. Ambient Intell. Humaniz. Comput. : 1–16. (2022).
Xu, Z., Chen, J. & Wu, J. Clustering Algorithm Intuitionistic Fuzzy sets. Inform. Sci. 178, 19 : 3775–3790. (2008).
Pelekis, N., Iakovidis, D. K., Kotsifakos, E. E. & Kopanakis, I. Fuzzy clustering intuitionistic fuzzy data. Int. J. Bus. Intell. data Min. 3, 1 : 45–65. (2008).
Rusinek, H. et al. Manuel Mourino, and Henryk Kowalski. Alzheimer disease: Measuring loss of cerebral gray matter with MR imaging. Radiology 178, 1 : 109–114. (1991).
Acknowledgements
The authors extend their appreciation to the Zhejiang Provincial Natural Science Foundation Youth Fund Project (Grant No. LQ23F010004), the National Natural Science Youth Science Foundation Project (Grant No. 62201508), and to the King Saud University (KSU) for funding this work through researchers supporting project number (RSP2024R387), King Saud University (KSU), Riyadh, Saudi Arabia.
Funding
The authors extend their appreciation to the Zhejiang Provincial Natural Science Foundation Youth Fund Project (Grant No. LQ23F010004), the National Natural Science Youth Science Foundation Project (Grant No. 62201508), and to the King Saud University (KSU) for funding this work through researchers supporting project number (RSP2024R387), King Saud University (KSU), Riyadh, Saudi Arabia.
Author information
Authors and Affiliations
Contributions
All authors contributed equally to the conceptualization, formal analysis, investigation, methodology, and writing and editing of the original draft. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Saladi, S., Yepuganti, K., Chinthaginjala, R. et al. A unique unsupervised enhanced intuitionistic fuzzy C-means for MR brain tissue segmentation. Sci Rep 14, 29804 (2024). https://doi.org/10.1038/s41598-024-81648-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-81648-9