Abstract
Antimicrobial resistance in Mycobacterium tuberculosis (M.tb) strains presents a significant challenge to global tuberculosis (TB) control efforts. This study was conducted to explore the distribution and prevalence of mutations at various sites within the 81 bp Rifampicin (RIF) resistance-determining region (RRDR) of the rpoB gene in M.tb, as detected by the Xpert MTB/RIF assay. This retrospective analysis encompassed 9,867 non-repeating patients diagnosed with TB between 2021 and 2023. Cases with RR detected by the Xpert were included in further detailed analysis. The study utilized Chi-square tests or Fisher’s exact tests to identify statistically significant differences in demographic variables and the prevalence of rpoB gene mutations between RResistant TB (RR-TB) and non-RR-TB groups. Multiple logistic regression analysis was employed to examine the relationship between probe types and demographic variables, with a P-value of less than 0.05 considered statistically significant. Over the three-year study period, M. tb was identified in 2,927 cases, with 485 being RR-TB. While individuals aged ≥ 65 years constituted the largest absolute number of RR-TB cases, the highest relative risk was observed in children aged 5–14 years (OR = 2.68, 95% CI 1.16–6.22, P = 0.02) compared to the ≥ 65 reference group. probe E missing emerged as the predominant mutation site, particularly prevalent in pulmonary specimens and among individuals aged 55–64 years, with a statistically significant difference (P < 0.001). An upward trend in probe B mutations was also observed, reaching statistical significance (χ2 = 6.614, P = 0.037).This molecular epidemiological study has identified the mutation patterns within the rpoB gene that contribute to RR, as identified through the use of Xpert technology over a three-year span in Jiangxi Province. The insights gained are instrumental in informing individualized treatment regimens for RR-TB patients by correlating mutation locations with resistance levels (e.g., probe E mutations confer high-level resistance requiring second-line drugs, while probe B mutations like D435Y may confer low-level “disputed” resistance). This facilitates precision therapy, avoids unnecessary second-line treatments, and reduces transmission. Future advancements in technology, such as large-scale sequencing studies, could build upon these findings to further elucidate the genetic variations at play. Ultimately, these discoveries could be corroborated through rigorous in vitro and in vivo experimental research, reinforcing the foundation of our understanding and response to antimicrobial resistance in M.tb.
Similar content being viewed by others
Introduction
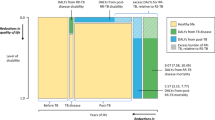
Tuberculosis (TB) is a chronic granulomatous disease that has been a major global health concern for centuries. Ranking among the top ten causes of death globally, TB continues to plague humanity1,2,3. The pathogen responsible for this disease, Mycobacterium tuberculosis (M.tb), was first isolated and identified by the pioneering Dr. Robert Koch in 1882, marking a significant milestone in the understanding of TB4,5. Despite being declared a global emergency by the World Health Organization (WHO) in 1993, TB persists as the foremost bacterial infection in terms of morbidity and mortality worldwide, with a staggering 1.3 million deaths recorded in 2022 6,7,8,9,10. Individuals with latent TB infection (LTBI) are asymptomatic and do not have active disease, with their immune system controlling M.tb replication11,12,13. Lifetime risk of progression to active TB among individuals with LTBI is approximately 5–10%, with the highest risk occurring within the first two years after infection and significantly increased by immunosuppression14. While progress have been made toward reducing the incidence and mortality rates of TB over the decades15,16,17,18the sheer number of individuals infected with M.tb remains substantially high. In 2022 alone, an estimated 7.5 million new cases were diagnosed19,20,21. The standard treatment for drug-susceptible TB is a rigorous six-month regimen involving four first-line antibiotics. Adherence to this protocol is paramount to prevent relapse, eradicate the bacteria, and avert the emergence of drug resistance22,23,24. The Mycobacterium bovis bacille Calmette-Guérin (BCG) vaccine is the only vaccine available against TB. While it can provide protection against severe forms of TB in children, its efficacy in preventing TB infection diminishes post-adolescence25,26,27. Rifampicin(RIF), also known as Rifampin, was discovered in 1965 and belongs to the antimycobacterial class of drugs28,29. RIF exerts bactericidal activity by inhibition of the bacterial DNA-dependent RNA polymerase (RNAP)30,31. The predominant antimycobacterial effect of RIF is to directly block the path of the elongating RNA transcript at the 5′ end beyond a length of 2–3 nt, arresting ongoing RNA synthesis32,33. However, it does not inhibit the mammalian RNAP enzyme34. RIF is a cornerstone of modern short-course, first-line TB treatment regimens30,35. Antimicrobial resistance in M.tb primarily arises from spontaneous mutations in specific genes that confer resistance to commonly used antibiotics, posing a major threat to global TB control36,37,38,39. In 2022, an estimated 410,000 (95% UI: 370 000–450 000) individuals developed RIF resistant TB (RR-TB)40,41. RR-TB patients require extended, costly therapies associated with lower cure rates and higher mortality, and higher rates of post-TB tissue damage42,43,44.
The rpoB gene encodes the RNA polymerase β subunit and is a critical target for detecting RR.The majority of missense mutations that confer resistance to RIF accumulate within the critical 81-base pair (bp) segment of the rpoB gene, known as the Rifampicin resistance determining region (RRDR)45,46,50. The Xpert® MTB/RIF assay stands out as a cutting-edge rapid nucleic acid amplification (NAA) test, adept at swiftly detecting M.tb and discerning RR, attributed to mutations in the rpoB gene, with remarkable accuracy within a mere two hours. Its suitability for point-of-care testing underscores its utility in clinical settings51,52,53. The insights gleaned from Xpert testing facilitates the selection of appropriate treatment regimens and initiating timely therapy, thereby reducing the further spread of M.tb infections54,55.
This study aimed to delve into the distribution and frequency of prevalent mutations at various probe sites within the 81 bp RRDR of M.tb, as elucidated by the Xpert MTB/RIF assay. By mapping these mutations, the study seeks to enhance our understanding of the genetic landscape associated with RR, potentially refining diagnostic and therapeutic strategies in the fight against M.tb.
Materials and methods
Study design, site, and population
This analytical study was carried out at the Jiangxi Province Chest Hospital, the sole provincial tertiary medical institution designated for TB in Jiangxi province, a region with a population of approximately 45.0 million and a high prevalence of M. tb in southern China. Individuals exhibiting symptoms suggestive of TB, identified either passively through referrals or actively via routine physician screenings, from January 2021 to December 2023, were deemed eligible for inclusion in the study. Patient data was retrospectively collected and assessed from archived records of specimens that underwent the Xpert MTB/RIF assay for detection.
Schematic overview showing the study selection process in this study.
Ethical considerations and compliance
All study procedures adhered to the ethical guidelines outlined in the Declaration of Helsinki (2013) and were conducted in compliance with local regulatory standards. Given the retrospective nature of the study, the Institutional Review Board (IRB) of Jiangxi Province Chest Hospital granted an exemption from full ethical review, as the data were anonymized and derived from routine diagnostic procedures.
