Abstract
Acrylamide (ACR) is a common food and environmental contaminant with potential carcinogenicity, but its molecular role in the development of breast cancer (BC) remains unclear. This study employed a network toxicology and molecular docking strategy to explore the potential association between ACR exposure and BC. Differentially expressed genes were first identified by comparing gene expression profiles between breast cancer and normal tissues. BC-related targets were then screened through integrated analysis of multiple databases and intersected with ACR-associated targets, resulting in 49 shared genes. Protein–protein interaction network construction and MCC algorithm analysis identified 10 core genes. GO and KEGG enrichment analyses revealed that these genes were mainly involved in pathways such as PI3K-Akt signaling, HIF-1 signaling, and ECM–receptor interactions. Molecular docking results suggested that ACR could bind to key targets including EGFR, FN1, JUN, and COL1A1. Molecular dynamics simulations demonstrated that the complexes remained structurally stable throughout the simulation. Pan-cancer and paired sample analyses further confirmed the aberrant expression of core genes in BC, and immunohistochemistry results supported their altered protein levels. These findings suggest that ACR may interfere with BC-related molecular mechanisms through multi-target and multi-pathway interactions, providing theoretical evidence for its potential carcinogenic role and future risk assessment.
Similar content being viewed by others
Introduction
Breast cancer (BC) stands as the leading form of cancer affecting women globally. Alarming trends show escalating incidences and mortality rates in both developing and developed nations alike1. Early symptoms of BC typically include palpable masses in the breast, aenlarged axillary nodes, and nipple discharge. During advanced phases, malignant cells can disseminate to lymph nodes, bones, hepatic tissue, and additional bodily systems. In China, BC ranks first in both incidence and mortality among malignant diseases in women. Notwithstanding advances in treatments, numerous women still fall victim to the illness, underscoring its status as one of the most widespread malignancies impacting women on a global scale2,3. BC is influenced by various factors, with 5–10% attributed to hereditary conditions, whereas the residual 90–95% can be traced back to environmental influences and lifestyle decisions, such as prolonged exposure to chemicals, diet, obesity, hormone therapy, and ionizing radiation4. Evidence suggests a positive link exists between exposure of acrylamide (ACR) and heightened chances of developing BC, with this correlation appearing more distinct in premenopausal women5.
ACR is a odorless, clear, highly soluble vinyl monomer, arises as a product of the acrylonitrile hydration reaction. In its monomeric form, ACR is a toxic compound with potential carcinogenic, neurotoxic, genotoxic, as well as reproductive and developmental toxic effects6,7. Present in a broad spectrum of industries like cosmetics, horticultural products, paper and pulp products, laboratory gel electrophoresis, coatings, and textiles, ACR has also made its presence known in the component list of cigarette smoke8. Research indicates that ACR along with its related compounds function as flocculants, capable of persisting within irrigation systems and drinking water supplies. Although traditional water treatment processes can remove ACR from environmental water, its high solubility allows residual amounts to persist in the water9. Investigations reveal that carbohydrate-rich, protein-poor plant-based foods are susceptible to producing considerable quantities of ACR when subjected to high-temperature processes exceeding 120 °C10. A notable concern of ACR exposure arises from cereal-based foods including crackers, cookies, coffee, breakfast cereals, bread, and particularly potato products like chips and french fries11. Large-scale human biomonitoring studies, such as those conducted in Taiwanese populations, have demonstrated that ACR exposure remains prevalent in the general population and is closely associated with dietary habits, including sweetened beverage and fried snack consumption12. In China, the estimated annual cancer risk attributable to dietary ACR exposure is 1.78 cases per 100,000 people, with a disease burden of approximately 157 disability-adjusted life years (DALYs). The average daily dietary ACR exposure has been estimated at 0.1531 μg/kg for males and 0.1554 μg/kg for females, corresponding to an annual cancer burden exceeding 23,000 DALYs, indicating that ACR exposure poses a significant public health risk in large populations, potentially comparable to or exceeding that observed in Europe and the United States13,14. Animal trials have illustrated a direct correlation between ACR exposure and tumor growth across numerous body parts, especially in hormone-sensitive tissues like the uterus and breast glands. For instance, feeding ACR orally to test subjects has proven to instigate cancerous developments, specifically leading to BC in female rats during animal experiments15,16. Consequently, the long-term health impacts and adverse outcomes of ACR exposure have garnered widespread attention.
Network toxicology (NT), an amalgamation of multiple disciplines, encompasses related technologies, genomics, big data analysis, and bioinformatics. At its core, it aims to scrutinize the toxicological routes of substances and deciphering the molecular dynamics linked to pathogenic processes. The application of NT in examining toxicological molecular mechanisms stands out due to its velocity, ease-of-use, broad coverage, and integrative viewpoint. Evaluating toxic effects within a large network context helps broaden the scope of toxicological research17,18. Molecular docking (MD) represents a computational method employed to forecast and scrutinize the binding interactions and affinities between ligands and their targets at the molecular scale. There’s an inverse relationship between the thermodynamic steadiness of the ligand-receptor conjugate and the binding free energy, whereby lesser values denote greater stability and energetic preference for the ligand-receptor pairing19. Based on this, the present study integrates novel analytical strategies to explore the complex relationship between ACR exposure and BC pathogenesis. This offers significant enlightenment for the formulation of efficacious preventive actions and risk mitigation tactics for BC.
