Abstract
To enhance thrombolysis eligibility in acute ischemic stroke, we developed a deep learning model to estimate stroke onset within 4.5 h using diffusion-weighted imaging (DWI) and fluid-attenuated inversion recovery (FLAIR) images. Given the variability in human interpretation, our multimodal Res-U-Net (mRUNet) model integrates a modified U-Net and ResNet-34 to classify stroke onset as < 4.5 or ≥ 4.5 h. Using DWI and FLAIR images from patients scanned within 24 h of symptom onset, the modified U-Net generated a DWI–FLAIR mismatch image, while ResNet-34 performed the final classification. mRUNet was evaluated against ResNet-34 and DenseNet-121 on an internal test set (n = 123) and two external test sets: a single-center (n = 468) and a multi-center (n = 1151). mRUNet achieved an area under the receiver operating characteristic curve (AUC-ROC) of 0.903 on the internal set and 0.910 and 0.868 on external sets, significantly outperforming ResNet-34 and DenseNet-121. Our mRUNet model demonstrated robust and consistent classification of the 4.5-h onset-time window across datasets. By leveraging DWI and FLAIR images as a tissue clock, this model may support timely and individualized thrombolysis in patients with unclear stroke onset, such as those with wake-up stroke, in clinical settings.
Similar content being viewed by others
Introduction
Acute ischemic stroke, resulting from sudden disruption of focal brain perfusion, imposes a significant burden on global mortality and disability1,2. The prognosis of acute ischemic stroke heavily depends on earlier reperfusion through timely intravenous thrombolysis, which is recommended within 4.5 h of symptom onset3,4,5. Yet approximately 35% of patients who experience strokes during sleep are unaware of the precise onset time6,7,8,9, resulting in their exclusion from thrombolysis according to international guidelines5,10. Previous clinical trials have reported inconsistent outcomes regarding the efficacy of thrombolysis beyond the 4.5-h window, underscoring the heterogeneity of patient responses and the importance of individualized decision-making in cases of unclear onset time11,12. The TRACE-III trial demonstrated improved functional outcomes with imaging-guided tenecteplase administration beyond the midpoint of sleep11, whereas the TIMELESS trial found no significant benefit of thrombectomy plus tenecteplase up to 24 h from last known well11,12.
Multiparametric magnetic resonance imaging (MRI) encompassing sequences differentially responsive to tissue pathology has aided in the estimation of unknown stroke-onset time13,14,15. Diffusion-weighted imaging (DWI) can rapidly reveal a reduced apparent diffusion coefficient (ADC) in ischemic lesions within minutes of stroke onset, whereas fluid-attenuated inversion recovery (FLAIR) detects increased water content within 1–4 h16,17,18. Per the 2019 guidelines, DWI–FLAIR mismatch (evidence level IIa), where an acute lesion is visible on DWI but not on FLAIR, can indicate that stroke onset occurred within 4.5 h or not, identifying potential candidates for thrombolysis6,15,19,20,21. Yet, human interpretation of mismatched DWI–FLAIR signals, which relies on dichotomized lesion visibility with inter-rater variability, may not consistently indicate stroke onset within 4.5 h15,19,21.
Machine and deep learning methods have been increasingly used to identify unclear stroke onset based on medical images22,23. Previously, machine learning algorithms predicted time since stroke onset using radiomic features extracted from DWI and FLAIR sequences as well as segmented lesions on DWI22,23. The MR-based deep learning algorithms classified time since stroke onset comparably to radiologists’ assessments on DWI–FLAIR mismatch and outperformed three machine learning models24,25. The convolutional neural network (CNN) is a recognized deep learning model for efficiently interpreting the complex brain imaging data26,27. Specifically, U-Net is renowned for its effectiveness in semantic segmentation and spatial localization for medical imaging analysis28,29. ResNet-34, which enables deep residual learning through skip connections and minimizes the vanishing-gradient problem, facilitates enhances diagnostic perforamance through image recognition30,31.
We postulated that leveraging deep learning using U-Net and ResNet-34 architectures could effectively uncover hidden clinical features in DWI and FLAIR images, aiding in estimating the unknown onset of ischemic stroke. We proposed a multimodal Res-U-Net (mRUNet) model that combines a modified U-Net and ResNet-34 to classify the 4.5-h stroke-onset window for thrombolysis. Using DWI and FLAIR within 24 h of symptom onset (n = 487), U-Net—modified by eliminating copy-and-crop connections—generates a DWI–FLAIR mismatch image, and ResNet-34 performs the final classification. The performance of the mRUNet model was thoroughly evaluated compared to those of ResNet-34 and DenseNet-121 in the internal test set (n = 123). Our proposed model’s performance was further validated on the external test set 1 (single-center, n = 468) and external test set 2 (multi-center, n = 1151).
Results
Onset time classification in the training set
The classification performance of our mRUNet model, along with two other deep learning models—ResNet-34 and DenseNet-121—on the training set (n = 487), are summarized in Table 1. In the training set, a fivefold cross-validation of the proposed mRUNet model yielded superior performance compared to ResNet-34 and DenseNet-121 with respect to area under the receiver operating characteristic curve (AUC-ROC), accuracy, specificity, negative predictive value (NPV), positive predictive value (PPV), and F1 score. The mRUNet model attained a mean AUC of 0.951 (95% CI 0.905–0.989), with a sensitivity of 0.921 and a specificity of 0.840. Conversely, the ResNet-34 model had a mean AUC-ROC of 0.888 (95% CI 0.806–0.951), with a sensitivity of 0.947 and a specificity of 0.680, and DenseNet-121 had a mean AUC-ROC of 0.835 (95% CI 0.756–0.908), with a sensitivity of 0.952 and a specificity of 0.609 (Table 1 and Fig. 1a).
