Abstract
Microbial transglutaminase (MTG) is an enzyme that catalyzes the cross-linking of glutamine and lysine residues in proteins. Because of its ability to modify proteins, MTG has various applications in the medical and food industries. Most studies have aimed to enhance the thermal stability of MTG by focusing only on point mutations. Introducing a disulfide (S-S) bond in the N-terminal region has been found to be effective, whereas S-S bonds in other regions were considered ineffective. Therefore, this study aimed to evaluate the impact of introducing an additional S-S bond on the thermal stability of an MTG mutant. We found that adding S-S bonds to regions other than the N-terminal, in conjunction with the N-terminal S-S bond, significantly enhanced thermal stability. This finding demonstrates the importance of reinforcing the weakest part of the protein first, followed by strengthening other regions for optimal thermal stability. The MTG variant with two S-S bonds retained its catalytic activity and substrate specificity towards protein substrates, making it a promising candidate for industrial applications. Thus, introducing S-S bonds could be an effective strategy to increase thermal stability of MTG and other industrial enzymes, thereby contributing to their potential industrial applications.
Similar content being viewed by others
Introduction
Transglutaminase (TG: protein-glutamine γ-glutamyltransferases, EC 2.3.2.13) is a family of enzymes that catalyzes the formation of covalent bonds between the γ-carboxyamide group of the glutamine residue and the ε-amino group of the lysine residue in a peptide or protein, leading to the cross-linking of the ε-(γ-glutamyl) lysine bridge1. TG is widely distributed in various mammalian cells and tissues, and its physiological characteristics have been studied previously2. The enzymatic activity of TG is crucial in various biological processes, including blood clotting, skin formation, and wound healing3. TG in mammals is calcium-dependent; however, Streptomyces mobaraensis TG is calcium independent4,5. Compared to TG derived from mammalian cells, microbial TG (MTG) has a shorter amino acid sequence, lower homology, and a different three-dimensional structure6. MTG is expressed in a form where the pro-sequence is combined (Pro-MTG), which matures after being cleaved by a protease7. The pro-sequence reportedly promotes efficient protein folding and secretion, and suppresses its enzymatic activity8. Since MTG is easy to purify, it is being used in various research and industrial applications, replacing TG obtained from animal tissues and organs9. Specifically, MTG has significant applications in the food, pharmaceutical, and textile industries owing to their ability to modify protein structures and enhance the functional properties of proteins10,11,12,13.
The thermal stability of an enzyme determines its applicability in industrial processes that often require high temperatures. Enzymes with higher thermal stability can maintain their activity over a broader range of temperatures, showing their versatility and cost-effectiveness for industrial applications. Several methods use protein engineering techniques to identify thermostable enzymes. Notably, various proteins have been stabilized through random mutagenesis, directed evolution, and rational mutagenesis using the three-dimensional structure. Studies have been conducted to enhance the thermal stability of MTG. For example, studies have reported multiple mutants obtained through random mutagenesis, DNA shuffling, and saturation mutagenesis10,11,12,13. In addition, rational modification of flexible regions (S2P-S23V-Y24N-E28T-S116A-S179L-S199A-A265P-A287P-K294L) has produced a thermostable MTG mutant, TGm2A14,15,16. Moreover, introducing disulfide (S-S) bonds in the N-terminal region can significantly enhance the thermal stability of proteins. For instance, the D3C/G283C or T7C/E58C mutant of MTG reportedly improved thermal stability compared to the wild-type (WT)17,18,19. However, the introduction of S-S bonds in other regions was considered ineffective17.
By systematically introducing S-S bonds into different regions of MTG, we aim to identify the most effective strategies for enhancing its thermal stability. We hypothesize that introducing S-S bonds into regions of this enzyme that are inherently less thermally stable will significantly improve their overall thermal stability. Therefore, the current study aims to evaluate how the introduction of an additional S-S bond to the D3C/G283C MTG mutant will impact its thermal stability. Moreover, this strategy might provide insights into improving the thermal stability of other industrial enzymes.
