Abstract
The ability of methicillin-resistant Staphylococcus aureus (MRSA) to adapt to environmental stressors is crucial for its survival and persistence. This study used shotgun proteomics to analyze the protein profiles of MRSA strains ST398 and JE2 under three experimental conditions: EC1 (control, 37 °C, pH 7), EC2 (35 °C, pH 6), and EC3 (35 °C, pH 6 with 5% NaCl). Proteins were extracted and digested with trypsin using filter-aided sample preparation (FASP), followed by LC–MS/MS analysis. Raw data were analyzed using Proteome Discoverer software with label-free quantification (LFQ), identifying an average of 2640 ± 70 and 2832 ± 154 proteins in ST398 and JE2, respectively. Qualitative and quantitative differences in protein expression were observed between strains and conditions. Under EC2, ABC transporters increased and arginine metabolism was upregulated, suggesting metabolic adaptation to acidic stress. Purine metabolism was downregulated, indicating a metabolic shift. Under EC3 condition, NaCl addition induced glycine-betaine biosynthesis and proteins involved in potassium limitation. JE2 exhibited a more pronounced oxidative and stress response under salt stress, indicating a strain-specific adaptation. These findings highlight strain-specific proteomic responses in MRSA, highlighting the roles of arginine, glycine-betaine, and purine metabolism, along with stress-response proteins, in coping with environmental stressors.
Similar content being viewed by others
Introduction
Staphylococcus aureus (S. aureus) is a gram-positive opportunistic bacterium commonly found in the skin and nasal cavity of many healthy individuals1. It can cause a range of infections, from localized skin and soft tissue infections (SSTIs) to severe systemic diseases such as endocarditis, pneumonia, and toxic shock syndrome2. Its pathogenicity is driven by various virulence factors, and its adaptability to environmental changes such as changes in pH, oxygen, CO2 levels, and temperature that it encounters during infections3.
S. aureus is a significant cause of both hospital-acquired and community-acquired infections, particularly among critically ill and immunocompromised patients, and presents a major challenge in clinical settings. Methicillin-resistant Staphylococcus aureus (MRSA), first identified in 19614, has become a major concern owing to its resistance to common antibiotics and its rapid transmission, contributing to increased morbidity and mortality.
These pathogenic bacteria encounter diverse environmental conditions, depending on the infection site, route, and phase. For instance, skin temperature is generally 2–4 °C lower than the core body temperature5, while the pH is slightly acidic, ranging from 4 to 66; however, it varies according to the wound state from acidic (pH 5.7–7.6) in early infections to alkaline (pH 7.5–8.9) in chronic infections7. NaCl concentration often increases following trauma8 and increased bacterial replication, leading to tissue hypoxia at the infection sites9.
Understanding these metabolic adaptations is crucial for identifying new therapeutic targets against antibiotic-resistant pathogens10,11. Proteomic studies of microbial protein expression in changing environments are essential to uncover metabolic and resistance mechanisms12,13.
This study investigated the protein expression of two MRSA strains under three experimental conditions (ECs) to simulate variations in pH, temperature, and NaCl concentration expected during wound infection. Livestock-associated MRSA (LA-MRSA) ST398, which often colonizes pigs and other animals, is significantly associated with animal-to-human transmission, posing zoonotic and public health risk14,15. The other strain was the Community Associated (CA-MRSA) JE2 strain (USA300 isolate), a primary SSTI source, and CA infections in the USA16.
The selected ECs for this study were designed to replicate the stressors encountered on the skin during S. aureus infections. EC1 served as the control (pH 7 at 37 °C), while EC2 (pH 6 at 35 °C) simulated mild stress, and EC3 (pH 6 at 35 °C with 5% NaCl) introduced additional osmotic stress to mimic the dynamic environment of wound sites. This approach, although involving multiple parameters, aims to emulate realistic environmental shifts in the wound environment, where factors such as pH and temperature fluctuate concurrently. EC2 represents a mild stress adjustment, whereas EC3 introduces targeted osmotic stress, enabling the examination of distinct bacterial proteomic adaptations under progressive levels of stress.
Results
MRSA growth in different experimental conditions
Figure 1A and B illustrate the growth curves of the MRSA strains ST398 and JE2, respectively, under different ECs. ST398 achieved a mid-exponential phase OD600 of 0.5 at approximately 130, 200, and 260 min in EC1, EC2, and EC3, respectively. In contrast, the JE2 strain exhibited a prolonged duration to reach a mid-exponential phase, with an OD600 of 0.55, taking 150 min in EC1, and extending to 180 and 300 min in EC2 and EC3, respectively. These observations suggest that the growth of both MRSA strains was partially influenced by changes in pH, temperature, and the presence of NaCl in TSB. Additionally, the data revealed slight growth discrepancies between strains under these conditions.
Growth curves of MRSA ST398 strain (A) and MRSA JE2 strain (B) in EC1 (blue), EC2 (orange) and EC3 (green). Each curve is obtained from the mean of the three replicates at each point.
Identification metrics and multivariate statistics
Following the observation of strain-specific growth patterns under different conditions, we sought to understand the underlying proteomic changes that might explain these variations. We characterized the bacterial cellular proteome of MRSA ST398 and JE2 strains by separating the peptide mixtures by long-gradient liquid chromatography and analyzing them using a Q-Exactive mass spectrometer. Proteins were identified by a tandem mass spectrometry database search engine, SEQUEST-HT, included in the Proteome Discoverer (PD) software. The identifications were obtained by applying an FDR significance threshold of 1%. PD allowed the identification and quantification of similar numbers of proteins in the two strains, with 2685 ± 83, 2693 ± 18, and 2541 ± 257 ST398 proteins in EC1, EC2, and EC3 respectively, and 2854 ± 29, 2749 ± 44, and 2899 ± 17 JE2 proteins in EC1, EC2, and EC3, respectively, as shown in Fig. 2.
Histograms show the total number of proteins quantified in ST398 and USA300 JE2 strains grown under EC1 (blue), EC2 (orange) and EC3 (green). Every bar represents the average deviation standard obtained from three biological replicates. The asterisk (*) indicates the statistically significant differences (p-value ≤ 0.05) obtained after applying the t-test.
Analysis showed no significant difference in protein numbers in the ST398 strain. A significant difference was observed in the JE2 strain between EC1 and EC2 and EC2 and EC3, partly due to the lower variability among the EC biological replicates analyzed for the JE2 strain. A high number of shared proteins were observed in all conditions for both strains, as shown in Fig. S1 (Supplementary Material 2). Unique proteins, exclusively expressed in one condition across all replicates, ranged from 1.24% in EC1-grown JE2 to 2.6% in EC2-grown ST398. These unique proteins could play a potential role in proteomic adaptations under the experimental conditions considered.
We performed multivariate statistics, specifically principal component analysis (PCA), using the normalized protein abundances quantified by Proteome Discoverer (PD). This analysis was carried out to evaluate whether the samples could be discriminated according to the growth conditions based on their proteome profile. Figure 3 shows a clear separation between the two strains associated with the first component, accompanied by a further clustering based on the three ECs related to the second component.
PCA plot of all samples analyzed in this study. A circle represents samples belonging to the JE2 strain, while a square represents those belonging to the ST398 strain. Blue, orange, and green colors mark EC1, EC2, and EC3, respectively.
Proteins associated with different growth conditions in the ST398 strain
EC2 versus EC1
Protein abundance data were further parsed to perform differential analysis between the different ECs, focusing initially on the ST398 strain. Comparing EC2 to EC1, we identified 387 differentially expressed proteins (DEPs), of which 220 exhibited significantly higher abundance and 167 showed significantly lower abundance in EC2 compared to EC1 as shown in Fig. 4A. A comprehensive list of all DEPs, including their abundance, log2-transformed abundance ratio (LogAR), biological process, and adjusted p-values, is provided in Supplementary Material 1 (ST1 sheet).