Methods
BC gene chip and differentially expressed gene screening
The GEO database (https://www.ncbi.nlm.nih.gov/geo/) was searched using the keyword “Breast cancer” with the species set to “Homo sapiens” and gene expression datasets with a sample size of ≥ 20 selected. Gene expression data were obtained using the GEOquery package in R, and normalization was performed using the normalizeBetweenArrays function from the limma package to minimize batch effects and improve data consistency. Sample grouping information was extracted from the dataset annotation files, and probe annotation was performed using official Bioconductor annotation packages to map probe IDs to gene symbols. Differentially expressed genes were identified based on thresholds of absolute log2 fold change (|logFC|> 0.585) and p value < 0.05. In this study, the GSE15852 dataset from the GEO database was used, which includes 43 breast cancer tissue samples and 43 paired normal breast tissue samples analyzed on the Affymetrix Human Genome U133A Array (GPL96 platform). This dataset represents the first gene expression profiling study of multi-ethnic Malaysian breast cancer patients (Malays, Chinese and Indian), aimed at exploring transcriptomic changes associated with the transition from normal cells to malignant BC cells.
Screening of potential gene targets for BC
The GeneCards database (https://www.genecards.org/) was used to retrieve gene targets associated with BC. The retrieved targets were standardized and deduplicated to construct a library of potential BC-related gene targets.
Download and acquisition of ACR data
The potential targets of ACR were retrieved and downloaded from the GeneCards, SwissTargetPrediction (http://swisstargetprediction.ch), and TTD (https://db.idrblab.net/ttd/) databases. The union of all retrieved targets was taken, and duplicate entries were removed to construct a library of ACR-related potential targets.
Identification of common targets for ACR and BC
The obtained ACR-related and BC-related targets were analyzed using the online bioinformatics platform (http://www.bioinformatics.com.cn) to construct a Venn diagram of intersecting targets. The overlapping targets identified in the Venn diagram were considered the common targets of ACR and BC.
Construction of protein–protein interaction network and screening of core targets
The list of common targets was uploaded to the STRING database (https://cn.string-db.org/) to construct a protein–protein interaction (PPI) network of the shared targets. Cytoscape 3.7.1 software was used for visualizing and analyzing the PPI network, where each node represents a target gene and the edges indicate interactions between biomolecules. The connectivity of each node in the network was evaluated by calculating node parameters. The CytoHubba plugin with the MCC algorithm was applied to analyze the network and identify core targets.
GO functional annotation and KEGG pathway enrichment analysis
To improve the identification of relevant targets and enhance the understanding of the mechanisms by which ACR promotes BC development, the intersecting targets were submitted to the DAVID database (https://david.ncifcrf.gov/) for functional enrichment analysis. The analysis was performed with “Homo sapiens” as the selected species and a significance threshold of P < 0.05. Gene Ontology (GO) annotation included biological processes (BP), cellular components (CC), and molecular functions (MF). KEGG pathway enrichment analysis was also conducted, and the results were interpreted based on KEGG pathway definitions20,21,22. All enrichment results were visualized using an online bioinformatics platform.
Molecular docking and molecular dynamics simulation
The three-dimensional structure of ACR was obtained from the PubChem database, and the crystal structures of the target proteins were retrieved from the RCSB Protein Data Bank (https://www.pdbus.org/). Prior to docking, water molecules and original ligands were removed from the protein structures using PyMOL. The ligand and receptor files were then converted into PDBQT format using AutoDock Tools 1.5.6. Molecular docking was performed using AutoDock 4.2.6, and the docking conformation with the lowest binding energy was selected as the optimal binding pose. The docking results were visualized using PyMOL. To further evaluate the stability of the protein–ligand complexes, molecular dynamics (MD) simulations were conducted using GROMACS 5.1.4. The Amber-ff99SB force field was applied to the protein, and the topology parameters for the ligand were generated using the Tleap module of the AmberTools20 package, also based on the Amber-ff99SB force field. The TIP3P water model was used for solvation. Each complex system underwent a 200-ns MD simulation. After the simulation, the root mean square deviation (RMSD) values of the heavy atoms in the complexes were calculated, and RMSD-time fluctuation curves were plotted to assess the structural stability of the docked complexes during the simulation.
Expression analysis of key targets in various cancers
An online bioinformatics platform was used to compare the overall expression distribution of four core targets (EGFR, FN1, JUN, COL1A1) across 33 cancer types relative to normal samples. Statistical differences between tumor and normal groups were assessed. For patients with paired samples, further analysis was conducted to examine the expression levels of core targets in tumor tissues compared to their corresponding normal tissues, highlighting differential expression patterns between tumors and normal tissues.
Expression analysis and validation of key targets in BC
The expression levels of the four core targets in BC tissues and normal tissues were analyzed using the GEPIA database (http://gepia.cancer-pku.cn/), which integrates data from TCGA and GTEx databases. The analysis parameters were set as |log FC|= 1 and p-value = 0.01. Boxplots were generated, exported, and saved for further analysis. Additionally, the Human Protein Atlas (HPA) database was utilized to verify the expression of the four core target markers in BC and normal tissues through immunohistochemistry results.