Results
In silico mutant design
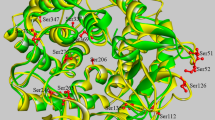
The three-dimensional structure of MTG is formed from three domains, namely, the α-helix domain 1 starting from the N-terminus, β-sheet domain, and α-helix domain 2 (Fig. 1A). Previous research has shown that the S2C/G283C, D3C/G283C, or T7C/E59C mutations introduced into the α-helix domain 1 improved thermal stability17,18. Conversely, it has been found that the E93C/V112C and A106C/D213C mutations introduced in the α-helix domain 2 did not improve thermal stability17. Our results identified α-helix 1 as the weakest region in MTG, emphasizing the need to strengthen this region. In other words, we suggest that strengthening α-helix 2 without strengthening α-helix 1 would not contribute to thermal stability because α-helix 1 would unfold. Furthermore, introducing two S-S bonds into α-helix 1 would have no effect because a single S-S bond in α-helix 1 sufficiently enhances thermal stability. Therefore, we first strengthened the α-helix domain 1 using D3C/G283C, which has been identified in previous studies as the most effective disulfide bond18 and introduced S-S bonds into the β-sheet domain or α-helix domain 2. To introduce S-S bonds into the protein, it is necessary to mutate the amino acid residue pairs to cysteines. These amino acid residue pairs were predicted using the crystal structure of MTG and a computer program and then verified experimentally. First, we calculated the relative positions of all the amino acid pairs in MTG and extracted those in which the distance between Cβ is within 2Å. We then removed the mutants that were expected to cause significant steric hindrance due to mutation to Cys residues based on the energy values. In general, the principle of introducing S-S bonds to confer thermal stability aims to rigidify and stabilize the protein structure. However, mutations to nearby residues or those located at the tip of the loop in the hairpin structure are considered unlikely to achieve the rigidifying effect and were therefore excluded from the candidate sequence.
Sequence and structure of MTG mutants. (A) The structure of MTG. Left: α-helix domain 1 is shown in blue, β-sheet domain in green, and α-helix domain 2 in red. Right: Colored according to B-factor values. Warmer colors indicate higher B-factor, while cooler colors indicate lower B-factor. (B) The sequence of MTG. α-helix domain 1 is shown in blue, β-sheet domain in green, and α-helix domain 2 in red. The dashed lines indicate combinations of amino acid residues with S-S bonds introduced. (C) Structure models of the S-S mutant MTGs. Red color indicates the Cys residue introduced to form an S-S bond.
In addition, the hairpin, which contains residues 239–254 and forms a pocket for the active residue Cys64, has a high B-factor and is highly flexible. However, we assumed that fixing the catalytic pocket would reduce enzyme activity and narrow substrate recognition. Hence, we did not introduce S-S bonds to these sites. Finally, five additional mutations were each introduced into the D3C/G283C mutant and evaluated. These five mutants were A81C/V311C, E93C/V112C, A106C/D213C, E107C/Y217C, and A160C/G228C, and the positions of the mutations in the sequence and structure are shown in Fig. 1B and C.
In this study, the S-S bonds that showed improved thermal stability are summarized in Table 1. The distance between each S-S bond ranged from 3.3 to 4.4 Å, and the dihedral angle values varied.
Improved thermal stability was observed upon the introduction of S-S bonds, irrespective of the secondary structure (α-helix, β-sheet, or coil). Similarly, enhanced thermal stability was achieved regardless of whether the S-S bonds were introduced into regions with high or low B-factors (Cα). While it is generally assumed that stabilizing regions with high B-factors is critical for improving thermal resistance, the findings of this study did not support this assumption. Instead, the results demonstrated that improved thermal stability can be achieved when two Cβ atoms are positioned at an appropriate distance and angle for S-S bond formation, regardless of the secondary structure or B-factor.
Thermostability evaluation of MTG S-S mutants
Designed mutants were expressed and purified (Supplementary Fig. S1) prior to the evaluation of their enzyme activity. To assess the thermostability of the WT MTG and various S-S mutants, purified proteins were incubated at 37, 45, 55, 60, 65, 70, 75, and 80 °C for 10 min, followed by subsequent analysis of their enzymatic activity. Among the S-S bond-introduced mutants based on the WT, only the D3C/G283C mutant exhibited improved thermal stability (Fig. 2A). In contrast, no significant change in thermal stability was observed for the other S-S bond-introducing mutants (A81C/V311C, E93C/V112C, A106C/D213C, E107C/Y217C, and A160C/G228C). However, when additional S-S bonds were introduced into the D3C/G283C mutant (rather than the WT background), a further improvement in thermal stability was observed compared to D3C/G283C alone.
