Abstract
The genetic characteristics of six wild populations of Ormosia microphylla were investigated and their soil properties were analyzed by using Hyper-seq sequencing technology. Based on the findings, targeted protection strategies are proposed. Genetic diversity analysis revealed moderate overall genetic diversity of O. microphylla populations, with the ZD showing the highest and the JX the lowest genetic diversity. Compared to the populations outside the tiankeng, the LY, located in a Karst tiankeng, exhibited lower genetic diversity. This reduced genetic diversity of LY was possibly influenced by the combined influence of selective pressures unique to the tiankeng environment as well as the founder effect. Genetic structure analysis further indicated that the optimal number of clusters for the six populations was K = 7. The LY exhibited a relatively pure genetic constitution, whereas the other populations exhibited more complex genetic structures. Phylogenetic analysis and assessments of genetic differentiation coefficients further demonstrated significant genetic distance of the LY from the other populations, with a high degree of differentiation. Overall, there was a significant level of genetic differentiation among the six populations. Soil analysis revealed that total organic content and Fe were significantly higher compared to other elements across the six populations. In addition, compared to other populations, the LY exhibited notable variations in soil characteristics, such as higher levels of Fe, K, TK, and Ca. Additionally, the pH of the soil harboring LY was neutral, whereas that of other populations was acidic. IBD analysis, after removing the LY, revealed that the outcomes changed from non-significant to significant, highlighting the critical role of the isolation effect of Karst tiankeng and geographical barriers in promoting genetic differentiation among populations. IBE analysis further indicated a significant impact of K and TK content in the soil on genetic differentiation between populations.
Similar content being viewed by others
Introduction
There are about 130 species of Ormosia in the world, which is a large genus of Fabaceae, mainly distributed in tropical America, Southeast Asia and northwestern Australia1. According to the latest survey statistics, there are 35 species of Ormosia plants in China, which are mainly distributed in subtropical regions such as Guangxi, Yunnan and Guizhou, and are the most distributed countries of Ormosia plants in Asia2. Due to the particularity of the required ecological factors, many plants of this genus are endemic to some countries and regions. Among them, Ormosia microphylla is one of the endemic species in China, and its geographical distribution is narrow. It is only distributed in evergreen broad-leaved forests and evergreen deciduous broad-leaved mixed forests in Guangxi, Guizhou, Fujian, Guangdong and other regions of China3. O. microphylla is also commonly referred to as “Guangxi Zitan”, a name derived from its primary distribution in Guangxi Province, and “Xiaoye Zitan”, which translates to “small-leaved zitan” in Chinese. Both terms are widely used in local contexts and traditional wood markets. Its sapwood and heartwood are clearly distinguished. The sapwood is light yellow–brown, and the heartwood accounts for a large proportion. It is deep purple-red to purple-black, with straight texture, fine structure, strong corrosion resistance, and beautiful color and pattern. It is a special precious material for manufacturing high-level furniture, musical instruments, and art crafts4. Due to the high price of logs and finished furniture, the theft of O. microphylla wild resources in recent decades has been very serious, and the predatory excavation has seriously damaged its population structure. In addition to the external environmental pressure, in the natural environment, the flowering and fruiting of O. microphylla adult plants are unstable, and the amount of fruit is small and there is a phenomenon of large and small years5. At the same time, the seeds of the plant also have the characteristics of poor propagation ability, hard seed coat, difficult to germinate, difficult to absorb water after drying and irregular germination5. These reasons together hinder the natural regeneration of O. microphylla population and the expansion of population size. Studies have found that the O. microphylla population belongs to the declining type, which is extremely lack of self-renewal ability3. At the same time, the habitat fragmentation of the population is serious, and its spatial distribution pattern belongs to the random distribution type, which seriously affects the survival of the species. Many of the original records are distributed in places, and no living body has been found. In the ' Red List of Biodiversity in China—Higher Plant Volume ' jointly compiled by the Ministry of Environmental Protection of China and the Chinese Academy of Sciences in 2013, O. microphylla was only classified as Near Threatened (NT). However, in China 's newly revised ' National Key Protected Wild Plants List ' in 2021, O. microphylla was quickly listed as a state-level key protected plant, while the other plants of the genus Ormosia were listed as second-level key protected plants6. However, there is still a big gap in the research on O. microphylla, which is limited to ecological protection research such as resource investigation5, community structure7 and breeding and reproduction8.
Although these studies have played a positive role in species conservation, the evolutionary history, causes of endangerment and future conservation strategies of species are still unclear. The study of genetic diversity is of great significance in the protection of endangered plants. This study can effectively reveal the environmental adaptability and evolutionary potential of species, and provide scientific and effective basis for the formulation of protection measures for related species and the screening of core germplasm resources. Unlike animals, plants do not adapt to the changing natural environment by means of migration. The diversity of ecological environment is an important driving factor for the diversity of species genetic resources. Therefore, under the selection pressure of special ecological factors in different ecological environments, individuals of certain genotypes in the population have more competitive advantages, thus gradually occupying the ecological high position, and finally causing genetic differentiation among different populations9. As a natural medium for plants, soil is not only the main provider of plant nutrition, but also its physical and chemical properties and resource availability also affect the physiological and ecological characteristics of plants. At the same time, the adaptability to different soils drives the genetic differentiation among populations and affects the level of genetic diversity of populations. Turner’s study on the adaptability of Arabidopsis lyrata to different soils showed that differences in different soil environments can promote the genetic differentiation of populations of the same species by affecting specific adaptive genes, such as genes related to calcium-magnesium ratio or ion transport, thereby enhancing their adaptability to their respective growth conditions10. Other studies have found that areas with high soil potassium content can enhance the adaptability of some plants to arid environments, resulting in a certain genetic differentiation between these populations and other regional populations11. In the study of the Qinghai-Tibet Plateau, it was also found that the available nitrogen content in the soil had a significant effect on plant diversity and genetic structure at different altitudes12. Studies on the competition and adaptation between invasive plant communities and native plants have also shown that the physical and chemical properties of soil can change the distribution pattern and diversity of plants. Invasive plants can affect the population structure of native plants by changing the available potassium and nitrogen levels in soil, thus changing the level of genetic diversity of native plants13. Therefore, studying the relationship between environmental factors such as soil physical and chemical properties and population genetic diversity can not only reveal the adaptation mechanism of plants under different environmental conditions, but also provide important theoretical basis for the protection strategies of endangered species14. However, at present, the research on the genetic diversity of endangered species is mostly focused on a single molecular biology research, and there are few joint studies of molecular biology such as ecological factors and genetic diversity. Therefore, we investigates the genetic characteristics of six wild populations of O. microphylla using Hyper-seq sequencing technology, which is a novel, effective and flexible marker-assisted selection and genotyping method, aiming to uncover the molecular mechanisms underlying its endangered status. At the same time, the soil characteristics of different populations were compared and analyzed. Finally, an integrated analysis was conducted to explore the relationship between soil and other ecological factors and population genetic diversity, providing deeper insights into the interactive mechanisms between ecological factors and genetic diversity. This study not only helps to reveal the ecological adaptability and evolutionary potential of O. microphylla, but also provides a scientific basis for future ecological restoration, species ex situ conservation and germplasm resource management.
