Abstract
This study aimed to explore the causal relationships between multiple blood amino acids (BAAs) and the Parkinson’s disease (PD). We downloaded genome-wide association study (GWAS) data for BAAs and PD from the OpenGWAS database, screened single nucleotide polymorphisms (SNPs) from the data, and evaluated the causal relationship between BAA levels and PD using the inverse variance weighted (IVW) method. The sensitivity analysis was also conducted. After SNP screening, three amino acid indicators were identified: met-a-308 (phenylalanine), met-a-584 (X-12100 hydroxytryptophan), and met-a-337 (5-hydroxyproline), which showed significant causal relationship with the occurrence of PD. There was no significant heterogeneity or horizontal pleiotropy, and the results were stable. The multivariate MR analysis showed that the mediating effects generated by the introduction of multiple variables were not significant. In conclusion, phenylalanine, X-12,100 hydroxytryptophan, and 5-hydroxyproline have a causal relationship with the occurrence of PD and may be potential early screening biomarkers and blocking targets.
Similar content being viewed by others
Introduction
Parkinson’s disease (PD) is a common degenerative disease of the nervous system that is clinically characterized by static tremors, motor delays, muscle rigidity, and postural balance disorders1,2. PD has a high incidence rate, high disability rate, and poor prognosis in middle-aged and older individuals3,4. A 2021 meta-analysis5 of studies from China included 24,117 people aged > 60 years and found that the prevalence of PD in the older population was 1.37% (95% confidence interval [CI] 1.02–1.73%). PD is a chronic progressive disease whose incidence increases with age, and there are currently no fully cured cases. As patients with PD experience damage to the motor function, those with late-stage disease lose their ability to take care of themselves, have difficulty moving, and often die from complications such as respiratory or circulatory failure6,7. Together, these factors seriously affect the quality of life of patients and increase the burden on their families and society8,9. Therefore, it is necessary to identify risk factors for PD in the early stages and initiate treatment as soon as possible to delay disease progression.
The main pathological feature of PD is extensive loss of dopaminergic neurons in the substantia nigra, which is associated with enhanced intracellular oxidative stress and mitochondrial dysfunction10,11. A previous pathological report12 confirmed that the concentration of glutathione in the substantia nigra, which has antioxidant effects, decreases, whereas that of lipid peroxides increases. No such changes were observed in other parts of the brain. Therefore, the metabolism of cysteine, glutamic acid, and aminoacetic acid is closely associated with oxidative stress in the substantia nigra13,14. These amino acids also participate in signal transmission as neurotransmitters, and any change leading to an imbalance in neurotransmitters could alter the metabolic pathways in the brain, potentially leading to diseases of the central nervous system. Another study15 suggested that homocysteine directly damages neurons and participates in the degeneration of dopaminergic neurons in patients with PD. In a cohort study by Figura et al.16, the serum amino acid metabolism of patients with PD at different stages of progression was compared, and significant differences were found between groups for alanine, arginine, phenylalanine, and threonine, indicating that changes in the metabolism of these amino acids are related to the occurrence and progression of PD. However, there is currently no consensus on which amino acids independently affect the development of PD.
The expression levels of blood amino acids (BAAs) show dynamic fluctuations during synthesis and decomposition, which are related to various factors (including genetics)17,18. Wiklund et al.19 reported that the serum leucine and isoleucine levels in children can be used to predict hypertriglyceridemia in early adulthood and serve as markers to identify populations at high risk for cardiovascular disease. A study by Hu et al.20 suggested that PPM1K rs1440581 and rs7678928 single nucleotide polymorphisms (SNPs) are associated with elevated levels of serum branched chain amino acids, which could increase the risk of cardiovascular disease. Because of the known genetic pattern of randomly assigning parental alleles to offspring, some genotypes can be used to determine the serum amino acid phenotype and infer the relationship between the phenotype and disease occurrence21,22,23. However, research on this causal relationship is still very rare at present.
In this study, we performed a double-sample genome-wide association study (GWAS) of BAAs and PD to analyze key SNPs and conducted a Mendelian randomization (MR) study to determine the impact of the expression levels of specific BAA on the pathogenesis of PD in search of biomarkers and blocking targets in the early stages of PD.
Results
Roadmap of the analysis process
The roadmap is shown in Fig. 1.
