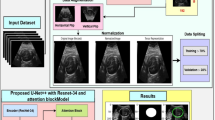
Abstract
B-ultrasound results are widely used in early pregnancy loss (EPL) prediction, but there are inevitable intra-observer and inter-observer errors in B-ultrasound results especially in early pregnancy, which lead to inconsistent assessment of embryonic status, and thus affect the judgment of EPL. To address this, we need a rapid and accurate model to predict pregnancy loss in the first trimester. This study aimed to construct an artificial intelligence model to automatically extract biometric parameters from ultrasound videos of early embryos and predict pregnancy loss. This can effectively eliminate the measurement error of B-ultrasound results, accurately predict EPL, and provide decision support for doctors with relatively little clinical experience. A total of 630 ultrasound videos from women with early singleton pregnancies of gestational age between 6 and 10 weeks were used for training. A two-stage artificial intelligence model was established. First, some biometric parameters such as gestational sac areas (GSA), yolk sac diameter (YSD), crown rump length (CRL) and fetal heart rate (FHR), were extract from ultrasound videos by a deep neural network named A3F-net, which is a modified neural network based on U-Net designed by ourselves. Then an ensemble learning model predicted pregnancy loss risk based on these features. Dice, IOU and Precision were used to evaluate the measurement results, and sensitivity, AUC etc. were used to evaluate the predict results. The fetal heart rate was compared with those measured by doctors, and the accuracy of results was compared with other AI models. In the biometric features measurement stage, the precision of GSA, YSD and CRL of A3F-net were 98.64%, 96.94% and 92.83%, it was the highest compared to other 2 models. Bland-Altman analysis did not show systematic deviations between doctors and AI. The mean and standard deviation of the mean relative error between doctors and the AI model was 0.060 ± 0.057. In the EPL prediction stage, the ensemble learning models demonstrated excellent performance, with CatBoost being the best-performing model, achieving a precision of 98.0% and an AUC of 0.969 (95% CI: 0.962–0.975). In this study, a hybrid AI model to predict EPL was established. First, a deep neural network automatically measured the biometric parameters from ultrasound video to ensure the consistency and accuracy of the measurements, then a machine learning model predicted EPL risk to support doctors making decisions. The use of our established AI model in EPL prediction has the potential to assist physicians in making more accurate and timely clinical decision in clinical application.
Similar content being viewed by others
Introduction
Early pregnancy loss (EPL) also called early miscarriage is the most common complication of pregnancy. It is estimated that there are 44 miscarriages per minute, with 23 million miscarriages in total worldwide each year1. About 80% of miscarriages occur in the first trimester. Excluding human factors, there are many risk factors for miscarriage, among which abnormal ultrasound video presentation is highly predictive of early pregnancy loss2,3,4 Several studies have shown that the size of the gestational sac (GS), oversized or missing yolk sac (YS), and the crown-rump length (CRL) in early pregnancy embryos are associated with the final pregnancy outcome5,6,7. It has also been shown that fetal heart rate is a strong predictor of embryonic developmental outcome8. Therefore, in the ultrasound videos of early pregnancy embryos obtained by TVS (Transvaginal Sonography), the biological features of primary clinical interest are gestational sac area (GSA), CRL yolk sac diameter (YSD) and fetal heart rate (FHR)9. GSA and CRL can predict the gestational age of the embryo in early pregnancy10YSD can be used as an indicator of the observation of the development of the embryo in early pregnancy11and FHR is a marker of embryo survival in early pregnancy embryos8. As mentioned in Prenatal Ultrasonographic Diagnosis of Fetal Abnormalities12the gestational sac area (GSA), yolk sac diameter (YSD), and crown-rump length (CRL) should be measured on the largest plane of the gestational sac, which is also the standard practice in clinical settings. For the measurement of GSA, CRL and YSD in the clinic, it is necessary to select the key frame of GS, CR and YS in the video, i.e., the measurements at the maximum of GSA, CRL and YSD are used as the actual feature values, and for the measurement of FHR, it is necessary for doctors to count the frequency of the fetal heartbeats in the video. However, early pregnancy ultrasound video presents inherent diagnostic challenges due to technical limitations including acoustic noise, artifact interference, and low tissue contrast, which collectively impair the visualization of critical soft tissue boundaries13. These image quality constraints, compounded by the subjective nature of manual biometric measurements from ultrasound video analysis, not only increase clinical workload but also introduce measurement variability that may compromise the accuracy of early pregnancy loss (EPL) risk assessment. Furthermore, the interpretation of embryonic development parameters derived from first-trimester ultrasound examinations poses significant clinical decision-making challenges, particularly for less experienced practitioners. This diagnostic complexity arises from frequent occurrences of conflicting biometric parameters - where some measurements fall within normal ranges while others deviate - requiring advanced pattern recognition skills and clinical correlation that typically develop through extensive supervised practice. Therefore, an intelligent model that is able to solve problem raised above and improve segmentation accuracy and clinical decision-making is in urgent need.
