Abstract
The Scorpaeniformes order encompasses a diverse array of teleost fish, including commercially important and venomous species. Fish venoms offer significant pharmacological potential, but incomplete phylogenetic understanding has hindered research. Resolving relationships among venomous fish families is crucial for studying venom evolution and discovering novel bioactive compounds. To address these phylogenetic uncertainties, we generated and assembled the complete mitochondrial genomes of Paracentropogon rubripinnis (Tetrarogidae) and Inimicus japonicus (Synanceiidae), two representative venomous species. The circular mitogenomes, 16,465 bp and 16,676 bp in length, respectively, contain the typical vertebrate mitochondrial gene complement. Comparative analyses revealed a highly conserved gene order and orientation across Scorpaeniformes, with slight variations in the Notothenioidei outgroups. We identified three novel conserved sequence blocks in the control regions and characterized structural features of protein-coding genes, tRNAs, and non-coding elements. Phylogenetic analyses using 13 mitochondrial protein-coding genes from 71 Scorpaeniformes and three outgroup species provided a higher-resolution phylogeny of the order, including 12 families and 31 genera. Our results support the monophyly of Tetrarogidae and Synanceiidae, placing them in an early-diverging position within the Scorpaeniformes phylogeny. This study provides insights into the phylogenetic positions of venomous fish families and lays a foundation for future research on fish venom evolution and applications.
Similar content being viewed by others
Introduction
The Scorpaeniformes order is one of the largest and most morphologically diverse groups of ray-finned fish, comprising more than 1400 species classified into 38 families, including commercially important and venomous fish1,2. Understanding the polyphyletic relationships within this order is crucial for effective conservation efforts, sustainable fisheries management, and potential pharmacological applications of venomous species. However, the complexity and diversity of Scorpaeniformes have presented significant challenges in resolving their evolutionary history, leading to unclear taxonomic boundaries and debates about the monophyly of the order3,4,5. These uncertainties persist despite numerous studies employing diverse approaches such as morphological analyses, partial mitochondrial gene sequencing, and nuclear gene examinations6,7. A prime example of this ongoing controversy is the unresolved phylogenetic relationships among families such as Synanceiidae, Tetrarogidae, Scorpaenidae, and Sebastidae. Previous studies using molecular sequence data or morphological features4,6,8,9,10,11,12,13 have yielded two competing hypotheses: ((Synanceiidae + Tetrarogidae) + (Scorpaenidae + Sebastidae)) (Fig. 1A) and ((Synanceiidae + Tetrarogidae) + Scorpaenidae) + Sebastidae) (Fig. 1B).
Competing phylogenetic hypotheses for venomous fish families within Scorpaeniformes. (A) The hypothesis shows a close relationship between Synanceiidae and Tetrarogidae, separate from Scorpaenidae and Sebastidae. (B) The alternative hypothesis suggests a closer relationship for Synanceiidae, Tetrarogidae, and Scorpaenidae, with Sebastidae as a more distant relative. Silhouettes represent characteristic body shapes for each family.
Phylogenetic analyses face several challenges that can lead to discrepancies between gene and species trees14. These discrepancies may arise from various factors, including incomplete lineage sorting, horizontal gene transfer, gene duplication and loss, and hybridization15,16,17,18,19. Furthermore, sampling errors and biases in outgroup selections can significantly influence phylogenetic reconstruction20. Meanwhile, increasing the number of genes and sequence length can help mitigate some of these issues by providing more phylogenetic signals; however, it also introduces new challenges. Obtaining and analyzing large-scale genomic data, especially for large eukaryotic genomes, is often costly, time-consuming, and computationally intensive21,22. Mitochondrial genomes represent essential evolutionary repositories for phylogenetic studies, including Scorpaeniformes phylogenetics9,23,24,25. Mitochondrial genomes, which are relatively easier to obtain and analyze compared to nuclear genomes, typically range from 15 to 20 kb in fish and contain a compact informative set of genes: 13 protein-coding genes (PCGs), two ribosomal RNAs (12s rRNA and 16s rRNA), 22 transfer RNAs (tRNAs), and two non-coding regions (control region and origin of the light strand (OL))—these are essential for transcription and replication26. Moreover, mitogenomes possess unique characteristics that make them suitable for evolutionary and phylogenetic studies, including conserved gene content across species, relatively high mutation rate, maternal inheritance, and low levels of recombination27,28,29. These features collectively contribute to the effectiveness of the mitochondrial genome as molecular markers in evolutionary biology and phylogenetics.
Among the diverse families within the Scorpaeniformes order, Tetrarogidae (also known as waspfish) and Synanceiidae (stonefish) are particularly interesting. Tetrarogidae comprises approximately 17 genera and 41 species, while Synanceiidae includes about nine genera and 36 species, and both are primarily distributed in the Indo-West Pacific. These bottom-dwelling fish are notorious for their venomous dorsal spines, which can inflict painful injuries on humans30,31. The venom of these fish has garnered attention for its potential pharmacological and medical applications32. Consequently, numerous studies have investigated the ecological distribution, morphological features, venom proteins, and evolutionary history of Scorpaeniformes fish11,13,33,34,35. However, taxonomic studies based on molecular characteristics remain relatively scarce. To date, according to the MitoFish database36among more than 4200 species with complete mitochondrial genomes, approximately 100 species belong to Scorpaeniformes. Notably, the family Tetrarogidae was absent from these mitogenomic studies. Paracentropogon rubripinnis (Tetrarogidae) and Inimicus japonicus (Synanceiidae) are two representative species of these families, inhabiting the coastal waters of southern Japan and the southern Korean peninsula in the Northwest Pacific. While three mitogenomes from the Synanceiidae family have recently been sequenced10,37,38no mitogenomic data are currently available for any Tetrarogidae species.
