Abstract
Analyze the correlation between the DNA methylation levels of Protocadherin 17 (PCDH17) cg03865667 and rheumatoid arthritis (RA), and evaluate its potential as a biomarker for diagnosing RA. Peripheral blood samples were collected from a cohort of 370 individuals, comprising patients diagnosed with RA, ankylosing spondylitis (AS), psoriatic arthritis (PsA), gout, systemic lupus erythematosus (SLE), dermatomyositis (DM), primary Sjögren’s syndrome (SS), and healthy controls (HC), for subsequent analysis.DNA methylation sequencing techniques were employed to evaluate the methylation levels of the PCDH17 cg03865667 locus. Relative to the HC, AS, and SS groups, PCDH17 cg03865667 was significantly downregulated in RA patients (P = 0.0403; p = 0.0290; p = 0.044). Compared to the HC group, the methylation levels at CpG sites 57,631,544, 57,631,571, and 57,631,581 were significantly downregulated in RA patients (p = 0.0078; p = 0.0123; p = 0.0309). For the TTCCTT and TTTCTT haplotypes, methylation levels were significantly lower in RA patients than in HC (p = 0.0188; p = 0.0053), particularly for the TTTCTT haplotype. Significant differences were observed between the CCP(-) RF(-) group, the CCP(+) / RF(+) group, and the HC group among the RA subgroups. No significant differences were found within the double-positive subgroup.The average methylation level of PCDH17 was negatively correlated with C-reactive protein (CRP) (r=-0.28, p = 6.9e-4). This study indicates that PCDH17 cg03865667 methylation may function as a potential biomarker for RA diagnosis. The subgroup analysis suggests that it may serve as a potential biomarker for diagnosing RA, indicating a capacity to enhance the diagnostic accuracy for seronegative RA and reduce the likelihood of missed and incorrect diagnoses.
Similar content being viewed by others
Introduction
Rheumatoid arthritis (RA) is described as a chronic, destructive, and progressive systemic autoimmune disease, characterized by symmetrical polyarthritis and synovitis. It represents one of the most common autoimmune diseases1. Common symptoms in RA patients include morning joint stiffness, fatigue, joint tenderness, subcutaneous rheumatoid nodules, fever, and symmetrical inflammation of small joints. Persistent inflammation may result in joint pain, progressive destruction, and joint dysfunction2. Additionally, inflammation is associated with various complications, including cardiovascular diseases3. Therefore, the effective control of inflammation is crucial to the treatment of RA and may also serve as an indicator for evaluating treatment efficacy. Physicians may adjust treatment plans based on the progression of inflammation. Investigating the specific molecular biological mechanisms of rheumatoid arthritis and identifying inflammation-related biomarkers is crucial.
A body of evidence suggests that both genetic and environmental factors are involved in the pathogenesis of RA. However, these factors alone do not completely elucidate the mechanism of RA onset. Recent evidence indicates that epigenetics plays a crucial role in the development of RA. DNA methylation, the most commonly studied epigenetic modification, has been utilized to characterize various aspects of RA pathology4. Additionally, RA-associated DNA methylation patterns demonstrate site-specific differences in immune pathways. Investigation of the relationship between DNA methylation and rheumatoid arthritis may lead to the discovery of new biomarkers and therapeutic targets for the disease.
Protocadherin 17 (PCDH17) is classified as a member of the protocadherin family, a component of the cadherin superfamily known for mediating calcium-dependent cell-cell adhesion5,6. It is recognized as a potential tumor suppressor gene and is commonly methylated7. Research has demonstrated that PCDH17 is frequently methylated in various types of cancers, including esophageal squamous cell carcinoma (ESCC)8, gastric cancer9, and nasopharyngeal carcinoma10, and correlates with a poor prognosis. Evidence suggests that PCDH17 plays a critical role in various biological activities, such as cell cycle regulation, apoptosis, autophagy, and signal transduction11,12,13.