Given these diagnostic challenges, AI has emerged as a promising solution. Deep learning, in particular, has demonstrated remarkable success in medical imaging. There are many impressive examples of deep learning applications in medical imaging14,15,16. Among lots of deep learning network models, U-Net17,18 is the most commonly used in medical image segmentation. There is also a lot of research that applies different machine learning methods to medical diagnosis5,19,20,21,22,23,24. Due to the small size of the yolk sac and embryo in early pregnancy, they occupy only a minimal area in ultrasound videos, making our segmentation particularly challenging. There are fewer studies on segmentation of early pregnancy ultrasound images using deep learning algorithms, and they mainly focus on the region of the gestational sac. For example, Samuel Sutton et al.25 segmented the germ region by U-Net, and the final classification accuracy Dice was 0.95, but the data used were foetuses from 11 to 19 weeks, which makes the task of segmenting the germ easier compared to this paper. At present, most deep learning-based segmentation studies on embryonic ultrasound imaging are concentrated on mid-to-late pregnancy. These studies typically focus on segmenting fetal structures such as the fetus itself, the fetal heart, or the fetal head circumference. For example, Looney et al.26 constructed OxNNet based on 3D U-Net to segment the placenta based on 3D ultrasound imaging of embryos, achieving an average dice coefficient of 0.84. There are fewer studies on the prediction of pregnancy outcomes in early pregnancy through machine learning, and most of them are based on machine learning modelling of biological indicators obtained from obstetric tests performed on pregnant women. For example, Reza Arabi Belaghi et al.27 predicted by logistic regression with a final AUC of 0.80. Tamar Amitai et al.28 predicted early pregnancy miscarriage using XGBoost and Random Forest with an AUC of only 0.68. etc.
In order to solve the challenges mentioned earlier, this study adopts a hybrid artificial intelligence approach. It modifies U-Net, overcomes the measurement difficulties caused by the small biological features and fuzzy boundaries in the ultrasound image, measures each frame in the ultrasound video accurately, and then inputs the features extracted from the video into a decision model established through machine learning to distinguish whether the early pregnant woman is at risk of EPL. Experimentally, the Dice of our model can achieve 98.82%, 96.02%, and 88.66% for the measurement tasks of GS, YS, and CR in ultrasound videos, respectively, and an AUC value of 0.969 (95%CI:0.962–0.975) for the prediction of EPL.
Methods
The purpose of the EPL predictive model is to build a decision model that automatically predicts the risk of spontaneous abortion from 6 to 10 weeks of ultrasound video. The process of the model construction includes: data collection, pre-processing and model construction.
Data collection and pre-processing
We used data from pregnant subjects in early pregnancy from the Longgang District Maternity & Child Healthcare Hospital of Shenzhen City, and the institutional review board approved the study prior to data collection. All patients provided written consent to store their medical information in the hospital database. In addition, all patient details were de-identified to protect patient privacy. The embryo pregnancy videos in this study were all ultrasound videos of singleton early pregnancy (6–10 weeks) examined using TVS technology. This included 1619 early pregnancy embryo ultrasound videos. Four features can be extracted from the ultrasound video for comprehensive analysis: FHR, GSA, CRL, YSD.
All ultrasound video data were collected by doctors with extensive experience based on a single acquisition process used in routine ultrasound examinations. The procedure is shown below: a trained sonographer performs an ultrasound examination using a GE VOLUSON E8/730 (General Electric Tech Co., Ltd., New York, USA) equipped with a 5–9 MHz transvaginal probe. The operator held the probe with the marker oriented upwards. The probe was moved close to the measured area, with the contact point as the fulcrum, swung at a constant speed to observe different sections. It was first scanned from left to right to visualize a complete longitudinal section of the uterus, then rotated 90 degrees counterclockwise to capture a complete transverse section. The whole process is recorded on video, the length of the video is not less than 15 s. All included ultrasound videos were 1920 × 1080 pixels in size.
All videos were categorized according to pregnancy outcomes recorded by the hospital. Cases that resulted in smooth delivery were classified as negative, while cases of spontaneous abortion were classified as positive. Cases involving miscarriage due to other factors or those without recorded outcomes were excluded from the analysis.
All data were manually annotated by senior doctors who have been in practice for more than 10 years to draw the outline of the corresponding area. The annotations were performed using LabelImage, a commonly used open-source software for image annotation.
Biometric parameters extracting model
Ultrasonic results usually include GSA, CRL, YSD and FHR. As mentioned above, these values are measured from the ultrasonic image. From an image processing perspective, obtaining these measurements can be framed as an image segmentation task, that is, the region of interest, such as GS, YS, etc. can be separated from the ultrasonic image to obtain the corresponding measured values. U-Net samples the semantic information of different layers jointly in the segmentation process, however, continuous down-sampling can cause information loss. Moreover, the biological features in ultrasound video are smaller and the boundary is fuzzier, making accurate segmentation and fitting to real features particularly challenging. Therefore, the deep learning model A3F-net (Attention Feature Fusion Filtering network) was established through modifying and rebuilding U-Net to adapt to the image data of this paper. The architecture of the proposed network is illustrated in Fig. 1.
Structure of A3F-net. St-Measure: Static measurement module, which is responsible for the measurement of static parameters such as GSA, CRL and YSD; Dy-Measure: Dynamic measurement module, which is responsible for the measurement of FHR.