This study aimed to elucidate the phylogenetic positions of Tetrarogidae and Synanceiidae and their evolutionary relationships within the Scorpaeniformes order. Thus, we sequenced and assembled the complete circular mitochondrial genomes of P. rubripinnis and I. japonicus to achieve this. We comprehensively analyzed their nucleotide composition, gene content, genomic organization, and codon usage profiles of protein-coding genes (PCGs). Additionally, we examined the structure of non-coding elements, including tRNAs, and the replication origin and identified novel conserved sequenced blocks in the control region. To provide a broader phylogenetic context, we performed phylogenetic analyses using 71 Scorpaeniformes and three outgroup species, focusing on the phylogenetic positions of Tetrarogidae and Synanceiidae within the order.
Results
Mitochondrial genome organization and structure
We sequenced and assembled the complete mitochondrial genomes of P. rubripinnis and I. japonicus using Illumina sequencing technology, yielding 59.8 and 33.2 Gb of raw data, respectively (Table S1). The resulting circular mitogenomes were 16,465 bp for P. rubripinnis and 16,676 bp for I. japonicus. Both mitogenomes exhibited the typical vertebrate mitochondrial gene composition, containing 13 protein-coding genes (COI–III, ND1–6, ND4L, Cytb, ATPase6, and ATPase8), two ribosomal RNA genes (12 S rRNA and 16 S rRNA), 22 transfer RNA genes (including two each for serine and leucine, and one for the other amino acids), and a control region (D-loop) (Fig. 2). Most genes (28 out of 37) were encoded on the heavy strand (H-strand). In comparison, the remaining nine genes (ND6 and eight tRNAs) were located on the light strand (L-strand) (Fig. 2; Table 1). To assess the conservation of genes in the order across Scorpaeniformes, we compared the mitogenome structures of P. rubripinnis and I. japonicus with those of 71 other Scorpaeniformes species and three Notothenioidei species (Table 2). Our analysis revealed that the order and orientation of all 37 genes and the control region in P. rubripinnis and I. japonicus are identical to those of other sequenced Scorpaeniformes species9,37,39,40. This high degree of conservation in gene arrangement appears to be a characteristic feature of Scorpaeniformes mitogenomes. Interestingly, we observed slight differences in the region between ND5 and the D-loop in the Notothenioidei species, highlighting a potential order-specific variation in mitogenome structures (Fig. S1).
Circular representation of the complete mitochondrial genomes of (A) P. rubripinnis and (B) I. japonicus. The outer circle shows the gene arrangement, with protein-coding genes, rRNAs, and tRNAs indicated. The inner circle represents the GC content, with the dark gray and light gray indicating GC and AT content, respectively. Genes transcribed on the heavy strand are shown outside the circle, while those on the light strand are shown inside. The total genome size is indicated for each species.
A comprehensive analysis of the structural and genomic features of P. rubripinnis and I. japonicus mitogenomes revealed several distinctive characteristics. Subsequently, we checked the intergenic regions, overlapping genes, nucleotide composition, and skewness of two mitogenomes. Intergenic spacer sequences were identified in nine areas, totaling 25 bp in P. rubripinnis and 30 bp in I. japonicus, with lengths ranging from 1 to 9 bp (Table 1). Concurrently, we observed gene overlaps in both species: six regions in P. rubripinnis (24 bp total) and seven in I. japonicus, involving various genes such as tRNAs, ATPase, ND, and Cytb. Nucleotide composition analysis showed a high A+T content in both mitogenomes (57.88% in P. rubripinnis and 58.66% in I. japonicus; Table 2), notably exceeding the average found in the Scorpaeniformes species (53.97 ± 1.7%). This elevated A+T content aligns with previous findings in S. verrucosa (59.35%), another member of the Synanceiidae family37.
Further examination of nucleotide bias through AT and GC skew analyses revealed interesting patterns. The AT skew was slightly negative for P. rubripinnis (− 0.0059) but positive for I. japonicus (0.0083), reflecting the broader trend in Scorpaeniformes, where 57 of 71 species exhibited positive AT skew values (average 0.0112 ± 0.017). Notably, Triglidae, Scorpaenidae, Sebastidae, and Synanceiidae fish demonstrated high AT skew (family average > 0.02), contrasting with the low AT skew (< − 0.01) observed in Stichaeidae and Anarhichadidae. The GC skew values were consistently negative across all Scorpaeniformes fishes, indicating a higher content of Cs than Gs, with P. rubripinnis and I. japonicus showing values of − 0.2242 and − 0.2285, respectively. These values are less negative than the Scorpaeniformes average (− 0.2487 ± 0.018), aligning with an increased GC skew observed in families such as Stichaeidae, Anarhichadidae, Gasterosteidae, Anoplopomatidae, Scorpaenidae and Tetrarogidae, in contrast to the lower GC skew seen in Cottidae, Hexagrammidae, and Triglidae. These findings collectively highlight the unique genomic features of P. rubripinnis and I. japonicus within the Scorpaeniformes mitochondrial genome evolution.
Protein-coding genes
The total length of 13 protein-coding genes (PCGs) was 11,428 bp, encoding 3800 codons in both P. rubripinnis and I. japonicus mitogenomes. Most PCGs utilized ATG as the start codon, except for COI, which possessed GTG, an accepted canonical mitochondrial start codon in vertebrates41,42,43 (Table 1). For termination, six PCGs (ND1, COI, ATPase8, ND4L, ND5, and ND6) contained the TAA stop codon, four (COII, ND3, ND4, and Cytb) used the incomplete T stop codon, and three (ND2, ATPase6, and COIII) used the incomplete TA stop codon. This pattern of stop codon usage was identical to that of S. verrucosa, another member of the Synanceiidae family37. The incomplete stop codons are likely completed to TAA by post-transcriptional polyadenylation44. The A+T content of the PCGs was 58.08% in P. rubripinnis and 58.43% in I. japonicus, while the A + T content at the third codon positions was 67.8% and 67.6%, respectively. Analysis of the relative synonymous codon usage (RSCU) revealed a preference for NNA and NNT codons over NNC and NNG (Fig. 3), consistent with the observed A and T bias at the third codon positions, which is typical in metazoan mitochondria25,45. Most PCGs showed negative AT skew values, indicating a higher A and T content, except for ATPase8 in P. rubripinnis and ND2 and ATPase8 in I. japonicus (Fig. 4). GC skew values were negative for most PCGs except for ND6, indicating a higher C content than G. ND6 exhibited the highest GC skew and lowest AT skew in both species. These skew patterns are consistent with those observed in other Scorpaeniformes mitogenomes9,24.
