Abstract
Breast cancer is a leading cause of cancer mortality in women with triple-negative breast cancer (TNBC) presenting greater treatment challenges due to its aggressive disease progression. Understanding TNBC’s unique cell signaling and gene expression profiles will reveal novel therapeutic strategies. Non-coding RNAs, including microRNAs (miRNAs) and long non-coding RNAs (lncRNAs), have emerged as key regulators of gene expression and potential therapeutic targets. This study focuses on a TNBC-enriched lncRNA, non-coding RNA in the aldehyde dehydrogenase 1A pathway (NRAD1, previously LINC00284), which promotes progression in multiple cancers. Our analysis reveals that NRAD1 is central to miRNA–mRNA networks in TNBC cells, mediating cancer-promoting gene expression changes. Fractionation studies showed that NRAD1 is primarily located in the nucleus and mitochondria, with some cytoplasmic presence allowing for transcript-specific competitive endogenous RNA (ceRNA) interactions with miRNAs. However, NRAD1 primarily effects miRNAs independently of ceRNA activity, instead upregulating DICER (a miRNA biogenesis protein), altering sub-cellular distribution, and reducing biogenesis of mitochondria-localized miRNA (i.e., miR-4485-3p). These findings demonstrate novel regulatory interactions between the cancer-promoting lncRNA NRAD1 and miRNAs that alter gene expression in TNBC, expanding our understanding of regulatory lncRNA–miRNA effects, TNBC biology, and highlighting future therapeutic strategies for targeting non-coding RNAs.
Similar content being viewed by others
Introduction
Breast cancer is the most diagnosed cancer in women and accounts for over 600,000 deaths globally per year1. Treatment regimens are largely based on the expression of three receptors, which define the three clinical breast cancer subtypes2. Hormone receptor positive (HR+) breast cancers express estrogen receptors (ER) and progesterone receptors (PR), and account for 70–75% of cases, and are treated with endocrine therapies, combined with chemotherapy in high-risk cases. Overexpression of the human epidermal growth factor receptor 2 (HER2) is present in 15–25% of breast cancers. Patients with HER2+ breast cancers are treated with HER2-targeting drugs in combination with chemotherapy. Triple-negative breast cancer (TNBC) lacks the three receptors, which prevents their treatment with these receptor-targeting drugs and makes chemotherapy the primary treatment.
In addition to having different treatment regimens, the three major clinical breast cancer subtypes have distinct disease progression profiles, with TNBC being the most aggressive and likely to recur3. The subtypes present with distinct gene expression and cell signaling pathways4,5,6 Recent advancements in treatments for TNBCs by targeting genetic factors and molecular pathways associated with TNBC are improving outcomes (e.g., pembrolizumab and olaparib)7. This illustrates how understanding the unique cell signaling and gene expression profiles associated with TNBC can be harnessed into new therapeutic interventions that improve patient outcomes.
Key regulators of gene expression and novel proposed targets are the non-coding RNAs, microRNAs (miRNAs) and long non-coding RNAs (lncRNAs)8. According to Genecode’s latest release, there are 1879 miRNAs and 34,914 lncRNAs in the human genome and many have now been implicated in cancer progression and treatment responses8,9. MiRNAs are small non-coding RNAs (21–22 bases) that regulate gene expression through their incorporation into the RNA-induced silencing complex (RISC)10. They bind complementary messenger RNAs (mRNAs) causing repression of translation and mRNA degradation. MiRNAs can act in a subtype-specific manner in breast cancer, resulting in distinct gene expression effects in HR+, HER2+, and TNBC subtypes11.
LncRNAs also display breast cancer subtype-specific expression12, and this has been especially studied in the context of TNBC. Expression of 300 lncRNAs is elevated in TNBC13. Functional characterization of some of these TNBC-specific lncRNAs has revealed their roles in various cellular processes. For instance, LINP1 is involved in DNA repair regulation13, while another TNBC-enriched lncRNA, non-coding RNA in aldehyde dehydrogenase 1A pathway (NRAD1), previously called LINC00284, promotes breast cancer stem cell properties14. The breast cancer subtype-specific lncRNA expression profiles suggest their potential involvement in the progression of different breast cancer subtypes and highlight their promise as therapeutic targets12. This is especially relevant given that there are increasing examples of lncRNAs affecting cancer-promoting gene expression through multiple mechanism, including through interactions with chromatin and miRNAs15. Review of the literature suggests that competitive endogenous RNA (ceRNA) interactions with miRNA (commonly referred to as “sponging”) is the most often-reported mechanism of lncRNA gene regulation8. The 2019 release of DIANA-LncBase catalogued 240,000 unique tissue and cell-type specific miRNA–lncRNA interactions, including 730 miRNA–lncRNA entries from manual curation of publications16.
