Abstract
As social changes accelerate, the incidence of psychosomatic disorders has significantly increased, becoming a major challenge in global health issues. This necessitates an innovative knowledge system and analytical methods to aid in diagnosis and treatment. Here, we establish the ontology model and entity types, using the BERT model and LoRA-tuned LLM for named entity recognition, constructing the knowledge graph with 9668 triples. Next, by analyzing the network distances between disease, symptom, and drug modules, it was found that closer network distances among diseases can predict greater similarities in their clinical manifestations, treatment approaches, and psychological mechanisms, and closer distances between symptoms indicate that they are more likely to co-occur. Lastly, by comparing proximity scores, it was shown that symptom-disease pairs in primary diagnostic relationships have a stronger association and are of higher referential value than those in diagnostic relationships. The research results revealed the potential connections between diseases, co-occurring symptoms, and similarities in treatment strategies, providing new perspectives for the diagnosis and treatment of psychosomatic disorders and valuable information for future mental health research and practice.
Similar content being viewed by others
Introduction
In recent years, due to rapid economic development and a faster pace of life, the number of psychosomatic disorder patients has been increasing annually1. Traditional treatment for psychosomatic disorders involves professional psychological counseling. However, most patients harbor fears and a resistant attitude towards it, and both patients and their relatives lack professional knowledge related to psychological healthcare. Additionally, the concealment and complexity of psychosomatic disorders themselves make it difficult for these conditions to be promptly detected and intervened2,3. This necessitates an innovative knowledge system and analytical method, serving as a specialized data foundation for a medical information intelligent decision-making system, to assist in diagnosis and treatment4,5.
A knowledge graph (KG) is a robust semantic network that can be used to establish a framework for complex psychosomatic disorder data. The classic KG is a graph data structure composed of knowledge points6,7,8. Each knowledge point is represented by a subject-predicate-object triplet (SPO), where the subject (S) and object (O) describe entities, and the predicate (P) represents the relationship. The SPO triplet structure is suitable for expressing qualitative knowledge (for example, the triplet < S: olanzapine, P: treats, O: schizophrenia > can express that the drug Olanzapine can treat schizophrenia).
KG plays a crucial role in the medical field, especially in areas such as drug discovery, clinical decision support systems, and drug recommendations9,10. By integrating multi-source heterogeneous data, KG can assist researchers in identifying potential drug targets, optimizing treatment plans, and supporting personalized medical decisions8,11. However, despite significant advancements in the application of KG in the medical field, its utilization in psychiatry remains relatively limited12. This limitation primarily stems from the unique characteristics of psychiatric data: firstly, psychiatric texts often include lengthy passages and sparse data, which increase the difficulty of knowledge extraction and modeling13,14; secondly, psychiatric knowledge is often fragmented and lacks systematic organization, making it challenging for traditional general-purpose medical KGs to be directly applied to psychiatric diagnostic and treatment processes15.Moreover, existing KG construction methods primarily focus on visualization and basic information retrieval, with insufficient emphasis on in-depth graph structure analysis. This limitation hampers their ability to support complex clinical decisions16.
The current KG construction methods face two major issues in their application to psychiatry:
Lack of specificity in ontology models: Existing KG construction methods are mostly based on ontology models designed for general medical domains, failing to adequately account for the unique characteristics of psychiatric diagnostic processes. For instance, the diagnosis of psychiatric disorders often relies on subjective symptom descriptions and unstructured texts, which are challenging for existing models to capture effectively.
Insufficient graph structure analysis: Current research largely focuses on the construction and visualization of KGs while neglecting in-depth graph structure analysis. This shortcoming limits the effectiveness of KGs in supporting complex clinical decisions, such as disease diagnosis and treatment recommendations.
These issues restrict the accuracy and practicality of KG-based question-answering systems in psychiatry, and they also hinder effective support for doctors’ diagnostic processes.
To address these issues, this study proposes a KG construction and analysis method tailored for the psychiatric domain. Specifically, the study employs the following methods:
Employing BERT and large language models (LLMs) for named entity recognition (NER) of psychiatric texts, and constructing SPO triples to create a high-quality psychiatric KG13,17,18,19,20,21,22.
Utilizing graph theory analysis methods to perform an in-depth analysis of the graph structure of the KG, with a particular focus on the topological relationships among disease, symptom, and drug nodes23.
Related work
The construction of biological KGs from plain text has emerged as a critical task in biomedical research, enabling the integration of heterogeneous data and facilitating knowledge discovery. However, extracting structured knowledge from unstructured text poses significant challenges, including entity recognition, relation extraction, and ontology alignment. In this section, we review existing approaches and tools for constructing KGs in the biomedical domain, highlighting their strengths and limitations.
Entity extraction methods in KG construction
When constructing a KG from text, entity extraction is a key step. Traditional entity extraction methods mainly rely on Named Entity Recognition (NER), a core task in the field of Natural Language Processing (NLP). NER aims to identify and classify named entities in the text, such as diseases, symptoms, and drugs.
Early entity extraction methods were mainly based on dictionary matching and rule-based methods, which typically relied on medical knowledge bases (e.g., UMLS (Unified Medical Language System)24, MeSH (Medical Subject Headings)) for term matching25. Although these methods performed well in specific domains, their adaptability was limited, and they struggled to handle out-of-vocabulary (OOV) words or the contextual information in the text. To address this, researchers introduced statistical learning methods, such as Hidden Markov Models (HMM)26 and Conditional Random Fields (CRF)27, to model sequence labeling tasks, improving the generalization ability of entity recognition. However, traditional statistical learning methods are highly dependent on feature engineering, requiring manual design of features such as part-of-speech tags, context windows, and character n-grams, making them difficult to adapt to complex structures in large-scale data.
In recent years, with the rise of deep learning, neural network-based NER methods have made significant progress. Bidirectional Long Short-Term Memory (BiLSTM) combined with CRF further improved the performance of sequence labeling tasks28, while Convolutional Neural Networks (CNN) enhanced entity recognition by extracting local features22. The emergence of pre-trained language models (PLM) has greatly enhanced the contextual modeling ability for NER tasks. BERT (Bidirectional Encoder Representations from Transformers) with its bidirectional attention mechanism can capture richer semantic information, achieving state-of-the-art performance in NER13. In the biomedical domain, due to the specialized nature and complexity of medical texts, domain-specific pre-trained models, such as BioBERT29 and SciBERT30, have been proposed. These models are further trained on medical literature data (e.g., PubMed, PMC, Medline), significantly improving the accuracy of entity recognition.
Recently, large language models (LLMs) have also been applied to Named Entity Recognition (NER) tasks, leveraging their extensive pre-trained knowledge to improve entity extraction31. Methods based on LLMs utilize contextual embeddings to capture complex relationships and dependencies between entities, offering a more flexible and powerful approach to NER.
LangChain in retrieval-augmented generation (RAG)
LangChain is a framework designed to streamline the integration of retrieval and generative models, playing a key role in the implementation of Retrieval-Augmented Generation (RAG) techniques. It simplifies the process of combining large language models (LLMs) with information retrieval, enabling systems to access external knowledge to enhance the relevance and accuracy of generated content31. LangChain offers a flexible interface to integrate various retrieval sources, such as databases or documents, into the generative process, allowing models to retrieve relevant information before generating responses. This framework has been widely adopted for tasks like question answering and summarization, where external knowledge is crucial.
Recent studies have shown that LangChain’s modular design and integration with retrieval engines make it an effective tool for building scalable RAG systems32. For instance, its ability to seamlessly combine dense retrieval methods with generative models has been demonstrated to improve performance in knowledge-intensive tasks33.
Ontologies for biomedical KGs
In the biomedical domain, ontologies play a vital role in organizing and standardizing knowledge across various medical and scientific disciplines. Key ontologies in this field include UMLS which integrates multiple biomedical terminologies and provides a comprehensive framework for linking diverse medical vocabularies24. Another important resource is SNOMED CT (Systematized Nomenclature of Medicine – Clinical Terms)34, a standardized clinical terminology that includes a wide range of medical concepts, enabling the integration of health information systems and enhancing data sharing. Additionally, the Gene Ontology (GO)35 provides a structured vocabulary for representing gene functions and biological processes, facilitating research in genomics and molecular biology. The Disease Ontology (DO)36 classifies diseases in a hierarchical structure, enabling the integration of disease-related data across disciplines and fostering collaboration in biomedical research.
In the context of mental health, ontologies play a critical role in standardizing the complex and heterogeneous terminology of psychiatric concepts. The Mental Disease Ontology (MDO), developed by the European Bioinformatics Institute (EBI), provides a formalized representation of mental disorders, including their classifications, causes, symptoms, and treatments. This ontology helps organize the complex relationships between psychiatric conditions and enables accurate mapping of mental health data. Alongside MDO, the Relation Ontology (RO)37 defines the relationships between biomedical entities, such as “is_a,” “part_of,” and “treats,” which are essential for linking mental disorders with broader biomedical concepts.
Current medical konwledge graph and graph analysis
Medical Knowledge Graphs (MKGs) have undergone significant evolution, transitioning from classical rule-based systems to advanced, data-driven frameworks. Early efforts, such as UMLS24 and SNOMED CT34, established the foundational standards for organizing medical terminology and ontologies. These systems excelled in interoperability and semantic consistency, but they faced limitations in capturing the complex, dynamic relationships and contextual nuances of medical data. Building upon these foundations, OpenBEL38 introduced a framework integrating biological and clinical knowledge, enabling a more detailed representation of disease mechanisms and drug interactions. Additionally, SemMedDB39 further enriched medical knowledge representation by extracting semantic relationships from the literature.