Relative synonymous codon usage (RSCU) in the mitochondrial genomes of (A) P. rubripinnis and (B) I. japonicus. The x-axis shows the 20 amino acids and their corresponding codons. The y-axis represents the RSCU values, with values greater than 1 indicating codons used more frequently than expected. The specific codons are listed below each amino acid in both (A) and (B). The color of each bar indicates particular codons.
AT and GC skew values for protein-coding genes in the mitochondrial genomes of (A) P. rubripiniis and (B) I. japonicus.
Transfer RNA genes and ribosomal RNA genes
The mitogenomes of P. rubripinnis and I. japonicus each contained 22 tRNA genes, including two for leucine and serine. Fourteen tRNA genes were located on the plus strand and eight on the minus strand (Table 1). In P. rubripinnis, tRNA gene lengths ranged from 66 bp (tRNA-Cys) to 74 bp (tRNA-Leu1 and tRNA-Lys), and in I. japonicus they ranged from 67 bp (tRNA-Cys) to 74 bp (tRNA-Leu). Secondary structure predictions revealed that 21 tRNA genes displayed canonical cloverleaf structures, while tRNA-Ser2 lacked a dihydrouridine (DHU) stem in both species (Fig. 5). This tRNA-Ser2 feature is consistent with observations in other vertebrate mitogenomes, including Scorpaenifromes9,24,46. All amino acid acceptor stems in the tRNA genes were conserved at 7 bp, including non-Watson–Crick base pairs. Unmatched base pairs, exclusively T–G base pairs, were present in stem regions, a common phenomenon that can be resolved by post-transcriptional editing47. The concatenated sequence of all tRNA genes showed positive AT skew (0.0289 in P. rubripinnis; 0.0166 in I. japonicus) and GC skew (0.0523 in P. rubripinnis; 0.0783 in I. japonicus), indicating a bias towards A and G nucleotides.
Predicted secondary structures of the 22 tRNA genes in the mitochondrial genomes of (A) P. rubripinnis and (B) I. japonicus. Each tRNA is shown in a cloverleaf secondary structure, with the amino acid it carries indicated above. Blue dots represent Watson–Crick base pairs, while yellow dots indicate non-canonical base pairings.
The 12 S and 16 S rRNA genes were located between tRNA-Phe and tRNA-Leu1, separated by tRNA-Val. In P. rubripinnis, the 12 S gene was 952 bp long with 55.15% A + T content, while the 16 S rRNA gene was 1682 bp with 58.32% A + T content. In I. japonicus, the 12 S rRNA gene was 954 bp (56.71% A + T), and the 16 S rRNA gene was 1688 bp (57.94% A + T).
Non-coding regions
The mitogenomes of P. rubripinnis and I. japonicus contained two major non-coding regions: the origin of light-strand replication (OL) and the control region (CR). These regions contain regulatory sequences essential for mitochondrial transcription and replication initiation48. The OL was located between tRNA-Asn and tRNA-Cys, with lengths of 32 bp in P. rubripinnis and 31 bp in I. japonicus. Both formed hairpin secondary structures (Fig. 6A), consistent with typical vertebrate OL characteristics. However, the I. japonicus OL exhibited an atypical structure with relatively lower predicted scores, warranting further investigation into its structural significance.
Analysis of non-coding regions in mitochondrial genomes. (A) Predicted secondary structures of light strand replication origin (OL) in P. rubripinnis and I. japonicus. Colors indicate base-pairing probabilities. (B) Multiple sequence alignment of control regions from 13 Scorpaeniformes species. Conserved sequence blocks (CSBs) are highlighted in different colors: CSB-F (orange), CSB-E (navy), CSB-D (yellow), CSB-1 (grey), CSB-2 (blue), and CSB-3 (green). Three novel conserved regions, Region-1 (purple), Region-2 (red), and Region-3 (brown), are also identified. Species names are abbreviated on the left.
The CR was positioned between tRNA-Pro and tRNA-Phe. In P. rubripinnis, it was 817 bp long with 61.57% A + T content and negative AT (− 0.0099) and GC (− 0.0764) skews. The I. japonicus CR was 1014 bp long with 64.99% A + T content and negative AT (− 0.0137) and GC (− 0.1324) skews. Multiple sequence alignment and conserved sequence analysis of the CRs from 13 Scorpaeniformes species revealed the presence of six previously described conserved sequence blocks (CSBs)49 in both P. rubripinnis and I. japonicus: CSB-1, -2, -3, -D, -E, and -F (Fig. 6B and Table S2). Additionally, we identified three novel CSBs (Region-1, Region-2, and Region-3), with Region-1 located upstream of CSB-F, and Region-2 and Region-3 situated between CSB-D and CSB-1 in Scorpaeniformes. While these regions were highly conserved across all examined Scorpaeniformes species. These novel CSBs may have functional roles similar to previously known CSBs in Scorpaeniformes.
Subsequently, two other I. japonicus mitogenome sequences were reported (accession numbers: MT604162 and MT375601)50,51. We compared our I. japonicus sequence with these two sequences and observed over 99.8% identity with both. While general mitogenomic features were consistent across studies, our analysis provides additional insights into tRNA and OL secondary structures, PCG codon usage, and novel Scorpaeniformes-specific CSBs.
