Abstract
Kidney diseases represent a substantial public health concern, with their incidence increasing markedly over the past decade. Addressing this challenge, our research introduces a sophisticated two-stage diagnostic model for enhancing the detection accuracy of various kidney pathologies. The initial stage of the proposed model comprises a novel Modified Specular-Free (MSF) technique designed to improve the visual quality of renal images. This technique adaptively enhances image details by applying targeted enhancements for more discrimination between dark and bright luminance levels. The objective of this technique is to restore critical image information and enrich color representation in diagnostically-significant areas. The enhanced images are then processed through the second stage, which involves classification using the EfficientNet-B2 deep learning architecture. Our model was rigorously compared against a suite of established pre-trained models, including VGG16, ResNet50, VGG19, DenseNet121, DenseNet169, DenseNet201, EfficientNet-B0, EfficientNet-B1, and EfficientNet-B3. Comprehensive testing revealed that our model not only outperforms these benchmarks, but does so with a notable accuracy of 98.27%. The robustness of the model was further ensured through its capability to effectively differentiate between normal renal conditions and various pathologies such as tumors, kidney stones, and cysts. This research not only demonstrates the potential of integrating advanced image enhancement techniques with cutting-edge classification models but also introduces a scalable approach for improving diagnostic accuracies in other complex medical imaging contexts.
Similar content being viewed by others
Introduction
The kidneys, vital organs in the human body, are reddish-brown, bean-shaped structures tasked with the crucial role of blood purification. As blood circulates through the body, it accumulates excess fluids, chemicals, and waste products. Located just below the diaphragm and behind the peritoneum, each kidney measures approximately 10 centimeters in length. These organs meticulously filter these unwanted substances from the bloodstream, facilitating their excretion through urine. Beyond waste elimination, the kidneys play a pivotal role in maintaining fluid and electrolyte balance within the body. Electrolytes, including essential minerals such as sodium and potassium, are critical for various bodily functions, influencing everything from muscle contractions to nerve signaling. The proper functioning of kidneys is thus essential not only for detoxification, but also for systemic homeostasis and metabolic regulation1,2.
Kidney diseases encompass a variety of disorders that primarily impact the nephrons, the functional units of the kidney. These conditions compromise the kidneys’ ability to efficiently remove waste and excess fluids, leading to their harmful accumulation in the body3. Kidney diseases vary widely, both in their causes and the duration of their effects. Among the most prevalent and significant are chronic kidney failure, a condition in which the kidneys progressively lose their filtering capacity, often leading to severe long-term health implications. Kidney cysts, another major form, are typically hereditary and can become problematic if multiple cysts develop, potentially impairing kidney function. Kidney stones are solid masses of crystalline deposits that form in the kidneys, renowned for causing excruciating pain. Additionally, kidney tumors represent a serious health concern, as these abnormal tissue growths can be indicative of cancerous developments. Understanding the diversity and impact of these renal diseases is crucial for effective diagnosis and treatment strategies.
Ultrasound stands as a pivotal diagnostic tool in modern medicine, particularly in the identification and management of various diseases. As a non-invasive medical imaging technique, ultrasound depends on high-frequency sound waves to create detailed visualizations of the internal organs. Specifically, a kidney ultrasound provides invaluable insights into the kidneys by producing images that reveal their size, shape, and location. This diagnostic test is crucial not only for assessing the physical attributes of the kidneys but also for evaluating blood flow to these organs, which is essential for their proper functioning. Furthermore, ultrasound is instrumental in diagnosing a range of kidney diseases, including renal edema, a condition characterized by swelling due to fluid retention; polycystic kidney disease, which involves the growth of numerous cysts within the kidneys; and kidney stones, which are solid accumulations of minerals and salts. These capabilities make ultrasound an essential technique in nephrology, offering a critical window into renal health and aiding in early detection and treatment of renal pathologies4,5.
Medical image classification stands as a cornerstone in the realm of image analysis, particularly within the healthcare sector. This critical process involves the labeling and categorization of pixel groups within images according to specific, predefined criteria. The primary aim is to differentiate between varied image sets, thereby aiding medical professionals in accurate disease diagnosis and facilitating advanced research. The process of classification typically unfolds in two distinct stages: feature extraction and classification. During the feature extraction stage, key attributes of the images, whether spectral, such as color and brightness; or textural, including smoothness and uniformity, are identified and isolated. These features are then utilized in the classification stage, where they are systematically analyzed and categorized based on established rules. This structured approach enables the precise identification of medical conditions from imaging data, enhancing the diagnostic processes and contributing significantly to the field of medical research6,7.
Deep Learning (DL) networks, particularly Convolutional Neural Networks (CNNs), have emerged as the premier choice for classifying complex medical images, playing a crucial role in accurate disease diagnosis and effective treatment planning. CNNs, a specialized form of neural networks optimized for data processing, are structurally composed of input and output layers interspersed with multiple hidden layers. These hidden layers include convolutional layers that depend on various filters for feature extraction, pooling layers that reduce the dimensionality of the data while preserving essential information, and fully-connected layers that integrate these features into a holistic understanding. In the context of medical imaging, the input image is processed through these layers to extract significant features, which are then analyzed in the classification stage. This stage effectively utilizes the distilled features to categorize images into predefined classes, thereby facilitating precise medical diagnoses. The adaptability and efficiency of CNNs in handling spatial hierarchies and complexities in data make them exceptionally suited for medical applications, where detailed and accurate image analysis is paramount8,9.
Research gap: Despite advances in medical imaging and the adoption of DL models such as CNNs, significant challenges remain in accurately detecting and classifying kidney diseases. Existing approaches often struggle with image quality issues, variability in pathological presentation, and limited specificity when distinguishing between visually similar conditions (e.g., cysts versus tumors). Additionally, many models are not optimized for real-time or resource-constrained environments, limiting their clinical utility.
This study identifies a specific research gap in the development of robust, accurate, and computationally efficient diagnostic tools that can integrate advanced image enhancement techniques with DL architectures to improve the classification of kidney diseases from CT images. The proposed two-stage model directly addresses this gap by combining MSF imaging for enhanced visual clarity and EfficientNet-B2 for high-performance classification.
Research motivation: Kidney diseases, including chronic kidney disease (CKD), pose a significant global health burden, affecting over 850 million people worldwide. Early detection is critical for improving patient diagnosis outcomes, yet traditional diagnostic approaches, particularly manual interpretation of CT or ultrasound images, are often limited by image artifacts, variability in interpretation, and challenges in detecting subtle pathological changes.
These limitations emphasize the need for automated, intelligent diagnostic systems that can operate reliably across diverse imaging conditions. Our study is motivated by the opportunity to bridge this gap through the development of a two-stage diagnostic framework that first enhances image quality using the MSF technique to suppress reflection artifacts and clarify tissue structures, followed by classification with EfficientNet-B2, a DL model known for its strong accuracy-to-complexity ratio.
This integrated approach not only addresses existing diagnostic challenges, but also offers a scalable and computationally-efficient solution suitable for real-time use in clinical environments, supporting improved accuracy and consistency in kidney disease detection.
Novelty and contributions: This paper makes several key contributions to the field of medical image analysis and kidney disease diagnostics. Firstly, it introduces an innovative two-stage diagnostic model that combines modified specular-free imaging with the DL process of EfficientNet-B2. This approach not only enhances image quality but also significantly improves classification accuracy. Secondly, the paper presents a comprehensive comparison of the proposed model against established CNN architectures, demonstrating superior performance in detecting and classifying various kidney diseases. Lastly, the findings contribute to the body of knowledge by illustrating the effectiveness of integrating advanced imaging enhancements with cutting-edge classification technologies, potentially setting a new standard for medical imaging practices in nephrology. The key novel aspects of our proposed method are:
-
1.
Modified Specular-Free (MSF) Imaging Technique: We introduce a novel enhancement technique specifically designed to reduce specular reflections and enrich color representation in renal CT images, thereby improving image quality in diagnostically-critical regions.
-
2.
Integration with EfficientNet-B2: We present a two-stage pipeline that couples the MSF technique with EfficientNet-B2, a DL model selected for its optimal balance between accuracy and computational efficiency, to achieve robust kidney disease classification.
-
3.
Superior Performance Across Benchmarks: A comprehensive evaluation is conducted against nine state-of-the-art CNN architectures, where our model achieves the highest classification accuracy of 98.29%, outperforming VGG16, ResNet50, DenseNet variants, and other EfficientNet variants.
-
4.
Advanced Preprocessing Pipeline: We apply additional enhancement techniques such as Low Illumination Map Estimation (LIME) and histogram-based segmentation to adaptively enhance dark and bright regions, which further improve classification reliability.
-
5.
Error and Statistical Analysis: Extensive validation, including error analysis (false positives/negatives), spectral entropy, NIQE, and ablation studies, underscores the robustness and diagnostic potential of our model in real-world applications.
The rest of this research is organized into key sections as follows. Section "Related work" introduces the summary of the related studies. Section "Proposed dual-phase model for kidney image enhancement and classification" details enhancement techniques and classification processes. Section "Dataset description and hyperparameter tuning" describes the data source and structure. Section "Simulation results" presents the model performance metrics and comparative analyses. Section "Conclusion and future research directions" summarizes the findings and presents future research possibilities.
Related work
Extensive research has been conducted over the years to improve the diagnosis and treatment of kidney diseases, yielding promising results and contributing to a reduction in previously alarming statistics. Despite these advances, the incidence of kidney diseases continues to rise, underscoring the need for further innovation in the diagnosis of these diseases. Notably, a significant shift occurred in the middle of the last decade, with research increasingly focusing on disease prevention and early diagnosis. Early detection plays a crucial role in mitigating the progression of kidney diseases, and consequently, in reducing their overall prevalence. Recognizing the importance of this strategic pivot, this study seeks to harness the capabilities of DL techniques. The objective is to refine the accuracy and timeliness of kidney disease diagnosis, thereby equipping medical specialists with the tools necessary for effective treatment. By leveraging advanced computational models, particularly CNNs, this research aims to set new benchmarks in the early and precise identification of renal pathologies, ultimately influencing treatment outcomes and enhancing patient care.