To overcome the limitations of classical methods, recent MKGs leverage machine learning and graph-based techniques to enhance knowledge representation and reasoning capabilities. For example, BioKG40 integrates multiple biomedical data sources, such as drug-target interactions and gene-disease associations, unifying them into a graph structure that allows for more accurate predictions in tasks like drug repurposing. Similarly, DRKG41 employs advanced embedding methods to model the interactions between drugs, diseases, and genes, supporting precision medicine applications. Furthermore, Hetionet42 combines heterogeneous data types from pharmacology and genomics to facilitate large-scale biomedical discoveries. In recent years, PrimeKG43 has further expanded this field by integrating multimodal data (such as patient electronic health records and genomic data), creating a comprehensive KG that provides stronger support for personalized medicine.
In recent years, the analysis of MKGs has also become increasingly in-depth. For instance, investigates the treatment modules (subgraphs) formed by the COVID-19 proteins and the proteins affected by other diseases, studying their proximity to predict potential COVID-19 therapeutic drugs44. By analyzing the topological relationships between disease symptoms23, Chinese medicine targets, and human protein-protein interactions (PPI), it was found that proteins related to symptoms tend to cluster within local PPI modules44,45. The network proximity between herbal medicine targets and symptom modules suggests the effectiveness of herbal treatments for symptoms46,47.
Materials and methods
Konwledge graph construction process
We adopts a top-down approach to construct the mental illness KG through four key steps:
First, the ontology schema layer is constructed. As the foundation of the KG, this step involves defining entities, attributes, and relationships, thus laying the basic framework for the entire graph. The ontology schema was confirmed by integrating expert opinions from doctors, hospital admission records, doctor-patient communication records, and diagnostic conclusions.
Next, data collection and preprocessing are carried out. Based on the ontology schema designed in the previous step, this study collected real patient data from the Psychiatry Department of Zhongda Hospital, Southeast University. After strict screening and cleaning to remove redundant and invalid information, each case was segmented into 16 paragraphs according to different tasks, laying a solid foundation for subsequent knowledge extraction.
Subsequently, knowledge extraction is implemented. This step focuses on identifying named entities and their relationships from the preprocessed data. BERT and fine-tuned LLM models were employed for named entity recognition, combined with rule-based relationship extraction methods, to effectively extract triplet information from large volumes of data, completing the knowledge extraction process.
Finally, knowledge storage and querying are performed. We selected the Neo4j graph database as the storage tool, utilizing Cypher query language for data retrieval and visualization. This step not only realized the storage of the mental illness KG but also provided a foundation for its future application and expansion.
In terms of storage and management, our study adopts RDF (Resource Description Framework) as the storage format for the KG, rather than traditional relational database management systems (DBMS). Given the complex relationships and non-structured data, such as doctor-patient communication records, involved in the mental illness KG, RDF is particularly suitable for representing these multidimensional relationships in a graph model, supporting semantic querying and reasoning. In contrast, traditional DBMSs are less effective at expressing such complex associations, making RDF more advantageous in this application scenario.
Dataset
The field of psychosomatic disorders currently lacks standardized annotated data resources. Our data comes from 262 patient medical records in the psychiatric department of Zhongda Hospital in Nanjing, China, including 172 female patients. All samples were from inpatients with a hospital stay of more than 10 days and a primary diagnosis of psychiatric disorders. Patients were excluded if they had mental disorders caused by substance abuse or organic diseases, severe abnormalities in heart, liver, or kidney function, or were pregnant or lactating women. We performed data cleaning and preprocessing on the medical records and produced two differently formatted NER datasets. The first type of NER dataset uses the “BIO” (Begin: the beginning character of an entity, Inside: a middle or ending character of an entity, Outside: a character that is not part of an entity) scheme for annotation48,49. Each character in the text sequence is tagged with a “B-”, “I-” and “O” label to indicate whether the character is part of a named entity. This dataset divides entities into 7 types. See Table 1.
An example from our dataset is presented below:
J + 01013897 has been experiencing low mood for half a year. The patient developed low mood after a friend dropped out of school half a year ago, showing a lack of interest in activities and difficulty concentrating. The patient visited an external hospital for a CT scan of the head and sinuses, which showed no significant abnormalities. The patient is conscious, mentally alert, has poor appetite, normal bowel movements, and lacks pleasure with low mood. Based on the medical history and current presentation, the current diagnosis of mild depression is agreed upon. It should be differentiated from conduct disorder. Treatment involves the administration of the antidepressant sertraline.
The BIO label for each Chinese character in this sentence is as shown in Table 2.
The second dataset is used for NER tasks with LLM. Initially, medical cases are divided into 16 sections based on different tasks, then dataset is constructed using each section separately. Examples of the dataset are shown in Fig. 2.B. The 16 sections include Patient number, Chief complaint, Course of disease, Severity, Summary of primary symptoms, Admission status, Mental examination, Medical history, Risk factors, Inducement, Treatment plan, Primary diagnostic basis, Primary diagnosed disease, Differential diagnostic basis, Differential diagnosed disease and Summary of signs. An example of 16 sections is shown as follows:
-
(1)
Patient number: 0101447813.
-
(2)
Chief complaint: Unable to feel happy, gloomy and joyless. Unable to concentrate. Fatigued and weak, lack of energy. Having had negative thoughts. Irritable and anxious, feeling of panic and chest tightness. Trembling hands, headache. Irregular sleep patterns, day-night reversal. Poor appetite.
-
(3)
Course of disease: A year ago, the patient began to experience low mood due to family issues and gastrointestinal disease, feeling persistently unhappy and unable to cheer up, with difficulty concentrating, unable to stop thinking about past sad events, feeling let down by parents, often self-blaming, feeling tired and weak, lacking energy, having passive thoughts but never acting on them, accompanied by symptoms of irritability, panic, chest tightness, trembling hands, and headaches. Irregular sleep patterns, reversal of day and night, poor appetite.
-
(4)
Severity: Unable to live and study normally, took a leave of absence to recuperate the gastrointestinal tract at home, afterward the gastrointestinal disease gradually improved, but emotional problems worsened, preventing a normal return to study.
-
(5)
Summary of primary symptoms: Symptoms include palpitations, chest tightness, trembling hands, and headaches. Insight is present. Always feeling down, unable to cheer up, difficulty concentrating, unable to control thoughts of past sad events, often self-blaming, feeling tired and weak, lacking energy, having negative thoughts but never acting on them, accompanied by symptoms of irritability, palpitations, chest tightness, trembling hands, and headaches. Sleep is irregular, with a reversed sleep cycle and poor appetite. Bowel and bladder functions are normal, sleep is average, and there has been no significant recent weight change.
-
(6)
Admission status: Yesterday, visited the psychiatric outpatient clinic at Zhongda Hospital, where they diagnosed a “depressive episode” and admitted for treatment.
-
(7)
Mental examination: Psychiatric evaluation: Conscious and oriented, passive engagement, rapid speech, no perceptual disturbances elicited, slightly active thinking, presence of pathological circumstantiality and obsessive thoughts, unstable emotions, decreased interest, poor concentration, impaired memory, reduced volition, poor appetite, sleep rhythm disturbances, with symptoms of palpitations, chest tightness, trembling hands, and headaches. Insight is present.
-
(8)
Medical history: One year ago, the patient began to experience low mood due to family issues and gastrointestinal disease.
-
(9)
Risk factors: Family factors: there are significant conflicts within the father’s family, and the grandparents argue daily.
-
(10)
Inducement: no obvious inducement.
-
(11)
Treatment plan: The treatment plan should include antidepressants and mood stabilizers, supplemented with psychotherapy.
-
(12)
Primary diagnostic basis: The patient speaks quickly because it is their natural manner, not because of the typical excitability associated with excessive talking.
-
(13)
Primary diagnosed disease: Diagnosed with a depressive episode with mixed features.
-
(14)
Differential diagnostic basis: The patient speaks quickly because it is their natural manner, not because of the typical excitability associated with excessive talking.
-
(15)
Differential diagnosed disease: bipolar disorder, mania.
-
(16)
Summary of signs: Sleep patterns are irregular, with day and night reversed, and appetite is poor. Emotional issues have gradually worsened. The patient is conscious and oriented, interacts passively, speaks quickly, exhibits no perceptual disturbances, has slightly active thoughts, pathologically circumstantial and obsessive thinking, unstable emotions, decreased interest, poor concentration, impaired memory, reduced volitional actions, poor appetite, disordered sleep rhythm, and symptoms including palpitations, chest tightness, trembling hands, and headaches. Insight is present.
The third dataset is utilized for querying text content. Initially, we structure the text content and subsequently query the medical records. Ultimately, we hope that the LLM can use the content of the medical records or its own summarized statements to respond.
BIO-labeled dataset and name entity recognition by RoBERTa model
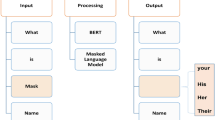
The BERT language model13,17, which utilizes the Transformer as a feature extractor50,51,52,53, is a deeply trained bidirectional language model based on attention mechanisms. It has achieved outstanding results in various tasks in the Natural Language Processing (NLP) field, such as question answering systems, natural language inference, and NER, among others. This study uses the RoBERTa Chinese pre-trained character vector model, which is improved based on Google’s official BERT pre-trained model for Chinese, and fine-tuned according to the NER task.