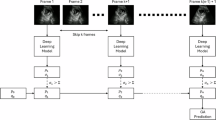
To further improve the segmentation performance for small regions of YS and CR in the video, the attention feature fusion filtering module29 was added between each convolution layer and upsampling layer. The module combines the convolutional feature map with the up-sampled feature map to enhance the resolution of the feature maps and sharpen blurred edges. The static measurement module processes segmented static objects to extract GS, YS, and CR parameters. GS: GSA is calculated via pixel counting; YS: Given its near-circular morphology, YSD is derived by fitting a minimal enclosing circle; CR: CRL is determined using principal component analysis (PCA) to identify the embryo’s longitudinal axis. Each frame of the ultrasound video is segmented and measured, generating a series of values. The final result is defined as the maximum value within the sequence. The Dynamic measurement module focuses on the segmented results of the CR region, where the fetal heart is in this area. According to the characteristics of periodic flickering in the fetal heart region, a sequence of difference maps was obtained by using frame-by-frame differencing. Fourier transform30 is done within the CR target obtained by segmentation, and the number of wave peaks in the waveform maps obtained within the unit between is measured as the FHR. The FHR extraction process consists of the following steps: Firstly, Fetal Heart Localization: The embryonic region is first segmented to approximate the fetal heart position, thereby constraining the search domain for cardiac activity; Secondly, Cardiac Motion Representation: Heartbeat dynamics are captured through temporal variations in the embryonic region’s cross-sectional area; thirdly, Temporal Signal Extraction: Frame differencing is applied to consecutive ultrasound frames to enhance area variations, generating a 3D time-domain matrix with: X-axis: Horizontal spatial dimension; Y-axis: Vertical spatial dimension; Z-axis: Temporal sequence; Next, Frequency-Domain Transformation: The differential sequences undergo Fourier transformation to convert time-domain signals into frequency components, enabling separation of high-frequency signals (fetal cardiac activity) and low-frequency artifacts (background tissue motion); Finally, Signal Refinement & FHR Determination: a low-pass filter suppresses residual noise in the frequency domain, after which the dominant spectral peak is identified as the FHR.
In addition, some image enhancement technologies were used for the ultrasound videos in order to improve the measurement accuracy, include cropping, random scaling, elastic deformation, random rotation and flip up and down, etc29,31. This study uses offline synthetic copies for image enhancement with a multiplication factor of 2.5. In this study, the test set is not involved in data augmentation, and only the training set is augmented. In this study, all experiments were performed on the Ubuntu 20.04 LTS operating system, utilizing four NVIDIA GeForce RTX 3080Ti GPUs.
EPL risk prediction model
Methods in scikit-learn which is a machine learning package for python namely linear classifier, tree model, ensemble learning model, were selected for experimentation, including Logistic Regression, Decision Tree, Random Forest, XGBoost and CatBoost. In this section, five representative machine learning models were selected to complete the prediction task. The selection was based on key criteria such as whether the model is linear and interpretable, and whether it uses ensemble techniques.
Logistic Regression (LR) is a linear classification algorithm with good interpretability and is currently the simplest and most effective linear classifier. It is one of the most commonly used methods to establish a medical decisions model.
Decision Tree (DT) is a tree structure that performs the selection of features at each internal node point and divides the subset in a recursive manner until it stops when the conditions are satisfied. In recent years, decision trees have been increasingly used in medical data processing for their simplicity.
Random Forest (RF) is an integrated learning algorithm that ensembles multiple decision tree models. It’s a bagging model, which constructs different decision trees by randomly selecting a subset of features and samples, and votes or averages the results of all the decision trees, with good robustness, generalization and accuracy.
XGBoost (XB) is an algorithm based on gradient boosting of tree models that can transform weak classifiers into strong ones. Iterative training is performed on the residuals of the previous tree. The log-likelihood function is used as the loss function, its negative gradient is used as an approximation of the residuals, and a regular term is introduced to improve the generalization of the model and to prevent overfitting. XB handles high-dimensional features in a distributed manner with a high degree of efficiency.
CatBoost (CB) is a gradient boosting algorithm, similar to XGBoost, consisting of Categorical and Boosting. Compared to XB, CB is able to deal with categorical features directly, can automatically complete missing values, uses a combination of categorical features, and can take advantage of the links between features, which greatly enriches the feature dimensions, and has high robustness and generalization ability.
Because the number of EPL is smaller than the number of normal pregnancies, the positive-to-negative sample ratio in the dataset is approximately 1:2. The imbalance of data may cause the decision result to be biased towards the majority class. In order to solve this problem, the positive samples were upsampled by synthetic minority over-sampling technique (SMOTE)32 to balance the data. All the feature values obtained from the measurements were normalized with min-max method33which is a normalization operation of the dataset by scaling the data to a value between 0 and 1. The formula for SMOTE is shown in formula (1).
Model evaluation index
The commonly used evaluation indexes of segmentation results are Intersection Over Union (IOU), Dice coefficient (Dice), and Precision. Both IOU and Dice are metrics that evaluate segmentation results, with Dice tending to measure average performance and IOU tending to measure worst performance. The calculation methods are shown in formulas (2)-(4).
The intersection area between automatically segmented region and ground truth is the true positive prediction area (TP), automatically segmented region minus TP is the false positive prediction area (FP), and ground truth minus TP is the false negative prediction area (FN). Both Dice and IOU measure the degree of overlap between ground truth and the machine segment to assess the detection capability of the target area. Precision evaluates whether the segmented object is correct. Each of these metrics has a value between 0 and 1, with larger values indicating better performance.