Phylogenetic analysis
To elucidate the relationships among Scorpaeniformes families and reconstruct a higher-resolution interrelationship of the Scorpaeniformes species, we collected 13 mitochondrial PCGs from 71 Scorpaeniformes and three Notothenioidei species (outgroups), including 12 families and 31 genera (Table 2). Phylogenetic trees were constructed using Bayesian inference (Fig. 7A) and maximum-likelihood (Fig. 7B) methods, resulting in highly congruent topologies with strong posterior probabilities and bootstrap values. Our analysis revealed that Tetrarogidae and Synanceiidae, including P. rubripinnis and I. japonicus, respectively, formed a monophyletic clade that occupied an early-diverging position within the Scorpaeniformes phylogeny. This finding builds upon previous studies, which identified monophyly between these families yet could not determine their exact phylogenetic location6. A recent study using mitochondrial PCGs placed Synanceiidae at a basal position within Scorpaeniformes, albeit without including Tetrarogidae10. Our analysis, which incorporates both families, supports and extends these findings, providing additional insights into their phylogenetic placement.
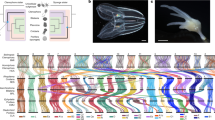
Phylogenetic trees of Scorpaeniformes based on 13 mitochondrial protein-coding genes. (A) Bayesian inference tree with posterior probability values indicated by colored circles at nodes. (B) Maximum likelihood tree with bootstrap support values shown by colored circles at nodes. The newly sequenced species (P. rubripinnis and I. japonicus) are highlighted in red. Family names are labeled on the right. Scale bars represent 0.1 substitutions per site.
The Scorpaenidae family, which includes most marine venomous fish with 26 genera and 223 species, has been subject to different phylogenetic interpretations13. Our analysis incorporates newly sequenced mitochondrial genomes from Tetrarogidae and Synanceiidae, supporting the phylogenetic relationship depicted in Fig. 1A rather than Fig. 1B. This result is particularly significant as it includes, for the first time, mitochondrial genome data from Tetrarogidae. While the overall topology of our phylogenetic tree is consistent with some previous studies8,9,10,13,24the inclusion of these new data provides a higher resolution of the interrelationships within Scorpaeniformes, especially regarding the phylogenetic positions of Tetrarogidae and Synanceiidae.
Discussion
Our comprehensive analysis of the complete mitochondrial genomes of P. rubripinnis and I. japonicus has provided significant new insights into the evolution of Scorpaeniformes, particularly regarding the phylogenetic placement of venomous fish families. While the mitogenomes exhibit the typical vertebrate mitochondrial structure52our findings extend beyond structural characterization. This study presents the first complete mitochondrial genome from the family Tetrarogidae, filling a crucial gap in Scorpaeniformes evolution. Although mitogenome organization is generally conserved across teleosts40complete mitogenome sequences are essential for robust phylogenetic analyses. Interestingly, the mitochondrial A + T content in both P. rubripinnis and I. japonicus is substantially higher than the Scorpaeniformes average, similar to what has been observed in S. verrucosa37. This shared characteristic among these species might reflect common evolutionary pressures or shared ancestry. In the mitochondrial control region, which is typically characterized by high sequence variability49we identified three novel conserved sequence blocks specific to Scorpaeniformes. The presence of these conserved elements within a traditionally variable region is noteworthy, and its universal conservation across all examined Scorpaeniformes species suggests it represents an ancestral characteristic of the order.
To further investigate the basal relationships within Scorpaeniformes, we expanded our analysis to include 13 mitochondrial PCGs from two Platycephalidae species, as a recent study suggested that Platycephalidae and Synanceiidae might occupy basal positions in the Scorpaeniformes phylogeny24. However, our results revealed that the phylogenetic position of Platycephalidae is highly sensitive to outgroup selection, as evidenced by contrasting topologies when using Notothenioidei versus Perciformes as outgroups (Fig. S2A and S2B). When using Notothenioidei as outgroups, Platycephalidae appeared in the most basal position, while with Perciformes outgroups, they clustered with Tetrarogidae and Synanceiidae in a non-basal clade. The consistently low bootstrap values (below 70) for deeper nodes in both analyses indicate fundamental uncertainty in these basal relationships, reflecting insufficient phylogenetic signal rather than technical limitations. To address this uncertainty, we investigated additional mitogenomic data, including the recently sequenced Inimicus didactylus38but this addition did not alter the tree topology since it represents another species within the already-sampled Synanceiidae family, thus providing limited new information for resolving basal Scorpaeniformes relationships. The current phylogenetic uncertainty likely stems from insufficient taxonomic sampling of sister groups, and the inclusion of mitogenomic data from closely related suborders (Cirrhitoidei, Serranoidei, Percoidei)13 would likely provide the phylogenetic signal necessary to resolve these basal relationships with greater confidence and reduce sensitivity to outgroup selection. Despite these uncertainties in deep relationships, our main phylogenetic analysis (Fig. 7) strongly supports the monophyly of Tetrarogidae and Synanceiidae, with these venomous fish families forming an early-diverging clade within Scorpaeniformes.
Recent advances in understanding venom evolution in terrestrial animals have been significantly aided by comparative genomic and transcriptomic approaches. For instance, studies of snake venoms have revealed how gene duplication and neofunctionalization contribute to toxin diversity53and similar approaches in scorpions have illuminated the molecular evolution of venom peptides54. Following these successful frameworks, our phylogenetic analysis of Tetrarogidae and Synanceiidae, despite uncertainties in their exact basal relationships, provides important evolutionary context for similar comparative analyses in venomous fish. The potential early divergence of these families, as suggested by our main phylogenetic analysis, indicates possible ancient origins of venom systems in Scorpaeniformes, making them valuable models for studying venom evolution in marine environments55. Comparative analyses of venom-related genes across these phylogenetic relationships will reveal patterns of molecular evolution and adaptation specific to fish venom systems. These evolutionary insights will guide future transcriptomic and proteomic investigations of fish venom diversity. Our findings highlight several key areas for future investigation. First molecular characterization of venom components across the Scorpaeniformes phylogeny is essential. Second, comparative analysis of venom transcriptome and proteome patterns will provide deeper insights into toxin evolution. Finally, expanded sampling of Tetrarogidae species will enhance our understanding of venom evolution within this family.