Given a sentence\(\:\left[{W}_{1},\:{W}_{2},\:\:...,{\:W}_{N}\right]\), where Wi represents a Chinese character in the sentence, and N represents the maximum length of the sentence. This sentence needs to be prefixed with a [CLS] token and suffixed with a [SEP] token before input. The [CLS] token (short for “classification”) is a special token used to aggregate information from the entire sequence, often serving as the representation for classification tasks.The [SEP] token (short for “separator”) is used to mark the end of a sentence and to separate two sentences in tasks. Following this, character embedding is performed by querying a character vector table, representing each character as a one-dimensional vector. For the NER task, the RoBERTa model is trained to predict the “BIO” labels for each character in the sequence. Therefore, we fine-tune the RoBERTa model with the first dataset, adding a classification layer to determine the label type of each character. The final pipeline is depicted in Fig. 1.
Fine-tuning the RoBERTa model for the NER task.
Fine-tuning dataset and name entity recognition by fine-tuned LLaMA model
LoRA is a low-rank adaptation fine-tuning method that significantly reduces the number of trainable parameters for downstream tasks and achieves excellent results54. For the pre-trained weight matrix\(\:\:{W}_{0}\in\:{R}^{d\times\:k}\), it can be updated via low-rank decomposition \(\:{W}_{0}+\varDelta\:W={W}_{0}+BA\) where \(\:B\in\:{R}^{d\times\:r}\), \(\:A\in\:{R}^{r\times\:k}\) and \(\:r\ll\:min\left(d,k\right)\). During the training process, \(\:{W}_{0}\) is frozen and does not receive gradient updates, while \(\:A\) and \(\:B\) contain trainable parameters. Figure 2A is a schematic diagram of the LoRA principle, where matrix\(\:\:A\) is initialized with random Gaussian values, and matrix \(\:B\) is initialized to zero. When the input is \(\:x\), for \(\:h={W}_{0}x\), the modified forward propagation:
We use both the pre-trained LLaMA model and the LLaMA model fine-tuned with LoRA from the KnowLM55,56,57,58,59,60 for entity extraction. This method uses the second type of dataset we created. When performing NER tasks with this project, it requires inputs of “Instruction” and “Input”. “Instruction” provides detailed information about the task to be performed by the model. “Input” is the text information received by the LLM, forming the basis for the model to perform calculations and generate outputs. Subsequently, the LLM returns an “Output” result, which is determined by the “Instruction” and “Input” provided by the user. In this research, we define “Instruction” as “Extract possible entities and their types from the given text, with optional entity types are [‘patient’, ‘symptom’, ‘drug’, ‘diagnosed disease’, ‘differential diagnosed disease’, ‘risk factor’, ‘severity’], answer in the format of (entity, entity type)”. The final pipeline is depicted in in Fig. 2B.
(A) Schematic diagram of the LoRA principle. (B) Process for LLM to extract entities and their types. First, the LLM is fine-tuned with LoRA using the second dataset, and upon completion of training, the fine-tuned LLM is used to perform the NER task.
Entity alignment
Through entity extraction, we obtained approximately 30,000 entities, which had a significant amount of duplication (for instance, if both case A and case B contain the entity “schizophrenia”, then the entity “schizophrenia” would be extracted from both cases) and errors. Therefore, we performed data cleaning and integration on the extracted entities24. Then, we undertook a crucial data processing step, namely the standardization and unification of entity names61,62,63. This step was accomplished by calculating the degree of approximation between entities, including textual and semantic similarity of entities. Textual similarity of entities was determined by calculating the proportion of identical characters between two entities to decide if they represent the same expression64. The similarity was calculated using the Jaccard coefficient:
Where\(\:\:{sim}_{t}\left(A,B\right)\) represents the textual similarity between entity \(\:A\) and entity \(\:B\), \(\:|A\cap\:B|\) represents the number of identical characters is between the two entities, and \(\:|A\cup\:B|\) represents the total number of characters for both entities.
Semantic similarity calculates the similarity between two entities based on their approximation in contextual semantics. The method involves calculating the cosine of the word vectors between entities. The closer the cosine value is to 1, the smaller the vector angle distance, and the greater the approximation between the two entities:
Here, \(\:{sim}_{s}\left(A,B\right)\) represents the semantic similarity between entity A and entity B, \(\:A=[{A}_{1},{A}_{2},...,{A}_{n}]\) represents the word vector set of entity A, and \(\:B=[{B}_{1},{B}_{2},...,{B}_{n}]\) represents the word vector set of entity B.
Due to the excessive similarity in the text of some different entities, relying solely on textual similarity can easily lead to errors in entity alignment. For example, “bipolar disorder” and “MDD” are two different diseases, but they have a high textual similarity in Chinese. To address this issue, we calculate entity similarity by combining textual similarity and semantic similarity with weighted contributions:
Where \(\:sim\left(A,B\right)\) represents the entity similarity between entity A and entity B. \(\:{\omega\:}_{t}\) denotes the weight assigned to the textual similarity \(\:{sim}_{t}\left(A,B\right)\), and \(\:{\omega\:}_{s}\:\)denotes the weight assigned to the semantic similarity \(\:{sim}_{\text{s}}\left(A,B\right)\), with the constriant that \(\:{\omega\:}_{t}+{\omega\:}_{s}=1\).
To determine the optimal weight allocation(\(\:{\omega\:}_{t}\) and \(\:{\omega\:}_{s}\)), we selected 1000 entity pairs from approximately 30,000 entities using stratified random sampling based on entity type distribution, ensuring the representativeness of the sample. Each pair consisted of entities of the same type. Three domain experts independently annotated these entity pairs, and the final labels were determined by majority voting.
For weight optimization, we employed a grid search approach with a step size of 0.01, exploring all possible weight combinations within the range [0, 1]. The evaluation metrics included accuracy, precision, recall, and F1-score, with the F1-score serving as the comprehensive evaluation metric.
Evaluation metrics under different textual similarity weights.
As shown in Fig. 3, the experimental results indicate that accuracy reached its peak of 0.92 at a textual similarity weight of 0.45, precision achieved its maximum value of 0.92 at a weight of 0.38, and recall peaked at 0.63 with a weight of 0.41. A notable characteristic of entity alignment tasks is the extreme imbalance between positive and negative samples: the number of non-matching entity pairs typically far exceeds that of matching pairs. This imbalance limits the usefulness of accuracy as a sole evaluation metric, as a model that predicts all entity pairs as non-matching could still achieve high accuracy without reflecting its true alignment capability. Therefore, in entity alignment tasks, relying solely on accuracy is insufficient, and it is essential to incorporate precision, recall, and F1-score for a comprehensive evaluation.
In our experiments, we particularly focused on balancing precision (to minimize false matches) and recall (to reduce missed matches). The results demonstrate that when the textual similarity weight is set to 0.4, the model achieves an optimal balance between precision (0.91) and recall (0.63), with an F1-score of 0.74, significantly outperforming other weight combinations.
Therefore, we calculate entity similarity by assigning weights of 0.4 and 0.6 to textual similarity and semantic similarity, respectively:
In this study, we opted not to link entities directly to ontology, instead relying on the combined textual and semantic similarity approach described above. This choice was motivated by the nature of our Chinese mental health clinical records, which often use natural language descriptions that do not fully align with standardized ontology terms. For instance, expressions like “low mood”and “depressed mood” are semantically similar, but linking them directly to an ontology might introduce mapping errors due to contextual nuances or incomplete ontology coverage in Chinese. Our similarity-based method preserves the diversity of these expressions without requiring external knowledge bases, offering greater flexibility. This is particularly advantageous for our small, domain-specific dataset, enabling rapid KG construction tailored to our analysis needs.
Relying on similarity-based alignment, our KG may retain a greater number of distinct entities, potentially resulting in a larger graph size compared to ontology-based methods, which tend to consolidate such entities into a single standardized term. However, this increased scale preserves subtle distinctions, enhancing downstream analyses—such as symptom clustering—by capturing a broader spectrum of clinical expressions. In contrast, ontology linking simplifies the KG by minimizing redundancy and enforcing uniform terminology, which could facilitate tasks like cross-dataset integration or standardized querying, though it risks oversimplifying contextual variations critical to our specific domain.
For entities with high similarity, their entity names are standardized and unified. For example, the expression “bad temper” is used to replace similar terms like " lose temper easily” and “irritable”. “diagnosed disease” refers to the condition identified through initial examination upon hospital admission; “primary diagnosed disease” and “differential diagnosed disease” are conclusions reached after thorough discussion and analysis by the medical team. Given that all three types essentially fall under the category of diseases, we have consolidated them into a unified “disease” entity node, distinguishing their specific types through three distinct relationships: “primary diagnosis”, “diagnosis”, and “differential diagnosis”. Ultimately, 3285 entities were obtained. Through such standardized processing, the KG can organize and link information more effectively. This improves the accuracy of queries and the search experience for users. It also enhances the quality of the graph, laying a solid foundation for subsequent analysis of the graph structure.