The volcano plot displays the DEPs derived from comparing EC2 and EC1 in strain ST398, highlighted in different colors based on their associated biological processes (A). The other volcano plots illustrate the biological processes with the gene names labeled for each process: ABC transporter (B), Cell redox homeostasis (C), Arginine metabolism (D), Purine metabolism (E), and Glycine-betaine biosynthesis (F). The x-axis shows the binary logarithm of the abundance ratio, while the y-axis represents the P-value as -log10.
We subsequently clustered these DEPs based on their biological processes (GO-BP), molecular functions (GO-MF), cellular component (GO-MF) and KEGG Orthology (KO) to gain insight into their roles under each growth condition. DEPs were clustered to different biological processes and pathways obtained by KEGG. As illustrated in Fig. 4B and Supplementary Material 1 (ST1 sheet), we identified nine DEPs involved in ABC transporters, seven of which were significantly more abundant in EC2. These included: iron citrate ABC transporter substrate-binding protein (HtsA), nickel import ATP-binding protein (Opp-1D), nickel import system ATP-binding protein (OppD2), peptide ABC transporter (SACOL2476), peptide ABC transporter (Opp1F), iron ABC transporter substrate-binding protein (SirA), and PTS family glucose/glucoside porter component IIABC (GlcC). The activities associated with these DEPs primarily involved ATP binding (opp1F), ion binding (SACOL2476, modA), transmembrane transport (oppD2), and enzymatic functions such as reductase (GAY51_04075), and dehydrogenases (opp-1D).
We also identified 13 DEPs associated with cell redox homeostasis (Fig. 4C and Supplementary Material 1 (ST1 sheet)), of which nine were more abundant in EC2, including pyruvate oxidase (PoxB), NADH-dependent dehydrogenase (YcjS_3), superoxide dismutase (SodA), disulfide bond protein A (DsbA), catalase (KatA), alkyl hydroperoxide reductase protein C (AhpC), peptide-methionine (S)-S-oxide reductase (msrA_2), alkyl hydroperoxide reductase subunit F (AhpF_1), and ferredoxin–NADP reductase (SAS2264). These DEPs associated with cell redox homeostasis primarily exhibited enzymatic activities, including catalase (katA), ribonucleotide synthase (dsbA), phosphatase (adh1), beta-lactamase (msrA_2), dehydrogenase (ahpC, poxB), hydrolase (SAS2264), and methyltransferase (trxB).
Additionally, we found six proteins involved in arginine metabolism (Fig. 4D and Supplementary Material 1 (ST1 sheet)), with five showing increased expression in EC2: arginine deiminase (ArcA), arginine repressor (ArcR), ornithine carbamoyltransferase (ArgF), carbamate kinase (ArcC), and urease subunit alpha (UreC). In arginine metabolism, these DEPs demonstrated amino acid (argF), NADP (arcA), and ion binding (ureC) activities, along with enzymatic functions such as carbamoyltransferase (argF) and oxidoreductase (arc).
Conversely, we identified 11 DEPs involved in purine metabolism (Fig. 4E and Supplementary Material 1 (ST1 sheet)), ten of which exhibited decreased abundance in EC2, namely phosphoribosylformylglycinamidine synthase (PurL), phosphoribosylformylglycinamidine cyclo-ligase (PurM), bifunctional purine biosynthesis (PurH), phosphoribosylformylglycinamidine synthase subunit (PurQ), phosphoribosylformylglycinamidine synthase (PurS), phosphoribosylaminoimidazole-succinocarboxamide synthase (PurC), phosphoribosylamine–glycine ligase (PurD), N5-carboxyaminoimidazole ribonucleotide mutase (PurE), purine nucleoside phosphorylase DeoD (pnp), and N5-carboxyaminoimidazole ribonucleotide synthase (PurK). These DEPs were involved in ATP (purl, purD, purC), GTP (purQ), and metal ion binding activities, as well as various enzymatic functions such as GTPase (purQ), ribonucleotide synthase (purK), IMP cyclohydrolase (purH), and purine-nucleoside phosphorylase kinase (purK, pnp).
Finally, two proteins involved in glycine-betaine biosynthesis (Fig. 4F and Supplementary Material 1 (ST1 sheet)), namely betaine aldehyde dehydrogenase (GbsA) and oxygen-dependent choline dehydrogenase (BetA), were both significantly less abundant in EC2. GbsA and BetA are involved in the activity of betaine-aldehyde dehydrogenase (NAD +).
The DEPs identified in these pathways were mainly localized in the cytoplasm (63%), membrane (33%), and extracellular region (4%) proteins.
EC3 versus EC1
By comparing EC3 to EC1, we identified 476 DEPs, with 300 showing significantly higher abundance and 176 showing significantly lower abundance in response to adding 5% NaCl (Fig. 5A). A comprehensive list of these DEPs, including their abundance, LogAR, adjusted p-values, GO-BP, GO-MF, GO-CC, and KO, is provided in Supplementary Material 1 (ST2 sheet).
Volcano plot shows the DEPs obtained by comparing EC3 and EC1 in strain ST398, marked with different colors according to the biological processes in which they are involved (A). The other volcano plots illustrate biological processes in ST398 with gene names labeled for each process: ABC transporter (B), Cell redox homeostasis (C), Lipid metabolism (D), Glycine-betaine biosynthesis (E), and Purine metabolism (F). The x-axis represents the binary logarithm of the abundance ratio, while the y-axis shows the P-value as -log10.
As with previous comparison, DEPs were grouped by biological processes and metabolic pathways. ABC transporters were represented by 12 DEPs. Eight of them, all different from those found to be increasing in the EC2 versus EC1 comparison, showed a significant decrease in response to NaCl (Fig. 5B and Supplementary Material 1 (ST2 sheet)), including molybdenum ABC transporter (ModA), ABC-type nitrate/sulfonate/bicarbonate transporter (TauA), quaternary amine transport ATP-binding protein (OpuCB), ABC transporter permease (NsaB), ABC transporter ATP-binding protein (MacB_1), heme ABC transporter ATP-binding protein (YjjK), ABC transporter permease protein (NikB), and ferrous iron transporter B (FetB). In particular, molybdenum ABC transporter (ModA) was found to be significantly more abundant in EC1 than in EC2. The DEPs linked to the ABC transporter process involved ATP (opuCB, macB_1, yjjK) binding, metal ion binding (modA), transmembrane transport (oppA, fetB, phnC, metN2), and hydrolase functions (tauA).
Additionally, several proteins related to cell redox homeostasis showed a significant decrease in response to NaCl. (Fig. 5C and Supplementary Material 1 (ST2 sheet)). Of the 20 DEPs identified, 15 were significantly decreased in EC3, including superoxide dismutase (SodA), alkyl hydroperoxide reductase subunit F (AhpF_1), peptide-methionine (S)-S-oxide reductase (msrA_2), iron-sulfur cluster regulator (iscR_1), alkyl hydroperoxide reductase protein C (AhpC), FMN-dependent NADH, oxidoreductase (AzoR), FAD-dependent oxidoreductase (G6Y24_14705), glutathione peroxidase homolog (BsaA), glutaredoxin (SAUSA300_0985), catalase (KatA_2), thioredoxin (TrxA), monooxygenase (CzcO), acetyl esterase (SAUSA300_0915), glyoxalase/bleomycin resistance protein/dioxygenase superfamily protein (SAUSA300_1458), and peroxide-responsive repressor (PerR). The first three proteins are also shared with those showing an increasing trend in EC2 when compared to EC1. These DEPs associated with cell redox homeostasis primarily exhibited enzymatic activities, including reductase (msrA_2, trxA, trxB, SAOUHSC_02460), superoxide dismutase (sodA), dehydrogenase (ipdA), peroxidase (ahpC_1, bsaA), oxidoreductase (SAUSA300_1183), catalase (katA_2), hydrolase (SAUSA300_0915) and dioxygenase (SAUSA300_1458). Some DEPs showed activities correlated with flavin adenine dinucleotide binding (czcO, ahpF_1), DNA and ion binding (perR, poxB), and electron transfer (azoR).