Thermal stability and temperature dependence of MTG mutants. (A) Residual activity of each mutant after thermal treatment (n = 3). Error bars indicate standard errors. (B) Specific activity of each mutant measured at each temperature (n = 3). Error bars indicate standard errors. (C) Relative activity of each mutant measured at each temperature (n = 3). Error bars indicate standard errors. (D) Melting temperatures of each mutant were determined using nanoDSF, with each sample measured in duplicate.
To evaluate the optimal temperature for the WT MTG and the various S-S mutants, MTG activity was measured using a hydroxamate assay. As anticipated from the thermal stability results, following the introduction of S-S bonds into the WT background, only the D3C/G283C mutant exhibited an increase in the optimum temperature from 55 to 60 °C. In contrast, no change in the optimum temperature was observed for the other S-S bond-introduced mutants (A81C/V311C, E93C/V112C, A106C/D213C, E107C/Y217C, and A160C/G228C) (Fig. 2B C). However, when these five additional mutant pairs were introduced into the D3C/G283C mutant, the optimum temperature for activity was further improved compared to that of the D3C/G283C mutant alone. Notably, an increase in specific activity at the respective temperatures was also observed for these constructs.
The melting temperature (Tm) of the WT MTG and various S-S mutants was measured in the presence and absence of a reducing agent (Fig. 2D). In the absence of a reducing agent, the Tm values revealed a clear trend. The WT protein exhibited a Tm of 61.84 °C, while individually introduced S-S mutants (A81C/V311C, E93C/V112C, A106C/D213C, E107C/Y217C, and A160C/G228C), which did not show significant stabilization effects, displayed Tm values comparable to that of WT. In contrast, the previously reported D3C/G283C mutant, known for its strong stabilizing effect, exhibited a significantly higher Tm of 68.93 °C. Importantly, introducing additional S-S bonds into the D3C/G283C mutant further increased the Tm to above 70 °C. Under reducing conditions, where the S-S bonds were cleaved, the Tm values of both WT and all S-S mutants converged to levels similar to that of WT. These results indicate that the stabilizing effects observed under non-reducing conditions are directly attributable to the presence of S-S bonds, which are abolished upon reduction.
Characterization of the enzyme kinetics of S-S mutants
The enzyme kinetics of the WT MTG and the various S-S mutants were measured by a glutamate dehydrogenase (GDH)-coupled enzyme assay. The enzymatic kinetics of each S-S mutant were similar to those of the WT MTG. A81C/V311C showed a decreased Km, D3C/G283C showed a slightly decreased Km, whereas E93C/V112C, A106C/D213C, E107C/Y217C, and A160C/G228C showed 5–21% higher Km than wild-type MTG, and thus 71–88% reduced kcat/ Km values (Table 2). The effects on Km and kcat by combining S-S bonds were minimal. These results indicate that the introduction of S-S bonds makes it possible to acquire an enzyme with enhanced thermal stability while maintaining comparable catalytic activity. Such differences in Km and kcat are thought to be due to the introduced S-S mutations affecting the active site in some way. In particular, the A81C/V311C mutant, which exhibited a decrease in Km, may have influenced the catalytic residue, as V311 is located on the β-sheet where D255, one of the catalytic residues, resides.
Evaluation of protein substrate reactivity
The reactivity of MTG and various S-S mutants with different protein substrates was investigated. To explore their potential application in food processing, the reactivity of MTG to sodium caseinate, fish gelatin, soy protein isolate, and α-lactalbumin was evaluated. We determined the amounts of ammonia released after incubating 2% solutions of these proteins with MTG for 1 h at 37 °C. Although variations in reactivity were observed depending on the protein substrate type, no substantial differences in reactivity were noted between each mutant and the WT (Fig. 3). This indicates that the introduction of S-S bonds did not significantly affect the reactivity toward polymeric substrates. Therefore, it is possible to develop industrially useful enzymes with enhanced thermal stability, comparable catalytic activity, and unaltered reactivity toward polymeric substrates.