The subcellular localization of lncRNAs is a determinant factor of their function and impacts potential regulation of miRNAs17,18,19. Cytoplasmic lncRNAs are often reported to be involved in post-transcriptional regulation through miRNA sponging. Some lncRNAs exhibit dynamic localization, shuttling between the nucleus and cytoplasm, which allows them to perform diverse functions. A smaller subset of lncRNAs are mitochondrial non-coding RNAs (mtncRNA). These are typically transcribed from mitochondrial DNA (mtDNA) and play roles in mitochondrial gene expression and function. Recent studies have also identified nuclear-encoded lncRNAs that localize to mitochondria, suggesting a complex interplay between nuclear and mitochondrial gene regulation15,20.
Some lncRNAs are reported to have both chromatin and miRNA interactions, such NRAD1/LINC00284. For example, in breast cancer, NRAD1-mediated gene expression changes were partly explained through chromatin interactions14. However, in other cancers, NRAD1/LINC00284-mediated gene expression changes have been explained through miRNA ‘sponging’ interactions21,22,23,24,25,26. For example, in colorectal cancer, LINC00284 sponged miR-361-5p which led to the increase in transforming growth factor beta (TGF-β) signaling and promotion of colorectal cancer24.
Here in we performed a comprehensive analysis of NRAD1 gene expression regulation in TNBC, focusing on characterizing miRNA effects. We find that NRAD1 is at the hub of miRNA–mRNA networks in TNBC cells and mediates the cancer-promoting gene expression changes through altering miRNA levels. We find evidence of ceRNA regulation of some miRNAs that is concentration dependent and NRAD1-transcript-specific. However, the effects of NRAD1 on miRNAs are predominately independent of ceRNA effects. NRAD1 upregulates key miRNA biogenesis protein DICER, alters the sub-cellular distribution of some miRNAs, and affects the biogenesis of mitochondria-localized miRNA miR-4485-3p. Together, this study demonstrates unique modes of interaction between the cancer-promoting lncRNA NRAD1 and miRNAs ultimately leading to changes in the gene expression landscape associated with the aggressive TNBC subtype.
Materials and methods
Cell culture
MDA-MB-468 cells were obtained from American Type Culture Collection (ATCC, Manassas, VA, USA) and SUM149 cells which were obtained from BioIVT (previously Asterand, Westbury, NY, USA). MDA-MB-468 cells were cultured in Dulbecco’s Modified Eagle Medium (DMEM; Life Technologies, ThermoFisher Scientific, Burlington, Canada) supplemented with 10% fetal bovine serum (FBS, Life Technologies, ThermoFisher), with the inclusion of antibiotic antimycotic (AA; Life Technologies, ThermoFisher Scientific). SUM149 cells were cultured in Ham’s F-12 Nutrient Mix (F-12; Life Technologies, Life Technologies, ThermoFisher Scientific) supplemented with 5% FBS, AA, HEPES buffer, 1 μg/mL hydrocortisone (Millipore Sigma, Mississauga, Canada), and 5 μg/mL human insulin (Millipore Sigma).
NRAD1 knockdown
Transient in vitro knockdown of NRAD1 in cancer cells was achieved through previously described protocol14. Briefly, NRAD1 knockdown was achieved by mixing antisense oligonucleotides with that consist of a central DNA “gap” flanked by LNA (Locked Nucleic Acid) regions (LNA GampeRs)27. NRAD1-specific GampeR’s (GapmeR #3 or #4) or negative control GapmeRs (Supplementary Table S1) were purchased from Qiagen (Qiagen, formerly Exiqon, Toronto, Canada) with OptiMEM reduced serum media (Life Technologies, ThermoFisher) and TransIT-BRCA transfection reagent (MJS Biolynk, Brockville, Canada) to a final concentration of 15 nM. Knockdown was confirmed 48 h post transfection with quantitative polymerase chain reaction (qPCR) as described below.