Currently, there is limited research on PCDH17, especially concerning its relationship with rheumatoid arthritis (RA). In this study, we investigated the association between PCDH17, whole blood methylation haplotypes, average methylation levels, and specific methylation sites with clinical indicators in RA patients. This research may offer potential clinical applications for predicting the degree of inflammation in RA patients. Analyzing RA subgroups could positively impact the diagnostic accuracy for seronegative RA, reduce misdiagnoses and missed diagnoses, thereby facilitating early diagnosis and treatment.
Materials and methods
Study design and patients
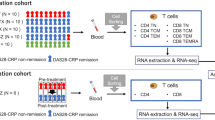
Prior to the commencement of this study, it was reviewed and approved by the Ethics Committee of Shanghai Guanghua Integrative Medicine Hospital (2018-K-12). All participants provided written informed consent prior to inclusion. All methods are conducted in accordance with relevant guidelines and regulations. Participants in this study were drawn from the Precision Medicine Research Cohort (PMRC) at Guanghua Hospital, affiliated with Shanghai University of Traditional Chinese Medicine, under clinical trial number ChiCTR2400083234. The cohort consisted of 370 participants, including 30 healthy controls (HC) and 340 patients with various rheumatic diseases. The spectrum of rheumatic diseases studied encompassed RA, ankylosing spondylitis (AS), psoriatic arthritis (PsA), gout, Sjögren’s syndrome (SS), systemic lupus erythematosus (SLE), and dermatomyositis (DM). Patient classifications included 166 RA cases, 24 SS cases, and 30 cases of other diseases. RA patients were categorized into three groups based on serum rheumatoid factor (RF) and anti-cyclic citrullinated peptide (anti-CCP) antibodies: CCP(-)RF(-), CCP(+)/RF(+), and CCP(+)RF(+). Disease inclusion criteria adhered to several standards, including the 2010 ACR criteria for RA, the 1984 revised New York criteria for AS, the 2015 ACR/EULAR criteria for gout, the 2006 ACR criteria for PsA, the 2016 ACR/EULAR classification for SS, the 2019 EULAR/ACR criteria for SLE, and the 2017 ACR/EULAR criteria for DM. Patients presenting with comorbid diseases, severe liver or kidney damage, cardiovascular disease, or a history of malignancy were excluded from the study. Comprehensive clinical information for all participants was documented, and whole blood samples were collected.
Targeted DNA methylation analysis
Peripheral venous blood samples were collected from all participants, and genomic DNA was extracted using the Blood Genomic DNA Extraction Kit (Concert, RC1001). The preprocessing step involved adding 40 µL of Proteinase K solution (10 mg/mL) to a 1.5 mL sample tube, followed by 400 µL of whole blood. The pretreated samples underwent automated DNA extraction using a nucleic acid purifier (Concert, H8/HF16), with rigorous quality control ensuring a DNA concentration of ≥ 20 ng/µL and a total amount of ≥ 400 ng. Sample purity was determined by an OD260/280 ratio between 1.7 and 1.9, and an OD260/230 ratio of ≥ 2.0. The target region was amplified by polymerase chain reaction (PCR) using a forward primer (PrimerF) with the sequence TTGGAATTAAATTGTTTGGAGAG and a reverse primer (PrimerR) with the sequence ACCACAACCTAATCAACATTT. The study focused on the cg03865667_43 site, with a sequencing length of 181 bp, located between chr13:57631468 and chr13:57631648. Following primer amplification, genomic DNA was treated with bisulfite to convert unmethylated cytosines (C) to uracils (U). Indexed PCR amplification introduced specific barcode sequences compatible with the Illumina platform at the ends of the library. Finally, high-throughput sequencing was conducted on the Illumina HiSeq platform (Illumina, CA, USA) using a paired-end 2 × 150 bp sequencing mode to generate FastQ data.
Statistical methods
Statistical analyses were conducted using IBM SPSS 27.0, GraphPad Prism software (version 10.2.3), and Sangerbox. The Kruskal-Wallis rank sum test was utilized to evaluate differences among groups, with multiple comparisons adjusted using the two-stage step-up method of Benjamini, Krieger, and Yekutieli. Spearman rank correlation analysis was applied to investigate the relationship between DNA methylation levels and clinical data in RA patients. P < 0.05 was considered statistically significant. The clinical relevance of PCDH17 methylation levels was determined through receiver operating characteristic (ROC) curve analysis.
