Abstract
This study investigated the impact of alkaline pretreatment on the biomethane yield of Xyris capensis experimentally and computationally using machine-learning (ML)-based techniques. Despite extensive studies on the anaerobic digestion of lignocellulosic biomass, the integration of a robust nexus of advanced data analytics, including explainable AI (XAI) based on SHapley Additive exPlanations (SHAP) and ML techniques, with experimental investigations has not been explored. The biomass was subjected to varying NaOH concentrations and exposure times, then digested anaerobically for 35 days. A comprehensive data-driven insight was gained through correlation-mapping, SHAP-based XAI for feature-ranking, cluster analysis for bio-digestion operational dataset using k-means integrated with Principal Component Analysis (PCA). Optimal hyperparameter settings in four different ML models, namely Artificial Neural Network (ANN), Random Forest (RF), Support Vector Machine (SVM), and Decision Tree (DT), were conducted for predicting the biomethane yield. NaOH pretreatment improved biomethane yield by 91–143%, with optimal yield at higher NaOH concentration and short exposure time. SHAP analysis revealed exposure time as the most influential feature with a strong negative impact on biomethane yield, retention time and NaOH concentration were identified as key positive contributors, while PCA captured 86% of the total data variance in the principal components (PCs) 1–3. K-means cluster analysis revealed 3 distinct groups, with cluster-0 exhibiting optimal NaOH pretreatment conditions connected to the highest biomethane yield. The RF model gave the best prediction with RMSE, MAE, MAD, MAPE, and VAF values of 3.1480, 2.0737, 1.7569, 5.7488, and 99.07, respectively, at the training phase. This research demonstrates the potential of data-driven approaches as powerful standalone tools and vital complements to experimental investigations of biomethane yield from lignocellulose biomass.
Similar content being viewed by others

Introduction
The energy sector in the world is demanding sustainable solutions to reduce the present energy crisis. Population growth, urbanization, industrialization, and technological development are the primary attributes of the rise in energy demand globally1. This finite nature of fossil fuels, the attendant environmental concerns, as well as the surge in global energy demands, have caused a shift towards renewable energy resources. Among these renewable energy resources, biomass is unique because it can be converted into solid, liquid, or gaseous fuels, offering a versatile and sustainable alternative for electricity generation, heating, and transportation. Biogas production from organic materials provides a promising opportunity to manage this challenge by harnessing renewable energy sources through anaerobic digestion. Lignocellulose materials are identified as the most available source of feedstock for biogas production globally, and they are available in the form of grasses, softwood, agricultural residues, hardwood, and energy crops. This feedstock has comparative advantages over other biogas feedstocks because of its availability, low cost, not competing with food supply, and relatively high yield2.
Lignocellulose materials have the potential to produce biogas during anaerobic digestion. However, they exhibit a limitation in terms of their efficiency owing to the slow rate of breaking down complex organic materials. Lignocellulose feedstocks comprise lignin, hemicellulose, and cellulose that are strongly bonded together and make the organic matter accessible to microbes during hydrolysis3. Therefore, pretreatment is required to break down the bonds within the lignocellulose, increase the surface area, and improve crystallinity and depolymerization before anaerobic digestion. Alkali pretreatment is a chemical pretreatment that addresses this challenge by altering the morphological arrangement of lignocellulose materials, improving biomethane yield, and reducing retention time4. The literature is replete with several studies on anaerobic digestion (AD) of lignocellulosic biomass subjected to alkaline pretreatment. Alkali pretreatment of rice straw was reported to accelerate methane production and increase methane yield by 87.1%5. Corn stover was pretreated with 0.5% KOH at 60 °C for 12 h, and the methane released was increased by 56.40% compared to the untreated substrate6. When NaOH was applied to rice straw and date palm, the optimum improvement was recorded at 6% w/w and 18% w/w, respectively7,8. Integrating alkali pretreatment into the anaerobic digestion of lignocellulose feedstock enhances the biogas release, reduces the retention time, and maximizes the potential of the process.
Feedstock composition, process parameters, and microbial dynamics greatly impact biogas production performance, rendering optimization a challenge. This challenge threatens the economic viability of the anaerobic digestion process. Hence, the intelligent model has demonstrated efficacy in intelligent feedstock management and real-time decision-making. Various mathematical models have been investigated for the prediction and optimization of the anaerobic digestion process, all showing the intricate nature of the process. These models include Schnute, transfer, Gompertz, modified Gompertz, cone, logistic, first-order, superimposed, etc9,10,11,12. , and are characterized by the combination of statistical, theoretical, and analytical methods. These models serve varying purposes in determining hydrolysis kinetics, bacterial growth, inhibition rate, lag phase duration, and biogas prediction9,13. The major challenge with these traditional models is that they are deterministic and require prior knowledge to ensure accurate prediction, hence failing to capture the non-linearity in biogas data14. This limitation has motivated the need to explore more intelligent approaches based on artificial intelligence (AI) and machine learning (ML) to enhance the accuracy of anaerobic digestion process modeling15. ML is an AI model that can learn and adjust from data, properly capturing the hidden trends in digestion and providing a more effective predictive model for anaerobic digestion16. Adaptive Neuro-fuzzy Inference System (ANFIS) is a data-driven machine learning model that can potentially address the non-linearity and complicated relationships between the input variables17. The ANFIS model processed and analyzed data trends to forecast biogas release using statistical and advanced algorithm models. This model is trained with historical data using different process parameters like pH, C/N ratio, pH, pretreatment conditions, mixing ratio, and concentration. Then, the model identifies patterns and relationships in the data and uses them to predict biogas yield based on new input data18. Fajobi et al. developed an ANFIS model to optimize and predict biogas yield from anaerobic co-digestion of mango pulp, cow dung, and Chromolaena odorat using pressure and temperature as the input parameters. The lowest Root Mean Square Error (RMSE) of 14.37 and coefficient of correlation (R2) of 0.9978 were reported, indicating that the biogas predicted is 99.78% accurate19. Anaerobic digestion and biogas yield from municipal solid waste were modeled using three models: kinetic, artificial neural network (ANN), and ANFIS, using digestion time, pH, moisture content, and volatile solids as the input parameters. The three models were compared using their performance metrics, and ANFIS has the best metric with an RMSE of 0.670 and an R2 value of 0.99920. Similarly, Chong et al. compared three different response surface methodology (RSM), ANN, and ANFIS models to predict and optimize biogas and methane yield from palm oil mill effluent. The study considered reticulation ratio, temperature, and pH as the input parameters, and it was reported that the ANFIS model has the best prediction with R2 of 0.9791, mean absolute error of 0.0730, and RMSE value of 0.143821. However, studies on optimizing and predicting biomethane yield are limited when the pretreatment conditions are considered input parameters.
Interest in advanced computational techniques for extensive data-driven insights into biogas research technology has grown in recent years, owing to the complex microbial interactions in the bio-digestion process and the pretreatment dynamics. These computational approaches can comprehend and interpret system behavior, identify hidden patterns, and facilitate biomethane process optimization, thus aiding data-driven decision-making and intelligence as a validation method for experimental investigations. While previous studies have extensively researched the AD of lignocellulosic biomass with generic ML models in their black-box nature, the robust framework of experimental investigations with advanced data analytics including explainable AI (XAI) based on SHapley Additive exPlanations (SHAP) and advanced ML techniques quantify the individual and cumulative influence of these variables on biomethane yield remains less explored in biogas research. These advanced techniques enhance bio methane predictions as well as explainable prediction outcomes, which deepens process understanding, providing actionable data-driven insights for the design and control of bioenergy systems. This study develops a novel integration of experimental and multimodal ML-based computational analysis to provide in-depth data-driven insights into the AD of Xyris capensis subjected to alkaline (NaOH) pretreatment. The experimental dataset was utilized for further statistical analysis, correlation-based parameters profiling, SHAP-based features ranking, cluster analysis and dimensionality reduction, and ML-based predictive models. This study aims at investigating the impact of alkaline pretreatment on the biomethane yield of Xyris capensis through the following objectives (i) experimental investigations of biomethane yield under different NaOH concentration and exposure time, digestion retention time in mesophilic anaerobic conditions for 35 days (ii) assessment of the linear correlation between digestion parameters, pretreatment conditions and biomethane yield (iii) statistical assessment and visualization of the impact of alkaline pretreatment on biomethane yield using a two-sample independent t-test (iv) SHAP-based feature ranking of digestion and pretreatment parameters (v) cluster analysis for bio-digestion operational dataset using k-means clustering integrated with Principal Component Analysis (PCA) (vi) develop ANN, Support Vector Machine (SVM), random forest and decision tree models for biomethane yield prediction. This research demonstrates the potential of data-driven approaches as powerful standalone tools and as vital complements to experimental investigations. By offering actionable intelligence, the study contributes to improved energy recovery and enhanced process control in the anaerobic digestion of lignocellulosic biomass.