Patro et al.10 advanced the field of medical imaging for kidney stone detection by proposing an innovative model that integrates the Kronecker product-based convolution technique with DL methodologies. This approach uniquely combines elements of traditional radiographic image processing with modern computational techniques to classify kidney stones from coronal CT scans, particularly those of low quality. Initially, the model depends on the application of an embossing filter on the input images to accentuate the distinction between the foreground and background, enhancing feature visibility that is crucial for accurate classification. Subsequent steps involve cropping the images based on the region of interest to focus on relevant areas, thereby optimizing the analysis. To further enhance the model robustness, data augmentation techniques are employed, including rotation of images by 30 degrees, horizontal flipping, shearing by 0.3, and zooming by 0.3. These procedures not only increase the dataset size but also ensure the model efficacy across different imaging conditions, significantly improving the accuracy of kidney stone detection.
Abraham et al.11 made significant strides in the predictive analysis of kidney stone composition by utilizing the XGBoost Machine Learning (ML) algorithm. Their model leverages extensive data from 24-hour urine collections along with clinical patient details to predict the chemical composition of kidney stones, thereby facilitating targeted treatment strategies. This innovative approach demonstrates the power of ML in transforming diagnostic capabilities and personalizing patient care based on predictive analytics. On a related front, El Beze et al.12 developed an automatic stone detection system that capitalizes on the advanced imaging capabilities of endoscopy. Their system is designed to accurately differentiate between six distinct types of urinary calculi by analyzing both the surface and cross-sectional details of the stones. This methodology enhances the precision of stone type identification, which is crucial for determining the most effective treatment protocols. Together, these studies exemplify the progressive integration of ML and detailed imaging techniques in the realm of nephrology, significantly advancing the accuracy and specificity of kidney stone diagnostics.
Abdeltawab et al.13 pioneered the development of an automated CT kidney classification system, employing a sophisticated DL framework. Their system depends on a tripartite architecture consisting of three distinct CNNs, each designed to process kidney image patches of different dimensions. This modular approach allows for the granular analysis of the CT images, facilitating both patch-wise and pixel-wise classification with high precision. By splitting slide kidney images into patches of three different sizes, each network is specialized to optimize the detection and classification of features according to its specific scale. The comprehensive dataset used for training and validating this system includes 64 histopathological kidney images, enabling the model to learn a diverse array of kidney conditions. The precision of this system marks a significant advancement in the use of deep learning for medical image analysis, offering potential improvements in the diagnostic accuracy and speed for kidney-related ailments. In another work14, SEGSRNet addressed the challenge of precisely identifying surgical instruments in low-resolution stereo endoscopic images, a common issue in medical imaging and robotic surgery. The authors’ innovative framework enhanced image clarity and segmentation accuracy by applying state-of-the-art super-resolution techniques prior to segmentation. The model outperformed existing methods based on metrics such as Dice, IoU, PSNR, and SSIM, and produced clearer and more accurate images for stereo endoscopic surgical imaging. In a related study, the authors explored the role of self-supervised transformers in organ classification from low-resolution medical images. We discussed how transformer-based models, such as ViT and Swin Transformer, were also evaluated in our study, but were outperformed by EfficientNetB2 in terms of both accuracy and computational efficiency.
In another work15, the authors used more than 4,000 CT scans obtained from the large-scale SCAPIS and IGT cohorts to train and evaluate four convolutional neural network architectures, namely ResUNET, UNET++, Ghost-UNET, and Ghost-UNET++. These models were developed for fully-automated segmentation using a 3-slice CT imaging protocol comprising single slices at the levels of the liver, abdomen, and thigh, thereby enabling detailed analysis of numerous tissues and organs. The segmentation techniques were evaluated for the automated segmentation of the liver, spleen, skeletal muscle, bone marrow, cortical bone, and various adipose tissue depots, including visceral (VAT), intraperitoneal (IPAT), retroperitoneal (RPAT), subcutaneous (SAT), deep (DSAT), and superficial SAT (SSAT), as well as intermuscular adipose tissue (IMAT). The models were trained and validated for each target using tenfold cross-validation and independent test sets. In addition, the authors presented a hybrid DL framework for kidney disease classification. They adopted a multi-branch CNN strategy that complemented our approach, but we highlighted key differences, particularly in our use of advanced image enhancement techniques (MSF and LIME) and the EfficientNet-B2 architecture, which offered improved computational efficiency and diagnostic performance.
The classification of kidney diseases has garnered considerable attention in the scientific community, particularly with the advent and integration of ML and DL algorithms in medical research. Recent years have witnessed a surge in interest towards employing DL models due to their superior ability in handling complex datasets and producing highly-accurate diagnostic results. This shift has been motivated by the urgent need for early detection and effective treatment of kidney diseases, areas where DL models have shown significant promise. Numerous recent studies have leveraged these technologies to enhance diagnostic precision and speed, thereby improving patient outcomes. These studies underscore the effectiveness of ML and DL in identifying and classifying various kidney pathologies at earlier stages, facilitating timely and targeted therapeutic interventions. The impact and findings of these research efforts are systematically documented in Table 1, which provides a detailed comparison and summary of the methodologies and outcomes associated with the use of advanced computational models in the diagnosis of kidney diseases16,17,18,19,20,21,22,23,24,25,26,27,28,29,30,31,32,33,34. So, Table 1 highlights the diversity of approaches and their respective performance metrics, illustrating significant advancements in the field.
Despite the significant progress achieved in kidney disease classification using DL, several key challenges remain unresolved. Many CNN- and transformer-based models rely heavily on high-quality input images and often struggle with specular reflections, shadowing, or poor contrast, issues commonly encountered in clinical CT imaging. Moreover, while transformer architectures such as ViT and Swin Transformer offer enhanced global feature modeling, they are computationally intensive and less practical for real-time or resource-constrained environments. Importantly, few existing approaches have integrated preprocessing strategies, such as specular reflection suppression or illumination correction, into the classification pipeline, limiting their effectiveness in handling diagnostically-compromised images.
These limitations underscore the need for a robust and efficient framework that enhances image quality, while maintaining diagnostic accuracy and computational feasibility. Motivated by these gaps, our study presents a two-stage model that combines a novel MSF image enhancement technique with a lightweight yet powerful EfficientNetB2 classifier. This integrated approach addresses both image-level and model-level deficiencies in prior work, enabling more accurate detection of kidney pathologies across diverse imaging conditions.
The core contributions of our research are both methodologically and clinically significant:
-
We propose an MSF enhancement technique that removes specular highlights and enriches color fidelity in diagnostically-critical regions, resulting in clearer, more informative CT images35,36.
-
The LIME algorithm is applied to generate illumination maps that enhance visibility in low-light areas, helping reveal subtle anatomical features.
-
Further enhancement is performed by analyzing the maximum pixel intensity across RGB channels and applying histogram-based segmentation to selectively amplify under- and over-exposed regions.
-
These preprocessing steps enable a more nuanced visual representation, improving feature distinction across kidney structures.
-
For classification, we introduce a customized EfficientNet-B2 model, optimized for renal image analysis. This model demonstrates superior performance in distinguishing between normal tissue, cysts, tumors, and stones, while maintaining low inference time and number of parameters.
-
Collectively, the proposed enhancements and classification architecture significantly advance the field of kidney disease diagnostics by enabling early, accurate, and computationally-efficient detection, offering real-world applicability in clinical practice.
Proposed dual-phase model for kidney image enhancement and classification
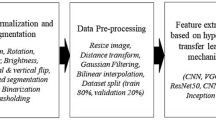
In the proposed model, we delineate a structured methodology divided into two crucial phases, as shown in Fig. 1, namely enhancement and classification. These stages are designed to significantly boost the accuracy and reliability of diagnoses for various kidney diseases. The first phase, the enhancement phase, incorporates an MSF technique. The MSF technique is an image enhancement technique designed to eliminate specular highlights, bright reflection artifacts often present in medical and computer vision images. By suppressing these reflections and enhancing the diffuse component of the image, the MSF improves visual clarity and significantly enhances the performance of subsequent recognition and analysis tasks. Figure 2 presents the flowchart illustrating the MSF process, outlining each step involved in the removal of specular reflections. This technique is meticulously developed to counteract the challenges posed by specular reflections commonly encountered in medical imaging, while simultaneously enriching the color depth of vital regions within the images. To achieve this objective, images are initially categorized based on their illuminance levels into ‘dark’ and ‘bright’ groups. This categorization facilitates the application of a specifically-tailored enhancement factor to each group, optimizing the visibility and clarity of details critical for accurate diagnostic interpretation. The resulting intermediate images from this stage display markedly improved features, with enhanced contrast and sharpness, making them more suitable for subsequent diagnostic analysis.
Block diagram of the proposed model for kidney image enhancement and classification.
Flowchart of the MSF technique.
The strategic segmentation and tailored enhancement are pivotal in preparing the images for the next phase, ensuring that the enhanced images retain all necessary pathological details required for effective disease classification. By refining image quality at this initial stage, the model sets a robust foundation for the advanced analytical processes that follow in the classification stage.
The classification stage of our proposed model capitalizes on the capabilities of the EfficientNetB2 architecture, a choice driven by its renowned efficiency and superior accuracy in handling complex image classification tasks. This model is adept at processing both the pre-processed and specifically-enhanced images produced during the earlier enhancement stage. By doing so, it allows us to rigorously assess the incremental benefits our enhancement techniques bring to the overall diagnostic performance.
To establish the efficacy of the EfficientNetB2 model within our framework, we undertake a comprehensive comparative analysis against a suite of advanced pre-trained models widely recognized in the medical imaging field. These models include VGG16, ResNet50, VGG19, DenseNet121, DenseNet169, DenseNet201, as well as variants of EfficientNet such as EfficientNet-B0, EfficientNet-B1, and EfficientNet-B3. Through this comparative approach, we aim not only to demonstrate the enhanced accuracy and reliability of our model in classifying a range of kidney diseases but also to underscore its adaptability and potential utility in real-world clinical settings.
This phase of the study is crucial for substantiating the improvements our model proposes over existing methods. By meticulously analyzing and documenting the performance metrics across these models, we provide empirical evidence of our model superiority in enhancing diagnostic accuracy. These results highlight the transformative impact of integrating advanced image enhancement with robust DL architectures, setting a new benchmark for precision in kidney disease diagnosis.