Specimen collection
All specimens were collected from patients with clinical suspicion of TB and categorized as: (1) Purulent specimens (e.g., sputum, purulent bronchoalveolar lavage fluid, purulent pleural fluid); (2) Non-purulent fluids specimen (e.g., CSF, urine, clear pleural fluid); (3) Solid specimens (e.g., tissue biopsies, lymph node aspirates).
Xpert MTB/RIF assay
The specimens were processed in biosafety cabinet. The purulent specimens were digested with equal volume of 4% NaOH for 10 min, and 1 ml of digested Specimen was mixed with 1:2 ratio using Specimen reagent (SR) of the kit for 10 min at room temperature. The specimen was vortexed for 10 s and left to incubate for an additional 5 min. The solid specimens were ground with equal volume of sterile water, and 1 ml tissue homogenate was mixed at 1:2 ratio using SR for 10 min at room temperature. The specimen was then vortexed for 10 s and left to incubate for an additional 5 min. The non-purulent fluids specimens were centrifuged at 10000 g for 10 min, and the centrifugal sediment was mixed with 2 ml SR for 10 min at room temperature; the specimen was then vortexed for 10 s and left to incubate for an additional 5 min. 2 ml of diluted specimen was transferred into single-use disposable cartridge of Xpert.
Statistical analysis
Data were statistically analyzed using the Statistical Package for Social Sciences (SPSS) version 27.0 (IBM Corp, USA). Categorical data were presented as frequencies and percentages. Chi-square tests or Fisher’s exact tests were performed to identify any statistically significant differences in variables of general characteristics between TB groups with probe hybridization failure and probe hybridization rpoB gene, and determine the trend of rpoB gene hybridization failure rate. The demographic variables (age and group) with respect to probe types were analyzed by multiple logistic regression. P-value less than 0.05 was considered to be statistically significant.
Results
Socio-demographic and clinical information
In the comprehensive analysis of 9,867 nonrepeating patients, the GeneXpert system identified M. tb infections in 2,927 individuals, constituting 29.7% of the sample. Within this infected cohort, 485 cases (16.6%) exhibited resistance to RIF, characterized by the detection of specific gene probe hybridization failure s in the rpoB gene. These mutations manifested either as an absence of wild type probe hybridization or through a significant delta Ct max (> 4.0), indicating a Ct difference exceeding 4.0 between the earliest and latest probe readings. Conversely, for 2,442 cases (83.4%), RR was NOT detected by the Xpert assay. Additionally, 55 cases where the Xpert assay yielded an indeterminate RR result were excluded from subsequent analyses (see Fig. 1). Applying univariate logistic regression, a comparison between the ≥ 65 years’ group and other age brackets revealed significant differences in RR-TB prevalence. Notably, the 55~64 years’ group exhibited a higher odds ratio (OR) of 1.55 (95% CI 1.17–2.07, P = 0.003), indicating a greater propensity for RR-TB compared to the oldest age group. the 25~34 years’ group showed an increased risk with an OR of 1.53 (95% CI 1.12–2.08, P = 0.007). Similarly, the 55~64 years’ group had an even higher OR of 1.85 (95% CI 1.34–2.57, P < 0.001). In contrast, the 15~24 and 35~44 years’ groups did not demonstrate statistically significant differences in RR-TB prevalence. Despite the ≥ 65 years group having the highest case count (n = 113), the 5–14 years group had the highest odds ratio (OR = 2.68, 95% CI 1.16–6.22, P = 0.02), indicating 2.68-fold greater risk than the ≥ 65 reference group.The analysis also confirmed that gender did not correlate with the incidence of RR-TB (Table 1).
Probe hybridization failure in rpoB probes among different age categories
rpoB mutations within the RRDR were identified by detecting hybridization signal loss at five codon-specific probe binding sites, enabling mutation localization. Probes A–E target specific RRDR segments: Probe A (codons 426–430), Probe B (430–437), Probe C (437–442), Probe D (442–448), and Probe E (448–452). Upon stratifying the individual probe level data according to both age category and probe type, a Chi-square statistical analysis revealed a hybridization failure frequency at the probe E (codons 448–452) site that demonstrated a statistically significant difference (χ2 = 22.28, P < 0.001) as depicted in Table 2. Notably, the age group of 65 years and older (n = 72, 22.6%) exhibited the highest frequency of probe hybridization failure, with subsequent frequencies decreasing in the following age groups: 55–64 years (n = 71, 22.3%), 45–54 years (n = 59, 18.5%), 25–34 years (n = 52, 16.3%), and the lowest frequency observed in the 0–4 years’ group (n = 0). In contrast, no significant differences in hybridization failure frequency were discerned across the age categories for probes A-D, with P values of 0.417, 0.539, 0.155, and 0.335, respectively. No significant difference (χ2 = 7.488; P = 0.11) in the Sex-wise distribution of RR-TB in relation to hybridization failure at different probe sites, as presented in Supplementary Table 1.
Hybridization failure in rpoB probes between pulmonary and extrapulmonary TB
This study utilized stratified analysis to assess rpoB hybridization failure s across pulmonary and extrapulmonary TB cases (Supplementary Table 2). Clinical specimens (N = 9,867) comprised three cohorts: 8,135 pulmonary (82.4%), 1,509 extrapulmonary (15.3%), and 223 pulmonary-extrapulmonary co-infections (2.3%). Among extrapulmonary specimens, 325 were MTB-positive, with 295 showing no RR and 30 RR cases. Comparative analysis revealed a disparity in resistance profiles, with pulmonary specimens exhibiting a higher prevalence of RR, confirmed in 445 cases compared to 30 in extrapulmonary cases. Conversely, extrapulmonary cases showed a lower resistance rate, with 9.2% (30 out of 325 cases), compared to 17.6% in pulmonary specimens (445 out of 2,530). The investigation revealed a notably high hybridization failure frequency at probe E among pulmonary specimens, a finding was statistically significant (X2 = 8.28, P = 0.02) when compared among extrapulmonary, pulmonary, and pulmonary-extrapulmonary co-infections TB (Table 3). Conversely, Probe A exhibited the highest rate of hybridization failure (missing data) within the pulmonary-extrapulmonary co-infections group, a difference that was statistically significant (X²=7.561, P = 0.02) when compared to pulmonary or extrapulmonary TB cases independently. In contrast, the variations observed for probes B-D were not statistically significant. Notably, among the 485 RR-TB cases identified by Xpert (RR ‘Detected’), 38 (7.8%) showed no missing probe hybridization. Resistance in these cases was inferred solely based on a maximum ΔCt value > 4.0 cycles, requiring supplementary tests (e.g., sequencing) for validation. Furthermore, the frequency of specimens with no missing probes did not show a statistically significant correlation with age (P = 0.88), sex (P = 0.38), bacillary load (P = 0.44), or the type of clinical specimens collected (P = 0.62), as presented in Supplementary Table 3. This suggests that these factors may not influence the detection rate of probe hybridization failure s in the studied context.