Results
Differences in PCDH17 gene methylation levels between RA and other diseases
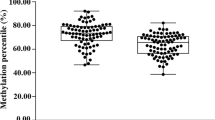
The Kruskal-Wallis rank sum test calculated P-values between the RA group and other groups, including AS, gout, PsA, SS, SLE, DM, and HC. Whole blood methylation studies revealed that PCDH17 (cg03865667) was downregulated in RA patients compared to the HC, AS, and SS groups (P = 0.0403; p = 0.0290; p = 0.044) (Fig. 1A). Conversely, PCDH17 (cg03865667) methylation levels were significantly elevated in RA patients relative to the DM group (P < 0.0001) (Fig. 1A). Subsequent analysis identified six CpGs within PCDH17, namely 57,631,502 (cg03865667_35), 57,631,504 (cg03865667_37), 57,631,526 (cg03865667_59), 57,631,544 (cg03865667_77), 57,631,571 (cg03865667_104), and 57,631,581 (cg03865667_114). The same statistical test was applied to calculate P-values between the RA group, other disease groups, and the HC group. Differential analysis across all six CpGs indicated significantly higher methylation levels in RA patients compared to the DM group (Fig. 1B-G). Relative to the HC group, RA patients exhibited significantly lower methylation levels at CpGs 57,631,544, 57,631,571, and 57,631,581 (p = 0.0078; p = 0.0123; p = 0.0309) (Fig. 1E-G). For CpGs 57,631,504, 57,631,544, and 57,631,571, methylation levels in RA patients were significantly reduced compared to those in the AS group (p = 0.0473; p = 0.0326; p = 0.0375) (Fig. 1C, E, F). Additionally, RA patients demonstrated significantly reduced methylation levels at CpGs 57,631,571 and 57,631,581 compared to the SS group (p = 0.0123; p = 0.0093) (Fig. 1F-G).
Methylation levels at the PCDH17 cg03865667 CpG site. (A-E) Multiple comparisons of methylation levels at different CG sites of PCDH17 cg03865667 across multiple groups, with P < 0.05 being statistically significant.
Significant changes in PCDH17 haplotype methylation levels between RA and other diseases
In the PCDH17 cg03865667 region, 10 methylated haplotypes were identified, and changes in their methylation levels were analyzed between RA and other diseases. Methylation levels in the TTCCTT and TTTCTT haplotypes were significantly reduced in RA patients compared to HC (p = 0.0188; p = 0.0053) (Fig. 2B-C), with a more pronounced decrease observed in the TTTCTT haplotype. DM patients exhibited significantly lower methylation levels in the CCCCTT, TTCCTT, and TTTCTT haplotypes compared to HC and RA (Fig. 2A-C). Additionally, significant differences in methylation levels in the TTTCTT haplotype were observed across the six groups, except for GOUT patients, compared to the HC group. The TTTTTTT haplotype is characterized by an absence of methylation in the PCDH17 gene. In this haplotype, methylation levels were significantly elevated in the SLE and DM groups compared to the HC group (p = 0.0256; p < 0.0001) (Fig. 2D). Methylation levels were significantly lower in the AS group compared to the RA group (p = 0.0483) (Fig. 2D) and significantly higher in the DM group compared to the RA group (p < 0.0001) (Fig. 2D).
Analysis of Haplotype Methylation Levels of PCDH17 cg03865667 and Methylation Level Differences in RA Subgroups. (A-D) Multiple comparisons of the haplotypic methylation levels of PCDH17 cg03865667 among multiple groups, with P < 0.05 considered statistically significant. (E) Multiple comparisons of methylation levels were performed across RA subgroups, and a P-value of less than 0.05 was considered statistically significant.
Differences in methylation levels of PCDH17 gene at different sites and haplotypes between RA subgroups and HC group
In this study, subgroup analysis was conducted on RA patients using the Kruskal-Wallis rank sum test to calculate p-values between the CCP(-) RF(-) group, CCP(+)/RF(+) group, CCP(+) RF(+) group, and the healthy control group. It was found that there were significant differences in the average methylation levels of PCDH17 among the CCP(-) RF(-) group, the CCP(+)/RF(+) group, and the healthy control group (p = 0.0349; p = 0.0172)(Fig. 2E).