Metrics
In this section, we describe the various metrics used to evaluate the performance of our models and the relationships within the psychosomatic disorder KG. The evaluation includes both traditional graph-based metrics, such as LCC (Largest Connected Component) and network distance, and commonly used machine learning metrics, such as precision, recall, and F1 score. Additionally, we introduce BLEU and ROUGE metrics for assessing the quality of model-generated text, as well as efficiency metrics like samples per second and steps per second.
LCC and LCC z-score
The z-score of the Largest Connected Component (LCC)65 of a set of nodes is used to describe the positioning of node sets in the psychosomatic disorder knowledge network. The LCC refers to the largest subset of nodes in a network where each node is reachable from any other node within the subset, and no additional nodes outside the subset can be included while maintaining this connectivity. We calculate the size of the LCC formed by the set of nodes, and then compare the calculated LCC size with the expected LCC of a randomly selected set of nodes. The LCC z-score(\(\:{z}_{LCC}\)) is the difference between the LCC size and the mean of randomization µ(random LCC), divided by the SD of the randomization σ(random LCC):
An LCC z-score greater than the expected mean indicates that the observed LCC is significantly larger than expected, meaning the node set aggregates into a local module.
Network distance Dab and network separation Sab
We measure the network relationship between two sets of nodes using network distance \(\:{D}_{ab}\) and network separation \(\:{S}_{ab}\)46,66. Network distance \(\:{D}_{ab}\), also denoted as < dab>, is the average of the shortest network distance between all pairs of nodes in the two sets of nodes:
Where \(\:\left|A\right|\) and \(\:\left|B\right|\) represents the number of nodes in two sets of nodes. Network separation compares the average shortest distance within each node set < daa> and < dbb> with the average shortest distance < dab> between node sets \(\:A\) and \(\:B\):
The random expectation of Sab is zero; a negative Sab indicates that the two sets of nodes are in the same network neighborhood, while a positive Sab indicates that the two sets of nodes are topologically separated.
Semantic similarity
We define semantic similarity to evaluate the biological and psychological mechanism similarities between diseases and between drug67,68. we constructed an association matrix. Taking the symptom module of diagnosed diseases as an example, where each row represents a disease, each column represents a symptom, and the elements in the matrix (0 or 1) indicate whether there is an association between them. Using this association matrix, we calculate semantic similarity using the Wang method69:
Where \(\:g1\:\)and \(\:g2\) are two diseases, \(\:{T}_{g1}\:\)and \(\:{T}_{g2}\) represent the sets of symptoms or drug corresponding to diseases \(\:g1\) and \(\:g2\), respectively, and \(\:IC\left(g\right)\) denotes the number of symptoms or drug associated with disease \(\:g\).
Network proximity distance and z-score
We define the network proximity metric (referred to in the text as “proximity distance d”) as the average distance from all points in node set \(\:A\) to the nearest points in node set B among two sets of nodes46,66:
Where \(\:\left|A\right|\) represents the number of nodes in node set \(\:A\), \(\:dist\left(a,b\right)\) represents the shortest distance between nodes a and b. Then, we simulated and obtained the expected distances between randomly selected disease-symptom pairs. We denote the expected mean distance as\(\:\:{\mu\:}_{rand}\left(A,B\right)\), the SD as\(\:{\:\sigma\:}_{rand}\left(A,B\right)\), and define the proximity z-score:
The proximity z-score measures the difference between the proximity distance and the expectation, with z < 0 indicating closer than random, and z > 0 indicating farther than random. For the proximity distance d and the proximity z-score, lower metric values indicate closer distances between the two sets of nodes in the network. Since it’s based on random simulation, the proximity z-score is a stochastic measure, meaning the same repeated calculation can produce different proximity z-score.
Precision, recall and F1 score
Precision, Recall, and F1 score are commonly used metrics for evaluating classification tasks. In our work, these metrics are used to assess the performance of Named Entity Recognition (NER) models.
Precision (P) measures the proportion of correct positive predictions (TP) relative to all positive predictions made by the model (TP + FP). A high precision indicates that the model is effective at predicting positive cases without many false positives.
Recall (R) measures the proportion of correct positive predictions (true positives) relative to all actual positive instances in the data (true positives + false negatives). High recall indicates that the model is capable of identifying most of the positive instances.
F1 score is the harmonic mean of precision and recall, providing a single metric that balances both concerns. It is particularly useful when there is an uneven class distribution, such as in NER tasks.
These metrics provide insights into the effectiveness of our models in identifying relevant entities in unstructured text.
Results
Comparison of NER results from different models
We evaluate the NER task performance on the RoBERTa, LLaMA, and LLaMA fine-tuned with LoRA. We measured the NER task quality in terms of precision (P), recall (R), and the F1 score. Table 3 shows the evaluation result of different models in the NER task. It indicates that the LLaMA fine-tuned with LoRA has a clear advantage in the NER task, showing that this method can more effectively identify entities in unstructured text.
Construction of the psychosomatic disorder KG
The KG on psychosomatic disorders comprises 3285 entities and 9668 triplets. The categories and quantities of entities are as follows: patient 189, symptom 2693, risk factor 75, severity 136, disease 94, drug 98. To distinguish different patient entities, we use hospital-assigned identification numbers for naming (e.g., “J + 0101389333”). There are 7 types of relationships: < patient, suffer, disease>, < drug, treat, disease>, < symptom, diagnose, disease>, < symptom, primary diagnose, disease>, < symptom, differential diagnose, disease>, < risk factor, cause, disease>, < severity, diagnose, disease>. See 4.A.
The KG displays the relationships between diseases and related entities, visually characterizing the features of the psychosomatic disorder domain. In the graph, dots represent different entities, the color of the dots indicates the types of entities, and the lines between dots represent the relationships between entities. Taking “major depressive disorder(MDD)” as an example from the disease category. See Fig. 4.B. The subgraph highlights key entities related to the disease type ‘MDD’ and their relationships, such as the < symptom: bad mood, diagnose, disease: MDD>, < drug: sertraline, treat, disease: MDD>, < risk factor: high pressure, cause, disease: MDD>. See Table 4. Figure 4C and Fig. 4D show examples of the structural network formed by the KG of symptom-disease pairs and drug-disease pairs, with subsequent graph structure analysis based on the structural network formed by the nodes of the KG.
The KG is formalized as a directed graph \(\:G=(E,R,T)\) in the RDF format70, where: \(\:E\) denotes the entity set (nodes), which includes six types: patients, symptoms, risk factors, severity, diseases, and drugs. \(\:R\) represents the set of directed relationships (edges), instantiated as RDF triples of the form \(\:⟨\text{s}\text{u}\text{b}\text{j}\text{e}\text{c}\text{t},\text{p}\text{r}\text{e}\text{d}\text{i}\text{c}\text{a}\text{t}\text{e},\text{o}\text{b}\text{j}\text{e}\text{c}\text{t}⟩\). \(\:T\) is the set of triplets. This structure adheres to the RDF framework, ensuring a systematic and interoperable representation of psychosomatic disorder knowledge.
Construction of KG. (A) Illustration of the meta-graph (schema) of the KG, defining the overall structure and relationships between different entity types. Rectangular nodes represent six distinct entity categories, while directed edges (arrows) indicate possible relationships between entities, with edge labels specifying the relationship types. For instance, a directed edge from the ‘patient’ entity to the ‘disease’ entity with the label “suffers” represents the fact triple (patient, suffers, disease) in the KG. The schema comprises seven unique relationship types that govern how different entities can be connected in the knowledge base. (B) Display of some entities and relationships in the “MDD” disease category. Dots represent entities, different entity types are indicated by different colors, and the names of the entities are written on the dots. Arrows represent relationships. (C,D) Examples of partial KG structural network. Figure (C) shows a partial network of nodes related to symptom-disease pairs, and Figure (D) shows a partial network of nodes related to drug-disease pairs. The complex relationships within the KG form a vast network structure of psychosomatic disorder knowledge.
Cypher-based triple validation
This section validates the integrity of critical symptom–disease diagnostic relationships and drug–disease therapeutic relationships within the knowledge graph using Cypher queries (Neo4j’s graph query language). By directly matching node-relationship patterns, we ensure semantic associations align with the intended design.
Symptom–disease triple verification
To confirm diagnostic associations between symptoms and diseases, the following Cypher query matches nodes connected via the diagnose relationship:
MATCH (s: Symptom)-[: diagnose]-> (d: Disease) RETURN s.name AS Symptom, d.name AS Disease |
The query returns symptom–disease pairs linked by diagnose relationship. As shown in Table 5, results include expected associations (e.g., “Loss of Interest diagnose Schizophrenia”, “Dirty Speech diagnose Depressive Episode”), confirming these relationships are correctly encoded.
Drug–disease triple verification
Similarly, therapeutic associations between drugs and diseases are verified via the treat relationship:
MATCH (dr: Drug)-[: treat]-> (d: Disease) RETURN dr.name AS Drug, d.name AS Disease |
Results (Table 6) demonstrate drug–disease therapeutic pairs (e.g., “Lamotrigine treat Bipolar Disorder”, “Seroquel treat Schizophrenia”), validating the completeness of these triples.