Protein substrate reactivity of MTG S-S mutants. The activity after incubation at 37 °C for 60 min (n = 3). Error bars indicate standard errors.
Selection of effective sites for introducing S-S bonds utilizing sequence information
To verify the possibility of predicting effective sites for S-S bond introduction based on sequence information, we analyzed the evolutionary conservation of the sites where S-S bonds were introduced. Using 406 sequences of MTG homologs, the amino acid conservation at the mutation sites was analyzed. Regardless of the conservation level of the surrounding sequences, the results showed that both mutants with improved and decreased thermal stability were observed (Supplementary Fig. S2).
Discussion
The introduction of S-S bonds is a simple and effective method for improving thermal stability without requiring multiple amino acid substitutions. In this study, we investigated further improvements in thermal stability using a MTG mutant by introducing the most important S-S bond (D3C/G283C). Although thermal stability improvements through random mutagenesis have been reported19we successfully created a MTG mutant with a higher thermal stability than those in previous studies14,15,17,18,19. We achieved this by introducing additional S-S bonds as a new approach. It is difficult to predict the appropriate site for introducing S-S bonds from sequence information containing evolutionary information. Previously, it was essential to utilize experimental structural information; however, the introduction of AlphaFold2 or RoseTTAfold reduced this limitation20,21.
From the S-S bonds mutants that has acquired thermal stability, it was found that the thermal stability was improved in the five mutants in which D3C/G283C on the α-helix domain 1 was strengthened and added to the β-sheet domain or α-helix domain 2 by S-S bonding. Furthermore, we confirmed that introducing a S-S bond to the β-sheet domain or α-helix domain 2 without introducing it to the α-helix domain 1 did not improve thermal stability. This indicates that the stability of the entire protein cannot be improved unless the weakest region of the protein, the α-helix domain 1, is strengthened alongside others.
The activity of this heat-resistant enzyme did not reduce, showing its significance. It was generally believed that there was a trade-off between thermal stability and activity22,23. However, many unknowns exist, and recent reports have questioned this trade-off24. There was no trade-off between thermal stability and activity in the mutants obtained in the present study. The strategy of avoiding S-S bond introduction within the active site yielded favorable results.
The results of this study suggested that in order to improve thermal stability without reducing enzyme activity, it is important to introduce S-S bonds in the region that is the weakest in the enzyme and not too close to the active domain. Structural information is essential for this, and obtaining structural information has become much easier with tools such as AlphaFold 2. On the other hand, it is still laborious to investigate the heat-sensitive parts of a protein’s structure using instrumental analysis, so it is probably easier to use in silico methods such as molecular dynamics or to screen a certain number of mutants.
MTG is being considered for use in various industries, but thermal stability can sometimes be an issue. The thermal stabilized MTG discovered in this study is expected to be useful in the food industry, textile industry, and leather processing field, which require high-temperature processing18,25.
Conclusion
The thermal stability of MTG was improved by introducing S-S bonds. When heated at 65 °C for 10 min, the wild-type MTG had no remaining activity. However, the D3C/G283C mutant, which had one S-S bond introduced, had approximately 40% of its activity retained17,18,19. In this study, a second S-S bond was introduced at a different site to the first S-S bond (A81C/V311C, E93C/V112C, A106C/D213C, E107C/Y217C, or A160C/G283C). These mutants improved their residual activity to approximately 80%. They maintained the same activity as the WT, and the substrate specificity for the protein was also maintained, it can be used in high-temperature range, which was difficult to do with WT MTG, thus enhancing their applicability in the food, textile and medical industries.
Methods
Mutants design
The Schrödinger software (Schrödinger, Inc.) was used for in silico S-S and site prediction. The three-dimensional structure of MTG was obtained from Protein Data Bank (PDB code 1iu4)6. First, the missing side chains and protons were added using the Protein Preparation Program, and the structure was optimized26. The amino acid pairs that were in a position relationship and could form an S-S bond were extracted using the S-S prediction program27. We used a program’s weighted score of 1,000 or less to determine whether the distance and angle were suitable for an S-S bond. Furthermore, we selected five mutants by excluding those located near the catalytic pocket, those with amino acid sequences differing by ten or fewer residues, and those with mutations in the same domain as the D3C/G283C mutant.
