Abstract
Inonotus hispidus, a fungal species morphologically closely related to the medicinal genus Sanghuang,has garned considerable attention due to its potential health-prompting benefits. This study presents the whole-genome sequence of I. hispidus ZA-14, a monokaryotic strain isolated from wild fruiting bodies growing in Morus alba (hereinafter referred to as MA). The 37.68 Mb genome was assembled from 5771 Mb of raw data, resulting in 51 contigs with a GC content of 48.48%. A total of 8924 protein-coding genes were predicted, with 223 non-coding RNAs identified. The genome analysis revealed a high density of genes associated with secondary metabolite biosynthesis, including those involved in terpenoid, polysaccharide, and flavonoid pathways—key contributors to its medicinal attributes. Comparative genomic analysis with other related fungal species provided insights into the evolutionary dynamics and speciation of I. hispidus. The study also identified a significant enrichment of glycoside hydrolase family genes, suggesting a critical role in nutrient absorption and utilization. These findings enhance our understanding of the genetic basis of I. hispidus’s medicinal value and its evolutionary relationships, providing a foundation for future breeding.
Similar content being viewed by others
Introduction
As documented in the ancient text Shen Nong Materia Medica, macrofungi have been utilised as traditional Chinese medicine by ancient Chinese people for millennia1,2. Notably, a number of renowned medicinal fungal species, including “Dongchong Xiaocao” (Ophiocordyceps sinensis), “Lingzhi” (Ganoderma lingzhi) and “Fuling” (Wolfiporia cocos), are gaining global recognition, with China playing a pivotal role in this trend. In recent years, “Sanghuang”, a valuable medicinal macrofungi in China,has also emerged as a focal point in medicinal, scientific, and industrial research3. Taxonomically classified within the kingdom Fungi, phylum Basidiomycota class Agaricomycetes, order Hymenochaetales, family Hymenochaetaceae, it comprises multiple species within the genera Inonotus, Fomitiporia, and Phellinus.
Inonotus hispidus, a species in the genus Inonotus, primarily infects hosts such as Morus alba, Fraxinus mandshurica, Ulmus macrocarpa, Ziziphus jujuba and Malus pumila. The strain I. hispidus ZA-14, which specifically parasitizes M. alba(white mulberry)—a tree widely distributed across China—has been studied for its ecological and medicinal relevance. Based on its morphological characteristics, habitat analysis, and herbal research, Chinese scholars have infered M. alba as authentic sources of the traditional Chinese medicine Sanghuang4,5. This fungus exhibits notable attributes: nutritional value comparable to that of other edible fungi, as well as immune-modulatory, antitumour, antioxidant, and cholesterol-lowering activities6,7,8. Additionally, it demonstrates antibacterial, anti-inflammatory, and other medicinal properties9.
I. hispidus is rich in a variety of bioactive compounds such as polysaccharides, polyphenols, steroids, terpenoids, pigments and amino acids10. The traditional use of Sanghuang in Chinese medicine, as documented in ancient pharmacopeias, underscores its therapeutic potential. Modern genomic studies have built upon this foundation by identifying specific bioactive compounds, such as triterpenoids and polysaccharides—whose medicinal properties were first recognized through traditional practices. Hispolon and hispidin, two natural polyphenols isolated from genus Inonotus and analogous to those in Sanghuang, exhibit anticancer, immunomodulatory, antioxidant, and antiviral activities. These properties were initially identified through ethnopharmacological studies and later validated through molecular analyses11,12,13,14,15. Different species within the genus Sanghuangporus contain protocatechuic acid and ergosterol, both of which have notable antibacterial and anticancer effects16. Among them, ergosterol—a signature steroidal compound in fungi—represents a key bioactive ingredient in Sanghuang, contributing to metabolic regulatory functions, hormone level modulation, and antitumour activity17.
With the development of DNA sequencing technologies, medicinal studies on macrofungi have expanded beyond the extraction of bioactive compounds and functional characterization to focus on genomic sequencing. For species within the genera Sanghuangporus and Inonotus, genome sequences of four distinct taxa—Sanghuangporus baumii, S. sanghuang, S. vaninii and I. hispidus (comprising two distinct genomes) —have been deposited in the National Centre for Biotechnology Information (NCBI, BioProject: PRJNA304358, PRJNA731629, PRJNA564179, PRJNA973857, PRJNA1060777). Analyses of these genomic datasets provide insights into the gene regulatory networks, biosynthetic pathways and pharmacological properties of these macrofungi, laying a solid foundation for their subsequent commercial production and enhancement of their medical value.
To advance the medicinal and industrial exploitation of I. hispidus ZA-14, we sequenced the genome of a monokaryotic strain following precise species identification. In addition to providing insights into medicinal application, comparative genomic analyses—particularly through cross-species comparisons with related fungal taxa, revealed the evolutionary and speciated dynamics of I. hispidus in Sanghuang.
Materials and methods
Strain culture and DNA isolation
The monokaryotic strain ZA-14 was isolated from 1-year-old wild fruiting bodies collected in Yulin, Shaanxi Province, China, in September 2019. For purification, ZA-14 mycelia were cultivated on sterile cellophane-covered potato dextrose agar (PDA) plate under dark conditions at 28 °C, with apical tip isolation used to ensure clonal purity. Genomic DNA was extracted from the harvested single spore mycelia using the cetyl trimethyl ammonium bromide (CTAB) method. High-quality genomic DNA was assessed for purity, concentration and integrity via NanoDrop™ 2000 spectrophotometer, Qubit® 2.0 Fluorometer (Thermo Scientific, Waltham, MA, USA) and 0.35% agarose gel electrophoresis.
Species identification
The internal transcribed spacer(ITS) region of strain ZA-14 was amplified from genomic DNA using primers ITS1 and ITS418. The PCR amplification protocal was as follows: initial denaturation at 94 °C for 2 min, followed by 36 cycles of denaturation at 94 °C for 30 s, annealing at 50 °C for 30 s and extension at 72 °C for 50 s, with a final extension at 72 °C for 10 min. Amplification products were analyzed by 1% agarose gel electrophoresis followed by sequencing. The generated ITS sequence was queried against the GenBank database (https://www.ncbi.nlm.nih.gov/genbank/) for homology searching, and species identification was performed via phylogenetic tree construction. The phylogenetic tree was inferred using the maximum likelihood (ML) method in MEGA6.0 software, with bootstrap values calculated from 1000 replicates.
Genome sequencing and assembly
The raw data generated by Nanopore PromethION sequencing are stored in binary fast5 format, which contains all raw sequencing signals, with a single read corresponding to a single fast5 file. The fast5 format data were converted to fastq format through base calling using the Albacore software in the MinKNOW19. Following adapter trimming, removal of low-quality reads, and filtering of short fragments (length < 2000 bp), the final dataset was obtained. Genome assembly was performed using NECAT software with default parameters, and Pilon software20 was subsequently used to correct assembly errors using second-generation sequencing data, yielding a highly accurate final genome.
Integrity of the fungal genome assembly was assessed using the alignment rate of the sequencing reads and BUSCO21, respectively. alignment rate refers to the proportion of clean reads that can be compared to the genome to the total number of clean reads, the alignment rate is related to the quality of the genome assembly and the quality of the reads sequencing, the more complete the genome assembly and the higher the quality of the sequenced reads, the higher the alignment rate. The Burrows-Wheeler Aligner (BWA) software22 was used for the alignment. The fungi_odb9 database in BUSCO contains 290 conserved core genes of the fungus. We used the BUSCO v2.0 software to assess the integrity of this fungal genome assembly.