Trend of the rpoB probe hybridization failure rate over three years
Upon conducting a retrospective analysis of observational data spanning three years, it was observed that although the absolute number of M.tb patients and RR-TB cases increased annually, the proportion of bacteriologically-positive individuals classified as RR-TB did not exhibit a statistically significant variation across the years (P = 0.06). Specifically, the proportion stood at 17.5% (128/731) in 2021, 16.9% (131/776) in 2022, and 15.3% (226/1475) in 2023, as detailed in Supplementary Table 4. Further stratification by the male-to-female ratio (M/F ratio) for TB incidence revealed no statistically significant differences among the years (P = 0.55). Regarding the distribution of hybridization failure across different sites, probe E consistently exhibited the highest hybridization failure frequency each year, registering 10.9% in 2021, 11.6% in 2022, and 10.5% in 2023. probe D followed in frequency, while probe C recorded the lowest hybridization failure rate, with 0.5% in 2021, 0.5% in 2022, and 0.2% in 2023. Chi-square statistical analysis was performed to assess the trends in rpoB probe hybridization failure rates. Notably, probe B hybridization failure showed an upward trend over the years, with a significant difference (χ2 = 6.614, P = 0.037), increasing from 1.2% in 2021 to 1.2% in 2022 and then to 2.4% in 2023 (Table 4. Conversely, hybridization failure rates for probes A, C, D, and E did not exhibit significant year-to-year variations.
Discussion
Drug-resistant M.tb has emerged as a formidable global health threat, posing a pressing challenge for developing countries that are grappling with the long-term and high pill-burden regimens required to combat it. China, which is one of the 30 high-TB burden countries in the world56is no exception. In an effort to halt the spread of drug-resistant M.tb, a range of rapid and reliable diagnostic techniques have been employed for the detection of TB and RR-TB, with GeneXpert being one of the most prominent. The GeneXpert system is equipped with five probes that are capable of discerning the difference between the conserved wild-type sequence and the mutations in the rpoB gene region, which are associated with RR. In this study, a total of 2,982 bacteriologically confirmed TB cases were identified using the GeneXpert assay from January 2021 to December 2023. The detection rate of RR-TB was found to be 16.3% (n = 485), which is significantly higher than the reported rate of 5.9% from Zhejiang Province in East China57. A systematic review and meta-analysis conducted in Ethiopia, which reported that male gender is frequently a risk factor for the development of drug-resistant TB in extrapulmonary TB patients58. However, a country-level analysis study using sex-disaggregated data from the World Health Organization (WHO) revealed that the risk of drug resistance among those with TB was the same for males as for females59. Our multiple logistic regression findings are in line with the latter study, indicating that there was no statistically significant difference in the risk of RR between sexes. While individuals aged ≥ 65 years constituted the largest absolute number of RR-TB cases—likely reflecting higher baseline TB incidence in this demographic—multivariable analysis revealed significantly elevated relative risks in younger populations. Compared to the ≥ 65 reference group, the highest adjusted odds ratio was observed in children aged 5–14 years (OR = 2.68, 95% CI 1.16–6.22, P = 0.02), followed by adults aged 25–34 years (OR = 1.85, 95% CI 1.34–2.57, P < 0.001).The pronounced RR-TB risk in children aged 5–14 years warrants particular attention. This aligns with evidence suggesting unique epidemiological drivers of pediatric DR-TB, including delayed diagnosis, suboptimal drug dosing, and immune responses facilitating rapid progression from recent infection to active drug-resistant disease60,61. The results of this study underscore the urgent need for effective strategies to combat the rise of RR-TB, particularly in high-burden countries like China.
In this research, the most frequent mutations within the 81 bp RRDR of the rpoB gene were detected at the probe E site (codons 448–452), followed by the probe D site (codons 442–448), and the least at the probe C site (codons 437–442). These findings align with studies conducted in Ethiopia and Nigeria62,63. However, they diverge from the Indian study that reported the most prevalent mutation at the probe A site (codons 426–430)64. A study from Guizhou province in West China revealed that the probe E site was the most common mutation gene, with the probe D site being the second most common mutation65echoing the results of this study. Nonetheless, in Zhejiang Province, probe D site was the most common, while probe E was less frequent56. The mutation pattern varied significantly within the same country. The diverse RR mutations might be attributed to geographical differences in M.tb lineage, transmission, and resistance acquisition rates66. The most conserved region was identified at the probe C site (codons 437–442), which is consistent with various studies across different regions of the world67,69. The study also identified 7.1% of RR-TB cases with no missing probes, as determined by cycle threshold values > 4 cycles. Similar findings were observed in other studies, where 8.3% and 6.0% of cases exhibited resistance in the rpoB gene without a missing probe type in the research conducted by Munir et al. and Alemu et al.70,71. The presence of all probes positive RR-TB might be triggered by mixed strain infections, polyclonal infections involving concomitant or sequential infection by genetically distinct strains, microevolution, or subpopulations that differ in drug resistance-related variants72. This study contributes to the understanding of mutation patterns in the rpoB gene, which can aid in the development of targeted interventions and improved diagnostics for RR-TB. Critically, while the Xpert MTB/RIF assay detects mutations in the rpoB gene, not all mutations confer clinically relevant rifampicin resistance. For instance, synonymous (“silent”) mutations may not alter the protein structure or drug binding affinity, while certain missense mutations (e.g., D435Y in probe B) confer only low-level resistance (termed “disputed mutations”) that might be overcome with optimized first-line regimens73. Without drug sensitivity testing (DST) or whole-genome sequencing (WGS), all RR-TB cases identified by Xpert are uniformly managed with second-line regimens—prolonged, toxic, and costly therapies that may be unnecessary for patients with non-resistance-conferring or low-resistance mutations. However, integrating DST/WGS with mutation pattern analysis allows clinicians to: (1) distinguish resistance-conferring mutations (e.g., S450L in probe E: high-level resistance)74 from silent/low-resistance variants; (2) stratify patients into tailored treatment pathways (e.g., maintaining first-line therapy for low-resistance cases such as D435Y); and (3) avoid misuse of second-line drugs, mitigating toxicity and delaying further resistance. Thus, elucidating rpoB mutation profiles, including their specific probe locations, refines precision medicine for RR-TB, optimizing outcomes while conserving resources.