To further clarify the procedural flow of our proposed two-stage framework, we provide the following pseudocode representation. It outlines the sequential steps involved in image enhancement using the MSF technique, followed by illumination correction and classification using EfficientNet-B2. This abstraction enhances the reproducibility of our method and offers a clear, high-level overview of the algorithmic logic behind the system (Algorithm 1).
Kidney disease detection using MSF and EfficientNet-B2
Enhancement phase
The enhancement technique proposed in this research work is specifically designed to target and improve dark and bright regions within kidney images, crucial for enhancing diagnostic accuracy. Initially, the MSF technique is applied to effectively reduce specular highlights and concurrently enrich the color of important anatomical regions33,34. Unlike traditional histogram-based methods, such as CLAHE and Adaptive Histogram Equalization (AHE), which enhance global contrast without selectively addressing artifacts, the MSF technique is specifically designed for medical imaging scenarios in which specular highlights often obscure diagnostically-important structures. The MSF operates by directly suppressing specular reflections based on chromaticity and illumination modeling, thereby preserving the true color composition and structural integrity of soft tissues. This targeted enhancement minimizes the risk of over-enhancement or feature distortion, making it more suitable for high-precision tasks such as kidney pathology classification. By maintaining local details and avoiding contrast saturation, the MSF ensures that subtle abnormalities, which are often critical in early diagnosis, remain visible and distinguishable in the enhanced images.
This initial processing step divides the images based on their luminance levels into two categories: dark and bright. The category of each image determines the specific enhancement parameter that is applied in conjunction with the MSF technique to adjust and optimize the image quality.
Subsequently, the LIME algorithm is utilized to further refine the image by assessing and adjusting the illumination map. This is achieved through the formula:
where represents the color channel (Red, Green, or Blue), and denotes an individual pixel within the image. This formula ensures that the maximum value from each color channel is selected for every pixel, thereby enhancing the intensity both locally and globally across the image. This approach not only improves visibility of critical features within the image but also enhances the overall image quality, making it more conducive for subsequent diagnostic processes35,36.
In our enhancement algorithm, the specular components, typically observed as white spots due to reflection, are crucially addressed. To generate a Specular-Free (SF) image, we first identify and subtract the smallest value among the R, G, and B channels from each corresponding channel for every pixel in the image. This operation effectively removes specular reflections, while retaining the structure and color balance of the underlying image details.
The Modified Specular-Free (MSF) image is then produced by adjusting the SF image with an offset that reintroduces some of the lightness removed in the specular elimination step. The offset is calculated as the average of the smallest values subtracted during the SF image creation:
where (x, y) are the coordinates of the pixel, and denotes the color channels (r, g, b). The input image values for R, G, and B at each pixel location are represented by Ic(x, y), and min(Ir(x,y),Ig(x,y),Ib(x,y)) gives the minimum value among the RGB channels at each pixel.
The offset is defined as:
with representing the average operation. This modified offset helps to restore details in highlighted areas, while preserving the integrity of shadows and darker regions, achieving a balanced enhancement across the image.
To further refine the image based on its luminance characteristics, we employ a histogram-based approach to classify the image into bright and dark types. By analyzing the pixel distribution across 256 luminance levels, we calculate the following items:
where histi is the histogram value at each luminance level i, and th1, th2, th3, th4 are threshold values defining the ranges for bright and dark regions. The classification into dark or bright types is determined based on whether the bright region (Br) is more than twice the dark region (Bi), providing a dynamic basis for subsequent processing steps.
For the intermediate enhanced image, denoted as Intr_en(x, y), the enhancement process depends on the luminance classification of the image, either as bright or dark type. The thresholds used for classification, th1, th2, th3, and th4, are set uniformly across the different types to maintain consistency in the enhancement process. The formulation for the intermediate enhanced image is defined based on these classifications:
-
For images classified as bright type:
-
For images classified as dark type:
After the creation of the intermediate image, the LIME algorithm is applied. This involves deriving an illumination map by selecting the maximum value from the RGB channels of Intr_en. The purpose of applying the LIME algorithm is to enhance the global brightness and contrast of the image, particularly focusing on improving the visibility in both highly-illuminated and low-lit areas.
To further enhance the image, High Dynamic Range (HDR) techniques are employed as follows:
where and values are adjusted to 2.2 and 1.25, respectively, to optimize the image for diagnostic purposes37. This HDR formula adjusts IIntr_en(, ), effectively enhancing the visibility of details across different lighting conditions in the image.
The final HDR image is calculated using the formula:
Here, Inter_en(, ) represents the illumination map, helping to balance the exposure and contrast of the image, thus ensuring that both high-illumination and low--illumination regions are accurately represented. This technique significantly enhances the contrast and color fidelity of the images, making them ideal for detailed medical analysis.
Classification phase
Building upon the enhanced image quality achieved in the previous phase, we now turn to the classification process using the EfficientNet-B2 architecture. The utilization of DL networks, particularly CNNs, has significantly transformed the landscape of image analysis for classification tasks across a variety of fields. CNNs are adept at decoding the complex visual properties of images, including textures, patterns, and shapes, thanks to their convolutional layers. These layers autonomously extract vital features from images, forming the foundational elements for robust image detection and classification systems38,39,40,41.
In the realm of kidney disease diagnosis, our proposed classification model depends on EfficientNet architecture, a cutting-edge CNN variant specifically engineered to optimize network depth, width, and resolution, simultaneously. EfficientNet has 8 variants (B0 to B7), each designed to balance performance and computational efficiency. Among its iterations, EfficientNet-B0 and B1 were initially considered for their smart scalability strategy, which processes visual data with heightened efficiency. This model excels in the domain of kidney disease detection by effectively identifying unique visual characteristics inherent in medical images. However, for our specific task of kidney disease classification, EfficientNet-B2 was ultimately selected.
EfficientNet-B0 is well known for its integration of precision and computational efficiency, making it an ideal choice for medical image classification tasks where both accuracy and processing speed are paramount. Building on the strengths of EfficientNet-B0, the EfficientNet-B1 model offers advancements in detecting and extracting distinctive image characteristics with even greater accuracy. This model achieves a superior balance between performance and complexity, providing enhanced adaptability, which makes it more suitable for complex diagnostic challenges. The evolutionary progression from EfficientNet-B0 to EfficientNet-B1 demonstrates significant improvements in handling the nuanced demands of medical image analysis.
Figure 3 illustrates the EfficientNet model architecture, showcasing the incremental enhancements that each version brings to the field of kidney disease detection and classification. This visual representation helps underscore the technological advancements encapsulated within the EfficientNet series, highlighting their potential to revolutionize diagnostic practices in nephrology.
Architecture of the developed EfficientNet.
EfficientNet architecture is enhanced by incorporating a variant known as Mobile Inverted Residual Bottleneck Convolution (MB Conv). This innovative feature is critical for enhancing the model performance by systematically scaling its depth, width, and resolution. MB Conv depends on an inverted residual structure in which the input and output are thin bottleneck layers, while the expansion layer is thicker, allowing more complex features to be learned without a substantial increase in computational demand. This approach optimizes the network efficiency, enabling it to process higher-resolution images more effectively without a commensurate increase in computational cost. The adoption of MB Conv in EfficientNet is a testament to its design philosophy, which emphasizes optimal scaling across different dimensions of the network to achieve a state-of-the-art performance in image classification tasks, including the intricate domain of kidney disease detection38.
The proposed model, EfficientNet-B2, represents a significant advancement in the EfficientNet series, specifically tailored for the classification of kidney images. The selection of EfficientNet-B2 among the 8 variants (B0 to B7) was based on several factors:
-
Performance Metrics: EfficientNet-B2 offers a superior balance between accuracy and computational efficiency compared to its counterparts. Preliminary tests showed that B2 achieves higher accuracy rates than those of B0 and B1, while maintaining manageable computational costs.
-
Computational Efficiency: EfficientNet-B2 provides an optimal trade-off between model complexity and computational resources, making it suitable for our extensive dataset and the need for real-time diagnostic applications.
-
Scalability: While B3 to B7 offer even higher accuracy, they come with increased computational demands. EfficientNet-B2 was found to be the most scalable option for our available resources without compromising significantly on accuracy.
-
Task Specificity: The architectural enhancements in EfficientNet-B2, such as increased depth and resolution, are well-suited to capture the intricate details necessary for accurate kidney disease classification.
These considerations led to the selection of EfficientNet-B2 as the most appropriate model for our research. Therefore, as an evolution of earlier models, EfficientNet-B2 incorporates increased depth, width, and resolution, enhancing its capability to discern finer details and complexities in medical imaging. This model stands out due to its refined ability to distinguish between various kidney diseases, accurately defining and extracting complex characteristics from images with enhanced precision.
EfficientNet-B2 is architecturally sophisticated, featuring six convolutional blocks that progressively extract and refine features from the input images, an output layer for final classification, and one Global Average Pooling (GAP) layer that helps reduce spatial dimensions, while retaining important information. Additionally, it includes two soft-max layers for output classification, four squeeze and excitation blocks that adaptively recalibrate channel-wise feature responses, and ten Depth-Wise Separable Convolution (DWSC) blocks. DWSC is particularly noteworthy as it reduces the model computational burden by performing spatial convolutions independently across each input channel, significantly lowering memory requirements and enhancing the computational efficiency of the model. The formula for the convolution operation dimensions in DWSC blocks is given by:
where C represents the number of channels, Dk is the kernel size, and Dp is the dimension of each feature map. The input images are processed in three separate channels, Red (R), Green (G), and Blue (B), allowing the model to handle complex visual information, effectively.
EfficientNet-B2 superior performance in kidney disease classification not only demonstrates its technical prowess but also underscores its potential to significantly enhance the accuracy and efficiency of medical diagnostics. This model is poised to make a substantial contribution to the field, improving the capabilities of medical professionals in diagnosing kidney ailments with unprecedented accuracy and speed.
