Abstract
In this multicenter study we investigated whether echocardiography-derived left ventricular global longitudinal strain (LV GLS), an indicator of myocardial fibrosis, independently predicts a positive genotype in hypertrophic cardiomyopathy (HCM). We performed a cross-sectional analysis including HCM patients with genetic testing results and echocardiographic data from two Austrian HCM registries. Echocardiographic parameters were measured in post-processing analysis by a blinded investigator. Among 125 patients with HCM, a positive genotype was present in 39%. Worse LV GLS was associated with a positive genotype in univariate analysis (Odds Ratio [OR] 95% CI 1.141, 1.018–1.279, p = 0.023). In multivariate regression analysis adjusted for genotype predictors (age at diagnosis < 45 years, arterial hypertension, positive family history of HCM, maximal to posterior wall thickness [MWTH: PWTH], reverse curve septal phenotype), the reverse curve septal phenotype remained as a single independent predictor of genotype-positive HCM (OR 6.948, 2.342–20.614, p < 0.001). Adding LV GLS to established Toronto and Mayo genotype prediction scores did not improve their performance. To conclude, worse LV GLS was not independently associated with genotype-positive HCM and did not improve the diagnostic yield of genetic testing in HCM in a multivariate model. Our study highlights the reverse curve septal phenotype as the strongest genotype predictor in HCM.
Similar content being viewed by others
Introduction
Hypertrophic cardiomyopathy (HCM) is defined as left ventricular hypertrophy not solely explained by abnormal loading conditions. HCM is the most common hereditary cardiomyopathy affecting up to 0.6% in the general population1. According to the novel 2023 ESC guidelines on cardiomyopathies, genetic testing is recommended to confirm the diagnosis in every patient and to enable cascade genetic testing in relatives, which should facilitate closer clinical follow-ups, earlier diagnosis and prevention of cardiac complications2. Moreover, genotype-positive (i.e. carriers of a pathogenic or likely pathogenic [P/LP] variant) compared to genotype-negative HCM patients are at higher risk for subsequent cardiovascular events according to the Sarcomeric Human Cardiomyopathy (ShaRe) Registry3. However, in clinical practice a positive genotype can only be identified in approximately 40% of HCM patients2. Genetic testing is a challenge to the health care system, as it is costly and capacities at genetic institutes are limited. Gruner et al. developed the Toronto Hypertrophic Cardiomyopathy Genotype Score for Prediction of a Positive Genotype in HCM („Toronto Score“). Predictors of a positive genotype were younger age at diagnosis, female sex, the absence of arterial hypertension, a positive family history of HCM, a reverse curve or neutral septal phenotype, and a higher ratio of maximal to posterior wall thickness (MWTH: PWTH)4. Bos et al. were able to confirm similar genotype predictors in a large cohort of 1053 HCM patients presenting the Phenotype-Based Genetic Test Prediction Score for Unrelated Patients („Mayo Score“)5. Both scores demonstrated good to excellent diagnostic accuracy4,5,6,7. Yet, both scores show limited diagnostic accuracy in their intermediate ranges4,5, and underline the necessity of novel parameters to improve pretest probability in those patients.
In genotype-positive HCM, a higher degree of myocardial fibrosis revealed by cardiac magnetic resonance imaging (cMRI) was shown in several studies8,9,10. Echocardiography-derived myocardial deformation analysis is an in-expensive and readily available modality delivering left ventricular global longitudinal strain (LV GLS) as readout. LV GLS serves well as an indicator of myocardial fibrosis in HCM, but previous studies reported conflicting results regarding the associations between LV GLS and a positive genotype11,12,13.
In this multicenter study we aimed to investigate whether echocardiography-derived LV GLS independently predicts a positive genotype and improves the diagnostic yield of the Toronto and Mayo Genotype Prediction Score for patients with HCM.