Correlation of PCDH17 methylation levels with common clinical indicators in RA patients
We explored the correlations between different sites and haplotype methylation levels of PCDH17 cg03865667 and common clinical indicators in RA patients. These indicators included sex, age, height, weight, ESR, CRP, RF, anti-CCP, and the presence of comorbidities such as interstitial lung disease, hypertension, and osteoporosis. Spearman rank correlation analysis assessed the correlations among these variables. Visualization and analysis of the data were performed using network data matrices and the Sangerbox tool. The correlation analysis revealed strong associations between sites 1544, 1526, 1571, and the average methylation level of PCDH17 (|r|≥0.9) (Fig. 3A). Additionally, osteoporosis showed a strong correlation with both RF and CCP (|r|=0.44, p = 0.01; |r|=0.52, p = 0.02) (Fig. 3A-B).
Correlation of PCDH17 cg03865667 methylation level with clinical indicators and results of clinical modeling. (A-B) Correlation of PCDH17 cg03865667 methylation level with clinical indicators. (C) Logistic regression was used to evaluate the roc curves of PCDH17 cg03865667.
In the correlation analysis, PCDH17 methylation levels were negatively correlated with CRP (r= −0.28, p = 6.9e-4) (Figure S1 A). With the exception of the 57,631,581 site, the other five CpGs showed the same trend (Table 1) (Figure S1 B-F). This suggests that PCDH17 may have significant clinical value in predicting CRP levels in these subgroups.
Correlation of PCDH17 haplotype methylation levels with common clinical indicators
Similarly, significant differences were identified in the correlation analysis between haplotype methylation levels and clinical indicators (Table 2). Correlation analysis revealed that TTCCTT and TTCTTT haplotype methylation levels were negatively correlated with ESR (r= −0.19, p = 0.02; r= −0.20, p = 0.01) (Figure S1 G-H). The TTTTTTT haplotype, indicating an absence of methylation in the PCDH17 gene, was positively correlated with CRP (r = 0.24, p = 0.0036) (Figure S1I), yet it exhibited no significant correlation with ESR.
The methylation levels of PCDH17 may assist in the diagnosis of seronegative RA
We conducted logistic regression analysis to investigate the specificity and accuracy of PCDH17 methylation levels in diagnosing RA. Univariate logistic regression results from the seronegative RA subgroups, compared with the HC group, indicated an area under the curve (AUC) of approximately 0.65 (Fig. 3C).
Discussion
RA is the most prevalent systemic autoimmune disease, leading to functional impairment and premature death. Consequently, early diagnosis and intervention are essential to mitigate the risks associated with this disease. Studies indicate that approximately 70–80% of RA patients possess anti-citrullinated peptide antibodies (ACPA) and RF14,15. However, 20–25% of RA cases are seronegative for RF and ACPA, leading to delays in diagnosis and the initiation of disease-modifying antirheumatic drugs (DMARDs) treatment for these patients16.ESR and CRP are commonly used tests for RA, but their sensitivity and specificity are relatively low17. ESR is correlated with disease activity and typically elevates in RA patients, particularly during active phases of the disease. However, an elevated ESR is not specific to RA and may also be observed in other inflammatory and infectious conditions. CRP, a highly responsive acute-phase serum reactant, is produced by the liver. It is produced in response to various pro-inflammatory cytokines from monocytes or macrophages. CRP levels are frequently used to gauge disease activity in RA, in conjunction with assessing joint swelling and discomfort.
DNA methylation is one of the most extensively studied epigenetic alterations in human cancers, occurring predominantly in the CpG islands of gene promoters. Abnormal promoter methylation leads to the silencing of tumor suppressor genes, playing a crucial role in the development and progression of many cancers18,19,20. The PCDH17 gene, located on human chromosome 13q21.2, is often inactivated due to promoter methylation in various cancers9. PCDH17 has been identified as an inhibitor of the Wnt/β-catenin signaling pathway in breast cancer21. The Wnt/β-catenin signaling pathway plays a significant role in the pathophysiology of RA22. Studies have demonstrated that this pathway is pivotal in the pathogenesis of RA, particularly in the aberrant activation of synovial fibroblasts (FLS) and joint destruction. Research also suggests that the Wnt/β-catenin signaling pathway is closely associated with inflammatory responses, and targeting this pathway may offer a novel strategy for treating RA23.