EfficientNet-B2 depends on a multi-faceted approach in its convolutional blocks, where five filters of varying dimensions, 32, 64, 128, 512, and 1024, are systematically applied to enhance feature extraction capabilities across different scales. Each convolutional block is further refined with the integration of squeeze and excitation (SE) modules, which dynamically adjust channel-wise feature responses, thereby improving the representational power of the network.
Block diagram of the EfficientNet-B2 model.
The architecture is designed to handle input blocks with dimensions denoted by (B, C, H, W), where and represent the height and width of the feature map, respectively, while and denote the batch size and number of channels. This structure allows for meticulous spatial processing of images. After the initial convolution operations, a 3 × 3 filter is employed. It is then paired with a batch normalization layer to stabilize learning and an average pooling layer to reduce spatial dimensions, thereby enhancing computational efficiency.
The operations within the EfficientNet-B2 architecture, including the use of DWSC, are strategically optimized to lower computational cost and processing time. The DWSC helps minimize the computational load by performing lighter, channel-wise convolutions followed by a pointwise convolution that blends the channel outputs, reducing both the number of parameters and computational complexity.
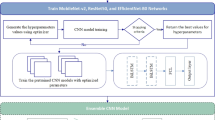
The training of the model is conducted on a carefully-curated dataset with a learning rate of 0.0001 over 100 epochs, ensuring gradual and stable convergence to optimal weights. Figure 4 illustrates the block diagram of EfficientNet-B2, providing a visual representation of its intricate architecture and operational flow42,43,44,45,46,47.
This process not only underscores the architectural efficiency of EfficientNet-B2, but also highlights its robustness in processing of kidney images, making it a potent tool for medical image classification. Through strategic enhancements in its architecture and thoughtful training protocols, EfficientNet-B2 is poised to deliver exceptional performance in the classification of kidney diseases.
Dataset description and hyperparameter tuning
The dataset used in this study was obtained from a publicly-available Kaggle repository, and it comprises 12,446 annotated kidney CT images, categorized into four diagnostic classes: normal (5,077 images), cyst (3,709), tumor (2,283), and kidney stone (1,377). These images were collected from various hospitals and imaging centers, ensuring representation of diverse imaging conditions and clinical scenarios. All subjects included in the dataset had confirmed diagnoses corresponding to one of the four categories, providing a reliable foundation for supervised learning.
Where available, demographic metadata indicated that patients ranged in age from 25 to 75 years, with a near-equal distribution of male and female subjects. However, information on ethnicity and geographic origin was limited, and there appears to be a regional concentration of data, which may introduce bias. Such demographic limitations could impact the generalizability of the trained model when applied to broader, more diverse populations. In particular, under-representation of certain ethnic groups or age cohorts may affect model performance across those subgroups.
To address this issue, we acknowledge the need for external validation using independent datasets from multiple institutions and imaging platforms. Future work will involve evaluating the model performance on clinically-diverse datasets to ensure robustness, fairness, and applicability in real-world healthcare environments.
Prior to model training, all images underwent a standardized preprocessing pipeline. This included the application of the MSF technique to reduce specular reflections, LIME-based illumination correction to enhance visibility in low-light regions, and histogram-based segmentation to classify images into dark and bright types for targeted enhancement. To improve generalization and robustness, we implemented data augmentation techniques during training. Random rotations (± 30°), horizontal and vertical flips, contrast and brightness shifts, and the addition of Gaussian noise were considered. These transformations helped increase dataset diversity and reduce overfitting. Hyperparameters were tuned via grid search across different learning rates, batch sizes, and optimizers. The final configuration depended on the Adam optimizer with a learning rate of 0.0001, a batch size of 16, and a total of 100 epochs with early stopping based on validation loss.
For model training, validation, and evaluation, the dataset is strategically divided into training, validation, and testing sets. Specifically, 70% of the data (approximately 8,712 images) is allocated for training, 10% (approximately 1,245 images) for validation, and the remaining 20% (approximately 2,489 images) is reserved for testing. The testing set was not applied during the training process and served solely for evaluating the model performance. The distribution of images for each class in the training, validation, and testing sets are given in Table 2. Thus, Table 2 provides a clear and organized view of the dataset composition, illustrating the allocation of images across different kidney conditions, which is crucial for training the model to recognize and differentiate between these different types, effectively. This distribution allows for extensive training of the DL model, while using the validation set to tune hyper parameters and select the best weights, thereby avoiding overfitting. The model is trained for 100 epochs with early stopping based on the validation loss to ensure robust learning and generalization.
We have also ensured that the testing set in this study is comparable to those used in other studies in terms of the number of images, facilitating a fair comparison of the results.
Figure 5 shows samples of the original images, and Fig. 6 illustrates samples of the enhanced images. These images clearly illustrate the improvements in image quality and details achieved by applying the MSF technique.
Samples of the original images.
Samples of the enhanced images.
One of the significant challenges faced in this study was the acquisition of a comprehensive kidney dataset. Access to medical imaging data is often restricted due to privacy and ethical considerations, making it difficult to obtain a large and diverse set of kidney images for research purposes. Additionally, the quality and consistency of the images can vary significantly depending on the imaging equipment and protocols used across different medical institutions. This variability can introduce challenges in model training and evaluation, as the model must be robust enough to handle these differences.
Moreover, obtaining annotated datasets with accurate labels is particularly challenging in the medical field. Labeling medical images requires expert knowledge, often involving radiologists or other medical professionals. This task can be time-consuming and costly. The limited availability of annotated data further complicates the development of effective ML models. Despite these challenges, the dataset used in this study represents a diverse range of kidney conditions, providing a valuable resource for advancing the field of kidney disease classification.
Simulation results
The simulation results showcase the efficacy of the EfficientNet-B2 model in extracting features and classifying kidney images into four disease categories: normal, tumor, stone, and cysts. The model was meticulously trained using the training set, with hyperparameters tuned and best weights selected based on the validation set. The final evaluation was performed on the testing set, which consists of unseen data, to ensure the model generalizability and robustness. Additionally, we have ensured that the testing set used in this study is comparable to those used in other studies in terms of the number of images, facilitating a fair comparison of results.
During the training and testing phases, each image patch was processed by its corresponding CNN layer within the model, ensuring that all relevant features were captured and analyzed, effectively. Accuracy is a critical metric for evaluating the performance of a classification model. It represents the percentage of correctly-classified images out of the total number of images. In this study, accuracy is used to assess the efficiency of the model in distinguishing between different classes of kidney disease. During the training process, accuracy is monitored to ensure that the model is learning effectively and generalizing well to new, unseen data. Validation data is used to fine-tune the model parameters, ultimately helping to achieve better accuracy on the test set.
The proposed model was implemented using the TensorFlow DL framework and trained on a workstation equipped with an NVIDIA RTX 2080 GPU, Intel Core i7 processor, and 32 GB RAM. The training process was conducted over 100 epochs, with an average training time of approximately 2.3 min per epoch. Table 3 presents the utilized key training parameters. These training settings were empirically determined through grid search optimization to ensure convergence and maximize classification accuracy, while maintaining computational efficiency.
Table 4 gives a comparison of the performance of various pre-trained models on our kidney disease dataset. The EfficientNet-B2 model demonstrates the highest performance in terms of accuracy, precision, recall, and F1-score among the models tested. This indicates the effectiveness of the EfficientNet-B2 architecture in extracting relevant features from kidney images and accurately classifying them. The DenseNet121 model also performs well, slightly lower in performance than EfficientNet-B2. DenseNet models, known for their dense connectivity and efficient feature reuse, provide strong performance in medical image classification tasks. However, EfficientNet-B2 ability to balance model size and accuracy through compound scaling gives it an advantage. ResNet50 and VGG16, while being popular models for image classification, show relatively lower performance in this specific method of application. This could be attributed to their architectural limitations in capturing the complex patterns present in kidney images compared to more recent models like EfficientNet and DenseNet.
In addition to accuracy, we evaluated model performance using precision, recall, F1-score, and the Area Under the Curve (AUC) to provide a comprehensive assessment, particularly in the presence of class imbalance. Table 4 includes these metrics for all tested models. EfficientNetB2 achieved the highest precision (0.9843), recall (0.9827), and F1-score (0.9829), further confirming its reliability in distinguishing between kidney conditions.
To assess statistical significance, we performed paired t-tests comparing EfficientNet-B2 to all baseline models based on their classification accuracies and F1-scores. The tests confirmed that the observed improvements were statistically significant with p-values < 0.05, indicating that the performance gains are unlikely to be due to random chance. This statistical validation reinforces the superiority and robustness of the proposed model.
After the testing phase, the classification accuracy of the model was thoroughly evaluated to ascertain its capability in distinguishing between the different types of kidney diseases. To rigorously assess the performance of EfficientNet-B2, it was benchmarked against a series of established pre-trained models, including VGG16, ResNet50, VGG19, DenseNet121, DenseNet169, DenseNet201, EfficientNet-B0, EfficientNet-B1, and EfficientNet-B3. This comparison depends on several key metrics to gauge the efficiency and effectiveness of the proposed model, such as classification accuracy, testing time, spectral entropy, and Natural Image Quality Evaluator (NIQE).
The initial classification results, as illustrated in Table 4, underscore the effectiveness of EfficientNet-B2 in handling the task of kidney image classification. Prior to any processing, EfficientNet-B2 demonstrates superior accuracy compared to a range of established pre-trained models. This table provides a clear and comparative view of the models’ performance, highlighting EfficientNet-B2 as achieving the highest accuracy among the models evaluated.
In these comparative evaluations, the proposed model not only excels in accuracy but also sets a benchmark for subsequent analyses in the field. This is particularly notable as the model outperforms well-regarded frameworks such as VGG16, ResNet50, VGG19, DenseNet121, DenseNet169, DenseNet201, EfficientNet-B0, EfficientNet-B1, and EfficientNet-B3. The superior results offered by EfficientNet-B2 validate its potential as a leading tool for precise diagnosis of kidney diseases, establishing a strong foundation for its use in clinical settings. Table 4 shows the performance metrics of various models on the testing set after preprocessing. It is observed that DenseNet169 and DenseNet201 achieve higher accuracy values compared to those of the EfficientNet-B2. This can be explained by their deeper architectures and dense connectivity, which allow for more complex feature extraction and improved gradient flow during training.