Results
Processing of GSE15852 dataset and identification of differentially expressed genes
The raw expression matrix of the GSE15852 dataset was imported and processed using R. The data were first log2-transformed and normalized. Density plot assessment showed that the expression distributions across samples were relatively consistent, with similar median levels, indicating good data quality suitable for further analysis (Fig. 1A). Differential expression analysis was performed using the limma package with thresholds of p value < 0.05 and |logFC|> 0.585. A total of 792 differentially expressed genes (DEGs) were identified, including 430 upregulated and 362 downregulated genes in breast cancer tissues. A volcano plot was generated using the ggplot2 package to visualize the overall distribution of DEGs (Fig. 1B), and a heatmap of the top 100 DEGs was plotted to illustrate the clustering patterns among different samples (Fig. 1C).
Data preprocessing and differential gene screening of GSE15852. (A) Violin plot showing the distribution of gene expression levels between the normal and cancer groups after data normalization. (B) Volcano plot of significantly differentially expressed genes (DEGs); each point represents a gene, with red indicating upregulated genes, blue indicating downregulated genes, and gray representing non-significantly changed genes. (C) Heatmap of the top 100 DEGs; hierarchical clustering was performed with samples grouped into normal and cancer categories to enhance intra-group aggregation and improve visualization clarity.
Results of BC and ACR target identification
The potential targets of ACR were screened using the GeneCards, SwissTargetPrediction, and TTD databases. The screening results were consolidated, and duplicates were removed, resulting in a final set of 597 potential ACR-related targets. Additionally, 14,193 potential targets for BC were obtained from the GeneCards database.
Identification of common targets for BC and ACR
A Venn diagram of the intersecting targets between ACR and BC was constructed using an online bioinformatics platform. A total of 49 intersecting targets were identified (Fig. 2), representing the common targets of ACR and BC.
Venn diagram of the targets of ACR and BC.
Construction of protein–protein interaction network and target screening results
The 49 intersecting targets were uploaded to the STRING database (species: Homo sapiens), and a protein–protein interaction (PPI) network was constructed by hiding free nodes. The network, consisting of 45 nodes and 176 edges, was visualized using Cytoscape 3.7.1. Node color depth reflects degree centrality (Fig. 3A). To identify potential hub targets, the MCC algorithm in the CytoHubba plugin was applied, and the top 10 core genes were extracted (Fig. 3B, Table 1), including fibronectin (FN1), fibroblast growth factor 2 (FGF2), epidermal growth factor receptor (EGFR), collagen alpha-1(I) chain (COL1A1), vimentin (VIM), twist-related protein 1(TWIST1), osteopontin (SPP1), Transcription factor SOX-9 (SOX9), transcription factor jun (JUN), and protein c-Fos (FOS). To further interpret their functional significance, we performed KEGG pathway enrichment analysis, as shown in Fig. 6, which clearly illustrates the involvement of these hub genes in multiple classical tumor-related signaling pathways. Together, Figs. 3 and 6 provide a structural–functional framework for understanding the toxicological mechanisms of acrylamide exposure in BC.
Intersection target PPI network interaction and target screening. (A) Visualization of the PPI network constructed from the 49 intersecting differentially expressed genes. Node color intensity indicates degree centrality, with darker colors representing higher interaction connectivity. (B) Identification of the top 10 hub genes using the CytoHubba plugin based on the MCC algorithm. These hub genes were further analyzed for their involvement in classical tumor-related pathways.
Results of GO functional and KEGG pathway enrichment analyses
Using the DAVID database, GO enrichment analysis was conducted on the cohort of 49 intersecting targets. A total of 114 biological process (BP) terms were significantly enriched, mainly involving the positive regulation of phosphatidylinositol 3-kinase/protein kinase B (PI3K-Akt) signaling transduction, cellular response to mechanical stimulus, and positive regulation of miRNA transcription. For cellular component (CC) analysis, 36 terms were identified, predominantly associated with the extracellular region, extracellular space, and extracellular exosome. Molecular function (MF) analysis revealed 29 enriched terms, primarily related to collagen binding, identical protein binding, and general protein binding. The top 10 significantly enriched entries from each GO category were visualized based on enrichment scores and gene counts. These enriched biological functions are closely associated with key pathological processes in BC development, including microenvironment remodeling, aberrant signal transduction, and dysregulation of transcription (Fig. 4A, B). KEGG pathway enrichment analysis identified 38 significantly enriched pathways, with the top 10 visualized based on P-value rankings. Notably, these included the HIF-1 signaling pathway, relaxin signaling pathway, and PI3K-Akt signaling pathway, which are closely associated with angiogenesis, extracellular matrix remodeling, and tumor cell proliferation in BC. These findings suggest that ACR exposure may influence BC progression through the dysregulation of multiple oncogenic signaling pathways (Fig. 5A, B).
GO functional annotation analysis of intersection targets. (A) Bar charts showing the top enriched GO terms based on enrichment score across three categories: BP, CC, and MF. The height of each bar reflects the enrichment score, with higher scores indicating more significant enrichment. (B) Bar charts showing the number of genes enriched in each GO term. The color gradient represents the −log10(p-value), with darker colors indicating higher statistical significance. These enriched biological functions are potentially associated with the mechanisms of ACR exposure in BC.
KEGG pathway enrichment analysis of intersection targets (Top 10). (A) Bubble plot of the top 10 significantly enriched KEGG signaling pathways. Each bubble represents a specific pathway, where the bubble size indicates the number of enriched genes and the color scale reflects the statistical significance. (B) Bar plot showing the number of genes enriched in each pathway, categorized by biological function. Pathways are grouped into Environmental Information Processing, Cellular Processes, Organismal Systems, and Human Diseases, providing insight into the possible mechanisms by which ACR exposure may contribute to BC progression. Pathway information was obtained from the KEGG database (www.kegg.jp/kegg/kegg1.html).