Results
Single nucleotide polymorphism (SNP) screening and genetic diversity
A total of 246,838 effective SNPs were obtained from 101 samples of O. microphylla, and upon filtering, 3,399 high-quality SNPs were obtained. The obtained SNP data were used to calculate the genetic diversity indices of the six populations (Table 1). The final dataset comprised 3,399 high-quality SNPs with no missing genotypes among the 101 individuals. Despite the absence of a reference genome, these SNPs were evenly distributed across all six populations, with each contributing a similar number, thus ensuring balanced representation and minimizing bias in downstream analyses of genetic diversity and structure. The private alleles (Np) of the six populations were in the range of 15,040 to 27,231, with the smallest Np in the JX population and the largest Np in the ZD population. The observed heterozygosity (HO) was in the range of 0.00144 to 0.0022, among which the LY population had the least HO and the ZD population had the largest expected heterozygosity (HE). The HE was in the range of 0.00094 and 0.00148, among which the HE of the JX population was the least and that of the ZD population was the highest. The nucleotide diversity (π) ranged from 0.00170 to 0.00240, with the smallest π was noted in the JX population and the largest in the ZD population. The genetic diversity level of the ZD population was the highest, and that of the JX population was the lowest. The genetic diversity of the six populations of O. microphylla was in the order of ZD > ZP > HS > YQ > LY > JX. To assess differences in genetic diversity (π) among populations, non-parametric comparisons were conducted using the Kruskal–Wallis test. The results revealed that the ZD population exhibited significantly higher genetic diversity than the other populations (P < 0.05), whereas the difference in π values between the LY and JX populations was not statistically significant (P > 0.05). Additionally, the LY population located within the Karst tiankeng exhibited significantly lower genetic diversity compared to the average level of populations outside the tiankeng (P < 0.05). The HE of six populations was higher than their HO, reflecting that selfing or inbreeding was common in the O. microphylla population; the FIS values of all six populations were > 0, further corroborating this phenomenon. Moreover, a positive FIS value suggests potential deviations from Hardy–Weinberg equilibrium within populations. One possible explanation is the Wahlund effect, which arises when undetected genetic substructure exists within a population. Differences in allele frequencies among these subpopulations can lead to an overall reduction in heterozygosity, resulting in an apparent heterozygote deficiency.
Phylogeny and genetic structure
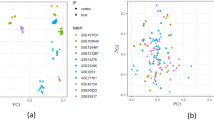
The UPGMA tree at the population level exhibited a separate branch of the LY population, while the other five populations were clustered into one branch, indicating a distant phylogenetic relationship of the O. microphylla individuals of the LY population with other populations (Fig. 1a). Bootstrap values were added to key nodes in the phylogenetic tree. Several major branches, including those involving the LY and ZP populations as well as the JX population, showed support values between 76 and 99, suggesting a moderate to high level of confidence in the tree topology. This phenomenon was also corroborated by the results of the principal component analysis. A clear population division was noted between individuals in the LY population and those in the other five populations (Fig. 1c, d). Additionally, the phylogenetic tree analysis at levels of population and individuals, showed a closer phylogenetic relationship between the two geographically close HS populations and YQ populations (Fig. 1a). Likewise, the two geographically close populations, JX and ZP, also showed a close phylogenetic relationship (Fig. 1a).
Phylogenetic analysis and principal component analysis of six populations of O. microphylla. (a): UPGMA tree among populations. (b): NJ tree among individuals. (c, d): Phylogenetic analysis and PCoA.
The mixed analysis showed that when K = 2, the six populations could be categorized into two groups, of which the LY population was one group and the other five populations were clustered into the second group (Fig. 2b). Only two individuals (LY6, LY12) in the LY population exhibited a mix of a small amount of genetic component of other groups. In contrast, within the second group comprising the other five populations, a small amount of genetic component of the LY population were also observed, but the hybrid component accounted for a large proportion of the genome of the individual. As the CV error reached its minimum at K = 7 (Fig. 2a), this value was considered the optimal number of genetic clusters for all O. microphylla individuals. The six O. microphylla populations were genetically divided into seven clusters. Although the K value exceeded the actual number of populations, this result may reflect underlying substructure within populations or differentiation from historical gene pools. This finding is consistent with the inference of positive FIS values, which also suggest the presence of genetic subdivision. Further analysis of the phylogenetic tree shows the categorization of the HS population into two groups, while the other populations belonged to one group each (Fig. 1b), indicated the presence of unrecognized genetic heterogeneity within the population. When K = 7, the genetic structure of individuals in LY population was also relatively simple compared with the other five populations, and the genetic components were relatively pure, with only a small degree of gene confounding observed (Fig. 2b). The population structure of the other five populations was complex, with distinct genetic components observed across them. The ZD population had the most complex genetic structure. Except for the primary genetic components of the LY population (blue module), the other four populations exhibited mixed genetic components (Fig. 2b). In the HS population, the red modules were the primary genetic component, and a significant proportion of red modules in the YQ and ZD populations were close to the geographical location of the population. In the ZP population, the green module constituted a significant proportion. Apart from the JX population, which is geographically close to ZP, the green module was present to a certain extent, whereas its proportion in other populations was minimal (Fig. 2b). Although both the phylogenetic tree (Fig. 1a) and population clustering analysis (Fig. 1b) assigned the LY population as a distinct group, results from PCA and admixture analysis revealed that certain LY individuals exhibited genetic admixture with other populations. This suggests a degree of discordance between individual-level gene trees and population-level assignments. Such inconsistency may be attributed to historical gene flow among populations or the retention of ancestral polymorphisms. These findings suggest that the differences in geographical location or living environment may potentially impact O. microphylla population structure and genetic composition.
Analysis of population structure. (a): CV error distribution for K from 1 to 10. K with the smallest CV value is marked in red. (b): The maximum likelihood phylogenetic tree for 101 O. microphylla.
Chemical properties of soil
In the soil environment harboring the six O. microphylla wild populations, the total organic carbon (TOC) and Fe levels were significantly higher than those of other substances (Fig. 3a). The soil of the YQ population had the highest TOC content (134.4 g/Kg), while the JX population had the lowest content (39.0 g/Kg). In addition, the contents of total nitrogen (TN) (5.887 g/Kg) and available nitrogen (AN) (0.587 g/Kg) in the soil harboring the YQ population were also significantly higher than those in soil harboring other populations (P < 0.05) (Fig. 3a, c). The Fe content was highest in the soil of the LY population (75.133 g/Kg), followed by the ZP population (58.367 g/Kg), and the lowest in the soil of the JX population. At the same time, the Ca content in the soil of the LY population (13.233 g/Kg) was also much higher than the soil harboring other populations, and the gap with the ZD population was nearly 15-fold (0.900 g/Kg), and ranked second in Ca content. The soil pH of the LY population was also differed significantly from that of other populations (Fig. 3b). The soil of other five populations was strongly acidic with pH between 3.6 and 4.3, while that harboring the LY population was neutral (pH = 7.1). The contents of K elements such as total potassium (TK) (20.767 g/Kg) and potassium (K) (18.500 g/Kg) in the soil harboring the ZD population were significantly higher than soil harboring other populations (P < 0.05) (Fig. 3a). In addition, the content of total phosphorus (TP) (0.746 g/Kg) in the soil of this population was also significantly higher than that harboring other populations (P < 0.05) (Fig. 3c). Thus, the chemical properties of the soil harboring the LY population were quite different from those of other populations, suggesting potential adoptive evolution of the O. microphylla population in this area to adapt to the special soil environment of Tiankeng. Likewise, compared with other populations, similar chemical properties were also noted for the soils of the two geographically close populations of YQ and HS.
Soil chemical properties of 6 O. microphylla populations. (a): Overview of soil physical and chemical properties of six O. microphylla populations. (b): soil pH of six O. microphylla populations. (c): Available potassium (AK), available nitrogen (AN), total phosphorus (TP) content in soil of six populations.