The high frequency of mutations observed in the rpoB gene can be attributed to its G + C-rich sequences, which are inherently more prone to mutation75. In the current investigation, the most prevalent mutation associated with RR was identified at the probe E site. Although the Gene Xpert assay did not pinpoint the exact location of the mutation, sequencing studies confirmed that codon 450, a critical site for resistance, was encompassed by probe E76. Additionally, mutations at codons 435 and 445 were found to be associated with probe B and probe D, respectively. Notably, the amino acid alteration at codon 450, where a polar serine is replaced by a non-polar leucine, diminishes the proper binding of RIF to the rpoB gene, thereby conferring resistance77,78. Interestingly, the probe E site exhibited the highest mutation rate among pulmonary specimens when compared to pulmonary-extrapulmonary co-infections specimens, with a statistically significant difference (P < 0.001). In contrast, no instances of probe C missing were observed in extrapulmonary specimens, which aligns with the findings of Sailo et al., who reported a higher mutation frequency for probes C and E in pulmonary specimens64. However, it is worth noting that probe A site presented the most common mutation across patients with pulmonary-extrapulmonary co-infections TB. This observation contrasts with the report by Admassu et al., which indicated a higher prevalence of RR in extrapulmonary TB patients compared to those with pulmonary TB79. In summary, our study observed differential mutation patterns across various probe sites between pulmonary and extrapulmonary specimens. However, these differences may reflect sampling variations rather than inherent biological influences, given the relatively small number of extrapulmonary TB cases (n = 325) compared to pulmonary cases (n = 2,530) in our cohort. The absence of probe C mutations in extrapulmonary specimens and the predominance of probe A mutations in co-infections warrant further investigation with larger sample sizes. These findings highlight the need for comprehensive mutation profiling across diverse clinical specimens to optimize TB management strategies.
The virulence of the infecting strains, host genetic factors, comorbidities, and non-compliance with treatment regimens collectively fuel the outbreak of drug-resistant M.tb36,80. A thorough multiple logistic regression analysis revealed that, across various age groups, the prevalence of mutations at the probe E site was significantly highest (P < 0.001) among the 55–64 years group in this study. This finding indicates a higher incidence of RR-TB attributable to mutations at the probe E site in this age group. It should be noted that the majority of RR-TB cases result from primary infection with resistant strains rather than acquired resistance through spontaneous mutations during treatment. Therefore, the observed association may reflect differential exposure risks or susceptibility to resistant strains in this demographic. The emergence of multidrug-resistant TB (MDR-TB) may involve numerous factors, including detrimental habits such as alcohol abuse and smoking, comorbidities like diabetes mellitus and HIV infection, and exposure to hazardous chemicals and materials such as tobacco81,82. Additionally, health challenges faced by women aged 55–64 years, such as menopause-related conditions, may increase vulnerability to TB progression. The degree of RR associated with rpoB mutations hinges on the site and amino acid alteration. Mutations within the rpoB gene can disrupt the intermolecular interaction between the rpoB protein and RIF, including the loss of hydrogen bonds, hydrophobic interactions, steric hindrance, and electrostatic repulsion, leading to a reduced affinity between the rpoB gene and RIF83. Notably, mutations at position 450 (S450L, S450F, or S450W) conferred high levels of RR, whereas substitution of a valine residue for an aspartic acid at position 435 (D435V) was associated with moderate-level RR83,84,85,86. Conversely, mutations D435Y fell into the low-level or susceptible categories83. One of the limitations of this present study is the inability to detect the exact rpoB gene mutation sites directly through PCR and sequencing. Future research may employ more advanced technologies and large-scale sequencing studies to further identify genetic variations based on these findings.
A significant increase in Probe B failure was observed between 2022 and 2023 (1.2–2.4%, P = 0.04). This temporal trend coincided with the widespread community transmission of SARS-CoV-2 in China following the relaxation of the ‘dynamic zero-COVID’ policy in December 2022 87. However, without pre-2021 data for comparison, we cannot definitively conclude that the observed increase is solely attributable to the COVID-19 pandemic. It is possible that the pandemic conditions, including SARS-CoV-2 coinfection including SARS-CoV-2 coinfection which can induce severe immunosuppression (particularly CD4 + T-cell depletion and granuloma disruption), enabling uncontrolled M. tb replication88may have contributed. The expanded bacterial population size could increase opportunities for spontaneous rpoB mutations during cell division. Concurrently, widespread corticosteroid use (e.g., dexamethasone for COVID-19) might further compromise macrophage function, accelerating mycobacterial proliferation89. Crucially, pandemic-related healthcare disruptions—including TB diagnostic delays, interrupted RIF regimens, and empirical antibiotic misuse—could created selective pressure that favored pre-existing resistant mutants, particularly those at Probe B-associated codons 430–437 (e.g., D435V)90. Moreover, inadequate TB diagnosis, inappropriate treatment, and disregard for TB in COVID-19 patients might have exacerbated the susceptibility to mutations conferring RR91. Mutations in Probe B’s region (codons 430–437, e.g., D435V) typically confer moderate-level RIF resistance64. This level could be particularly favored under the partial treatment pressure and suboptimal TB management prevalent during COVID-19 disruptions (e.g., interrupted regimens, empirical antibiotics), whereas high-level resistance mutations (like Probe E’s S450L) were likely already established and less affected by these transient pressures92. While these factors provide a plausible biological rationale for the observed trend, the lack of pre-pandemic baseline data limits causal inference. Future studies with extended temporal analysis (including pre-2021 data) are needed to validate this association. While not implying direct viral mutagenesis, this synergy between immunosuppression, bacterial burden amplification, and suboptimal TB management likely amplified resistance acquisition. Nonetheless, this study’s findings are confined to the molecular epidemiology of rpoB gene mutation patterns as assessed by Xpert and require further validation through in vitro and in vivo experimental research.
Future studies should integrate parameters such as treatment adherence, healthcare access, socioeconomic factors (e.g., family income, geographic barriers), comorbidities (e.g., diabetes, HIV), and psychosocial support to comprehensively explore drivers of RR.
Conclusions
In this molecular epidemiology study, while individuals ≥ 65 years contributed the largest absolute number of RR-TB cases, the highest relative risk was observed in children aged 5–14 years and adults aged 25–34 years. The study revealed a notable pattern in the mutations associated with RR-TB, with the most common mutation occurring at the probe E site. This was followed by mutations at the probe D site, while the least frequent mutations were observed at the probe C site. Notably, the probe E site exhibited the highest rate of mutation among pulmonary specimens, a finding that was statistically significant (P < 0.001). Further analysis indicated that the prevalence of probe E mutations was significantly highest among the 55–64 years’ group (P < 0.001). A retrospective examination of observational data over a three-year period unveiled an upward trend in the occurrence of probe B mutations, which was statistically significant (χ2 = 6.614, P = 0.037). This trend increased, with a significant increase from 1.2–2.4% in RR-TB cases between 2022 and 2023 within the study area. This surge may be temporally associated to the co-infection with SARS-CoV-2. To deepen our understanding of these genetic variations, future research employing more advanced technologies, such as large-scale sequencing, is recommended. These findings should also be validated through rigorous in vitro and in vivo experimental research. This approach will not only improve understanding of the genetic underpinnings of RR-TB but also potentially guide the development of more effective therapeutic strategies. The location of mutations within the RRDR (e.g., Probe E vs. Probe B) dictates resistance levels and treatment strategies. While Probe E mutations (high-level resistance) necessitate second-line drugs, Probe B mutations (e.g., D435Y) may represent “disputed resistance” where first-line regimens could remain effective. Future diagnostics must integrate mutation location to optimize therapy.”