Furthermore, a few proteins involved in lipid metabolism were significantly decreased in response to NaCl, such as putative glycerophosphoryl diester phosphodiesterase (SAUSA300_1667), glycerophosphodiester phosphodiesterase (SAUSA300_0222), triacylglycerol lipase (Lip1), acyl carrier protein (AcpP), and lipoprotein (ST398NM01_1887) (Fig. 5D and Supplementary Material 1 (ST2 sheet)). Specifically, SAUSA300_1667 and SAUSA300_0222 were engaged in phosphoric diester hydrolase activity. When considering glycine betaine biosynthesis, two proteins that were increased in EC2 (compared to EC1), namely betaine aldehyde dehydrogenase (GbsA) and oxygen-dependent choline dehydrogenase (BetA), showed a significant increase in response to NaCl (Fig. 5E and Supplementary Material 1 (ST2 sheet)). GbsA and BetA are involved in the activity of betaine-aldehyde dehydrogenase (NAD +).
Finally, several proteins involved in purine metabolism displayed a significant increasing trend in response to NaCl (Fig. 5F and Supplementary Material 1 (ST2 sheet)), reversing their previous pattern. These included adenylosuccinate synthetase (PurA), phosphoribosylformylglycinamidine cyclo-ligase (PurM), and other enzymes involved in purine biosynthesis such as amidophosphoribosyltransferase (PurF), phosphoribosylformylglycinamidine synthase subunit PurL (PurL), phosphoribosylamine–glycine ligase (PurD), and orotidine 5’-phosphate decarboxylase (PyrF).
The DEPs in the described pathways were primarily localized in cytoplasmic (57%), membrane (38%), and cell envelope (5%) proteins (Supplementary Material 1 (ST2 sheet)).
Proteins associated with different growth conditions in the JE2 strain
EC2 versus EC1
By comparing EC2 to EC1, we identified 419 DEPs. Among these, 194 exhibited significantly higher abundance and 225 showed significantly lower abundance in EC2 compared to EC1 (as shown in Supplementary Material 1 (ST3 sheet)). Figure 6A illustrates the DEPs, that belong to several relevant biological processes and KEGG pathways.
Volcano plot shows the DEPs obtained by comparing EC2 and EC1 in strain JE2, marked with different colors according to the biological processes in which they are involved (A). The other volcano plots illustrate biological processes in JE2 with gene names labeled for each process: ABC transporter (B), Arginine metabolism (C), Lipid metabolism (D), Purine metabolism (E), Cell redox homeostasis (F), and Stress response (G). The x-axis represents the binary logarithm of the abundance ratio, while the y-axis shows the P-value as -log10.
We identified 16 DEPs categorized as ABC transporters (Fig. 6B and Supplementary Material 1 (ST3 sheet)), ten of which were significantly increased in EC2. These included ferrous iron transporter B (FetB), amino acid transporter (OpuCC_1), phosphonates import ATP-binding protein (PhnC), phenol-soluble modulin export ABC transporter (PmtA), peptide ABC transporter ATP-binding protein (QU38_16520), ABC transporter substrate-binding protein (GO814_11290), ABC transporter, permease protein (HMPREF3211_00118), phosphotransferase system, fructose-specific IIABC component (SAUSA300_2576), quaternary amine transport ATP-binding protein (OpuCA), and glycine betaine/carnitine/choline ABC transporter (OpuCC). The DEPs linked to the ABC transporter process involved ATP binding (QU38_16520, GAY51_10340, opuCA, pmtA) and ion binding (fetB), transmembrane transport (opuCC_1, opuCC, GO814_11290), ABC-type phosphonate transport (phnC), carbohydrate:proton symporter (SAUSA300_2576).
Furthermore, we identified 11 DEPs associated with arginine metabolism (Fig. 6C and Supplementary Material 1 (ST3 sheet)), with ten of them showing increased expression in EC2, namely arginine deiminase (ArcA), urease accessory proteins (UreA, UreB, UreC, UreD, UreE, UreF, UreG), ornithine carbamoyltransferase (ArgF), and arginine repressor (ArgR). The activities of these DEPs were of binding as nickel cation (ureD, ureF, ureE, ureC), GTP (ureG, ureB), arginine (arcA, argF) and of deiminase (arcA) and reductase (argC).We also found 12 DEPs involved in lipid metabolism (Fig. 6D and Supplementary Material 1 (ST3 sheet)), nine of which, namely putative lipoprotein (SAUSA300_0100), biotin carboxyl carrier protein acetyl-CoA carboxylase (accB_1), triacylglycerol lipase (Lip1), putative lipoprotein (Lpl2), tandem-type lipoprotein (Lpl5), phosphate acyltransferase (PlsX), putative glycerophosphoryl diester phosphodiesterase (SAUSA300_1667), capsular polysaccharide biosynthesis protein Cap5I (cap5I), and lipoprotein (SAUSA300_1439), exhibited increased abundance in EC2. GZ156_01970 and SAUSA300_1667 are associated with phosphoric diester hydrolase activity, as detailed by molecular function in the Gene Ontology (GO).
In addition, seven proteins related to purine metabolism showed significant changes in expression in the EC2 versus EC1 comparison (Fig. 6E and Supplementary Material 1 (ST3 sheet)). Five DEPs with enzyme activity were decreasing in EC2, including phosphoribosylformylglycinamidine synthase (PurL), bifunctional purine biosynthesis protein (PurH), N5-carboxyaminoimidazole ribonucleotide mutase (PurE), phosphoribosylglycinamide formyltransferase (PurN), and phosphoribosylformylglycinamidine synthase subunit (PurQ).
In contrast to ST398, only five proteins involved in cell redox homeostasis were significantly more abundant in EC2; instead, we observed a decrease in eight proteins (Fig. 6F and Supplementary Material 1 (ST3sheet)), including iron-sulfur cluster repair protein (ScdA), dihydrolipoyl dehydrogenase (lpdA), oxidoreductase, short-chain dehydrogenase/reductase family (SACOL2321), NAD(P)/FAD-dependent oxidoreductase (SAUSA300_1701), SDR family NAD(P)-dependent oxidoreductase (GAY51_09180), FAD-dependent urate hydroxylase HpyO/Asp monooxygenase (SAOUHSC_01890), oxidoreductase (SAUSA300_1966), and thioredoxin reductase (TrxB). These DEPs have shown activities associated with enzymatic functions such as dehydrogenase (lpdA), oxidoreductase (SACOL2321), reductase (trxB), and DNA binding (SAUSA300_1966).
In addition, we identified 28 DEPs related to the stress response (Fig. 6G and Supplementary Material 1 (ST3sheet)), with 14 of them showing increased expression in EC2. These were fluoride-specific ion channel (crcB_1), DnaK protein (DnaK), transcription-repair coupling factor (mfd_1), sensor protein kinase (WalK), DNA repair exonuclease (YhaO), Metalloregulator ArsR/SmtB family transcription factor (YozA), ATP-dependent protease subunit (HslV), protein GrpE (GrpE), replication initiation protein (RepC), endoribonuclease (YabJ), nickel/cobalt efflux system (NixA), protein VraX (VraX), small heat shock protein (M1K003_0928), molecular chaperone GroEL (GroEL), and co-chaperonin GroES (GroES). Furthermore, we identified three proteins responsible for potassium limitation (part of the two-component system) within the proteins decreasing in EC2: TrkA family potassium uptake protein (KtrA), potassium-transporting ATPase ATP-binding subunit (KdpB), and response regulator protein (KdpE).