Precision, sensitivity, specificity, F1-Score and area under the curve (AUC) are used to evaluate the performance of decision model. The calculation methods are shown in formulas (5)-(8).
Precision is the proportion of positive samples that are predicted correctly, out of all positive samples predicted.
Sensitivity is the ratio of positive samples for predicted pairs to all true positive samples, and it is used as a commonly assessed metric in medical research, and it has a greater effect on reducing the rate of missed diagnosis.
Specificity is the ratio of negative samples for predicted pairs to all true negative samples, and it has a greater effect on reducing the rate of misdiagnosis.
The F1-score generally indicates the proportion of positive samples of the prediction pair out of the total positive samples, and is used to evaluate the accuracy of the binary classification model.
.
AUC is defined as the area under the Receiver Operating Characteristic (ROC) curve and the larger the AUC value, the better the performance of the prediction algorithm.
Results
Datasets
The ultrasound videos of 1619 women in early pregnancy were collected. Due to the effects of ultrasound imaging and maternal abdominal fat, this may result in more artifacts and noise in the ultrasound videos. To address the impact of noise, we excluded videos: 306 videos deemed uninterpretable by physicians. Besides that, 215 videos with missing pregnancy outcome data, and 468 videos obtained outside the first trimester.
We ultimately retained a total of 630 ultrasound videos from the first trimester, including 214 positive and 416 negative cases. Among them, 80% proportion of pregnant women including 172 positive and 332 negative cases were randomly selected as the training and validation set for 5-fold cross-validation, and 20% proportion of pregnant women including 42 positive and 84 negative cases were selected as the test set. The specific data cleaning process is shown in Fig. 2. All the videos were annotated by experienced experts, outline the gestational sacs, yolk sacs, and germs.
The patient inclusion workflow.
Model workflow
The workflow of the entire model is shown in Fig. 3. Video set was inputted to the model to train a deep neuro network named A3F-net to get GSA, YSD, CRL and FHR. Among them, the first three are separated from each frame in the video, and FHR needs to be associated with the spatio-temporal characteristics of the video to calculate. Therefore, the results will also be presented in two parts.
The construction process of the Model.
Biological index extraction
Two other networks U-Net and Attention U-Net were chosen for comparison with A3F-net. They were chosen because A3F-net is an improved version of U-Net, integrating not only the attention mechanism, but also the guide filter. So it was compared with the basic U-Net and the attention version of U-Net. The segmentation and comparison results with other networks are shown in Table 1.
For the segmentation of the three targets, A3F-net results were the best. Compared to U-Net, the IOU, Dice and Precision values of the three targets increased by an average of 9.31%, 3.86% and 8.3%, respectively. Especially for CR, the improvement of segmentation results was the most significant, and CR is the smallest and most difficult to segment among the three. Both the performance for GS and YS of Attention U-Net were better comparing to U-Net, but worse for CR region. However, A3F-net consistently demonstrated superior performance.
Figure 4. shows the results of A3F-net compared to other networks and doctor manual segmentation. For the same frame in the video, the results of A3F-net could be largely aligned with the boundaries of the expert’s manual segmentation, even for the small target areas like CR and YS. At the same time, because the A3F-net network traverses all the frames in the ultrasound video, it could find the best measurement frame more accurately than the doctor’s manual measurement, to obtain more accurate measurement values.
The segmentation results of A3F-net compared to other models, as well as the doctor’s manual segmentation.
The measurements results of GSA, YSD, and CRL were obtained by image processing according to the segmentation results. The result of FHR was obtained after processing the segmentation result of CR in continuous frames. Table 2 shows the absolute error and relative error between the result of model and that of doctors.
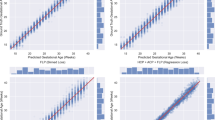
Figure 5. shows four Bland-Altman plots between automated and manual measurements for GSA, YSD, CRL and FHR. In these plots, the horizontal coordinate is the mean of the model measurements and the doctor’s measurements, and the vertical coordinate is the difference between the two. The middle line represents the average of the differences, and the upper and lower lines are the upper and lower limits of the 95% consistency limits. It shows that most of the difference scatter points are within the 95% consistency interval, which indicates that our measurement results have a high consistency with the manual measurement results.
Bland-Altman plot of our model measurements. (a) GSA (b) YSD (c) CRL (d) FHR.
Prediction of EPL
The results of the five decision models are shown in Table 3. The result demonstrated that CB shows the best AUC, and has advantages in various evaluation indicators except sensitivity. The precision and specificity are above 98% and the sensitivity is also in the top two models. For the other 4 machine learning models, both DT and LR belong to a single classifier, the performance of DT is mediocre and the specificity of LR ranked second. RF, XB and CB are all ensemble learning model. Their overall performance is better than that of the single classifier model. XB has the best sensitivity, but the advantage is not obvious compared with CB, and CB has a much higher specificity. Overall, ensemble learning models all exhibit the best performance, with CB performing the best. Figure 6 shows the ROC curves and AUC values of the five machine learning models. CatBoost shows the highest performance, with an AUC of 0.969 (95%CI:0.962–0.975), followed by RF, XB, DT, and LR, with AUC values of 0.967 (95%CI:0.961–0.972), 0.963 (95%CI:0.958–0.968), 0.947 (95%CI:0.941–0.953) and 0.935 (95%CI:0.927–0.943), respectively. Figure 7. shows the confusion matrix demonstrated by all the machine learning models, with darker colors on the main diagonal indicating better classification ability, which shows that ensemble learning models such as CB, RF, and XG exhibit the best performance.