Pathway enrichment analysis of core targets
Pathway enrichment analysis was performed for the 10 core targets identified via the MCC algorithm. Using the R packages ggplot2 and reshape2, we generated a horizontal gradient bar chart to visualize the enrichment profile. In this chart, the length of each bar represents the relative enrichment significance of each KEGG pathway, while the presence of orange squares indicates the involvement of core targets within these pathways (Fig. 6). Notably, EGFR, FN1, JUN, and COL1A1 were frequently enriched across all major pathways, suggesting that these genes may serve as key regulatory hubs in ACR-induced BC progression.
KEGG enrichment analysis of intersection targets (Top 10) overlapping gene horizontal gradient bar chart. (A) KEGG pathway enrichment analysis of all differentially expressed genes, with the top 10 significantly enriched pathways displayed. The bar length represents − log10(P-value), indicating the statistical significance of enrichment. (B) Heatmap showing the distribution of the 10 hub genes (identified via the MCC algorithm) across the top enriched KEGG pathways. Orange squares indicate that the gene is involved in a given pathway, with the color intensity reflecting the enrichment level. The frequent presence of EGFR, FN1, JUN, and COL1A1 across multiple pathways suggests their central regulatory role in BC progression.
Molecular docking and molecular dynamics simulation results
Molecular docking analysis was conducted to evaluate the binding affinity between ACR and the core target proteins. The results showed that ACR formed hydrogen bonds with key amino acid residues of multiple targets, with all binding free energies less than 0 kcal/mol, indicating spontaneous binding to the proteins. Specifically, ACR interacted with EGFR at residues Asp855 and Lys745 with a binding energy of − 3.16 kcal/mol; with FN1 at residues Tyr1573 and Ser1597 with a binding energy of − 3.14 kcal/mol; with JUN at residues Arg270 and Lys273 with a binding energy of − 3.01 kcal/mol; and with COL1A1 at residues Asn1372 and Thr1419 with a binding energy of − 4.43 kcal/mol (Table 2). Furthermore, molecular dynamics simulations were performed to assess the binding stability of the four ACR–protein complexes. The results demonstrated that all complexes maintained low and relatively stable RMSD values throughout the 200 ns simulation, indicating that ACR remained stably bound to the target proteins under dynamic conditions, thereby supporting the reliability of the molecular docking results (Fig. 7). Overall, these findings suggest that ACR can spontaneously bind and stably interact with the core targets, supporting its potential role in modulating related signaling pathways.
Molecular docking and molecular dynamics simulation validation of core targets with ACR. (A–D) The molecular docking poses of ACR with EGFR, FN1, JUN, and COL1A1, respectively. Left panels show the overall structure of the protein–ligand complex; middle panels provide close-up views of ACR binding at the active sites; right panels show the interaction diagrams highlighting key amino acid residues forming hydrogen bonds with ACR. (E–H) The RMSD trajectories of the four protein–ligand complexes during 200 ns molecular dynamics simulations, indicating the stability of the complexes over time. The low and relatively stable RMSD values suggest that ACR forms stable interactions with the target proteins under dynamic conditions.
Expression analysis of key targets across multiple cancer types
Expression levels of the four core targets were analyzed across 33 cancer types, comparing their overall expression distributions between tumor and normal samples. Boxplots were generated to visualize the results. EGFR expression was downregulated in seven cancer types, including BC, and upregulated in four types. FN1 expression was downregulated in three cancer types and upregulated in 10 types, including BC. JUN expression was downregulated in 11 cancer types, including BC, and upregulated in two types. COL1A1 expression was upregulated in 15 cancer types, including BC.
Further analysis was conducted on 15 cancer types with paired tumor and normal samples to assess changes in core target expression. Results showed that EGFR was downregulated in three cancer types, including BC; FN1 was upregulated in seven cancer types, including BC; JUN was downregulated in eight cancer types, including BC; and COL1A1 was upregulated in 10 cancer types, including BC (Supplementary Fig. 1, 2).
Expression analysis and validation of key targets in BC
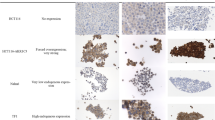
To further validate the expression of core targets in BC, their expression levels were analyzed using the GEPIA database. Significant differences were observed in the expression levels of the four core genes between primary tumor samples and normal tissue samples. Notably, EGFR and JUN were downregulated in tumor tissues compared to normal tissues, while FN1 and COL1A1 were significantly upregulated in tumor tissues (Fig. 8A), consistent with the previous analysis results. To confirm these observations at the protein level, immunohistochemistry results from the HPA were examined. As shown in Fig. 8B, EGFR and JUN exhibited lower staining intensity in tumor tissues, whereas FN1 and COL1A1 showed enhanced staining in tumor samples. These consistent transcriptomic and proteomic findings reinforce the potential regulatory significance of these targets in BC pathogenesis.
Expression analysis and immunohistochemical validation of core targets in BC. (A) Differential expression analysis of EGFR, FN1, JUN, and COL1A1 between normal and tumor tissues in BC, based on data from the GEPIA database. The y-axis represents log2-transformed transcript per million (TPM + 1) values. (B) Immunohistochemical staining results of the four core proteins in normal and breast tumor tissues, obtained from the HPA database. The protein-level expression trends were consistent with transcriptomic data, supporting their potential functional roles in BC progression. *P < 0.05 represents the comparison between the BC group and the normal group.