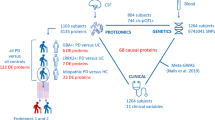
A. Schematic diagram of multivariate Mendelian randomization and mediation analysis. The basic assumptions of Mendelian randomization analysis include (1) the assumption of relevance, which states that the selected instrumental variable must be significantly correlated with the exposure factor; (2) The assumption of independence, which states that instrumental variables must have no significant correlation with potential confounding factors that may affect exposure or outcome; (3) Exclusivity limitation means that instrumental variables can only affect outcomes through the path of “instrumental variables → exposure → outcome”. B. The analytic process in this study. (SNP, single nucleotide polymorphism. IVW, inverse variance weighted. MR, Mendelian randomization. GWAS, genome-wide association study. LD, Linkage disequilibrium.)
Instrumental variable screening
SNP sites related to 86 BAAs were selected from the compiled SNP data for preliminary causal analysis. After matching the GWAS data of PD (finn-b-G6_PARKINSON), the effect values of all instrumental variables were obtained. After normalization, the instrumental variables associated with BAAs and PD were obtained and included in the MR analysis. The specific instrumental variable data for the BAA exposure factors are shown in Table 1, which includes only indicators with significance (P < 0.05). The F-test statistics of the instrumental variables were all > 10, indicating that the SNPs selected in this study were mostly strong-effect instrumental variables with minimal bias.
MR causal effect estimation of the BAA index and PD
A total of five models were used for calculation, while only the inverse variance weighted (IVW) model results showed a significant causal relationship between the BAA indices, met-a-308 (phenylalanine) (odds ratio [OR] = 0.001, 95% confidence interval [CI]: 0.000–0.106, P = 0.00478), met-a-337 (5-hydroxyproline) (OR = 4.177, 95% CI: 1.125–15.518, P = 0.0327), and met-a-584 (X-12100 hydroxytryptophan) (OR = 0.276, 95% CI: 0.082–0.934, P = 0.0385), and occurrence of PD (Table 2). The MR results are presented in a forest map (Fig. 2a). Figure 2b–d shows the linear relationship between the instrumental variables of the BAA index and occurrence of PD. It was shown that the effect size of met-a-337 estimated by the simple model deviated significantly from that of the IVW model, and the effect sizes of met-a-308 and met-a-584 estimated by all the other four models were consistent with those of the IVW model.
Mendelian randomization analysis of the causal relationship between Blood amino acid index Iron and Parkinson’s disease (PD). (A). Blood amino acids (Blood amino acid index) of Parkinson’s disease (Parkinson 's diseases, the pathogenesis of PD) Mendelian randomization of the results of the analysis of forest graph display. (B) - d. Blood amino acids (Blood amino acid index) met - a - 308 (B), met - a - 337 (C), met - a - 584 (D) and Parkinson’s disease (Parkinson 's diseases, The results of Mendelian randomization analysis between PD) are shown in scatter plots. (E) - g. Blood amino acids (Blood amino acid index) met - a - 308 (E), met - a - 337 (F), met - a - 584 (G) and Parkinson’s disease (Parkinson 's diseases, Funnel plot showing the heterogeneity test of the results of Mendelian randomization analysis of causality between PD). (H)-(j). Blood amino acid index met-a-308 (H), met-a-337(I), met-a-584(J) and Parkinson’s disease (PD, P < 0.05). PD) using a single SNP loci analysis of the causal relationship between each effect of estimated results of forest.
Sensitivity analysis
Cochran’s Q-test was used to detect heterogeneity in the IVW model results. The P-values for met-a-308, met-a-337, and met-a-584 were 0.651, 0.545, and 0.636, respectively, all of which were > 0.05, indicating no significant heterogeneity (Table 3).
We also used the MR-Egger regression to test the variable-level pleiotropy of BAAs (met-a-308, met-a-337, and met-a-584). The test P-values for each indicator were > 0.05, and the intercept was close to 0, indicating that the causal effects of the BAA indices on the development of PD were not affected by horizontal pleiotropy (Table 4).
Funnel plots of instrumental variables for the BAA indices are shown in Fig. 2e–g. The scatter plot of met-a-308 (Fig. 2e) was not symmetrically distributed, indicating a potential bias in the results. The scatter plots of met-a–337 (Fig. 2f) and met-a-584 (Fig. 2g) were symmetrical, indicating that there was no potential bias in the results for the two indices.
Through leave-one-out analysis, each instrumental variable was removed individually to analyze the causal effects of met-a-308, met-a-337, and met-a-584 on the occurrence of PD (Fig. 2h–j). The total effects of the met-a-308, met-a-337, and met-a-584 instrumental variable sets did not show significant deviations.