Cluster analysis and principal component analysis
Based on the soil characteristics data, cluster analysis and PCA were conducted on six O. microphylla populations to show the uniqueness and difference of soil chemical characteristics of different O. microphylla populations more intuitively and clearly. The cluster analysis showed that the soil characteristics harboring the three populations of HS, ZD, and YQ were nearly similar, and were first clustered together in one group (Fig. 4a). Then, these three populations were clustered with the ZP population, indicating similar soil characteristics of the HS, ZD, and YQ populations with those of the ZP population. LY and JX populations were clustered separately on one group each, suggesting that the chemical properties of the soil harboring the two populations differed significantly from those of the other four populations; concurrently, the two populations differed significantly. PCA also showed high similarity between the three populations of HS, ZD, and YQ, while LY and JX populations were segregated far from the other four populations (Fig. 4b). Among them, the distance of the LY population from other populations in PC1 (57.62%) and PC2 (24.33%) was large, which further confirmed the particularity of soil environment in Tiankeng.
Cluster analysis (a) and principal component analysis (PCA) (b) of soil chemical characteristics of six O. microphylla populations.
Genetic differentiation, isolation by distance (IBD) and isolation by environment (IBE) detection
The range of genetic differentiation coefficient between the six O. microphylla populations was from 0.220 (HS-YQ, HS-ZD, and ZD-YQ) to 0.620 (JX-LY), among which the LY population had the highest degree of differentiation from the other five populations (FST = 0.520–0.620); furthermore, the lowest degree of differentiation from other populations was observed for ZD (FST = 0.22–0.520) (Fig. 5a). IBD detection and IBE detection were performed to further explore the driving factors of genetic differentiation.
Genetic differentiation analysis (a), isolation by distance (IBD) test (b) and isolation by environment (IBE) test (c).
IBD detection of the six O. microphylla populations showed no significant correlation among populations in terms of genetic distance and geographical distance (P = 0.169, R = 0.300). However, IBD detection was carried out on the other five populations in this study due to the special Tiankeng habitat of LY population. After excluding LY populations, a significant positive correlation was noted between genetic and geographical distances among the other five populations (P = 0.034, R = 0.697) (Fig. 5c). The findings indicated that the greater the geographical distance among O. microphylla populations in ordinary habitats, the greater was their degree of genetic differentiation. The IBD test revealed that the special habitat of Tiankeng as well as long-distance geographical isolation were the primary reasons for the genetic differentiation of O. microphylla population. IBE results showed a significantly positive correlation of FST with TK and K in soil (R ≥ 0.4, P < 0.01) (Fig. 5b), also suggesting that K content in soil possibly promotes genetic differentiation among O. microphylla populations.
Discussion
Dual impacts of tiankeng environments and population history on genetic diversity in O. microphylla
Population genetics states that the species with a wider distribution have higher genetic diversity and are accordingly highly adaptable to the environment. In contrast, the narrower the species distribution, the lower the genetic diversity, the lower is the ability to adapt to the environment15. Karst tiankeng is a special geological environment distributed in the karst area. The tiankeng has large humidity, low temperature, and high negative oxygen ion concentration because of the cliffs around the tiankeng or most of the surrounding walls, facilitating the formation of unique hydrothermal conditions different from the surrounding areas16. However, compared with the changeable and complex external environment, that inside the tiankeng is relatively stable and single. In this study, we compared the genetic diversity of O. microphylla populations inside and outside the tiankeng. The genetic diversity of LY O. microphylla population inside the tiankeng (π = 0.00172) was lower than that outside (π = 0.00202), but the difference in genetic diversity compared to the JX population (π = 0.00170) was not statistically significant. This suggests a strong selection pressure of the unique living environment inside the tiankeng on the O. microphylla population. During the adaptation process, only a small number of genotypes were retained, consequently reducing the overall genetic diversity of the population. The ‘island effect’ due to the negative topographic environment impedes the communication between populations and makes the update of the population gene pool in the tiankeng difficult, and this may also be one of the reasons of lower genetic diversity of O. microphylla population in the tiankeng than that outside. Furthermore, the level of genetic diversity of species may also be potentially affected by the diversity of population history17. The species outside the Karst tiankeng migrate in succession with tiankeng’s natural evolution18,19; besides, the founder effect of this accidental migration on the genetic diversity of the population cannot be undermined. The founder effect occurs when a small group is separated from the original population and results in the establishment of a new population in a new, isolated environment; the genetic diversity of the new population is generally low due to the limited gene pool of the founder group20. Therefore, the limited gene pool of the O. microphylla Tiankeng population may be one of the possible reasons for its low genetic diversity. Zhu et al.21 studied the genetic characteristics of Paphiopedilum concolor and P. hirsutissimum, and found that most of the genetic diversity indices of P. concolor were less than those of P. hirsutissimum; in fact, there were significant bottleneck effects in some populations of P. concolor. Therefore, this study suggests that the bottleneck experienced by the P. concolor population may be the cause of the decrease of its genetic diversity. Notably, all six O. microphylla populations examined in this study exhibited positive Tajima’s D values. Combined with field survey data indicating that natural populations have drastically declined due to illegal logging and habitat destruction-with many historical distribution sites now lacking surviving individuals and existing populations showing clear signs of fragmentation-these results suggest that the species has undergone a pronounced anthropogenic bottleneck. In addition to this human-induced bottleneck, the positive Tajima’s D values, significant genetic differentiation revealed by phylogenetic and population structure analyses, and the lack of a significant IBD pattern among non-LY populations collectively suggest that the unique evolutionary trajectory of the LY population is largely driven by geographic isolation imposed by the Karst tiankeng landform. The tiankeng acts as a natural barrier, restricting gene flow and amplifying the effects of genetic drift, ultimately leading to marked genetic divergence between the LY population and those outside the tiankeng. These findings indicate that the evolutionary history of the LY population may be shaped by a combination of founder effects and geographic isolation. Within this unique microhabitat, multiple factors-including strong environmental filtering, founder effects, anthropogenic bottlenecks, and restricted gene flow-may have jointly contributed to the reduced genetic diversity observed in the LY population.
The transmission characteristics also indicated prevalent selfing or inbreeding in the six O. microphylla populations. In the ex-situ germplasm conservation nursery, long-term observations revealed that some O. microphylla individuals were able to undergo normal flowering and fruit set even under solitary planting conditions or when only a single flowering individual was present. This suggests that the species may possess a certain degree of self-fertilization capability. This mating strategy usually led to reduced genetic diversity and inbreeding depression22,23. For example, offspring derived from self-fertilized populations may exhibit a certain proportion (approximately 4%) of albino seedlings24. Zhou et al.25 also reported the prevalence of selfing (FIS > 0) in the population of O. henryi in southern China. Interestingly, there was moderate genetic diversity of the six O. microphylla populations. This finding is consistent with observations reported by Li et al.24. in Ormosia hosiei, where relatively high levels of genetic diversity (HE > 0.720) were maintained despite the presence of inbreeding within populations. Similarly, Zhang et al.26 noted that even populations of O. hosiei with small census sizes did not exhibit a marked decline in genetic diversity, suggesting that this species possesses a strong capacity to retain genetic variation under conditions of population reduction. Tajima’s D > 0 suggests that the population may have experienced a bottleneck effect27, and also that the population may have encountered a balanced selection effect28. The balanced selection effect (Tajima’s D > 0) exhibited a positive effect on maintaining the genetic diversity level of the population29. Intra-population selfing or inbreeding increases the proportion of homozygotes; however, the balanced selection effect may maintain the genotypes of heterozygotes with adaptive advantages, consistently maintaining certain genetic diversity of the population30. The balanced selection effect can also reduce the impact of genetic drift by maintaining multiple alleles, suggesting that some alleles may not be completely lost even in inbred populations, due to the presence of balanced selection.