The integration of novel mitogenomic data with existing molecular evidence has significantly advanced our understanding of Scorpaeniformes evolution. Our findings, including the first complete mitochondrial genome from Tetrarogidae and the identification of distinct mitogenomic features, contribute to the growing body of molecular markers useful for fish phylogenetics. This phylogenetic framework, while acknowledging uncertainties in deep relationships, establishes a foundation for investigating various aspects of fish evolution, from molecular adaptation to morphological innovation. Moreover, the discovery of novel conserved sequence blocks and the supported monophyly of venomous fish families provides crucial molecular tools and evolutionary context for future research. As genomic and transcriptomic techniques continue to advance, our finding will be invaluable for future investigations of biodiversity and adaptation in Scorpaeniformes, particularly in understanding venom evolution and innovation in marine environments.
Materials and methods
Sample collection and DNA extraction
P. rubripinnis and I. japonicus specimens were collected from Geomun-do (34.1 N, 127.18 E), Yeosu, Republic of Korea. Total genomic DNA was extracted from 100 mg of muscle tissues using the cetyltrimethylammonium bromide (CTAB)-based method. Briefly, tissue samples were incubated in CTAB buffer (2% cetyltrimethylammonium bromide, 1.4 M NaCl, 20 mM EDTA, 100 mM Tris–HCl, and pH 8.0) at 56 °C for 1 h. Then, DNA was isolated using phenol/chloroform extraction followed by ethanol precipitation. The extracted DNA was resuspended in TE buffer and stored at − 20 °C until further use. All experimental protocols in this study were approved by the Institutional Animal Care and Use Committee of Chonnam National University (CNU IACUC-YB-2023-363). All methods were carried out in accordance with relevant guidelines and regulations, including the ARRIVE guidelines for reporting animal research.
Sequencing, mitochondrial genome assembly, and annotation
Genomic DNA libraries with 550 bp insert were constructed for P. rubripinnis and I. japonicus using the Illumina standard protocol (San Diego, USA). Paired-end sequencing generated 2 × 101 bp reads for P. rubripinnis, and 2 × 151 bp reads for I. japonicus. Raw reads were preprocessed using Trimmomatic (v0.36)56to remove adapter sequences, poly-N sequences, and low-quality bases. The parameters were set as follows: ILLUMINACLIP: TruSeq3-PE-2.fa:2:30:10 LEADING:3 TRAILING:3 SLIDINGWINDOW:4:15 MINLEN:36 for the 101 bp library and MINLEN:50 for the 151 bp library. Mitochondrial genome assembly was performed using two different approaches: Norgal (v1.0.0)57 with default parameters and NOVOPlasty (v2.7.2)58. For NoVOPlasty, we used COI coding sequences of each species (GeneBank: KU216091.1 for P. rubripinnis; KU199176.1 for I. japonicus) as seed sequences with parameters set to ‘Type = mito, K-mer = 39, Genome range = 12,000–22,000’. The assembled genomes from both methods showed high concordance, with only a 1 bp difference in length and > 99.9% sequence identity. We selected the NOVOPlasty assembly for subsequent analyses. Mitochondrial genome annotation was performed using MitoAnnotator52 to generate a gene map and identify genome features.
Bioinformatics analysis
The 13 protein-coding genes (PCGs) of P. rubripinnis and I. japonicus were extracted based on their annotation information, and relative synonymous codon usage (RSCU) scores were calculated using CAIcal59. Secondary tRNA gene structures were predicted using tRNAscan-SE60 with default parameters and visualized using PseudoViewer61. To analyze nucleotide composition bias, AT and GC skew values were calculated using the following formulas: AT skew = (A – T)/(A + T) and GC skew = (G – C)/(G + C)62.
To analyze the origin of light-strand replication (OL), the sense strand sequence forming the regular OL, located between tRNA-Asn (N) and tRNA-Cys (C), was extracted. The secondary OL structure was predicted using RNAstructure software63. To examine the control region, we aligned the control regions from mitogenomes of 12 species using the MUSCLE algorithm implemented in MEGA X software64 with default settings. Conserved sequence blocks within the control region were identified using DnaSP (v.6.12.03)65 with a window length of 8 and a conservation score of 6. Regions with a homozygosity score above 0.9 were designated as conserved sequence blocks.
Phylogenetic analysis
Phylogenetic relationships were investigated using mitogenomes of 71 Scorpaeniformes species and 3 Notothenioidei species, available in the MitoFish database (Table 2). The mitogenomes of the 3 Notothenioidei species (Parachaenichthys charcoti, Chionodraco hamatus, and Pagothenia borchgrevinki) were selected as the outgroup based on comprehensive phylogenomic analysis13 that demonstrated clear evolutionary separation between these groups using mitochondrial genome-scale data. Nucleotide sequences of the 13 protein-coding genes (PCGs) were extracted from the mitochondrial genome sequences using their annotation information. Multiple sequence alignment was performed using MAFFT (v7.407)66 with an --auto parameter. The resulting alignments were quality-checked and formatted using trimAl (v1.2rev59)67. A supermatrix was generated by concatenating the alignments of all 13 PCGs using phyutility (v2.7.1)68 for phylogenetic tree reconstruction. Bayesian inference analysis was conducted using MrBayes69 and implemented in the TOPALi (v2.5) platform70. The GTR + I + G model was applied with 300,000 generations. For the maximum-likelihood (ML) analysis, the best-fit model was determined using ModelFinder71. Then, the ML tree was inferred based on the concatenated alignment using IQTREE (v1.6.12)72 with 1000 bootstrap replicates and the GTR + F + R6 model.
Data availability
Whole genome sequencing data of P. rubripinnis and I. japonicus were deposited to the NCBI database with the following SRA accession numbers: SRR11802645 and SRR11802644, and BioProject accession numbers: PRJNA632998 and PRJNA633000, respectively. The complete mitogenomes of P. rubripinnis and I. japonicus are available with GenBank accession numbers MT506029 and MT506030, respectively.
References
Nelson, J. S. Fishes of the World (Wiley, 1994).