Reverse causal assessment of the risk of PD on the BAA index
We used the same method to screen the SNP sites, and nine instrumental variables were included. A total of five models were used for the reversal analysis, and the IVW model results showed that there was no significant causal relationship between the occurrence of PD and BAA indices, met-a-308 (OR = 1.0069, 95% CI: 0.998–1.016, P = 0.127), met-a-337 (OR = 0.995, 95% CI: 0.977–1.013, P = 0.5616), and met-a-584 (OR = 1.005, 95% CI: 0.978–1.032, P = 0.7249). The occurrence of PD did not result in a decrease in the value of the BAA index (met-a-308, met-a-337, and met-a-584) (Table 5; Fig. 3).
Parkinson’s disease (Parkinson 's diseases, PD) and amino acid of Blood (Blood amino acid index) between Iron causality analysis of the Mendelian randomization. Forest plot of the results of Mendelian randomization analysis of Blood amino acid indexes met-a-308, met-a-337, met-a-584 for Parkinson’s disease (PD).
Multivariate MR analysis of the BAA index in the pathogenesis of PD
We first conducted univariate MR analysis of BAA (met a-308, met a-337, and met a-584) levels, and 61 PD-related exposure factors were identified (Table S1). The factors with a significant causal relationship (P < 0.05) with the BAAs were ukb-a-469, ukb-b-11,361, ukb-b-14,461, ukb-b-18,700, ukb-b-18,786, ukb-b-12,227, ukb-d-30270_irnt, ukb-d-1508_3, and ukb-d-2654_2. These factors were included in multivariate MR studies combined with the BAA (met a-308, met a-337, and met a-584) levels (Table 6). As shown in Table 6, we obtained nine multivariate MR models that met the requirements (all multivariate MR model results are shown in Table S4). The indirect effects of exposure factors were adjusted for in the nine multivariate MR models, and the results showed that the BAA (met-a-308, met-a-337, and met-a-584) levels had a significant and direct impact on the onset of PD.
Mediation effect analysis
Multivariate analysis revealed that all nine multivariate MR models had mediating effects of PD-related exposure factors on BAA levels with 95% CIs (0.501, 0.805, 0.434, 0.041, -0.042, -0.089, -0.111, -0.016, and − 0.182) (Table 7). However, the 95% CI relm of the mediating effects included 0; thus, they were not significant, indicating all these nine models showed no mediating effects.
Discussion
In this MR study, the levels of three BAAs were found to have a significant causal relationship with the development of PD. Among them, the high-expression levels of met-a-308 (phenylalanine) and met-a-584 (X-12100 hydroxytryptophan) could reduce the risk of PD and were protective factors, while that of met-a-337 (5-hydroxyproline) increased the risk of PD and was a risk factor. We also found that the levels of other amino acids, such as aspartic acid, serine, lysine, and cysteine, were not independent exposure factors that could affect the risk of developing PD.
Phenylalanine (chemical name 2-amino-3-phenylalanine) is an organic compound that is one of the essential amino acids in the human body. It is catalyzed by hydroxylases in the human body to form arginine, participates in the synthesis of neurotransmitters and hormones, and plays an important role in sugar and fat metabolism24,25,26. However, high concentrations in the cerebrospinal fluid can cause phenylketonuria due to neurotoxicity, ultimately leading to cognitive impairment. High concentrations of phenylalanine in the plasma may also affect the blood-brain barrier transport of other neutral amino acids, leading to a deficiency in other types of amino acids in the brain27. Phenylalanine may also be involved in the synthesis of serotonin and catecholamines, which affect the brain function28. Further, it is an independent protective factor against the development of PD; this was discovered for the first time in this study. Sanayama et al.29 reported a negative correlation between phenylalanine levels and glutathione peroxidase activity, indicating that higher phenylalanine levels led to lower oxidative stress responses in patients. Patients with PD are highly sensitive to oxidative stress; under a low oxidative stress response, the risk of developing PD is low30,31. However, the regulatory pathway of phenylalanine in the oxidative stress response remains unclear.