Geographic isolation and ecological selection drive genetic differentiation and structural diversity in O. microphylla populations
The exploration of genetic structure is highly significant to the formulation of strategies for endangered species protection31. Both phylogenetic and genetic analyses indicate that the LY population within the Karst tiankeng has undergone substantial genetic differentiation from the other five populations, with which it shares a more distant genetic relationship. However, PCA and admixture analyses revealed that a few individuals within the LY population still exhibit close genetic affinities and signs of admixture with individuals from other populations. Such patterns are commonly observed in natural populations, particularly in cases where geographic isolation or ecological barriers coexist with historical gene flow. Gene flow is usually more frequent among populations that have closer geographical location, while gene exchange is limited when populations are located at farther geographical distance, and genetic differentiation usually increases when the with increasing distance32. However, we found that the tiankeng population still exhibited substantial genetic differentiation from geographically proximate populations. Based on the results of the IBD analysis, we suggest that the unique environmental conditions of the tiankeng intensify geographic isolation among O. microphylla populations, while reducing the influence of geographic distance alone on genetic differentiation.
In addition, population structure analysis indicated that K = 7 was the optimal number of genetic clusters for the six populations. Furthermore, all populations exhibited positive FIS values, and we hypothesize that the observation of K exceeding the actual number of populations may be partially attributed to the Wahlund effect, which reflects potential genetic substructure within populations. However, the pronounced genetic differentiation of the LY population from the others, along with its distinct phylogenetic branch (Fig. 1a, b), provides stronger evidence for the dominant role of geographic isolation rather than within-population substructure. Therefore, although the Wahlund effect may have introduced some degree of clustering noise, we conclude that the primary drivers of genetic differentiation in O. microphylla are geographic isolation and adaptive evolution. Except for the LY population, the individual genetic components of the other five populations were more complex, with multiple populations within one population. The PCA analysis also showed the chaotic distribution of the individuals of other populations except for the tiankeng population (LY), which was significantly different from other populations.
Except for the tiankeng population, which exhibits a clearly distinct genetic composition from the other populations, the remaining populations show relatively complex genetic backgrounds. We propose two possible explanations for this pattern. While the O. microphylla population was distributed on a large scale in Guangxi, but due to subsequent climate change or human intervention, the scale gradually shrunk to establish many small populations. Although the populations were gradually separated in the case of species, their gene banks still retained a large amount of similar genetic information. This historical association shows that the population seems to have no significant distinction in genetic structure and PCA. Forgiarini et al.33 also noted this phenomenon when their study explored the genetic characteristics of Iris plants endemic to South America. Even in the case of severe fragmentation of the habitat, some populations still maintained similar genetic structure. However, as the population size shrinks, there is a gradual increase in the effect of random genetic drift on these small populations, eventually increasing genetic differentiation among populations34. The significant IBD results also suggest that the gene flow of the fragmented O. microphylla population decreases with the increase in geographical distance, making the complete elimination of the cumulative effects of this genetic differentiation difficult35. On the other hand, this phenomenon may also be due to the adaptive evolution of the population and the balanced selection effect (Tajima’s D > 0). The IBE results in this study showed significant effect of three ecological factors on the genetic differentiation of O. microphylla population.
This result indicates that different ecological factors exert varying selection pressures among different populations, driving the adaptive differentiation of specific gene loci in each population under different directions36, thus enhancing the genetic differentiation among populations. The endangered plant populations are often more vulnerable to genetic drift and random events37, although balanced selection can maintain the diversity of different alleles to a certain extent38. The maintenance of this diversity increase the complexity of genetic components among different individuals. Therefore, when analyzing genetic structure, the population structure shows a mixed phenomenon. In summary, the combined effects of adaptive differentiation and geographical isolation in O. microphylla population have driven gene frequency divergence between populations and increased the genetic differentiation among populations. The gene frequency diversity resulting from equilibrium effect and random drift account for the complex genetic structure of the population.
Iron deficiency stress and selection of iron tolerance
In this study, the study of the soil characteristics of O. microphylla wild populations revealed primary distribution of O. microphylla populations mostly in soils with relatively high TOC and Fe content. The soil can be broadly divided into four grades based on the TOC content as: TOC content < 1%, the soil nutrients are very poor, making it poorly fertile; TOC = 1–2%, the soil has certain fertility to meet the healthy growth of plants; TOC = 2–4%, the soil has a strong nutrient supply capacity; TOC > 4%, the soil has excellent fertility and water retention capacity, this is usually found in undisturbed natural ecosystems, including forests, or organic fertilized soils39. In the six O. microphylla populations examined in this study, the TOC in the soil was > 4% (40 g/kg), suggesting the high content of soil organic matter as an important basis for the growth and development of O. microphylla. Therefore, for the introduction and cultivation work to be performed in the future, soil with high TOC should be selected for cultivation or should be regularly supplemented with organic matter.
In addition to the high TOC content, the soil Fe content of the six wild populations was also significant. Although Fe is the third most abundant mineral element in the crust of the earth and is highly abundant in the soil, it mainly exists in the form of Fe3+ which has low bioavailability, making it difficult for plants to absorb and utilize40. The activity of Fe in soil is significantly impacted by the redox potential (Eh). In long-term flooded soil, the Eh decreases due to the lack of dissolved oxygen, leading to the abundance in Fe2+ with high solubility and better bioavailability41. Most areas of Lingnan, China, where O. microphylla grows, is primarily red loam with poor water permeability. The region experiences abundant precipitation, causing flooding of the soil, leading to serious soil acidification over time42. This study also confirmed that the soil harboring the other five populations was strongly acidic, except that harboring the LY population. The O. microphylla populations surviving in this acidic soil environment may already have strong Fe tolerance and adaptability, enabling them to avoid toxic reactions caused by excessive iron. The extent and molecular mechanism involved in plant tolerance to Fe toxicity are critical not only for the growth and development, but also possibly influence the genetic differentiation of the population. In agricultural practices, K fertilizer is commonly used to improve the symptoms of Fe toxicity in plants; however, its molecular mechanism is not yet fully elucidated41. Zhang et al.43 reported that the steady-state balance of K ions in the root tip region is an important mechanism facilitating the plants roots to tolerate Fe toxicity. Likewise, Wu et al.44 found that the K channel gene OsAKT1 (Arabidopsis K+ transporter 1) expressed in rice roots affects the process of tolerance of rice plants to to Fe toxicity by influencing the K+ concentration gradient inside and outside the root xylem. Interestingly, the IBE test also revealed that K and TK in the soil significantly promoted the genetic differentiation among O. microphylla populations in. The findings suggest that the soil K potentially enhances the tolerance of O. microphylla to Fe in the soil by regulating a few physiological processes, such as the ionic equilibrium in the roots, while the variations in soil K content of different populations render the populations tolerant to Fe toxicity. Different environmental selection pressures are exerted on distinct populations, which may lead to adaptive genetic variation and promote genetic differentiation among populations.
The characteristics of the soil harboring the LY population were significantly different from those harboring other populations, as reflected in the soil characteristics cluster analysis (Fig. 4a) and PCA analysis (Fig. 4b). The soil harboring this population was neutral, and the Fe3+ content remained at a high level. In alkaline calcareous soils, high pH and carbonate content significantly reduced Fe solubility, making the absorption of the required Fe difficult for plants. Under these conditions, Fe usually exists in its insoluble form, as Fe3+, which is difficult to be absorbed and utilized by plants, consequently increasing the vulnerability of plants to Fe deficiency stress45. Fe is an important trace element required for photosynthesis, respiration, chlorophyll synthesis, redox reaction, nitrogen fixation, as well as other physiological and biochemical reactions that help in the growth and development of plants46,47. Nonetheless, the strategies of plants to cope with an environment deficient of Fe vary with different species. Usually, dicotyledonous plants and non-gramine monocotyledonous plants (such as Glycine max, Arachis hypogaea, etc.) adopt the reduction mechanism for absorption. In the Fe-deficient environment, the low-soluble Fe3+ content in the soil is reduced to Fe2+ by Fe-chelating reductase (FCR), which is absorbed by plants easily48. This study also demonstrates that the K and TK content in the soil harboring this population was significantly lower than that harboring other populations. The above results lead us to speculate that the individuals of this population were not affected by Fe toxicity compared to other populations, however, they were in a state of Fe deficiency stress. The individuals belonging to LY population may have eliminated the individuals lacking the mechanism to overcome the Fe-deficient environment through natural selection, thus adapting to this unique environment. The study findings also suggest that the LY population may have undergone significant genetic differentiation compared to other O. microphylla populations.