Due to the relatively low conservatism of repetitive sequences among species, the prediction of repetitive sequences for a specific species requires the construction of a specific database of repetitive sequences. Therefore, with the help of four software (LTR_FINDER v1.05, MITE-Hunter, RepeatScout v1.0.5, and PILER-DF v2.4), we constructed a database of repetitive sequences of this fungal genome based on the principles of structural prediction and de novo prediction23,24,25,26. The database was classified by PASTEClassifier v1.0 and then merged with the database of Repbase (19.06) to form the final repetitive sequence database27,28. Next, RepeatMasker v4.0.629 was used to predict the repetitive sequences of this fungus based on the constructed repetitive sequence database. Finally, the characteristics of the de novo assembly genomic features of ZA-14 were visualised using the Circos software(v0.69-8)30.
The GFF format is utilized for the description of repetitive sequences, incorporating information pertaining to the origin of repetitive sequence elements (predominantly transposons), the location in the genome, the type, characteristics and other pertinent information. For further details, please refer to the GFF format description, which can be found at the following URL: http://www.sequenceontology.org/gff3.shtml.
Gene prediction and annotation
The primary approach to gene structure prediction involved the implementation of three distinct methods: de novo prediction, homologous protein-based prediction, and transcriptome-based prediction. Subsequently, the results obtained from these three predictions were integrated to generate a comprehensive dataset. Genscan, Augustus v2.4, GlimmerHMM v3.0.4, GeneID v1.4, SNAP (version 2006-07-28) were used for de novo prediction30,31,32,33,34; Homologous protein-based prediction was done using GeMoMa v1.3.135; Unigene sequences were formed by using PASA v2.0.236. Finally, the prediction results obtained from the above three methods are integrated using EVM v1.1.137 and modified with PASA v2.0.2. The following statistics were provided: the number and average length of protein-coding genes, as well as the number and average length of introns and exons. Non-coding RNAs encompass a variety of functional categories, including microRNAs, rRNAs and tRNAs. Various methodologies have been developed to predict different non-coding RNAs based on their structural characteristics. The tRNAs in the genome were predicted using the software tRNAscan-SE38, and the rRNAs in the genome as well as other ncRNAs besides tRNAs and rRNAs were predicted using the software Infernal 1.1 based on the Rfam database39,40.
The predicted gene sequences were then subjected to a BLAST alignment with the functional databases such as Eukaryotic Orthologous Groups (KOG), Kyoto Encyclopedia of Genes and Genomes (KEGG), Swiss-Prot, TrEMBL, Non-Redundant Protein Database (Nr) and the results of gene function annotation were obtained41,42,43,44,45. Functional annotation of the Gene Ontology (GO) based on the Nr database alignment results with the application software Blast2GO46,47. The function of the protein is annotated using the protein families (Pfam) database with the software HMMER48,49. In addition, metabolic pathway enrichment analysis was determined using the Clusters of Orthologous Genes (COG) and KEGG database. GO function enrichment analysis predicted the biological function of the protein sequence encoded by the gene.
Carbohydrate enzyme annotation
Carbohydrate-active enzymes (CAzymes) are divided into six families: Glycoside Hydrolases (GHs), Glycosyl Transferases (GTs), Polysaccharide Lyases (PLs), Carbohydrate Esterases (CEs), Auxiliary Activities (AAs), Carbohydrate-Binding Modules (CBMs). Classification of enzymes for synthesis, metabolism and transport of carbon compounds and related information can be obtained by CAzymes. Therefore, we used the software HMMER for functional annotation of carbohydrate enzyme genes based on the Carbohydrate Associated Enzyme Database (CAZy)50.
Cytochrome P450 annotation
Cytochrome P450 (CYP450) is a superfamily of proteins involved in the metabolism of organisms including fungi. CYP450 not only catalyses the biosynthetic metabolism of a number of endogenous substances with important physiological functions, but also participates in the biooxidation of many exogenous substances. The target protein sequence was annotated using BlastP, with the Fungal Cytochrome P450 Database (FCPD) serving as the reference51.
Transporter classification database annotation
Transporter Classification Database (TCDB) is a database for the classification of membrane transporter proteins. It has developed a Transporter Classification (TC) System, which is similar to the EC System for the classification of enzymes. The TC System provides TC Numbers consisting of 5 numbers or letters, each of which actually represents a particular level of classification. The protein sequences of all the predicted genes were used to performe BLAST alignment with the TCDB. Signal peptides, short peptide chains (typically 5–30 amino acids in length) that direct newly synthesised proteins towards the secretory pathway. The protein sequences of all the predicted genes were analysed using the software SignalP 4.0 to identify proteins containing signal peptides52. Protein sequences of all the predicted genes were analysed using TMHMM software53 to identify proteins containing transmembrane helices as transmembrane proteins. After the proteins containing transmembrane helices were eliminated from the predicted proteins containing signal peptides, the remaining proteins were secreted proteins. Finally, we further analysed the secreted proteins using EffectorP54 to predict the fungal effector proteins.
Prediction of gene clusters involving secondary metabolites
The antiSMASH (Antibiotics & Secondary Metabolite Analysis Shell, https://antismash.secondarymetabolites.org/) is a tool for identifying and analysing biosynthetic gene clusters (BGCs) in microorganisms. The antiSMASH analyses were run with default parameters (relaxed) to predict the secondary metabolite biosynthesis gene clusters in ZA-14.
Phylogenomics analysis
In order to explore the evolutionary dynamics of ZA-14, the genome sequences of an additional 19 fungal species were downloaded from NCBI for phylogenomics analysis (Supplementary Table S9). The inference of single-copy orthologous genes from the 20 fungal species was conducted using OrthoFinder v2.5.4 with the MAFFT option to facilitate subsequent multiple sequence alignment55. The maximum likelihood tree was constructed on the basis of the resulting alignment using RAxML v8.2.12 with the PROTGAMMAJTT model56. Statistical support values were obtained using non-parametric bootstrap with 1000 replicates. The divergence times of the 20 fungal species were estimated using r8s v1.8157. The time of divergence of 203–245 Mya between Inonotus and Agaricus was selected as normal priors to restrain the age of the nodes58.
The expansion and contraction of gene families were determined using CAFE5 with the following parameters: Poisson distribution, Gamma model59.
Comparative genomics analysis
In order to explore the dynamics of speciation in ZA-14, the genome sequences of S. baumii, S. sanghuang, S. vaninii and I. hispidus were aligned in pairs (Supplementary Table S9) using MCScan (Python version)60. Based on the resulting blocks, genomic synteny map among the five species was drawn using jcvi package in Python 360. The syntenic, translocation, inversion and duplication genomic blocks between ZA-14 and S. baumii, S. sanghuang, S. vaninii or I. hispidus were analysed using MUMmer 4. The results were then visualised by SyRI61,62.
In order to identify the differences and similarities of medicinal application in Sanghuangporus with a uniform standard, the genome sequences of S. baumii, S. sanghuang, S. vaninii and I. hispidus were reannotated using the same pipeline as that of ZA-14.
Results
Species identity
The isolated ZA-14 strain was identified as Inonotus hispidus using rDNA-ITS sequence analysis in our previous study63.