The differences between CB and RF were tested using the Delong algorithm, and the results showed that the differences were not significant. Further tests revealed that there were no significant differences among the ensemble learning models used, including CB, RF, and XG, all of which exhibited excellent AUC values. In contrast, the differences between single classifiers, such as LR and DT, and ensemble learning models were more pronounced. These finding suggest that ensemble learning models offer good applicability and high predictive effectiveness for pregnancy outcome prediction. Among all the ensemble learning models, CatBoost performed the best, achieving an AUC of 0.969.
ROC curve and AUC of 5 machine learning models.
The abscissa represents the prediction results of the model, and the ordinate represents the experimental results. (a) LR, (b) DT, (c) XB, (d) RF, and (e) CB.
Discussion
Principal findings
This study’s two-stage model simulates the process of manual measurement and subjective prediction by clinicians. It not only handles the measurement of small target areas such as CRL and YSD, but also leverages the excellent performance of machine learning, significantly enhancing the efficiency of the embryonic ultrasound video prediction in the clinic.
The clinical interpretation and diagnosis of embryo ultrasound images in early pregnancy are complicated. For the obtained embryonic ultrasound video, doctors need to select the frame with the largest cut-off of each biometric feature in the video for measurement34. And the initial prediction of pregnancy outcome is made based on the measured features. In this process, the pregnancy outcome is greatly influenced by the subjectivity of the doctors, on the one hand, the wrong selection of the largest cut-off and inaccurate measurement of the biometric features, and on the other hand, the prediction of the pregnancy outcome is in error due to the difference in experience of different doctors. Meanwhile the process of manual measurement and prediction is very time consuming. Therefore, it is important to introduce reliable artificial intelligence models into the prediction of embryonic pregnancy outcomes.
Although several studies have applied AI to predict adverse pregnancy outcomes from ultrasound, the specific approach of using ultrasound videos for automatic biometric extraction and subsequent EPL prediction appears to be underexplored. Most existing research focuses on static image segmentation or uses demographic and biochemical markers for prediction. Moreover, current fetal ultrasound studies lack integration of video frame analysis with tailored ensemble learning models for EPL risk assessment. This study addresses these gaps by introducing a novel methodological contribution.
We have noticed that our study shares certain methodological similarities with the work reported in PMID: 36,140,62835; however, there are significant differences in several key aspects. First, the segmentation tasks differ: our study focuses on the segmentation of GS, YS, and CR in dynamic ultrasound videos during early pregnancy (6–10 weeks), whereas the referenced study targets fetal head structures in static images from the mid-to-late pregnancy period, which are easier to identify. Second, we leverage dynamic video data to select frames with the maximum GSA, YSD, and CRL, and extract FHR for early pregnancy outcome prediction—an approach not employed in the referenced work. Third, our focus lies in predicting early pregnancy outcomes, while the referenced study primarily aims at estimating gestational age and fetal weight. Lastly, in terms of prediction methods, we utilize ensemble learning models rather than traditional empirical formulas. Taken together, our work is not an extension of the referenced study but represents a novel exploration toward early pregnancy risk assessment.
Discussion of results
In this study, a deep neural network A3F-net was designed to extract biometric features. In the segmentation task of GS, YS and CR, the IOU, Dice and Precision evaluation of A3F-net model were all optimal compared with the other two models. For the measurement of FHR, the results were also not significantly different from actual clinical counts. The error analysis shows that the absolute error of FHR between the model and the reported data is relatively large, close to 10 bpm. This is attributable to the weak fetal heart fluctuations characteristic of early pregnancy, which complicate accurate measurement. To obtain the FHR, doctors need to measure it by observing the waveform as shown in Fig. 8, first locating the fetal heart, then selecting a stable video segment, and determining the FHR based on changes in brightness. The whole process makes it difficult to determine the start and end points, and the weak brightness changes of the fetal heart cause measurement instability. Additionally, measurement errors exist among different doctors. Additionally, since FHR is typically measured on selected segments rather than across the entire ultrasound video, this contributes to discrepancies between measured and true values. Our automatic measurement model measures based on the entire video, which can eliminate some measurement errors caused by different observers. Furthermore, Fig. 5. shows good consistency between the model’s measurement results and the doctor’s measurements. Therefore, our model can more accurately reflect the true FHR.
Illustration of fetal heart rate measurement performed by a physician.
Then Logistic Regression, Decision Tree, Random Forest, XGBoost and CatBoost machine learning algorithms were used to build five models. The experimental results showed that the established machine learning model, especially the ensemble learning models could predict EPL with high accuracy. Among them, CatBoost performed best, the AUC could achieve 0.969 (95%CI:0.962–0.975).