Although we did not include a common garden experiment and thus cannot directly distinguish the relative contributions of genetic differentiation and phenotypic plasticity, the significant genetic structure observed among populations and the results of environmental association analyses suggest that the differentiation of O. microphylla populations is more likely driven by long-term natural selection rather than solely by phenotypic plasticity. First, phylogenetic and population structure analyses revealed that the LY population within the tiankeng has formed an independently evolving lineage, consistent with the "Island effect," and its genetic homogeneity aligns closely with the geographic isolation that limits gene flow. Second, variation in soil potassium (K) and total potassium (TK) levels may impose selective pressures-such as those acting on genes involved in iron or potassium metabolism-leading to the fixation of adaptive alleles. For instance, the neutral soil environment in the LY population may have selected for genotypes tolerant to iron deficiency, whereas populations in acidic soils may have retained mechanisms adapted to iron toxicity. Additionally, population-specific signatures of balancing selection (Tajima’s D > 0) further support historical selective pressures as the underlying cause of differentiation, rather than short-term phenotypic responses. However, due to the absence of a fully assembled reference genome for O. microphylla, we are currently unable to precisely localize or functionally annotate candidate loci potentially under selection, nor can we perform standard outlier tests based on genomic coordinates. In future studies, we plan to integrate whole-genome assembly and annotation with genome-wide scans for selection, and combine these analyses with common garden experiments and gene expression profiling. This will enable a more comprehensive understanding of the molecular mechanisms by which soil factors drive adaptive population divergence.
Methodological caveats: revisiting the neutral theory framework
Our study adopts commonly used population genetic parameters such as FST and Tajima’s D to assess genetic diversity and differentiation among O. microphylla populations. These methods, however, are based on assumptions derived from the neutral theory of molecular evolution. In recent years, the explanatory power of this theory has been increasingly questioned. For instance, Lynch et al.49 provided strong evidence that observed levels of genetic diversity in real populations are inconsistent with neutral expectations. Moreover, recent findings suggest that the human mitochondrial genome is predominantly functional rather than neutral50, and that short tandem repeats (STRs), once thought to be evolutionarily neutral, are involved in transcriptional regulation51.
As the neutral theory was historically linked to the molecular clock hypothesis—which itself has lost much empirical support—the continued use of neutrality-based models in population genetics calls for caution. While our analyses provide valuable insights into genetic structure and differentiation, the interpretations should be considered provisional given these theoretical limitations. We acknowledge that the observed patterns may not be fully explained under a neutral framework alone. Future studies could benefit from integrating functional genomic approaches and considering alternative models, such as the Maximum Genetic Diversity (MGD) theory, to better understand the evolutionary mechanisms shaping diversity in natural populations.
Protection suggestions and measures
In light of the increasingly severe population crisis, implementing scientific protection measures based on genetic diversity and genetic structure is crucial for guiding practical efforts52. Therefore, based on the research findings. Accordingly, this study puts forward relevant suggestions for the protection of O. microphylla population: (1) For the introduction and cultivation of O. microphylla, using soils with high organic matter content and high drainage is recommended, such as the mixture of humus and sandy soils, coconut coir, perlite, and compost mixture. These combinations can ensure a high content of organic matter, concurrently also increasing the drainage and K content of the soil and avoiding the Fe toxicity due to root hypoxia. The introduction of K-rich plants in the introduction and cultivation area can help enhance the potassium content in the soil. Nevertheless, in the natural environment, the application of K fertilizer to the O. microphylla population should be avoided and supplemented only when there is a significant lack of K content in the soil, to safeguard the natural selection process and ecological adaptability of the population from excessive K content. (2) The LY population, characterized by unique genetic distinctiveness, warrants prioritized in-situ conservation. Meanwhile, the ZD population serves as a critical genetic diversity reservoir, where tissue culture and cutting propagation should be scaled to boost population size. Coupled with artificial pollination, ex-situ conservation, and genetic monitoring, these strategies will enhance inter-population gene flow and adaptive resilience. This integrative approach establishes an evolutionarily informed conservation network, ensuring ecological sustainability for this endangered species. (3) In the cases of population fragmentation or isolation, the selfing characteristics of O. microphylla population may result in reduced genetic diversity. To enhance genetic diversity and adaptability in O. microphylla populations, it is essential to select genetically similar populations such as ZD, YQ, and HS for artificial or off-site pollination and introducing exogenous genes. This method can help to improve the genetic diversity and adaptability of offspring. Additionally, implementing ex-situ conservation strategies for different populations, establishing a long-term detection system for the population, regularly evaluating its genetic diversity and genetic structure changes, and adjusting protection measures in time are crucial. Concurrently, the genetic structure analysis in this study also proved only a little difference in the gene pool between different O. microphylla populations, which also provided the possibility of hybrid breeding. (4) The preliminary investigation of the O. microphylla population in this study revealed the distribution of most populations in the economic forest outside the reserve, making them susceptible to damage from artificial logging. Additionally, the stem borer infestation of the O. microphylla has been very severe, with more than 90% of plants showing damage (Fig. S1). When the trunk is invaded by pests, the gum secreted by the trunk can easily cause the foraging of other insects, such as Vespa, Polistes sp (Fig. S2). Based on this characteristic, the local villagers often place beehives near the tree species. However, excessive collection of gum not only causes the trees to secrete gum repeatedly, depleting their own nutrition and water, affecting normal growth and reproduction, but also interferes with their self-healing process, weakening the tree defense and causing the aggravation of pests. Therefore, establishing protected areas, preventing and controlling pests and diseases, and implementing protection and publicity work are also top priorities.
Conclusion
Integrating high-throughput Hyper-seq genotyping and soil physicochemical analyses, we systematically investigated six wild O. microphylla populations across southern China. Our results revealed moderate species-level genetic diversity with substantial interpopulation heterogeneity. The ZD population exhibited the highest genetic diversity, contrasting sharply with the lowest levels in JX. Notably, the Karst tiankeng-confined LY population showed reduced genetic diversity compared to extra-tiankeng populations, suggesting combined effects of habitat-specific selection pressures and founder effects. Genetic structure analysis demonstrated significant divergence of LY population, reflecting its homozygous genetic composition shaped by prolonged environmental stability in this isolated ecosystem. Soil analyses revealed distinct edaphic conditions for LY population, particularly in pH and Fe content, indicating potential iron deficiency stress. Conversely, other populations in strongly acidic soils faced elevated iron toxicity risks. IBE analysis identified soil potassium (K, TK) as key drivers of genetic differentiation. All populations showed moderate inbreeding, potentially constraining evolutionary adaptive capacity. Accelerating habitat fragmentation, anthropogenic disturbances, and genetic erosion collectively threaten population viability, necessitating urgent conservation interventions.