Materials and methods
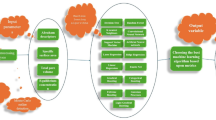
The methodological framework and approach used in this study are presented in Fig. 1. It encompasses experimental, statistical, and multimodal machine learning based analysis for investigating the impact of alkaline pretreatment on the optimal methane yield from anaerobic digestion of Xyris Capensis.
Methodological framework of this research.
Materials sourcing
Xyris capensis, which Nilsson first discovered in 189222, used for the research was sourced locally in Limpopo Province, South Africa (24°40′S 30°20′E), chopped into smaller sizes (2–4 cm), and sun-dried to 25% moisture content. The dried sample was kept in a plastic bag in a well-ventilated and controlled environment in the laboratory (about 4 °C). The sample was then subjected to alkaline pretreatment before the pretreated and untreated samples were characterized for ultimate and proximate composition according to the Association of Official Analytical Chemists (AOAC) procedure23. Liquid digestate from the previous anaerobic digester, where lignocellulose feedstock and wastewater were co-digested, was collected and used as the inoculum for the experiment. The inoculum was also stored in a controlled environment in the laboratory before characterization and anaerobic digestion.
Alkali pretreatment
Xyris capensis was pretreated with dilute NaOH to alter the recalcitrant characteristics of the substrate and improve the biomethane production. The alkali used was purchased locally from a supplier in Johannesburg, South Africa. The choice of NaOH concentration, exposure time, and temperature was selected based on previous studies with small adjustments considering the morphological structure of Xyris capensis4. As presented in Table 1, NaOH pellet was dissolved in water at different concentrations, and the chopped Xyris capensis were soaked in the solution at the set temperature for the predetermined exposure times. The substrate was dipped at 1: 10 (w/v) and stirred continuously using a magnetic stirrer set at 200 rpm. When the treatment times are completed, the solution is filtered to remove the solid from the liquid, and then washed with water until a neutral pH is achieved using a digital pH meter. The pretreated feedstock was oven-dried for 6 h at 50 °C to remove the moisture to an acceptable level before it was then stored in plastic bags and kept in a fridge set at 4 °C before characterization and anaerobic digestion.
Experimental setup
The experiment to investigate the biomethane potential of NaOH-pretreated and untreated Xyris capensis was carried out according to the VDI 4630 standard using the Automatic Methane Potential Testing System II (AMPTS II)24. Twelve 500-ml digester bottles were charged with 400 g of stable inoculum, and the pretreated and untreated substrate was added. The feedstock added to each digester was calculated using Eq. 1, which was determined based on volatile solids (VS) at 2: 1 of substrate to inoculum. The digesters were loaded and labeled, as shown in Table 1. The experiment was conducted at mesophilic conditions; therefore, the water bath temperature where the digesters were arranged was set at 37 ± 2 °C. The experiment was duplicated twice, and the average value for each treatment was recorded. Two reactors filled with only inoculum were also run simultaneously to ascertain the gas remaining in the inoculum and used for overhead correction. The gas generated from the digesters with only inoculum was deducted from another yield to determine the actual volume of biomethane produced by the substrate alone. The AMPTS II software was set at 60 s on and 60 s off for the mixer, 10% CO2 flush gas, and 80% of the stirrer speed for the experiment. The headspace of the reactor was set at 100 ml, and the biomethane generation was projected at 60%25. To remove the trapped oxygen and set anaerobic conditions in the digester, each digester was purged with nitrogen gas for about 60 s. To purify the gas released, 75 ml NaOH (3 M) solution in a 100 ml screw bottle was used. Silicon tubes were used to transfer the gas produced from the digesters directly to the purification unit before being linked to the measuring unit, where the volume of biomethane released was recorded. The experiment was terminated on day 35 of the retention period when it was established that the daily biomethane release was below 1%.
Where \(\:{M}_{i}\) is the Inoculum mass (g), \(\:{C}_{i}\) is the inoculum concentration (%), \(\:{M}_{s}\) is the substrate mass (g), while \(\:{C}_{S}\) =is the substrate concentration (%)24.
Statistical and machine learning computational framework
Correlation analysis for the biodigestion and pretreatment parameters
The linear interrelationship between the variables of anaerobic digestion and pretreatment conditions was analyzed using a Pearson correlation matrix and visualized using the correlation heat map. This expresses the potential co-linearity amongst the variables and the positive and negative correlation of the key biodigestion and pretreatment variables to the methane yield.
T-test analysis for assessment of pretreatment impact on biomethane yield
The impact of the alkaline (NaOH) pretreatment on the methane yield was statistically validated using a 2-sample independent t-test. The t-test assessed the average biomethane yields between the no-treatment and NaOH-pretreatment conditions, with the null hypothesis (H₀) assuming that no significant difference existed between the means. A significance level of p-value = 0.05 was employed. The test statistics were calculated according to Eq. 2.
Where \(\:{n}_{1},\:{n}_{2}\) are sample sizes, \(\:{\stackrel{-}{X}}_{1}\) and \(\:{\stackrel{-}{X}}_{2}\) are samples meanwhile \(\:{{s}_{1}}^{2}\:and\:{{s}_{2}}^{2}\) are sample variance. A boxplot depicting the mean, variance, and standard deviation of biomethane yield across the two treatment categories was further used to visualize the result of the t-test
SHapley additive explanations (SHAP)
SHAP is an additive feature identification technique based on co-operative game theory. It measures the impact of every feature on the prediction outcome of a machine learning model by assigning an importance value to all model features through SHAPley values26,27. In this study, SHAP provided a robust model-agnostic approach to evaluate the contribution of pretreatment condition and bio-digestion parameters to the prediction of the biomethane yield of Xyris capensis. The SHAPley value , based on \(\:n\) number of model input features \(\:i\), is calculated using Eq. 3.
Where \(\:{\varphi\:}_{i}\) is the SHAPley values representing the importance of each feature, \(\:n\) denoted the number of features while \(\:N\) represents the group input in the dataset. \(\:S\) is a subset of \(\:N\). The SHAP algorithm’s basic principle is that the sum of all feature contributions is obtained by subtracting the baseline \(\:{\varphi\:}_{o}\) and the model’s predicted value \(\:f\left(x\right)\) as in Eq. 427
The value of ϕi(x) indicates how much the feature affects the model prediction from the baseline \(\:{\varphi\:}_{o}\) for the data instance \(\:x\). The baseline value, \(\:{\varphi\:}_{o}\), represents the expected output. Accurate estimation of SHAP values is time-consuming, especially for high-dimensional datasets. However, numerous ways have been established to make SHAP more practical in real-world applications. Examples of algorithms are gradient SHAP, “Kernel SHAP”, “Tree SHAP”, and “Deep SH”28.
Principal component analysis
Principal Component Analysis (PCA) is a mathematical method employed to effectively reduce the dimensions required to represent the features of data matrices. This approach represents the original matrix through an array of new uncorrelated variables known as principal components (PC), which retain the most variance in the biodigester dataset. The co-variant matrix C is estimated from the averaged dataset using Eq. 5.
where \(\:\text{x}\) is the mean data matrix, C is the covariant matrix, and N is the number of observations. The eigen decomposition can be solved using Eq. 6.
where \(\:{{\uplambda\:}}_{i}\) is the eigenvalue of the \(\:\text{i}\text{t}\text{h}\) PC, while \(\:{\text{v}}_{i}\) is the corresponding eigen-vector. Each PC accounts for a segment of the total variance, and the explained variance ratio (EVR) is computed as in Eq. 7.
Where p is the number of features. To determine the PC scores, project the mean data onto the chosen eigenvectors using Eq. 9.
where \(\:{\text{v}}_{k}\) is the matrix of top k eigenvectors.
Biodigester operational clusters analysis (k-means clustering)
k-means clustering was used to identify distinct operational clusters within the biodigester operational dataset, considering the key input variables involved. The k-means clustering partitions the dataset into \(\:k\) non-overlapping clusters by minimizing the within-cluster sum of squares (WCSS), effectively grouping operational states with similar characteristics. In the iterative process, each data point \(\:{\text{x}}_{\text{j}}\) is assigned to the cluster with the nearest centroid \(\:{{\upmu\:}}_{\text{i}}\), using the Euclidean distance in Eq. 9, while the cluster assignment is formalized using Eq. 11.