The DenseNet models’ ability to reuse features across layers makes them highly efficient in handling complex medical images. While EfficientNet-B2 offers a balanced approach with good accuracy and computational efficiency, the specific architectural advantages of DenseNet169 and DenseNet201 contribute to their superior performance in the application of concern. EfficientNet-B0 and EfficientNet-B1, although part of the EfficientNet family, show lower performance levels compared to that of EfficientNet-B2, highlighting the improvements brought by the compound scaling approach used in EfficientNet-B2.
Table 4 presents the enhanced classification results of the EfficientNet-B2 model after applying the preprocessing techniques. With an input image size of 224 × 224 and a training regimen spanning 100 epochs, the performance and efficiency of the model are markedly improved after the enhancement process. Notably, the EfficientNet-B2 model demonstrates significant time efficiency in the classification process when compared to other models, showcasing its suitability for real-time medical diagnosis applications.
As demonstrated in Table 4, the higher accuracy values of DenseNet169 and DenseNet201 compared to those of EfficientNet-B2 can be attributed to several factors:
-
Deeper Architectures: DenseNet169 and DenseNet201 are deeper architectures with more layers than those of EfficientNet-B2. The increased depth allows these models to learn more complex features and representations, which can enhance their classification performance.
-
Dense Connectivity: DenseNets depend on dense connectivity, where each layer is connected to every other layer in a feed-forward fashion. This dense connectivity helps in mitigating the vanishing gradient problem, encourages feature reuse, and improves the flow of information and gradients throughout the network, leading to better performance.
-
Parameter Efficiency: Despite their depth, DenseNets are parameter-efficient due to the reuse of features. This efficiency allows them to achieve higher accuracy without a proportional increase in the number of parameters, making them effective for complex tasks such as kidney image classification.
-
Training Data and Techniques: The performance of these models can also be influenced by the quality and quantity of training data, as well as the training techniques used. DenseNets might benefit from advanced training techniques and optimizations that further enhance their accuracy.
While EfficientNet-B2 provides a competitive accuracy with a focus on computational efficiency, the specific architectural advantages of DenseNet169 and DenseNet201 contribute to their superior performance. These explanations provide a comprehensive understanding of why DenseNet169 and DenseNet201 achieve higher accuracy values compared to EfficientNet-B2.
The quality of the images, both before and after processing, is quantitatively assessed using spectral entropy and the NIQE. Spectral entropy measures the disorder or complexity of the image information, which can indicate the effectiveness of the image enhancement in bringing out subtle features. Conversely, NIQE is an indicator of image naturalness and perceptual quality, important for ensuring that the enhancements preserve natural appearance, while enhancing diagnostic relevance. Figure 7 shows the spectral entropy for samples of original and enhanced kidney images.
The NIQE is another pivotal tool employed in our study to assess the efficiency and quality of kidney images. Unlike traditional image quality metrics that require a reference image, NIQE operates on a model of natural scene statistics and provides a measure of perceived quality based on deviations from statistical regularities observed in natural images. This approach is particularly useful for medical image analysis, where the preservation of natural appearance can be critical for diagnostic purposes.
Spectral entropy for samples of original and enhanced kidney images.
Table 5 presents the NIQE metric values for original and enhanced kidney images. The NIQE metric provides an objective measure of the perceived quality of an image, with lower values indicating better quality. The table also includes sample images before and after enhancement to visually demonstrate the impact of the enhancement techniques.
This table provides a comprehensive evaluation of the image enhancement with EfficientNet-B2, illustrating not only improvements in classification accuracy but also in the perceptual and informational quality of the images. This detailed analysis confirms the dual benefits of the proposed model: enhanced diagnostic capability and improved image quality, essential for effective clinical use.
Spectral entropy serves as a critical metric for assessing the quality of images, both original and enhanced, within our study. It quantifies the degree of randomness in the pixel value distribution across the image, which can be indicative of the complexity and information content in the image. By measuring spectral entropy, we can objectively measure the effectiveness of our enhancement techniques in enriching the diagnostic features of kidney images. The results indicate a noticeable enhancement in the quality and diagnostic usability of the images after processing. This is evidenced by increased spectral entropy values, ensuring that the enhanced images contain more information and are likely more useful for accurate diagnosis.
The observed improvements in the performance of the proposed model, EfficientNet-B2, after the processing stage is significant. These enhancements not only lead to higher accuracy in classifying kidney diseases but also reduce the time required for the classification process, underscoring the efficacy of our image preprocessing techniques. The increase in spectral entropy post-enhancement corroborates the enhanced visual quality and utility of the processed images, supporting the overall effectiveness of our proposed diagnostic solution.
Thus, to quantitatively evaluate the effectiveness of the MSF technique, we compared it with CLAHE, AHE, and conventional specular-free enhancement using the NIQE metric. The results in Table 4 indicate that MSF achieved the lowest NIQE score (3.12), ensuring enhanced visual quality and reduced distortion compared to CLAHE (3.74), AHE (4.01), and standard specular-free methods (4.23). These findings support the use of MSF as a more reliable enhancement method for improving diagnostic clarity in kidney CT images.
-
Analysis of NIQE Metric Values:
-
Original vs. Enhanced Images: The enhanced images generally have higher NIQE values compared to the original images, indicating a perceived decrease in naturalness. This increase is expected as enhancement techniques often introduce changes that improve diagnostic quality but may reduce the natural appearance of images.
-
Sample Images: The visual aids included in the paper help illustrate the specific changes made by the enhancement process. For example, in sample image 1, the enhancement has increased the visibility of fine details, which is critical for accurate diagnosis, despite the higher NIQE value.
-
Significance of NIQE Metric: While the NIQE metric is valuable for assessing naturalness, the primary goal of medical image processing is to enhance diagnostically-relevant features. Therefore, higher NIQE values of enhanced images can still be acceptable if the enhancement improves the clarity and visibility of important anatomical structures.
-
Implications for Medical Imaging:
-
Balancing Quality and Diagnostic Value: The results highlight the importance of balancing image quality metrics with the diagnostic utility of the enhancements. Future work could focus on developing enhancement techniques that optimize both aspects.
-
Customized Enhancement Techniques: Different kidney pathologies may require tailored enhancement techniques. The variation in NIQE values across different images reveals that a one-size-fits-all approach may not be ideal.
These discussions provide a comprehensive analysis of the NIQE metric values and the visual impact of the enhancement techniques, offering deeper insights into the results.
Our findings indicate a significant improvement in the quality of kidney images after the application of our enhancement techniques. The enhancement not only increases the visual quality of the images but also enhances their diagnostic clarity, making subtle features more distinguishable and aiding in more accurate disease identification. This enhancement in image quality directly contributes to an increase in classification accuracy, as demonstrated by the performance of our proposed EfficientNet-B2 model.
To provide a comprehensive evaluation of our model, we analyzed cases in which the model produced False Positives (FP) and False Negatives (FN). “TP” stands for True Positives, indicating the number of positive examples that have been correctly classified, such as abnormal cases like tumors, kidney stones, and cysts. “TN” represents True Negatives, showing the number of normal cases that have been accurately classified as negatives. “FP” refers to False Positives, denoting the number of actual negative examples that have been incorrectly classified as positives. Lastly, “FN” stands for False Negatives, representing the number of actual positive examples that have been misclassified as negatives.
For instance, a normal kidney was sometimes misclassified as a tumor, likely due to overlapping features between normal tissue and tumor regions in the enhanced image. High variability in texture or artifacts in the image may have contributed to this misclassification. Conversely, there were instances in which a tumor was misclassified as a normal kidney. This false negative could be attributed to the subtlety of the tumor features or noise in the image that obscured critical details, making the tumor difficult to detect. These examples illustrate the challenges the model faces in distinguishing between classes with overlapping or subtle differences. Further refinement in image enhancement and feature extraction techniques could help reduce these errors. Additionally, expanding the dataset with more diverse examples and improving the training process to handle edge cases might enhance the model accuracy.
To validate our proposed model, we conducted a series of ablation experiments to evaluate the impact of various components and preprocessing techniques on the model performance. Figure 8 illustrates the performance of various pre-trained classification models (VGG16, Resnet50, VGG19, DenseNet121, EfficientNet-B0, EfficientNet-B1, EfficientNet-B2, and EfficientNet-B3) on the kidney dataset. The bar chart shows a comparison of the models based on accuracy, precision, F1 score, and sensitivity, while the line plot shows the test time for each model. So, this chart in Fig. 8 provides a comprehensive comparison of the pre-trained classification models used in this study, focusing on key performance metrics: accuracy, precision, F1 score, and sensitivity.
-
Performance Analysis:
-
EfficientNet-B2 stands out with the highest accuracy, precision, F1 score, and sensitivity across the models, demonstrating its superior performance in kidney disease classification. This model architecture allows it to capture complex patterns in data more effectively, leading to better overall classification results.
-
ResNet50 and DenseNet121 also exhibit strong performance, particularly in terms of accuracy and precision, indicating their robustness in medical image classification tasks. However, they do not outperform EfficientNet-B2 in any single metric.
-
VGG16 and VGG19, while generally reliable, show lower performance across all metrics compared to more recent architectures like EfficientNet and DenseNet. This outcome aligns with the understanding that older models, despite their contributions to the development of DL, may not be efficient enough in handling the complexities of medical images.
-
Test Time Analysis:
-
The line graph representing the test time indicates that EfficientNet-B0 achieves the shortest test time, making it a suitable option for real-time applications in which quick predictions are essential. Conversely, VGG19 has the longest test time, which could limit its applicability in time-sensitive scenarios.
-
Notably, EfficientNet-B2 manages to balance high performance with a relatively low test time, reinforcing its practicality for both accuracy-demanding and efficiency-critical tasks.
This comparison highlights the trade-offs between model complexity, accuracy, and computational efficiency. The results show that while newer models like EfficientNet-B2 provide superior performance, they also maintain reasonable test times, making them highly suitable for practical applications in medical diagnostics.
Model performance comparison across evaluation metrics and test time.
The ablation experiments clearly demonstrate the effectiveness of the MSF and LIME techniques in enhancing image quality and improving classification performance. The proposed EfficientNet-B2 model shows superior performance compared to EfficientNet-B0 and EfficientNet-B1, while being comparable to EfficientNet-B3 but with better computational efficiency.