Cell treatments with miRNA mimic
SUM149 cells were seeded in 6-well plates at 20% confluency and after 18h post seeding, the cells were transfected with miRCURY LNA mimic or negative controls (Qiagen). The mimics were mixed with OptiMEM reduced serum media (Life Technologies, ThermoFisher) and TransIT-BRCA transfection reagent (MJS Biolynk) and added to sub-confluent cells to a final treatment concentration of 25 nM as per the manufacturer’s instructions. Treatment efficacy was confirmed by TaqMan assay at 48 h post transfection as described below.
Quantitative polymerase chain reaction
For all transcript expression analyses by qPCR, cells were collected in TRIzol (Life Technologies, ThermoFisher Scientific) and total RNA was purified using a PureLink RNA kit (Life Technologies, ThermoFisher Scientific) as per the manufacturer’s instructions. Equal amounts of harvested total RNA were reverse transcribed with iScript cDNA Synthesis Kit (Bio-Rad, Mississauga, Canada) as per the manufacturer’s instructions. QPCR was performed using SsoAdvanced Universal SYBR Super-mix (Bio-Rad) and transcript-specific primers (primer sequences are listed in Supplementary Table S2) as per the manufacturer’s recommended protocol using a CFX96 or CFX384 Touch RealTime PCR Detection System (Bio-Rad). Primer efficiencies, determined by standard curves of diluted cDNA samples, were incorporated into the CFX Manager software (Bio-Rad). Gene expression for all samples was calculated relative to two or three reference genes and relative to the negative control or NRAD1-targeting GapmeR-treated samples.
Gene arrays
SUM149 cells were treated with NRAD1-targeting antisense oligonucleotides GapmeRs #3 or #4, or negative control GapmeR for 48 h and then collected in TRIzol reagent (n = 3). RNA purification was performed as described above and sent to the Centre for Applied Genomics (TCAG, The Hospital for Sick Kids, Toronto, Canada) for Affymetrix Human Gene 2.0 ST gene chip platform analysis and raw data files are available on the Gene Expression Omnibus (GEO), GSE294997. The data was processed using limma R package v3.54.2 to reveal differential gene expression. MDA-MB-468 cells had been similarly treated with anti-NRAD1 GapmeR #4 or negative control GapmeR and extracted RNA processed for Affymetrix Human Gene 2.0 ST gene chip platform analysis was accesses from GEO (GSE118710)14. The fold-change cutoff of greater than or equal to 1.4 or less than or equal to - 1.4 and had a p-value below 0.05 were used in further analyses (Supplementary File 1).
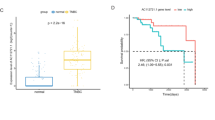
Analysis of patient tumor gene expression data
The RNA sequencing (RNA‐seq) log2 V2 RSEM expression data for the Breast Invasive Carcinoma (Cell 2015) from The Cancer Geneome Atlas (TCGA were accessed via cBioportal and patients were specifically identified from the clinical data28,29. Gene array expression data from the Breast Cancer Molecular Taxonomy of Breast Cancer International Consortium (METABRIC) dataset was also accessed via cBioportal and TNBC patients specifically identified30.
MicroRNA arrays
SUM149 and MDA-MB-468 cells were with NRAD1-targeting GapmeRs #3 or #4 or negative control GapmeR for 48 h and then collected using the mirVana™ miRNA Isolation Kit (Life Technologies, ThermoFisher Scientific) as per manufactures instructions for small miRNAs. Total RNA from cell lysates was used to confirm knockdown of NRAD1 using qPCR as described. Small miRNA purification was sent to the Centre for Applied Genomics (TCAG, The Hospital for Sick Kids, Toronto, Canada) for Affymetrix GeneChip miRNA 4.0 array. Data was collected in raw CEL format and differential miRNA expression was visualized using the Affymetrix Transcriptome Analysis Console (TAC). MiRNA names adjusted to the most recent version of miRbase using the miRbaseConverter in R to version V22. Mature miRNAs were identified and those that met the fold-change cut-off of ± 1.4 along with having a p-value less than or equal to 0.05 were used in further analysis (Supplementary File 1). The raw data is deposited on GEO (GSE295075).