Nodes representing symptom clusters for diagnosing diseases form modules within the network structure
We rely on our psychosomatic disorder KG to extract all symptom nodes and their diagnostically related diseases (referred to as “diseases” in this section). We focused on 54 diseases, with a total of 107 types of symptom cluster nodes. We found that for 46 of these 54 diseases, the connectivity components formed by their related symptom clusters were significantly larger than random expectation (\(\:{z}_{LCC}\) > 1.5). See Fig. 5.A and Fig. 5.E. Where y-axis represents the number of diseases corresponding to specific LCC z-score shown on the x-axis. This indicates that symptoms related to diseases cluster into a local module. Additionally, we analyzed the network separation metric between diseases (Sab), where disease pairs overlapping between modules have Sab < 0, and those topologically separated between modules have Sab > 0. We found the average network separation metric Sab between diseases and their related symptom modules to be 0.99. See Fig. 5.B. Numerous studies suggest that the random expected value of Sab = 0 serves as a threshold for assessing network separation[23; 44; 47]. The value greater than 0, indicating that symptom modules related to different diseases are distantly separated from each other. We systematically evaluated how Sab values correlate with symptom sharing between diseases. At Sab = 1.1, the number of shared symptoms decreased by 45% compared to Sab = 0, with diminishing reductions at higher Sab values. This nonlinear relationship suggests Sab = 1.1 represents a critical threshold for substantial module separation. While our observed Sab = 0.99 falls just below this threshold, its proximity to 1.1—combined with the sharp symptom-sharing decline in the Sab > 0 regime—strongly indicates that disease-associated symptom modules are functionally distinct and topologically distant in the network.
We also explored whether the network distances between symptom modules corresponding to these diseases could reveal clinical relationships between the diseases. For this purpose, we calculated the network distances (Dab) between diseases, centered around diseases, with symptom clusters forming modules. We used 5027 symptom-disease pairs to calculate the Co-symptoms count between disease pairs. We used 5027 symptom-disease pairs to calculate the Co-symptoms count between disease pairs. Here, ‘Co-symptoms’ refers to symptoms shared by two different diseases, such as insomnia, fatigue, and difficulty concentrating being common to both compulsive disorder and depressive episode. We found that the Co-symptoms count between two diseases was negatively correlated with the network distance Dab of their symptom modules (Pearson’s correlation = -0.39, P = 2.26 × 10–95), indicating that a closer network distance between diseases can predict their clinical manifestations to be more similar. See Fig. 5.C and Fig. 5.E. We also studied whether the network distance between diseases could predict their similarity in psychiatry. For this, we defined semantic similarity of symptoms48. We found that the overall semantic similarity of disease pairs negatively correlates with their average network distance Dab (Pearson’s correlation = -0.80, P < 1 × 10–100). See Fig. 5.D. In summary, we found that two diseases with closer network distances share more symptoms and have stronger similarities. The visualization of the symptom modules of diseases is shown in Fig. 5.F.
For example, the network distance Dab=1.25 between “Mood disorders” and “recurrent depressive disorder” is substantially lower than the average network distance < Dab > = 2.04 for diseases, with 99 Co-symptoms count, greatly surpassing the average Co-symptoms count across diseases. “Mood disorders” and “recurrent depressive disorder” share many symptoms, such as “irritability” and “sleep disturbances”. Other pairs of diseases with high similarity (highlighted in red in Fig. 5.C) include “compulsive disorder” and “depressive episode” (Dab=1.67, Co-symptoms count = 85), among others. Conversely, pairs of diseases with a higher network distance exhibit less Co-symptoms count and are not considered similar within the field of psychiatry, such as “acute stress psychosis” and “Tourette’s symptoms” (highlighted in green in Fig. 5.C), which have a larger Dab=3.73 and a lower comorbidity count of 3.
Symptom module centered on diagnosed diseases. (A) Distribution of the LCC z-score of the largest connected component formed by the symptoms of 54 diseases. Symptoms of 46 out of the 54 diseases form significantly clustered local modules (\(\:{z}_{LCC}\) > 1.6). The red dashed line represents \(\:{z}_{LCC}\) = 1.6. (B) Distribution of network separation (Sab) for symptom groups possessed by all disease pairs, with an average network separation < Sab > > 0, indicating that different diseases form symptom modules that are distant from each other. (C) The network distance (Dab) of interaction between disease pairs and the clinical similarity of diseases (Co-symptoms count) are negatively correlated, with Pearson’s correlation − 0.39. Each dot represents a disease pair. Examples of similar diseases, such as ”Mood disorders” and “recurrent depressive disorder” (Dab=1.25, Co-symptoms count=99), “compulsive disorder” and “depressive episode” (Dab=1.67, Co-symptoms count=85) are highlighted in red. We also highlight in green an example with a farther network distance and fewer shared symptoms, namely “acute stress disorder” and “Tourette’s symptoms” (Dab=3.73, Co-symptoms count=3). (D) The interaction network distance (Dab) between disease pairs is negatively correlated with the semantic similarity of symptoms. (E) Examples of disease modules and the network distance between disease pairs. Taking “Mood disorders”, “recurrent depressive disorder”, and “Tourette’s symptoms” as examples. (F) Visual representation of the symptom module for the disease “Mood disorders”.
Nodes of disease groups diagnosed through symptoms cluster into modules in the network structure
Following the previous section, we again extracted all nodes of symptoms and their diagnosed diseases (referred to as “diseases” in this section). However, we will focus on symptoms as central nodes to observe the local modules formed by disease group nodes. We focused on 105 symptoms, among which there were 54 types of disease group nodes. We found that the connectivity components formed by their related disease groups were significantly larger than random expectation (\(\:{z}_{LCC}\) > 1.03). See Fig. 6.A, indicating that diseases related to symptoms can also form local modules, meaning there is a many-to-many correspondence between diseases and symptoms. Furthermore, we also found that the < Sab> between the symptom pairs is 0.09, a value close to 0, indicating a high degree of similarity between some symptoms of disease, with some symptom clusters able to reflect a single disease simultaneously. See Fig. 6.B.
We also explored whether the network distances between disease modules corresponding to these symptoms could reveal the co-occurrence relationships between symptoms. For this purpose, we calculated the Dab between symptoms, centered on the symptoms, with disease groups forming modules. We calculated the number of concurrent diseases between symptom pairs. We found a negative correlation between Co-diseases count for two symptoms and their disease modules’ network distance Dab (Pearson’s correlation = -0.63, P < 1 × 10–100). Here, ‘Co-diseases’ refers to diseases that exhibit both of two symptoms. For example, generalized anxiety disorder and major depressive disorder are considered “Co-diseases” for the symptoms sleep disturbances and fatigue, because both diseases commonly exhibit these symptoms. See Fig. 6.C. Subsequently, to further validate this co-occurrence relationship, we calculated the relative risk (RR) between each pair of symptoms. RR is a standard measure of the strength of association (in this study, the simultaneous occurrence of two symptoms). We then found a negative correlation between the RR of symptom pairs and their network distance Dab (Pearson’s correlation = -0.29, P = 2.1 × 10–57). See Fig. 6.D. This indicates that a closer network distance between symptoms suggests they are more likely to occur together. For example, “decreased interest” and “Emotional discomfort” (Dab=0.36, RR = 10.7), “decreased interest” and “Vomit” (Dab=0.72, RR = 9.7) (highlighted in red in Fig. 6.D). Conversely, symptom pairs with longer network distances have lower RR and are less likely to co-occur, such as “hypochondriacal delusions” and “Emotional discomfort” (Dab=1.83, RR = 1.9) (highlighted in green in Fig. 6.D). These results validate our hypothesis that the network distances of disease modules associated with symptoms can reflect the co-occurrence relationships between symptoms. The visualization of the disease modules associated with symptoms is shown in Fig. 6.E and Fig. 6.F.
Symptom-centered diagnostic disease modules. (A) Distribution of LCC z-score for the largest connected components formed by related diseases of 105 symptoms. (B) Distribution of network separation (Sab) for all symptom pairs, with an average network separation < Sab> very close to 0, indicating a high degree of similarity between some symptoms of psychosomatic disorders. (C) The network distance (Dab) between symptom pairs and the co-occurrence (Co-diseases count) between symptom pairs are negatively correlated (Pearson’s correlation = -0.63). (D) There is a negative correlation between the relative risk (RR) of symptom pairs and the network distance Dab (Pearson’s correlation = -0.29), confirming that shorter network distances between symptoms can predict their co-occurrence. (E,F) Visualization of the disease modules associated with the symptoms “Disorientation” and “depression”.
Nodes representing groups of drug for diagnosed diseases form modules within the network structure
We rely on our psychosomatic disorder KG to extract all drug nodes and their diseases with a therapeutic relationship (referred to as “diseases” in this section). We focused on 37 diseases, among which there were 98 types of drug group nodes. We found that for 35 of these 37 diseases, the connectivity components formed by their related drug clusters were significantly larger than random expectation (\(\:{z}_{LCC}\) > 2.9). See Fig. 7.A. This indicates that drug related to disease treatment cluster into a local module. Furthermore, we also found that the < Sab> between diseases and drug modules treating diseases was 1.15, a value greater than 0, indicating that drug modules related to different diseases are distantly separated from each other. See Fig. 7.B.
We also explored whether the network distances between drug modules corresponding to these diseases could reveal the treatment principles between diseases. For this purpose, we calculated the Dab between diseases, centered on the diseases, with drug groups forming modules. We used 279 drug-disease pairs to calculate the number of drug shared between pairs of diseases. We found a negative correlation between the Co-drug count for two diseases and their drug modules’ network distance Dab (Pearson’s correlation = -0.34, P = 2.08 × 10–18), indicating that a closer network distance between diseases can predict their treatment plans to be more similar. Here, ‘Co-drug’ refers to drugs that are used to treat both of two diseases. For example, lithium and quetiapine are ‘Co-drug’ for bipolar disorder and depressive episode. See Fig. 7.C. We also studied whether the network distances between diseases could predict their similarity in psychological mechanisms. For this, we defined the semantic similarity of drug23.