Subsequent to the assignment, cluster centroids are adjusted by calculating the mean of all data points within each cluster with Eq. 11.
Where \(\:{c}_{i}\:\)represents the collection of data points allocated to cluster , while \(\:\left|{c}_{i}\right|\) is the data points within the cluster. The total within-cluster sum of squares (WCSS) is minimized as follows in Eq. 12.
Iterations continue until the change in WCSS between iterations falls below a threshold. Clustering results were further validated by projecting the clustered data onto the first two principal components derived from PCA, enabling an intuitive interpretation of the operational regime.
Artificial neural network
An artificial neural network (ANN) is a machine learning model that mimics the human brain with interconnected nodes that transform data into an output29. The neurons are typically organized in layers or vectors, and the output of one-layer functions as the input for the subsequent layer and potentially further layers. The feed-forward neural network (FFNN) is a specific type of ANN where input layer data is transmitted directly to the output layer without any feedback. Neurons in ANN have three layers: the input layer, the hidden layer(s), and the output layer. The input layer receives inputs \(\:{x}_{j}:(j=\text{1,2}\dots\:n)\), the hidden layer(s) consist of neurons \(\:{n}_{j}:(j=\text{1,2}\dots\:n)\), and the output layer produces outputs\(\:{o}_{j}:(j=\text{1,2}\dots\:n).\) They represent neuron output in the first hidden layer of a two-hidden neural network. The first hidden layer has \(\:{m}_{1}\) neurons, the second has \(\:{m}_{2}\:\)neurons. Weights linking the first hidden layer to the input layer are labelled \(\:{{w}_{il}}^{1}\:\)and those connecting the second hidden layer to the first are labelled \(\:{{w}_{ij}}^{2}\) and expressed in Eqs. 13 and 14. Activation function for neurons in the first hidden layer is \(\:{\varphi\:}_{i}\), and for neurons in the second layer, \(\:{\psi\:}_{j}\)30,31.
Support vector machine
SVM is a supervised learning technique employed for classification and regression tasks. The primary advantages of SVM are its simplicity, computational efficiency, and capability to be trained with a small number of samples. Nonetheless, identifying the ideal kernel and its parameters presents the greatest obstacle. The basic idea of SVM is to maximize the geometric margin between two datasets while simultaneously minimizing the empirical classification error32. For regression purposes, the function \(\:f\left(x\right)\) is estimated to illustrate the relationship between the input and output variables. The objective function is defined as in Eq. 15
Subjected to
Where \(\:{\text{y}}_{\text{i}}\) denotes the biomethane yield and \(\:{\text{x}}_{\text{i}}\) represents the input variables (digestion and pretreatment). The slack variables for points outside the insensitive tube are represented as \(\:{\xi\:}_{i}\) and \(\:{\xi\:}_{i}^{*}\), \(\:\epsilon\) is the tube width, while \(\:C\) represents the regularization parameters. This study used a radial basis function (RBF) kernel function. Thus, the non-linearity of the data is expressed as in Eq. 17.
Where \(\:\gamma\:\:\)is a kernel parameter, while the Euclidean distance between \(\:{x}_{i}\:and\:{x}_{j}\:\)samples is \(\:||{x}_{i}-{x}_{j}||\). The final regression function to compute the biomethane yield is given by \(\:f\left(x\right)\) in Eq. 18.
Where \(\:b\:\)is the bias term while \(\:{\alpha\:}_{i}\:\text{a}\text{n}\text{d}\:{\alpha\:}_{i}^{*}\) are training Lagrange multiplier.
Decision tree
A decision tree (DT) is a non-parametric supervised machine learning model characterized by a hierarchical structure with a root node, branches, internal nodes, and leaf nodes for both regression and classification tasks. It is a decision support recursive partitioning structure that models’ decisions and their outcomes, including chance event outcomes, resource costs, and efficiency. Each internal node represents a test on an attribute (e.g., whether a coin flip comes up heads or tails), each branch represents the test outcome, and each leaf node represents a class label. At each stage, a DT uses information entropy to select the next appropriate variable for separating the set of objects. DT ignores the dependence assumption and classification sequence, unlike Bayesian models. DT generates simple classification rules, which is a major benefit. These rules help analyze sensor performance and extract features.
Random forest
Random forest (RF) is an ensemble learning technique that generates numerous DTs during the training process and produces the mean prediction of the individual trees for regression tasks. The outcome of the RF is the class chosen by the majority of trees during classification, while in regression tasks, the result is the mean of the predictions from the trees. The rationale behind the RF model is that numerous uncorrelated models collectively exhibit superior performance compared to their isolated functioning. For a classification problem, each tree provides a classification or a “vote.” The forest selects the classification based on the predominant “votes.” However, for the regression task, the ensemble computes the mean of the outputs from all individual trees. The regression model is expressed in Eq. 19.
Where \(\:{f}_{i}\left(x\right)\) denotes the prediction from \(\:t-th\) tree, and \(\:y\) represents the predicted biomethane yield. The aggregate number of DT in the forest is represented as \(\:T\). \(\:{x}_{i\:}\in\:{\mathbb{R}}^{4}\) since there are 4 input variables in the biodigester dataset.
Model development, evaluations, and hyper-parameter settings
A careful selection of key control parameters of machine learning models is an important step in achieving optimal prediction performance. Some of the key hyper-parameters of the four models developed are presented in Table 2. A 2-hidden-layer architecture with 10 neurons in each layer was selected for the ANN. Owing to the complex non-linear relationship involved in the corrosion problem, a single hidden layer architecture may be insufficient in capturing the complex chemical reactions with different varying parameters involved31. The RBF kernel was selected based on its effectiveness in capturing nonlinear relationships in SVM-based corrosion rate prediction. The Gamma and epsilon values were set at 0.1, respectively, to control the influence of individual data points and ensure smooth decision boundaries.
The prediction performance of the developed machine learning was evaluated using Root Mean Square Error (RMSE), Mean Absolute Error (MAE), Mean Absolute Percentage Error (MAPE), and Variance Accounted For (VAF), and computed using Eqs. 20–23.
Results and discussion
Effect of pretreatment on feedstock composition and cumulative biomethane yield
The result of the compositional analysis of the NaOH-pretreated and untreated feedstock is presented in Table 3. It was observed that NaOH pretreatment influences the composition of the feedstock, and the impact varies based on the treatment conditions. The total solids (TS) of the feedstock were affected differently, but none of the treatment conditions reduced the TS beyond the acceptable level. It was noticed that treatments P, R, and T have the same value of TS, while treatments Q and S produce the same value of TS. However, treatment U, which is the untreated sample, has a different value from the pretreated samples, which shows that pretreatment has an impact on the feedstock. Compared with previous studies, all the treatment conditions produced higher TS6, which is a good result for biomethane production. Volatile solids (VS) are the available organic matter for the microbes’ degradation during biomethane production, and it was noticed that pretreatment applied influences the VS of the feedstock. However, all the values were more than 90%, which is an indication of good buffering capacity of the samples. These higher percentages are expected to produce higher biomethane, and they are more than what was reported for some lignocellulose biomasses3,33. The elemental composition of the substrate was also noticed to have been affected by pretreatment when pretreated samples are compared with the untreated feedstock, as shown in Table 3. C/N ratio is a major component of biomass that significantly influences the biomethane release. Carbon is the food source for microbes, while a certain quantity of nitrogen is also needed for their growth. An imbalance in the C/N ratio can lead to digester failure because of the overaccumulation or underproduction of volatile fatty acids34. A C/N ratio of 20–30 is recommended for optimum production35. It can be observed from Table 3 that all the treatment conditions have a good C/N ratio. Pretreatment application was observed to improve the C/N ratio of the feedstock compared to the untreated sample. The result of characterization of this study aligned with the previous study when different chemical pretreatment was experimented on another lignocellulose biomass36.