Classification performance of the proposed multimodal Res-U-Net model. (A) In the training set (n = 487), mRUNet achieved an AUC of 0.951 (95% CI 0.905–0.989), outperforming ResNet-34 (AUC, 0.888; 95% CI 0.806–0.951) and DenseNet-121 (AUC, 0.835; 95% CI 0.756–0.908). (B) In the internal test set (n = 123), mRUNet achieved an AUC of 0.903 (95% CI 0.835–0.959), significantly higher than ResNet-34 (AUC, 0.768; 95% CI 0.670–0.853; p = 0.007) and DenseNet-121 (AUC, 0.790; 95% CI 0.760–0.867; p = 0.011). (C) In the external test set 1 (n = 468, single-center cohort), mRUNet achieved an AUC of 0.910 (95% CI 0.883–0.935), significantly higher than ResNet-34 (AUC, 0.790; 95% CI 0.744–0.833; p < 0.001) and DenseNet-121 (AUC, 0.814; 95% CI 0.775–0.853; p < 0.001). (D) In the external test set 2 (n = 1151, multi-center cohort), mRUNet achieved an AUC of 0.868 (95% CI 0.848–0.888), significantly outperforming ResNet-34 (AUC, 0.805; 95% CI 0.777–0.830; p < 0.001) and DenseNet-121 (AUC, 0.808; 95% CI 0.783–0.832; p < 0.001). AUC-ROC, area under the receiver operating characteristic curve; CI, confidence interval; mRUNet, multimodal Res-U-Net.
Onset time classification in the internal test set
The classification results of the proposed mRUNet model in the internal test set (n = 123) are summarized, along with those of ResNet-34 and DenseNet-121 in Table 1 and Fig. 1b. For the internal test set (n = 123), the classification performance was higher for our mRUNet model than for ResNet-34 and DenseNet-121 regarding evaluation metrics other than sensitivity and NPV (Table 1). The mean AUC-ROC for the mRUNet model was 0.903 (95% CI 0.835–0.959), which significantly higher than for that for ResNet-34 (0.768; 95% CI 0.670–0.853; p = 0.007) and DenseNet-121 (0.790; 95% CI 0.760–0.867; p = 0.011), as shown by the DeLong test (Table 1 and Fig. 1b).
The mRUNet model had an accuracy of 0.854, with a sensitivity of 0.825 and a specificity of 0.883, an NPV of 0.828, a PPV of 0.881, and an F1 score of 0.852. In contrast, the ResNet-34 algorithm achieved an accuracy of 0.707, with a sensitivity of 0.984 and a specificity of 0.417, an NPV of 0.962, a PPV of 0.639, and an F1 score of 0.775. The DenseNet-121 algorithm achieved an accuracy of 0.732, with a sensitivity of 0.841 and a specificity of 0.617, an NPV of 0.787, a PPV of 0.697, and an F1 score of 0.763.
Onset time classification in the external test sets
Our mRUNet model demonstrated outstanding and consistent performance of classification across two external tests sets (external test set 1, n = 468; external test set 2, n = 1151) as indicated in Table 1 and Fig. 1c,d.
The clinical trial using the single center-based external test set 1 (n = 468) showed that the mRUNet model attained a mean AUC-ROC of 0.910 (95% CI 0.883–0.935), which was significantly higher than that of ResNet-34 (0.790; 95% CI 0.744–0.833; p < 0.001) and DenseNet-121 (0.814; 95% CI 0.775–0.853; p < 0.001), as determined by the DeLong test. The mRUNet attained a sensitivity of 0.863 and a specificity of 0.863, an accuracy of 0.863, an NPV of 0.863, a PPV of 0.863, and a F1 score of 0.863 (Table 1 and Fig. 1c).
In the multi center-based external test set 2 (n = 1151), the proposed mRUNet model achieved a mean AUC-ROC of 0.868 (95% CI 0.848–0.888), which was significantly higher than for that of ResNet-34 (0.805; 95% CI 0.777–0.830; p < 0.001) and DenseNet-121 (0.808; 95% CI 0.783–0.832; p < 0.001), as determined by the DeLong test. The mRUNet model attained a sensitivity of 0.826 and a specificity of 0.813, an accuracy of 0.820, a NPV of 0.803, a PPV of 0.836, and a F1 score of 0.831 (Table 1 and Fig. 1d).
Critical regions for onset-time classification using Grad-CAM
The Grad-CAM results showed that the proposed model focused on the ischemic lesions when classifying stroke-onset time. The model’s attention to the lesions of the DWI images progressively expanded to include those in both the DWI and FLAIR images as the time since stroke onset increased (Fig. 2).