Proteins involved in these processes and pathways were primarily cytoplasmic (58%), membrane (33%), and associated with the ATP-binding cassette (ABC) transporter complex (9%) (Supplementary Material 1 (ST3 sheet)).
EC3 versus EC1
Comparing EC3 to EC1, we identified a total of 397 DEPs. Among these, 180 exhibited a significantly lower abundance, while217 showed a significantly higher abundance in response to the addition of 5% NaCl (Fig. 7A and Supplementary Material 1 (ST4 sheet)).
Volcano plot shows the DEPs obtained by comparing EC2 and EC3 in strain JE2, marked with different colors according to the biological processes in which they are involved (A). The other volcano plots illustrate biological processes in JE2 with gene names labeled for each process: ABC transporter (B), Lipid metabolism (C), Purine metabolism (D), Cell redox homeostasis metabolism (E), and Stress response (F). The x-axis represents the binary logarithm of the abundance ratio, while the y-axis shows the P-value as − log10.
Several ABC transporters showed a decrease in response to NaCl (Fig. 7B and Supplementary Material 1 (ST4 sheet)), such as amino acid transporter (OpuCC_1), ferrous iron transporter B (FetB), peptide ABC transporter ATP-binding protein (QU38_11560), ABC-type transport system permease (NCTC7972_01260), and ABC-type transport system ATPase component (EcsA_2). The first two were also found to be increased in EC2 compared to EC1 with the JE2 strain. These proteins mainly involved in ATP binding (ecsA_2, QU38_11560) and transmembrane transport (opuCC_1, FetB).
In addition, three proteins involved in lipid metabolism were significantly less abundant and four were significantly more abundant upon exposure to NaCl (Fig. 7C Supplementary Material 1 (ST4 sheet)). Among the latter, we can mention biotin carboxyl carrier protein acetyl-CoA carboxylase (accB_1), putative lipoprotein (Lpl2), lipoprotein (LpL), and acyl carrier protein (AcpP). The first two proteins were also found to be increased in EC2 compared to EC1 (considering the JE2 strain).
Consistent with the ST398 strain, several proteins involved in purine metabolism were significantly increased in abundance in response to the addition of NaCl (Fig. 7D Supplementary Material 1 (ST4 sheet)). These DEPs include purine nucleoside phosphorylase (PgeF), phosphoribosylglycinamide formyltransferase (PurN), phosphoribosylformylglycinamidine synthase subunit (PurQ), and N5-carboxyaminoimidazole ribonucleotide mutase (PurE).
Furthermore, various proteins involved in cell redox homeostasis were affected by NaCl addition (Fig. 7E Supplementary Material 1 (ST4 sheet)). Specifically, nine out of 14 proteins exhibited a significant increase in their abundance in EC3, unlike ST398. These DEPs include NAD(P)H-dependent oxidoreductase (FrP), aldo/keto reductase (SAUSA300_2159), alkyl hydroperoxide reductase subunit F (AhpF), FAD-dependent oxidoreductase (G6Y24_14705), catalase (KatA), short-chain dehydrogenase (CpnA), thioredoxin reductase (TrxB), NAD(P)/FAD-dependent oxidoreductase (SAUSA300_1701), and FAD-dependent urate hydroxylase (SAOUHSC_01890). The last three were also found to be decreased in EC2 compared to EC1 with the JE2 strain. These DEPs are implicated in NAD binding (ahpF) and enzymatic activities, including catalase (katA), reductase (trxB, SAUSA300_2159), oxidoreductase (frp), and dehydrogenase (cpnA, G6Y24_14705).
Additionally, 13 proteins involved in stress response increased significantly their abundance in response to NaCl (Fig. 7F Supplementary Material 1 (ST4 sheet)), including heat shock protein (groES_2), acetyltransferase (SAI7S6_1019950), chaperone protein (ClpB_5), RNA polymerase sigma factor (SigB), arsenical pump membrane protein (ArsB), probable nitronate monooxygenase (SAUSA300_0825), HTH-type transcriptional regulator (SarS_1), Probable inorganic carbon transporter subunit (DabA), DEAD-box ATP-dependent RNA helicase (CshB), pyruvate kinase (pyK). Moreover, three subunits of potassium-transporting ATPase (KdpA, KdpB, and KdpC) known to participate in potassium homeostasis (and therefore likely influenced by potassium limitation), also showed increased expression following NaCl addition. The activities associated with DEPs involve ATP binding (clpB_5, cshB, groES_2, kdpB), DNA binding (sigB), ion binding (dabA), transmembrane transport (arsB, kdpA), and enzymatic activities such as N-acetyltransferase (SAI7S6_1019950) and monooxygenase (SAUSA300_0825).
Consistent with our findings in ST398 under EC3, we identified two proteins associated with glycine betaine biosynthesis that showed a significant increase in response to NaCl, namely glycine dehydrogenase (decarboxylating) subunit 1 (GcvPA) and oxygen-dependent choline dehydrogenase (BetA).
In this comparison, the DEPs examined were primarily localized in the membrane (54%), cytoplasm (42%), and ATP-binding cassette (ABC) transporter complex (4%) (Supplementary Material 1 (ST4 sheet)).
Shared features between the strains and the ECs
Differential expression analysis was further examined to identify DEPs shared between the two strains or unique to each. Comparing EC2 to EC1 in both strains, we identified 26 shared DEPs (Fig. 8A). Among these shared proteins, three were related to arginine metabolism (UreC, ArgF, ArcA), three were related to pathogenesis (SdrC, EntQ, Tet(L)), two to glycolysis/gluconeogenesis (PgK, TpI), one to lipid metabolism (SAUSA300_1667), and was one was an ABC transporter (OppD2).
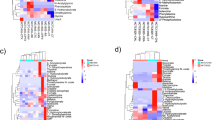
The heat map shows the proteins shared between ST398 and JE2 in EC2 compared to EC1 (A) and EC3 compared to EC2 (B). The heatmap is represented by the binary logarithm of fold change (Log2FC) for each strain.
As shown in Fig. 8B, 39 proteins showed increased expression in EC3 compared to EC2 in both the strains analyzed in this study. These proteins were primarily involved in purine metabolism (PurQ, PurN, PurE), cell redox homeostasis (TrxB, SAUSA300_1701), potassium homeostasis (KdpC), glycine-betaine biosynthesis (BetA), valine-leucine biosynthesis (IlvC), and sulfur metabolism (IscS_1).
Discussion
The persistent threat of S. aureus infections remains a significant public health challenge, stemming from the ability of this bacterium to adapt its metabolism to the environmental conditions during commensalism, colonization, and deep tissue infections25. Metabolic changes are also expected to be strain-dependent, yet few recent studies have addressed this issue26. Previous studies have employed proteomic approaches to investigate these adaptations, either by testing the effects of different antibiotic levels27 or by comparing clinical strains of MRSA28. In this study, we characterized the proteomic profiles of two different MRSA strains (ST398 and JE2) under stress conditions, first by varying the pH and temperature (EC2), as they usually fluctuate together during the infection process, and then by assessing the additive effect of saline stress (EC3). In the tested ECs, a decrease in pH and temperature was meant to mimic the "acid mantle of healthy human skin”29,30. These represent typical environmental stressors encountered by pathogens such as S. aureus. Adding NaCl to EC2 allowed us to assess the combined effect of salinity on bacterial metabolism. This condition simulates the skin environment more closely, providing insights into how S. aureus adapts its metabolic processes in response to increased salinity on top of acidic conditions31. Furthermore, S. aureus is known to possess the ability to deal with osmotic pressure and adapt to salinity stress by orchestrating a complex metabolic shift, including biofilm formation32.