The cumulative biomethane yield released from NaOH-pretreated and untreated Xyris capensis after 35 days of retention time is presented in Fig. 2. A total methane yield of 258.68, 287.80, 304.02, 328.20, 310.20, and 135.06 ml CH4/ gVSadded was recorded for treatments P, Q, R, S, T, and U, respectively. This result indicates that pretreatment enhanced the biomethane yield by 91.53, 113.09, 125.10, 143.00, and 129.68% for treatments P, Q, R, S, and T, respectively, compared to the untreated Xyris capensis (treatment U). This finding agrees with the previous studies that reported that the pretreatment increases the biomethane yield of lignocellulose feedstock37,38,39. Biogas yield from anaerobic co-digestion of cow dung and food waste was improved by 28.1, 20.23, and 13.27% when pretreated with ultrasonic, autoclave, and microwave pretreatment methods, respectively40. Enzymatic pretreatment of corn stover increases the biomethane yield by 36.9% compared to the untreated substrate41. The optimum biomethane yield of 328.20 ml CH4/ gVSadded was observed when 4% w/w of NaOH was utilized for 20 min at 90 °C, which indicates that higher concentration with shorter exposure time favours the pretreatment of Xyris capensis. It was observed that beyond that period, further increases in the alkali concentration do not translate into further biomethane yield. This can be traced to the release of inhibitory compounds like acetic acid, furfural, and 5-hydroxymethylfurfural, which can hinder the activities of the microorganisms and subsequent biomethane release37. The reduction in biomethane yield beyond this condition could be the over-degradation of hemicellulose in the substrate, thereby reducing the available organic content for biomethane release. The pretreatment decrystallized and expanded lignin-carbohydrate bonds of the substrate, thereby increasing the accessible surface area for enzymatic hydrolysis. The properties of NaOH alter the lignin structure and significantly redistribute/eliminate the lignin content from acetyl groups, hemicellulose, and uronic acid. The performance of the NaOH pretreatment depends on the percentage of lignin in the feedstock. It was noticed from the study that treatment alters the hydrolyzable links and improves saccharification, which increases solubilization and reduces crystallinity and polymerization, as reported in a previous study42. This study aligned with what was reported when rice straw, Napier grass, and hazelnut were pretreated with NaOH. It was observed that the optimum yield was when 4% NaOH was used, but at different treatment times and temperatures43,44,45. The variation in the treatment time and temperature can be linked to the variation in the morphological arrangement of the feedstock and the purity of the chemical used.
Cumulative biomethane yield of NaOH-pretreated and untreated feedstock.
Statistical evaluation of digestion process parameters and pretreatment conditions
An in-depth statistical insight into the relationship between the vital biodigester parameters and pretreatment conditions is vital to understand the complex biochemical processes within the biodigester and the biomethane production trends. A correlation analysis was conducted between the critical operating and pretreatment parameters. The discovered correlations highlight the multifactorial nature of the anaerobic digestion process, where no single parameter solely determines methane output but rather a collective interaction of physical, chemical, and biological elements that play a crucial role. Table 4 presents the statistical summary of the relevant parameters.
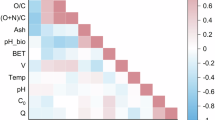
Figure 3 shows the magnitude and direction of the linear correlation between the key operating conditions and methane yield. The retention time of the digestion exhibits a weak positive correlation (0.13) with the methane yield. This indicates that prolonged digestion facilitates more thorough hydrolysis and methanogenesis. The very weak (-0.04) correlation value of temperature depicts an essentially negligible linear correlation with methane yield. The NaOH concentration exhibits a marginally mild negative correlation with methane yield. This may result from non-linear effects since excessive NaOH might suppress microbial activity, whilst appropriate concentrations improve digestibility. Consequently, the general trend is that the overall trend becomes weak or slightly negative. Likewise, exposure time exhibits a weak negative correlation with methane yield. Prolonged exposure to NaOH results in over-degradation or the production of inhibitory chemicals, hence diminishing yield. The optimal duration of exposure is crucial. These findings underscore the significance of controlling both pretreatment duration and intensity to enhance biogas production from Xyris capensis.
Correlation heat map of the anaerobic digestion of Xyris capensis.
SHAP feature importance analysis
The non-linear interactions among digestion and pretreatment parameters such as exposure time, retention time, NaOH concentration, and temperature are further analyzed using SHAP-based feature-ranking. This approach reveals the contribution of each variable to the model output while giving insight into the operational factors that significantly affect biomethane yield. Figure 4 presents the SHAP values of each feature, indicating their contributions to the prediction (either positive or negative) and the degree of variability over various scenarios. Exposure times have the most significant effect, with a broad SHAP range of around − 200 to + 50. Extended exposure time generally lowers model output, while shorter exposure time marginally enhances it. This inverse contribution corresponds with its mean SHAP value of -14.23, affirming it as a negatively impactful variable. The distribution indicates that long periods of exposure lead to decreasing biomethane production, potentially due to excessive breakdown or structural inhibition during pretreatment. Furthermore, retention is the second most significant feature, exhibiting a positive SHAP mean of + 13.81. Enhanced model output is obtained at a high retention time, indicating a direct association with biomethane generation. Extended digestion durations generally facilitate more thorough breakdown and methanogenesis, hence supporting essential anaerobic digestion dynamics.
The concentration of NaOH has a moderate positive SHAP mean of + 2.07. The trend in the figure depicts elevated concentrations and their positive influence in enhancing the output, although with less consistency across the data points. This indicates a threshold or ideal range beyond which increased concentration may not benefit biomethane generation. The temperature exhibits a moderate positive SHAP value of + 0.94, featuring a little variability in Fig. 4. This indicates that temperature exerted a slight positive effect on biomethane prediction. This is consistent with the assumption that mesophilic temperatures promote anaerobic microbial activity. However, the model does not assign significant variability to temperature.
SHAP values plot of the digestion and pretreatment parameters.
Beyond the individual average importance presented in the SHAP values in Fig. 4, a good understanding of the cumulative importance of each parameter gives useful insight into how feature effects vary across cases. Presented in Fig. 5 is the SHAP decision plot illustrating the cumulative importance of each feature. A trend that aligns with the observation in Fig. 4 was noted for each feature. The high exposure time began at a high level and then swiftly dropped, resulting in a significant decline in predictive value, hence reinforcing its earlier established strong negative impact. Furthermore, retention time and NaOH concentration augment model predictions, affirming their combined influence on the enhancement of biomethane from Xyris capensis. In numerous instances, temperature results in a negligible deviation from the baseline value, corresponding with its low average SHAP value.
SHAP decision plot of the digestion and pretreatment parameters.
Dimensionality reduction of anaerobic digestion parameters using PCA
Both individual contributions and relationships and interdependence among digestion and pretreatment variables are reflected in the PCA results. This further reinforces the insights gained from the DTR-based feature importance by reducing the dataset’s dimensionality while retaining the bio-digestion dataset’s core structure. The PCA efficiently illustrated the variance inherent in the digestion dataset by vividly identifying the dominant principal component (PC) accountable for significant variability. The result of the PCA is presented in Table 4 and visually illustrated in Fig. 5. From Table 5, the first principal component (PC1) accounts for about 36.64% of the variance in the data, while 28.71% of the variance is attributed to the second principal component (PC2). The combined cumulative variance between PC1 and PC2 (65.35%) indicated that they were insufficient to represent the dataset’s structure. While PC3 and PC4 contribute less variance, the combined cumulative variance between PC1 and PC3 (86.64%) shows that the 3 PCs are enough to capture the variance and the structure in the dataset.
Figure 6 (a) presents the explained variance and the cumulative variance for each component expressed in percentage. The cumulative variance plot reveals that the first three PCs (PC1 to PC3) together capture nearly 87% of the variability. While PC4 contributes less, it is still considered significant. However, the dimensionality of the bio digestion dataset has been reduced to 3 (PC1 to PC3), implying that these 3 PCs can substantially capture the variance in the dataset. Figure 6 (b) is the scree plot depicting each principal component’s eigenvalues with a reference line at eigenvalue = 1 to indicate significant components. The eigenvalue for only PC1 and PC2 substantially exceeds 1, whilst the other components are approximately below 1. According to Kaiser Criterion, only PCs with eigenvalues above 1 are significant46. This establishes PC1 and PC2 as substantial contributors to the dimensional structure. The PCA outcome has a significant implication for the monitoring efforts in the biodigester operation by focusing on parameters that strongly load on PC1 and PC2. From the PCA result, PC1 positively correlates with NaOH concentration and negatively with the NaOH contact time. PC2 is positively influenced by retention time and negatively related to NaOH concentration. PC1 is negatively influenced by retention time and temperature, while PC4 is negatively influenced by time and temperature.