Hopkins, B. J. & Hodgson, W. C. Enzyme and biochemical studies of stonefish (Synanceja trachynis) and soldierfish (Gymnapistes marmoratus) venoms. Toxicon. 36, 791–793. https://doi.org/10.1016/S0041-0101(97)00167-0 (1998). https://doi.org:.
Edwards, O. & Wiley, G. D. J. A teleost classification based on monophyletic groups. (2010).
Wm Leo, S. & Matthew, T. C. Casting the percomorph net widely: The importance of broad taxonomic sampling in the search for the placement of serranid and percid fishes. Copeia. 2007, 35–55. https://doi.org/10.1643/0045-8511(2007)7[35:CTPNWT]2.0.CO;2 (2007).
Melanie, L. J. S. & David Johnson, L. R. P. G. Interrelationships of Fishes. (Academic, 1996).
Imamura, H. Phylogenetic relationships and new classification of the superfamily scorpaenoidea (Actinopterygii: Perciformes). Species Divers. 9, 1–36. https://doi.org/10.12782/specdiv.9.1 (2004).
Smith, W. L. & Wheeler, W. C. Polyphyly of the mail-cheeked fishes (Teleostei: Scorpaeniformes): evidence from mitochondrial and nuclear sequence data. Mol. Phylogenet. Evol. 32, 627–646. https://doi.org/10.1016/j.ympev.2004.02.006 (2004).
Lautredou, A. C. et al. New nuclear markers and exploration of the relationships among serraniformes (Acanthomorpha, Teleostei): The importance of working at multiple scales. Mol. Phylogenet. Evol. 67, 140–155. https://doi.org/10.1016/j.ympev.2012.12.020 (2013).
Cui, L. et al. The first two complete mitochondrial genomes for the family Triglidae and implications for the higher phylogeny of Scorpaeniformes. Sci. Rep. 7, 1553. https://doi.org/10.1038/s41598-017-01654-y (2017).
Xu, Z., Ye, Y., Yang, H., Xu, M. & Guo, B. The complete mitochondrial genome of Minous monodactylus (Minoinae: synanceiidae: Scorpaeniformes) and phylogenetic studies of Scorpaeniformes. Mitochondrial DNA Part. B. 4, 1670–1671. https://doi.org/10.1080/23802359.2019.1606684 (2019).
Smith, W. L., Elizabeth, E. & Clara, R. Phylogeny and taxonomy of flatheads, scorpionfishes, sea robins, and stonefishes (Percomorpha: Scorpaeniformes) and the evolution of the lachrymal saber. Copeia. 106, 94–119. https://doi.org/10.1643/CG-17-669 (2018).
Ishida, M. N. N. F. R. I. & Ono Hiroshima (Japan)). Phylogeny of the suborder Scorpaenoidei (Pisces: Scorpaeniformes). (1994).
Betancur, R. R. et al. Phylogenetic classification of bony fishes. BMC Evol. Biol. 17, 162. https://doi.org/10.1186/s12862-017-0958-3 (2017).
Vankan, M., Ho, S. Y. W. & Duchene, D. A. Evolutionary rate variation among lineages in gene trees has a negative impact on Species-Tree inference. Syst. Biol. 71, 490–500. https://doi.org/10.1093/sysbio/syab051 (2022).
Woese, C. R. On the evolution of cells. Proc. Natl. Acad. Sci. U. S. A. 99, 8742–8747. https://doi.org/10.1073/pnas.132266999 (2002).
Nussbaum, R. L. Genética En Medicina (Elsevier Health Sciences, 2008).
Arnold, M. L. Natural Hybridization and Evolution (Oxford University Press, 1997).
Maddison, W. P. Gene trees in species trees. Syst. Biol. 46, 523–536 (1997).
Alberts, B. et al. Molecular Biology of the Cell, vol. 3 (Garland, 1994).
Rokas, A., Williams, B. L., King, N. & Carroll, S. B. Genome-scale approaches to resolving incongruence in molecular phylogenies. Nature. 425, 798–804. https://doi.org/10.1038/nature02053 (2003).
Yandell, M. & Ence, D. A beginner’s guide to eukaryotic genome annotation. Nat. Rev. Genet. 13, 329–342. https://doi.org/10.1038/nrg3174 (2012).
Ekblom, R. & Wolf, J. B. A field guide to whole-genome sequencing, assembly and annotation. Evol. Appl. 7, 1026–1042. https://doi.org/10.1111/eva.12178 (2014).
Simon, C., Buckley, T. R., Frati, F., Stewart, J. B. & Beckenbach, A. T. Incorporating molecular evolution into phylogenetic analysis, and a new compilation of conserved polymerase chain reaction primers for animal mitochondrial DNA. Annu. Rev. Ecol. Evol. Syst. 37, 545–579. https://doi.org/10.1146/annurev.ecolsys.37.091305.110018 (2006).
Cui, L. et al. The first complete mitochondrial genome of the Flathead Cociella crocodilus (Scorpaeniformes: Platycephalidae) and the phylogenetic relationships within Scorpaeniformes based on whole mitogenomes. Genes. 10 https://doi.org/10.3390/genes10070533 (2019).
Zhang, B., Zhang, Y. H., Wang, X., Zhang, H. X. & Lin, Q. The mitochondrial genome of a sea anemone bolocera sp. exhibits novel genetic structures potentially involved in adaptation to the deep-sea environment. Ecol. Evol. 7, 4951–4962. https://doi.org/10.1002/ece3.3067 (2017).
Shi, X., Tian, P., Lin, R., Huang, D. & Wang, J. Characterization of the complete mitochondrial genome sequence of the globose head whiptail Cetonurus globiceps (Gadiformes: Macrouridae) and its phylogenetic analysis. PLoS One. 11, e0153666. https://doi.org/10.1371/journal.pone.0153666 (2016).
Wolstenholme, D. R. Genetic novelties in mitochondrial genomes of multicellular animals. Curr. Opin. Genet. Dev. 2, 918–925. https://doi.org/10.1016/s0959-437x(05)80116-9 (1992).