There are few studies on EPL prediction from early embryo ultrasound video. In fact, due to the very small size of the early embryo, the task of segmentation is more difficult and more sensitive to error, for example, Wang Y et al.36 used CNN to segment the gestational sac region in the early pregnancy embryo, and the Dice obtained was only 0.91. At present, most of the studies on embryo ultrasound video segmentation were based on embryos in the second and third trimesters of pregnancy. Sobhaninia Z37 and K Rasheed38 both segmented ultrasound images of embryos using a deep neural network to get the fetal head circumference. That means both their research worked on middle and late stage embryos not early stage. Anquez J39 studied segmentation of the utero-fetal unit (UFU) from 3DUS volumes, with an average overlap measure of 0.89 when comparing with manual segmentations. But their task was parted the image into two classes of tissues: the amniotic fluid and the fetal tissues. Veronika A. Zimmer et al.40 applied a multi-task learning model to segment the placental region in TVS images, which means their task was also for second and third trimesters, and the final Dice obtained was 0.86.
Most of the current research on AI in pregnancy outcome prediction has been modelled after various biological indicators obtained from more complex physical examinations of pregnant women in the middle or late stages of pregnancy. Even though some studies used some ultrasound results in predicting pregnancy outcomes, for example, GS was used in the prediction model established by Fu K et al.21crown-rump length, yolk sac diameter, and heart rate etc. were used in the research of Yan Ouyang41their values were measured manually. In addition, there are still a few studies in pregnancy outcome prediction has been modelled after various biological indicator in early pregnancy. For example, Wang Y et al. ‘s study used gestational sac for prediction. Among them, the study of Y Ouyang et al.41 and Wang Y et al.36 were similar to this paper. Y Ouyang et al. used logistic regression to establish a prediction model with age, crown-rump length, yolk sac diameter, heart rate etc., but its AUC value was only 0.890. Wang Y et al.36 used CNN to identify the morphologic characteristics of gestational sac in early pregnancy to predict pregnancy outcomes, but the AUC value of them was only 0.885. According to the results of references, Table 4 lists the comparison. The different studies used different features to establish the decision model, among these, the performers of our results are the best, followed by that of Wang Y et al.36 and Y Ouyang et al.41. All these studies used the ultrasound parameters to predict the risk of miscarriage, this further confirms the importance of ultrasonography.
Clinical implications
We have provided for clinicians a whole-process intelligent model that covers measurement and decision making. Since the biometric parameters obtained during ultrasound are very important for predicting EPL, accurate measurements are required, especially in early embryos. Compared with the inefficiency of manual measurement and the high risk of prediction errors stemming from some doctors’ limited clinical experience, an automated, machine-performed measurement method minimizes observer error and enables more accurate decision making.
Limitations and future directions
This study still has some limitations, the source of the dataset we used for training lacks the support of multicentre and multimodal diagnosis data, and we hope to include data from more hospitals for research and validation to improve the model’s generalizability. In addition, the structure of the two-stage model used in this study is relatively complex, and further research on end-to-end models is needed. Despite these limitations, the experiments still demonstrate that the proposed two-stage method is effective and feasible for automatically predicting pregnancy outcomes based on ultrasound videos.
Conclusion
In this paper, a two-stage artificial intelligence model was proposed to enable the prediction of EPL from ultrasound videos of early pregnancy embryos. The model can simulate the process of the clinician’s first measurement and then prediction of the ultrasound image of the embryo in early pregnancy. In the first stage, the precision of GSA, YSD and CRL of A3F-net were 98.64%, 96.94% and 92.83%; the AUC of the decision in the second stage was 0.969 (95%CI:0.962–0.975). It shows great potential for clinical application, improves the consistency of measurement, and provide high-precision decision support for EPL prediction in ultrasound diagnosis.
Data availability
The datasets generated and analysed during the current study are not publicly available due to hospital regulation restrictions and patient privacy concerns, but are available from the corresponding author on reasonable request. The network implementation and prediction code can be accessed at https://github.com/zyaaalex/A3FNet.git.
References
Hristova, M. & Angelova, M. International miscarriage rates: A comprehensive overview of prevalence and causes. J. IMAB. 30, 5703–5709 (2024).
DeVilbiss, E. A. et al. Prediction of pregnancy loss by early first trimester ultrasound characteristics. Am J Obstet Gynecol 223, doi:ARTN 242.e1-e2210.1016/j.ajog.02.025 (2020). (2020).
Pollie, M. P., Romanski, P. A., Bortoletto, P. & Spandorfer, S. D. Combining early pregnancy bleeding with ultrasound measurements to assess spontaneous abortion risk among infertile patients. Am. J. Obstet. Gynecol. 229 https://doi.org/10.1016/j.ajog.2023.07.031 (2023).
Yin, C. H. et al. An accurate segmentation framework for static ultrasound images of the gestational sac. J. Med. Biol. Eng. 42, 49–62. https://doi.org/10.1007/s40846-021-00674-4 (2022).
Doubilet, P. M., Phillips, C. H., Durfee, S. M. & Benson, C. B. First-Trimester prognosis when an early gestational sac is seen on ultrasound imaging: logistic regression prediction model. J. Ultrasound Med. 40, 541–550. https://doi.org/10.1002/jum.15430 (2021).
Detti, L. et al. Early pregnancy ultrasound measurements and prediction of first trimester pregnancy loss: A logistic model. Sci. Rep. 10, 1545. https://doi.org/10.1038/s41598-020-58114-3 (2020).