Genome sequence assembly and annotation
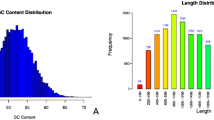
The complete genome of I. hispidus ZA-14 was sequenced from the monokaryotic strain ZA-14 using the Nanopore sequencing platform. The 37.68 Mb genome was assembled from 5771 Mb raw data (153.12 × genome coverage), yielding 51 contigs with a GC content of 48.48% (Table 1). Of the 51 contigs, the longest measured 5.22 Mb, and the N50 length was 2.80 Mb. The GC skew showed no obvious distribution pattern across the whole genome (Fig. 1). Genome assembly completeness assessments revealed that annotation of ZA-14 was very complete, with 92.07% of BUSCOs in the RefSeq annotation set being complete and 2.76% fragmented. A total of 8924 genes were predicted, with an average sequence length of 2420.47 bp, accounting for 57.32% of the whole genome sequence length, which indicated a high gene density in the genome (Fig. 1 and Table 1). In total, 223 non-coding RNAs (ncRNAs) were identified, showing low genomic density, including 96 tRNAs, 76 rRNAs and 51 other ncRNAs. The total length of the repeat sequences was 7,672,428 bp, covering 20.36% of the genomic length. Transposable elements (TEs) constituted approximately 18.03% of the genome, with long terminal repeats (LTRs) and non-LTR transposons accounting for 11.59% and 6.44% of the genome, respectively (Table 1).
Characteristics of the de novo assembly genomic features of I. hispidus ZA-14. From the outside to the inside are: I, contigs, the different colors represent different contigs (> 1 Mb in length); II, GC content: calculated as the percentage of G + C in 20 kb sliding windows and 1 kb non-overlapping windows. The inward blue part represents the GC content in the region lower than the average genome GC content, while the outward purple part represents the opposite; III, GC skew: calculated as the percentage of (G − C)/(G + C) in 20 kb sliding windows and 1 kb non-overlapping windows. The inward green part represents G/C < 1, while the outward pink part represents G/C > 1; IV, Gene density: three circles starting from the dark red color to inside, respectively, represent the numbers of coding genes, tRNAs, rRNAs and coverage of repetitive sequences in 100 kb sliding windows. The intensity of the color positively correlates with gene density; V, Gene density of collinear blocks. The height of the bars and folds is positively correlated with gene density.
The figure was independently created by the authors using genome data generated in this study. The visualization was produced with software circos (v0.69-8) and finalized in Adobe Illustrator2024. The figure design and annotations were completed by Dingbang Ding.
Gene function annotation
All 8924 predicted protein-encoding genes were annotated in this study (Supplementary Table S1). A total of 2882 genes were assigned to the KEGG pathways, of which 22.83% (658) were involved in metabolic processes, accounting for the major proportion (Supplementary Fig. 1 and Supplementary Table S2). In the KOG analysis, 53.89% (4809) of the genes were classified into functional categories; 25.39% (1221) of these genes were associated with metabolism. Among metabolism-related genes, those involved in “amino acid transport and metabolism”, “carbohydrate transport and metabolism” and “Secondary metabolites biosynthesis, transport and catabolism” were the most abundant (Fig. 2 and Supplementary Table S3).
KOG functional classification of predicted protein-encoding genes in the I. hispidus ZA-14 genome.
The biosynthesis of bioactive compounds
The fungus Sanghuang produces a variety of secondary bioactive metabolites such as triterpenoids, polysaccharides and flavonoids, which contribute to its immunomodulatory, antioxidant and antitumour activities64,65. Consequently, genes responsible for the biosynthesis of these secondary metabolites were identified within the fungal genome.
Terpenoid biosynthesis
A total of 11 key enzymes involved in the mevalonate pathway (MVA) were identified in I. hispidus ZA-14 (Supplementary Fig. 2). Among these, diphosphomevalonate decarboxylases were encoded by double-copy, while the remaining 10 enzymes were encoded by single-copy genes (Supplementary Table S4). In addition to the 11 key enzymes in the MVA pathway, genes putatively associated with terpenoid biosynthesis were identified. One gene (ID: EVM0008296.1) encodes lanosterol synthase, which catalyzes the cyclization of the triterpenes (e.g., squalene or 2,3-oxidosqualene) to a protosterol cation and ultimately to lanosterol, the precursor of all steroids66. In addition, two other genes (IDs: EVM0005747.1 and EVM0006066.1) were found involved in the biosynthesis of sesquiterpenoids and triterpenoids (Supplementary Fig. 3).
Polysaccharide biosynthesis
A total of 19 enzymes, encoded by 38 distinct genes, were identified to participate in the biosynthesis of polysaccharides (starch and sucrose metabolism) (Supplementary Fig. 4 and Supplementary Table S4). These enzymes were identified in the I. hispidus ZA-14. Most of these enzymes were encoded by single- or double-copy genes, while alpha-glucosidase synthase and beta-glucosidase were encoded by three-copy and ten-copy genes, respectively. Furthermore, analysis of glucose biosynthesis processes revealed the participation of five enzymes, encoded by six distinct genes. These include 1,3-β-glucan synthase, 1,4-α-glucan branching enzyme, glucose-6-phosphate isomerase, phosphoglucomutase, and UTP-glucose-1-phosphate uridylyltransferase.
Flavonoid biosynthesis
No key enzyme directly involved in the flavone and flavonol biosynthesis was enriched from I. hispidus ZA-14.
Functional annotation of putative CAZymes
We identified CAZymes (Carbohydrate-active enzymes) in the I. hispidus ZA-14 genome and compared CAZyme repertoires with those of other fungi. A total of 381 genes were assigned to CAZyme families, as defined in the CAZy database (Carbohydrate Associated Enzyme Database) (Supplementary Table S6). All CAZymes were classified into six major modules: 66 genes assigned to Auxiliary Activities (AA), 8 genes to carbohydrate-binding modules (CBM), 24 genes to carbohydrate esterases (CE), 198 genes to glycoside hydrolases (GH), 70 genes to glycosyl transferases (GT), and 15 genes for polysaccharide lyases (PL) (Supplementary Table S6).
To analyze the characteristics of CAZymes of I. hispidus ZA-14, we compared its CAZyme repertoires with those of Sanghuangporus baumii, S. sanghuang, S. vaninii and I. hispidus strain. I. hispidus ZA-14 exhibited similar total numbers of genes encoding CAZymes and comparable gene counts across the six classes of CAZymes when compared to two other Inonotus hispidus and three Sanghuangporus species (Fig. 3A). Furthermore, the number of GHs was significantly higher than the number of GTs, indicating that fungal survival is contingent on lignocellulose decomposition. These results suggest that polysaccharide degradation plays a more significant role than polysaccharide synthesis in the growth and metabolism of I. hispidus ZA-14.
Comparative analysis of CAZymes from I. hispidus ZA-14 and other fungi. (A) The comparative analysis of fungal CAZymes. The numerical representation of CAZyme modules or domains is expressed as horizontal bars. (B) Heatmap of CAZyme families found in I. hispidus ZA-14, I. hispidus, S. baumii, S. sanghuang and S. vaninii. The x and y axes represent species and CAZyme families, respectively. The color of each box is indicative of the scaled values of the gene numbers encoding CAZyme families. The color of the boxes is used to indicate the increase in gene numbers encoding CAZyme families, with blue indicating low numbers and red denoting high numbers.