Methods
Study design
This is an Austrian multi-center cross-sectional analysis of the HCM Registries Graz and Innsbruck. The study was conducted in compliance with Good Clinical Practice and the Declaration of Helsinki. All patients provided written informed consent to participate and both registries were approved by the ethics committees of the competent Medical Universities of Graz (EC-No 30–286 ex 17/18) and Innsbruck (EC-No 1192/2023). For this study, we included patients with a diagnosis of HCM, availability of genetic testing results and a stored transthoracic echocardiography study with adequate image quality of apical chamber views allowing for post-hoc offline myocardial deformation analysis. In our study, HCM was defined as either an interventricular septal thickness ≥ 13 mm and guideline-defined cardiomyopathy-specific red flags, or ≥ 15 mm not explainable by loading conditions, or ≥ 17 mm. Patients were enrolled between 2019 and 2023. Demographic and vital parameters, laboratory values, signs and symptoms of HCM, past medical history, and genetic testing results were retrieved from registry data. Family histories of HCM and sudden cardiac death (SCD) were re-assessed via individual phone calls. Echocardiographic studies were performed within 6 months before or after blood sampling.
Genetic testing
Genetic testing was conducted in all patients either with (1) panels that included candidate genes coding for sarcomeric HCM; or (2) segregation analysis if the familial HCM-causing variant was known. Genotype was classified as positive in case of a pathogenic ([P], class V) or likely pathogenic ([LP], class IV) variant in accordance with international criteria14.
Echocardiographic analysis
Echocardiographic analyses were performed off-line from digitally stored electrocardiogram (ECG)-gated images and loops using the post-processing software TomTec (TomTec Imaging Systems, Munich, Germany), by a blinded investigator (VS) and as described previously15,16,17,18,19. Left ventricular ejection fraction (LVEF) was assessed via Simpson´s biplane method. Extent of hypertrophy was assessed by measuring end-diastolic interventricular septum and end-diastolic posterior wall thickness (PWTH) in parasternal long axis view16,20. Maximal left ventricular wall thickness (MWTH) was defined as the thickest end-diastolic wall measurement at any segment21,22,23. Septal phenotype was classified assessing apical four-, three-, and two-chamber views, into reverse curve, sigmoid, apical, or neutral17,24. Left ventricular outflow tract obstruction (LVOTO) was defined as a peak LVOT gradient at rest ≥ 30 mmHg2,25. Myocardial deformation analysis assessing LV GLS was performed using the automatically generated auto-strain as described previously15,18. Briefly, endocardial borders were tracked automatically defining enddiastole and endsystole via ECG. If contours were not tracked correctly by the software, readjustment was performed by the blinded investigator. Peak systolic strain was assessed in apical four-, three- and two-chamber views resulting in the average peak systolic LV GLS. For simplification purposes, higher LV GLS values are described as worse, while lower LV GLS is described as better in this study.
Statistical analysis
Continuous variables were reported as medians [25–75th percentile], and count data as absolute frequencies (column%). Group comparisons were performed with Chi-Square Test, Student´s t-Test or Mann-Whitney U Test, as appropriate. Correlations between continuous variables were assessed by Spearman´s rank correlation. Associations between LV GLS and genotype were analyzed in univariate and multivariate binary regression analysis. Parameters that were associated with a positive genotype in the Toronto and Mayo Score were included in univariate analysis, and the strongest predictors in multivariate analysis4,5. These confounders included age at diagnosis, positive family history of HCM, positive family history of SCD, MWTH: PWTH ratio, reverse curve septal phenotype, and arterial hypertension. Diagnostic accuracies of both Toronto and Mayo Score (Supplementary Table S1) and the added value of LV GLS were analyzed using receiver-operating characteristics (ROC) analyses, and expressed as area under the ROC curve (AUC) with 95% confidence intervals (CI)4,5.
All statistical analyses were performed with IBM SPSS statistics version 29 and a P-value < 0.05 was considered statistically significant.