Data availability
The data supporting this study’s findings are not publicly available due to ethical and privacy considerations, as they contain sensitive information that could compromise the confidentiality of research participants. However, anonymized or aggregated data may be made available upon reasonable request to the corresponding author, subject to compliance with institutional ethics protocols and data protection agreements.
Abbreviations
- BCG:
-
Mycobacterium bovis bacille Calmette-Guérin
- CSF:
-
Cerebrospinal fluid
- LTBI:
-
Latent TB infection
- M. tb :
-
Mycobacterium tuberculosis
- NAA:
-
Nucleic acid amplification
- RIF:
-
Rifampicin
- RNAP:
-
DNA-dependent RNA polymerase
- rpoB gene:
-
Encoding the beta-subunit of RNA polymerase
- RR:
-
Rifampicin resistance
- RRDR:
-
Rifampicin resistance determining region
- RR-TB:
-
Rifampicin-resistant tuberculosis
- SPSS:
-
Statistical package for social sciences
- SR:
-
Specimen reagent
- TB:
-
Tuberculosis
- WHO:
-
World Health Organization
References
Koch, A. & Mizrahi, V. Mycobacterium tuberculosis. Trends Microbiol. 26 (6), 555–556. https://doi.org/10.1016/j.tim.2018.02.012 (2018).
Ali, A. M. et al. Primary gastroduodenal tuberculosis presenting as gastric outlet obstruction: A case report and review of literature. World J. Clin. Cases. 12 (8), 1536–1543. https://doi.org/10.12998/wjcc.v12.i8.1536 (2024).
Alzayer, Z. & Al Nasser, Y. Primary lung tuberculosis. In: StatPearls. Treasure Island (FL): (StatPearls Publishing, 2023).
Negi, K., Bhaskar, A. & Dwivedi, V. P. Progressive Host-Directed strategies to potentiate BCG vaccination against tuberculosis. Front. Immunol. 13, 944183. https://doi.org/10.3389/fimmu.2022.944183 (2022). Published 2022 Jul 28.
Faroug, R. et al. Diagnosis and treatment of tuberculosis of the foot and ankle-A literature review. Foot (Edinb). 37, 105–112. https://doi.org/10.1016/j.foot.2018.07.005 (2018).
GyimahFT & Dako-Gyeke, P. Perspectives on TB patients’ care and support: a qualitative study conducted in Accra metropolis, Ghana. Global Health. 15 (1), 19. https://doi.org/10.1186/s12992-019-0459-9 (2019). Published 2019 Mar 5.
Chaudhry, L. A., Zamzami, M., Aldin, S. & Pazdirek, J. Clinical consequences of non-compliance with directly observed therapy short course (DOTS): story of a recurrent defaulter. Int. J. Mycobacteriol. 1 (2), 99–103. https://doi.org/10.1016/j.ijmyco.2012.05.003 (2012).
Abebe, M. et al. TB case detection: can we remain passive while the process is active? Pan Afr. Med. J. 11, 50 (2012).
Salgueiro, V. C. et al. Extracellular vesicles in mycobacteria: new findings in biogenesis, host-pathogen interactions, and diagnostics. mBio Published Online April. 3 https://doi.org/10.1128/mbio.02552-23 (2024).
da Costa, C. et al. Perspectives on development and advancement of new tuberculosis vaccines. Int. J. Infect. Dis. Published Online Febr. 26 https://doi.org/10.1016/j.ijid.2024.106987 (2024).
Boom, W. H., Schaible, U. E. & Achkar, J. M. The knowns and unknowns of latent Mycobacterium tuberculosis infection. J. Clin. Invest. 131 (3), e136222. https://doi.org/10.1172/JCI136222 (2021).
Lewinsohn, D. M. et al. Official American thoracic society/infectious diseases society of america/centers for disease control and prevention clinical practice guidelines: diagnosis of tuberculosis in adults and children. Clin. Infect. Dis. 64 (2), 111–115. https://doi.org/10.1093/cid/ciw778 (2017).
Yang, Q. et al. The interaction of macrophages and CD8 T cells in Bronchoalveolar lavage fluid is associated with latent tuberculosis infection. Emerg. Microbes Infect. 12 (2), 2239940. https://doi.org/10.1080/22221751.2023.2239940 (2023).
Escalante, P. et al. New diagnostics for the spectrum of asymptomatic TB: from infection to subclinical disease. Int. J. Tuberc Lung Dis. 27 (7), 499–505. https://doi.org/10.5588/ijtld.23.0032 (2023).
Ledesma, J. R. et al. Global-, Regional-, and national-level impacts of the COVID-19 pandemic on tuberculosis diagnoses, 2020–2021. Microorganisms 11 (9), 2191. https://doi.org/10.3390/microorganisms11092191 (2023). Published 2023 Aug 30.
Shaweno, D., Horton, K. C., Hayes, R. J. & Dodd, P. J. Assortative social mixing and sex disparities in tuberculosis burden. Sci. Rep. 11 (1), 7530. https://doi.org/10.1038/s41598-021-86869-w (2021). Published 2021 Apr 6.
Kak, N. et al. Strategic priorities for TB control in Bangladesh, Indonesia, and the Philippines - comparative analysis of national TB prevalence surveys. BMC Public Health. 20(1) 560. Published 2020 Apr 25. (2020). https://doi.org/10.1186/s12889-020-08675-9
Subbaraman, R., Jhaveri, T. & Nathavitharana, R. R. Closing gaps in the tuberculosis care cascade: an action-oriented research agenda. J. Clin. Tuberc Other Mycobact. Dis. 19, 100144. https://doi.org/10.1016/j.jctube.2020.100144 (2020). Published 2020 Jan 11.
Singh, V. Tuberculosis treatment-shortening. Drug discov today. Published Online March. 26 https://doi.org/10.1016/j.drudis.2024.103955 (2024).
Zhu, J., Liu, Y. J. & Fortune, S. M. Spatiotemporal perspectives on tuberculosis chemotherapy. Curr. Opin. Microbiol. 72, 102266. https://doi.org/10.1016/j.mib.2023.102266 (2023).
Moradi, M. et al. Liposomal delivery system/adjuvant for tuberculosis vaccine. Immun. Inflamm. Dis. 11 (6), e867. https://doi.org/10.1002/iid3.867 (2023).
Alsayed, S. S. R., Gunosewoyo, H. & Tuberculosis Pathogenesis, current treatment regimens and new drug targets. Int. J. Mol. Sci. 24 (6), 5202. https://doi.org/10.3390/ijms24065202 (2023). Published 2023 Mar 8.
Fatima, S., Bhaskar, A. & Dwivedi, V. P. Repurposing Immunomodulatory drugs to combat tuberculosis. Front. Immunol. 12, 645485. https://doi.org/10.3389/fimmu.2021.645485 (2021). Published 2021 Apr 13.