In this study, we found that high expression of X-12,100 hydroxytryptophan reduces the risk of PD while 5-hydroxyproline increases the risk of PD. This is consistent with the significant decrease in the TRP level and increase in the proline level in the serum of patients with PD32. TRP is an essential amino acid in the human body that is mainly metabolized via the kynurenine pathway. The main metabolite of kynurenine (KYN), kynurenic acid (KYNA), plays an important role in maintaining normal physiological functions of the central nervous system and has neuroprotective effects in neurodegenerative diseases such as PD, cerebral ischemia, stroke, and epilepsy33,34,35. Plasma-free TRP and branched-chain amino acids (leucine, isoleucine, and valine) participate in regulating the concentration of TRP in the brain. When the concentration of plasma-free TRP increases, the amount of TRP entering the brain increases and the activation of the KYN metabolic pathway in the brain is enhanced, thereby regulating the concentration of KYNA36. Notably, in this study, TRP (met-a-304) and c-glycosyltryptophan (met-a-502) were not independent influencing factors for PD and only X-12,100-hydroxytryptophan reduced the risk of PD.
Proline is an essential amino acid in the human body that undergoes hydroxylation by catalytic enzymes resulting in the formation of hydroxyproline37,38,39. It is an important component of collagen in animals. Hydroxyproline is also present in various plant proteins, particularly in cell walls40. Plewa et al.41 adopted a metabolomics approach and observed an increase in the serum concentration of hydroxyproline in patients with PD compared with healthy controls, which is consistent with the results of this study. These serological changes may be associated with the development of neurodegenerative diseases. Collagen contains a large amount of proline and hydroxyproline, and an increase in the hydroxyproline content in the serum is partly due to increased degradation of collagen, which might be caused by the activity of matrix metalloproteinases, which are considered potential pathogenic factors for PD42,43,44.
The heterogeneity detection results showed that the P-values of the IVW model for all three amino acids were > 0.05, indicating the absence of heterogeneity. Further, the MR-Egger regression analysis showed that the results of this study were not affected by horizontal pleiotropy; the scatter plot showed no significant bias, and the leave-one-out analysis showed no significant deviation in the results. We also used five models to reverse-deduce the effect of PD on BAAs and found no significant reverse causal relationship. We also introduced a multivariate analysis and found that the mediating effects in the multivariate MR model were not significant, indicating that external factors had no significant impact on the results of this study. Therefore, the results are stable and reliable.
In Cheng JY et al.'s study45, MR model was also built to predict the causal role of 9 amino acids on 6 neurodegenerative diseases, which was somewhat similar to this study in methodology. But in this study, 86 amino acids were included, and the target disease was only PD, with 5 regression models used for analysis, which is more reliable than their study using only IVW method. In this study, mediation effect analysis was also carried out to exclude the influence of exposure factors, strengthening the evidence levels for the influencing factors.
This study has some limitations. First, although the results of this study are stable and reliable, the underlying causes of the impact of the BAA indices on the pathogenesis of PD still need to be explored. Second, the results obtained in this study are all based on data analysis, and their effectiveness still needs to be verified in both in vivo and in vitro experiments.
In conclusion, the MR results in this study suggest a causal relationship between the three blood amino acid indices (phenylalanine, X-12100 hydroxytryptophan, and 5-hydroxyproline) and the risk of developing Parkinson’s disease. These findings indicate that alterations in these amino acids may play a role in the disease process, although further research is needed to clarify the mechanisms underlying these associations and their clinical implications.
Methods
Study design
This study used public datasets to investigate the impact of the BAA indices on the development of PD, including the mediating effects of other exposure factors. This study was conducted in accordance with the STROBE-MR Statement46.
Data sources
GWAS data on BAA levels from the OpenGWAS database (https://gwas.mrcieu.ac.hk/) were downloaded. The human blood metabolite (met-a) ID list provided by Shin et al.47 and circulating metabolite (met-a) ID list provided by Kettunen et al.48 were intersected, and only BAA-related IDs were retained. Finally, 86 BAA IDs were obtained (specific BAA IDs and names are listed in Table S2) and used for subsequent analyses. The data of these 86 BAAs were standardized and organized using the R package “TwoSampleMR”.
The GWAS data for PD (finn-b-G6_PARKINSON) were obtained from a meta-analysis49 of a Finnish database (https://www.finngen.fi/en/access_results), with data sourced from European ethnicities including both male and female individuals. We downloaded correlation summary statistical data for this analysis from the GWAS Catalog50.
The GWAS data for PD-related exposure factors, including all possible factors, were also downloaded from the OpenGWAS database (https://gwas.mrcieu.ac.uk//). We performed univariate MR analysis with PD data using these factors and retained only factors that had P values of < 0.05. We identified 61 qualified PD-related exposure factors (Table S3).