Results
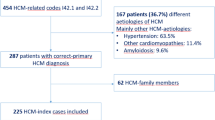
The study cohort comprised 125 patients with HCM with a mean age at diagnosis of 53 ± 16 years and 38% women. Median NT-proBNP was 543 (193, 1396) pg/ml, and median eGFR was 79 (64, 94) ml/min/1,73 m². Genotype was positive in 49 patients (39%), with 34 patients (70%) carrying a P/LP variant in MYBPC3, 10 patients (20%) in MYH7, 2 patients (4%) in MYL3, and one patient (2%) each in MYH6, TPM1 and TNNI3. Arterial hypertension was present in 83 patients (67%), dyslipidemia in 58 (47%), atrial fibrillation in 27 (22%), coronary artery disease in 18 (15%), and diabetes mellitus in 9 (7%) patients. Mean LVEF was 59 ± 7%. Median MWTH was 19.9 (18.2, 22.4) mm with a median PWTH of 11.2 (9.7, 12.6) mm. Resting LVOT gradient was available in 110 patients with a median gradient of 10 (5, 35) mmHg and 35 (32%) patients had a resting LVOT gradient ≥ 30 mmHg (Table 1).
Left ventricular global longitudinal strain
Median LV GLS was − 19.1 (-21.0, -16.0) and correlated significantly with LVEF (Spearman r=-0.560, p < 0.001), NT-proBNP (r = 0.445, p < 0.001), and Troponin T (r = 0.303, p = 0.002). Patients with prior implantable cardioverter-defibrillator (ICD) implantation had worse LV GLS values compared to those without ICD (-17.2 [-18.5, -13.1] vs. -19.4 [-21.4, -16.2], p = 0.005). LV GLS was worse in patients with compared to those without reverse curve septal phenotype (-18.1 [-20.2, -15.1] vs. -19.4 [-21.5, -16.4], p = 0.035), and was better in those with compared to those without sigmoid septal phenotype (-20.1 [-21.6, -18.1] vs. -18.1 [-20.6, -15.2], p = 0.006). There was no correlation between LV GLS and other genotype predictors as age at diagnosis, arterial hypertension, positive family history of HCM or SCD, MWTH or PWTH.
Determinants of a positive genotype
LV GLS was significantly worse in genotype-positive HCM, while LVEF did not differ between genotype-positive and genotype-negative HCM. Genotype-positive HCM patients were significantly younger at diagnosis, had higher rates of ICD implantation, more often a positive family history of HCM, and lower prevalences of LVOTO, arterial hypertension, dyslipidemia and coronary artery disease. The reverse curve septal phenotype was more prevalent in genotype-positive HCM, while the sigmoid phenotype was associated with genotype-negative HCM. MWTH did not differ between groups, while PWTH was significantly increased in genotype-negative HCM leading to a lower MWTH: PWTH ratio in those patients (Table 1).
In univariate analyses, a positive genotype was associated with worse LV GLS (OR 1.141, CI 1.018–1.279, p = 0.023), younger age at diagnosis (OR 0.947, CI 0.922–0.973, p < 0.001), the absence of arterial hypertension (OR 0.176, CI 0.078–0.396, p < 0.001), a positive family history of HCM (OR 3.378, CI 1.454–7.846, p = 0.005), a reverse curve septal phenotype (OR 18.700, CI 7.207–48.523, p < 0.001), presence of either reverse curve or neutral septal phenotype (OR 17.571, CI 5.021–61.497, p < 0.001), and a higher MWTH: PWTH ratio (OR 9.604, CI 2.858–32.279, p < 0.001), p = 0.023). Stratifying MWTH and age by evidence-based cut-offs of 20 mm and 45 years, respectively, both variables were predictive of a positive genotype (OR 2.171, CI 1.043–4.520, p = 0.038 and OR 5.797 CI 2.510–13.390, p < 0.001) (Table 2).