While this study systematically investigated the genetic-environment interactions in O. microphylla populations, three critical limitations warrant attention. First, the absence of common garden experiments precludes definitive discrimination between genetically driven and phenotypically plastic trait variations, thereby constraining mechanistic validation of adaptive differentiation. Second, restricted sampling coverage may underestimate genetic diversity in cryptic or peripheral populations. Third, functional linkages between candidate genes and adaptive traits remain unresolved. To address these gaps, we propose: (1) Multi-site common garden trials coupled with transcriptomics profiling to disentangle genetic adaptation from epigenetic regulation; (2) Expanded population sampling incorporating edge habitats; (3) Longitudinal genetic monitoring to track spatiotemporal dynamics of population structure. This work establishes foundational data for conserving O. microphylla (a Category I nationally protected species), informing regional habitat restoration strategies and national conservation frameworks. Globally, it advances understanding of genetic diversity maintenance in karst ecosystems, particularly regarding extreme edaphic selection and isolation-driven evolutionary trajectories.
Materials and methods
Materials
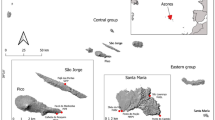
Six natural populations of O. microphylla were distributed in southern China, of which only one population was distributed in Guizhou Province, and the other five populations were distributed in Guangxi Zhuang Autonomous Region (Fig. 6 and Table 2). It is worth noting that the LY population is located in the Karst tiankeng, and its geographical environment is significantly different from other populations. The plant materials used in this study were collected and identified by associate researcher Jianmin Tang and Wei Xiao (Guangxi Botanical Research Institute, China). Permission for the collection of plant materials was obtained from the local Forestry Bureaus and relevant nature reserve authorities prior to sampling. One sample of each population was preserved in the Herbarium of Guangxi Botanical Research Institute (China). The identification numbers are assigned sequentially as 20230411GXOMJT01-20230411GXOMJT06.
The distribution map of 6 populations of O. Microphylla. (LY populations are distributed in Karst tiankeng. LY, HS, YQ, ZD, JX and ZP are the abbreviation of sampling site, the full names are shown in Table 2. The map was generated using ArcGIS Desktop v.10.8.1 (Esri Inc., Redlands, CA, USA; https://www.esri.com/en-us/arcgis/products/arcgis-desktop/overview). All spatial data were manually processed and visualized by the authors. The base layers used in this map are open-source and publicly available. No copyrighted materials were used or modified in the creation of this figure.)
Collection of plant samples
The leaf samples were collected from 101 O. micropylla individuals from six populations. All the samples were collected from healthy, mature individuals to best represent the genetic diversity of each population. To minimize the likelihood of sampling clonally propagated individuals or closely related neighbors, we maintained a minimum distance of 10–20 m between sampled plants wherever possible, depending on the terrain and plant density. Sampling was conducted to capture the spatial and ecological range of each population, covering both central and marginal microhabitats. Fresh leaves were immediately dried in discolored silica gel to preserve DNA quality for downstream molecular analyses. The plant materials collected in this study are in compliance with Chinese and international guidelines and laws, including the IUCN Policy Statement on Research Involving Species at Risk of Extinction and the Convention on the Trade in Endangered Species of Wild Fauna and Flora.
DNA extraction and Hyper-seq library construction
Genomic DNA was extracted from 101 O. micropylla leaf samples using the Tissue DNA Kit (Omega Bio-Tek, USA) following the manufacturer’s protocol. The quality of the extracted DNA was evaluated by 1% agarose gel electrophoresis using a Bio-Rad PowerPac™ Basic electrophoresis system (Model: 164–5050), which allowed visualization of DNA integrity and potential degradation. The quantity and purity of DNA were measured using both a NanoDrop 2000 spectrophotometer (Thermo Fisher Scientific, USA) and a Qubit 3.0 Fluorometer (Thermo Fisher Scientific, USA) to ensure accuracy. Only samples that met the following criteria were retained for library construction: total DNA mass > 2 μg, concentration > 30 ng/μL, and OD260/OD280 ratio between 1.80 and 2.00. Barcode adaptors were then ligated to the qualified DNA samples, followed by fragment size selection and PCR amplification to construct a Hyper-seq library. The constructed Hyper-seq library was sequenced by Wuhan Benagen Technology Co., Ltd. (Wuhan, China) using the Illumina NovaSeq 6000 platform (Illumina, San Diego, USA), generating 150-bp paired-end reads.
SNP calling
SNP calling was performed using the ustacks, cstacks, sstacks, tsv2bam, and gstacks modules of Stacks v2.6.4 in turn53. As no reference genome is available for O. microphylla, the analysis followed a de novo assembly pipeline. The main parameters are as follows: the minimum depth required for the original reads to form stacks is 3; the maximum number of base differences between different stacks at the formation site was 5; the number of allowed mismatches between sample sites when constructing a catalog is 3. Finally, the gstacks module of Stacks v2.6.4 was used to obtain a total of 246,838 raw SNPs from 101 O. microphylla samples. VCFtools v0.1.11 was used to filter SNPs by removing loci with a population missing rate > 40%, minor allele frequency (MAF) ≤ 0.05, and sequencing depth < 3, resulting in 3,399 high-quality SNPs used for downstream analyses53. Quality control metrics for each sequencing sample, including raw and clean read counts, data volume, and base quality scores (Q20/Q30), are summarized in Table S1.
Genetic diversity analysis
The population module in Stacksv2.6.4 software was used to calculate the genetic diversity indexes of various groups, including nucleotide diversity (π), number of private alleles (Np), expected heterozygosity (HE), observed heterozygosity (HO), inbreeding coefficient (FIS), and the average value of each index was calculated using all loci (variant and fixed). The Tajima 'D value of each population was calculated using Vcftools v0.1.13 software, and the sliding window was set to 3,000 bp. To assess whether π differed significantly among O. microphylla populations, we performed a nonparametric Kruskal–Wallis H test followed by pairwise post hoc comparisons when applicable. All statistical analyses were conducted using SPSS v. 26.0 (IBM Corp., Armonk, NY, USA), with a significance threshold set at P < 0.05.
Phylogenetic analysis, PCA analysis and population genetic structure analysis
The maximum likelihood algorithm of IQtree v2.0 is used to construct the phylogenetic tree between individuals and groups, with the best-fit nucleotide substitution model selected automatically using ModelFinder (parameter: -m MFP), and branch support evaluated by 1,000 ultrafast bootstrap replicates (-bb 1000) and SH-aLRT test (-alrt 1000); the resulting tree is processed and visualized by Figtree software54. The principal component was performed by VCF2PCACluster software with default settings, using the filtered SNP matrix and eigen decomposition to derive the major principal components55. The population structure of the samples was analyzed by Admixture v1.3.0 software, with linkage disequilibrium (LD) pruning performed using PLINK v1.9 (–indep-pairwise 50 5 0.2). Clustering was performed by assuming the number of clusters (K value = 1–10), each repeated five times, and the CV error under different K values was calculated56.
Collection of soil samples
In each O. microphylla plot, five healthy and well-grown adult individuals were randomly selected for soil sampling. One independent soil sample was collected from the rhizosphere of each selected tree, resulting in a total of 30 soil samples across all populations (6 populations × 5 individuals per population). Before sampling, the surface vegetation and debris were removed, and the surface soil around the O. microphylla individual (10–30 cm from the surface) was dug using a pollution-free shovel. The soil samples in the habitat were mixed and marked. After the soil samples are naturally dried in the air, they are ground and sifted to remove coarse particles and debris. The treated samples are stored in a dry, shady environment to prevent moisture and deterioration.
Determination of soil physical and chemical properties
The pH of the soil was measured using a pH meter (pHS-25, China Quzhou Aipu Measurement Instrument Co., Ltd.), which is usually measured in a soil–water suspension with a water-soil mass ratio of 2.5:1. The physical and chemical properties of the soil were determined according to the ' soil agrochemical analysis method’57: the total organic carbon (TOC) in the soil was determined by potassium dichromate oxidation method ; the total nitrogen (TN) content in soil was determined by Kjeldahl method. Alkaline-N (AN) in soil was determined by alkaline hydrolysis diffusion method. Soil total phosphorus (TP) was determined by sodium hydroxide alkali fusion-molybdenum antimony anti-colorimetric method. Soil total potassium (TK) was determined by NaOH fusion-flame photometer. Available potassium (AK) in soil was determined by neutral ammonium acetate extraction (1 mol·L-1) -flame photometer method. The contents of K (K2O) and Na (Na2O) in soil were determined by flame photometric method. The contents of Ca (CaO), Mg (MgO) and Fe (Fe2O3) were determined by atomic absorption spectrometry.