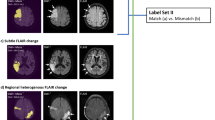
Critical regions for onset-time classification using Grad-CAM. Grad-CAM was applied to the lesion images extracted from the b1000 volumes of the diffusion-weighted imaging (DWI) or fluid-attenuated inversion recovery (FLAIR) images. Lesion regions marked with higher values, closer to red on the map, indicate a greater contribution to onset-time classification. (A) The Grad-CAM results are indicated for the stroke-onset time of 2.3 h (< 4.5 h). (B) The Grad-CAM results are indicated for the stroke-onset time of 5.8 h (> 4.5 h). (C) The Grad-CAM results are indicated for the stroke-onset time of 18.7 h (> 4.5 h). Grad-CAM, gradient-weighted class activation mapping; hrs, hours; DWI, diffusion-weighted imaging; FLAIR, fluid-attenuated inversion recovery.
Discussion
In our mRUNet model, a DWI–FLAIR mismatch image is generated by the U-Net, modified by eliminating copy-and-crop connections, and ResNet-34 performs the classification of stroke onset within 4.5 h. The proposed mRUNet model demonstrated superior performance in classifying the 4.5-h stroke-onset time window compared to that of the previously validated ResNet-34 and DenseNet-121 deep learning algorithms. Our mRUNet model showed high reliability and generalizability in classifying stroke onset within or beyond the 4.5-h treatment window across diverse clinical datasets, enabling rapid identification of thrombolysis-eligible patients in emergency settings. By leveraging DWI and FLAIR as a tissue clock, our model enhances stroke-onset estimation especially in cases of unknown or unclear onset, such as wake-up strokes, facilitating timely and individualized interventions for acute ischemic stroke.
The observed high AUC-ROC, accuracy, and F1 score (> 0.85) underscore our model’s reliable and balanced classification of stroke-onset time. In contrast, although the ResNet-34 and DenseNet-121 models had high sensitivity (> 0.80), they demonstrated low specificity (< 0.65), leading to their relatively high false-positive rates. Incorrectly identifying stroke-onset time as within 4.5 h (false positive) may lead to an inappropriate decision to administer thrombolysis, increasing the risk of intracranial hemorrhage1,2. In the classification of stroke-onset time, our mRUNet model (AUC-ROC, 0.903; sensitivity, 0.825, specificity, 0.883) outperformed both human visual assessment of DWI–FLAIR mismatch and previously validated algorithms21,22,23,25,32. Previous classification based on visual inspection of DWI–FLAIR mismatch yielded sensitivity values of 0.49–0.62 and specificity values of 0.78–0.9121,22.
Previous machine learning models for classifying stroke onset within 4.5 h —including random forest (RF), logistic regression (LR), and support vector machine (SVM)—have reported a wide range of performance, with AUC-ROCs between 0.62 and 0.89, sensitivities between 0.51 and 0.95, and specificities between 0.29 and 0.9122,23,25,32,33,34,35. Specifically, RF, LR, and SVM models analyzing DWI and FLAIR images showed higher sensitivity (up to 75.8%) compared to human readers22. An SVM with a radial basis function kernel achieved high internal (AUC-ROC, 0.896) and external (AUC-ROC, 0.895) validation performance for classifying wake-up stroke patients33. Similarly, a radiomics-based RF model achieved an AUC-ROC of 0.840 with a specificity of 0.91425,32,34. A radiomics-based model using DWI and ADC maps achieved an AUC-ROC of 0.754 and a sensitivity of 0.952, enhancing clinical applicability via a prototype web platform for real-world validation35. An automated machine learning approach combining cross-modal CNN-based lesion segmentation and feature extraction from DWI and FLAIR achieved an accuracy of 0.805, outperforming human assessments23. Compared with previous machine learning models, the mRUNet model achieved higher AUC-ROCs (0.868–0.910) and maintained balanced sensitivity (0.825–0.863) and specificity (0.813–0.883) across internal and external datasets, suggesting more robust and clinically applicable performance.
Recent deep learning advances in neuroimaging have automated and improved the classification of the 4.5-h thrombolysis window, demonstrating performance exceeding that of human experts, particularly in cases with uncertain onset times and varied imaging modalities25,32,36,37. A deep learning model using perfusion-weighted imaging improved stroke-onset classification with an AUC-ROC of 0.68, compared to a clinical baseline of 0.5836. A ResNet-34-based model attained an accuracy of 0.726, a sensitivity of 0.712, and a specificity of 0.74125. Semi-supervised learning with a tissue clock framework detected DWI–FLAIR mismatch with an AUC-ROC of 0.74332. A synchronized dual-stage network, which jointly optimized infarct segmentation and time-since stroke classification using DWI and FLAIR in a parallel architecture, achieved an AUC-ROC of 0.879 and an accuracy of 0.800, outperforming sequential models37. Deep learning models offer the advantage of automatically extracting complex, non-linear relationships from high-dimensional inputs, while enhancing interpretability through gradient-based or feature-based visualization techniques36,38.