In multivariate analysis, significant parameters from univariate analyses as age at diagnosis < 45 years, positive family history of HCM, arterial hypertension, MWTH: PWTH ratio and a reverse curve septal phenotype were included in addition to LV GLS. In this model, a reverse curve septal phenotype remained as the only predictor of a positive genotype in HCM (OR 6.948 CI 2.342–20.614, p < 0.001) (Table 3).
When reverse curve septal phenotype was omitted from the multivariate model, family history of HCM (OR 2.876, CI 1.035–7.991, p = 0.043), arterial hypertension (OR 0.357, CI 0.128–0.990, p = 0.048), MWTH: PWTH ratio (OR 5.666, CI 1.351–23.757, p = 0.018), and age at diagnosis < 45 years (OR 3.843, CI 1.356–10.891, p = 0.011) independently predicted a positive genotype, while LV GLS did not (OR 1.141, CI 0.994–1.310, p = 0.061).
Sensitivity analyses excluding patients with prior septal reduction therapy (n = 7) yielded comparable results. The association between LV GLS and genotype remained statistically significant in univariate analysis (p = 0.028) but not in the multivariate model (p = 0.313).
Genotype-prediction scores
Both the Toronto Score (short and long model) and the Mayo Score were applied and their diagnostic accuracy assessed (Supplementary Table S2). Both scores showed excellent accuracies in predicting a positive genotype (Toronto Score: AUC 0.834, p < 0.001; Mayo Score: AUC 0.806, p < 0.001). The short model of the Toronto Score demonstrated acceptable accuracy (AUC 0.760, p < 0.001).
LV GLS was stratified by -19.0% as the value demonstrating best fit with a positive genotype. Regarding both scores, addition of LV GLS did not improve diagnostic accuracy, respectively. Within the intermediate score range between 0 and 6 points, the long model of the Toronto Score demonstrated poor diagnostic accuracy (AUC 0.530, p = 0.718) which was, however, not improved by adding LV GLS (AUC = 0.523, p = 0.784) (Supplementary Fig. 1).
Discussion
In this study, worse LV GLS was not independently associated with a positive genotype in HCM patients in univariate analysis and did not enhance information on the HCM genotype in addition to known genotype predictors. To our knowledge, this is the first study aiming to improve diagnostic accuracy of the Toronto and Mayo Score by adding LV GLS. Our results highlight the reverse curve septal phenotype as the strongest genotype predictor in HCM. It extends previous studies investigating associations between LV GLS and genotype-positive HCM in a multicenter cohort from a mid-European country.
Associations of LV GLS
LV GLS is an in-expensive and easily accessible technique, and serves well as an indicator of myocardial fibrosis both in HCM and other cardiac diseases12,26,27. In our cohort, worse LV GLS was associated with surrogates of heart failure severity, including a lower LVEF, higher NT-proBNP and higher Troponin T levels. Extensive late gadolinium enhancement (LGE) in cMRI is an additional risk factor for SCD2,28. In line with this, patients with previous ICD implantation in our study had worse LV GLS values than those without. Moreover, LV GLS was worse in patients with reverse curve septal phenotype compared to those without, and better in those with sigmoid septal phenotype compared to those without. The same associations between both reverse curve and sigmoid septal phenotype and LV GLS were described previously by Geske et al. The authors also reported associations between higher maximum wall thickness and worse LV GLS, which were not observed in our cohort11.