Moreover, CAZymes in gene families from the four selected fungal species were identified and compared with those of I. hispidus ZA-14 (Fig. 3B). The results showed that the copy numbers of CE16, GH179, GH28, GH30, GT57, GT69 and PL42 were higher in these fungi than in I. hispidus ZA-14.
Protein structure prediction
A total of 807 genes were annotated to encode CYPs (Cytochrome P450) in I. hispidus ZA-14. We only obtained 86 membrane transport-related proteins by TCBD annotation. And we identified 721 signal peptide proteins, 1825 transmembrane proteins, 439 secreted proteins and 18 effector proteins (Supplementary Table S7).
Gene cluster
A total of 269 functional genes are involved in regulating the production of secondary metabolites. Of them, 48 functional genes regulated the production of NRPS-like (non-ribosomal peptide synthetases), 73 genes regulated the production of TS (terpene synthases), 8 genes regulated the production of T1PKS (iterative type I polyketide synthases) and 10 genes regulated the production of NRPS (Supplementary Table S8).
Comparative genomic analysis
The phylogenomics tree inferred from an alignment of the single-copy orthologous genes from I. hispidus ZA-14 and the other 20 fungal species with full bootstrap support (Fig. 4A). We found that I. hispidus ZA-14, two species in Inonotus hispidus and three species in Sanghuangporus were phylogenetically separated. Inonotus was estimated to have emerged at a mean crown age of 1.85 Mya. Of the species in Sanghuangporus, those with a mean crown age of 9.51 Mya had a closer phylogenetic relationship. The mean crown age between them is 46.06 Mya.
Comparative genomic analysis of I. hispidus ZA-14 and other 20 fungi. (A) Maximum clade credibility tree inferred from the single-copy orthologous genes. The numbers of gene family expansion and contraction in each species are indicated by plus (in red color) and minus (in green color) symbols, respectively. (B) Gene family (Single-Copy Orthologs, the number of single-copy homologous genes in the species common gene families; Multiple-Copy Orthologs, the number of multiple-copy homologous genes in the species common gene families; Unique genes, genes in specific gene families; Unclustered genes, genes that have not been clustered into any families).
Gene family contraction occurred with gene family expansion during the evolution of the 21 fungal species studied (Fig. 4A). Regarding Inonotus and Sanghuangporus, 222, 68, 323, 67, 142 and 326 gene families had, respectively, expanded in I. hispidus ZA-14, I. hispidus MA, I. hispidus QDAU, S. baumii, S. sanghuang and S. vaninii, corresponding to 551, 135, 42, 728, 54 and 85 gene families being contracted. A total of 5193 orthologous groups were identified from three species of Inonotus hispidus, I. hispidus ZA-14 shared almost orthologous groups with I. hispidus MA (21) and I. hispidus QDAU (29), whereas I. hispidus MA and I. hispidus QDAU shared the maximum orthologous groups (653) (Supplementary Fig. 5A). In contrast to the three Sanghuangporus species, 21 of the 218 genes specific to the I. hispidus ZA-14 (IDs: EVM0003284.1, EVM0003204.1, EVM0003093.1, etc.) were annotated to the guanine nucleotide-binding protein gamma subunit (Supplementary Fig. 5B and Supplementary Table S1). Furthermore, 4752 orthologous groups were identified from I. hispidus ZA-14 and Sanghuangporus, I. hispidus ZA-14 shared almost orthologous groups with S. baumii (20), S. sanghuang (26) and S. vaninii (20). Similarly, the number of single-copy homologous genes and multi-copy homologous genes in I. hispidus ZA-14 was approximately the same as in the two species of I. hispidus and three species of Sanghuangporus (Fig. 4B).
We further performed the genome synteny analysis of I. hispidus ZA-14, I. hispidus MA and three Sanghuangporus species. The shared synteny between I. hispidus ZA-14 and I. hispidus MA was the highest among I. hispidus and the other three species of Sanghuangporus (Fig. 5). Of the shared synteny between I. hispidus ZA-14 and I. hispidus MA, S. vaninii, the region of translocation and inversion was much higher than that of the synteny accounting only for a small proportion of the whole genome sequence (Supplementary Fig. 6).
Genomic synteny of I. hispidus ZA-14, I. hispidus, S. baumii, S. sanghuang and S. vaninii. Shared blocks of synteny are linked by different color lines between I. hispidus ZA-14 and the other selected four fungi. The figure was independently created by the authors using genome data generated in this study. The dynamics of speciation among species was analyzed using MCScan (Python version), the visualization was produced with jcvi package and finalized in Adobe Illustrator 2024. The figure design and annotations were completed by Dingbang Ding.
Discussion
Limited genetic information on I. hispidus has hampered its applications in the molecular mechanisms of host function and interactions with plants, although the fungus is a key commercial products in agricultural productio67. To address this, the development of a whole-genome map of I. hispidus was undertaken to facilitate the exploration of its genetic information and the annotation of related functional genes. These limitations underscore the necessity of a comprehensive genomic framework, which our study provides through the assembly and annotation of the ZA-14 genome. The classification of Sanghuang remains controversial due to the fact that no national or industrial standards exist for the evaluation of its active substances. Although the establishment of the genus Sanghuangporus (containing 15 species) has partially resolved this issue, it has not been completely resolved68,69,70,71. I. hispidus in the genus hispidus is still referred to as Sanghuang in traditional medicine due to its similarity in morphology and medicinal properties to Sanghuangporus species72. To address this gap, our whole-genome analysis provides a foundation for exploring how genetic architecture underpins its ecological adaptability. Notably, our phylogenetic analysis reveals that several well-known medicinal macrofungi, including G. sinense, S. commune and W. cocos, are distantly related to I. hispidus ZA-14 and each other. This challenges the hypothesis that medicinal properties necessarily evolve through convergent evolution across different clades73, suggesting that such properties may not play a significant role in shaping the life cycles of these fungi. Unexpectedly, I. hispidus ZA-14 was found to be closely related to two species of I. hispidus and three species of Sanghuangporus, implying medicinal traits may be inherited from a common ancestor rather than arising independently. At the generic level, our findings suggest that the genetic toolkit for producing bioactive compounds could be present in the ancestor of the Sanghuang clade. The speciation of I. hispidus and Sanghuangporus is likely driven primarily by genome sequence translocations and inversions from their common ancestor. The overlapping emergence times of these fungi with their specific hosts may have further contributed to speciation by driving adaptation to the distinct lignocellulose composition and defense mechanisms of each host74. Notably, the delayed speciation event in I. hispidus ZA-14 occurred compared to the other two I. hispidus species is attributed to prolonged environmental adaptation, explaining the observed metabolic differences between the species75. For instance, I. hispidus ZA-14 shows specific annotation to the guanine nucleotide-binding protein gamma subunit, which is distinct from the three Sanghuangporus species and is a G-protein family that regulate inflammatory signaling cascade76. This genetic variation could explain why I. hispidus ZA-14 is associated with lower uric acid and xanthine oxidase levels77, providing new genomic evidence supporting this pharmacological link.