Instrumental variables
Because of the small sample size of the GWAS data on BAA levels in this study, SNPs were screened using the following criteria during the MR analyses: SNPs with P < 5 × 10− 6 and those in linkage imbalance (r2 < 0.001 SNPs with a physical distance between every two genes > 10000 kb) were excluded. Next, we calculated the F-statistics of the screened instrumental variables to remove weak instrumental bias. F < 10 indicated that the genetic variation used was a weak instrumental variable that might have biased the results51. The specific formula for calculating the F-statistic is
where N represents the sample size, K represents the number of instrumental variables used, and R2 reflects the degree of exposure to the inverse variances (IVs). R2 = 2 × (1 - MAF) × MAF × β2, where MAF is the minimum allele frequency, and β is the allele effect value.
MR causal effect estimation
Five types of two-sample MR (IVW, MR-Egger, weighted median, simple mode, and weight mode) methods were used to evaluate the causal effects of exposure on the outcome. However, according to a previous study52, the IVW method fits better for the evaluation of causal effects. In this study, the IVW method was treated as the main method in the MR analysis, while the other methods were treated as a supplement. If pleiotropy was present, the MR-Egger method was used to calculate the results.
Sensitivity analysis
Heterogeneity investigation
Cochran’s Q test was used to evaluate heterogeneity between various SNP estimates. If significant heterogeneity existed, the random-effects model was adopted during IVW53. I2 statistics were used to reflect the proportion of heterogeneity in the total variation of instrumental variables; I2 > 50% indicated high heterogeneity, whereas I2 < 25% indicated mild or no heterogeneity. The formula for I2 calculation is
Horizontal pleiotropy
We used the MR-Egger method to conduct a pleiotropy test on the instrumental variables. If the intercept of the MR Egger met P < 0.05, this indicated significant horizontal pleiotropy of genetic variation.
Leave-one-out verification
The MR results of the remaining instrumental variables were calculated by removing individual SNPs. A significant change after excluding the SNP indicated that the MR effect estimation was sensitive to the SNP.
Multivariate MR analysis and estimation of mediating effects
Multivariate MR is an extension of MR that uses genetic variations related to multiple possible exposures to estimate their impact on a single outcome. Before conducting the multivariate MR analysis, we first conducted a univariate MR analysis of BAA levels and exposure factors. Significant exposure factors were included in the multivariate MR analysis to construct the model. By identifying the direct effects of BAA levels and PD-related exposure factors on PD through multivariate MR analysis, the indirect effect of BAA levels on PD-related exposure factors can be obtained through the BAA → PD-related exposure factors → PD pathway axis. The effect value and standard error of the mediating effect were calculated using the following formula:
where βM is the value of the mediating effect, βA is the MR effect value of BAA levels on PD-related exposure factors, βB is the direct effect value of PD-related exposure factors on PD, SEM is the standard error of mediating effects, SEA is the standard error of the MR analysis of BAA levels on PD-related exposure factors, and SEB is the standard error of the MR analysis of PD-related exposure factors on PD. The proportion of mediating effect is calculated as |βM/βC| × 100%, where βC is the effect value of BAA levels on PD in univariate MR.
Statistical analyses
All data calculations and statistical analyses were performed using the R software (https://www.r-project.org/, version 4.2.2). The “TwoSampleMR” package was used for the analyses54. Cochran’s Q test and leave-one-out analysis were used to evaluate the robustness and reliability of the results. The MR-Egger intercept method was used to test horizontal pleiotropy. ORs and 95% CIs were calculated for the effect size. All statistical P-values were tested bilaterally, and statistical significance was set at P < 0.05.
Data availability
The datasets generated and analysed during the current study are available included in this article and its supplemental information files.
References
Kacprzyk, K. W. et al. Prevalence of Malnutrition in patients with Parkinson’s disease. Syst. Rev. Nutrients. 14 (23), 5194. https://doi.org/10.3390/nu14235194 (2022). Published 2022 Dec 6.
Wong, S. L., Gilmour, H. & Ramage-Morin, P. L. Parkinson’s disease: prevalence, diagnosis and impact. Health Rep. 25 (11), 10–14 (2014).
Baiano, C. et al. Prevalence and clinical aspects of mild cognitive impairment in Parkinson’s disease: a meta-analysis. Mov. Disord. 35 (1), 45–54. https://doi.org/10.1002/mds.27902 (2020).
Kacprzyk, K. W. et al. Prevalence of Malnutrition in patients with Parkinson’s disease: a systematic review. Nutrients 14 (23), 5194. https://doi.org/10.3390/nu14235194 (2022). Published 2022 Dec 6.