In this study, correlation analysis between PCDH17 methylation levels and clinical indicators revealed a negative correlation between PCDH17 and C-reactive protein (CRP) levels (r= −0.28, p = 6.9e-4), confirming a close association between PCDH17 methylation levels and CRP. Additionally, among six CpG sites analyzed, five exhibited the same trend, with the exception of the site at position 57,631,581. RF and anti-citrullinated protein antibodies (ACPA) are autoantibodies used in the diagnosis of RA patients. Their discovery has greatly facilitated the early diagnosis and treatment of RA24. Building on this theoretical research, ROC analysis of PCDH17 with CCP(-) RF(-) was conducted, showing that methylation levels of PCDH17 can enhance the diagnostic accuracy for seronegative RA, thereby avoiding misdiagnosis.
Additionally, this study comprised 30 healthy individuals and 340 patients diagnosed with seven distinct rheumatic and autoimmune diseases. Methylation differential analysis identified significant differences between the RA group and the HC, AS, SS, and DM groups, offering novel insights into clinical differentiation. Furthermore, correlation analysis demonstrated a strong association between osteoporosis and both RF and CCP, suggesting clinicians should remain vigilant regarding the potential for osteoporosis in RA patients.
This study conducted an analysis of PCDH17 methylation levels to preliminarily explore the relationship between RA and the methylation of the PCDH17 gene, potentially offering clinical applications for predicting the inflammation severity in RA patients. Additionally, it positively impacts the diagnostic accuracy for seronegative RA and reduces the likelihood of misdiagnoses and missed diagnoses. Finally, the study analyzed a limited set of clinical indicators. Future research should consider including a broader range of relevant indicators and increasing the sample size to further investigate the relationship between PCDH17 cg03865667 methylation and the onset and activity of RA.
Data availability
The datasets presented in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found below: SRA, PRJNA1094652.
References
Rosik, J., Kulpa, J., Szczepanik, M. & Pawlik, A. The role of semaphorins in the pathogenesis of rheumatoid arthritis. Cells 13 (7), 618 (2024). Published 2024 Apr 2.
Scott, D. L., Wolfe, F. & Huizinga, T. W. Rheumatoid arthritis. Lancet 376 (9746), 1094–1108 (2010).
Jiang, P. et al. DNA methylation change of HIPK3 in Chinese rheumatoid arthritis and its effect on inflammation. Front. Immunol. 13, 1087279 (2023).
Shi, Y. et al. Circulating DNA methylation level of CXCR5 correlates with inflammation in patients with rheumatoid arthritis. Immun. Inflamm. Dis. 11 (6), e902 (2023).
Kim, S. Y., Yasuda, S., Tanaka, H., Yamagata, K. & Kim, H. Non-clustered Protocadherin. Cell. Adh Migr. 5 (2), 97–105 (2011).
Frank, M., Kemler, R. & Protocadherins Curr. Opin. Cell. Biol. ;14(5):557–562. (2002).
Hu, X. et al. ET al. Protocadherin 17 acts as a tumour suppressor inducing tumour cell apoptosis and autophagy, and is frequently methylated in gastric and colorectal cancers. J Pathol. ;229(1):62–73. (2013).
Haruki, S. et al. Frequent Silencing of Protocadherin 17, a candidate tumour suppressor for esophageal squamous cell carcinoma. Carcinogenesis 31 (6), 1027–1036 (2010).
Costa, V. L. et al. ET al. TCF21 and PCDH17 methylation: An innovative panel of biomarkers for a simultaneous detection of urological cancers. Epigenetics 6(9), 1120–1130 (2011).
Yang, Y., Liu, J., Li, X. & Li, J. C. PCDH17 gene promoter demethylation and cell cycle arrest by genistein in gastric cancer. Histol. Histopathol. 27 (2), 217–224 (2012).