Current phylogenomic studies have demonstrated significant gene family loss in fungi. Indeed, this phenomenon of gene loss has been observed across all life forms and is recognized as a fundamental evolutionary force78. Given the lack of significant difference in genes and gene clusters related to nutritional strategies and medicinal properties between I. hispidus ZA-14 and species of Sanghuangporus, we speculate that the expansion and contraction of gene families, particularly those undergoing rapid evolution, play a significant role in the host specificity of Sanghuang species. Cytochrome P450 (CYP450) enzymes represent a superfamily of monooxygenases pivotal to numerous biological processes, including primary and secondary metabolism, across a diverse array of life forms79,80. In this study, 807 CYPs were obtained through annotating the I. hispidus ZA-14 for cytochrome P450. Fungi have evolved in response to the limitations of their environment, diversifying their CYP450 to facilitate survival as a living resource81,82. It is hypothesised that fungal CYP450 are involved in the detoxification of natural and environmental contaminants, thus facilitating survival in diverse ecological niches83. Moreover, the potential of fungal CYP450 as an antifungal drug target is a promising approach for the control of pest and plant pathogenic fungi.
Such dynamic gene family evolution may directly influence the functional specialization of I. hispidus, as evidenced by its enriched secondary metabolite biosynthesis pathways. Secondary metabolites (e.g.,terpenoids, polysaccharides) derived from the Sanghuang fungus, I. hispidus, and other medicinal macrofungi offer novel strategies for addressing various diseases84. A comparative analysis of secondary metabolite gene clusters shows that filamentous fungi are characterized by an increased prevalence of T1PKS and NRPS gene clusters, whereas basidiomycetes are distinguished by a preponderance of TS gene clusters85. The terpene synthase family, a medium-sized gene family, catalyzes the biosynthesis of monoterpene, sesquiterpene, and diterpene backbones86. Correspondingly, the I. hispidus ZA-14 has more TS gene clusters (73) that participates in the biosynthesis of terpenoids than other gene clusters. Triterpenoids extracted from S. baumii and S. sanghuang show a high antioxidant, anti-inflammatory, and anti-tumor activities87,88,89,90. Genes in the conserved MVA pathway involved in the biosynthesis of terpene backbones were annotated in the I. hispidus ZA-14 and these genes are involved in the synthesis of triterpenoids. Further homology analysis demonstrated that the fungi and the three species of Sanghuangporus showed high genetic similarity. This suggests that similar genomes have similar biological activity.
Polysaccharides are the most studied metabolites in macrofungi and their different molecular weights, branching structures, conformations and chemical modifications provide the structural basis for a wide range of biological activities, including anti-tumour, antioxidant, anti-inflammatory and immunomodulatory activities91,92. We found 38 protein-coding genes involved in the pathway of polysaccharide biosynthesis in I. hispidus ZA-14, viz. the starch and sucrose metabolism pathway. In the same way, a similar number of genes that are involved in the polysaccharide biosynthesis pathway were identified in each of the three Sanghuangporus species93. The results suggest that polysaccharides may also be the main medicinal metabolites of Inonotus, but further experiments are needed to confirm this hypothesis.
The absence of key enzymes in the flavonoid biosynthesis pathway in I. hispidus ZA-14 raises intriguing questions about the metabolic flexibility of this fungus. It is possible that alternative pathways or novel enzymes compensate for the lack of traditional biosynthetic routes. Future studies combining transcriptomic and metabolomic analyses could provide deeper insights into these metabolic adaptations and their role in the medicinal properties of I. hispidus.
The genomic insights into CAZymes and nutrient utilization pathways lay a foundation for optimizing cultivation strategies to enhance yield and bioactive compound production. Artificial cultivation is a practical solution to the scarcity of available mature fruiting bodies, which can be facilitated by genomic information94. By analyzing the CAZymes of I. hispidus ZA-14 in comparison with I. hispidus and the three species of Sanghuangporus, we found that the GH (Glycoside hydrolase) family genes were significantly enriched in the fungi. Glycoside hydrolase-related genes are essential for mushroom growth and development, particularly in cap expansion and overall senescence95. They facilitate nutrient absorption by degrading starch and other substrates, enabling mushrooms to utilize diverse energy sources, with lignocellulose degradation being a critical component of this process96,97. The evidence suggests that the enrichment of GH family genes at the gene level enhances the growth of I. hispidus ZA-14 by improving its ability to absorb and utilize nutrients from diverse substrates as energy sources. Consequently, the knowledge of CAZymes has the potential to promote the breeding and cultivation of I. hispidus ZA-14, thereby providing a theoretical foundation for enhancing the yield at the genetic level. The results obtained can be utilized to subsequently combine transcriptomics in order to compare the key genes involved in these processes and thereby produce I. hispidus that is both high-yielding and high-quality.
Conclusion
The comprehensive genomic analysis of I. hispidus ZA-14 has provided valuable insights into its genetic architecture and evolutionary history. The identification of a robust repertoire of genes involved in the biosynthesis of secondary metabolites, such as terpenoids and polysaccharides, corroborates the traditional use of I. hispidus as a medicinal fungus with immunomodulatory, antioxidant, and antitumor properties. Notable features include a high gene density and the presence of multiple gene families associated with secondary metabolism, such as terpene synthases (e.g., terpene synthases) and glycoside hydrolases, which may contribute to the fungus’s ecological adaptability and nutritional strategies. Comparative genomics with other Sanghuangporus and Inonotus species revealed distinct evolutionary trajectories and gene family expansions. These findings not only enhance our understanding of the medicinal potential of I. hispidus but also provide a genetic basis for strain cultivation and commercial production. The genomic dataset generated herein serves as a critical resource for future research, including efforts to enhance yield and quality, as well as explore its roles in ecosystem services and biotechnological applications.
Data availability
The raw sequencing data of this study has been deposited in National Center for Biotechnology Information (NCBI) with BioProject accession number PRJNA1209412.
References
Anonymous. Shen Nong Materia Medica, 102-200 A.D. In: Han, E. (ed) Reprinted (People’s Hygiene Press, Beijing, 1955)
Yuan, Y. et al. Archaeological evidence suggests earlier use of Ganoderma in Neolithic China. Chin. Sci. Bull. 63, 1180–1188. https://www.sci-hub.ru/10.13341/j.jfr.2014.1171 (2018).
Zhou, L.-W., Ghobad-Nejhad, M., Tian, X.-M., Wang, Y.-F. & Wu, F. J. F. R. I. Current status of ‘Sanghuang’as a group of medicinal mushrooms and their perspective in industry development. Food Rev. Int. 38, 589–607. https://doi.org/10.1080/87559129.2020.1740245 (2022).
Dai, Y., Qin, G. & Xu, M.. The forest pathogens of root and butt rot in Northeast China. Forest Research.13, 15–22. https://www.sci-hub.ru/10.13275/j.cnki.lykxyj.2000.01.003 (2012)
Bao, H., Yang, S., Li, Q., Bau, T. & Li, Y. J. J. F. R. Supplementary textual research on “Sanghuang”. Fungal Res. 4, 264–270. https://doi.org/10.13341/j.jfr.2014.1171 (2017).
Gründemann, C. et al. Influence of Inonotus hispidus on function of human immune cells. Eur. J. Integr. Med. 1, 54. https://doi.org/10.1016/j.eujim.2016.08.128 (2016).
Islam, M. T. et al. Anticancer perspectives on the fungal-derived polyphenolic hispolon. Anti-Cancer Agent Me. 20, 1636–1647. https://doi.org/10.2174/1871520620666200619164947 (2020).