Plasmids and bacterial strains
Corynebacterium glutamicum YDK010 was used as an expression host for MTGs28. pPSPTG1 is a plasmid for expressing pro-MTG29. Genes coding for MTG mutants were synthesized by optimizing codon usage for C. glutamicum (Genescript) and inserted at the KpnI and BamHI sites of pPSPTG1.
Culture medium for protein production
C. glutamicum was grown at 30 °C in the medium (5 g/L glucose, 10 g/L tryptone, 10 g/L yeast extract, 5 g/L NaCl, and 0.2 g/L DL-methionine, pH 7.2) containing 25 mg/mL kanamycin (NACALAI TESQUE, INC.). To produce the pro-form MTG mutants, C. glutamicum was grown at 30 °C in the medium (60 g/L glucose, 1 g/L MgSO4, 30 g/L (NH4)2SO4, 1.5 g/L KH2PO4, 0.01 g/L FeSO4·7H2O, 0.01 g/L MnSO4·7H2O, 450 µg/L biotin, 0.15 g/L DL-methionine, and 50 g/L CaCO3, pH 7.5) containing 25 mg/mL kanamycin. To produce MTG mutants containing an S-S bond, 3 mM dithiothreitol was added 5 h after the culture ensued, as described previously18.
Determination of protein concentration
Reverse-phase HPLC was performed using Proteonavi C4 column (4.6 mm id ×15 cm, Osaka Soda) as described previously18. Purified WT MTG was used as a standard. It was purified from the Streptoverticillium spp. S-8112 supernatant culture, as described previously4. The concentration of the standard WT MTG was determined using the Bradford assay, with bovine serum albumin (Bio-Rad) serving as the protein standard.
Purification of MTGs
MTG was purified as described previously18. In summary, culture supernatant was obtained by removing bacterial cells from the culture solution through centrifugation (10,000 x g, 4 °C, 10 min) and filtration. The supernatant was subjected to buffer exchange using a Sephadex G-25 (M) (Cytiva) column equilibrated with 20 mM MES (pH 5.5). After adjusting the pH to 7.0 with sodium hydroxide, Pro-MTG was activated by protease from Bacillus licheniformis (Sigma-Aldrich) treatment, which cleaved and removed the pro-sequence. Then, 0.5-1.0% v/w of protease was added, and the MTG concentration was determined using UHPLC (LC-2040 C 3D, SHIMADZU CORPORATION). After adjusting the pH to 5.5 with 10% acetic acid, the sample was loaded onto a cation-exchange column RESOURCE™ S 6 mL (Cytiva) equilibrated with 20 mM MES (pH 5.5). After the column was washed with 10-column volumes of the same buffer, MTG was eluted with a linear gradient of sodium chloride from 0 to 500 mM over 10-column volumes at a flow rate of 6 mL/min. Fractions at the top were collected and subjected to buffer exchange. For buffer exchange, samples were subjected to a gel filtration column HiPrep™ 26/10 Desalting (Cytiva) equilibrated with 20 mM sodium phosphate (pH 6.0).
Colorimetric hydroxamate assay
A colorimetric hydroxamate assay using N-carbobenzoxy-L-glutaminylglycine (Z-QG) was employed to measure the specific MTG activity, as described previously30. The reaction solution A comprised 30 mM Z-QG, 100 mM NH2OH, 10 mM reduced glutathione, and 50 mM MES (pH 6.0). Reactions were quenched by adding stop solution B (1 N HCl, 4% TCA, and 5% FeCl3・6H2O). The assays were performed as follows: A 50 µL aliquot of the enzyme sample was added to 500 µL of the reaction solution A and incubated for 10 min at 37 °C. The enzymatic reaction was quenched by adding 500 µL of stop solution B, and the absorbance was measured at 525 nm using SpectraMax M2 Microplate Reader (Molecular Devices). One unit of enzyme activity was defined as the formation of 1 µmol of hydroxamic acid/min, using L-glutamic acid γ-monohydroxamate as the standard.