Genotype-positive HCM and LV GLS
There is a strong rationale linking worse LV GLS with a positive genotype in HCM. In large cohorts, genotype-positive compared to genotype-negative HCM patients had more advanced heart failure, cardiac fibrosis and a higher risk of subsequent HCM complications, including arrhythmias and heart failure3,8,9,10. In that regard, LV GLS may not only serve as an indicator of myocardial fibrosis, but also of worse cardiac function. Interestingly, in our sturdy there was no difference between genotype-positive and -negative patients regarding LVEF, NT-proBNP and Troponin T, which is in line with a previous study indicating that LV GLS delivers information on myocardial disease beyond these parameters12. Still, results from previous studies on associations between LV GLS and genotype in HCM are conflicting11,13. In 2014 Geske et al. reported worse regional strain values mid-inferoseptal, basal anteroseptal and basal inferoseptal in genotype-positive HCM, while LV GLS was similar in genotype-positive and genotype-negative HCM even in univariate analysis. They reported a younger age at diagnosis compared to our cohort (43 ± 18 vs. 53 ± 16 years), a higher number of prior ICD implantation (27% vs. 12%), more patients with positive family history of HCM (43% vs. 25%), and higher LVEF (68 ± 8 vs. 59 ± 7%), while the prevalence of positive genetic testing was identical, with 39%. In the cohort of Geske et al., one third of all patients had prior septal reduction therapy (SRT), which might have influenced regional deformation analysis (6% prior SRT in our cohort). Geske et al. associated worse regional strain values with the reverse curve septal phenotype, and better values with the sigmoid phenotype, which is similar to univariate associations with LV GLS observed in our cohort. While distribution of septal phenotypes was overall comparable in both cohorts (sigmoid 39% vs. 36%; reverse curve 33% both), genetic testing results in septal phenotype subgroups differed substantially. In our cohort, only three patients with sigmoid phenotype were genotype positive, which may partly explain the differing results in univariate analysis regarding global strain values11. In 2023 Zhang et al. reported conflicting results showing better LV global circumferential strain (LV GCS), LV global radial strain (LV GRS) and LV GLS values in genotype-positive HCM in their cohort of 117 Chinese patients. Characteristics from their cohort differ from ours with younger age at diagnosis and lesser comorbidities as arterial hypertension, diabetes mellitus and coronary artery disease. Moreover, they adjusted for known genotype predictors, but their analyses are limited by omitting septal phenotype as a covariate13.
Zhang et al. hypothesized that LV GLS might be increased in early stages of genotype-positive HCM indicating hypercontractility, and may decrease during the disease course as a marker of myocardial fibrosis and heart failure progression13. In our study and the study by Geske and colleagues, the reverse curve and sigmoid septal phenotype, as a visualization of differences in septal wall thickness at different positions of the septum, were strongly associated with LV GLS11. Our study suggests that worse LV GLS is rather a function of septal phenotype than of the underlying genotype. Integrating existing literature with our findings, evidence is likely insufficient to consider LV GLS as a valid surrogate for genotype-positive HCM.
Genotype prediction scores
According to the 2023 cardiomyopathy guidelines genetic testing is recommended in every HCM patient to enable diagnosis, prognosis, therapy and cascade genetic testing2. With growing awareness among clinicians, ubiquitously applied genetic testing is becoming a challenge to the health care system since it is costly and in Austria only available at selected genetic institutes which are constantly strained with assignments and unable to deliver reports timely. The Toronto and Mayo Score are easily applicable in a clinical setting and are useful in tailoring genetic testing4,5. Validation of both scores has been performed previously and indicates generalizability to the overall HCM populations4,5,6,7. Both scores proved to be of excellent accuracy in our cohort and can be used to support physicians in clinical decision-making. Still, diagnostic accuracy in the intermediate score group remains low and could not be improved by adding LV GLS in our study. In those patients, the decision on whether a genetic testing should be performed still remains difficult and needs to be assessed individually in clinical practice.
Limitations
The results of our study are limited by the small sample size increasing the risk of type II error. As HCM is mainly a regional myocardial disease, the lack of regional strain analysis is one of the major limitations of our study. Moreover, since cohorts were enrolled from referral centers, referral bias may preclude generalizability to the overall HCM population. Yet, baseline characteristics are overall comparable to larger HCM cohorts such as the ShaRe registry3.