Liu, S. et al. PCDH17 increases the sensitivity of colorectal cancer to 5-fluorouracil treatment by inducing apoptosis and autophagic cell death. Signal. Transduct. Target. Ther. 4, 53 (2019).
Wu, J. C. et al. Se-Allylselenocysteine induces autophagy by modulating the AMPK/mTOR signaling pathway and epigenetic regulation of PCDH17 in human colorectal adenocarcinoma cells. Mol. Nutr. Food Res. 59 (12), 2511–2522 (2015).
Xiang, Y., Yang, Y., Lin, C., Wu, J. & Zhang, X. MiR-23a-3p promoted G1/S cell cycle transition by targeting protocadherin17 in hepatocellular carcinoma. J. Physiol. Biochem. 76 (1), 123–134 (2020).
Kim, H. & Sung, Y. K. Epidemiology of rheumatoid arthritis in Korea. J. Rheum. Dis. 28 (2), 60–67 (2021).
Rantapää-Dahlqvist, S. et al. Antibodies against Cyclic citrullinated peptide and IgA rheumatoid factor predict the development of rheumatoid arthritis. Arthritis Rheum. 48 (10), 2741–2749 (2003).
Coffey, C. M., Crowson, C. S., Myasoedova, E., Matteson, E. L. & Davis, J. M. 3rd Evidence of Diagnostic and Treatment Delay in Seronegative Rheumatoid Arthritis: Missing the Window of Opportunity. Mayo Clin Proc. ;94(11):2241–2248. (2019).
Colglazier, C. L. & Sutej, P. G. Laboratory testing in the rheumatic diseases: a practical review. South. Med. J. 98 (2), 185–191 (2005).
Chao, C., Chi, M., Preciado, M. & Black, M. H. Methylation markers for prostate cancer prognosis: a systematic review. Cancer Causes Control. 24 (9), 1615–1641 (2013).
Jerónimo, C. & Henrique, R. Epigenetic biomarkers in urological tumors: A systematic review. Cancer Lett. 342 (2), 264–274 (2014).
Neuhausen, A., Florl, A. R., Grimm, M. O. & Schulz, W. A. DNA methylation alterations in urothelial carcinoma. Cancer Biol. Ther. 5 (8), 993–1001 (2006).
Yin, X. et al. Protocadherin 17 functions as a tumor suppressor suppressing Wnt/β-catenin signaling and cell metastasis and is frequently methylated in breast cancer. Oncotarget 7 (32), 51720–51732 (2016).
Rabelo Fde, S. et al. The Wnt signaling pathway and rheumatoid arthritis. Autoimmun. Rev. 9 (4), 207–210 (2010).
Huang, Y. et al. Wilforine inhibits rheumatoid arthritis pathology through the Wnt11/β-catenin signaling pathway axis. Arthritis Res. Ther. 25 (1), 243 (2023).
Smolen, J. S. et al. Rheumatoid arthritis. Nat. Rev. Dis. Primers. 4, 18001 (2018).
Funding
This work was funded by the National Natural Science Funds of China (82074234 and 82073901), Shanghai Municipal Health Commission (202340274).
Author information
Authors and Affiliations
Contributions
FZ, MZ and JL are responsible for sample and data collection, and wrote the main manuscript text. CC, PJ, KW, JZ, YS, YZ for sample and data collection, review and editing. YS, YL prepared Figs. 1, 2 and 3.YZ, QL, LW, HQ and LL for review and editing. SG for conceptualization, review and editing. XL and QZ are responsible for revision, and manuscript review. DH is responsible for the funding acquisition, revision, and manuscript review.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Consent to participate
All authors listed have contributed to the entire writing process of this manuscript and have given their informed consent for its publication.
Consent for publication
The authors assert that none of the material in this paper has been published, nor is it under consideration for publication elsewhere.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhao, F., Zhou, M., Liu, J. et al. Correlation analysis of circulating PCDH17 DNA methylation changes with rheumatoid arthritis patients. Sci Rep 15, 17939 (2025). https://doi.org/10.1038/s41598-025-02236-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-02236-z