We found that the overall semantic similarity of disease pairs negatively correlates with their average network distance Dab (Pearson’s correlation = -0.82, P < 1 × 10–100). See Fig. 7.D. In summary, we found that two diseases with closer network distances have more similar treatment plans and psychological mechanisms. The visualization of the drug modules associated with diseases is shown in Fig. 7.E and Fig. 7.F.
For example, the network distance Dab between “bipolar disorder” and “depressive episode” is 0.89, much lower than the average network distance of diseases < Dab > = 2.14, with 19 shared drug, far above the average number of shared drug for diseases, which is 4. “bipolar disorder” and “depressive episode” share many drug, such as “seroquel” and “olanzapine”. Other disease pairs with high similarity (highlighted in red in Fig. 7.C) include “schizophrenia” and “depressive episode” (Dab=1.19, Co-drug = 15), among others. Conversely, disease pairs with higher network distances have fewer shared drug and are not considered similar in psychological mechanisms, such as “recurrent depressive disorder” and “Mood disorders” (highlighted in green in Fig. 7.C), which have a larger Dab=2.04 and a smaller Co-drug count of 2.
Drug modules centered on treating diseases. (A) Distribution of LCC z-score for the largest connected component of treatment-related drug for 37 diseases. drug for 35 out of these 37 diseases form significantly clustered local modules (\(\:{z}_{LCC}\) > 2.9). The red dashed line represents \(\:{z}_{LCC}\) = 2.9. (B) Distribution of network separation (Sab) for drug groups corresponding to all disease pairs, with an average network separation < Sab > > 0, indicating that different diseases form modules that are distant from each other. (C) The network distance (Dab) between disease pairs and the similarity of treatment principles (Co-drug count) are negatively correlated, with a Pearson’s correlation of -0.34. Each point represents a disease pair. Examples of similar diseases highlighted in red, such as “bipolar disorder” and “depressive episode” (Dab=0.89, Co-drug count=19), “schizophrenia” and “depressive episode” (Dab=1.19, Co-drug count=15). We also highlighted in green an example with a farther network distance and fewer shared symptoms, namely recurrent “recurrent depressive disorder” and “Mood disorders” (Dab=2.04, Co-drug count=2). (D) The network distance between disease pairs and the semantic similarity of drug are negatively correlated, indicating that diseases with closer network distances have similar psychological mechanisms. (E,F) Visualization of the drug modules associated with “schizophrenia” and “depressive episode”.
The modules formed by symptom cluster nodes for differential diagnosis of diseases and the degree of association between diseases and symptoms
Besides diagnosed diseases, we rely on our psychosomatic disorder KG to extract all symptoms and their disease nodes with a differential diagnosis relationship. Differential diagnosis means excluding the possibility of a disease based on certain symptoms. We focused on 53 diseases with differential diagnosis, among which there were 243 types of symptom cluster nodes. We found that for 36 of these 53 diseases with differential diagnosis, the connectivity components formed by their related symptom clusters were significantly larger than random expectation (\(\:{z}_{LCC}\) > 1.1). See Fig. 8.A. This indicates that symptoms of diseases with differential diagnosis cluster into a local module. Furthermore, we also found the < Sab> between symptom modules corresponding to diseases with differential diagnosis to be 2.26, a value greater than 0, indicating that symptom modules corresponding to different differential diagnosis diseases are far apart from each other. See Fig. 8.B.
In our psychosomatic disorder KG, diagnostic relationships are divided into diagnosis, primary diagnosis and differential diagnosis. These relationships are used to distinguish between different stages of disease determination. The symptoms connected by diagnosis are mostly patient-reported and colloquial, whereas those connected by primary diagnosis or differential diagnosis are summarized by psychiatrists and are terms standardized in the field of psychiatric medicine. To explore the degree of association between symptoms and their diseases for both types of relationships, we observed the network structure of symptoms and diseases under primary diagnosis relationships and diagnosis relationships, respectively. To quantify the degree of association, we used two metrics: (1) proximity distance d, the average shortest distance between nodes within symptom and disease modules. (2) the proximity z-score, measuring the difference between proximity distance d and random expectation. For these metrics, lower values indicate a higher degree of association between diseases and symptoms. For the proximity z-score, z-score < 0 indicates closer than random, z-score > 0 indicates further than random. We plotted box plots for both types of relationships, comparing symptom-disease pairs in primary diagnosis relationships (orange bars) and diagnosis relationships (blue bars). See Fig. 8.C. We found that under both metrics, the orange bars are consistently lower than the blue bars, indicating that symptom-disease pairs in primary diagnosis relationships have a stronger degree of association and are of higher reference value. Specific examples are shown in Fig. 8.D.
Analysis of primary diagnosis, diagnosis, and differential diagnosis. (A) Distribution of LCC z-score for the largest connected components formed by symptoms corresponding to 53 differential diagnosis diseases. Of these, symptoms of 36 diseases form significantly clustered local modules (\(\:{z}_{LCC}>\)1.1). The red dashed line represents \(\:{z}_{LCC}\:\)= 1.1. (B) Distribution of network separation (Sab) for symptom clusters corresponding to all differential diagnosis disease pairs, with an average network separation < Sab> greater than 0, indicating that different differential diagnosis diseases form modules distant from each other. (C) Disease-symptom pairs are divided into primary diagnosis relationships and diagnosis relationships. Symptom-disease pairs in the primary diagnosis relationship (orange bars) show a shorter network distance than those in the diagnosis relationship (blue bars), indicating a stronger association in primary diagnosis relationships. (D) For example, symptoms such as “decreased interest” and “low mood” primarily diagnose “depressive episode” (with a very low network proximity z-score). They are included in the main clinical manifestations of “depressive episode” in real life. However, the symptom of “weird behavior”, having a higher network proximity z-score, indicates that it is not a main symptom of “depressive episode” but a main symptom of “schizophrenia”.
Discussion
In this study, we defined entity and relation types based on clinical terminology widely used in the field of psychiatry. Through communication and discussion, we developed a comprehensive set of standardized concepts that can precisely represent various clinical entities and their relationships. To accommodate the demands of real- some mod world psychiatric applications, we also sought opinions from domain experts and made ifications and adjustments to the entity and relation types. Ultimately, we obtained 6 types of entities and 7 types of relationships.
The extracted knowledge on psychosomatic disorders inherently contains some redundancy due to repeated information. To address this, we applied entity alignment methods to eliminate duplicate entities and ensure their standardization and unification. As a result, we constructed a high-quality psychosomatic disorder KG. Compared to existing mental health KGs and ontologies, our KG demonstrates two distinctive advantages: firstly, it innovatively incorporates clinical practice data, including doctor-patient dialogue records, which significantly enhances the practical utility and clinical relevance of the KG; secondly, the schema design strictly adheres to the standardized diagnostic workflow in psychiatric practice, ensuring professional and systematic knowledge representation.
After constructing the KG, we analyzed its structure, revealing the connections between diseases, symptoms, and drug in the field of psychosomatic disorders.
We found that in 54 diseases, the symptoms corresponding to 46 diseases, diseases related to 105 symptoms, and the drug required for 35 of 37 diseases all formed significantly clustered local modules. This phenomenon indicates that there is a close interrelation between symptoms and pharmacological treatments in the diagnosis and treatment of psychosomatic disorders, suggesting that the network structure of psychosomatic disorders might have a decisive influence on symptom and drug selection. The discovery of this correlation not only unveils the intrinsic link between the symptoms and treatment of psychosomatic disorders but also provides a theoretical basis for using bioinformatics methods to predict potential treatment strategies for mental diseases. For example, by analyzing the symptom cluster modules, we can identify which symptoms are key manifestations of a particular psychosomatic disorder and thereby infer the most effective medication combinations.
The symptom modules of diseases reflect their clinical similarity. For instance, the diseases “Mood disorders” and “recurrent depressive disorder” have close network distances Dab in their symptom modules and a high number of shared symptoms, hence a high clinical similarity between them. Conversely, “acute stress disorder” and “Tourette syndrome” have a high network distance Dab and very few shared symptoms, therefore are not considered similar by the field of psychiatry. By analyzing the clinical similarities between diseases, it aids doctors in more accurately distinguishing and identifying similar diseases during the early diagnostic stages, especially in cases of overlapping or unclear symptoms, thus improving diagnostic accuracy. Moreover, it provides a symptom-based method of disease classification that may complement traditional etiological classification methods, offering data support for new classification standards in psychiatric disorders.
The disease modules of symptoms reflect the co-occurrence relationships among symptoms. For example, the symptoms “hallucinations” and “anhedonia” have close network distance Dab in their disease modules and a high number of concurrent diseases, indicating a high degree of co-occurrence between the symptoms. Conversely, “elevated mood” and “lack of insight” have a high network distance Dab and very few concurrent diseases, thus exhibiting a low degree of co-occurrence. Subsequently, we validated this conclusion using the relative risk (RR) of symptoms, showing that the co-occurrence of symptoms is negatively correlated with their network distance between modules. By analyzing the co-occurrence relationships of symptoms between different diseases, it helps to understand why certain symptoms frequently appear together, potentially indicating a common pathophysiological basis. Furthermore, in the management of patients with multiple psychosomatic disorders, it enables a better understanding and prediction of symptom progression, optimizing treatment plans.