Liu, X. et al. Extraction, characterization and antioxidant activity analysis of the polysaccharide from the solid-state fermentation substrate of Inonotus hispidus. Int. J. Biol. Macromol. 123, 468–476. https://doi.org/10.1016/j.ijbiomac.2018.11.069 (2019).
Angelini, P. et al. A comparative study of the antimicrobial and antioxidant activities of Inonotus hispidus fruit and their mycelia extracts. Int. J. Food Prop. 22, 768–783. https://doi.org/10.1080/10942912.2019.1609497 (2019).
Politi, M. et al. Current analytical methods to study plant water extracts: The example of two mushrooms species, Inonotus hispidus and Sparassis crispa. Phytochem. Anal. PCA. 18, 33–41. https://doi.org/10.1002/pca.949 (2007).
Zan, L.-F. et al. Antioxidant hispidin derivatives from medicinal mushroom Inonotus hispidus. Chem. Pharm. Bull. 59, 770–772. https://doi.org/10.1248/cpb.59.770 (2011).
Ali, N. A., Jansen, R., Pilgrim, H., Liberra, K. & Lindequist, U. J. P. Hispolon, a yellow pigment from Inonotus hispidus. Phytochemistry 41, 927–929. https://doi.org/10.1016/0031-9422(95)00717-2 (1996).
Fan, H.-C. et al. Dehydroxyhispolon methyl ether, a hispolon derivative, inhibits WNT/β-catenin signaling to elicit human colorectal carcinoma cell apoptosis. Int. J. Mol. Sci. 21, 8839. https://doi.org/10.3390/ijms21228839 (2020).
Sarfraz, A. et al. Hispolon: A natural polyphenol and emerging cancer killer by multiple cellular signaling pathways. Environ. Res. 190, 110017. https://doi.org/10.1016/j.envres.2020.110017 (2020).
Benarous, K. et al. Harmaline and hispidin from Peganum harmala and Inonotus hispidus with binding affinity to Candida rugosa lipase: In silico and in vitro studies. Bioorg. Chem. 62, 1–7. https://doi.org/10.1016/j.bioorg.2015.06.005 (2015).
Thanh, N. T. et al. Chemical constituents from the fruiting bodies of Phellinus igniarius. Nat. Prod. Res. 32, 2392–2397. https://doi.org/10.1080/14786419.2017.1413572 (2018).
Maertens, J. A. History of the development of azole derivatives. Clin. Microbiol. Infect. 10, 1–10. https://doi.org/10.1111/j.1470-9465.2004.00841.x (2004).
White, T. J. PCR protocols: A guide to methods and applications. 8, 335. https://www.sci-hub.ru/10.1016/0167-7799(90)90215-J (1990).
Payne, A., Holmes, N., Rakyan, V. & Loose, M. BulkVis: A graphical viewer for Oxford nanopore bulk FAST5 files. Bioinformatics 35, 2193–2198. https://doi.org/10.1093/bioinformatics/bty841 (2019).
Walker, B. J. et al. Pilon: An integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 9, e112963. https://doi.org/10.1371/journal.pone.0112963 (2014).
Simao, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: Assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212. https://doi.org/10.1093/bioinformatics/btv351 (2015).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760. https://doi.org/10.1093/bioinformatics/btp324 (2009).
Xu, Z. & Wang, H. LTR_FINDER: An efficient tool for the prediction of full-length LTR retrotransposons. Nucleic Acids Res. 35, W265–W268. https://doi.org/10.1093/nar/gkm286 (2007).
Han, Y. J. & Wessler, S. R. MITE-Hunter: A program for discovering miniature inverted-repeat transposable elements from genomic sequences. Nucleic Acids Res. 38, e199. https://doi.org/10.1093/nar/gkq862 (2010).
Price, A. L., Jones, N. C. & Pevzner, P. A. Identification of repeat families in large genomes. Bioinformatics 21, I351–I358. https://doi.org/10.1093/bioinformatics/bti1018 (2005).
Edgar, R. C. & Myers, E. W. PILER: Identification and classification of genomic repeats. Bioinformatics 21, I152–I158. https://doi.org/10.1093/bioinformatics/bti1003 (2005).
Bao, W. D., Kojima, K. K. & Kohany, O. Repbase Update, a database of repetitive elements in eukaryotic genomes. Mobile DNA-Uk 6, 11. https://doi.org/10.1186/s13100-015-0041-9 (2015).
Wicker, T. et al. A unified classification system for eukaryotic transposable elements. Nat. Rev. Genet. 8, 973–982. https://doi.org/10.1038/nrg2165 (2007).
Tarailo-Graovac, M. & Chen, N. Using RepeatMasker to identify repetitive elements in genomic sequences. Curr. Protocols Bioinform. https://doi.org/10.1002/0471250953.bi0410s25 (2009).
Blanco, E., Parra, G. & Guigó, R. Using geneid to identify genes. Curr. Protocols Bioinform. https://doi.org/10.1002/0471250953.bi0403s18 (2007).
Majoros, W. H., Pertea, M. & Salzberg, S. L. TigrScan and GlimmerHMM: Two open source eukaryotic gene-finders. Bioinformatics 20, 2878–2879. https://doi.org/10.1093/bioinformatics/bth315 (2004).
Burge, C. & Karlin, S. Prediction of complete gene structures in human genomic DNA. J. Mol. Biol. 268, 78–94. https://doi.org/10.1006/jmbi.1997.0951 (1997).
Stanke, M. & Waack, S. Gene prediction with a hidden Markov model and a new intron submodel. Bioinformatics 19(Suppl 2), ii215–ii225. https://doi.org/10.1093/bioinformatics/btg1080 (2003).
Korf, I. Gene finding in novel genomes. Bmc Bioinform. 5, 59. https://doi.org/10.1186/1471-2105-5-59 (2004).
Keilwagen, J. et al. Using intron position conservation for homology-based gene prediction. Nucleic Acids Res. 44, e89. https://doi.org/10.1093/nar/gkw092 (2016).
Campbell, M. A., Haas, B. J., Hamilton, J. P., Mount, S. M. & Buell, C. R. Comprehensive analysis of alternative splicing in rice and comparative analyses with Arabidopsis. Bmc Genom. 7, 327. https://doi.org/10.1186/1471-2164-7-327 (2006).
Haas, B. J. et al. Automated eukaryotic gene structure annotation using EVidenceModeler and the program to assemble spliced alignments. Genome Biol. 9, R7. https://doi.org/10.1186/gb-2008-9-1-r7 (2008).
Lowe, T. M. & Eddy, S. R. tRNAscan-SE: A program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 25, 955–964. https://doi.org/10.1093/nar/25.5.955 (1997).
Nawrocki, E. P. et al. Rfam 1.20: Updates to the RNA families database. Nucleic Acids Res. 43, D130–D137. https://doi.org/10.1093/nar/gku1063 (2015).
Nawrocki, E. P. & Eddy, S. R. Infernal 1.1: 100-fold faster RNA homology searches. Bioinformatics 29, 2933–2935. https://doi.org/10.1093/bioinformatics/btt509 (2013).
Boeckmann, B. et al. The SWISS-PROT protein knowledgebase and its supplement TrEMBL in 2003. Nucleic Acids Res. 31, 365–370. https://doi.org/10.1093/nar/gkg095 (2003).