Barr, C. M., Neiman, M. & Taylor, D. R. Inheritance and recombination of mitochondrial genomes in plants, fungi and animals. New. Phytol. 168, 39–50. https://doi.org/10.1111/j.1469-8137.2005.01492.x (2005).
Dellaporta, S. L. et al. Mitochondrial genome of Trichoplax adhaerens supports placozoa as the basal lower metazoan phylum. Proc. Natl. Acad. Sci. U S A. 103, 8751–8756. https://doi.org/10.1073/pnas.0602076103 (2006).
Haddad, V., Stolf, H. O., Risk, J. Y., França, F. O. S. & Cardoso, J. L. C. Report of 15 injuries caused by Lionfish (pterois volitans) in aquarists in brazil: a critical assessment of the severity of envenomations. J. Venom. Anim. Toxins Including Trop. Dis. 21, 8. https://doi.org/10.1186/s40409-015-0007-x (2015).
Ngo, S. Y. & A, O. S. H. J. Ponampalam R stonefish envenomation presenting to a Singapore hospital. Singap. Med. J. 50, 506–509 (2009).
Church, J. E. & Hodgson, W. C. The Pharmacological activity of fish venoms. Toxicon 40, 1083–1093. https://doi.org/10.1016/S0041-0101(02)00126-5 (2002).
Chungthanawong, S. & Motomura, H. Two new species of the Waspfish genus Ablabys (Scorpaeniformes: Tetrarogidae) from the Western Pacific ocean. Ichthyol. Res. 66, 114–128. https://doi.org/10.1007/s10228-018-0665-0 (2019).
Shinohara, M. et al. A novel chemoattractant lectin, karatoxin, from the dorsal spines of the small scorpionfish Hypodytes rubripinnis. J. Pharmacol. Sci. 113, 414–417. https://doi.org/10.1254/jphs.10058SC (2010).
Smith, W. L. & Wheeler, W. C. Venom evolution widespread in fishes: a phylogenetic road map for the bioprospecting of piscine venoms. J. Hered. 97, 206–217. https://doi.org/10.1093/jhered/esj034 (2006).
Sato, Y., Miya, M., Fukunaga, T., Sado, T. & Iwasaki, W. MitoFish and mifish pipeline: A mitochondrial genome database of fish with an analysis pipeline for environmental DNA metabarcoding. Mol. Biol. Evol. 35, 1553–1555. https://doi.org/10.1093/molbev/msy074 (2018).
Wang, Q., Wang, J., Luo, J. & Chen, G. The complete mitochondrial genome of the Synanceia verrucosa (Scorpaeniformes: Synanceiidae). Mitochondrial DNA DNA Mapp. Seq. Anal. 27, 4234–4235. https://doi.org/10.3109/19401736.2015.1022756 (2016).
Li, Q., Li, H., Peng, Z., Yu, W. & Wei, S. Complete mitochondrial genome of Inimicus didactylus (Pallas, 1769) and its phylogenetic analysis. Mitochondrial DNA B Resour. 10, 304–308. https://doi.org/10.1080/23802359.2025.2482250 (2025).
Wu, Z., Wang, D., Hu, J. & Wang, Q. Complete mitochondrial genome of the Scorpaenopsis cirrhosa (Scorpaeniformes: Scorpaenidae). Mitochondrial DNA Part. A. 27, 3449–3450. https://doi.org/10.3109/19401736.2015.1063136 (2016).
Miya, M. et al. Major patterns of higher teleostean phylogenies: a new perspective based on 100 complete mitochondrial DNA sequences. Mol. Phylogenet. Evol. 26, 121–138. https://doi.org/10.1016/s1055-7903(02)00332-9 (2003).
Desjardins, P. & Morais, R. Nucleotide sequence and evolution of coding and noncoding regions of a quail mitochondrial genome. J. Mol. Evol. 32, 153–161. https://doi.org/10.1007/BF02515387 (1991).
Yue, G. H., Liew, W. C. & Orban, L. The complete mitochondrial genome of a basal teleost, the Asian Arowana (Scleropages formosus, Osteoglossidae). BMC Genom. 7, 242. https://doi.org/10.1186/1471-2164-7-242 (2006).
Yang, X. et al. Characterization of the complete mitochondrial genome sequence of Homalogaster Paloniae (Gastrodiscidae, Trematoda) and comparative analyses with selected digeneans. Parasitol. Res. 115, 3941–3949. https://doi.org/10.1007/s00436-016-5160-9 (2016).
Temperley, R. J., Wydro, M., Lightowlers, R. N. & Chrzanowska-Lightowlers, Z. M. Human mitochondrial mRNAs—like members of all families, similar but different. Biochim. Et Biophys. Acta (BBA) Bioenergetics. 1797, 1081–1085. https://doi.org/10.1016/j.bbabio.2010.02.036 (2010).
Yang, H. et al. Characterization of the complete mitochondrial genome sequences of three croakers (Perciformes, Sciaenidae) and novel insights into the phylogenetics. Int. J. Mol. Sci. 19 https://doi.org/10.3390/ijms19061741 (2018).
Watanabe, Y., Suematsu, T. & Ohtsuki, T. Losing the stem-loop structure from metazoan mitochondrial tRNAs and co-evolution of interacting factors. Front. Genet. 5, 109 (2014).
Lavrov, D. V., Brown, W. M. & Boore, J. L. A novel type of RNA editing occurs in the mitochondrial tRNAs of the centipede Lithobius forficatus. Proc. Natl. Acad. Sci. 97, 13738. https://doi.org/10.1073/pnas.250402997 (2000).
Fernández-Silva, P., Enriquez, J. A. & Montoya, J. Replication and transcription of mammalian mitochondrial Dna. Exp. Physiol. 88, 41–56. https://doi.org/10.1113/eph8802514 (2003).
Lee, W. J., Conroy, J., Howell, W. H. & Kocher, T. D. Structure and evolution of teleost mitochondrial control regions. J. Mol. Evol. 41, 54–66. https://doi.org/10.1007/BF00174041 (1995).