Temperature characteristics of MTG
To evaluate the thermal stability, the enzyme solution was dissolved in 20 mM MES buffer (pH 6.0) solution to a final concentration of 1.5 U/mL and incubated for 10 min at 50, 60, 65, 70, and 75 °C. After thermal incubation, the enzyme solution was quickly transferred to an ice, and the enzyme activity was subsequently measured.
To evaluate the optimal temperature, MTG solution (50 µL, 0.05 mg/mL) was added to 500 µL of hydroxamate assay reaction solution A. The solution obtained was reacted for 10 min at 37, 45, 55, 60, 65, 70, 75, and 80 °C. Then, 500 µL of hydroxamate assay stop solution B was added to the reaction mixture, and the amount of the product was measured by recording the absorbance at 525 nm.
To evaluate the melting temperatures (Tm), the enzyme solution was prepared in 20 mM sodium phosphate (pH 6.0) containing either 1 mM TCEP or none and incubated at 25 °C for 1 h. The intrinsic tryptophan fluorescence at 330 nm and 350 nm was monitored in real time using Prometheus Panta (NanoTemper) across a temperature range of 25–95 °C, with a gradient of 1 °C per minute. All experiments were performed in duplicate. Protein melting temperatures (Tm) were calculated as the peaks of the first derivatives of the 350/330 nm fluorescence ratio, analyzed using the Prometheus Panta software.
Kinetic measurement of MTG and their mutants by GDH-coupled enzyme assay
Kinetic parameters of MTG were determined through a GDH-coupled enzyme assay31. The reaction solution consisted of 200 mM MOPS-NaOH buffer (pH 7.2), 10 mM α-ketoglutarate, 1 mM EDTA, 0.5 mM NADH, varying concentrations of Z-QG as the acyl-donor substrate (0, 0.63, 2.5, 10 and 40 mM), 10 mM PEG-NH2 (SUNBRIGHT MEPA-20 H, NOF Corporation, Tokyo, Japan) as an acyl-acceptor substrate, 20 U/mL of glutamate dehydrogenase (Sigma-Aldrich, G7882), and 10 µg/mL MTG. The assays were performed as follows: A 20 µL aliquot of purified enzyme sample was added to 180 µL of reaction solution. The enzyme assay was carried out at 37 °C in a 96-well plate, and the decline in the absorbance at 340 nm by oxidation of NADH was monitored using SpectraMax M2 Microplate Reader (Molecular Devices).
Substrate reactivity of the enzyme
To measure the reactivity with various proteins, a 50 mM MES buffer (pH 6.0) was prepared containing 2% (w/v) of sodium caseinate (Sigma-Aldrich), α- lactalbumin (Sigma-Aldrich), fish gelatin (Sigma-Aldrich), and soy protein isolate (Merck). The MTG solution (10 µL, 2 U/mL) was added to 100 µL of each protein solution and incubated for 1 h at 37 °C. Reaction was quenched by adding 110 µL of 12% TCA, and the ammonia was measured using a LabAssay™ Ammonia (FUJIFILM Wako Pure Chemical Corporation).
Data availability
All data generated or analysed during this study are included in this published article and its supplementary information files.
Abbreviations
- GDH:
-
Glutamate dehydrogenase
- MTG:
-
Microbial transglutaminase
- TG:
-
Transglutaminase
- S-S:
-
Disulfide
References
Ikura, K. et al. Amino acid sequence of guinea pig liver transglutaminase from its cDNA sequence. Biochemistry 27, 2898–2905. https://doi.org/10.1021/bi00408a035 (1988).
Folk, J. E. & Transglutaminases Annu. Rev. Biochem. 49, 517–531 (1980). https://doi.org/10.1146/annurev.bi.49.070180.002505.
Duarte, L., Matte, C. R., Bizarro, C. V. & Ayub, M. A. Z. Transglutaminases: Part I-origins, sources, and biotechnological characteristics. World J. Microbiol. Biotechnol. 36, 15. https://doi.org/10.1007/s11274-019-2791-x (2020).