Kanehisa, M., Goto, S., Kawashima, S., Okuno, Y. & Hattori, M. The KEGG resource for deciphering the genome. Nucleic Acids Res. 32, D277–D280. https://doi.org/10.1093/nar/gkh063 (2004).
Yangyang, D. et al. Integrated NR database in protein annotation system and its localization. Comput. Eng. 32, 71–72. https://www.sci-hub.ru/10.1109/INFOCOM.2006.241 (2006).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402. https://doi.org/10.1093/nar/25.17.3389 (1997).
Tatusov, R. L., Galperin, M. Y., Natale, D. A. & Koonin, E. V. The COG database: A tool for genome-scale analysis of protein functions and evolution. Nucleic Acids Res. 28, 33–36. https://doi.org/10.1093/nar/28.1.33 (2000).
Conesa, A. et al. Blast2GO: A universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21, 3674–3676. https://doi.org/10.1093/bioinformatics/bti610 (2005).
Ashburner, M. et al. Gene Ontology: Tool for the unification of biology. Nat. Genet. 25, 25–29. https://doi.org/10.1038/75556 (2000).
Eddy, S. R. Profile hidden Markov models. Bioinformatics 14, 755–763. https://doi.org/10.1093/bioinformatics/14.9.755 (1998).
Finn, R. D. et al. The Pfam protein families database: Towards a more sustainable future. Nucleic Acids Res. 44, D279–D285. https://doi.org/10.1093/nar/gkv1344 (2016).
Cantarel, B. L. et al. The carbohydrate-active EnZymes database (CAZy): An expert resource for glycogenomics. Nucleic Acids Res. 37, D233–D238. https://doi.org/10.1093/nar/gkn663 (2009).
Park, J. et al. Fungal cytochrome P450 database. BMC Genom. 9, 1–11. https://www.sci-hub.ru/10.1186/1471-2164-9-402 (2008).
Petersen, T. N., Brunak, S., von Heijne, G. & Nielsen, H. SignalP 4.0: Discriminating signal peptides from transmembrane regions. Nat. Methods 8, 785–786. https://doi.org/10.1038/nmeth.1701 (2011).
Krogh, A., Larsson, B., von Heijne, G. & Sonnhammer, E. L. L. Predicting transmembrane protein topology with a hidden Markov model: Application to complete genomes. J. Mol. Biol. 305, 567–580. https://doi.org/10.1006/jmbi.2000.4315 (2001).
Sperschneider, J. et al. EffectorP: Predicting fungal effector proteins from secretomes using machine learning. New Phytol. 210, 743–761. https://doi.org/10.1111/nph.13794 (2016).
Emms, D. M. & Kelly, S. OrthoFinder: Phylogenetic orthology inference for comparative genomics. Genome Biol. 20, 238. https://doi.org/10.1186/s13059-019-1832-y (2019).
Stamatakis, A., Hoover, P. & Rougemont, J. A rapid bootstrap algorithm for the RAxML web servers. Syst. Biol. 57, 758–771. https://doi.org/10.1080/10635150802429642 (2008).
Sanderson, M. J. r8s: Inferring absolute rates of molecular evolution and divergence times in the absence of a molecular clock. Bioinformatics 19, 301–302. https://doi.org/10.1093/bioinformatics/19.2.301 (2003).
Kumar, S., Stecher, G., Suleski, M. & Hedges, S. B. TimeTree: A resource for timelines, timetrees, and divergence times. Mol. Biol. Evol. 34, 1812–1819. https://doi.org/10.1093/molbev/msx116 (2017).
Han, M. V., Thomas, G. W. C., Lugo-Martinez, J. & Hahn, M. W. estimating gene gain and loss rates in the presence of error in genome assembly and annotation using CAFE 3. Mol. Biol. Evol. 30, 1987–1997. https://doi.org/10.1093/molbev/mst100 (2013).
Wang, Y. P. et al. A toolkit for detection and evolutionary analysis of gene synteny and collinearity. Nucleic Acids Res. 40, e49. https://doi.org/10.1093/nar/gkr1293 (2012).
Marçais, G. et al. MUMmer4: A fast and versatile genome alignment system. PLoS Comput. Biol. 14, e1005944. https://doi.org/10.1371/journal.pcbi.1005944 (2018).
Goel, M., Sun, H. Q., Jiao, W. B. & Schneeberger, K. SyRI: Finding genomic rearrangements and local sequence differences from whole-genome assemblies. Genome Biol. 20, 277. https://doi.org/10.1186/s13059-019-1911-0 (2019).
Wang, H. et al. Isolation and identification of a wild Sanghuang strain in northern Shaanxi, optimization of solid medium and determination of medicinal component content. Acta Sericologica Sinica. 48, 69-76. https://www.sci-hub.ru/10.13441/j.cnki.cykx.2022.01.010 (2022).
Lin, W.-C. et al. Anti-inflammatory activity of Sanghuangporus sanghuang mycelium. Int. J. Mol. Sci. 18, 347. https://doi.org/10.3390/ijms18020347 (2017).
Liu, K. et al. Polyphenolic composition and antioxidant, antiproliferative, and antimicrobial activities of mushroom Inonotus sanghuang. LWT-food Sci. Technol. 82, 154–161. https://doi.org/10.1016/j.lwt.2017.04.041 (2017).
Yuan, Y., Wu, F., Si, J., Zhao, Y.-F. & Dai, Y.-C.J.G. Whole genome sequence of Auricularia heimuer (Basidiomycota, Fungi), the third most important cultivated mushroom worldwide. Genomics 111, 50–58. https://doi.org/10.1016/j.ygeno.2017.12.013 (2019).
Prasad Singh, P., Srivastava, D., Jaiswar, A. & Adholeya, A. Effector proteins of Rhizophagus proliferus: Conserved protein domains may play a role in host-specific interaction with different plant species. Brazil. J. Microbiol. 50, 593–601. https://doi.org/10.1007/s42770-019-00099-x (2019).
Wu, S. H., Chang, C. C., Wei, C. L., Jiang, G. Z. & Cui, B. K. Sanghuangporus toxicodendri sp. nov. (Hymenochaetales, Basidiomycota) from China. MycoKeys 57, 101–111. https://doi.org/10.3897/mycokeys.57.36376 (2019).
Zhu, L., Song, J., Zhou, J. L., Si, J. & Cui, B. K. Species diversity, phylogeny, divergence time, and biogeography of the Genus Sanghuangporus (Basidiomycota). Front. Microbiol. 10, 812. https://doi.org/10.3389/fmicb.2019.00812 (2019).
Wu, S. H., et al. Species clarification for the medicinally valuable ‘sanghuang’ mushroom. Bot. Stud. 53, 135–149 (2012).
Wu, S. H. & Dai, Y. C. Species clarification of the medicinal fungus Sanghuang. Mycosystema. 39, 781-794. https://www.sci-hub.ru/10.13346/j.mycosystema.190354 (2020).
Wang, Z. X., Feng, X. L., Liu, C., Gao, J. M. & Qi, J. Diverse metabolites and pharmacological effects from the basidiomycetes Inonotus hispidus. Antibiotics (Basel, Switzerland) https://doi.org/10.3390/antibiotics11081097 (2022).
Nagy, L. G. et al. Latent homology and convergent regulatory evolution underlies the repeated emergence of yeasts. Nat. Commun. 5, 4471. https://doi.org/10.1038/ncomms5471 (2014).