Scree plot showing (a) explained and cumulative variance (b) eigenvalues relating to each principal component.
Operational cluster analysis of the digestion process
A k-means cluster analysis visualized via the PCA further revealed the dynamics of anaerobic digestion of Xyris capensis under NaOH pretreatment, as illustrated in Fig. 7. The spatial disparity across the clusters along PC1 and PC2 indicates significant underlying feature diversity, hence supporting the fact that the operational parameters of Cluster 0 and Cluster 2 are the most different. Cluster 1 bridges the traits of the two extremes in a more transitional area. Every centroid is the average location of all observations in that cluster. Hence, it is practically the “typical digestion regime” for that group. Every cluster shows a unique mix of NaOH content, digestion temperature, and exposure time, all affecting biomethane yield. Cluster 0 is a high-performing NaOH-pretreated condition. The position of Cluster 0 in PCA space shows a different biomethane-enhancing system, likely related to ideal NaOH concentration, balanced exposure, and adequate retention duration. Conversely, Cluster 2 probably contains untreated or low-intensity pretreatment conditions defined by lesser NaOH exposure and low biomethane yields. Applied under optimal operational windows, alkaline (NaOH) pretreatment clusters into separate, high-performing clusters (e.g., Cluster 0). PCA provides a clear dimensional reduction that facilitates the visualization of the underlying variations in digesting tactics. This knowledge offers a data-driven framework for targeted operational optimization, particularly for scaling up pretreatment operations or creating an adaptive digestion protocol. Cluster 1 might be intermediate procedures where moderate pretreatment was used but did not produce the synergistic effects seen in Cluster 0.
Operational clusters of the anaerobic digestion of Xyris capensis.
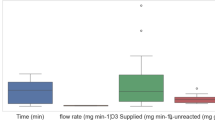
Figure 8 illustrates the distribution and variability of key operating parameters across the 3 clusters. Across the 3 clusters, temperatures vary from 19 °C to 31 °C. Operating in the higher mesophilic range (29–31 °C), Cluster 0 is known to host active microbial populations. While Cluster 2 is more diverse, Cluster 1 shifts somewhat cooler. When paired with NaOH pretreatment, higher operating temperatures synergistically enhance hydrolysis rates and microbial metabolism, thereby explaining why Cluster 0 conditions can correspond to better yield. With a median retention length of 30 days, Cluster 0 indicates long-duration digesting processes. With the shortest retention times (10 days), Cluster 1 suggests rapid-cycle digestion, presumably tuned for quicker throughput. A hybrid approach is implied by Cluster 2, which spans a wider spectrum yet focuses on 16 days. Longer retention durations (Cluster 0) often allow more complete digestion of substrates, which is especially advantageous when lignin content is lowered by NaOH pretreatment, hence facilitating deeper microbial action. In all three clusters, NaOH levels range from 0 to 5% w/w. With medians at 3–4% NaOH concentration, clusters 0 and 1 reveal wider, overlapping NaOH ranges (0–5%). Skewed more toward low or no pretreatment, Cluster 2 could indicate control or little pretreatment situations. This verifies that Cluster 2 may include untreated samples with predicted decreased biomethane yield, consistent with previous findings. With far longer NaOH exposure times (50–60 min), Cluster 2 may compensate for its lower NaOH concentration. With shorter exposure times (10–30 min), clusters 0 and 1 imply more aggressive pretreatment in less time. Though combining both ideal concentration and duration (as probably observed in Cluster 0) enhances performance, longer exposure time at lower NaOH concentration (Cluster 2) may still produce moderate efficacy.
Variation of bio-digestion parameters and NaOH pretreatment conditions across the operational clusters.
Statistical insight into the effect of alkaline pretreatment on the biomethane yield
The impact of the alkaline pretreatment of the Xyris capensis biomass sample on its biomethane yield was validated statistically beyond the experimental investigation. A two-sample independent t-test was carried out to compare the average yield of biomethane between the pretreated and untreated categories, with a null hypothesis that there is no significant difference in the biomethane yield between the untreated and pretreated categories. The statistical analysis demonstrates the effectiveness of NaOH pretreatment in improving biomethane output from Xyris capensis. Based on t-test’s p-value \(\:<\) 0.05, we can ignore the null hypothesis, assuming no difference between the two categories. Hence, the improvement in biomethane yield is statistically significant. Figure 9 shows the variation in biomethane yield and the mean and standard deviation of biomethane yield values across the untreated and pretreated categories. These charts further establish an enhanced methane generation after NaOH pretreatment. The figures prove that NaOH pretreatment greatly increases biomethane output compared to untreated conditions. The ability of NaOH to decompose complicated cell wall structures, lessen lignin shielding, and enhance methanogenesis and enzymatic hydrolysis could be attributed to this observation47. During anaerobic digestion, optimal circumstances with 6% NaOH and 0.10 g·g−1 cellulase produced a notable rise in gas generation48. Compared to untreated samples, solid-state NaOH pretreatment at 4% concentration produced 144% more methane49.
Biomethane yield changes under NaOH pretreatment and no-treatment conditions.
Performance evaluation of the developed model
The model developed for predicting the biomethane yield of NaOH-treated Xyris capensis was evaluated for accuracy and reliability using important performance metrics. Table 6 presents the statistical metrics value of the ANN, SVM, RF, and DT models at the training phase. The table revealed a significant variation in their prediction performance across all metrics. Based on RMSE, the RF model outperformed other models with the lowest RMSE of 3.1480, showing it had the least prediction error and better accuracy at training. This implies that the RF exhibits little variation in predicting the biomethane yield at the training phase. The accuracy of DT follows the RF, which had an RMSE of 5.8544, while ANN and SVM recorded higher RMSE values of 7.2026 and 8.3786, respectively, indicating less reliable prediction during training. A similar trend was noted based on MAE values, with the RF exhibiting the lowest MAE value of 2.0737, affirming its reliability. The MAE values of SVM \(\:{(\text{R}\text{M}\text{S}\text{E}}_{\text{S}\text{V}\text{M}}=\:6.0038\)), ANN \(\:{(\text{R}\text{M}\text{S}\text{E}}_{\text{A}\text{N}\text{N}}=\:6.2926,)\), DT (\(\:{\text{R}\text{M}\text{S}\text{E}}_{\text{D}\text{T}}=4.5435\:)\:\)indicates that DT predictions are on average, significantly closer to the actual values. Furthermore, the lower MAD further confirms that RF predictions are more stable and consistent. RF had the smallest MAD value of 1.7569, while the MAD values of DT, ANN, and SVM are 4.7654, 6.1675, and 6.2029, respectively. Considering the MAPE values of the models, which indicate the percentage prediction accuracy of the model, RF was also noted to have the best prediction accuracy based on the MAPE value of 5.7488. This indicates that the RF model is about 94% accurate at predicting the biomethane yield during training. This was better than DT \(\:{(\text{M}\text{A}\text{P}\text{E}}_{\text{D}\text{T}}=\:\)6.9543), SVM \(\:{(\text{M}\text{A}\text{P}\text{E}}_{\text{S}\text{V}\text{M}}=\:\)9.0657), and ANN \(\:{(\text{M}\text{A}\text{P}\text{E}}_{\text{A}\text{N}\text{N}}=\)10.6767). Based on VAF, all models were noted to capture all the variance in the actual bio-methane output. However, RF gave the highest VAF of 99.0680%, followed closely by DT \(\:{(\text{V}\text{A}\text{F}}_{\text{D}\text{T}}=\:\)98.7887%), SVM \(\:{(\text{V}\text{A}\text{F}}_{\text{S}\text{V}\text{M}}=\:\)96.9575%), and ANN \(\:{(\text{V}\text{A}\text{F}}_{\text{A}\text{N}\text{N}}=\:\)93.9540%). Generally, the RF model predicted biomethane yield optimally across all metrics. While ANN and SVM approximated well, their larger errors suggest suboptimal parameter tuning and learning structure. Though slightly inferior to RF, DT performed well and may be used in practice.