Advances in stroke therapy have introduced bridging therapy, involving the sequential application of intravenous thrombolysis followed by mechanical thrombectomy to optimize reperfusion outcomes39,40. Mechanical thrombectomy, performed with or without prior thrombolysis, has extended the treatment window to 6–24 h after onset and significantly improved outcomes in patients with acute ischemic stroke39,40. Developments in computed tomography (CT) perfusion, CT/MR angiography, and MRI penumbra imaging, coupled with the integration of artificial intelligence, have refined patient selection and expedited procedural planning41,42. Deep learning models, such as a CNN trained on CT perfusion features, have achieved high performance in predicting endovascular thrombectomy eligibility (AUC-ROC, 0.935), outperforming traditional machine learning methods41. The present study focused on identifying candidates for intravenous thrombolysis within the 4.5-h therapeutic window based on DWI and FLAIR imaging, constrained by the absence of vascular imaging modalities and limited documentation of vessel status and mechanical thrombectomy outcomes. Future work will aim to integrate vascular imaging techniques along with clinical outcome data to extend the model’s applicability to mechanical thrombectomy selection and bridging therapy strategies39,41,42.
Notably, our modified U-Net architecture, with copy-and-crop connections eliminated from the traditional architecture, effectively integrated six preprocessed DWI and FLAIR images co-registered with lesion maps into a single representative DWI–FLAIR mismatch image. This DWI–FLAIR mismatch image, functioning as a tissue clock, encompasses the multidimensional trajectory and pathophysiology of ischemic stroke lesions, facilitating stroke-onset classification through the ResNet-34 architecture6,15,19,20,21. Our mRUNet model, which combined the modified U-Net and ResNet-34 deep neural architectures, is designed to interpret complicated multimodal images through deep computational layers, for precise stroke-onset classification26,27. Our architecture incorporates the strengths of the traditional U-Net in semantic feature extraction of medical images and those of ResNet-34 in deep residual learning with skip connections and computation efficiency28,29,30,31. Confirming the DWI-FLAIR mismatch as a tissue clock, the model’s attention to lesions in the DWI images progressively extended to those in both the DWI and FLAIR images as the time since stroke-onset increased.
Our proposed mRUNet model has several notable advantages. First, it showed high generalizability and reliability, achieving robust stroke-onset classification within the 4.5-h time window across three test sets. In a single-center retrospective study, an unbiased researcher confirmed the model’s high classification performance (AUC-ROC, 0.910) using external test set 1. Additionally, the model exhibited satisfactory performance (AUC-ROC, 0.868) in the multi-center external test set 2, which included younger patients and those with more severe stroke symptoms than those in external test set 1. These results highlight the model’s effectiveness in reliably classifying stroke-onset time across heterogeneous datasets with varying demographics, clinical characteristics, and imaging conditions43,44. Second, our model has high applicability in real-world clinical settings, particularly for patients with ischemic stroke lesions of varying locations and sizes. Our dataset included patients with multiple lesions, lesions outside the middle cerebral artery territories and even those with very small lesion volumes. Our proposed model, designed for enhanced efficacy, relies exclusively on clinically essential MR images, particularly the DWI and FLAIR sequences acquired within 24 h of onset. This tissue clock may facilitate reliable onset-time classification in clinical practice by minimizing subjective bias in data acquisition arising from variations in patient reports and clinician evaluation25,32,36,37.
The following limitations should be considered when interpreting the study results. First, we excluded patients with severe leukoaraiosis or suboptimal image quality, which may have produced incorrect FLAIR signals. This exclusion may have introduced selection bias and limited the model’s applicability to stroke populations with comorbid white matter disease45,46. Second, restricting the study cohort to individuals who underwent both DWI and FLAIR imaging within 24 h of symptom onset ensured accurate ground truth labeling; however, it may reduce the generalizability of the findings to broader stroke populations in which such imaging protocols are not routinely performed45,46. Third, although DWI lesion boundaries were initially delineated by an extensively trained neurological nurse specialist and subsequently reviewed by two board-certified neurologists to minimize inter-observer variability and emulate real-world clinical workflows, the initial reliance on a non-physician observer may have introduced a degree of subjectivity45,47.
Although our model showed robust classification within the critical 4.5-h stroke-onset window —the established optimal therapeutic window for thrombolysis—future studies are needed to expand stroke-onset estimation across broader time windows to enable more individualized and flexible treatment strategies3,4. This need is underscored by recent clinical trials: TRACE-III showed improved outcomes with imaging-guided tenecteplase treatment beyond sleep onset, whereas TIMELESS did not demonstrate significant benefit for thrombectomy combined with tenecteplase up to 24 h after the last known well11,12. Additionally, further validation is required across a wider range of acute stroke interventions, including mechanical thrombectomy, bridging therapy, and real-time deployment in emergency settings39,40,48. Finally, as our model was developed using six multimodal inputs derived from preprocessed DWI, FLAIR, and manually delineated lesion maps, future efforts should focus on automating the entire workflow from raw imaging to output, thereby improving cost-effectiveness and clinical applicability.
Our proposed mRUNet model, utilizing integrated DWI and FLAIR mismatch images, outperformed ResNet-34 and DenseNet-121 in classifying stroke onset within the 4.5-h window, showing high generalizability and reliability across internal and external test sets. The mRUNet model allows emergency department clinicians to rapidly identify thrombolysis-eligible patients by determining whether stroke onset occurred within the previous 4.5 h, particularly in cases with unknown or unclear onset times, such as wake-up strokes. These findings suggest that DWI and FLAIR images capture the intricate pathophysiology of ischemic lesions, serving as a promising tissue clock for stroke-onset estimation. Our mRUNet model, utilizing clinical MR sequences enhances our understanding of individual lesion timing and trajectory, contributing to the accurate determination of symptom onset within the 4.5-h window. This capability will empower clinicians to make informed decisions on tailored interventions, emphasizing the clinical utility of DWI and FLAIR mismatch as a clinical tissue clock.