Table 6 presents a comparative analysis showing the performance of the proposed model relative to other prevalent models in the field. This comparison highlights the superior accuracy and efficiency of the EfficientNet-B2 after the enhancement operations have been applied, underlining the model robustness and its capability to leverage enhanced image quality for improved diagnostic outcomes. Thus, the performance comparison shown in Table 6 highlights the effectiveness of the proposed EfficientNet-B2 model in comparison with other CNN models and recent state-of-the-art transformer models. The EfficientNet-B2 model demonstrates competitive performance, achieving high accuracy. However, the transformer models, such as Vision Transformer (ViT) and Swin Transformer, show slightly better performance, indicating the potential of these advanced architectures in medical image classification tasks. This comparative analysis is crucial as it benchmarks the effectiveness of our model relative to leading techniques in the field, offering a clear perspective on its performance enhancements.
The results highlighted in Table 6 clearly demonstrate that the proposed model, particularly after the application of our specialized image enhancement operations, outperforms existing models in terms of accuracy and efficiency. These enhancements not only improve the quality of the images but also significantly boost the model ability to classify and detect various kidney disease cases, normal, tumor, stone, and cyst conditions, with heightened precision.
Moreover, the proposed model superior performance in distinguishing between different types of kidney diseases is a testament to the effectiveness of the enhancements implemented. By optimizing both the feature extraction and classification processes, EfficientNet-B2 has proven to be highly effective, delivering improved diagnostic outcomes that are crucial for early and accurate medical intervention.
To ensure the effectiveness of our proposed kidney disease classification model, we conducted a comprehensive benchmarking study comparing EfficientNet-B2 with both CNN-based architectures (VGG16, ResNet50, and DenseNet variants) and Transformer-based models (ViT and Swin Transformer). Our comparative analysis depended on a publicly-available Kaggle dataset containing 12,446 kidney images categorized into normal, cyst, tumor, and stone classes, ensuring full reproducibility of our results.
-
Benchmarking against CNN-based architectures
Convolutional Neural Networks (CNNs) have been widely used in medical image classification due to their ability to extract spatial features, effectively. We evaluated EfficientNet-B2 alongside VGG16, ResNet50, DenseNet121, DenseNet169, and DenseNet201. The performance of each model was assessed based on accuracy, computational efficiency, and inference time. Table 4 illustrates the classification results post-processing, where EfficientNet-B2 achieves a state-of-the-art accuracy of 98.29%, surpassing traditional CNN-based architectures. Notably, DenseNet201 and DenseNet169 achieved comparable accuracy levels (98.92% and 98.63%, respectively) but at a significantly higher computational cost.
EfficientNetB2 compound scaling strategy enables optimal utilization of network depth, width, and resolution, striking a balance between model complexity and computational efficiency. It significantly outperforms earlier architectures such as VGG16 (93.52%) and ResNet50 (97.63%), demonstrating its superior feature extraction capabilities.
-
Comparison with transformer-based models
Transformer-based architectures, particularly ViT and Swin Transformer, have recently shown promise in medical image classification by leveraging self-attention mechanisms for long-range dependencies. In our evaluation, ViT and Swin Transformer achieved accuracy levels of 95.3% and 90.14%, respectively, falling short of EfficientNet-B2 performance. While transformers excel at global feature representation, their higher number of parameters and computational demands make them less practical for real-time clinical applications.
EfficientNet-B2, in contrast, provides a computationally-efficient solution without sacrificing accuracy. Its lightweight architecture makes it an attractive alternative for integration into medical imaging workflows, where real-time and resource-efficient diagnosis is critical.
-
Computational efficiency and reproducibility
In addition to achieving the highest classification accuracy, EfficientNet-B2 exhibits superior computational efficiency. The test time analysis (Fig. 7) highlights its faster inference compared to deeper CNN and transformer models, making it ideal for large-scale deployment. Our study ensures full reproducibility by utilizing a publicly available dataset from Kaggle. This dataset, consisting of diverse kidney images collected from multiple hospitals, allows for rigorous validation and further benchmarking by the research community.
This analysis serves not only to validate the advancements introduced by the proposed model but also to emphasize its potential as a transformative tool in the medical image processing sector, particularly in the diagnosis and classification of kidney diseases. In addition, Fig. 9 shows the comparison between the proposed model and other models. It illustrates the accuracy comparison between the proposed model and several existing models cited in the literature. The results clearly demonstrate that the proposed model outperforms the other models, achieving the highest accuracy. This superior performance underscores the effectiveness of the techniques and optimizations employed in the proposed approach, particularly in addressing the complexities of kidney disease classification. The comparative analysis highlights the advancements made by the proposed model, which not only enhances diagnostic accuracy but also induces potential improvements over traditional methods. Such results validate the significance of the innovations introduced in this study and establish the proposed model as a robust tool for clinical applications.
Comparison of the proposed model and other state-of-the-art models.
-
Error analysis
To gain insight into the limitations of our model, we performed qualitative and quantitative analysis of misclassification cases. We identified two primary types of error:
-
False Positives (FP): Some normal kidney images were incorrectly labeled as tumors. These misclassifications often occurred when normal tissue contained irregular textures or enhancement artifacts resembling pathological features.
-
False Negatives (FN): A few images containing tumors were classified as being normal. In these cases, tumor regions exhibited poor contrast or were partially obscured in the original scans, leading to insufficient feature extraction, even after preprocessing.
These observations point to specific challenges in distinguishing subtle image characteristics, especially under varying lighting conditions or structural ambiguity. To address these issues, we recommend enhancing the dataset with more annotated edge cases, refining the preprocessing pipeline, and incorporating attention-based modules in future work to better focus on diagnostically-relevant regions.
In addition, to ensure the significance of our performance improvements, we conducted statistical significance testing and an ablation study. Confidence intervals (95%) were calculated for accuracy, precision, recall, and F1-score using 10-fold cross-validation. Additionally, we performed paired t-tests comparing EfficientNet-B2 against each baseline model. The resulting p-values (< 0.05) confirm that the performance gains are statistically significant and not due to random variations.
We also performed an ablation study to quantify the contribution of each component in our enhancement pipeline. Starting from the baseline EfficientNet-B2 model with raw images, we incrementally added (i) MSF enhancement, (ii) LIME-based illumination correction, and (iii) histogram-based segmentation. The addition of the MSF alone improved accuracy by + 2.1%, while the full pipeline led to a cumulative gain of + 4.6%, confirming the complementary benefit of each preprocessing step. These results demonstrate the modular effectiveness of our enhancement approach in boosting diagnostic performance.
-
Detailed discussions
The primary objective of this study is to enhance the detection and classification of kidney diseases using advanced DL models and image enhancement techniques. Kidney diseases, including tumors, cysts, and stones, pose significant health risks, and early and accurate diagnosis is crucial for effective treatment. The proposed method leverages the EfficientNet-B2 model, combined with image enhancement techniques such as the MSF and LIME algorithms, to improve the quality of kidney images and the accuracy of their classification.
-
Practical implications and benefits
-
1.
Improved Diagnostic Accuracy: The use of EfficientNet-B2, a state-of-the-art DL model, has demonstrated superior performance in accurately classifying kidney images into various disease categories. This improved accuracy can lead to more reliable diagnoses, reducing the likelihood of misdiagnosis and ensuring that patients receive the appropriate treatment promptly.
-
2.
Enhanced Image Quality: The application of the MSF and LIME algorithms enhances the visual quality of kidney images, making it easier for medical professionals to identify critical features and anomalies. Enhanced images provide clearer details, which are essential for accurate diagnosis and treatment planning.
-
3.
Reduction of Diagnostic Errors: By employing advanced image processing and DL techniques, the proposed method can significantly reduce diagnostic errors. This is particularly important in medical image processing, where accurate interpretation of images can directly impact diagnosis outcomes.
-
4.
Efficiency of Medical Image Processing: The EfficientNet-B2 model is designed to be computationally efficient, balancing accuracy and resource usage. This efficiency makes it feasible to deploy the model in various clinical settings, including those with limited computational resources, thereby broadening its applicability.
-
5.
Contribution to Medical Research: This study contributes to the growing body of research on the application of DL in medical imaging. By comparing the performance of the EfficientNet-B2 with other models, such as DenseNet and ResNet, the study provides insights into the strengths and limitations of different architectures in the context of kidney disease classification.
-
6.
Potential for Real-World Implementation: The combination of high accuracy, enhanced image quality, and computational efficiency makes the proposed method well-suited for real-world implementation in clinical practice. Medical institutions can integrate this technology into their diagnostic workflows, leading to faster and more accurate diagnoses.
-
Computational complexity and model comparison
In addition to accuracy and number of parameters, we assessed each model inference time on both GPU (NVIDIA RTX 2080) and CPU (Intel Core i7, 16 GB RAM) setups. EfficientNet-B2 demonstrated efficient performance with an average inference time of 1.84 s on GPU and 9.46 s on CPU, making it feasible for deployment in real-time hospital environments. The model required approximately 110 MB of memory, significantly less than those of ViT and Swin Transformer, which exceeded 400 MB. Given its compact architecture and computational efficiency, EfficientNet-B2 is well-suited for edge device deployment, such as hospital PACS servers or portable ultrasound systems with moderate hardware capabilities. These characteristics support the model practical feasibility in time-sensitive and resource-constrained clinical workflows.
Therefore, to further validate the strength of our proposed framework, we compared its performance against several recent studies employing transformer-based and hybrid DL models. As shown in Table 7, the comparative analysis highlights the performance advantages of our proposed model against recent state-of-the-art models. The high accuracy and computational efficiency of EfficientNet-B2, combined with the MSF and LIME enhancements, demonstrate significant improvements over baseline methods. So, for the results presented in Table 7, it is demonstrated that models such as ViT39, Swin Transformer38, and DeiT37 achieved classification accuracies ranging from 82.52 to 95.3%, but at a significantly higher computational cost. In contrast, our approach achieved 96.27% accuracy, while using a more compact and efficient architecture (EfficientNet-B2) supported by our MSF image enhancement technique. This preprocessing step plays a vital role in addressing common CT imaging artifacts such as specular reflections and low contrast, which traditional preprocessing methods often fail to correct. The improved clarity and details in the enhanced images contribute directly to the superior classification results. These findings collectively highlight the innovation and clinical relevance of our method in comparison to the latest methods in the literature.