Conclusions
Worse LV GLS was not independently associated with genotype-positive HCM in our study and did not improve the diagnostic accuracy of established genotype prediction scores. Our study highlights the reverse curve septal phenotype as the strongest genotype predictor in HCM. Further studies are warranted to optimize genetic testing strategies in HCM patients.
Data availability
The data underlying this article will be shared on reasonable request with the corresponding author.
References
Semsarian, C., Ingles, J., Maron, M. S. & Maron, B. J. New perspectives on the prevalence of hypertrophic cardiomyopathy. J. Am. Coll. Cardiol. 65(12), 1249–1254 (2015).
Arbelo, E. et al. 2023 ESC guidelines for the management of cardiomyopathies. Eur. Heart J. 44(37), 3503–3626 (2023).
Ho, C. Y. et al. Genotype and lifetime burden of disease in hypertrophic cardiomyopathy: insights from the sarcomeric human cardiomyopathy registry (SHaRe). Circulation 138(14), 1387–1398 (2018).
Gruner, C. et al. Toronto hypertrophic cardiomyopathy genotype score for prediction of a positive genotype in hypertrophic cardiomyopathy. Circ. Cardiovasc. Genet. 6(1), 19–26 (2013).
Bos, J. M. et al. Characterization of a phenotype-based genetic test prediction score for unrelated patients with hypertrophic cardiomyopathy. Mayo Clin Proc. ;89(6):727 – 37. (2014).
Murphy, S. L. et al. Evaluation of the Mayo clinic Phenotype-Based genotype predictor score in patients with clinically diagnosed hypertrophic cardiomyopathy. J. Cardiovasc. Transl Res. 9(2), 153–161 (2016).
Moriki, T. et al. A validation study of the Mayo clinic Phenotype-Based genetic test prediction score for Japanese patients with hypertrophic cardiomyopathy. Circ. J. 85(5), 669–674 (2021).
Neubauer, S. et al. Distinct subgroups in hypertrophic cardiomyopathy in the NHLBI HCM registry. J. Am. Coll. Cardiol. 74(19), 2333–2345 (2019).
Kim, H. Y. et al. Genotype-Related clinical characteristics and myocardial fibrosis and their association with prognosis in hypertrophic cardiomyopathy. J. Clin. Med. ;9(6), 1671 (2020).
Olivotto, I. et al. Microvascular function is selectively impaired in patients with hypertrophic cardiomyopathy and sarcomere myofilament gene mutations. J. Am. Coll. Cardiol. 58(8), 839–848 (2011).
Geske, J. B. et al. Deformation patterns in genotyped patients with hypertrophic cardiomyopathy. Eur. Heart J. Cardiovasc. Imaging. 15(4), 456–465 (2014).
Saito, M. et al. Clinical significance of global two-dimensional strain as a surrogate parameter of myocardial fibrosis and cardiac events in patients with hypertrophic cardiomyopathy. Eur. Heart J. Cardiovasc. Imaging. 13(7), 617–623 (2012).
Zhang, J. et al. Myocardial contractility characteristics of hypertrophic cardiomyopathy patients with and without sarcomere mutation. Heart Vessels. 39(11), 939–948 (2024).
Richards, S. et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American college of medical genetics and genomics and the association for molecular pathology. Genet. Med. 17(5), 405–424 (2015).
Holler, V. et al. Myocardial deformation analysis in MYBPC3 and MYH7 related sarcomeric hypertrophic cardiomyopathy-The Graz hypertrophic cardiomyopathy registry. Genes (Basel) 12(10), 1469 (2021).
Lang, R. M. et al. Recommendations for cardiac chamber quantification by echocardiography in adults: an update from the American society of echocardiography and the European association of cardiovascular imaging. Eur. Heart J. Cardiovasc. Imaging. 16(3), 233–270 (2015).
Verheyen, N. et al. Spatial relationship between mitral valve and ventricular septum assessed by resting echocardiography to diagnose left ventricular outflow tract obstruction in hypertrophic cardiomyopathy. Eur. Heart J. Cardiovasc. Imaging. 24(6), 710–718 (2023).