Ando, H. et al. Purification and characteristics of a novel transglutaminase derived from microorganisms. Agric. Biol. Chem. 53, 2613–2617. https://doi.org/10.1080/00021369.1989.10869735 (1989).
Nonaka, M. et al. Polymerization of several proteins by Ca2+-Independent transglutaminase derived from microorganisms. Agric. Biol. Chem. 53, 2619–2623. https://doi.org/10.1080/00021369.1989.10869736 (1989).
Kashiwagi, T. et al. Crystal structure of microbial transglutaminase from Streptoverticillium mobaraense. J. Biol. Chem. 277, 44252–44260. https://doi.org/10.1074/jbc.M203933200 (2002).
Pasternack, R. et al. Bacterial pro-transglutaminase from Streptoverticillium mobaraense–purification, characterisation and sequence of the Zymogen. Eur. J. Biochem. 257, 570–576. https://doi.org/10.1046/j.1432-1327.1998.2570570.x (1998).
Suzuki, M. et al. A back hydrogen exchange procedure via the acid-unfolded state for a large protein. Biochemistry 51, 5564–5570. https://doi.org/10.1021/bi300495p (2012).
Fuchsbauer, H. L. Approaching transglutaminase from Streptomyces bacteria over three decades. FEBS J. 289, 4680–4703. https://doi.org/10.1111/febs.16060 (2022).
Miwa, N. Innovation in the food industry using microbial transglutaminase: Keys to success and future prospects. Anal. Biochem. 597, 113638. https://doi.org/10.1016/j.ab.2020.113638 (2020).
Kolotylo, V., Piwowarek, K. & Kieliszek, M. Microbiological transglutaminase: Biotechnological application in the food industry. Open. Life Sci. 18 https://doi.org/10.1515/biol-2022-0737 (2023).
Duarte, L., Matte, C. R., Bizarro, C. V. & Ayub, M. A. Z. Review transglutaminases: Part II-industrial applications in food, biotechnology, textiles and leather products. World J. Microbiol. Biotechnol. 36, 11. https://doi.org/10.1007/s11274-019-2792-9 (2019).
Yokoyama, K., Nio, N. & Kikuchi, Y. Properties and applications of microbial transglutaminase. Appl. Microbiol. Biotechnol. 64, 447–454. https://doi.org/10.1007/s00253-003-1539-5 (2004).
Wang, X. et al. Significantly improving the thermostability and catalytic efficiency of Streptomyces mobaraenesis transglutaminase through combined rational design. J. Agric. Food Chem. 69, 15268–15278. https://doi.org/10.1021/acs.jafc.1c05256 (2021).
Yang, P. et al. Enhanced thermostability and catalytic activity of Streptomyces mobaraenesis transglutaminase by rationally engineering its flexible regions. J. Agric. Food Chem. 71, 6366–6375. https://doi.org/10.1021/acs.jafc.3c00260 (2023).
Ye, J., Yang, P., Zhou, J., Du, G. & Liu, S. Efficient production of a thermostable mutant of transglutaminase by Streptomyces mobaraensis. J. Agric. Food Chem. 72, 4207–4216. https://doi.org/10.1021/acs.jafc.3c07621 (2024).
Yokoyama, K. et al. Effect of introducing a disulfide Bridge on the thermostability of microbial transglutaminase from Streptomyces mobaraensis. Appl. Microbiol. Biotechnol. 105, 2737–2745. https://doi.org/10.1007/s00253-021-11200-6 (2021).
Suzuki, M., Date, M., Kashiwagi, T., Suzuki, E. & Yokoyama, K. Rational design of a disulfide Bridge increases the thermostability of microbial transglutaminase. Appl. Microbiol. Biotechnol. 106, 4553–4562. https://doi.org/10.1007/s00253-022-12024-8 (2022).
Suzuki, M. et al. Random mutagenesis and disulfide bond formation improved thermostability in microbial transglutaminase. Appl. Microbiol. Biotechnol. 108, 478. https://doi.org/10.1007/s00253-024-13304-1 (2024).
Jumper, J. et al. Highly accurate protein structure prediction with alphafold. Nature 596, 583–589. https://doi.org/10.1038/s41586-021-03819-2 (2021).