The drug modules of diseases reflect the similarity in treatment approaches among diseases. For example, the diseases “bipolar disorder” and “depressive episode” have close network distance Dab in their drug modules and share many drug, indicating a high similarity in their treatment approaches. Conversely, “recurrent depressive disorder” and “Mood disorders” have a high network distance Dab in their drug modules and share very few drug, thus their treatment plans are generally not comparable. By analyzing the similarity in treatment plans between different diseases, it is possible to discover potential new medication guidelines or alternative treatment methods. Furthermore, this can assist medical researchers in extending the known effects of drug to new disease areas, promoting drug repurposing research.
For diagnosed disease-symptom pairs and disease-drug pairs, we have also defined semantic similarity to analyze the degree of similarity between diseases. This allows for quantifying the associations between diseases and their symptoms, as well as between diseases and drugs. By using this method, doctors can more accurately identify and differentiate various psychosomatic disorders, which is crucial for improving diagnostic accuracy, particularly among conditions with overlapping symptoms, such as anxiety or mood disorders. Moreover, it can uncover potential similarities in drug responses across different diseases, providing personalized treatment options, which is especially valuable for psychosomatic disorders that often require trials of multiple medications to determine the most effective treatment.
For primary and diagnostic disease-symptom pairs, our analysis using proximity distance d and proximity z-score found that primary diagnosis relationships have stronger associations, confirming that network proximity can effectively predict disease-symptom pairs with stronger correlations. For example, the symptoms “reduced interest” and “lack of will” in the primary diagnosed disease “depressive episode” have a low proximity distance d and proximity z-score, and these are major clinical manifestations of “depressive episode” in real life. Meanwhile, the symptom “bizarre behavior” and the disease “depressive episode” have a high proximity distance d and z- score, but it has a low proximity distance d and z-score with “schizophrenia”, indicating it is not a main symptom of depressive episode but of schizophrenia. This analysis helps medical professionals more precisely identify the core symptoms associated with specific psychosomatic disorders. This is crucial for the diagnosis of psychosomatic disorders, especially in the early stages, as accurate symptom identification can significantly improve the success rate of treatment.
Our work lays the foundation for further development of smart medical information systems in psychiatry. Medical information intelligent systems often lack domain-specific knowledge bases that serve as reliable knowledge sources. In contrast to general encyclopedic knowledge, our knowledge is derived from summarized medical records provided by psychiatrists, enabling the creation of a highly specialized KG database. The KG we constructed helps guide psychiatric staff to engage more effectively with patients, analyze the proximity of knowledge modules in the dialogues formed, and enhance the quality of healthcare. Furthermore, by analyzing the knowledge network structure of psychosomatic disorder, we have delved into the connections between these diseases, symptoms, and drug, providing a technical roadmap and foundational data for developing applications that save psychiatrists’ time, enhance treatment efficacy and compliance, and improve patient quality of life. The current scale of the KG is relatively small, and the original dataset lacks sufficient diagnostic-related entities—such as severity levels, susceptibility factors, and past medical histories—which makes it challenging to accurately recommend personalized treatment plans based on entity relationships. Moving forward, we plan to expand the dataset to enrich the diversity of the KG and uncover a broader range of connections.
Conclusion
This research collected 262 cases from the Psychiatric Department of Zhongda Hospital in Nanjing, China and used BERT and LLM for entity extraction to build a psychosomatic disorder KG containing 3285 entities and 9668 relationships. Subsequently, graph theory was applied to analyze the structure of the constructed KG. The study found that symptoms of diseases, diseases related to these symptoms, and the drug required for these diseases form local clustering modules within the graph structure. Semantic similarity analysis was also defined to measure the degree of similarity between diseases. Through this definition, the associations between diseases and symptoms, and diseases and drug can be quantified. The research findings are as follows:
-
(1)
LLaMa, fine-tuned with LORA, improved in its ability to extract entities, achieving an accuracy close to that of the BERT model, up to 93%.
-
(2)
The average network separation Sab measure between disease and related symptom modules is 0.99, greater than 0, indicating that symptom modules associated with different diseases are far apart from each other. The Co-symptoms count between two diseases and the network distance Dab of their symptom modules are negatively correlated. The overall semantic similarity between disease pairs and their average network distance Dab is also negatively correlated, suggesting that closer network distances between diseases can predict more similar clinical presentations.
-
(3)
The average network separation Sab measure between symptoms and their related disease modules is 0.09, close to 0, indicating that a high degree of similarity among some symptoms can reflect the same disease. The RR between pairs of symptoms is negatively correlated with their network distance Dab, suggesting that symptoms are more likely to co-occur with closer network distances.
-
(4)
The average network separation Sab measure between diseases and their drug treatment modules is 1.15, greater than 0, indicating that drug modules associated with different diseases are significantly distant from each other. The Co-drug count between diseases and the network distance Dab of their drug modules are negatively correlated. The overall semantic similarity between disease pairs and their average network distance Dab is also negatively correlated, suggesting that diseases with closer network distances have more similar treatment regimens and psychological mechanisms.
-
(5)
By comparing the proximity d and proximity z-score metrics, it is shown that symptom-disease pairs in primary diagnostic relationships have a stronger association and higher referential value than those in diagnostic relationships.
The research results help medical professionals more accurately identify the core symptoms of diseases, not only revealing the interrelationships among psychosomatic disorders but also potentially providing a theoretical basis for developing new treatment methods and improving existing treatment strategies.
Data availability
The data that support the findings of this study are available at https://github.com/zzhzhouzihan016/Psychosomatic_Disease_Knowledge_Graph.
References
Organization, W. H. World Mental Health Report: Transforming Mental Health for all (World Health Organization, 2022).
Huang, Y. et al. Prevalence of mental disorders in china: A cross-sectional epidemiological study. Lancet Psychiatry. 6, 211–224 (2019).
Jiang, G. et al. The status quo and characteristics of Chinese mental health literacy. Acta Physiol. Sin. 53, 182 (2021).
Huang, Y. et al. The China mental health survey (cmhs): I. Background, aims and measures. Soc. Psychiatry Psychiatr. Epidemiol. 51, 1559–1569 (2016).
Huang, Y. Q. Epidemiological study on mental disorder in China. Zhonghua Liu Xing Bing Xue Za Zhi. 33, 15–16 (2012).
Hou, M., Wei, R., Lu, L., Lan, X. & Cai, H. Research review of knowledge graph and its application in medical domain. J. Comput. Res. Dev. 55, 2587–2599 (2018).
Li, L. et al. Real-world data medical knowledge graph: construction and applications. Artif. Intell. Med. 103, 101817 (2020).
Wang, Q., Mao, Z., Wang, B. & Guo, L. Knowledge graph embedding: A survey of approaches and applications. IEEE Trans. Knowl. Data Eng. 29, 2724–2743 (2017).
Zhao, Z., Han, S. K. & So, I. M. Architecture of knowledge graph construction techniques. Int. J. Pure Appl. Math. 118, 1869–1883 (2018).
Martinez-Rodriguez, J. L., Lopez-Arevalo, I. & Rios-Alvarado, A. B. Openie-based approach for knowledge graph construction from text. Expert Syst. Appl. 113, 339–355 (2018).
Jiang, G., Liu, H., Solbrig, H. R. & Chute, C. G. Mining severe drug-drug interaction adverse events using semantic web technologies: A case study. BioData Min. 8, 1–12 (2015).
Freidel, S. & Schwarz, E. Knowledge graphs in psychiatric research: potential applications and future perspectives. Acta Psychiatry. Scand. 151, 180–191 (2025).
Devlin, J., Chang, M. W., Lee, K., Toutanova, K. & Bert Pre-training of deep bidirectional transformers for language understanding. In Proceedings of the 2019 C0onference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies. 4171-4186 (2019).
Perlis, R. et al. Using electronic medical records to enable large-scale studies in psychiatry: treatment resistant depression as a model. Psychol. Med. 42, 41–50 (2012).
Castro, V. M. et al. Validation of electronic health record phenotyping of bipolar disorder cases and controls. Am. J. Psychiatry. 172, 363–372 (2015).
Hogan, A. et al. Knowledge graphs. ACM Comput. Surv. (Csur). 54, 1–37 (2021).
Cui, Y., Che, W., Liu, T., Qin, B. & Yang, Z. Pre-training with whole word masking for Chinese Bert. IEEE/ACM Trans. Audio Speech Lang. Process. 29, 3504–3514 (2021).
Naveed, H. et al. A comprehensive overview of large Language models. ArXiv (2023).
Vaswani, A. et al. Attention is all you need. Adv. Neural. Inf. Process. Syst. 30 (2017).
Wang, S. et al. Gpt-ner: named entity recognition via large Language models. ArXiv (2023).
Li, J., Sun, A., Han, J. & Li, C. A survey on deep learning for named entity recognition. IEEE Trans. Knowl. Data Eng. 34, 50–70 (2020).
Collobert, R. et al. Natural Language processing (almost) from scratch. J. Mach. Learn. Res. 12, 2493–2537 (2011).
Gan, X. et al. Network medicine framework reveals generic herb-symptom effectiveness of traditional Chinese medicine. Sci. Adv. 9, eadh0215 (2023).