Libardo, J., Boshoff, M. D. & Barry, H. I. 3 The present state of the tuberculosis drug development pipeline. Curr. Opin. Pharmacol. 42, 81–94. https://doi.org/10.1016/j.coph.2018.08.001 (2018).
Lange, C. et al. 100 years of Mycobacterium bovis Bacille Calmette-Guérin. Lancet Infect. Dis. 22 (1), e2–e12. https://doi.org/10.1016/S1473-3099(21)00403-5 (2022).
Okafor, C. N., Rewane, A. & Momodu, I. I. Bacillus calmette Guerin. In: StatPearls. Treasure Island (FL): StatPearls Publishing; July 3, (2023).
Zhu, B., Dockrell, H. M., Ottenhoff, T. H. M., Evans, T. G. & Zhang, Y. Tuberculosis vaccines: opportunities and challenges. Respirology 23 (4), 359–368. https://doi.org/10.1111/resp.13245 (2018).
Felker, I. & Trajman, A. Shorter regimens for multidrug-/rifampicin-resistant TB. Int .J. Tuberc. Lung Dis. 27 (1) 3–4. (2023). https://doi.org/10.5588/ijtld.22.0537.
Grobbelaar, M. et al. Evolution of rifampicin treatment for tuberculosis. Infect. Genet. Evol. 74, 103937. https://doi.org/10.1016/j.meegid.2019.103937 (2019).
Eckartt, K. A. et al. Compensatory evolution in NusG improves fitness of drug-resistant M. tuberculosis. Nature 628 (8006), 186–194. https://doi.org/10.1038/s41586-024-07206-5 (2024).
Campbell, E. A. et al. Structural mechanism for rifampicin Inhibition of bacterial Rna polymerase. Cell 104 (6), 901–912. https://doi.org/10.1016/s0092-8674(01)00286-0 (2001).
Walker, S. S. et al. Affinity Selection-Mass spectrometry identifies a novel antibacterial RNA polymerase inhibitor. ACS Chem. Biol. 12 (5), 1346–1352. https://doi.org/10.1021/acschembio.6b01133 (2017).
Lin, W. et al. Structural basis of Mycobacterium tuberculosis transcription and transcription Inhibition. Mol. Cell. 66 (2), 169–179e8. https://doi.org/10.1016/j.molcel.2017.03.001 (2017).
Smith, P. B. et al. Rifampin pharmacokinetics and safety in preterm and term infants. Antimicrob. Agents Chemother. 63 (6), e00284–e00219. https://doi.org/10.1128/AAC.00284-19 (2019).
Rubin, E. J. & Mizrahi, V. Shortening the short course of tuberculosis treatment. N Engl. J. Med. 384 (18), 1764–1765. https://doi.org/10.1056/NEJMe2104499 (2021).
Palomino, J. C. & Martin, A. Drug resistance mechanisms in Mycobacterium tuberculosis. Antibiot. (Basel). 3 (3), 317–340. https://doi.org/10.3390/antibiotics3030317 (2014).
Motavaf, B. et al. Detection of genomic mutations in katG and rpoB genes among multidrug-resistant Mycobacterium tuberculosis isolates from Tehran, Iran. New Microbes New Infect. 41 100879. (2021). https://doi.org/10.1016/j.nmni.2021.100879
Zumla, A. et al. Drug-resistant tuberculosis–current dilemmas, unanswered questions, challenges, and priority needs. J. Infect. Dis. 205 (Suppl 2), S228–S240. https://doi.org/10.1093/infdis/jir858 (2012).
Zhang, F. & Cheng, W. The mechanism of bacterial resistance and potential bacteriostatic strategies. Antibiot. (Basel). 11 (9), 1215. https://doi.org/10.3390/antibiotics11091215 (2022).
Ravikoti, S., Bhatia, V. & Mohanasundari, S. K. Recent advancements in tuberculosis (TB) treatment regimens. J. Family Med. Prim. Care. 14 (2), 521–525. https://doi.org/10.4103/jfmpc.jfmpc_1237_24 (2025).
Anirvan, P. The dire struggle: india’s unfulfilled promise to eliminate tuberculosis. Indian J. Med. Ethics. IX (4), 338–339. https://doi.org/10.20529/IJME.2024.060 (2024).
Malenfant, J. H. & Brewer, T. F. Rifampicin Mono-Resistant tuberculosis-A review of an uncommon but growing challenge for global tuberculosis control. Open. Forum Infect. Dis. 8 (2), ofab018. https://doi.org/10.1093/ofid/ofab018 (2021). Published 2021 Jan 28.
Dheda, K. et al. Multidrug-resistant tuberculosis. Nat Rev Dis Primers. 10 (1) 22. Published 2024 Mar 24. (2024). https://doi.org/10.1038/s41572-024-00504-2
Malefane, L. & Maarman, G. Post-tuberculosis lung disease and inflammatory role players: can we characterise the myriad inflammatory pathways involved to gain a better understanding? Chem. Biol. Interact. 387, 110817. https://doi.org/10.1016/j.cbi.2023.110817 (2024).
Adékambi, T., Drancourt, M. & Raoult, D. The RpoB gene as a tool for clinical microbiologists. Trends Microbiol. 17 (1), 37–45. https://doi.org/10.1016/j.tim.2008.09.008 (2009).
Qi, Y. et al. Utilization of the RpoB gene as a specific chromosomal marker for real-time PCR detection of Bacillus anthracis. Appl. Environ. Microbiol. 67 (8), 3720–3727. https://doi.org/10.1128/AEM.67.8.3720-3727.2001 (2001).
Zaw, M. T., Emran, N. A. & Lin, Z. Mutations inside rifampicin-resistance determining region of RpoB gene associated with rifampicin-resistance in Mycobacterium tuberculosis. J. Infect. Public. Health. 11 (5), 605–610. https://doi.org/10.1016/j.jiph.2018.04.005 (2018).
Bhembe, N. L. & Green, E. Characterization of mutations in the RpoB gene conferring rifampicin resistance in Mycobacterium tuberculosis complex isolated from lymph nodes of slaughtered cattle from South Africa. Braz J. Microbiol. 51 (4), 1919–1927. https://doi.org/10.1007/s42770-020-00356-4 (2020).
Li, M. C. et al. RpoB mutations and effects on Rifampin resistance in Mycobacterium tuberculosis. Infect. Drug Resist. 14, 4119–4128. https://doi.org/10.2147/IDR.S333433 (2021). Published 2021 Oct 5.
Portelli, S. et al. Prediction of rifampicin resistance beyond the RRDR using structure-based machine learning approaches. Sci. Rep. 10 (1) 18120. (2020). https://doi.org/10.1038/s41598-020-74648-y
Chakravorty, S. et al. The New Xpert M.TB/RIF Ultra: Improving detection of mycobacterium tuberculosis and resistance to Rifampin in an assay suitable for [oint-of-Care testing. mBio. 8 (4) e00812-17. (2017). https://doi.org/10.1128/mBio.00812-17
Nandlal, L., Perumal, R. & Naidoo, K. Rapid Molecular Assays for the Diagnosis of Drug-Resistant Tuberculosis [published correction appears in Infect Drug Resist. ;15:6081–6084]. Infect Drug Resist. 2022 15 4971–4984.https://doi.org/10.2147/IDR.S381643
Chen, K. et al. Clinical validation of urine-based Xpert® M.TB/RIF assay for the diagnosis of urogenital tuberculosis: A systematic review and meta-analysis. Int. J. Infect. Dis. 95, 15–21. https://doi.org/10.1016/j.ijid.2020.03.023 (2020).