Kim, H. W. et al. The complete mitochondrial genome sequence and phylogenetic analysis for Inimicus japonicus (Scorpaeniformes: Synanceiidae). Mitochondrial DNA B Resour. 5, 3141–3142. https://doi.org/10.1080/23802359.2020.1800424 (2020).
Li, W. et al. The complete mitochondrial genome of Inimicus japonicus and its phylogenetic analysis. Mitochondrial DNA B Resour. 5, 3537–3538. https://doi.org/10.1080/23802359.2020.1823270 (2020).
Iwasaki, W. et al. MitoFish and mitoannotator: a mitochondrial genome database of fish with an accurate and automatic annotation pipeline. Mol. Biol. Evol. 30, 2531–2540. https://doi.org/10.1093/molbev/mst141 (2013).
Rao, W. Q. et al. The rise of genomics in snake venom research: recent advances and future perspectives. Gigascience. 11 https://doi.org/10.1093/gigascience/giac024 (2022).
Sunagar, K. et al. Evolution stings: the origin and diversification of Scorpion toxin peptide scaffolds. Toxins (Basel). 5, 2456–2487. https://doi.org/10.3390/toxins5122456 (2013).
Harris, R. J. & Jenner, R. A. Evolutionary ecology of fish venom: adaptations and consequences of evolving a venom system. Toxins (Basel). 11. https://doi.org/10.3390/toxins11020060 (2019).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for illumina sequence data. Bioinformatics. 30, 2114–2120. https://doi.org/10.1093/bioinformatics/btu170 (2014).
Al-Nakeeb, K., Petersen, T. N. & Sicheritz-Pontén, T. Norgal: extraction and de Novo assembly of mitochondrial DNA from whole-genome sequencing data. BMC Bioinform. 18, 510. https://doi.org/10.1186/s12859-017-1927-y (2017).
Dierckxsens, N., Mardulyn, P. & Smits, G. NOVOPlasty: de Novo assembly of organelle genomes from whole genome data. Nucleic Acids Res. 45, e18–e18. https://doi.org/10.1093/nar/gkw955 (2016).
Puigbò, P., Bravo, I. G., Garcia-Vallve, S. & CAIcal A combined set of tools to assess codon usage adaptation. Biol. Direct. 3, 38. https://doi.org/10.1186/1745-6150-3-38 (2008).
Lowe, T. M. & Chan, P. P. tRNAscan-SE On-line: integrating search and context for analysis of transfer RNA genes. Nucleic Acids Res. 44, W54–W57. https://doi.org/10.1093/nar/gkw413 (2016).
Byun, Y. & Han, K. PseudoViewer: web application and web service for visualizing RNA pseudoknots and secondary structures. Nucleic Acids Res. 34, W416–W422. https://doi.org/10.1093/nar/gkl210 (2006).
Perna, N. T. & Kocher, T. D. Patterns of nucleotide composition at fourfold degenerate sites of animal mitochondrial genomes. J. Mol. Evol. 41, 353–358. https://doi.org/10.1007/BF00186547 (1995).
Reuter, J. S. & Mathews, D. H. RNAstructure: software for RNA secondary structure prediction and analysis. BMC Bioinform. 11, 129. https://doi.org/10.1186/1471-2105-11-129 (2010).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 35, 1547–1549. https://doi.org/10.1093/molbev/msy096 (2018).
Rozas, J. et al. DnaSP 6: DNA sequence polymorphism analysis of large data sets. Mol. Biol. Evol. 34, 3299–3302. https://doi.org/10.1093/molbev/msx248 (2017).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780. https://doi.org/10.1093/molbev/mst010 (2013).
Capella-Gutiérrez, S., Silla-Martínez, J. M. & Gabaldón, T. TrimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics. 25, 1972–1973. https://doi.org/10.1093/bioinformatics/btp348 (2009).
Smith, S. A. & Dunn, C. W. Phyutility: a phyloinformatics tool for trees, alignments and molecular data. Bioinformatics. 24, 715–716. https://doi.org/10.1093/bioinformatics/btm619 (2008).
Ronquist, F. & Huelsenbeck, J. P. MrBayes 3: bayesian phylogenetic inference under mixed models. Bioinformatics. 19, 1572–1574. https://doi.org/10.1093/bioinformatics/btg180 (2003).
Milne, I. et al. TOPALi: software for automatic identification of Recombinant sequences within DNA multiple alignments. Bioinformatics. 20, 1806–1807. https://doi.org/10.1093/bioinformatics/bth155 (2004).
Kalyaanamoorthy, S., Minh, B. Q., Wong, T. K. F., von Haeseler, A. & Jermiin, L. S. ModelFinder: fast model selection for accurate phylogenetic estimates. Nat. Methods. 14, 587–589. https://doi.org/10.1038/nmeth.4285 (2017).
Nguyen, L. T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: A fast and effective stochastic algorithm for estimating Maximum-Likelihood phylogenies. Mol. Biol. Evol. 32, 268–274. https://doi.org/10.1093/molbev/msu300 (2014).
Acknowledgements
This research was supported by Korea Institute of Marine Science & Technology Promotion (KIMST) funded by the Ministry of Oceans and Fisheries (RS-2025-02292973) and the Global-Learning & Academic research institution for Master’s · PhD students, and Postdocs (LAMP) Program of the National Research Foundation of Korea (NRF) grant funded by the Ministry of Education (No. RS-2024-00442775).
Author information
Authors and Affiliations
Contributions
S.-G.L. designed the study and performed the experiments. S.-G.L. and S.K. conducted the mitogenome analyses. S.-G.L., S.K., and C.P. wrote the manuscript. C.P. supervised and contributed to the project coordination. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lee, SG., Kim, S. & Park, C. Complete mitogenomes of venomous fish Paracentropogon rubripinnis and Inimicus japonicus elucidate phylogenetic relationships in Scorpaeniformes. Sci Rep 15, 28596 (2025). https://doi.org/10.1038/s41598-025-05085-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-05085-y