MicroRNA quantification by TaqMan PCR assays
MiRNA (small and total) was purified from cells using the mirVanna miRNA isolation kit (Life Technoloiges, ThermoFisher Scientific) as per manufactures instructions for small miRNAs. Total miRNA was isolated with modification utilizing miRNA columns and lysis extraction from mirVanna miRNA isolation and wash buffers from PureLink RNA kit (Life Technologies, ThermoFisher Scientific). cDNA was synthesized from the purified small miRNA using the pre-formulated reverse transcription primers and the reagents in the TaqMan microRNA reverse transcription kit (Life Technologies, ThermoFisher). TaqMan Fast Advanced Master Mix (ThermoFisher) was used for the qPCR. MiRNA levels detected using either the TaqMan miRNA assays (miR-4485, miR-484, miR-192-5p, miR-3929, miR-28-3p, miR-769-5p, miR-1303, and miR-452, cat#4427975) were calculated relative to 2 to 3 reference miRNAs (RNU48, miR-25-3p, and miR-93-5p) and relative to the negative control.
The miEAA V 2.0 platform was used to perform an over-representation analysis of miRNAs involved in biological processes31. Each cell line was performed separately only utilizing significantly differentiated miRNAs for GapmeR #4 versus negative control GapmeR treatment. The mRNA binding of differentially expressed miRNAs was predicted using MultiMir32. The resulting NRAD1/miRNA/mRNA networks were plotted using Cytoscape V 3.2 software33.
Cloning NRAD1-201 and NRAD1-202 transcripts from MDA-MB-468 cells
Standard techniques were used to clone the NRAD1-201 and 202 transcripts from MDA-MB-468 into the pcDNA3 plasmid. First-strand cDNA was synthesized from MDA-MB-468 cell RNA, using SuperScript III, and oligoDT12-18 (Life Technologies, Thermo Fisher Scientific), as per manufacturer’s instructions. The NRAD1-201 and 202 sequence were retrieved and amplified using specific primers (Supplementary Table S3) with flanking 5′-HindIII (forward) and 3′-BamHI (reverse) sites, in the presence of Platinum SuperFi enzyme (Life Technologies, Thermo Fisher Scientific). The amplified NRAD1 cDNA with the primer-introduced restriction sites were digested with BamHI and HindIII and ligated into double-digested (BamHI and HindIII) pcDNA3 plasmid using T4 ligase (New England Bioscience, Whitby, Ontario, Canada). Competent TOP10 E. coli cells were transformed and incubated overnight at 37 °C. The plasmids were purified with a plasmid purification kit, and the resulting plasmids. The clones were verified to have the correct insert by sequencing (Plasmidsaurus, Supplementary File 2).
Predicted miRNA binding to NRAD1 transcripts
To identify miRNAs predicted to bind NRAD1 transcripts, we utilized the LncBook 2.0 database, which integrates comprehensive lncRNA–miRNA interaction predictions34. We queried the database for all available NRAD1 transcripts that had transcript level support in Ensembl (Release 113). The LncBook tool employs three computational prediction tools (miRanda, TargetScan, and RNAhybrid) and we focused on interactions supported by at least two tools35,36,37. By filtering for high-confidence predictions, we generated a list of miRNAs with potential binding sites on the target NRAD1 lncRNA transcripts (Supplementary File 3).