Qi, S. et al. Prevalence of Parkinson’s Disease: A Community-based study in China. Mov. Disord. 36 (12), 2940–2944. https://doi.org/10.1002/mds.28762 (2021).
Tolosa, E. et al. Challenges in the diagnosis of Parkinson’s disease. Lancet Neurol. 20 (5), 385–397. https://doi.org/10.1016/S1474-4422(21)00030-2 (2021).
Jankovic, J. Parkinson’s disease: clinical features and diagnosis. J. Neurol. Neurosurg. Psychiatry. 79 (4), 368–376. https://doi.org/10.1136/jnnp.2007.131045 (2008).
Balestrino, R. & Schapira, A. H. V. Parkinson disease. Eur. J. Neurol. 27 (1), 27–42. https://doi.org/10.1111/ene.14108 (2020).
Elbaz, A. et al. Epidemiology of Parkinson’s disease. Rev. Neurol. (Paris). 172 (1), 14–26. https://doi.org/10.1016/j.neurol.2015.09.012 (2016).
Costa, H. N. et al. Parkinson’s Disease: a multisystem disorder. Neurosci. Bull. 39 (1), 113–124. https://doi.org/10.1007/s12264-022-00934-6 (2023).
Trist, B. G., Hare, D. J. & Double, K. L. Oxidative stress in the aging substantia nigra and the etiology of Parkinson’s disease. Aging Cell. 18 (6), e13031. https://doi.org/10.1111/acel.13031 (2019).
Bjørklund, G. et al. The glutathione system in Parkinson’s disease and its progression. Neurosci. Biobehav Rev. 120, 470–478. https://doi.org/10.1016/j.neubiorev.2020.10.004 (2021).
Molina-Holgado, E., Dewar, K. M., Grondin, L., van Gelder, N. M. & Reader, T. A. Changes of amino acid and monoamine levels after neonatal 6-hydroxydopamine denervation in rat basal ganglia, substantia nigra, and Raphe nuclei. J. Neurosci. Res. 35 (4), 409–418. https://doi.org/10.1002/jnr.490350408 (1993).
Dionísio, P. A., Amaral, J. D. & Rodrigues, C. M. P. Oxidative stress and regulated cell death in Parkinson’s disease. Ageing Res. Rev. 67, 101263. https://doi.org/10.1016/j.arr.2021.101263 (2021).
Fan, X. et al. Role of homocysteine in the development and progression of Parkinson’s disease. Ann. Clin. Transl Neurol. 7 (11), 2332–2338. https://doi.org/10.1002/acn3.51227 (2020).
Figura, M. et al. Serum amino acid profile in patients with Parkinson’s disease. PLoS One. 13 (1), e0191670. https://doi.org/10.1371/journal.pone.0191670 (2018). Published 2018 Jan 29.
Lakke, J. P. et al. Amino acid abnormalities in cerebrospinal fluid and blood serum of patients with Parkinson’s disease, other heredodegenerative disorders and head injuries. Adv. Neurol. 45, 243–247 (1987).
Cui, Z. et al. Relationship between serum amino acid levels and bone Mineral density: a mendelian randomization study. Front. Endocrinol. (Lausanne). 12, 763538. https://doi.org/10.3389/fendo.2021.763538 (2021). Published 2021 Nov 9.
Wiklund, P. et al. Serum amino acid profiles in childhood predict triglyceride level in adulthood: a 7-Year longitudinal study in girls. J. Clin. Endocrinol. Metab. 101 (5), 2047–2055. https://doi.org/10.1210/jc.2016-1053 (2016).
Hu, W. et al. Effects of PPM1K rs1440581 and rs7678928 on serum branched-chain amino acid levels and risk of cardiovascular disease. Ann. Med. 53 (1), 1316–1326. https://doi.org/10.1080/07853890.2021.1965204 (2021).
Barlow, D. P. & Bartolomei, M. S. Genomic imprinting in mammals. Cold Spring Harb Perspect. Biol. 6 (2), a018382. https://doi.org/10.1101/cshperspect.a018382 (2014). Published 2014 Feb 1.
Schnurr, T. M. et al. The influence of transmitted and non-transmitted parental BMI-associated alleles on the risk of overweight in childhood. Sci Rep. ;10(1):4806. Published 2020 Mar 16. doi: (2020). https://doi.org/10.1038/s41598-020-61719-3
Ma, W. et al. Sex-biased and parental allele-specific gene regulation by KDM6A. Biol. Sex. Differ. 13 (1), 40. https://doi.org/10.1186/s13293-022-00452-0 (2022). Published 2022 Jul 23.