Data processing and statistical analysis
Statistical analyses of soil physicochemical properties were conducted using SPSS v. 26.0 (IBM Corp., Armonk, NY, USA). Prior to analysis, the normality and homogeneity of variances were assessed using the Shapiro–Wilk test and Levene’s test, respectively. For variables that met the assumptions of normality and homoscedasticity, one-way analysis of variance (ANOVA) followed by Tukey’s HSD post hoc test was performed. For variables that violated these assumptions, the Kruskal–Wallis H test followed by Dunn’s post hoc test was applied. All statistical tests were two-tailed, with a significance threshold set at P < 0.05.
Principal component analysis (PCA) and hierarchical clustering were performed based on soil physicochemical properties to explore overall differences in soil characteristics among populations. Data visualization was conducted using the ggplot2 and factoextra packages in R.
Genetic differentiation analysis and IBD, IBE detection
The population module of Stacks was used to calculate the genetic differentiation coefficient (FST) between the two populations. Based on the geographic coordinate information of the 6 population, the actual geographic distance was calculated using the R package geosphere58. According to the MaxEnt model, the R was used to calculate the importance of 30 indicators such as 19 bioclimatic variablesdownloaded from the WorldClim database (http://worldclim.org/) and 11 soil characteristic indicators in this study21. At the same time, the Spearman correlation coefficient (R) between the 30 environmental variables was calculated by SPSS software to remove the redundant variables. When the absolute value of the correlation between the two variables was greater than 0.8, one of the more important variables was retained22. Finally, 18 representative environmental variables were determined : Annual mean temperature (BIO1), Maximum temperature of warmest month (BIO5), Minimum temperature of coldest month (BIO6), Temperature annual range (BIO7), Mean temperature of driest quarter (BIO9), Mean temperature of warmest quarter (BIO10), Precipitation of driest month (BIO14), Coefficient of variation precipitation seasonality (BIO15), Precipitation of wettest quarter (BIO16), Precipitation of warmest quarter (BIO18), Precipitation of coldest quarter (BIO19), AK, AN, TP, TN, pH, Ca, Na. Prior to conducting correlation analyses of soil variables, potential outliers were identified using boxplots and Z-score methods. In cases where extreme outliers were detected, robust regression based on the Theil-Sen estimator was applied as a supplementary validation step to ensure the robustness of the results. Finally, Mantel test was performed on FST, geographical distance, and 12 representative environmental variables, Spearman correlation coefficient was calculated, and 9999 permutation tests were performed.
Data availability
The data can be obtained by contacting Yishan Yang (yangyishan0113@163.com). Raw sequencing data are available at NCBI with the SRA accession number of PRJNA1214912 (https://www.ncbi.nlm.nih.gov/bioproject/PRJNA1214912).
References
Wu, Z. Y., Raven, P. H. & Hong, D. Y. Flora of China Vol. 10, 73–85 (Science Press, Beijing, 2010).
Wang, X. D., Liu, P., Liu, M. J., Xiao, X. Y. & Chen, F. S. Biology and ecology research status of Ormosia species in China. Plant Sci. J. 36, 440–451 (2018).
Yang, H. Y., Feng, B. X., Yuan, M. Q. & Yang, J. W. Analyses on population pattern and endangered causes of rare and endangered species Ormosia micropylla. Seed 32, 52–54 (2013).
Luo, Z. Y. & Yang, C. H. In Rare and precious tree species in Guizhou, 129–134. Guizhou People 's Publishing House, Guiyang (In Chiese).
Xiao, Z. H. Resource investigation and seeding rearing technology of Ormosia micropylla in Hunan Province. Master’s Thesis, Central South University of Forestry and Technology, Changsha, China, (2023).
National Forestry and Grassland Administration, Ministry of Agriculture and Rural Affairs. List of National Key Protected Wild Plant Species. Beijing (2021).
Feng, B. X., Wang, D. J., Yang, J. W. & Wu, K. M. Study on population composition and structure of Ormosia micropylla located in Nanshao. Jianhe County. Seed 31, 65–68 (2012).
Tang, J. M., Qin, H. Z., Liang, H. & Wei, X. Study on seedling quality classification of Ormosia micropylla. Spec. Econ. Anim. Plant 26, 40–42 (2023).
Silva-Arias, G. A., Caballero-Villalobos, L., Giudicelli, G. C. & Freitas, L. B. Landscape and climatic features drive genetic differentiation processes in a South American coastal plant. BMC Ecol. Evol. 21, 196 (2021).
Turner, T. L., von Wettberg, E. J. & Nuzhdin, S. V. Genomic analysis of differentiation between soil types reveals candidate genes for local adaptation in Arabidopsis lyrata. PLoS ONE 3, e3183 (2008).
Jadwiszczak, K. A., Mazur, M., Bona, A., Marcysiak, K. & Boratyński, A. Soil requirements, genetic diversity and population history of the Juniperus sabina L. Varieties in Europe and Asia. Forests 15, 866 (2024).
Han, W. et al. Effects of soil physico-chemical properties on plant species diversity along an elevation gradient over alpine grassland on the Qinghai-Tibetan Plateau, China. Front. Plant Sci. 13, 822268 (2022).
Wu, H., Liu, Y., Zhang, T., Xu, M. & Rao, B. Impacts of soil properties on species diversity and structure in Alternanthera philoxeroides-invaded and native plant communities. Plants 13, 1196 (2024).
Taberlet, P. et al. Genetic diversity in widespread species is not congruent with species richness in alpine plant communities. Ecol. Lett. 15, 1439–1448 (2012).
Bortoluzzi, C. et al. The type of bottleneck matters: Insights into the deleterious variation landscape of small managed populations. Evol. Appl. 13, 330–341 (2020).
Zhu, X. W. & Chen, W. H. Tiankengs in the Karst of China. Carsologica Sin. S1, 7–24 (2006).
Nybom, H. Comparison of different nuclear DNA markers for estimating intraspecific genetic diversity in plants: DNA-markers and plant genetic diversity. Mol. Ecol. 13, 1143–1155 (2004).
Zoltán, B. et al. The conservation value of karst dolines for vascular plants in woodland habitats of Hungary: Refugia and climate change. Int. J. Speleol. 43, 15–26 (2014).
Feng, H. Z. The Study on Origion and Evolution of Karst Tiankeng Flora in Dashiwei, Guangxi. Master’s Thesis, Guangxi Normal University, Guilin, China (2012).
Susan, M. M. et al. Isolation, small population size, and management influence inbreeding and reduced genetic variation in K’gari dingoes. Conserv. Genet. 25, 955–971 (2024).
Zhu, X. L. et al. Genetic Diversity of the Paphiopedilum concolor and P. hirsutissimum in Guangxi based on SSR. Guangxi Sci. 30, 1068–1078 (2023).
Furlan, E. et al. Small population size and extremely low levels of genetic diversity in island populations of the platypus, Ornithorhynchus anatinus. Ecol. Evol. 2, 844–857 (2012).
Zhu, X. L. et al. Genomic evidence reveals high genetic diversity in a narrowly distributed species and natural hybridization risk with a widespread species in the genus Geodorum. BMC Plant Bio. 23, 317–328 (2023).