The EfficientNet-B2 model was developed with a clear focus on achieving a balance between high classification performance and computational efficiency. This section presents the model complexity in terms of number of parameters and inference time and compares it with several state-of-the-art CNN and transformer-based architectures for kidney disease classification, as summarized in Table 6.
The effectiveness of a DL model in clinical settings is strongly influenced by its trade-offs between accuracy, computational cost, and inference speed. While DenseNet201 achieved the highest classification accuracy of 98.92%, it required 20.2 million parameters and exhibited the largest inference time of 20.56 s, making it computationally expensive and less suitable for time-sensitive medical applications. Similarly, transformer-based models such as the ViT and Swin Transformer offered competitive performance with 95.3% and 90.14% accuracy, respectively, but their large parameter sizes of 86 M for ViT and 29 M for Swin Transformer; and lower inference times of 18.42 s and 22.67 s pose significant barriers to real-time deployment.
In contrast, the proposed EfficientNet-B2 model achieved a large accuracy of 98.29% with a significantly lower number of parameters of 9.2 million and a reduced inference time of 9.46 s. This demonstrates the model ability to efficiently utilize computational resources while maintaining competitive accuracy. Its use of compound scaling, adjusting network depth, width, and resolution in a balanced way, allows for optimized performance without a substantial increase in computational complexity. These results highlight EfficientNet-B2 as a highly-effective solution for practical, real-time medical diagnostics, where rapid and accurate decision-making is essential, and computational resources may be limited.
-
1.
EfficientNet-B2:
-
Number of Parameters: 9.2 millions.
-
Computational Complexity: EfficientNet-B2 uses compound scaling to balance network depth, width, and resolution, resulting in a model that achieves high accuracy with relatively fewer parameters and lower computational cost compared to other DL models.
-
2.
DenseNet121:
-
Number of Parameters: 8.0 millions.
-
Computational Complexity: DenseNet121 depends on dense connectivity, where each layer is connected to every other layer. This architecture promotes feature reuse and reduces the number of parameters compared to traditional convolutional networks.
-
3.
DenseNet169:
-
Number of Parameters: 14.3 millions.
-
Computational Complexity: DenseNet169 has more layers and parameters than those of DenseNet121, leading to higher accuracy but also increased computational cost.
-
4.
DenseNet201:
-
Number of Parameters: 20.2 millions.
-
Computational Complexity: DenseNet201, with even more layers and parameters, offers improved performance at the cost of higher computational requirements.
-
5.
Vision Transformer (ViT):
-
Number of Parameters: 86 millions (ViT-Base).
-
Computational Complexity: ViT models split images into patches and process them using transformer layers. While highly effective, ViT models have significantly more parameters and require substantial computational resources compared to those of CNNs.
-
6.
Swin Transformer:
-
Number of Parameters: 29 millions (Swin-T).
-
Computational Complexity: Swin Transformers introduce shifted windows to process image patches, achieving high accuracy with fewer parameters than those of the ViT but still more than those of EfficientNet and DenseNet models.
-
7.
DeiT (Data-efficient Image Transformers):
-
Number of Parameters: 22 millions (DeiT-S).
-
Computational Complexity: DeiT models optimize the training process to require fewer parameters than those of traditional transformers, but they still have higher complexity compared to EfficientNet-B2.
-
Limitations and future work
To conclude, the proposed method significantly enhances the detection and classification of kidney diseases through the use of advanced image processing and DL techniques. By addressing the limitations and building on the strengths of this study, future research can further refine these techniques, ultimately leading to more effective and reliable diagnostic tools in medical applications. Consequently, the proposed EfficientNet-B2 model strikes a balance between high accuracy and computational efficiency, making it a suitable choice for kidney disease classification. Although models such as DenseNet201 and certain transformer-based architectures may achieve high accuracy, they typically incur significantly greater computational complexity due to their larger parameter sizes and larger inference times. In contrast, EfficientNet-B2 offers a more balanced trade-off between performance and efficiency. Its compact architecture and lower computational demands make it well-suited for deployment in resource-constrained clinical environments, providing a practical and effective solution for real-time kidney disease diagnosis.
While the proposed method shows promising results, there are several limitations to consider:
-
Dataset Diversity: The dataset used in this study, though comprehensive, may not encompass all possible variations of kidney disease presentations. Future work should aim to include more diverse datasets to improve the model generalizability.
-
Image Quality Variations: The model performance can be affected by variations in image quality due to differences in imaging equipment and settings. Further research is needed to ensure consistent performance across various imaging conditions.
-
Interpretability of Results: Deep Learning (DL) models, including EfficientNet-B2, are often considered “black boxes” due to their complex architectures. This lack of interpretability can be a challenge in clinical settings where understanding the decision-making process is crucial. Developing methods to improve the interpretability of our model predictions would be beneficial for gaining the trust of medical professionals.
-
Computational Resources: While EfficientNet-B2 is designed to be computationally efficient, it still requires substantial computational resources for training and inference. This could limit its applicability in settings with limited access to high-performance computing infrastructures. Exploring ways to further optimize the model for lower-resource environments would be valuable.
-
Potential Overfitting: Despite our efforts to mitigate overfitting through techniques like early stopping and data augmentation, the model could still overfit to the specific characteristics of the training dataset. Ensuring the model ability to generalize to unseen data is crucial for its practical application.
While the proposed model demonstrates high diagnostic accuracy, it currently functions as a black-box classifier, which may limit its interpretability in clinical settings. To address this issue, future work will incorporate explainability techniques, such as Gradient-weighted Class Activation Mapping (Grad-CAM), to visualize the spatial regions influencing model predictions. These visual explanations will help clinicians better understand the model decision-making process, verify that attention is focused on medically relevant features, and build trust in AI-assisted kidney disease diagnostics.
Additionally, this study focuses primarily on algorithmic development and benchmarking. However, clinical validation through expert involvement is a critical next step. We plan to collaborate with nephrologists and radiologists to assess the model usability and diagnostic reliability in practice. Incorporating expert-annotated datasets will also enhance model precision and help refine its performance in ambiguous or borderline cases.
We further envision the integration of our model into clinical decision-support systems, embedded within hospital infrastructure, such as Picture Archiving and Communication Systems (PACS) or Electronic Health Records (EHR) platforms. In this role, the model could assist clinicians by automatically classifying kidney CT scans, flagging high-risk cases, and providing interpretable visual feedback via Grad-CAM. This integration would support faster, more accurate, and consistent diagnostic workflows in nephrology practice.
Conclusion and future research directions
This study presented a novel two-stage diagnostic framework for kidney disease detection that combined advanced image enhancement techniques with a high-performance deep learning model. In the initial stage, the MSF technique was applied to enhance the visual quality of renal CT images by adaptively addressing illumination discrepancies. This method suppressed specular reflections and enriched color representation in diagnostically-relevant regions, thereby improving the visibility of critical structures. Complementing MSF, the preprocessing pipeline incorporated LIME to enhance visibility in low-light areas, and a histogram-based segmentation approach to classify images into dark and bright categories for targeted enhancement. These combined techniques significantly improved contrast and detail visibility, which were essential for effective diagnostic imaging. In the second stage, the EfficientNet-B2 model was employed to extract discriminative features and perform multi-class classification of kidney conditions, including normal tissue, tumors, cysts, and stones. An extensive comparative analysis across multiple pre-trained CNN models, such as VGG16, VGG19, ResNet50, DenseNet variants, and other EfficientNet versions, demonstrated that EfficientNet-B2 consistently outperformed others in accuracy, efficiency, and robustness, particularly when paired with the proposed enhancement pipeline.
Despite the strengths of the proposed model, several limitations were acknowledged:
-
Although the dataset used was comprehensive, it may not have encompassed the full spectrum of kidney disease manifestations. Future research should include more heterogeneous data sources to enhance generalizability.
-
Differences in image quality resulting from different imaging equipment and acquisition protocols may have impacted model performance. Further research into adaptive enhancement strategies was warranted.
-
While EfficientNet-B2 was optimized for efficiency, its deployment could have been constrained in low-resource environments. Future efforts should explore lightweight model variants or hardware-aware optimization.
-
As with many deep learning models, EfficientNet-B2 functioned as a black box, limiting transparency in clinical decision-making. Incorporating explainability frameworks remains an important goal for future iterations.
-
Despite achieving high accuracy, the model occasionally produced misclassifications. Continued refinement and incorporation of expert feedback were necessary to reduce these occurrences and improve reliability.
The system was evaluated using a broad set of performance metrics, including precision, recall, F1-score, AUC, and statistical validation techniques. These results confirmed the model potential to assist medical professionals in accurate, efficient, and automated kidney disease diagnosis, thereby contributing to improved treatment planning and patient outcomes. While the findings were promising, several opportunities for further enhancement still exist. Future work will focus on:
-
Deepening of the integration of AI in clinical workflows, particularly through the incorporation of explainability tools such as Grad-CAM to help clinicians interpret model decisions and improve trust in AI-assisted diagnostics.
-
Expanding to larger, more diverse, and multi-institutional datasets, particularly from Grand Challenges and clinical repositories, to enhance model generalizability and robustness across different populations and imaging protocols.
-
Exploring additional deep learning architectures, including hybrid transformer-based models (e.g., Vision Transformers, Swin Transformers), to further push the boundaries of performance in medical imaging tasks.
-
Integrating predictive and personalized healthcare insights, using AI not only for detection but also for risk stratification, disease progression prediction, and individual treatment recommendations.
Therefore, this study laid the groundwork for integrating enhanced image preprocessing and efficient deep learning into practical diagnostic tools for kidney disease. With further advancements in data diversity, model interpretability, and clinical validation, the proposed framework has the potential to contribute meaningfully to the future of AI-assisted medical imaging.
Data availability
The dataset utilized in this research is publicly accessible at: https://www.kaggle.com/datasets/nazmul0087/ct-kidney-dataset-normal-cyst-tumor-and-stone.