Our data confirmed a reduced growth rate of MRSA strains ST398 and JE2 when exposed to lowered pH, temperature, and increased salinity, especially in combination, as previously reported33. Furthermore, our ECs led to the reassortment of several metabolic pathways. Among these, purine metabolism was affected by both conditions in both strains. Specifically, we observed a reduction in several functions required for de novo biosynthesis of IMP that, in turn, is the substrate of the growth rate-limiting enzyme inosine 5′-monophosphate dehydrogenase (IMPDH), which is required to expand the cell guanine nucleotide pool and provide an interesting target for novel antimicrobial drugs34. These data suggest a rapid metabolic shift to actively slow down bacterial replication by acting on key limiting pathways. Contrary to our findings, Yee et al. reported an increase in purine metabolism proteins in MRSA under acidic conditions, underscoring the potential strain-specific metabolic adaptations. This divergence highlights the complexity of the MRSA response to environmental stress and underscores the need for targeted studies on different strains and conditions35. Interestingly, our findings agree with those of a previous study that showed reduced levels of PurE, PurQ, and PurK proteins when S. aureus was exposed to mupirocin, an antibiotic used to treat skin infections, which induced a stress response and led to the inactivation of proteins involved in purine metabolism36. These observations also consistent with those of another study that noted a reduction in the purine metabolism pathway in S. aureus when challenged with antimicrobials37, favoring other pathways that could be more essential for stress adaptation. A further reduction in growth was observed in EC3 compared to EC1, where purine metabolism proteins were also expressed at lower levels.
Both strains showed an increase in ABC transporter protein levels under both conditions. These proteins are associated with the transport of nickel, zinc, and iron, as well as with betaine/proline/choline family members. These transport systems are crucial for coupling ATP hydrolysis to the translocation of substrates across the membrane, facilitating the uptake of nutrients that are critical for bacterial survival and pathogenesis, or that can act as osmoprotectants. Nickel, zinc, and iron transporters are vital for several biological functions, including resistance to oxidative stress and the structural components of proteins. In addition, their importance in pathogenicity has been demonstrated in previous studies 38. Conversely, transport proteins associated with betaine, proline, and choline are primarily implicated as osmoprotectants. Their increased expression at slightly acidic pH suggests an adaptive mechanism to maintain cellular integrity against environmental stress38,39. We have also identified DEPs involved in the betaine/carnitine/choline/proline metabolic pathway, such as “OpuC”, which is recognized for its role as a primary osmoprotectant, facilitating the uptake of osmolytes such as glycine, betaine, and carnitine, which aid osmotic balance. These osmolytes play a vital role in the cellular response to osmotic stress, helping to maintain fluid balance while protecting cellular structures. Furthermore, the involvement of OpuC may extend beyond osmoprotection, potentially contributing to its pathogenicity40. This response indicates the adaptation of these MRSA strains to osmotic stress, aligning with slightly acidic pH adjustments under experimental conditions.
An increase in the arginine metabolism pathway under EC2 in both strains was also observed, highlighting the importance of arginine for S. aureus growth41 and maintaining intracellular pH homeostasis to fight acid shock42. In particular, the observed overexpression of arginine deiminase (ArcA) can be attributed to its crucial role in counteracting acidic stress by increasing intracellular pH through ammonia production. This adaptation is particularly relevant for MRSA survival in the acidic conditions of wound sites, suggesting a potential for targeted therapeutic interventions aimed at disrupting this pathway43. Moreover, bacteria can utilize arginine as an energy source when grown under oxygen-competent conditions44. In the JE2 strain, the DEPs associated with the arginine biosynthesis pathway included proteins belonging to the urease complex. They function synergistically with arginine to neutralize the medium. This observation is consistent with other studies45,46,47 that showed how both urease and arginine deiminase activities increase under acidic conditions, generating ammonia to buffer the medium. Specifically, urease catalyzes the conversion of urea into ammonia and carbon dioxide through a reaction dependent on nickel cations, thus acting as a key agent in medium neutralization39. Therefore, our findings demonstrate the importance of arginine in bacterial metabolism during stress adaptation. Given the overexpression of proteins involved in the arginine biosynthesis pathway under stress conditions, targeting this pathway may offer a novel antimicrobial strategy. For instance, inhibition of arginine deiminase could render MRSA strains more susceptible to acidic environments, potentially reducing their virulence and persistence in wound infections.
An increase in glycine betaine biosynthesis was observed in both S. aureus strains upon exposure to NaCl, suggesting a coordinated response to osmotic stress. Glycine betaine, an osmoprotectant, is synthesized and accumulated by S. aureus to counteract cellular dehydration and ionic imbalance caused by high salt concentrations. This osmolyte helps maintain osmotic balance and stabilizes macromolecules, effectively enabling bacteria to adapt and survive under hyperosmotic conditions. The ability to synthesize, accumulate, and store glycine betaine appears to be crucial for the resilience of S. aureus in challenging environments48,49.
Additionally, both S. aureus strains exhibited an increase in lipid metabolism under the test’s conditions. Proteins, such as lip, lpL, AcpP, and AccB, play pivotal roles in lipid biosynthesis and phospholipid metabolism, which are essential for membrane integrity and adaptation to stress. For instance, lip and AcpP are involved in fatty acid biosynthesis, which contributes to maintaining membrane fluidity and stability under osmotic stress, whereas lpL and AccB play critical role in phospholipid metabolism and fatty acid biosynthesis, respectively, enabling the bacterium to adjust its membrane composition and support cellular growth under challenging conditions50,51,52. Lipid metabolism is integral to the bacterial stress response as it aids in stabilizing the cell membrane and maintaining cell shape during growth. The overexpression of these proteins under stress conditions thus suggests that enhanced lipid metabolism may contribute to S. aureus resilience, support membrane function, and aid survival in high-salinity environments.
A clear contrast in the redox stress responses was observed between the ST398 and JE2 strains. The ST398 strain showed increased expression of proteins associated with maintaining cell redox homeostasis in EC2, and this elevated response persisted even in the presence of osmotic stress in EC3. In contrast, the JE2 strain displayed a reduced redox response in EC2, with expression levels rising only in EC3. This suggests that ST398 may be more proactive in upregulating redox defenses, possibly enabling it to withstand oxidative stress more effectively across various environmental challenges. Essential proteins involved in redox stress management include the catalase enzyme (KatA), which plays a central role in detoxifying hydrogen peroxide (H₂O₂). Catalase is crucial for scavenging high levels of H₂O₂, thereby protecting the cell from oxidative damage and enabling bacteria to resist the toxic effects of reactive oxygen species (ROS)53,54, Additionally, AhpC and AhpF, members of the peroxiredoxin family forming the alkyl hydroperoxide reductase system, were upregulated. This enzyme neutralizes a range of peroxides, including hydrogen peroxide, organic peroxides, and peroxynitrite, thereby offering broad protection against oxidative stress55. Thioredoxin reductase (TrxB) complements these roles by recycling oxidized thioredoxin, which supports various thiol-dependent enzymes involved in antioxidant defense, DNA synthesis, and repair. Overall, KatA, AhpC, AhpF, and TrxB form a robust defense network that is crucial for S. aureus resistance to oxidative stress, particularly in stressful environments encountered during nasal colonization and infection56. Their differential regulation between ST398 and JE2 highlights a potential advantage for JE2 in tailoring redox responses specifically to stress intensity, while ST398 maintains a more consistent yet less adaptive defense across conditions. Further validation showed a similar trend between proteins involved in stress response and those involved in cell redox homeostasis. These latter were significantly more abundant under EC3 than EC2 in JE2. Conversely, they were more abundant in EC2 compared to EC3 ST398. Furthermore, increased levels of proteins involved in potassium-transporting ATPase due to potassium limitation were observed in EC1 and EC3 compared with EC2 in the JE2 strain. Potassium transport is vital for chemiosmotic homeostasis; thus, when the pH is acidic, the upregulation of potassium uptake becomes crucial for maintaining its range under pH stress59,60. This suggests that the osmotic and pH imbalance caused potassium limitation within the cell, which was compensated by increasing potassium transport to achieve that balance again.