Kumar, S., Stecher, G., Suleski, M. & Hedges, S. B. TimeTree: A resource for timelines, timetrees, and divergence times. Mol. Biol. Evol. 34, 1812–1819. https://doi.org/10.1093/molbev/msx116 (2017).
Wang, Q. C., Bao, H. Y. & Li, Z. J. Genomic comparison between two strains isolated from growing in different tree species. Front. Genet. 14, 1221491. https://doi.org/10.3389/fgene.2023.1221491 (2023).
Li, J. et al. GNG12 regulates PD-L1 expression by activating NF-κB signaling in pancreatic ductal adenocarcinoma. FEBS Open Bio 10, 278–287. https://doi.org/10.1002/2211-5463.12784 (2020).
Sun, Z. et al. Anti-gouty arthritis and anti-hyperuricemia properties of Sanghuangporus vaninii and Inonotus hispidus in rodent models. Nutrients https://doi.org/10.3390/nu14204421 (2022).
Albalat, R. & Cañestro, C. Evolution by gene loss. Nat. Rev. Genet. 17, 379–391. https://doi.org/10.1038/nrg.2016.39 (2016).
Crešnar, B. & Petrič, S. Cytochrome P450 enzymes in the fungal kingdom. Biochem. Biophys. Acta. 29–35, 2011. https://doi.org/10.1016/j.bbapap.2010.06.020 (1814).
Hussain, R., Ahmed, M., Khan, T. A. & Akhter, Y. Fungal P(450) monooxygenases - the diversity in catalysis and their promising roles in biocontrol activity. Appl. Microbiol. Biotechnol. 104, 989–999. https://doi.org/10.1007/s00253-019-10305-3 (2020).
Ferrer-Sevillano, F. & Fernández-Canón, J. M. J. E. C. Novel phacB-encoded cytochrome P450 monooxygenase from Aspergillus nidulans with 3-hydroxyphenylacetate 6-hydroxylase and 3, 4-dihydroxyphenylacetate 6-hydroxylase activities. Eukaryot Cell 6, 514–520. https://doi.org/10.1128/ec.00226-06 (2007).
Podobnik, B. et al. CYP53A15 of Cochliobolus lunatus, a target for natural antifungal compounds. J. Med. Chem. 51, 3480–3486. https://doi.org/10.1021/jm800030e (2008).
Pratiwi, R. A., Yahya, N. S. W. & Chi, Y. Bio function of cytochrome P450 on fungus: A review. IOP Conf. Ser. Earth Environ. Sci. 959, 012023. https://doi.org/10.1088/1755-1315/959/1/012023 (2022).
Lu H , Lou H , Hu J ,et al. Macrofungi: A review of cultivation strategies, bioactivity, and application of mushrooms. Compreh. Rev. Food Sci. Food Saf. 19, 2333-2356. https://doi.org/10.1111/1541-4337.12602 (2020).
Wawrzyn, G. T., Held, M. A., Bloch, S. E. & Schmidt-Dannert, C. in Biosynthesis and Molecular Genetics of Fungal Secondary Metabolites, Volume 2 (eds Zeilinger, S., Martín, J.-F. & García-Estrada, C.) 43–65 (Springer New York, 2015).
Chen, F., Tholl, D., Bohlmann, J. & Pichersky, E. The family of terpene synthases in plants: A mid-size family of genes for specialized metabolism that is highly diversified throughout the kingdom. Plant J. 66, 212–229. https://doi.org/10.1111/j.1365-313X.2011.04520.x (2011).
Cai, C. S. et al. Extraction and antioxidant activity of total triterpenoids in the mycelium of a medicinal fungus. Sci. Rep.-Uk 9, 7418. https://doi.org/10.1038/s41598-019-43886-0 (2019).
Sun, Y. et al. Chemical structure and anti-inflammatory activity of a branched polysaccharide isolated from Phellinus baumii. Carbohyd. Polym. 268, 118214. https://doi.org/10.1016/j.carbpol.2021.118214 (2021).
Wang, T. et al. Effective isolation of antioxidant Phelligridin LA from the fermentation broth of Inonotus baumii by macroporous resin. Bioprocess Biosyst. Eng. 43, 2095–2106. https://doi.org/10.1007/s00449-020-02398-2 (2020).
Zheng, N. et al. Optimization of extraction process and the antioxidant activity of phenolics from Sanghuangporus baumii. Molecules 26, 3850. https://doi.org/10.3390/molecules26133850 (2021).
Wu, F. et al. Resource diversity of Chinese macrofungi: Edible, medicinal and poisonous species. Fungal Diversity 98, 1–76. https://doi.org/10.1007/s13225-019-00432-7 (2019).
Maity, P. et al. Biologically active polysaccharide from edible mushrooms: A review. Int. J. Biol. Macromol. 172, 408–417. https://doi.org/10.1016/j.ijbiomac.2021.01.081 (2021).
Jiang, J. H., Wu, S. H. & Zhou, L. W. The First Whole Genome Sequencing of Sanghuangporus sanghuang provides insights into its medicinal application and evolution. J. Fungi https://doi.org/10.3390/jof7100787 (2021).
Lu, H. Y., Lou, H. H., Hu, J. J., Liu, Z. J. & Chen, Q. H. Macrofungi: A review of cultivation strategies, bioactivity, and application of mushrooms. Compreh. Rev. Food Sci. Food Saf. 19, 2333–2356. https://doi.org/10.1111/1541-4337.12602 (2020).
Tao, Y. et al. Identification and expression analysis of a new glycoside hydrolase family 55 exo-β-1,3-glucanase-encoding gene in Volvariella volvacea suggests a role in fruiting body development. Gene 527, 154–160. https://doi.org/10.1016/j.gene.2013.05.071 (2013).
Fang, M. et al. Genome sequence analysis of Auricularia heimuer combined with genetic linkage map. J. Fungi (Basel, Switzerland) https://doi.org/10.3390/jof6010037 (2020).
Lin Z., Zhao R. Advances of genomics-assisted cultivation and breeding of edible and medicinal mushrooms. Acta Edulis Fungi 25, 93. https://doi.org/10.16488/j.cnki.1005-9873.2018.01.015 (2018).
Funding
This study was supported by the Fundamental Research Funds for the Central Universities (2452022006), the China Agriculture Research System of MOF and MARA (CARS-18) and Autonomous innovation project of Henan Academy of Agricultural Sciences (2024ZC122).
Author information
Authors and Affiliations
Contributions
DBD conceived and designed the study. DBD, HW, and JJL analyzed the data and wrote the manuscript. CS, MJZ, and FJ provided assistance for the study. HW, CS and LJB provided the funding. LJB and XW supervised the study and revised the manuscript. All authors have read and agreed to the version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ding, D., Wang, H., Liang, J. et al. Genomic analysis of Inonotus hispidus provides insights into its medicinal properties and evolutionary dynamics. Sci Rep 15, 23031 (2025). https://doi.org/10.1038/s41598-025-05528-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-05528-6
Keywords
This article is cited by
-
Bipolar, not tetrapolar: mating system determination in Inonotus hispidus through genomic and phenotypic analysis
Applied Microbiology and Biotechnology (2026)
-
Inonotus hispidus (Bull.) P. Karst from the Inner Western Anatolia part of Türkiye with a molecular analysis, phenotypic characters study, genus of host determination, and geo-hydrographic distribution factors analysis
Bulletin of the National Research Centre (2025)