The generalization capacity of the machine learning models was also assessed at the testing stage to determine their predictive consistency. As shown in Table 7, the performance metrics give a clearer picture of how each model responded to unseen data. The RF model achieves the most optimal performance, achieving the lowest RMSE of 5.6862, confirming its excellent generalization performance with unseen data outside the training phase. However, the ANN demonstrated the highest RMSE of 9.9094. This depicts a substantial divergence from the experimental values of biomethane yield, which diminished during the testing phase. The DT and SVM exhibited RMSE values of 5.9346 and 7.5069, respectively. Based on the MAE value, RF still predicts best with 4.2938, slightly higher than DT’s 3.5767, the lowest MAE across models. ANN again emerged with the least performance with an MAE of 7.8629, while SVM had a moderate performance with 5.9737. DT had the lowest MAD of 2.4676, followed by RF (3.8981), SVM (5.3245), and ANN (9.3768). With a MAPE of 7.0717%, RF made the most accurate percentage predictions, making it acceptable for operational bioenergy systems. SVM had the highest VAF (94.60%), ahead of RF (93.93%). DT had 89.80%, followed by ANN with 92.96%.
The RF model has the smallest area on the radar plot in Fig. 10, indicating lower RMSE, MAE, and MAPE relative to ANN, SVM, and DT models. Wider shapes created by the ANN, SVM, and DT models suggest larger error metrics. This verifies that RF provides improved learning capacity and error minimization during the building of models. The pattern stays constant as the RF models exceed the others with lower values on all axes. The radar charts confirm visually that RF is the most accurate model for predicting biomethane yield from Xyris capensis. Its minimal errors and higher accuracy throughout training and testing make it the perfect choice for practical bioenergy modeling and deployment. Though all models showed a decline in performance in testing (as expected because of unseen data), the RF model exhibited superior generalization and slight performance loss.
Statistical metrics for the best model (RF) at training and testing.
The experimental and predicted biomethane values at the training phase are compared graphically, as shown in Fig. 11. It shows the experimental biomethane yield values compared to the RF (most accurate) model-predicted values. Experimental and predicted values across all sample indices significantly overlap, suggesting great prediction accuracy during training. The model’s capacity to identify nonlinear patterns in the data is confirmed even more by the close clustering of points and trends between the experimental and predicted values. The error histogram shows the distribution of prediction errors during training. The distribution is centered around zero, indicating that the model does not show consistent bias. The bell-shaped error distribution closely resembles the overlay normal distribution curve, suggesting that the prediction errors are random50. The error distribution verifies that most prediction errors are within a small range and that large departures are uncommon.
Comparison plot of the actual and predicted methane yield using the RF (most accurate) model-predicted values at the training.
Similarly, at the testing phase, the experimental and predicted biomethane value was compared in Fig. 12. This chart shows the experimental biomethane yield values compared to the RF (most accurate) model-predicted values at the testing phase. Although certain areas show little variation, the general trend tracking stays consistent. The model accurately predicts biomethane yield during testing outside training data by effectively understanding the inherent nonlinear relationships. The histogram shows the spread of prediction errors throughout testing. Centered around zero, the error distribution indicates low bias in model predictions. The error distribution is somewhat more spread out than during training, and the fit with the normal distribution is less perfect, suggesting a broader error spread. Though more diverse than in training, the testing errors are still well-behaved and typically fall within a reasonable range.
Comparison plot of the actual and predicted methane yield using the RF (most accurate) model-predicted values at the training.
Figure 13 presents the scatter plot of experimental and RF (most accurate) model-predicted methane yield values at both the testing and training phases. These plots show a near-linear alignment of points, indicating good model accuracy and little residual error during training. This shows that the model has properly understood the connection between the input characteristics and biomethane yield, hence supporting the model’s previously reported low error values. Similarly, the model maintains a significant connection between experimental and predicted values at the testing stage despite introducing a novel dataset. Some minor line deviations are observable, which is expected in real-world situations because of natural noise and complexity in anaerobic digestion processes.
Scatter plot of actual and predicted methane yield using the RF model at the training and testing phase.
Overall, the RF model exhibited the best prediction performance than other models. The ensemble and random feature sampling nature of RF, which are noise and variance resistant and requires less parameter optimization could be accountable for its superior performance51. While the performance of non-ensemble-based DT is better than ANN and SVM, it is less accurate compared to RF. This is due to its training overfitting possibilities, particularly in the absence of pruning or complexity regulation, resulting in memorizing instead of learning, hence poor generalization on novel data52. The less accurate prediction of ANN could be attributed to its susceptibility to overfitting when dealing with a small training data set. The complexity and number of its parameters, as well as high model flexibility, could result in learning noise instead of the underlying patterns, especially without a strong mitigation strategy like regularization, data augmentation, among others53. In a similar study by Ahmad et al.54 RF consistently gave a better prediction accuracy than ANN in small-data contexts owing to their potential for overfitting. SVM, however, relies heavily on well-structured, low-noise datasets for good performance55. It is susceptible to hyperparameter settings, whose defective tuning could significantly result in underfitting or overfitting56. Comprehensive parameter optimization could be challenging for SVM when dealing with meagre or noisy datasets, owing to their quadratic or cubic training complexity57.
In a similar study, ANN was used to simulate the biogas yield of chemically treated grass, clover, and wheat straw co-digested during anaerobic digestion. Retention time, temperature, alkali concentration, and substrate composition were used as input parameters while the cumulative methane yield was the output. The performance metric of the model shows a varying level of performance based on the variation of input parameters, with a minimum RMSE value of 0.41058, which is lower than 9.9094 for ANN, and the minimum from RF (5.6862) observed in this study. An ML application for the optimization and prediction of fresh mass methane yield of particle size pretreated Arachis hypogea shells using ANFIS was reported to have RMSE, MAPE, and MAD values of 2.7875, 9.0643, and 1.7665, respectively. These values are lower compared to this study, which shows that the model performed better than this18. Biomethane release from the organic fraction of municipal solid waste was optimized and predicted with ANN, linear regression, XGBoost, RF, and SVM. Moisture content, C/N ratio, lignocellulose contents, and age of the waste were the input parameters with biomethane as the output. XGBoost and RF were reported as the best models with RMSE values of 305 and 496, respectively59, which are higher compared to the values recorded in this study. SVM was reported as the best model for the prediction of biogas and methane yield of solid-state anaerobic digestion, using existing biogas and methane data. The RMSE and MAE values of 3.21 and 1.93 were reported60, respectively; these values are lower compared to this study. It was observed that this study deviates from the existing studies, which can be traced to the difference in feedstock, input process parameters, model version, and type. It was also difficult to have a comprehensive comparison because the performance metrics considered were not the same, for instance, most of the studies do not consider VAF in their performance metrics.
Conclusion
The impact of alkaline (NaOH)-pretreatment on the process output of anaerobic digestion of Xyris capensis was investigated in this study through an integrated approach involving experimental and multi-modal statistical and ML-based analysis, including correlation analysis, SHapley Additive exPlanations (SHAP) for feature-ranking, cluster analysis for bio-digestion operational and pretreatment datasets, using k-means integrated with Principal Component Analysis (PCA). This provides in-depth insights into the operational dynamics of anaerobic digestion. From the experimental front, the NaOH pretreatment demonstrated a substantial enhancement in biomethane yield by breaking down the recalcitrant properties of Xyris capensis. Furthermore, all the pretreatment conditions investigated improve the yield to about 143% under optimal pretreatment conditions. SHAP analysis revealed exposure time as the most influential feature with a strong negative impact on biomethane yield, while retention time and NaOH concentration were identified as key positive contributors. PCA further verified the importance of these features, capturing over 86% of the variance in three principal components. ANN, SVM, DT, and RF models were developed to predict biomethane yield. The RF model outperformed others (ANN, DT, and SVM), achieving the lowest error margins during training and testing. The integrated experimental and data-driven computational techniques presented in this study contribute significantly to sustainable energy production by offering actionable intelligence toward improved energy recovery and enhanced process control in the anaerobic digestion of lignocellulosic biomass.
Specimen deposited in a public herbarium
Xyris capensis was deposited at Flora of Botswana on 15th June 1996, with record number 84221, recorded by MG Bingham and collector number 11045.
Permission to collect Xyris capensis sample
Xyris capensis is a common grass found in almost all the Provinces of South Africa; therefore, official permission is not required.
Feedstock identification for the study
Author Daniel M. Madyira identified the feedstock species used for this study.
Data availability
The datasets used and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Singh, D., Tembhare, M., Machhirake, N. & Kumar, S. Biogas generation potential of discarded food waste residue from Ultra-Processing activities at food manufacturing and packaging industry. Energy 263, 126138. https://doi.org/10.1016/J.ENERGY.2022.126138 (2023).