Discussion
Labeled as a potential human carcinogen within IARC’s Group 2A category since 1994, ACR—an organic soluble compound—has been determined to be possibly carcinogenic to humans based on substantial data gathered from animal experiments. However, evidence from human studies remains limited23. The predominant sources of human exposure to ACR include occupational exposure via skin contact and inhalation during its industrial processing, along with cigarette smoking, both considered principal means of ACR uptake. Emerging scientific evidence suggests that ACR forms as a dietary contaminant during high-heat preparation of various regular consumables. It is primarily generated as a byproduct of the Maillard reaction process, occurring under circumstances featuring high temperature and limited moisture conditions, using raw foods rich in reducing sugars and amino acids as precursors13,24. Remarkably, the disruptive impacts of ACR on hormonal balance, especially in hormone-sensitive regions such as the mammary glands, might elucidate its carcinogenic nature specifically in females, highlighting its role in breast cancer development25. Considering the significant toll of BC, exploring the link between human exposure to ACR and the risk of developing this disease holds considerable importance. A subgroup analysis limited to premenopausal women demonstrated a linear increase in BC risk with higher ACR intake, which also showed a relatively linear relationship with increased risks of endometrial and ovarian cancers26. However, the results of epidemiological studies remain mixed. Nonetheless, explicating the plausible biological underpinnings of how ACR contributes to BC remains essential for crafting exposure mitigation tactics and preventative measures. Against this backdrop, this study aims to leverage network toxicology and molecular docking techniques to clarify the molecular pathogenesis of ACR in BC.
Toxicity of compounds plays a pivotal role in gauging their ecological dangers. With the advancement of modern network technologies and the enrichment of toxicity datasets, network toxicology compared to network pharmacology focuses on constructing targeted network models of toxicological properties. This approach facilitates the prediction of the toxicity of compounds or toxic materials and their potential mechanisms of toxic action18,27,28. Renowned as a premier and extensively-utilized computational strategy grounded in molecular structure, molecular docking excels in forecasting interactions among biomolecules by initially pinpointing the ligand-receptor binding pose and then evaluating their compatibility through specialized scoring techniques29,30. Within this investigation, we harnessed the capabilities of network toxicology integrated with molecular docking methodologies to dissect and interpret the dataset comprehensively. Using a Venn diagram, 49 potential gene targets potentially associated with ACR’s impact on BC development were identified. Subsequently, 10 key potential targets related to ACR-induced BC were screened using the PPI network. Among these, EGFR, FN1, JUN, and COL1A1 were found to play central roles in pathway enrichment, highlighting their critical contributions. We analyzed the expression of four key targets across 33 cancer types and 15 paired cancer types. The results showed significant differences in the expression levels of these core targets between BC and normal tissue samples. Specifically, EGFR and JUN were significantly downregulated in BC, while FN1 and COL1A1 were significantly upregulated. Further validation through GEPIA and HPA databases using immunohistochemical results revealed consistent findings, indicating that these core targets play important regulatory roles in ACR-induced BC development.
EGFR, an integral component of the ErbB family of the receptor tyrosine kinase (RTKs), exerts a pivotal influence on the physiological processes of epithelial cells. It is involved in various pathological processes in multiple cancers, including programmed cell death, oncogenic transformation, cellular proliferation, therapeutic resistance, and metastatic dissemination31,32. Fibronectin (FN), a principal constituent of the extracellular matrix (ECM), is intricately linked to the metabolic reconfiguration of BC cells. This remodeling process significantly influences the metabolic diversity of cancer cells, which is strongly correlated with their invasive and metastatic capabilities33. FN1, as a direct participant in the assembly of the ECM, exerts a significant influence over tumor cell adherence, expansion, mobility, and invasive behavior34. A growing body of evidence corroborates the strong link between aberrant expression of FN1 and critical facets of malignancy, including stage progression, patient prognosis, and treatment efficacy in colorectal cancer (CRC) and ovarian cancer35,36,37. c-Jun is a protein product of the JUN proto-oncogene, a cellular counterpart to the human-derived transforming viral oncogene v-JUN. Its activity is governed by post-translational modifications and plays a pervasive role in critical cellular processes such as cell differentiation, proliferation, apoptosis, growth, migration, and transformation38. In human BC cells, excessive production of c-Jun catalyzes carcinogenesis and fosters resistance to hormonal therapies. The COL1A1 gene encodes the pro-α1 chain of type I collagen, the primary component of type I collagen, which is widely distributed in the stroma of parenchymal organs and connective tissues. By promoting extracellular matrix remodeling, COL1A1 enhances tumor cell migration, proliferation, and resistance to apoptosis, playing a critical role in the progression and development of various cancers39. Its overexpression is closely associated with poor prognosis in multiple cancers, making it a potential target for cancer diagnosis, treatment, and prognostic evaluation. By modulating the expression of stem cell factor and C–C chemokine ligand 5, it spurs the invasion of breast epithelial cells and the multiplication of BC stem cells40. Remarkably, the findings from molecular docking analyses revealed that ACR exhibits potent binding affinity towards all five proposed critical gene targets, forming stable hydrogen bonds with the amino acid residues of each protein ligand. The specific bond lengths formed play a critical role in stabilizing these interactions. Interestingly, these four prospective pivotal gene targets are intricately involved in orchestrating vital cellular functions, including proliferation, survival, differentiation, and metastasis. This suggests that they could serve as potential biomarkers for preventing or treating ACR-induced BC, further reinforcing their relevance as potential targets for preventive or therapeutic interventions.