The results of the study discussed so far demonstrate the adaptive processes employed by the two MRSA strains during the mid-log growth phase under various stress conditions. While our proteomic analysis provides valuable information, it is important to note that the stress conditions examined, although designed to mimic key aspects of the wound environment, offer insights into the evolution of the S. aureus proteome that must be verified and compared with an in vivo system capable of simulating an infectious process. Additionally, the study mainly focused on two MRSA strains; therefore, the metabolic adaptations observed may not represent the full spectrum of clinical isolates.
In conclusion, our study reveals critical adaptive mechanisms of MRSA to environmental stressors, highlighting that arginine biosynthesis is essential for acid stress and that glycine and betaine biosynthesis are important for osmotic homeostasis. Meanwhile, redox homeostasis serves as a variable factor in strain-specific responses. Further studies are necessary to understand how these pathways and the proteins highlighted in this study may represent promising targets for the development of new antimicrobial molecules.
Material and methods
Bacterial strains and growth conditions
MRSA strains ST39817 and JE218 were obtained from the Department of Veterinary and Animal Sciences at the University of Copenhagen, Denmark.
The in vitro growth of the bacteria was performed in different ECs as follows: EC1 (control condition) was performed in tryptic soy broth (TSB) at pH 7 at 37 °C, EC2 in TSB at pH 6 at 35 °C, and EC3 in TSB at pH 6 with 5% NaCl at 35 °C. Each condition involved growing both strains in three biological replicates, and the growth curves were reported as the mean of these replicates. In brief, a total 2–3 colonies of each strain were inoculated in 7 mL of TSB in a 15 mL tube and shaken at 120 RPM overnight in an incubator. Then, subcultures were prepared after 18 h by 1:66 dilution, starting at an optical density (OD)600 of 0.05 and OD600 was measured every 30 min afterwards using a UV–Vis spectrophotometer (SHIMADZU), until each culture reached the mid-log phase (OD₆₀₀ ≈ 0.5, corresponding to approximately 1 × 10⁷ CFU/mL). Although the incubation times varied between conditions, each culture was harvested when the mid-log phase was achieved, ensuring comparable physiological states across treatment groups. After reaching the target density, the cultures were immediately cooled on ice, centrifuged, and the bacterial pellets were washed three times with cold PBS. After reaching the target density, the cultures were immediately cooled on ice, centrifuged, and the bacterial pellets were washed three times with cold PBS. Bacterial counts were verified via serial dilutions in PBS and plating on tryptic soy agar (TSA) with overnight incubation. These meticulously standardized conditions allowed us to obtain reproducible proteomic samples for subsequent analysis.
Bacterial protein extraction
The proteins were extracted using an established protocol19. Briefly, bacterial pellets were dissolved in lysis buffer containing 2% sodium dodecyl sulfate (SDS), 20 mM Tris–HCl (pH 8.8), and 100 mM dithiothreitol (DTT), followed by agitation in a thermomixer (Eppendorf, Hamburg, Germany) at 600 rpm for 20 min at 95 °C. To disrupt the extracellular bacterial wall, mechanical force was applied using a TissueLyser LT mechanical homogenizer (Qiagen, Hilden, Germany). This process involved bead beating for 10 min at 30 cycles/s, combined with three freeze–thaw cycles, followed by a 15-min sonication step. Finally, the samples were centrifuged at 12,000 × g for 12 min at 4 °C to separate the bacterial proteins in the supernatant, which was then stored at -80 °C for later use.
Proteins digestion by filter-aided sample preparation (FASP)
Protein extracts were reduced with dithiothreitol (DTT), alkylated with iodoacetamide (IAA), and digested with trypsin, according to the FASP protocol to obtain peptide mixtures for mass spectrometry analysis, as previously described in20. Following digestion, peptide mixtures were collected, dried, and resuspended in 0.2% formic acid. Peptide concentrations were determined using a Nanodrop 2000 spectrophotometer (Thermo Fisher Scientific, San Jose, CA, USA) employing a HeLa protein digest standard (Thermo Fisher Scientific) to obtain a calibration curve.
LC–MS/MS analysis, data processing and statistical analysis
Peptide mixtures were analyzed by mass spectrometry using a Q-Exactive mass spectrometer coupled with an UltiMate 3000 RSLCnanoLC system (Thermo Scientific, San Jose, CA, USA). For each sample, 4 μg of the peptide mixture was concentrated and washed in a trap column (Acclaim PepMap C18, 75 μm × 2 cm nanoViper, 3 μm, 100 Å, Thermo Scientific) and then fractionated on a C18 RP column (Acclaim PepMap RSLC C18, 75 μm × 50 cm nanoViper, 2 μm, 100 Å, Thermo Scientific) using a linear gradient of 245 min from 5 to 37.5% eluent B (0.1% formic acid in 80% acetonitrile) in eluent A (0.1% formic acid)21. Spectral fragmentation was obtained by Higher Energy Collision Dissociation (HCD), with nitrogen as the collision gas. Raw MS files were subjected to database searching using Proteome Discoverer software (version 2.4, Thermo Scientific) with SEQUEST-HT as a search engine. Protein identifications were obtained using the Staphylococcus aureus 1280 database retrieved from UniProt Knowledgebase (UniProtKB), 2024_07 release), enzyme: trypsin, with two missed cleavages allowed; precursor mass tolerance: 10 ppm; MS/MS tolerance: 0.02 Da; charge states: + 2, + 3, and + 4; cysteine carbamidomethylation as static modification; methionine oxidation and acetylation (Acetyl), loss of Methionine (Met-loss) and loss of Methionine + Acetylation (Met-loss + Acetyl) on N-Terminus as dynamic modifications. Validation of identification significance was carried out using Percolator with a false discovery rate (FDR) threshold of 1%. A Consensus step was performed to compare two conditions at a time (EC1 vs. EC2 and EC2 vs. EC3), considering three biological replicates per condition. A standard consensus workflow was set on Proteome Discoverer 2.4 to evaluate label-free precursor ion quantification. The relative quantification of peptides across different samples was performed using the peak area under the curve of the detected peptide ions. This quantification was obtained by setting the “Minora feature detector node” in the processing workflow (Peak Feature Detection: min. trace length, 5; max. ΔRT of isotope pattern multiplets [min], 0.2; and Feature to ID Linking: PSM Confidence at least, high) to detect features in individual raw data files and map them to PSMs and the “Feature Mapper node” in the consensus workflow (Chromatographic Alignment: RT alignment, true; tuning, coarse; RT shift [min], 10; mass tolerance, 10 ppm; and Feature Linking and Mapping: RT tolerance [min], 0; mass tolerance, 0 ppm; min. S/N threshold, and 5) determine the most abundant features. Using the results of the Minora algorithm, the features were aligned to have the same retention time across all sample files. The abundance ratio (AR) between the two groups was log2-transformed (LogAR), and differential proteins were evaluated using a t-test (background-based) with the p-value adjusted using the Benjamini–Hochberg correction. Proteins were considered significantly differential if they had an abundance ratio of less than, or equal to − 1.2 and greater than, or equal to + 1.2 (− 1.2 ≤ Log2 AR ≥ + 1.2) with an adjusted p-value ≤ 0.0522.