Fasoulakis, Z. et al. The prognostic role of CRL discordance in first trimester ultrasound. J. Perinat. Med. 52, 294–297. https://doi.org/10.1515/jpm-2023-0132 (2024).
Liu, L. J., Jiao, Y. X., Li, X. H., Ouyang, Y. & Shi, D. N. Machine learning algorithms to predict early pregnancy loss after in vitro fertilization-embryo transfer with fetal heart rate as a strong predictor. Comput Meth Prog Bio 196, doi:ARTN 10562410.1016/j.cmpb.105624 (2020). (2020).
Taylor, T. J., Quinton, A. E., de Vries, B. S. & Hyett, J. A. First-trimester ultrasound features associated with subsequent miscarriage: A prospective study. Aust Nz J. Obstet. Gyn. 59, 641–648. https://doi.org/10.1111/ajo.12944 (2019).
Batmaz, G. et al. The early pregnancy volume measurements in predicting pregnancy outcome. Clin. Exp. Obstet. Gynecol. 43, 241–244 (2016).
Suguna, B. & Sukanya, K. Yolk sac size & shape as predictors of first trimester pregnancy outcome: A prospective observational study. J. Gynecol. Obstet. Hum. Reprod. 48, 159–164 (2019).
Li, S. & Luo, G. Prenatal ultrasonographic diagnosis of fetal abnormalities. Peking: People’s Military Med. Press 81 p (2004).
Park, S. et al. From traditional to cutting-edge: A review of fetal well-being assessment techniques. IEEE Trans. Biomed. Eng. 72, 1542–1552 (2025).
Kang, J., Ullah, Z. & Gwak, J. MRI-Based Brain Tumor Classification Using Ensemble of Deep Features and Machine Learning Classifiers. Sensors-Basel 21, doi:ARTN 222210.3390/s21062222 (2021).
He, S. J. et al. Deep learning radiomics-based preoperative prediction of recurrence in chronic rhinosinusitis. Iscience 26, doi:ARTN 10652710.1016/j.isci.106527 (2023). (2023).
Interlenghi, M. et al. A Radiomic-Based machine learning system to diagnose Age-Related macular degeneration from Ultra-Widefield fundus retinography. Diagnostics 13, ARTN (2023). 296510.3390/diagnostics13182965.
Tian, Y. et al. Multi-scale U-net with edge guidance for multimodal retinal image deformable registration. Annu. Int. Conf. IEEE Eng. Med. Biol. Soc. 2020, 1360–1363. https://doi.org/10.1109/EMBC44109.2020.9175613 (2020).
Zhang, S. Y. et al. CNN-Based Medical Ultrasound Image Quality Assessment. Complexity doi:Artn 993836710.1155/2021/9938367 (2021). (2021).
Petersen, J. F. & Lokkegaard, E. Fresh tools, familiar findings: machine learning in prediction of pregnancy loss. Fertil. Steril. 122, 78–79. https://doi.org/10.1016/j.fertnstert.2024.05.136 (2024).
Shamshuzzoha, M. & Islam, M. M. Early prediction model of macrosomia using machine learning for clinical decision support. Diagnostics 13, ARTN (2023). 275410.3390/diagnostics13172754.
Fu, K. et al. Development of a model predicting the outcome of in vitro fertilization cycles by a robust decision tree method. Front. Endocrinol. (Lausanne). 13, 877518. https://doi.org/10.3389/fendo.2022.877518 (2022).
Kadambi, A. et al. Random forests for accurate prediction of the risk of hypertensive disorders of pregnancy at term. Obstet. Gynecol. 139, 60s–61s (2022).
Wen, J. Y., Liu, C. F., Chung, M. T. & Tsai, Y. C. Artificial intelligence model to predict pregnancy and multiple pregnancy risk following fertilization-embryo transfer (IVF-ET). Taiwan. J. Obstet. Gyne. 61, 837–846. https://doi.org/10.1016/j.tjog.2021.11.038 (2022).
Hancock, J. T. & Khoshgoftaar, T. M. CatBoost for big data: an interdisciplinary review. J Big Data-Ger 7, doi:ARTN 9410.1186/s40537-020-00369-8 (2020).
Sutton, S., Mahmud, M., Singh, R. & Yovera, L. in International Conference on Applied Intelligence and Informatics. 231–247 (Springer).
Looney, P. et al. Fully automated 3-D ultrasound segmentation of the placenta, amniotic fluid, and fetus for early pregnancy assessment. IEEE Trans. Ultrason. Ferroelectr. Freq. Control. 68, 2038–2047 (2021).
Arabi Belaghi, R., Beyene, J. & McDonald, S. D. Prediction of preterm birth in nulliparous women using logistic regression and machine learning. PLoS One. 16, e0252025 (2021).
Amitai, T. et al. Embryo classification beyond pregnancy: early prediction of first trimester miscarriage using machine learning. J. Assist. Reprod. Genet. 40, 309–322 (2023).
Liu, L. J., Tang, D., Li, X. H. & Ouyang, Y. Automatic fetal ultrasound image segmentation of first trimester for measuring biometric parameters based on deep learning. Multimed Tools Appl. 83, 27283–27304. https://doi.org/10.1007/s11042-023-16565-6 (2024).