Zach, D. K. et al. Low-grade systemic inflammation and left ventricular dysfunction in hypertensive compared to non-hypertensive hypertrophic cardiomyopathy. Int. J. Cardiol. 399, 131661 (2024).
Mor-Avi, V. et al. Current and evolving echocardiographic techniques for the quantitative evaluation of cardiac mechanics: ASE/EAE consensus statement on methodology and indications endorsed by the Japanese society of echocardiography. J. Am. Soc. Echocardiogr. 24(3), 277–313 (2011).
Lang, R. M. et al. Recommendations for chamber quantification: a report from the American society of echocardiography’s guidelines and standards committee and the chamber quantification writing group, developed in conjunction with the European association of echocardiography, a branch of the European society of cardiology. J. Am. Soc. Echocardiogr. 18(12), 1440–1463 (2005).
Losi, M. A. et al. Echocardiography in patients with hypertrophic cardiomyopathy: usefulness of old and new techniques in the diagnosis and pathophysiological assessment. Cardiovasc. Ultrasound. 8, 7 (2010).
Wigle, E. D. et al. Hypertrophic cardiomyopathy. The importance of the site and the extent of hypertrophy. A review. Prog Cardiovasc. Dis. 28(1), 1–83 (1985).
Klues, H. G., Schiffers, A. & Maron, B. J. Phenotypic spectrum and patterns of left ventricular hypertrophy in hypertrophic cardiomyopathy: morphologic observations and significance as assessed by two-dimensional echocardiography in 600 patients. J. Am. Coll. Cardiol. 26(7), 1699–1708 (1995).
Binder, J. et al. Echocardiography-guided genetic testing in hypertrophic cardiomyopathy: septal morphological features predict the presence of myofilament mutations. Mayo Clin Proc. ;81(4):459 – 67. (2006).
Verheyen, N. et al. Austrian consensus statement on the diagnosis and management of hypertrophic cardiomyopathy. Wien Klin. Wochenschr. 136(Suppl 15), 571–597 (2024).
Cameli, M. et al. Left ventricular deformation and myocardial fibrosis in patients with advanced heart failure requiring transplantation. J. Card Fail. 22(11), 901–907 (2016).
Balciunaite, G. et al. Exploring myocardial fibrosis in severe aortic stenosis: echo, CMR and histology data from FIB-AS study. Int. J. Cardiovasc. Imaging. 38(7), 1555–1568 (2022).
Weng, Z. et al. Prognostic value of LGE-CMR in HCM: A Meta-Analysis. JACC Cardiovasc. Imaging. 9(12), 1392–1402 (2016).
Acknowledgements
We thank all our patients who consented to participate in this study.
Funding
The Graz HCM Registry was supported by unrestricted research grants from Pfizer, Bristol-Myers Squibb, Astra Zeneca, Takeda Pharmaceuticals, Boehringer Ingelheim, and Swedish Orphan Biovitrum AB. Dr. Santner was supported by the Austrian Society of Cardiology (Project-related Grant 2024).
Author information
Authors and Affiliations
Contributions
V.S. and N.V. wrote the main manuscript text. V.S., N.V., V.H., M.U., G.P., G.W., E.H., M.W. and K.A. developed the study design and defined the methodological implementation. V.H., N.S., D.Z., H.T., R.B., J.G. and E.K. performed data acquisition and analysis. V.S., N.V., P.P.R., D.vL., S.R-S. and A.Z. performed analysis and interpretation of the data. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Consent to participate
Informed consent was obtained from all individual participants included in the study.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Santner, V., Höller, V., Ungericht, M. et al. Left ventricular global longitudinal strain and diagnostic yield of genetic testing in hypertrophic cardiomyopathy in a multicenter registry analysis. Sci Rep 15, 22573 (2025). https://doi.org/10.1038/s41598-025-05696-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-05696-5