Venn diagrams were generated using the web tool InteractiVenn (https://www.interactivenn.net/). At the same time, principal component analysis (PCA) was performed using the Perseus bioinformatics tool, utilizing normalized abundance data from all proteins identified through Proteome Discoverer analysis. Normalization was based on the total peptide abundance in each sample. The annotation analysis was mainly performed in Excel, integrating the Gene Ontology (GO) for biological processes (GO-BP), molecular function (GO-MF), and cellular component (GO-CC) data obtained from the UniProt Knowledgebase (UniProtKB) alongside KEGG pathway data sourced57,58 from the eggNOG-mapper 2.1.1259. Permission has been obtained from Kanehisa Laboratories to use the KEGG pathway database58. Initially, a KEGG Orthology (KO) analysis was conducted to map the proteins. However, due to limitations in the annotation of the S. aureus database, KO data were integrated and compared with GO-BP data to enhance the annotation of DEPs. As a result, proteins were primarily clustered by their GO-BP and KO data to assign the pathways. Additional analyses were subsequently performed in R23. For clustering, we focused on clusters containing at least four proteins; however, biologically significant processes involving two proteins were also considered. We focused on differentially expressed proteins (DEPs) with a higher average abundance in a specific direction for comparison and relevance to the experimental conditions. Volcano plots were generated using the ggplot2 package in R24. Heatmaps were created using R.
Data availability
The datasets generated and analyzed during the current study are available through the ProteomeXchange Consortium via the PRIDE partner repository (http://proteomecentral.proteomexchange.org)61. The proteomics data is identified by the dataset identifier PXD058815. Additionally, other data generated to support this study are included in this article and in Supplementary Material 1, Tables 1–4.
References
Otto, M. Staphylococcus colonization of the skin and antimicrobial peptides. Expert Rev. Dermatol. 5, 183–195 (2010).
Tong, S. Y. C., Davis, J. S., Eichenberger, E., Holland, T. L. & Fowler, V. G. Staphylococcus aureus infections: Epidemiology, pathophysiology, clinical manifestations, and management. Clin. Microbiol. Rev. 28, 603–661 (2015).
Pal, M. et al. Distinguished researcher gate view project inter-professional health care view project epidemiology, pathogenicity, animal infections, antibiotic resistance, public health significance, and economic impact of Staphylococcus Aureus: A comprehensive review. Am. J. Public Health Res. 8, 14–21 (2020).
Jevons, M. P. “Celbenin”—resistant Staphylococci. BMJ 1(5219), 124–125. https://doi.org/10.1136/bmj.1.5219.124-a (1961).
Lenhardt, R. & Sessler, D. I. Estimation of mean body temperature from mean skin and core temperature. Anesthesiology 105, 1117–1127 (2006).
Proksch, E. pH in nature, humans and skin. J. Dermatol. 45, 1044 (2018).
Schneider, L. A., Korber, A., Grabbe, S. & Dissemond, J. Influence of pH on wound-healing: A new perspective for wound-therapy?. Arch. Dermatol. Res. 298, 413–420. https://doi.org/10.1007/s00403-006-0713-x (2007).
Alreshidi, M. M. et al. Changes in the cytoplasmic composition of amino acids and proteins observed in Staphylococcus aureus during growth under variable growth conditions representative of the human wound site. PLoS ONE 11, e0159662 (2016).
Hajdamowicz, N. H., Hull, R. C., Foster, S. J. & Condliffe, A. M. The impact of hypoxia on the host-pathogen interaction between neutrophils and Staphylococcus aureus. Int. J. Mol. Sci. 20, 5561. https://doi.org/10.3390/ijms20225561 (2019).
Shoaib, M. et al. MRSA compendium of epidemiology, transmission, pathophysiology, treatment, and prevention within one health framework. Front. Microbiol. https://doi.org/10.3389/fmicb.2022.1067284 (2023).
Lee, B. Y. et al. The economic burden of community-associated methicillin-resistant Staphylococcus aureus (CA-MRSA). Clin. Microbiol. Infect. 19, 528–536 (2013).
Peng, B., Li, H. & Peng, X. Proteomics approach to understand bacterial antibiotic resistance strategies. Exp. Rev. Proteom. 16(10), 829–839. https://doi.org/10.1080/14789450.2019.1681978 (2019).
Tsakou, F., Jersie-Christensen, R., Jenssen, H. & Mojsoska, B. The role of proteomics in bacterial response to antibiotics. Pharmaceuticals 13, 1–27. https://doi.org/10.3390/ph13090214 (2020).
Kinross, P. et al. Livestock-associated meticillin-resistant Staphylococcus aureus (MRSA) among human MRSA isolates European Union/European Economic Area countries. Eurosurveillance https://doi.org/10.2807/1560-7917.ES.2017.22.44.16-00696 (2017).
Kadlec, K., Feßler, A. T., Hauschild, T. & Schwarz, S. Novel and uncommon antimicrobial resistance genes in livestock-associated methicillin-resistant Staphylococcus aureus. Clin. Microbiol. Infect. 18, 745–755. https://doi.org/10.1111/j.1469-0691.2012.03842.x (2012).
King, M. D. et al. Emergence of community-acquired methicillin-resistant Staphylococcus aureus USA 300 clone as the predominant cause of skin and soft-tissue infections. Ann. Intern. Med. 144, 309 (2006).
van Rijen, M. M. L., Van Keulen, P. H. & Kluytmans, J. A. Increase in a Dutch hospital of methicillin-resistant Staphylococcus aureus related to animal farming. Clin. Infect. Dis. 46, 261–263 (2008).
Fey, P. D. et al. A genetic resource for rapid and comprehensive phenotype screening of nonessential staphylococcus aureus genes. MBio https://doi.org/10.1128/mBio.00537-12 (2013).
Tanca, A. et al. A straightforward and efficient analytical pipeline for metaproteome characterization. Microbiome https://doi.org/10.1186/s40168-014-0049-2 (2014).
Pisanu, S. et al. Neutrophil extracellular traps in sheep mastitis. Vet. Res. https://doi.org/10.1186/s13567-015-0196-x (2015).
Addis, M. F. et al. Comparative secretome analysis of Staphylococcus aureus strains with different within-herd intramammary infection prevalence. Virulence 13, 174–190 (2022).
Shah, A. D., Goode, R. J. A., Huang, C., Powell, D. R. & Schittenhelm, R. B. Lfq-Analyst: An easy-To-use interactive web platform to analyze and visualize label-free proteomics data preprocessed with maxquant. J. Proteome Res. 19, 204–211. https://doi.org/10.1021/acs.jproteome.9b00496 (2019).
{R Core Team}. R: A Language and Environment for Statistical Computing (2024).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis Vol. 35, 3–10 (Springer, 2016).
Wood, T. K., Knabel, S. J. & Kwan, B. W. Bacterial persister cell formation and dormancy. Appl. Environ. Microbiol. 79, 7116–7121. https://doi.org/10.1128/AEM.02636-13 (2013).
Choueiry, F., Xu, R. & Zhu, J. Adaptive metabolism of Staphylococcus aureus revealed by untargeted metabolomics. J. Proteome Res. 21, 470–481 (2022).
Liu, X. et al. Proteomic response of methicillin-resistant S. aureus to a synergistic antibacterial drug combination: A novel erythromycin derivative and oxacillin. Sci. Rep. https://doi.org/10.1038/srep19841 (2016).
Nikolic, P. Untargeted proteomic differences between clinical strains of methicillin-sensitive and methicillin-resistant Staphylococcus aureus. Microb Pathog 179, 106121 (2023).