Maraci, M. A., Bridge, C. P., Napolitano, R., Papageorghiou, A. & Noble, J. A. A framework for analysis of linear ultrasound videos to detect fetal presentation and heartbeat. Med. Image Anal. 37, 22–36. https://doi.org/10.1016/j.media.2017.01.003 (2017).
Muaad, A. Y., Davanagere, H. J., Hussain, J. & Al-antari, M. A. Deep ensemble transfer learning framework for COVID-19 Arabic text identification via deep active learning and text data augmentation. Multimed Tools Appl. https://doi.org/10.1007/s11042-024-18487-3 (2024).
Nedjar, I., Mahmoudi, S. & Chikh, M. A. A topological approach for mammographic density classification using a modified synthetic minority over-sampling technique algorithm. Int. J. Biomed. Eng. Tec. 38, 193–214. https://doi.org/10.1504/Ijbet.2022.120870 (2022).
Ali, P. J. M. Investigating the impact of Min-Max data normalization on the regression performance of K-Nearest neighbor with different similarity measurements. Aro 10, 85–91. https://doi.org/10.14500/aro.10955 (2022).
Liu, R. et al. Multifactor prediction of embryo transfer outcomes based on a machine learning algorithm. Front. Endocrinol. 12 ARTN 74503910.3389/fendo.2021.745039 (2021).
Alzubaidi, M. et al. Ensemble transfer learning for fetal head analysis: from segmentation to gestational age and weight prediction. Diagnostics 12, 2229 (2022).
Wang, Y. et al. Automated prediction of early spontaneous miscarriage based on the analyzing ultrasonographic gestational sac imaging by the convolutional neural network: a case-control and cohort study. Bmc Pregnancy Childb. 22 ARTN 62110.1186/s12884-022-04936-0 (2022).
Sobhaninia, Z. et al. Fetal ultrasound image segmentation for measuring biometric parameters using Multi-Task deep learning. Annu. Int. Conf. IEEE Eng. Med. Biol. Soc. 2019, 6545–6548. https://doi.org/10.1109/EMBC.2019.8856981 (2019).
Rasheed, K., Junejo, F., Malik, A. & Saqib, M. Automated fetal head classification and segmentation using ultrasound video. Ieee Access. 9, 160249–160267. https://doi.org/10.1109/Access.2021.3131518 (2021).
Anquez, J., Angelini, E. D., Grangé, G. & Bloch, I. Automatic segmentation of antenatal 3-D ultrasound images. Ieee T Bio-Med Eng. 60, 1388–1400. https://doi.org/10.1109/Tbme.2012.2237400 (2013).
Zimmer, V. A. et al. Placenta segmentation in ultrasound imaging: Addressing sources of uncertainty and limited field-of-view. Med Image Anal 83, doi:ARTN 10263910.1016/j.media.102639 (2023). (2022).
Ouyang, Y., Peng, Y. Q., Zhang, S. M., Gong, F. & Li, X. H. A simple scoring system for the prediction of early pregnancy loss developed by following 13,977 infertile patients after in vitro fertilization. Eur J Med Res 28, doi:ARTN 23710.1186/s40001-023-01218-z (2023).
Acknowledgements
We gratefully acknowledge the help and support provided by the Longgang District Maternity & Child Healthcare Hospital of Shenzhen City and the Scientific Research Project of Xiang Jiang Lab.
Funding
This work was supported by Chongqing Natural Science Foundation (Grant No. CSTB2024NSCQ-MSX0761), the Scientific Research Project of Xiang Jiang Lab (Grant No. 23XJ02002), the Science and Technology Program of Longgang District, Shenzhen (Grant No. LGKCYLWS2023019) and the Shenzhen Science and Technology Program (Grant No. JCYJ20240813144105007).
Author information
Authors and Affiliations
Contributions
Lijue Liu, Yuan Zang and Huimu Zheng have contributed equally to this work. Teng Pan, Jinhai Deng and Cheng supervised this work. Conceptualization, resources, and project administration were done by Lijue Liu, Teng Pan, Jinhai Deng, Cheng; Methodology, software, formal analysis, investigation, data curation and visualization were done by Yuan Zang, Huimu Zheng, Siya Li, Yu Song, Xue Feng, Xiyuan Zhang, Yaoxu Li, Lulu Cao, Guanglin Zhou, Tingting Dong and Qi Huang. Writing - original draft and writing - review & editing were done by Lijue Liu, Yuan Zang, Teng Pan and Jinhai Deng. All the authors reviewed and edited the manuscript. The authors reviewed and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
This retrospective study was approved by the Institutional Review Board of Longgang District Maternity & Child Healthcare Hospital of Shenzhen City (Approval Number: LGFYKYXMLL-2024-143). Given the retrospective nature of the study and the use of de-identified patient data, the requirement for informed consent was waived by the IRB. The study was conducted in accordance with the ethical standards of the Declaration of Helsinki and its later amendments.
Consent for publication
We have obtained informed consent to publish from the participant to report individual patient data.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Liu, L., Zang, Y., Zheng, H. et al. An AI method to predict pregnancy loss by extracting biological indicators from embryo ultrasound recordings in early pregnancy. Sci Rep 15, 25946 (2025). https://doi.org/10.1038/s41598-025-11518-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-11518-5