An exhaustive assessment utilizing GO functional annotation coupled with KEGG pathway enrichment was carried out for the set of 49 potential gene targeted by ACR affecting BC evolution. A total of 179 GO functional annotation entries and 38 KEGG pathway entries were enriched. Notably, the KEGG enrichment results revealed significant enrichment in pathways such as breast cancer, cancer pathways, the PI3K-Akt signaling pathway, and the HIF-1 signaling pathway. Under hypoxic environments, HIF-1 serves as a resilient transcription factor, consisting of HIF-1α and HIF-1β subunits. This complex activates the transcription of an extensive array of genes engaged in neovascularization, migration, invasion, extracellular matrix remodeling, and metastasis, all pivotal stages critical for both tumor advancement and distant spread41,42. Research has indicated that HIF-1α is excessively expressed in human breast malignancies, encompassing ductal carcinoma in situ (DCIS) and early-stage BC, and is closely associated with tumor grade and invasion43. The PI3K-Akt signaling cascade is implicated in diverse cellular operations, and its aberrant activation frequently drives uncontrolled cell proliferation and hinders programmed cell death. This pathway’s deregulation is intimately tied to the onset and progression of a multitude of cancer forms44. Research indicates that the misregulation of the PI3K-Akt signaling pathway is strikingly prevalent in BC, acting as a pivotal hub within multiple interconnected signaling networks. Its abnormal activation plays a critical role in cell survival, proliferation, growth, and metabolism45. These findings suggest that ACR exposure may promote BC development by directly regulating BC-specific pathways and activating key target genes, disrupting the balance of normal cellular survival, proliferation, and apoptosis. This ultimately leads to carcinogenesis, potentially involving cross-regulation through multiple pathways and targets.
Our investigation employed a network toxicology combined with molecular docking methodologies to decipher the prospective mechanisms whereby ACR instigates BC via multi-target and intricate pathway interplay. The findings indicate that ACR significantly affects key targets closely associated with tumor growth and invasion (EGFR, FN1, JUN, and COL1A1) and may disrupt normal cellular functions by activating signaling pathways such as HIF-1 and PI3K-Akt. Grounded on these outcomes, minimizing exposure to ACR through diet and environment can notably reduce the likelihood of developing BC, simultaneously presenting fresh targets and complementary avenues for current treatment methodologies. However, to further solidify the relationship between ACR exposure and BC development, longitudinal epidemiological studies are needed to track the impact of long-term exposure on disease progression and provide clearer causal evidence. Additionally, it is essential to prioritize the evaluation of population-specific sensitivities to ACR exposure, including premenopausal and postmenopausal women, to develop more targeted prevention strategies. Furthermore, enhancing food processing methods to curtail ACR generation and enforcing more stringent regulations ought to be cardinal concerns in the realm of public health policy. Through multidisciplinary collaboration, exploring practical pathways and intervention measures to reduce ACR exposure will be of great significance in alleviating the burden of cancer.
Conclusions
In summation, our study initially unveiled the probable connection between ACR exposure and the onset of BC, offering a novel scientific viewpoint for exploring the health hazards posed by environmental contaminants. Through network toxicology analysis and molecular docking techniques, we systematically analyzed how ACR might promote BC occurrence and progression by interfering with processes such as proliferation, apoptosis, cell growth, and metabolism via multi-pathway and multi-target synergistic effects. The findings highlight the critical importance of reassessing environmental exposure to ACR and its potential health impacts while offering a scientific basis for developing future risk control and prevention strategies. However, this study has certain limitations. Primarily anchored in bioinformatic data analysis, this approach lacks empirical corroboration from in vivo or in vitro trials. The identified associations warrant additional substantiation via cell-based or animal model experiments. Additionally, due to the limitations in the accuracy and coverage of the databases used, some potential targets or pathways may not have been fully captured, which could impact the comprehensiveness and generalizability of the results. Therefore, future research should focus on integrating experimental validation with multiple technological approaches to deepen our understanding of ACR’s carcinogenic mechanisms.
Data availability
Data is provided within the manuscript or supplementary information files.
References
Nagahashi, M. & Miyoshi, Y. Targeting sphingosine-1-phosphate signaling in breast cancer. Int. J. Mol. Sci. https://doi.org/10.3390/ijms25063354 (2024).
Wang, X. et al. Progress of breast cancer basic research in China. Int. J. Biol. Sci. 17, 2069–2079. https://doi.org/10.7150/ijbs.60631 (2021).
Sun, M. et al. The potential mechanism of liujunzi decoction in the treatment of breast cancer based on network pharmacology and molecular docking technology. Curr. Pharm. Des. 30, 702–726. https://doi.org/10.2174/0113816128289900240219104854 (2024).
Kolak, A. et al. Primary and secondary prevention of breast cancer. Ann. Agric. Environ. Med. 24, 549–553. https://doi.org/10.26444/aaem/75943 (2017).
Pelucchi, C., Bosetti, C., Galeone, C. & La Vecchia, C. Dietary acrylamide and cancer risk: An updated meta-analysis. Int. J. Cancer 136, 2912–2922. https://doi.org/10.1002/ijc.29339 (2014).