Western blotting
Collected cells were lysed in RIPA buffer and sonicated for 5 min (15S ON, 45S OFF, 40% amplitude, 4 °C) and quantified with Pierce BCA Protein Assay Kit (ThermoFisher scientific). Laemmli buffer was added to the lysates and the samples were boiled 10 min and 40 μg of MDA-MB-468 lysate samples and 20 μg of SUM149 lysate samples was run per lane in Mini-PROTEAN TGX Stain-Free Precast Gel (Bio-Rad) and run for 1 h at 100 V in Tris–Glycine-SDS buffer. The lysates were transferred onto PVDF membranes in transfer system (Bio-Rad), in transfer buffer (3 g TRIS, 144 g Glycine, 200 μL MetOH, 800 μL H2O), for 19 h 30 V at 4 °C. The membranes were blocked in 5% skim milk in TBST for 8 h at 4 °C. The membranes were incubated with 1:1000 anti-DICER (Dicer (D38E7) Rabit mAb #53625, Cell Signaling Technology, Danvers, Massachusetts, USA) diluted in 5% bovine serum albumin, overnight at 4 °C followed by peroxidase conjugated affiniPure goat anti-rabbit IgG (H + L, #111-035-144, Jackson Immunoresearch, West Grove, PA, USA) antibody (1:1000 in 5% milk TBST) for 1 h at room temperature. The chemiluminescence was imaged with the ChemiDoc imaging system (Bio-Rad) and the band intensities were calculated and plotted using Imagelab software (Bio-Rad) versus total protein.
Luciferase assay
Oligos specific to the wildtype (WT) NRAD1-miR-4485-3p predicted binding region and the mutated version of the sequence (MUT) are listed in Supplementary Table S4. To make double stranded sequences for cloning, the oligos were admixed into oligo annealing buffer and heated to 90 °C for 3 min, followed by cooling to 37 °C for 15 min. The WT and MUT annealed oligos (Life Technologies, ThermoFisher Scientific) were cloned into the multiple cloning sites of the pmirGLO Dual-Luciferase miRNA Target Expression Vector (ThermoFisher Scientific, using SacI and XhoI restriction enzymes (New England Biolabs, Ipswich, MA, USA). The confirmed vectors were co-transfected into MDA-MB-468 cells with the pRLTK vector (Promega, ThermoFisher Scientific), using TransIT-BRCA transfection reagent. 24 h later the mirVana miRNA negative control mimic or mimic-hsa-miR-4485-3p (Life Technologies ThermoFisher Scientific) was transfected into the cells using TransIT-BRCA. The resulting firefly and renilla luciferase activity in the cells were measured 24 h later using the Dual-Glo® Luciferase Assay System with a SpectraMax® M3 Multi-Mode Microplate Reader (Molecular Probes, San Jose CA, USA). Binding of the mimic sequence to the luciferase reporter vector would inhibit production of luminescence.
Nucleus, cytoplasm and mitochondria fractionation
Nuclear and cytoplasmic fractionation followed by RNA extraction was performed using the PARIS™ kit (Life Technologies, ThermoFisher Scientific) as per manufactures instructions. Confirmation of fractionation was determined through the comparison of qPCR levels of cytoplasmic lncRNA DANCR or for miRNAs miR-25-3p and nuclear lncRNA NEAT1 or for miRNAs RNU48 (Supplementary Table S2, for NEAT1 and DANCR). Mitochondria separation from nuclear-cleared cytoplasm was performed using the Mitochondria Isolation Kit for Cultured Cells (Life Technologies ThermoFisher Scientific) following the using the reagent-based methods with modifications. Briefly, cells were harvested by trypsin and pelleted by centrifugation at 500×g for 5 min. Cells were counted and resuspended in PBS and centrifuged in a 2.0 mL RNA/DNA free microfuge tube at 850×g for 2 min to remove nuclei. Supernatants were removed and cell pellets were resuspended in 800 µL Mitochondrial Isolation Reagent A with RNA inhibitor (SUPERase·In™ RNase Inhibitor (20 U/μL), cat# AM2696, Life Technologies ThermoFisher Scientific) and using a 25G × 1 ½” TW (0.5 mm × 40 mm, BD PrecisionGlide Needle) five times aspirations followed by five times with a 30G × ½” (0.3 mm × 13 mm, BD PrecisionGlide Needle). The remainder of the procedure was followed as suggested by manufacture, all buffers were supplemented with RNA inhibitor and mitochondria and cytoplasm fractions were isolated for RNA analysis. QPCR confirmed separation of mitochondria from cytoplasm using primers against mitochondria-specific COX-2 and MT-12S compared to cytoplasmic-localize DANCR (Supplementary Table S2). For miRNA analysis, the mirVanna miRNA isolation kit (Life Technologies, ThermoFisher Scientific) was applied to fractions, following the manufacturer’s instructions for small miRNA isolation.