Costa, F. G. & Horswill, A. R. Overcoming pH defenses on the skin to establish infections. PLoS Pathog. 18, e1010512 (2022).
Lenhardt, R. & Sessler, D. I. Estimation of mean body temperature from mean skin and core temperature. Anesthesiology 105(6), 1117–1121 (2006).
Belkaid, Y. & Segre, J. A. Dialogue between skin microbiota and immunity. Science 346(6212), 954–959 (2014).
Julia M Ross, N. I. & Marten, M. R. Proteome analyses of Staphylococcus aureus biofilm at elevated levels of NaCl. Clin. Microbiol. Open Acces https://doi.org/10.4172/2327-5073.1000219 (2015).
Rode, T. M. et al. Responses of Staphylococcus aureus exposed to HCL and organic acid stress. Can. J. Microbiol. 56, 777–792 (2010).
Juvale, K., Shaik, A. & Kirubakaran, S. Inhibitors of inosine 5′-monophosphate dehydrogenase as emerging new generation antimicrobial agents. Medchemcomm 10, 1290–1301 (2019).
Yee, R., Cui, P., Shi, W., Feng, J. & Zhang, Y. Genetic screen reveals the role of purine metabolism in Staphylococcus aureus persistence to rifampicin. Antibiotics 4, 627–642 (2015).
Reiß, S. et al. Global analysis of the Staphylococcus aureus response to mupirocin. Antimicrob. Agents Chemother. 56, 787–804 (2012).
Ren, Z. et al. A general map of transcriptional expression of virulence, metabolism, and biofilm formation adaptive changes of Staphylococcus aureus when exposed to different antimicrobials. Front. Microbiol. https://doi.org/10.3389/fmicb.2022.825041 (2022).
Akhtar, A. A. & Turner, D. P. The role of bacterial ATP-binding cassette (ABC) transporters in pathogenesis and virulence: Therapeutic and vaccine potential. Microb. Pathog. 171, 105734 (2022).
Bore, E., Langsrud, S., Langsrud, Ø., Rode, T. M. & Holck, A. Acid-shock responses in Staphylococcus aureus investigated by global gene expression analysis. Microbiology 153, 2289–2303. https://doi.org/10.1099/mic.0.2007/005942-0 (2007).
kiran-et-al-2018-opuc-an-abc-transporter-that-is-associated-with-staphylococcus-aureus-pathogenesis.
Audretsch, C., Gratani, F., Wolz, C. & Dandekar, T. Modeling of stringent-response reflects nutrient stress induced growth impairment and essential amino acids in different Staphylococcus aureus mutants. Sci. Rep. https://doi.org/10.1038/s41598-021-88646-1 (2021).
Guan, N. & Liu, L. Microbial response to acid stress: Mechanisms and applications. Appl. Microbiol. Biotechnol. 104, 51–65 (2020).
Cotter, P. D. & Hill, C. Surviving the acid test: Responses of gram-positive bacteria to low pH. Microbiol. Mol. Biol. Rev. 67, 429–453 (2003).
Makhlin, J. et al. Staphylococcus aureus ArcR controls expression of the arginine deiminase operon. J. Bacteriol. 189, 5976–5986 (2007).
Zhou, C. et al. Urease is an essential component of the acid response network of Staphylococcus aureus and is required for a persistent murine kidney infection. PLoS Pathog 15, e10075385 (2019).
Bäckhed, F. et al. Defining a healthy human gut microbiome: Current concepts, future directions, and clinical applications. Cell Host Microbe 12, 611–622. https://doi.org/10.1016/j.chom.2012.10.012 (2012).
Walker, K. A. & Miller, V. L. The intersection of capsule gene expression, hypermucoviscosity and hypervirulence in Klebsiella pneumoniae. Curr. Opin. Microbiol. 54, 95–102. https://doi.org/10.1016/j.mib.2020.01.006 (2020).
Vijaranakul, U., Nadakavukaren, M. J., de Jonge, B. L., Wilkinson, B. J. & Jayaswal, R. K. Increased cell size and shortened peptidoglycan interpeptide bridge of NaCl-stressed Staphylococcus aureus and their reversal by glycine betaine. J. Bacteriol. 177, 5116–5121 (1995).
Detkova, E. N. & Boltyanskaya, Yu. V. Osmoadaptation of haloalkaliphilic bacteria: Role of osmoregulators and their possible practical application. Microbiology (N Y) 76, 511–522 (2007).
Parsons, J. B. & Rock, C. O. Bacterial lipids: Metabolism and membrane homeostasis. Prog. Lipid Res. 52, 249–276 (2013).
Perez-Lopez, M. I. et al. Variations in carotenoid content and acyl chain composition in exponential, stationary and biofilm states of Staphylococcus aureus, and their influence on membrane biophysical properties. Biochim. Biophys. Acta Biomembr. 1861, 978–987 (2019).
Zamudio-Chávez, L. et al. Staphylococcus aureus Modulates carotenoid and phospholipid content in response to oxygen-restricted growth conditions, triggering changes in membrane biophysical properties. Int. J. Mol. Sci. 24, 14906 (2023).
Horsburgh, M. J., Ingham, E. & Foster, S. J. In Staphylococcus aureus, Fur is an interactive regulator with PerR, contributes to virulence, and is necessary for oxidative stress resistance through positive regulation of catalase and iron homeostasis. J. Bacteriol. 183, 468–475 (2001).
Gaupp, R., Ledala, N. & Somerville, G. A. Staphylococcal response to oxidative stress. Front. Cell Infect. Microbiol. 2, 33. https://doi.org/10.3389/fcimb.2012.00033 (2012).
Poole, L. B. Bacterial defenses against oxidants: mechanistic features of cysteine-based peroxidases and their flavoprotein reductases. Arch. Biochem. Biophys. 433, 240–254 (2005).
Cosgrove, K. et al. Catalase (KatA) and alkyl hydroperoxide reductase (AhpC) have compensatory roles in peroxide stress resistance and are required for survival, persistence, and nasal colonization in Staphylococcus aureus. J. Bacteriol. 189, 1025–1035 (2007).
Kanehisa, M. et al. Toward pathway engineering: A new database of genetic and molecular pathways. Sci. Technol. Jpn. 59(34–38), 59 (1996).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30. https://doi.org/10.1093/nar/28.1.27 (2000).
Yang, Y. et al. Transcriptomic analysis of Staphylococcus aureus under the stress condition caused by Litsea cubeba L. essential oil via RNA sequencing. Front. Microbiol. https://doi.org/10.3389/fmicb.2020.01693 (2020).
Gries, C. M. et al. Potassium uptake modulates Staphylococcus aureus metabolism. mSphere https://doi.org/10.1128/mSphere.00125-16 (2016).
Perez-Riverol, Y. et al. The PRIDE database at 20 years: 2025 update. Nucleic Acids Res. 53, D543–D553 (2025).
Acknowledgments
We would like to thank John E. Olsen for providing the bacterial strains from the University of Copenhagen.
Funding
This research has received funding from the European Union Horizon 2020 research and innovation program under Marie Skłodowska Curie grant agreement number 956154.
Author information
Authors and Affiliations
Contributions
Conceptualization, D.AN., S.P., S.U. and D.P.; methodology, D.AN., and S.P.; investigation, D.AN. and S.P.; analysis, D.AN., S.P. and A.T.; writing—original draft preparation, D.AN.; writing—review and editing, S.P., A.T., S.U. and D.P.; visualization, D.AN.; supervision, S.P., S.U., and D.P.; resources, B.P. and A.S. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Al Nahhas, D., Pisanu, S., Tanca, A. et al. Comparative proteomics analysis of MRSA under different experimental conditions. Sci Rep 15, 28656 (2025). https://doi.org/10.1038/s41598-025-04224-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-04224-9