Fan, M. et al. Toxicity, formation, contamination, determination and mitigation of acrylamide in thermally processed plant-based foods and herbal medicines: A review. Ecotoxicol. Environ. Saf. 260, 115059. https://doi.org/10.1016/j.ecoenv.2023.115059 (2023).
Xie, M., Lv, X., Wang, K., Zhou, Y. & Lin, X. Advancements in chemical and biosensors for point-of-care detection of acrylamide. Sensors. https://doi.org/10.3390/s24113501 (2024).
Kunnel, S. G., Subramanya, S., Satapathy, P., Sahoo, I. & Zameer, F. Acrylamide induced toxicity and the propensity of phytochemicals in amelioration: A review. Cent. Nerv. Syst. Agents Med. Chem. 19, 100–113. https://doi.org/10.2174/1871524919666190207160236 (2019).
Koszucka, A., Nowak, A., Nowak, I. & Motyl, I. Acrylamide in human diet, its metabolism, toxicity, inactivation and the associated European Union legal regulations in food industry. Crit. Rev. Food Sci. Nutr. 60, 1677–1692. https://doi.org/10.1080/10408398.2019.1588222 (2019).
Shipp, A. et al. Acrylamide: Review of toxicity data and dose–response analyses for cancer and noncancer effects. Crit. Rev. Toxicol. 36, 481–608. https://doi.org/10.1080/10408440600851377 (2008).
Sansano, M., Heredia, A., Peinado, I. & Andrés, A. Dietary acrylamide: what happens during digestion. Food Chem. 237, 58–64. https://doi.org/10.1016/j.foodchem.2017.05.104 (2017).
Liao, K.-W. et al. Associating acrylamide internal exposure with dietary pattern and health risk in the general population of Taiwan. Food Chem. https://doi.org/10.1016/j.foodchem.2021.131653 (2022).
Li, Y., Liu, J., Wang, Y. & Wei, S. Cancer risk and disease burden of dietary acrylamide exposure in China, 2016. Ecotoxicol. Environ. Saf. https://doi.org/10.1016/j.ecoenv.2022.113551 (2022).
Lin, Y.-S., Morozov, V., Kadry, A.-R., Caffrey, J. L. & Chou, W.-C. Reconstructing population exposures to acrylamide from human monitoring data using a pharmacokinetic framework. Chemosphere. https://doi.org/10.1016/j.chemosphere.2023.138798 (2023).
Riboldi, B. P., Vinhas, Á. M. & Moreira, J. D. Risks of dietary acrylamide exposure: A systematic review. Food Chem. 157, 310–322. https://doi.org/10.1016/j.foodchem.2014.02.046 (2014).
Benisi-Kohansal, S., Salari-Moghaddam, A., Seyed Rohani, Z. & Esmaillzadeh, A. Dietary acrylamide intake and risk of women’s cancers: A systematic review and meta-analysis of prospective cohort studies. Br. J. Nutr. 126, 1355–1363. https://doi.org/10.1017/s0007114520005255 (2021).
He, J., Zhu, X., Xu, K., Li, Y. & Zhou, J. Network toxicological and molecular docking to investigate the mechanisms of toxicity of agricultural chemical Thiabendazole. Chemosphere. https://doi.org/10.1016/j.chemosphere.2024.142711 (2024).
Liu, C. et al. Integrated metabolomics and network toxicology to reveal molecular mechanism of celastrol induced cardiotoxicity. Toxicol. Appl. Pharmacol. https://doi.org/10.1016/j.taap.2019.114785 (2019).
Huang, S. Analysis of environmental pollutant Bisphenol F elicited prostate injury targets and underlying mechanisms through network toxicology, molecular docking, and multi-level bioinformatics data integration. Toxicology https://doi.org/10.1016/j.tox.2024.153847 (2024).
Kanehisa, M., Furumichi, M., Sato, Y., Matsuura, Y. & Ishiguro-Watanabe, M. KEGG: Biological systems database as a model of the real world. Nucleic Acids Res. 53, D672–D677. https://doi.org/10.1093/nar/gkae909 (2025).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 28, 1947–1951. https://doi.org/10.1002/pro.3715 (2019).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30. https://doi.org/10.1093/nar/28.1.27 (2000).
Marques, C. et al. Association between dietary intake of acrylamide and increased risk of mortality in women: Evidence from the E3N prospective cohort. Sci. Total Environ. https://doi.org/10.1016/j.scitotenv.2023.167514 (2024).
Lyn-Cook, L. E. et al. Food contaminant acrylamide increases expression of Cox-2 and nitric oxide synthase in breast epithelial cells. Toxicol. Ind. Health 27, 11–18. https://doi.org/10.1177/0748233710380217 (2010).
Besaratinia, A. & Pfeifer, G. P. A review of mechanisms of acrylamide carcinogenicity. Carcinogenesis 28, 519–528. https://doi.org/10.1093/carcin/bgm006 (2007).
Adani, G. et al. Dietary intake of acrylamide and risk of breast, endometrial, and ovarian cancers: A systematic review and dose–response meta-analysis. Cancer Epidemiol. Biomark. Prev. 29, 1095–1106. https://doi.org/10.1158/1055-9965.Epi-19-1628 (2020).