Lawn, S. D., Nicol, M. P. & Xpert® M.TB/RIF assay: development, evaluation and implementation of a new rapid molecular diagnostic for tuberculosis and rifampicin resistance [published correction appears in Future Microbiol. ;7(8):1024]. Future Microbiol. 2011 6(9) 1067–1082. (2012). https://doi.org/10.2217/fmb.11.84
Ershova, J. V. et al. Impact of GeneXpert M.TB/RIF® on treatment initiation and outcomes of RResistant and RIF-susceptible TB patients in Vladimir TB dispensary, Russia. BMC Infect. Dis. 20 (1), 543. https://doi.org/10.1186/s12879-020-05243-9 (2020).
Yu, S., Ma, J. & Jia, Z. Estimating the incidence of Tuberculosis - Shanghai, china, 2025–2050. China CDC Wkly. 2 (52), 995–998. https://doi.org/10.46234/ccdcw2020.266 (2020).
Ren, Y. et al. Trends of rifampicin resistance in patients with pulmonary tuberculosis: A longitudinal analysis based on drug resistance screening in Eastern China between 2015 and 2019. Infect. Drug Resist. 15, 7707–7717. https://doi.org/10.2147/IDR.S394089 (2022).
Diriba, G. et al. Drug resistance and its risk factors among extrapulmonary tuberculosis in ethiopia: A systematic review and meta-analysis. PLoS One. 16 (10), e0258295. https://doi.org/10.1371/journal.pone.0258295 (2021).
McQuaid, C. F., Horton, K. C., Dean, A. S., Knight, G. M. & White, R. G. The risk of multidrug- or rifampicin-resistance in males versus females with tuberculosis. Eur. Respir J. 56 (3), 2000626. https://doi.org/10.1183/13993003.00626-2020 (2020).
Schaaf, H. S. & Hughes, J. Current treatment of Drug-Resistant tuberculosis in children. Indian J. Pediatr. 91 (8), 806–816. https://doi.org/10.1007/s12098-023-04888-z (2024).
Nolt, D. & Starke, J. R. Tuberculosis infection in children and adolescents: testing and treatment. Pediatrics 148 (6), e2021054663. https://doi.org/10.1542/peds.2021-054663 (2021).
Alemu, A. et al. Does Xpert® MTB/RIF assay give rifampicin resistance results without identified mutation? Review of cases from addis ababa, Ethiopia. BMC Infect. Dis. 20 (1), 87. https://doi.org/10.1186/s12879-020-4817-2 (2020).
Ochang, E. A. et al. Evaluation of rifampicin resistance and 81-bp rifampicin resistant determinant region of RpoB gene mutations of Mycobacterium tuberculosis detected with xpertmtb/rif in cross river state, Nigeria. Int. J. Mycobacteriol. 5 (Suppl 1), S145–S146. https://doi.org/10.1016/j.ijmyco.2016.09.007 (2016).
Sailo, C. V. et al. Distribution and frequency of common mutations in RpoB gene of Mycobacterium tuberculosis detected by Xpert MTB/RIF and identification of residential areas of rifampicin resistant-TB cases: A first retrospective study from mizoram, Northeast India. J. Clin. Tuberc Other Mycobact. Dis. 29, 100342. https://doi.org/10.1016/j.jctube.2022.100342 (2022).
Chen, L. et al. RpoB gene mutation profile in rifampicin-resistant Mycobacterium tuberculosis clinical isolates from guizhou, one of the highest incidence rate regions in China. J. Antimicrob. Chemother. 65 (6), 1299–1301. https://doi.org/10.1093/jac/dkq102 (2010).
Uddin, M. K. M. et al. Distribution and frequency of RpoB mutations detected by Xpert MTB/RIF assay among Beijing and Non-Beijing rifampicin resistant Mycobacterium tuberculosis isolates in Bangladesh. Infect. Drug Resist. 13, 789–797 (2020).
Ektefaie, Y., Dixit, A., Freschi, L. & Farhat, M. R. Globally diverse Mycobacterium tuberculosis resistance acquisition: a retrospective geographical and Temporal analysis of whole genome sequences. Lancet Microbe. 2 (3), e96–e104. https://doi.org/10.1016/s2666-5247(20)30195-6 (2021).
Ullah, I. et al. Rifampicin resistance mutations in the 81 bp RRDR of RpoB gene in Mycobacterium tuberculosis clinical isolates using Xpert MTB/RIF in Khyber pakhtunkhwa, pakistan: a retrospective study. BMC Infect. Dis. 16, 413. https://doi.org/10.1186/s12879-016-1745-2 (2016).
Mboowa, G., Namaganda, C. & Ssengooba, W. Rifampicin resistance mutations in the 81 bp RRDR of RpoB gene in Mycobacterium tuberculosis clinical isolates using Xpert® MTB/RIF in kampala, uganda: a retrospective study. BMC Infect. Dis. 14, 481. https://doi.org/10.1186/1471-2334-14-481 (2014).
Munir, M. K. et al. Comparison of genexpert® probe missing in hotspot RRDR of RPOB gene among primary and acquired drug resistant cases of pulmonary tuberculosis.Biol. Clin. Sci. Res. J., 2022: 165. https://doi.org/10.54112/bcsrj.v2022i1.165] 2., Does Xpert® MTB/RIF assay give rifampicin resistance results without identified mutation? Review of cases from Addis Ababa, Ethiopia. BMC Infect Dis. 2020 20(1):87. 10.1186/s12879-020-4817-2 (2022).
Meehan, C. J. et al. Whole genome sequencing of Mycobacterium tuberculosis: current standards and open issues. Nat. Rev. Microbiol. 17 (9), 533–545. https://doi.org/10.1038/s41579-019-0214-5 (2019).
Sinha, P., Srivastava, G. N., Tripathi, R., Mishra, M. N. & Anupurba, S. Detection of mutations in the rpoB gene of rifampicin-resistant Mycobacterium tuberculosis strains inhibiting wild type probe hybridization in the MTBDR plus assay by DNA sequencing directly from clinical specimens. BMC Microbiol. 20 (1):284. (2020). https://doi.org/10.1186/s12866-020-01967-5
Gopie, F. A. et al. Should treatment of low-level mono-resistant tuberculosis be different? [published correction appears in. J. Clin. Tuberc Other Mycobact. Dis. 24, 100245 (2021). 100245 J. Clin. Tuberc. Other Mycobact. Dis.