Li, F. Q., Zhou, Z. C. & Xie, Y. J. Genetic diversity and genetic differentiation of different populations of Ormosia hosiei in three watersheds. Mol. Plant Breed. 15, 4263–4274 (2017).
Zhou, C. C. et al. Genetic structure of an endangered species Ormosia henryi in southern China, and implications for conservation. BMC Plant Biol. 23, 220 (2023).
Zhang, R., Zhou, Z. & Du, K. Genetic diversity of natural populations of endangered Ormosia hosiei, endemic to China. Biochem. Syst. Ecol. 40, 13–18 (2012).
Yang, Y. S. et al. Evidence for the use of Karst Tiankengs as shelters: The effect of Karst Tiankengs on genetic diversity and population differentiation in Manglietia aromatica. Forests 15, 576 (2024).
DeGiorgio, M., Lohmueller, K. E. & Nielsen, R. A model-based approach for identifying signatures of ancient balancing selection in genetic data. PLoS Genet. 10, e1004561 (2014).
Lyimo, B. M. et al. Genetic polymorphism and evidence of signatures of selection in the Plasmodium falciparum circumsporozoite protein gene in Tanzanian regions with different malaria endemicity. Malar. J. 23, 139–139 (2024).
Hashiguchi, Y. et al. Draft genome of akame (Lates japonicus) reveals possible genetic mechanisms for long-term persistence and adaptive evolution with low genetic diversity. Genome Biol. Evol. 16, evae174 (2024).
Funk, W. C., McKay, J. K., Hohenlohe, P. A. & Allendorf, F. W. Harnessing genomics for delineating conservation units. Trends Ecol. Evol. 27, 489–496 (2012).
Slatkin, M. Gene flow and the geographic structure of natural populations. Science 236, 787–792 (1987).
Forgiarini, C. et al. Low genetic differentiation despite high habitat fragmentation in an endemic and endangered species of Iridaceae from South America: Implications for conservation. Bot. J. Linn. Soc. https://doi.org/10.1093/botlinnean/boae036 (2024).
Haag, T. et al. The effect of habitat fragmentation on the genetic structure of a top predator: Loss of diversity and high differentiation among remnant populations of Atlantic Forest jaguars (Panthera onca). Mol. Ecol. 19, 4906–4921 (2010).
Liu, Y., Wang, H., Yang, J., Dao, Z. & Sun, W. Conservation genetics and potential geographic distribution modeling of Corybas taliensis, a small ‘sky Island’ orchid species in China. BMC Plant Biol. 24, 11 (2024).
Bazakos, C. et al. Whole genome scanning of a Mediterranean basin hotspot collection provides new insights into olive tree biodiversity and biology. Plant J. 116, 303–319 (2023).
Ellstrand, N. C. & Elam, D. R. Population genetic consequences of small population size: Implications for plant conservation. Annu. Rev. Ecol. Evol. Syst. 24, 217–242 (1993).
Teixeira, J. C. & Huber, C. D. The inflated significance of neutral genetic diversity in conservation genetics. Proc. Natl. Acad. Sci. 118, e2015096118 (2021).
Ravikumar, P. & Somashekar, R. K. Evaluation of nutrient index using organic carbon, available P and available K concentrations as a measure of soil fertility in Varahi River basin, India. Proc. Int. Acad. Ecol. Environ. Sci. 3, 330–343 (2013).
Cui, D. M. et al. Progress and application of novel iron biochelates. J. Huazhong Agric. Univ. 42, 59–72 (2023).
Shi, W. M. et al. Advances in olecular mechanisms of plant adaptation to soil stress. J. Plant Nutr. Fertil. 30, 1329–1338 (2024).
Zhao, X. Q. et al. Scientific issues and strategies of acid soil use in China. Acta Pedol. Sin. 60, 1248–1263 (2023).
Zhang, L. et al. Excess iron stress reduces root tip zone growth through nitric oxide-mediated repression of potassium homeostasis in Arabidopsis. New Phytol. 219, 259–274 (2018).
Wu, L. B., Holtkamp, F., Wairich, A. & Frei, M. Potassium ion channel gene OsAKT1 affects iron translocation in rice plants exposed to iron toxicity. Front. Plant Sci. 10, 579 (2019).
Bashir, K., Takahashi, R., Nakanishi, H. & Nishizawa, N. K. The road to micronutrient biofortification of rice: progress and prospects. Front. Plant Sci. 4, 15 (2013).
Kroh, G. E. & Pilon, M. Regulation of iron homeostasis and use in chloroplasts. Int. J. Mol. Sci. 21, 3395 (2020).
Wang, Z. et al. Iron uptake, translocation, and regulation in higher plants. J. Shanghai Norm. Univ. Nat. Sci. 46, 729–739 (2017).
Zhang, N. N., Shangguan, Z. P. & Chen, J. Molecular physiological mechanism and regulation of plant responses to iron deficiency stress. J. Plant Nutr. Fertil. 24, 1365–1377 (2018).
Lynch, M., Wei, W., Ye, Z. & Pfrender, M. The genome-wide signature of short-term temporal selection. Proc. Natl. Acad. Sci. U.S.A. 121, e2307107121 (2024).
Lake, N. J. et al. Quantifying constraint in the human mitochondrial genome. Nature 629, 322–329 (2024).
Horton, C. A. et al. Short tandem repeats bind transcription factors to tune eukaryotic gene expression. Science 381, eadd1250 (2023).
Laikre, L. et al. Neglect of genetic diversity in implementation of the convention of biological diversity. Conserv. Biol. 24, 86–88 (2010).
Lu, R. K. Analysis Method in Soil Agricultural Chemistry (China Agricultural Science and Technology Press, Beijing, 2000).
Rochette, N. C., Rivera-Colón, A. G. & Catchen, J. M. Stacks 2: Analytical methods for paired-end sequencing improve RADseq-based population genomics. Mol. Ecol. 28, 4737–4754 (2019).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Minh, B. Q. et al. IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 37, 1530–1534 (2020).
He, W. et al. VCF2PCACluster: a simple, fast and memory-efficient tool for principal component analysis of tens of millions of SNPs. BMC Bioinform. 25, 173 (2024).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009).
Acknowledgements
This work was supported by the Guangxi Natural Science Foundation (2024GXNSFAA010452), the Business Expense Fund of the Guangxi Institute of Botany (Project No. Gui Zhi Ye 25009), the National Key Research and Development Program (No.2022YFF1300703), Chinese Academy of Sciences ‘Light of West China’ Program (2022), Guangxi Forestry Science and Technology Promotion Demonstration Project (2024GLKX10, 2023LYKJ03 and [2022]GT23), Guangxi Key Laboratory of Plant Functional Phytochemicals Research and Sustainable Utilization (No. ZRJJ2024-3 and ZRJJ2024-11), Hechi Science and Technology Foundation and Talent Project (Heke AC231113). This publication is partially based on work at Guangxi Key Laboratory of Plant Functional Substances and Resources Sustainable Utilization of Guangxi Institute of Botany. The plant materials used in this study were collected and identified by associate researcher Jianmin Tang and Wei Xiao (Guangxi Botanical Research Institute, China).
Author information
Authors and Affiliations
Contributions
J.T. and X.W., Conceptualization; Y.Y. and J.S., methodology; G.H. and R.Z., validation; L. G. and Y. J., formal analysis; D.L. and J.T., investigation(sampling); data analysis, Y.Y.; Y.Y. and G.H., writing—original draft preparation; Y.Y. and J.S., writing—review and editing; X.W., supervision; J.T., Y.Y., R.Z. and X.W. funding acquisition. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yang, Y., Sang, J., He, G. et al. Isolation effects and soil properties drive genetic differentiation in Ormosia microphylla populations. Sci Rep 15, 25824 (2025). https://doi.org/10.1038/s41598-025-09432-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-09432-x