Methods
Internal dataset
This study retrospectively evaluated patients with acute ischemic stroke who visited Asan Medical Center, Seoul, South Korea between January 2013 and November 2017 (n = 1974). Among these patients, those who underwent both DWI and FLAIR sequences of brain MRI within 24 h of symptom onset were included (n = 1055). Moreover, 45 patients with unclear onset time and 319 patients with unknown onset time—those lacking identical last known normal time and first observed abnormal time—were excluded, leading to the inclusion of 691 patients. After visually examining the MRI images, 81 patients having acute lesions within the extensive leukoaraiosis or low image quality for analysis were excluded. This resulted in the inclusion of 610 patients in the internal dataset (Fig. 3).
Flowchart of the study population in the internal dataset. A flowchart of the study population and process is presented for the internal dataset. This dataset included 610 patients with acute ischemic stroke who had undergone proper diffusion-weighted imaging (DWI) and fluid-attenuated inversion recovery (FLAIR) scans within 24 h of a clearly defined symptom onset. The patients were divided into training (n = 487) and test (n = 123) sets at an 8:2 ratio. DWI, diffusion-weighted imaging; FLAIR, fluid-attenuated inversion recovery.
The final 610 patients were split into a training set (n = 487) and a test set (n = 123) at an 8:2 ratio, without overlapping patients. The characteristics of the internal dataset (n = 610) are summarized in Table 2. The training (n = 487) and test (n = 123) sets showed nonsignificant differences regarding age, sex, National Institutes of Health Stroke Scale (NIHSS) score on admission, and time from stroke to MRI (Table 1). The study was conducted in accordance with the Declaration of Helsinki. The study protocol was approved by the Institutional Review Board of Asan Medical Center (IRB No. 2013-0162). The requirement for written informed consent was waived by the Institutional Review Board of Asan Medical Center due to the retrospective nature of the study.
External dataset
Two external test sets were used to validate the reproducibility of the proposed mRUNet model. The first external test set comprised 468 patients who visited Asan Medical Center from January 2017 to March 2022. A single-blind retrospective clinical trial (IRB No. 2023-0762) was conducted to assess the performance of our mRUNet model integrated into the Stroke Onset Time Artificial Intelligence (SOTA) software in classifying the 4.5-h time window for stroke onset. The second test set was retrospectively chosen from the Korean Stroke Neuroimaging Initiative (KOSNI) database, and comprised 1151 patients who had visited eight tertiary stroke centers in South Korea, excluding Asan Medical Center, between January 2013 and December 2016.
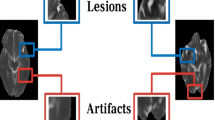
Participants in the external test sets were selected based on three specific inclusion criteria: age ≥ 20 years with a diagnosis of acute ischemic stroke by a medical professional; clearly defined symptom onset time; and DWI and FLAIR data free of substantial artifacts and acquired within 24 h of symptom onset.
The clinical characteristics of the two external test sets are summarized in Table 2. The external test set 2 included patients with older age (p = 0.005) and higher NIHSS scores on admission (p < 0.001) than those in the external test set 1. There were no significant differences in sex or time from stroke onset to MRI between the two external test sets (Table 2).
Image processing and infarct segmentation
Every patient with acute ischemic stroke underwent DWI and FLAIR sequences on brain MRI scans. ADC maps were generated automatically from the DWI scans using built-in software, and Python version 3.8.10 was used to extract the three-dimensional (3D) b0 and b1000 volumes from the 4D DWI images.
The ischemic stroke lesion for each patient was manually segmented and inspected by neurologists. Prior to the main annotation process, a neurological nurse specialist with extensive experience in stroke imaging (Gayoung Park), blinded to stroke-onset information, performed preliminary delineations on several sample cases, which were reviewed and approved by two board-certified stroke neurologists. She then manually delineated the lesion boundaries on b1000 images using MRIcron software, identifying lesions as hyperintense on b1000 and hypointense on ADC. Various tools within MRIcron, including the 3D fill tool, pen tool, and closed pen tool, were used for manual delineation. The resulting lesion masks were saved in both volume-of-interest and NIfTI file formats. Subsequently, two board-certified stroke neurologists (Eun-Jae Lee and Dong-Wha Kang) independently reviewed the lesion masks and reached a consensus after applying necessary corrections where discrepancies were identified. The manually delineated binary lesion masks were linearly registered to the Montreal Neurological Institute (MNI) 152 standard space.
DWI and FLAIR images underwent the following preprocessing steps: initially, b0, b1000, ADC, and FLAIR images were skull-stripped using FreeSurfer version 7.4.1 and corrected for N4 bias field inhomogeneity by ANTs version 2.4.4. Subsequently, FLAIR images were co-registered to the b0 image (FLAIR_b0). The midsagittal plane of each FLAIR_b0 image was registered to the standard space through warping based on the inverse deformation. To facilitate quantitative comparisons between infarct regions and the contralateral side, ratio maps were generated by mirroring the images along the fitted midsagittal plane (FLAIR_b0_ratiomap). Intensity normalization, using z-score transformation, was applied to b0, b1000, ADC, and FLAIR_b0 images. Additionally, b1000 images were subjected intensity-based k-means clustering for categorization into three subgroups (b1000_k3). Processed DWI (b0, b1000, b1000_k3, ADC) and FLAIR images (FLAIR_b0, FLAIR_b0_ratiomap) were then linearly registered into the MNI 152 standard space. The six images in the standard space were co-registered to the binary lesion mask in the standard space, serving as inputs for subsequent modeling.