Bodenreider, O. The unified medical Language system (umls): integrating biomedical terminology. Nucleic Acids Res. 32, D267–D270 (2004).
Névéol, A., Doğan, R. I. & Lu, Z. Semi-automatic semantic annotation of pubmed queries: A study on quality, efficiency, satisfaction. J. Biomed. Inform. 44, 310–318 (2011).
Bikel, D. M., Schwartz, R. & Weischedel, R. M. An algorithm that learns what’s in a name. Mach. Learn. 34, 211–231 (1999).
Lafferty, J., McCallum, A. & Pereira, F. Conditional random fields: Probabilistic models for segmenting and labeling sequence data. In International Conference on Machine Learning (2001).
Huang, Z., Xu, W. & Yu, K. Bidirectional lstm-crf models for sequence tagging. ArXiv (2015).
Lee, J. et al. Biobert: A pre-trained biomedical Language representation model for biomedical text mining. Bioinformatics 36, 1234–1240 (2020).
Beltagy, I., Lo, K., Cohan, A. & Scibert A pretrained language model for scientific text. arXiv (2019).
Zhang, B. & Soh, H. Extract, define, canonicalize: an llm-based framework for knowledge graph construction. ArXiv (2024).
Karpukhin, V. et al. Dense Passage Retrieval for Open-Domain Question Answering. In Proceedings of the 2020 Conference on Empirical Methods in Natural Language Processing 6769-6781 (2020).
Izacard, G. & Grave, E. Leveraging passage retrieval with generative models for open domain question answering. arXiv (2020).
Donnelly, K. Snomed-ct: The advanced terminology and coding system for ehealth. Stud. Health Technol. Inform. 121, 279-279 (2006).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Schriml, L. M. et al. Human disease ontology 2018 update: classification, content and workflow expansion. Nucleic Acids Res. 47, D955–D962 (2019).
Smith, B. et al. The Obo foundry: coordinated evolution of ontologies to support biomedical data integration. Nat. Biotechnol. 25, 1251–1255 (2007).
Catlett, N. L. et al. Reverse causal reasoning: applying qualitative causal knowledge to the interpretation of high-throughput data. BMC Bioinform. 14, 1–14 (2013).
Kilicoglu, H. et al. A pubmed-scale repository of biomedical semantic predications. Bioinformatics 28, 3158–3160 (2012).
Walsh, B., Mohamed, S. K., Nováček, V. & Biokg A knowledge graph for relational learning on biological data. In Proceedings of the 29th ACM International Conference on Information & Knowledge Management. 3173-3180 (2020).
Zhang, R. et al. Drug repurposing for covid-19 via knowledge graph completion. J. Biomed. Inform. 115, 103696 (2021).
Himmelstein, D. S. et al. Systematic integration of biomedical knowledge prioritizes drugs for repurposing. Elife 6, e26726 (2017).
Chandak, P., Huang, K. & Zitnik, M. Building a knowledge graph to enable precision medicine. Sci. Data. 10, 67 (2023).
Menche, J. et al. Uncovering disease-disease relationships through the incomplete interactome. Science 347, 1257601 (2015).
Barabási, A. L., Gulbahce, N. & Loscalzo, J. Network medicine: A network-based approach to human disease. Nat. Rev. Genet. 12, 56–68 (2011).
Guney, E., Menche, J., Vidal, M. & Barábasi, A. L. Network-based in Silico drug efficacy screening. Nat. Commun. 7, 10331 (2016).
Cheng, F., Kovács, I. A. & Barabási, A. L. Network-based prediction of drug combinations. Nat. Commun. 10, 1197 (2019).
Loukachevitch, N. et al. Nerel-bio: A dataset of biomedical abstracts annotated with nested named entities. Bioinformatics 39, btad161 (2023).
Artemova, E. et al. Runne-2022 shared task: recognizing nested named entities. ArXiv (2022).
Han, K. et al. Transformer in transformer. Adv. Neural Inf. Process. Syst. 34, 15908–15919 (2021).
Gillioz, A., Casas, J. & Mugellini, E. & Abou Khaled, O. Overview of the Transformer-based Models for NLP Tasks. In 2020 15th Conference on Computer Science and Information Systems (FedCSIS). 179-183 (2020).
Braşoveanu, A. M. & Andonie, R. Visualizing transformers for nlp: a brief survey. In 2020 24th International Conference Information Visualisation (IV) 270-279 (2020).
Tetko, I. V., Karpov, P., Van Deursen, R. & Godin, G. State-of-the-art augmented Nlp transformer models for direct and single-step retrosynthesis. Nat. Commun. 11, 5575 (2020).
Hu, E. J. et al. Lora: Low-rank adaptation of large language models. In International Conference on Learning Representations (2022).
Chen, X. et al. Knowprompt: Knowledge-aware prompt-tuning with synergistic optimization for relation extractio. In Proceedings of the ACM Web Conference 2022 2778-2788 (2022).
Yao, Y. et al. Editing large language models: Problems, methods, and opportunities. arXiv (2023).
Zhu, Y. et al. Knowagent: Knowledge-augmented planning for llm-based agents. ArXiv (2024).
Qiao, S. et al. Making Language models better tool learners with execution feedback. ArXiv (2023).
Yao, Y. et al. Schema-aware reference as prompt improves data-efficient knowledge graph construction. In Proceedings of the 46th International ACM SIGIR Conference on Research and Development in Information Retrieval. 911-921 (2023).
Chen, X. et al. Decoupling knowledge from memorization: Retrieval-augmented prompt learning. Adv. Neural. Inf. Process. Syst. 35, 23908–23922 (2022).
Zhao, X., Zeng, W., Tang, J., Wang, W. & Suchanek, F. M. An experimental study of state-of-the-art entity alignment approaches. IEEE Trans. Knowl. Data Eng. 34, 2610–2625 (2020).
Trisedya, B. D., Qi, J. & Zhang, R. Entity alignment between knowledge graphs using attribute embeddings. Proc. AAAI Conf. Artif. Intell. 33, 297-304 (2019).
Zhang, Q. et al. Multi-view knowledge graph embedding for entity alignment. arXiv (2019).
Liu, Z., Shen, Y., LI, Y. & SU, W. Construction of a psychological medical knowledge graph based on mfe-bert and fnnattention. Comput. Syst. Appl. 32, 108–119 (2023).
Zhou, X., Menche, J., Barabási, A. L. & Sharma, A. Human symptoms–disease network. Nat. Commun. 5, 4212 (2014).
Yıldırım, M. A., Goh, K. I., Cusick, M. E., Barabási, A. L. & Vidal, M. Drug—target network. Nat. Biotechnol. 25, 1119–1126 (2007).
Lin, D. An information-theoretic definition of similarity. Int. Conf. Mach. Learn. 98, 296-304 (1998).
Yu, G. et al. Gosemsim: an r package for measuring semantic similarity among go terms and gene products. Bioinformatics 26, 976–978 (2010).
Wang, J. Z., Du, Z., Payattakool, R., Yu, P. S. & Chen C.-F. A new method to measure the semantic similarity of go terms. Bioinformatics 23, 1274–1281 (2007).
Cavalleri, E. et al. An ontology-based knowledge graph for representing interactions involving Rna molecules. Sci. Data. 11, 906 (2024).
Acknowledgements
The author is very grateful for the guidance provided by his mentor Yihui Zhu, the encouragement given by his relatives, and the contributions of Tantao Su and Ruoliu Xu, Yihan Du and Shijie Peng to this research.
Funding
This work was supported by the College Students’ Innovative Entrepreneurial Training Plan Program (Grant No. 202410300249Y, XJDC202410300374) and the Startup Foundation for Introducing Talent of Nanjing University of Information Science and Technology, China (Grant No.2022r095).
Author information
Authors and Affiliations
Contributions
Z.Z., Z.Z. and Y.Z. conceptualized the study and developed the methodology. Z.Z. (Ziyi Zeng) conducted the formal analysis. Z.Z. (Ziyi Zeng), W.J., Y.Z., J.M., Y.Y., M.Y. and Y.C. curated the data. Z.Z. (Zihan Zhou) and Z.Z. (Ziyi Zeng) wrote the original draft. Z.Z. (Zihan Zhou), Z.Z. (Ziyi Zeng), Y.Z., W.J., J.M., Y.Y., M.X. and S.Z. reviewed and edited the manuscript. Z.Z. (Zihan Zhou), Z.Z. (Ziyi Zeng) and Y.Z. created the visualizations. M.X. and S.Z. supervised the project. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Institutional review board statement
Ethics Committee Approval given by the Clinical Research Ethics Committee, Zhongda Hospital Affiliated to Southeast University, Nanjing in accordance with the principles of the Helsinki Declaration of 2013 (protocol code 2021ZDSYLL349-P02, 29 April 2022).
Informed consent
Written informed consent was waived by the IEC for Clinical Research of Zhongda Hospital, Affiliated to Southeast University (Approval No. 2021ZDSYLL349-P02, 29 April 2022). This study was conducted in accordance with the principles of the Helsinki Declaration (2013).
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhou, Z., Zeng, Z., Jiang, W. et al. Research on the proximity relationships of psychosomatic disease knowledge graph modules extracted by large language models. Sci Rep 15, 20653 (2025). https://doi.org/10.1038/s41598-025-05499-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-05499-8