Dahadha, S., Amin, Z., Bazyar Lakeh, A. A. & Elbeshbishy, E. Evaluation of different pretreatment processes of lignocellulosic biomass for enhanced biomethane production. Energy Fuels. 31, 10335–10347. https://doi.org/10.1021/ACS.ENERGYFUELS.7B02045 (2017).
Olatunji, K. O. & Madyira, D. M. Comparative analysis of the effects of five pretreatment methods on morphological and methane yield of groundnut shells. Waste Biomass Valorization. 15, 469–486. https://doi.org/10.1007/S12649-023-02177-6/FIGURES/5 (2024).
Olatunji, K. O., Madyira, D. M., Ahmed, N. A. & Ogunkunle, O. Influence of alkali pretreatment on morphological structure and methane yield of Arachis hypogea shells. Biomass Convers. Biorefin. 14, 12143–12154. https://doi.org/10.1007/S13399-022-03271-W/METRICS (2024).
Li, X. et al. Quantitative visualization of subcellular lignocellulose revealing the mechanism of alkali pretreatment to promote methane production of rice straw. Biotechnol. Biofuels. 13, 1–14. https://doi.org/10.1186/S13068-020-1648-8/FIGURES/11 (2020).
Siddhu, M. A. H. et al. Improve the anaerobic biodegradability by copretreatment of thermal alkali and steam explosion of lignocellulosic waste. Biomed. Res. Int. 2016 https://doi.org/10.1155/2016/2786598 (2016).
He, Y., Pang, Y., Liu, Y., Li, X. & Wang, K. Physicochemical characterization of rice straw pretreated with sodium hydroxide in the solid state for enhancing biogas production. Energy Fuels. 22, 2775–2781. https://doi.org/10.1021/ef8000967 (2008).
Lahboubi, N. et al. Effect of Alkali-NaOH pretreatment on methane production from anaerobic digestion of date palm waste. Ecol. Eng. Environ. Technol. 23, 78–89. https://doi.org/10.12912/27197050/144846 (2022).
Karki, R. et al. Anaerobic Co-Digestion of various organic wastes: kinetic modeling and synergistic impact evaluation. Bioresour Technol. 343, 126063. https://doi.org/10.1016/J.BIORTECH.2021.126063 (2022).
Olugbemide, A. D., Lajide, L., Adebayo, A. & Owolabi, B. J. Optimization and kinetic study of biogas production from rice husk through Solid-State alkaline pretreatment method. Invertis J. Renew. Energy. 6, 175. https://doi.org/10.5958/2454-7611.2016.00024.2 (2016).
Jijai, S. & Siripatana, C. Kinetic model of biogas production from Co-Digestion of Thai rice noodle wastewater (Khanomjeen) with chicken manure. Energy Procedia. 138, 386–392. https://doi.org/10.1016/J.EGYPRO.2017.10.177 (2017).
Rani, P., Pathak, V. V. & Bansal, M. Biogas production from wheat straw using textile industrial wastewater by Co-Digestion process: experimental and kinetic study. J. Turkish Chem. Soc. Sect. A: Chem. 9, 601–612. https://doi.org/10.18596/JOTCSA.1009483 (2022).
Pramanik, S. K., Suja, F. B., Porhemmat, M. & Pramanik, B. K. Performance and kinetic model of a Single-Stage anaerobic digestion system operated at different successive operating stages for the treatment of food waste. Processes 2019. 7, 600. https://doi.org/10.3390/PR7090600 (2019).
Khashaba, N. H., Ettouney, R. S., Abdelaal, M. M., Ashour, F. H. & El-Rifai, M. A. Artificial neural network modeling of Biochar enhanced anaerobic sewage sludge digestion. J. Environ. Chem. Eng. 10, 107988. https://doi.org/10.1016/J.JECE.2022.107988 (2022).
Andrade Cruz, I. et al. Fernando Romanholo ferreira, L. Application of machine learning in anaerobic digestion: perspectives and challenges. Bioresour Technol. 345, 126433. https://doi.org/10.1016/J.BIORTECH.2021.126433 (2022).
Şenol, H., Elibol, E. A., Bianco, F. & Race, M. Enhancing methane production from pistachio skin via optimized Hydrothermal-Alkaline pretreatment and autoregressive integrated moving average modeling. J. Environ. Manage. 380, 125121. https://doi.org/10.1016/J.JENVMAN.2025.125121 (2025).
Najafi, B. & Faizollahzadeh Ardabili, S. Application of ANFIS, ANN, and logistic methods in estimating biogas production from spent mushroom compost (SMC). Resour. Conserv. Recycl. 133, 169–178. https://doi.org/10.1016/j.resconrec.2018.02.025 (2018).
Olatunji, K. O. et al. Performance evaluation of ANFIS and RSM modeling in predicting biogas and methane yields from Arachis hypogea shells pretreated with size reduction. Renew. Energy. 189, 288–303. https://doi.org/10.1016/J.RENENE.2022.02.088 (2022).
Fajobi, M. O. et al. Prediction of Biogas Yield from Codigestion of Lignocellulosic Biomass Using Adaptive Neuro-Fuzzy Inference System (ANFIS) Model. Journal of Engineering 2023, 9335814, (2023). https://doi.org/10.1155/2023/9335814
Avinash, L. S. & Mishra, A. Comparative evaluation of artificial intelligence based models and kinetic studies in the prediction of biogas from anaerobic digestion of MSW. Fuel 367, 131545. https://doi.org/10.1016/J.FUEL.2024.131545 (2024).
Chong, D. J. S., Chan, Y. J., Arumugasamy, S. K., Yazdi, S. K. & Lim, J. W. Optimisation and performance evaluation of response surface methodology (RSM), artificial neural network (ANN) and adaptive neuro-fuzzy inference system (ANFIS) in the prediction of biogas production from palm oil mill effluent (POME). Energy 266 https://doi.org/10.1016/J.ENERGY.2022.126449 (2023).
Xyris Capensis Var. Schoenoides (Mart.) L.A.Nilsson | Plants of the World Online | Kew Science. https://powo.science.kew.org/taxon/urn:lsid:ipni.org:names:. 77295336–77295331. Accessed 28 Apr 2025 (2025).
Official Methods of Analysis, 21st Edition. AOAC International. (2019). https://www.aoac.org/official-methods-of-analysis-21st-edition-2019/. Accessed 15 Oct 2021 (2021).
organischer Stoffe Substratcharakterisierung, V. Verein Deutscher Ingenieure Characterisation of the Substrate, Sampling, Collection of Material Data, Fermentation Tests VDI 4630 VDI-Richtlinien (2016).
Olatunji, K. O., Madyira, D. M., Ahmed, N. A. & Ogunkunle, O. Experimental evaluation of the influence of combined particle size pretreatment and Fe3O4 additive on fuel yields of Arachis hypogea shells. Waste Manag Res. https://doi.org/10.1177/0734242X221122560 (2022).
Sun, Y. & Ma, H. Interpretable analysis of transformer winding vibration characteristics: SHAP and Multi-Classification feature optimization. Int. J. Electr. Power Energy Syst. 166, 110585. https://doi.org/10.1016/j.ijepes.2025.110585 (2025).
Tempel, F., Ihlen, E. A. F., Adde, L. & Strümke, I. Explaining human activity recognition with SHAP: validating insights with perturbation and quantitative measures. Comput. Biol. Med. 188, 109838. https://doi.org/10.1016/j.compbiomed.2025.109838 (2025).
Lundberg, S. M. & Lee, S. I. A unified approach to interpreting model predictions. CoRR abs/1705.07874 (2017).
Sanni, O., Adeleke, O., Ukoba, K., Ren, J. & Jen, T. C. Application of machine learning models to investigate the performance of stainless steel type 904 with agricultural waste. J. Mater. Res. Technol. 20, 4487–4499. https://doi.org/10.1016/j.jmrt.2022.08.076 (2022).
Kilani, A. J., Adeleke, O. & Fapohunda, C. A. Application of machine learning models to investigate the performance of concrete reinforced with oil palm empty fruit brunch (OPEFB) fibers. Asian J. Civil Eng. 23, 299–320. https://doi.org/10.1007/s42107-022-00424-0 (2022).
Adeleke, O., Akinlabi, S. A., Jen, T. & Dunmade, I. Application of artificial neural networks for predicting the physical composition of municipal solid waste: An assessment of the impact of seasonal variation. https://doi.org/10.1177/0734242X21991642 (2021).
Lee, W. S. et al. Sensing technologies for precision specialty crop production. Comput. Electron. Agric. 74, 2–33. https://doi.org/10.1016/j.compag.2010.08.005 (2010).