Onset time classification model: multimodal Res-U-Net
We developed a mRUNet architecture that integrates a modified 3D U-Net and ResNet-34 to classify stroke-onset time within the 4.5-h therapeutic window. A schematic illustration of the proposed model is shown in Fig. 4.
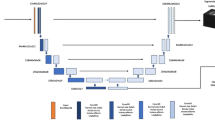
Overview of the proposed mRUNet architecture for stroke-onset time classification. Six preprocessed 3D multimodal images (DWI and FLAIR) are used as a six-channel input. A modified U-Net, consisting of contracting and expanding paths without copy-and-crop connections, generates a single 3D DWI–FLAIR mismatch image by focusing on global intensity differences. This mismatch image is then processed by a 3D ResNet-34, which extracts high-level semantic features through sequential residual blocks. After global average pooling and a fully connected layer with 300 units, a softmax output predicts whether stroke onset occurred within 4.5 h. 3D, three-dimensional; DWI, diffusion-weighted imaging; FLAIR, fluid-attenuated inversion recovery; mRUNet, multimodal Res-U-Net.
The input consists of six preprocessed 3D multimodal images (91 × 109 × 70), derived from DWI and FLAIR sequences and co-registered with lesion maps. These six images are combined as six input channels, and fed into the network.
The baseline U-Net architecture consists of a symmetric structure comprising a contracting path and an expanding path for semantic feature extraction and spatial information recovery28. The contracting path applies repeated 3 × 3 × 3 convolutions with rectified linear unit (ReLU) activations, followed by 2 × 2 × 2 max pooling layers for downsampling. The expanding path restores spatial resolution through 2 × 2 × 2 transposed convolutions, combined with 3 × 3 × 3 convolutions and ReLU activations, culminating in a 1 × 1 × 1 convolution at the output layer. Copy-and-crop connections are traditionally employed to concatenate high-resolution features from the contracting path to the expanding path, preserving spatial precision.
In the proposed modification, the copy-and-crop connections were removed to focus the model on capturing global intensity differences rather than fine-grained spatial structures. Preserving detailed spatial information was unnecessary for the DWI–FLAIR mismatch detection task and could introduce redundant noise. Removing these connections promoted deeper semantic feature learning and improved generalization across heterogeneous datasets. The modified U-Net outputs a single three-dimensional DWI–FLAIR mismatch image, which serves as input for the subsequent classification stage.
The DWI–FLAIR mismatch image is then processed by a 3D ResNet-34 network30. The ResNet-34 begins with a 7 × 7 × 7 convolutional layer followed by max pooling and consists of four sequential residual stages. The first residual stage contains three residual blocks with 64 filters, the second stage contains four residual blocks with 128 filters, the third stage includes six residual blocks with 256 filters, and the final stage consists of three residual blocks with 512 filters. Each residual block comprises two 3 × 3 × 3 convolutional layers, each followed by batch normalization and a ReLU activation function. Skip connections within residual blocks enable efficient gradient propagation, mitigating the vanishing-gradient problem commonly encountered in deep networks. After the final residual stage, global average pooling aggregates the feature maps into a compact representation, which is then passed through a fully connected layer with 300 hidden units. A softmax activation function produces the final probability estimate for stroke-onset time being within the 4.5-h window.
Model training and evaluation
The proposed model was implemented in PyTorch and trained on two RTX 3090 GPUs with a batch size of 4. Adaptive moment estimation (Adam) was used as the optimizer, with a learning rate of 0.000149. Early stopping was applied if the validation loss did not improve over 100 consecutive epochs, with a maximum training limit of 1,000 epochs.
For labeling, patients were categorized into two groups: those imaged at < 4.5 h from known symptom onset were assigned a positive label, and those imaged at ≥ 4.5 h were assigned a negative label. A fivefold cross-validation scheme was employed to compare the performance of different methods. In each fold, the dataset was randomly split into a training set (80%) and a hold-out evaluation set (20%), which we refer to as the internal test set to distinguish it from the separate external test sets used for generalizability assessment. Each algorithm was trained on four folds and evaluated on the remaining fold (internal test set). The optimal algorithm was selected by averaging performance metrics across the five internal test sets.
Classification performance for 4.5-h stroke-onset time window was assessed using multiple evaluation metrics, including the AUC-ROC, accuracy, sensitivity, specificity, NPV, PPV, and F1 score. The 95% confidence intervals (CIs) for the AUC-ROC were estimated using a bootstrapping method with 1000 resamples. The calculation formulas were defined as follows:
Model evaluation was conducted across three test sets: an internal test set (n = 123) from the same institution, and two external test sets, including a single-center cohort (external test set 1, n = 468) and a multi-center cohort (external test set 2, n = 1151). ROC curve thresholds were determined using the Youden index (sensitivity + specificity − 1). Pairwise comparisons of the AUC-ROCs between mRUNet and ResNet-34, and between mRUNet and DenseNet-121, were performed using the DeLong test.