Traoré, A. N. et al. Isoniazid and rifampicin Resistance-Conferring mutations in Mycobacterium tuberculosis isolates from South Africa. Pathogens 12 (8), 1015. https://doi.org/10.3390/pathogens12081015 (2023). Published 2023 Aug 4.
Vinogradov, A. E. & Anatskaya, O. V. DNA helix: the importance of being AT-rich. Mamm. Genome. 28 (9–10), 455–464. https://doi.org/10.1007/s00335-017-9713-8 (2017).
Mohajeri, P., Sadri, H., Farahani, A., Norozi, B. & Atashi, S. Frequency of mutations associated with rifampicin resistance in Mycobacterium tuberculosis strains isolated from patients in West of Iran. Microb. Drug Resist. 21 (3), 315–319. https://doi.org/10.1089/mdr.2014.0075 (2015).
Ameke, S. et al. Molecular epidemiology of Mycobacterium tuberculosis complex in the Volta region of Ghana. PLoS One. 16 (3), e0238898. https://doi.org/10.1371/journal.pone.0238898 (2021). Published 2021 Mar 17.
Alamgir, M., Sajjad, M., Baig, M. S. & Noori, M. Y. Mutational frequencies in mycobacterial RpoB gene using genexpert/mtb Rif assay in rifampicin resistant patients at a tertiary care setting in urban sindh, pakistan: analysis from a Five-Year period. Pak J. Med. Sci. 37 (4), 1151–1154. https://doi.org/10.12669/pjms.37.4.3875 (2021).
Admassu, W., Ayelign, B., Abebe, G. & Tadesse, M. Detection of Mycobacterium tuberculosis and rifampicin resistance by Xpert® MTB/RIF assay among presumptive tuberculosis cases at Jimma university medical center, Southwest Ethiopia. PLoS One. 17 (1), e0262929. https://doi.org/10.1371/journal.pone.0262929 (2022). Published 2022 Jan 27.
Khawbung, J. L., Nath, D. & Chakraborty, S. Drug resistant tuberculosis: A review. Comp. Immunol. Microbiol. Infect. Dis. 74, 101574. https://doi.org/10.1016/j.cimid.2020.101574 (2021).
Jones, R. M., Adams, K. N., Eldesouky, H. E. & Sherman, D. R. The evolving biology of Mycobacterium tuberculosis drug resistance. Front Cell Infect Microbiol. ;12:1027394. Published 2022 Oct 5. (2022). https://doi.org/10.3389/fcimb.2022.1027394
Xi, Y., Zhang, W., Qiao, R. J. & Tang, J. Risk factors for multidrug-resistant tuberculosis: A worldwide systematic review and meta-analysis. PLoS One. 17 (6), e0270003. https://doi.org/10.1371/journal.pone.0270003 (2022). Published 2022 Jun 16.
Li, M. C. et al. RpoB mutations and effects on Rifampin resistance in Mycobacterium tuberculosis. Infect. Drug Resist. 14, 4119–4128. https://doi.org/10.2147/IDR.S333433 (2021). Published 2021 Oct 5.
Jamieson, F. B. et al. Profiling of RpoB mutations and mics for Rifampin and Rifabutin in Mycobacterium tuberculosis. J. Clin. Microbiol. 52 (6), 2157–2162. https://doi.org/10.1128/JCM.00691-14 (2014).
Li, M. C. et al. RpoB mutations are associated with variable levels of Rifampin and Rifabutin resistance in Mycobacterium tuberculosis. Infect. Drug Resist. 15, 6853–6861. https://doi.org/10.2147/IDR.S386863 (2022). Published 2022 Nov 28.
Pang, Y. et al. Study of the Rifampin monoresistance mechanism in Mycobacterium tuberculosis. Antimicrob. Agents Chemother. 57 (2), 893–900. https://doi.org/10.1128/AAC.01024-12 (2013).
He, Y. et al. Clinical characteristics of mild patients with breakthrough infection of Omicron variant in China after relaxing the dynamic zero COVID-19 policy. Vaccines (Basel). 11 (5), 968. https://doi.org/10.3390/vaccines11050968 (2023). Published 2023 May 10.
Brunetti, N. S. et al. SARS-CoV-2 uses CD4 to infect T helper lymphocytes. Elife 12, e84790. https://doi.org/10.7554/eLife.84790 (2023). Published 2023 Jul 31.
Koshi, E. J., Young, K., Mostales, J. C., Vo, K. B. & Burgess, L. P. Complications of corticosteroid therapy: A comprehensive literature review. J. Pharm. Technol. 38 (6), 360–367. https://doi.org/10.1177/87551225221116266 (2022).
Silva, B. P. M. D. et al. Drug-Resistant Tuberculosis and COVID-19: A scoping review on a new threat to antimicrobial resistance. Rev. Bras. Enferm. 76 (Suppl 1), e20220803. https://doi.org/10.1590/0034-7167-2022-0803 (2023).
Bostanghadiri, N., Jazi, F. M., Razavi, S., Fattorini, L. & Darban-Sarokhalil, D. Mycobacterium tuberculosis and SARS-CoV-2 coinfections: A review. Front. Microbiol. 12, 747827. https://doi.org/10.3389/fmicb.2021.747827 (2022). Published 2022 Feb 3.
Nusrath Unissa, A., Hassan, S., Indira Kumari, V., Revathy, R. & Hanna, L. E. Insights into RpoB clinical mutants in mediating rifampicin resistance in Mycobacterium tuberculosis. J. Mol. Graph Model. 67, 20–32. https://doi.org/10.1016/j.jmgm.2016.04.005 (2016).
Acknowledgements
The authors would like to acknowledge all the participants of this study and the staffs from Jiangxi Province Chest Hospital.
Author information
Authors and Affiliations
Contributions
Conceptualization, Zhan Qiu Mao and Hui Qiong Yang; Data curation, Zhan Qiu Mao; Formal analysis, Zhan Qiu Mao and Huilie Zheng; Investigation, Zhan Qiu Mao, Zhen Qiong Liu,Fei Mei Li, Ling Shan Xu, Li Chao Liang, Ling Hua Shu; Methodology, Zhan Qiu Mao, Qi Long Zhang and Hui Qiong Yang; Resources, Zhan Qiu Mao; Software, Zhan Qiu Mao; Supervision, Hui Qiong Yang; Validation, Zhan Qiu Mao; Visualization, Zhan Qiu Mao; Writing – original draft, Zhan Qiu Mao;project administration, Zhan Qiu Mao; funding acquisition, Zhan Qiu Mao. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Institutional review board statement
Ethical approval was waived by the local Ethics Committee of Jiangxi Province Chest Hospital in view of the retrospective nature of the study and all the procedures had been performed were part of the routine care.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Mao, Z.Q., Zhang, Q.L., Zheng, H. et al. RpoB mutation patterns in Rifampicin-resistant tuberculosis: a Jiangxi Province study, 2021–2023. Sci Rep 15, 27988 (2025). https://doi.org/10.1038/s41598-025-11949-0
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-11949-0