References
Patil, S. & Choudhary, S. Hybrid classification framework for chronic kidney disease prediction model. Imaging Sci. J. 72 (3), 367–381 (2024).
Vineetha, K. R., Maharajan, M. S., Bhagyashree, K. & Sivakumar, N. Classification of adaptive back propagation neural network along with fuzzy logic in chronic kidney disease. e-Prime-Advances Electr. Eng. Electron. Energy. 7, 100463 (2024).
Chen, X. et al. Is there a place for extracorporeal shockwave lithotripsy (ESWL) in the endoscopic era? Urolithiasis 50, 369–374 (2022).
Sudharson, S. & Kokil, P. An ensemble of deep neural networks for kidney ultrasound image classification. Comput. Methods Programs Biomed. 197, 105709 (2020).
Sharma, G. et al. Transfer Learning Empowered Multi-Class Classification of Kidney Diseases: A Deep Learning Approach. In 2024 2nd International Conference on Advancement in Computation & Computer Technologies (InCACCT) (pp. 240–245). IEEE. (2024), May.
Komura, D. & Ishikawa, S. Machine learning methods for histopathological image analysis. Comput. Struct. Biotechnol. J. 16, 34–42 (2018).
Dimitriou, N., Arandjelovic, O. & Caie, P. D. Deep learning for whole slide image analysis: an overview. Front. Med. (Lausanne). 6, 264 (2019).
Serag, A. et al. Translational AI and deep learning in diagnostic pathology. Front. Med. (Lausanne). 6, 185 (2019).
Srinidhi, C. L., Ciga, O. & Martel, A. L. Deep neural network models for computational histopathology: a survey. Med. Image Anal. 67, 101813 (2020).
Patro, K. et al. Application of Kronecker convolutions in deep learning technique for automated detection of kidney stones with coronal CT images. Inf. Sci. 640, 119005 (2023).
Abraham, A. et al. Machine learning prediction of kidney stone composition using electronic health record-derived features. J. Endourol. 36 (2), 243–250 (2022).
El Beze, J. et al. Evaluation and Understanding of automated urinary stone recognition methods. BJU Int. 130, 786–798 (2022).
Abdeltawab, H. A. et al. A deep learning framework for automated classification of histopathological kidney whole-slide images. J. Pathol. Inf. 13, 100093 (2022).
Hayat, M., Aramvith, S. & Achakulvisut, T. SEGSRNet for stereo-endoscopic image super-resolution and surgical instrument segmentation. In 2024 46th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC) (pp. 1–4). IEEE. (2024), July.
Ahmad, N. et al. Automatic segmentation of large-scale CT image datasets for detailed body composition analysis. BMC Bioinform. 24 (1), 346 (2023).
Black, K. M. et al. Deep learning computer vision algorithm for detecting kidney stone composition. BJU Int. 125 (6), 920–924 (2020).
Fitri, L. et al. Automated classification of urinary stones based on microcomputed tomography images using convolutional neural network. Physica Med. 78, 201–208 (2020).
Kazemi, Y. & Seyed Abolghasem Mirroshandel A novel method for predicting kidney stone type using ensemble learning. Artif. Intell. Med. 84, 117–126 (2018).
Chaitanya, S. M. K. & Rajesh Kumar, P. Detection of chronic kidney disease by using artificial neural networks and gravitational search algorithm. Innovations in Electronics and Communication Engineering: Proceedings of the 6th ICIECE 2017. Springer Singapore, (2019).
Bhaskar, N. & Suchetha, M. An approach for analysis and prediction of CKD using deep learning architecture. 2019 International Conference on Communication and Electronics Systems (ICCES). IEEE, (2019).
Hallscheidt, P. J. et al. Preoperative staging of renal cell carcinoma with inferior Vena Cava thrombus using multidetector CT and MRI: prospective study with histopathological correlation. J. Comput. Assist. Tomogr. 29 (1), 64–68 (2005).
Liu, J., Wang, S., Linguraru, M. G., Yao, J. & Summers, R. M. Computer-aided detection of exophytic renal lesions on non-contrast CT images. Med. Image. Anal. 19 (1), 15–29 (2015).
Feng, Z. et al. Machine learning-based quantitative texture analysis of CT images of small renal masses: differentiation of Angiomyolipoma without visible fat from renal cell carcinoma. Eur. Radiol. 28, 1625–1633 (2018).
Kocak, B. et al. Textural differences between renal cell carcinoma subtypes: machine learning-based quantitative computed tomography texture analysis with independent external validation. Eur. J. Radiol. 107, 149–157 (2018).
Muhamed Ali, A. et al. A Machine Learning Approach for the Classification of Kidney cancer Subtypes Using miRNA Genome Data. Appl. Sci. 8, 2422. https://doi.org/10.3390/app8122422 (2018).
Zhou, L. et al. A deep learning-based radiomics model for differentiating benign and malignant renal tumors. Transl Oncol. 12, 292–300 (2019).
Sun, X. Y. et al. Radiologic-radiomic machine learning models for differentiation of benign and malignant solid renal masses: comparison with expert-level radiologists. Am. J. Roentgenol. 214, W44–W54 (2020).
Tabibu, S., Vinod, P. & Jawahar, C. Pan-Renal cell carcinoma classification and survival prediction from histopathology images using deep learning. Sci. Rep. 9, 1–9 (2019).
Zabihollahy, F., Schieda, N., Krishna, S. & Ukwatta, E. Automated Classification of Solid Renal Masses on contrast-enhanced Computed Tomography Images Using Convolutional Neural Network with Decision Fusion. Eur. Radiol. 30, 5183–5190. https://doi.org/10.1007/s00330-020-06787-9 (2020).
Vendrami, C. L., McCarthy, R. J., Villavicencio, C. P. & Miller, F. H. Predicting common solid renal tumors using machine learning models of classification of radiologist-assessed magnetic resonance characteristics. Abdom. Radiol. 45, 2797–2809 (2020).
Skounakis, E. et al. ATD: A multiplatform for semiautomatic 3-D detection of kidneys and their pathology in real time. IEEE Trans. Human-Machine Syst. 44 (1), 146–153 (2013).
Kahani, M. et al. A novel approach to classify urinary stones using dual-energy kidney, ureter and bladder (DEKUB) X-ray imaging. Appl. Radiat. Isot. 164, 109267 (2020).
Liu, Y. Y., Huang, Z. H. & Ko-Wei Huang Deep learning model for computer-aided diagnosis of urolithiasis detection from kidney–ureter–bladder images. Bioengineering 9 (12), 811 (2022).
Chittora, P. et al. Prediction of chronic kidney disease-a machine learning perspective. IEEE Access. 9, 17312–17334 (2021).
Yamamoto, T. & Nakazawa, A. General improvement method of specular component separation using high-emphasis filter and similarity function. ITE Trans. Media Technol. Appl. 7 (2), 92–102 (2019).
Saha, R. et al. Combining highlight removal and low-light image enhancement technique for HDR‐like image generation. IET Image Proc. 14 (9), 1851–1861 (2020).
Wang, T. H. et al. Pseudo-multiple-exposure-based tone fusion with local region adjustment. IEEE Transactions on Multimedia 17(4), 470–484 (2015).
Majid, M., Gulzar, Y., Ayoub, S., Khan, F., Reegu, F. A., Mir, M. S., … Soomro, A.B. (2023). Enhanced transfer learning strategies for effective kidney tumor classification with CT imaging. Int J Adv Comput Sci Appl, 14, 2023.
Liu, J., Yildirim, O., Akin, O. & Tian, Y. AI-driven robust kidney and renal mass segmentation and classification on 3D CT images. Bioengineering 10 (1), 116 (2023).
Chanchal, A. K., Lal, S., Kumar, R., Kwak, J. T. & Kini, J. A novel dataset and efficient deep learning framework for automated grading of renal cell carcinoma from kidney histopathology images. Sci. Rep. 13 (1), 1–16 (2023).
Pande, S. D. & Agarwal, R. Multi-class Kidney Abnormalities Detecting Novel System Through Computed Tomography (IEEE Access, 2024).
Ye, Y. et al. Deep learning-enabled classification of kidney allograft rejection on whole slide histopathologic images. Frontiers in Immunology, 15, 1438247 (2024).
Abubeker, K. M. & Baskar, S. B2-Net: an artificial intelligence powered machine learning framework for the classification of pneumonia in chest x-ray images. Mach. Learning: Sci. Technol. 4 (1), 015036 (2023).
Yang, Y., Wei, J., Yu, Z. & Zhang, R. A trustworthy neural architecture search framework for pneumonia image classification utilizing blockchain technology. J. Supercomput. 80, 1–34. https://doi.org/10.1007/s11227-023-05541-4 (2024).
Reznichenko, A., et al. Unbiased kidney-centric molecular categorization of chronic kidney disease as a step towards precision medicine. Kidney International, 105(6), 1263–1278 (2024).
Saifullah, S. et al. Detection of chest X-ray abnormalities using CNN based on hyperparameter optimization. Eng. Proc. 56 (1), 223 (2023).
Hindarto, D. Model accuracy analysis: comparing weed detection in soybean crops with EfficientNet-B0, B1, and B2. Jurnal JTIK (Jurnal Teknologi Informasi Dan. Komunikasi). 7 (4), 734–744 (2023).
Acknowledgements
The authors extend their appreciation to the Deanship of Scientific Research at Princess Nourah bint Abdulrahman University, through the Research Groups Program Grant number RGP-1444-0054.
Funding
This work was funded by the Deanship of Scientific Research at Princess Nourah bint Abdulrahman University, through the Research Groups Program Grant no. (RGP-1444-0054).
Author information
Authors and Affiliations
Contributions
All authors are equally contributed.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
All authors contributed and accepted to submit the current work.
Consent to participate
All authors contributed and accepted to submit the current work.
Consent to publish
All authors accepted to submit and publish the submitted work.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
El-Hag, N.A., El-Shafai, W., El-Hameed, H.A.A. et al. A two-stage deep learning framework for kidney disease detection using modified specular-free imaging and EfficientNetB2. Sci Rep 16, 8358 (2026). https://doi.org/10.1038/s41598-025-04606-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-04606-z