Explainability analysis
To provide interpretability of the model’s predictions, Gradient-weighted Class Activation Mapping (Grad-CAM) was applied50. Grad-CAM heatmaps were generated to visualize the ischemic regions contributing most significantly to the classification decision. Heatmaps were produced for lesion areas extracted from b1000 volumes of DWI and FLAIR images. Three representative cases with varying lesion sizes and stroke-onset times were selected. Grad-CAM scores were normalized between 0 and 1, with higher scores (visualized in red) indicating greater contributions to the onset-time classification.
Statistical analyses
Demographic and clinical characteristics were compared between the training and internal test sets as well as between external test set 1 and external test 2. Data normality was first assessed using Shapiro–Wilk test. Continuous variables, showing a non-parametric distribution, were compared using the Mann–Whitney U test, while categorical variables were compared using Fisher’s exact test. AUC-ROC values were compared between models using the DeLong test, for pairwise comparisons between mRUNet and ResNet-34, and between mRUNet and DenseNet-121. Statistical analyses were conducted using STATA SE version 16 (StataCorp, College Station, TX). All tests were two-sided, with a significance threshold of p < 0.05.
Data availability
Data collected for this study can be made available (in the form of any or all of the de-identified data in the study database, study protocol, statistical analysis plan, and analytic code) to researchers who provide a methodologically sound research proposal to assist with achieving the aims in the approved proposal. Data will be available from the time of publication of the Article in print upon reasonable request. Proposals should be directed to the corresponding author.
References
Brott, T. & Bogousslavsky, J. Treatment of acute ischemic stroke. N. Engl. J. Med. 343, 710–722 (2000).
Powers, W. J. Acute ischemic stroke. N. Engl. J. Med. 383, 252–260 (2020).
Bluhmki, E. et al. Stroke treatment with alteplase given 3·0–4·5 h after onset of acute ischaemic stroke (ECASS III): Additional outcomes and subgroup analysis of a randomised controlled trial. Lancet Neurol. 8, 1095–1102 (2009).
Hacke, W. et al. Thrombolysis with alteplase 3 to 4.5 hours after acute ischemic stroke. N. Engl. J. Med. 359, 1317–1329 (2008).
Powers, W. J. et al. 2018 guidelines for the early management of patients with acute ischemic stroke: A guideline for healthcare professionals from the American heart association/American stroke association. Stroke 49, e46–e99 (2018).
Kang, D.-W., Kwon, J. Y., Kwon, S. U. & Kim, J. S. Wake-up or unclear-onset strokes: Are they waking up to the world of thrombolysis therapy?. Int. J. Stroke 7, 311–320 (2012).
Kim, Y.-J., Kim, B. J., Kwon, S. U., Kim, J. S. & Kang, D.-W. Unclear-onset stroke: Daytime-unwitnessed stroke vs. wake-up stroke. Int. J. Stroke 11, 212–220 (2016).
Mackey, J. et al. Population-based study of wake-up strokes. Neurology 76, 1662–1667 (2011).
Rimmele, D. L. & Thomalla, G. Wake-up stroke: Clinical characteristics, imaging findings, and treatment options—An update. Front. Neurol. 5, 35 (2014).
Jauch, E. C. et al. Guidelines for the early management of patients with acute ischemic stroke. Stroke 44, 870–947 (2013).
Xiong, Y. et al. TRACE III: Tenecteplase reperfusion therapy in acute ischaemic cerebrovascular events. Stroke Vasc. Neurol. 9, 82–89 (2024).
Albers, G. W. et al. TIMELESS trial design: Thrombolysis in imaging-eligible, late-window patients. Int. J. Stroke 18, 237–241 (2023).
Kim, B. J. et al. Magnetic resonance imaging in acute ischemic stroke treatment. J. Stroke 16, 131 (2014).
Serena, J., Dávalos, A., Segura, T., Mostacero, E. & Castillo, J. Stroke on awakening: Looking for a more rational management. Cerebrovasc. Dis. 16, 128–133 (2003).
Thomalla, G. et al. Negative FLAIR identifies acute ischemic stroke within 3 hours. Ann. Neurol. 65, 724–732 (2009).
Hoehn-Berlar, M. et al. Temporal evolution of ADC and T1/T2 after MCA occlusion in rats. Magn. Reson. Med. 34, 824–834 (1995).
Mintorovitch, J. et al. DWI vs T2-weighted MRI for ischemia and reperfusion detection. Magn. Reson. Med. 18, 39–50 (1991).
Venkatesan, R. et al. Measuring water content in focal ischemia via MRI. Magn. Reson. Med. 43, 146–150 (2000).
Aoki, J. et al. FLAIR for estimating onset time in acute ischemic stroke. J. Neurol. Sci. 293, 39–44 (2010).
Kang, D.-W. et al. MRI-guided reperfusion therapy for unclear-onset stroke: Restore. Stroke 43, 3278–3283 (2012).
Thomalla, G. et al. DWI-FLAIR mismatch for stroke within 4.5 hours: PRE-FLAIR study. Lancet Neurol. 10, 978–986 (2011).
Lee, H. et al. Machine learning approach to identify stroke within 4.5 hours. Stroke 51, 860–866 (2020).
Zhu, H. et al. Automatic ML for ischemic stroke onset time via DWI/FLAIR. Neuroimage Clin. 31, 102744 (2021).