Riddell, N. et al. Characterization and biological potency of mono- to tetra-halogenated carbazoles. Environ. Sci. Technol. 49, 10658–10666. https://doi.org/10.1021/acs.est.5b02751 (2015).
Xiao, F., Qiu, J. & Zhao, Y. Exploring the potential toxicological mechanisms of vine tea on the liver based on network toxicology and transcriptomics. Front. Pharmacol. https://doi.org/10.3389/fphar.2022.855926 (2022).
Dong, Y. et al. Molecular mechanism of Epicedium treatment for depression based on network pharmacology and molecular docking technology. BMC Complement. Med. Therap. https://doi.org/10.1186/s12906-021-03389-w (2021).
Pinzi, L. & Rastelli, G. Molecular docking: Shifting paradigms in drug discovery. Int. J. Mol. Sci. https://doi.org/10.3390/ijms20184331 (2019).
Sigismund, S., Avanzato, D. & Lanzetti, L. Emerging functions of the EGFR in cancer. Mol. Oncol. 12, 3–20. https://doi.org/10.1002/1878-0261.12155 (2017).
Steelman, L. S. et al. Critical roles of EGFR family members in breast cancer and breast cancer stem cells: Targets for therapy. Curr. Pharm. Des. 22, 2358–2388. https://doi.org/10.2174/1381612822666160304151011 (2016).
Chen, C., Ye, L., Yi, J., Liu, T. & Li, Z. FN1 mediated activation of aspartate metabolism promotes the progression of triple-negative and luminal a breast cancer. Breast Cancer Res. Treat. 201, 515–533. https://doi.org/10.1007/s10549-023-07032-9 (2023).
Evans, K. W. et al. Oxidative phosphorylation is a metabolic vulnerability in chemotherapy-resistant triple-negative breast cancer. Can. Res. 81, 5572–5581. https://doi.org/10.1158/0008-5472.Can-20-3242 (2021).
Zhang, Z. et al. Identification of key biomarkers related to epithelial-mesenchymal transition and immune infiltration in ameloblastoma using integrated bioinformatics analysis. Oral Dis. 29, 1657–1667. https://doi.org/10.1111/odi.14173 (2023).
Bao, H., Huo, Q., Yuan, Q. & Xu, C. Fibronectin 1: A potential biomarker for ovarian cancer. Dis. Markers 2021, 5561651. https://doi.org/10.1155/2021/5561651 (2021).
Wang, J., Li, R., Li, M. & Wang, C. Fibronectin and colorectal cancer: Signaling pathways and clinical implications. J. Recept. Signal Transduct. Res. 41, 313–320. https://doi.org/10.1080/10799893.2020.1817074 (2021).
Shao, W. et al. Chemical genomics reveals inhibition of breast cancer lung metastasis by Ponatinib via c-Jun. Protein Cell 10, 161–177. https://doi.org/10.1007/s13238-018-0533-8 (2018).
Li, X. et al. COL1A1: A novel oncogenic gene and therapeutic target in malignancies. Pathol. Res. Pract. https://doi.org/10.1016/j.prp.2022.154013 (2022).
Jiao, X. et al. c-Jun induces mammary epithelial cellular invasion and breast cancer stem cell expansion. J. Biol. Chem. 285, 8218–8226. https://doi.org/10.1074/jbc.M110.100792 (2010).
Kachamakova-Trojanowska, N. et al. HIF-1 stabilization exerts anticancer effects in breast cancer cells in vitro and in vivo. Biochem. Pharmacol. https://doi.org/10.1016/j.bcp.2020.113922 (2020).
Liu, X. et al. HIF-1–regulated expression of calreticulin promotes breast tumorigenesis and progression through Wnt/β-catenin pathway activation. Proc. Natl. Acad. Sci. https://doi.org/10.1073/pnas.2109144118 (2021).
de Heer, E. C., Jalving, M. & Harris, A. L. HIFs, angiogenesis, and metabolism: Elusive enemies in breast cancer. J. Clin. Investig. 130, 5074–5087. https://doi.org/10.1172/jci137552 (2020).
Alves, C. L. & Ditzel, H. J. Drugging the PI3K/AKT/mTOR pathway in ER+ breast cancer. Int. J. Mol. Sci. https://doi.org/10.3390/ijms24054522 (2023).
Miao, W., Wang, Z., Gao, J. & Ohno, Y. Polyphyllin II inhibits breast cancer cell proliferation via the PI3K/Akt signaling pathway. Mol. Med. Rep. https://doi.org/10.3892/mmr.2024.13348 (2024).
Funding
The study was supported by the National Natural Science Foundation of China (grant no.82260537 and 82460478), the Association Foundation Program of Department of Yunnan Science and Technology and Kunming Medical University (grant no. 202501AY070001-006), Biomedical Projects of Yunnan Key Science and Technology Program (202302AA310046), and the scientific research fund project of Department of Education, Yunnan Province (grant no. 2023Y0804 and 2024Y219).
Author information
Authors and Affiliations
Contributions
Z.D. and C.Y.: conceive and conduct experiments, write the original draft articles. W.W.: investigation. Q.Z. and W.Y.: collate and analyze data. Q.R., R.Z: Data curation. Y.Z. and S.Y.: fund support and manuscript revision. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Duan, Z., Yu, C., Yang, W. et al. Network toxicology and molecular docking reveal the potential link between acrylamide exposure and breast cancer. Sci Rep 15, 21076 (2025). https://doi.org/10.1038/s41598-025-06964-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-06964-0