Ajayi-Banji, A. A., Rahman, S., Sunoj, S. & Igathinathane, C. Impact of corn Stover particle size and C/N ratio on reactor performance in Solid-State anaerobic Co-Digestion with dairy manure. J. Air Waste Manage. Assoc. 70, 436–454. https://doi.org/10.1080/10962247.2020.1729277 (2020).
Ilhan, Z. E., Marcus, A. K., Kang, D. W., Rittmann, B. E. & Krajmalnik-Brown, R. PH-Mediated microbial and metabolic interactions in fecal enrichment cultures. mSphere 2. https://doi.org/10.1128/MSPHERE.00047-17 (2017).
Kainthola, J., Kalamdhad, A. S. & Goud, V. V. Optimization of process parameters for accelerated methane yield from anaerobic Co-Digestion of rice straw and food waste. Renew. Energy. 149, 1352–1359. https://doi.org/10.1016/J.RENENE.2019.10.124 (2020).
Olugbemide, A. D., Oberlintner, A., Novak, U. & Likozar, B. Lignocellulosic corn Stover biomass Pre-Treatment by deep eutectic solvents (DES) for biomethane production process by bioresource anaerobic digestion. Sustain. 2021. 13, Page 10504, 10504. https://doi.org/10.3390/SU131910504 (2021).
Olatunji, K. O., Ahmed, N. A. & Ogunkunle, O. Optimization of biogas yield from lignocellulosic materials with different pretreatment methods: A review. Biotechnol. Biofuels 14, 1–34 https://doi.org/10.1186/S13068-021-02012-X (2021).
Dahunsi, S. O., Oranusi, S. & Efeovbokhan, V. E. Optimization of pretreatment, process performance, mass and energy balance in the anaerobic digestion of Arachis Hypogaea (Peanut) hull. Energy Convers. Manag. 139, 260–275. https://doi.org/10.1016/J.ENCONMAN.2017.02.063 (2017).
Şenol, H. Alkaline-Thermal and mild ultrasonic pretreatments for improving biomethane yields: impact on structural properties of chestnut shells. Fuel 354, 129373. https://doi.org/10.1016/J.FUEL.2023.129373 (2023).
Habib, S. S., Torii, S., Mol, K., Charivuparampil, A. & Nair, A. Optimization of the factors affecting biogas production using the Taguchi design of experiment method. Biomass 4, 687–703 (2024). https://doi.org/10.3390/BIOMASS4030038
Wang, S. et al. Enzyme pretreatment enhancing biogas yield from corn stover: feasibility, optimization, and mechanism analysis. J. Agric. Food Chem. 66, 10026–10032. https://doi.org/10.1021/ACS.JAFC.8B03086/ASSET/IMAGES/MEDIUM/JF-2018-03086Q_0014.GIF (2018).
Chen, Y. et al. Effects of acid/alkali pretreatments on lignocellulosic biomass Mono-Digestion and its Co-Digestion with waste activated sludge. J. Clean. Prod. 277, 123998. https://doi.org/10.1016/J.JCLEPRO.2020.123998 (2020).
Şenol, H. Anaerobic digestion of hazelnut (Corylus Colurna) husks after alkaline pretreatment and determination of new important points in logistic model curves. Bioresour Technol. 300, 122660. https://doi.org/10.1016/J.BIORTECH.2019.122660 (2020).
Budiyono, W.A. et al. The effect of pretreatment using sodium hydroxide and acetic acid to biogas production from rice straw waste. MATEC Web Conf. 101, 02011 (2017). https://doi.org/10.1051/MATECCONF/201710102011
Pinpatthanapong, K., Boonnorat, J., Glanpracha, N. & Rangseesuriyachai, T. Biogas production by Co-Digestion of sodium hydroxide pretreated Napier grass and food waste for community sustainability. Energy Sour. Part A Recover. Utilization Environ. Eff. 44, 1678–1692. https://doi.org/10.1080/15567036.2022.2055232 (2022).
Ďuriš, V., Bartková, R. & Tirpáková, A. Principal component analysis and factor analysis for an Atanassov IF data set. Mathematics 9. https://doi.org/10.3390/math9172067 (2021).
Olatunji, K. O., Ahmed, N. A. & Ogunkunle, O. Optimization of biogas yield from lignocellulosic materials with different pretreatment methods: A review. Biotechnol. Biofuels. 14, 159. https://doi.org/10.1186/s13068-021-02012-x (2021).
Li, X. et al. Enhancement of anaerobic digestion performance of corn straw via combined sodium Hydroxide-Cellulase pretreatment. Biochem. Eng. J. 187, 108652. https://doi.org/10.1016/j.bej.2022.108652 (2022).
Liang, Y. G. et al. Effect of Solid-State NaOH pretreatment on methane production from thermophilic Semi-Dry anaerobic digestion of Rose stalk. Water Sci. Technol. 73, 2913–2920. https://doi.org/10.2166/wst.2016.145 (2016).
Adeleke, O., Olatunji, O. O., Jen, T. C. & Olawuyi, I. Enhanced prediction of heating value of municipal solid waste using hybrid Neuro-Fuzzy model and decision Tree-Based feature importance assessment. Green. Energy Resour. 3, 100119. https://doi.org/10.1016/j.gerr.2025.100119 (2025).
Khan, A. A., Chaudhari, O. & Chandra, R. A. Review of ensemble learning and data augmentation models for class imbalanced problems: Combination, implementation and evaluation. Expert Syst. Appl. 244, 122778. https://doi.org/10.1016/j.eswa.2023.122778 (2024). .
Lan, T., Hu, H., Jiang, C., Yang, G. & Zhao, Z. A. Comparative study of decision tree, random forest, and convolutional neural network for Spread-F identification. Adv. Space Res. 65, 2052–2061. https://doi.org/10.1016/j.asr.2020.01.036 (2020).
Maier, H. R. et al. Exploding the myths: an introduction to artificial neural networks for prediction and forecasting. Environ. Model. Softw. 167, 105776. https://doi.org/10.1016/j.envsoft.2023.105776 (2023).
Ahmad, M. W., Mourshed, M. & Rezgui, Y. Trees vs neurons: comparison between random forest and ANN for High-Resolution prediction of Building energy consumption. Energy Build. 147, 77–89. https://doi.org/10.1016/j.enbuild.2017.04.038 (2017).
Wang, X., Huang, F. & Cheng, Y. Computational performance optimization of support vector machine based on support vectors. Neurocomputing 211, 66–71. https://doi.org/10.1016/j.neucom.2016.04.059 (2016).
Malashin, I., Tynchenko, V., Gantimurov, A., Nelyub, V. & Borodulin, A. Support vector machines in polymer science: A review. Polymers (Basel). 17 https://doi.org/10.3390/polym17040491 (2025).
Nalepa, J. & Kawulok, M. Selecting training sets for support vector machines: A review. Artif. Intell. Rev. 52, 857–900. https://doi.org/10.1007/s10462-017-9611-1 (2019).
Almomani, F. Prediction of biogas production from chemically treated Co-Digested agricultural waste using artificial neural network. Fuel 280, 118573. https://doi.org/10.1016/J.FUEL.2020.118573 (2020).
Singh, D., Tembhare, M., Pundalik, K., Dikshit, A. K. & Kumar, S. Machine learning based prediction of biogas generation from municipal solid waste: A Data-Driven approach. Process Saf. Environ. Prot. 192, 93–103. https://doi.org/10.1016/J.PSEP.2024.10.037 (2024).
Ganeshan, P., Bose, A., Lee, J., Barathi, S. & Rajendran, K. Machine learning for high solid anaerobic digestion: performance prediction and optimization. Bioresour Technol. 400, 130665. https://doi.org/10.1016/J.BIORTECH.2024.130665 (2024).
Author information
Authors and Affiliations
Contributions
OA – conceptualization, methodology, design, formal analysis, software, and writing original draft; KOO – conceptualization, methodology, design, investigation, data curation, visualization, and writing original draft; DMM – conceptualization, investigation, data curation, resources, supervision, and writing review and editing; TCJ – conceptualization, investigation, resources, visualization, supervision, and writing review and editing. All the authors read and agreed to publish the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Adeleke, O., Olatunji, K.O., Madyira, D.M. et al. Application of multimodal machine learning-based analysis for the biomethane yields of NaOH-pretreated biomass. Sci Rep 15, 24372 (2025). https://doi.org/10.1038/s41598-025-09527-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-09527-5