Baek, M. et al. Accurate prediction of protein structures and interactions using a three-track neural network. Science 373, 871–876. https://doi.org/10.1126/science.abj8754 (2021).
Sterner, R. & Liebl, W. Thermophilic adaptation of proteins. Crit. Rev. Biochem. Mol. Biol. 36, 39–106. https://doi.org/10.1080/20014091074174 (2001).
Feller, G. Protein stability and enzyme activity at extreme biological temperatures. J. Phys. Condens. Matter. 22, 323101. https://doi.org/10.1088/0953-8984/22/32/323101 (2010).
Miller, S. R. An appraisal of the enzyme stability-activity trade-off. Evolution 71, 1876–1887. https://doi.org/10.1111/evo.13275 (2017).
Zhu, Y. & Tramper, J. Novel applications for microbial transglutaminase beyond food processing. Trends Biotechnol. 26, 559–565. https://doi.org/10.1016/j.tibtech.2008.06.006 (2008).
Sastry, G. M., Adzhigirey, M., Day, T., Annabhimoju, R. & Sherman, W. Protein and ligand preparation: Parameters, protocols, and influence on virtual screening enrichments. J. Comput. Aided Mol. Des. 27, 221–234. https://doi.org/10.1007/s10822-013-9644-8 (2013).
Salam, N. K., Adzhigirey, M., Sherman, W. & Pearlman, D. A. Structure-based approach to the prediction of disulfide bonds in proteins. Protein Eng. Des. Sel. 27, 365–374. https://doi.org/10.1093/protein/gzu017 (2014).
Date, M., Yokoyama, K., Umezawa, Y., Matsui, H. & Kikuchi, Y. High level expression of Streptomyces mobaraensis transglutaminase in Corynebacterium glutamicum using a chimeric pro-region from Streptomyces cinnamoneus transglutaminase. J. Biotechnol. 110, 219–226. https://doi.org/10.1016/j.jbiotec.2004.02.011 (2004).
Kikuchi, Y., Date, M., Yokoyama, K., Umezawa, Y. & Matsui, H. Secretion of active-form Streptoverticillium mobaraense transglutaminase by Corynebacterium glutamicum: Processing of the pro-transglutaminase by a cosecreted subtilisin-Like protease from Streptomyces albogriseolus. Appl. Environ. Microbiol. 69, 358–366. https://doi.org/10.1128/AEM.69.1.358-366.2003 (2003).
Yokoyama, K. et al. In vitro refolding process of urea-denatured microbial transglutaminase without pro-peptide sequence. Protein Expr Purif. 26, 329–335. https://doi.org/10.1016/s1046-5928(02)00536-3 (2002).
Oteng-Pabi, S. K. & Keillor, J. W. Continuous enzyme-coupled assay for microbial transglutaminase activity. Anal. Biochem. 441, 169–173. https://doi.org/10.1016/j.ab.2013.07.014 (2013).
Acknowledgements
We received a lot of advice from K. Yokoyama, T. Kashiwagi, N. Miwa and M. Date about the content of past research on MTG. We are grateful to Y. Matsui and K. Takei for technical assistance with the experiments.
Funding
This work was supported by Ajinomoto Co., Inc.
Author information
Authors and Affiliations
Contributions
T.O., K.T., Y.H., Y.M., I.A., and M.S. contributed to the conception and design of the study. T.O. and K.T. designed the mutants, planned and performed the experiments. A.S. measured Tm value and interpreted the data. The manuscript was written by T.O. and K.T. All authors reviewed and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
K. Takahashi and T. Ono have competing interests related to this research. The details of the competing interests are as follows: WO/2022/071061. Other authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.

Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ono, T., Takahashi, K., Hirao, Y. et al. Introduction of multiple disulfide bonds increases the thermostability of transglutaminase. Sci Rep 15, 21105 (2025). https://doi.org/10.1038/s41598-025-07842-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-07842-5
Keywords
This article is cited by
-
Microbial transglutaminase in food biotechnology: from biochemical mechanisms to industrial applications
Applied Microbiology and Biotechnology (2026)