Richardson, M. B. et al. Synthesis and explosion hazards of 4-Azido-l-phenylalanine. J. Org. Chem. 83 (8), 4525–4536. https://doi.org/10.1021/acs.joc.8b00270 (2018).
Janas, K. M. The control of L-phenylalanine ammonia-lyase activity by phosphonate and aminooxy analogues of phenylalanine. Acta Biochim. Pol. 40 (4), 451–454 (1993).
Niu, P. P. et al. MIA-Directed 2-Pyridione-enabled selective Ortho-C-H arylation of phenylalanine: a mechanistic study. J. Org. Chem. 86 (3), 3096–3106. https://doi.org/10.1021/acs.joc.0c02872 (2021).
van Spronsen, F. J., Hoeksma, M. & Reijngoud, D. J. Brain dysfunction in phenylketonuria: is phenylalanine toxicity the only possible cause? J. Inherit. Metab. Dis. 32 (1), 46–51. https://doi.org/10.1007/s10545-008-0946-2 (2009).
Fernstrom, J. D. & Fernstrom, M. H. Tyrosine, phenylalanine, and catecholamine synthesis and function in the brain. J. Nutr. 137 (6 Suppl 1), 1539S–1548S. https://doi.org/10.1093/jn/137.6.1539S (2007).
Sanayama, Y. et al. Experimental evidence that phenylalanine is strongly associated to oxidative stress in adolescents and adults with phenylketonuria. Mol. Genet. Metab. 103 (3), 220–225. https://doi.org/10.1016/j.ymgme.2011.03.019 (2011).
Vallée, A., Vallée, J. N. & Lecarpentier, Y. Potential role of cannabidiol in Parkinson’s disease by targeting the WNT/β-catenin pathway, oxidative stress and inflammation. Aging (Albany NY). 13 (7), 10796–10813. https://doi.org/10.18632/aging.202951 (2021).
Robea, M. A. et al. Parkinson’s Disease-Induced zebrafish models: focussing on oxidative stress implications and sleep processes. Oxid. Med. Cell. Longev. 2020, 1370837. https://doi.org/10.1155/2020/1370837 (2020). Published 2020 Aug 18.
Jiménez-Jiménez, F. J. et al. Cerebrospinal and blood levels of amino acids as potential biomarkers for Parkinson’s disease: review and meta-analysis. Eur. J. Neurol. 27 (11), 2336–2347. https://doi.org/10.1111/ene.14470 (2020).
MaffeiME 5-Hydroxytryptophan (5-HTP): natural occurrence, analysis, biosynthesis, Biotechnology, Physiology and Toxicology. Int. J. Mol. Sci. 22 (1), 181. https://doi.org/10.3390/ijms22010181 (2020). Published 2020 Dec 26.
Wang, Q., Liu, D., Song, P. & Zou, M. H. Tryptophan-kynurenine pathway is dysregulated in inflammation, and immune activation. Front. Biosci. (Landmark Ed). 20 (7), 1116–1143. https://doi.org/10.2741/4363 (2015). Published 2015 Jun 1.
Geisler, S. et al. Neopterin levels and Kyn/Trp ratios were significantly increased in dengue virus patients and subsequently decreased after recovery. Int. J. Infect. Dis. 91, 162–168. https://doi.org/10.1016/j.ijid.2019.12.005 (2020).
Truyens, M. et al. Effect of 5-Hydroxytryptophan on fatigue in quiescent inflammatory bowel disease: a Randomized Controlled Trial. Gastroenterology 163 (5), 1294–1305e3. https://doi.org/10.1053/j.gastro.2022.07.052 (2022).
Hayat, S. et al. Role of proline under changing environments: a review. Plant. Signal. Behav. 7 (11), 1456–1466. https://doi.org/10.4161/psb.21949 (2012).
Mauger, A. B. Naturally occurring proline analogues. J. Nat. Prod. 59 (12), 1205–1211. https://doi.org/10.1021/np9603479 (1996).
Sebák, F., Szolomájer, J., Papp, N., Tóth, G. K. & Bodor, A. Proline cis/trans isomerization in intrinsically disordered proteins and peptides. Front. Biosci. (Landmark Ed). 28 (6), 127. https://doi.org/10.31083/j.fbl2806127 (2023).
Christgen, S. L. & Becker, D. F. Role of Proline in Pathogen and host interactions. Antioxid. Redox Signal. 30 (4), 683–709. https://doi.org/10.1089/ars.2017.7335 (2019).