Ho, K. C., Speier, W., El-Saden, S. & Arnold, C. W. Predicting stroke onset using imaging features. AMIA Annu. Symp. Proc. 892 (2017).
Polson, J. S. et al. Deep learning to identify thrombolysis-eligible stroke patients. J. Neuroimaging 32, 1153–1160 (2022).
Gu, J. et al. Recent advances in convolutional neural networks. Pattern Recognit. 77, 354–377 (2018).
Yamashita, R., Nishio, M., Do, R. K. G. & Togashi, K. CNNs in radiology. Insights Imaging 9, 611–629 (2018).
Ronneberger, O., Fischer, P. & Brox, T. U-Net: Convolutional networks for biomedical segmentation. In MICCAI 2015 234–241 (Springer, 2015).
Siddique, N., Paheding, S., Elkin, C. P. & Devabhaktuni, V. U-Net and variants in medical image segmentation. IEEE Access 9, 82031–82057 (2021).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. In CVPR 2016 770–778 (IEEE, 2016).
Talo, M. et al. CNNs for brain disease classification using MRI. Comput. Med. Imaging Graph. 78, 101673 (2019).
Polson, J. et al. Semi-supervised stroke detection using DWI/FLAIR. In EMBC 2021 2258–2261 (IEEE, 2021).
Jiang, L. et al. ML model for wake-up stroke onset time. Eur. Radiol. 32, 3661–3669 (2022).
Liu, Z. et al. Radiomics-based ML for posterior circulation stroke. Neuroradiology 66, 1141–1152 (2024).
Zhang, Y. Q. et al. Radiomic ML to classify ischemic stroke onset time. J. Neurol. 269, 350–360 (2022).
Ho, K. C., Speier, W., El-Saden, S. & Arnold, C. W. Classifying stroke onset using imaging features. AMIA Annu. Symp. Proc. 892–901 (2017).
Zhang, X. et al. SDS-Net: Deep network for thrombolysis window prediction. J. Imaging Inform. Med. https://doi.org/10.1007/s10278-024-01308-2 (2024).
Akay, E. M. Z. et al. DL stroke-onset prediction vs. DWI–FLAIR mismatch. Neuroimage Clin. 40, 103544 (2023).
Bakka, A. G. et al. Advances and challenges in stroke therapy. Cureus 17, e78288 (2025).
Yeo, L. L. L. et al. Updates on thrombectomy: Targets and devices. Front. Neurol. 12, 712527 (2021).
Gao, H. et al. Deep learning with CT perfusion for EVT eligibility. Front. Med. 10, 1085437 (2023).
Song, S. S. Advanced imaging in acute stroke. Semin. Neurol. 33, 436–440 (2013).
Arsava, E. M. et al. Predictive validity of stroke etiology classification. JAMA Neurol. 74, 419–426 (2017).
Wu, O. et al. Big data phenotyping in acute ischemic stroke. Stroke 50, 1734–1741 (2019).
Hernandez Petzsche, M. R. et al. ISLES 2022 MRI stroke lesion dataset. Sci. Data 9, 762 (2022).
Maier, O. et al. ISLES 2015 benchmark for stroke lesion segmentation. Med. Image Anal. 35, 250–269 (2017).
Ortiz-Ramón, R. et al. Texture analysis for ischemic stroke lesion detection. Comput. Med. Imaging Graph. 74, 12–24 (2019).
Haq, M. et al. FDA-approved AI for stroke triage: A review. Cureus 16, (2024).
Kingma, D. P. & Ba, J. Adam: A method for stochastic optimization. arXiv preprint arXiv:1412.6980 (2014).
Selvaraju, R. R. et al. Grad-CAM: Visual explanations from deep networks. In ICCV 2017 618–626 (IEEE, 2017).
Acknowledgements
This research was supported by a grant from the Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI) funded by the Ministry of Health & Welfare (HR18C0016), a grant from the National IT Industry Promotion Agency (NIPA) funded by the Korea government (MSIT) (No. S0252-21-1001, Development of AI Precision Medical Solutions (Doctor Answer2.0)), and a grant from the National Research Foundation of Korea (NRF) funded by the Korean government (MSIT) (2022R1F1A1060778), Republic of Korea.
Funding
The funders had no role in the design and conduct of the study; collection, management, analysis, and interpretation of the data; preparation, review, or approval of the manuscript; and decision to submit the manuscript for publication.
Author information
Authors and Affiliations
Contributions
E-JL and D-WK conceived the study and designed the experiments. EN and Y-SK performed the data modelling and data visualisation. EN, E-JL and D-WK wrote the original draft and reviewed and edited the manuscript. D-IC, HJC, JL, J-KC, M-SP, KHY, J-MJ, SHA, D-EK, J-HL, K-SH, S-IS, and K-PP contributed to data acquisition and clinical expertise. JYC, BJK, SU.K, GP, H-SJ, and JH performed the experiments and result interpretation. EN, Y-SK, E-JL and D-WK directly accessed and verified the data in the study. All authors reviewed and approved the manuscript. All authors had full access to all the data in the study and had final responsibility for the decision to submit for publication.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Namgung, E., Kim, Y.S., Lee, EJ. et al. Deep learning to identify stroke within 4.5 h using DWI and FLAIR in a prospective multicenter study. Sci Rep 15, 26262 (2025). https://doi.org/10.1038/s41598-025-10804-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-10804-6