Plewa, S. et al. The Metabolomic Approach reveals the alteration in human serum and cerebrospinal fluid composition in Parkinson’s Disease patients. Pharmaceuticals (Basel). 14 (9), 935. https://doi.org/10.3390/ph14090935 (2021). Published 2021 Sep 17.
Rempe, R. G., Hartz, A. M. S. & Bauer, B. Matrix metalloproteinases in the brain and blood-brain barrier: versatile breakers and makers. J. Cereb. Blood Flow. Metab. 36 (9), 1481–1507. https://doi.org/10.1177/0271678X16655551 (2016).
Li, P. & Wu, G. Roles of dietary glycine, proline, and hydroxyproline in collagen synthesis and animal growth. Amino Acids. 50 (1), 29–38. https://doi.org/10.1007/s00726-017-2490-6 (2018).
Langrock, T. & Hoffmann, R. Analysis of Hydroxyproline in Collagen Hydrolysates. Methods Mol. Biol. 2030, 47–56. https://doi.org/10.1007/978-1-4939-9639-1_5 (2019).
Cheng, J. Y., Deng, Y. T. & Yu, J. T. The causal role of circulating amino acids on neurodegenerative disorders: a two-sample mendelian randomization study. J. Neurochem. 166 (6), 972–981. https://doi.org/10.1111/jnc.15937 (2023).
Skrivankova, V. W. et al. Strengthening the reporting of Observational studies in Epidemiology using mendelian randomization: the STROBE-MR Statement. JAMA 326 (16), 1614–1621. https://doi.org/10.1001/jama.2021.18236 (2021).
Shin, S. Y. et al. An atlas of genetic influences on human blood metabolites. Nat. Genet. 46 (6), 543–550. https://doi.org/10.1038/ng.2982 (2014).
Kettunen, J. et al. Genome-wide study for circulating metabolites identifies 62 loci and reveals novel systemic effects of LPA. Nat. Commun. 7, 11122. https://doi.org/10.1038/ncomms11122 (2016). Published 2016 Mar 23.
Kurki, M. I. et al. FinnGen provides genetic insights from a well-phenotyped isolated population [published correction appears in Nature. ;:]. Nature. 2023;613(7944):508–518. doi: (2023). https://doi.org/10.1038/s41586-022-05473-8
Buniello, A. et al. The NHGRI-EBI GWAS catalog of published genome-wide association studies, targeted arrays and summary statistics 2019. Nucleic Acids Res. 47 (D1), D1005–D1012. https://doi.org/10.1093/nar/gky1120 (2019).
Pierce, B. L., Ahsan, H. & Vanderweele, T. J. Power and instrument strength requirements for mendelian randomization studies using multiple genetic variants. Int. J. Epidemiol. 40 (3), 740–752. https://doi.org/10.1093/ije/dyq151 (2011).
Bowden, J. et al. Consistent estimation in mendelian randomization with some Invalid instruments using a weighted median estimator. Genet. Epidemiol. 40 (4), 304–314. https://doi.org/10.1002/gepi.21965 (2016).
Sang, N. et al. Causal relationship between sleep traits and risk of systemic lupus erythematosus: a two-sample mendelian randomization study. Front. Immunol. 13, 918749. https://doi.org/10.3389/fimmu.2022.918749 (2022). Published 2022 Jun 17.
Xiao, G. et al. Causality of genetically determined metabolites on anxiety disorders: a two-sample Mendelian randomization study. J Transl Med. ;20(1):475. Published 2022 Oct 20. doi: (2022). https://doi.org/10.1186/s12967-022-03691-2
Acknowledgements
This study was supported by National Natural Science Foundation of China (82074355, 82074441) and Shanghai Municipal Health Commission (GWIV-28).
Author information
Authors and Affiliations
Contributions
Lei Cheng is responsible for project design, experiment division, data analysis, manuscript preparation and submission; Jing Zhang, Wenting Shi and Qingjun Dong complete data recording and analysis; Juntao Yan and Yiren Hu gives technical and writing guidance; Qingjun Dong, Qing Ye and Zhongyan Zhou participates in project design and provides funding support.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Cheng, L., Yan, J., Shi, W. et al. Exploring the causal link between serum amino acids and Parkinson’s disease: a Mendelian randomization approach. Sci Rep 14, 30271 (2024). https://doi.org/10.1038/s41598-024-81787-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-024-81787-z