ASncmtRNA2
Antisense non-coding mitochondrial RNA 2 (ASncmtRNA2) was quantified adapting a protocol by Fitzpatrick et al., with modifications38. Total RNA was isolated from TRIzol as described above. Briefly, 100 ng of total RNA was mixed with 1 μL of 50 μM Random Hexamers (Life Technologies ThermoFisher Scientific) and 1 μL of 10 mM dNTP (Applied Biosystems, ThermoFisher Scientific) to make a total volume of 14 μL contents were mixed and heated for 66 °C for 5 min, placed on ice for 1 min. This was then combined with 1 μL of 100 mM DTT (Life Technologies, ThermoFisher Scientific), 1 μL of Super Script IV Reverse Transcriptase (200 U/μL) (Life Technologies, ThermoFisher Scientific) and 4 μL 5 × Superscript IV RT buffer, mixed and heated at 23 °C for 10 min, 55 °C for 10 min, 80 oC for 10 min. The resulting cDNA was diluted 1 in 10 and PCR was performed using 4 μL of Diluted cDNA, 5 μL SYBR green (Bio-Rad) and 1 μL of primer mix 4 μM ASncmtRNA-2 forward and reverse, primers are listed in Supplemental Table S2. QPCR setting 95 °C for 30 s, 95 °C for 10 s, 60 °C for 30 s × 45 cycles. QPCR was compared to reference primes (18S rRNA and 16 s rRNA, Supplementary Table S2). Confirmation of product was done using 2% agarose DNA gel, 20 μL of qPCR product and 4 μL of TrackIt cyan/orange loading buffer (catalog #10482028, Life Technologies, ThermoFisher Scientific), TrackIt ladder (catalog# 10488085, Life Technologies, ThermoFisher Scientific). QPCR products were purified and sequenced by Sanger Sequencing to verify the amplification of ASncmtRNA2 (Supplementary File S4).
Statistics
All data points represent biological replicates, not technical replicates. For qPCR assays, multiple technical replicates were averaged into a single biological replicate data point. All statistical analyses were performed in the GraphPad Prism software (GraphPad Software, San Diego, CA, USA). In all cases where three or more groups are compared, a one-way or two-way ANOVA was performed (with Dunnett’s or Tukey’s multiple comparisons post-test as indicated in the figure legend). Comparisons between two groups were done using a two-tailed student’s t-test. For co-expression analyses, p values were determined by the cor.test() function with the method argument set to “spearman” in Rv4.2. LncBook data analysis was performed in R studio, Wilcoxon test, Cliff’s delta and adjusted p-value by false discovery rate was performed on each group. Significant p values are indicated as follows in the figures: p < 0.05 = *, p < 0.01 = **, p < 0.001 = ***, p < 0.0001 = ****.
Results
NRAD1 alters the TNBC transcriptome
We previously demonstrated that NRAD1 is highly expressed in TNBC compared to other breast cancer subtypes14. Its knockdown using GapmeR #3 and GapmeR #4 in TNBC SUM149 and MDA-MB-468 cells, reduces cell proliferation, survival and tumor growth, with more pronounced effects with GapmeR #414. We therefore used these cell lines as models to further investigate the effects of NRAD1 on gene expression in TNBC. Negative control or antisense oligonucleotide LNA GapmeRs targeting NRAD1 (GapmeR #3 or #4) were applied to SUM149 or MDA-MB-468 cells and effects on genome wide gene expression were assessed by Affymetrix human gene 2.0ST gene array analysis, which measures expression in 24,838 genes (Supplementary File 1). This analysis revealed significant alterations in gene expression in both cell lines, with over 600 genes upregulated and 900 genes downregulated upon NRAD1 knockdown in both cell lines (Fig. 1A, GapmeR #4 compared to negative control; Supplementary Fig. S1 GapmeR #3 compared to negative control). We noted many examples of commonly up and downregulated genes induced by NRAD1 knockdown in both cell lines, such fibronectin-leucine-rich transmembrane protein 3 (FLRT3), coiled-coil domain containing 68 (CCDC68) mitogen-activated protein kinase 4 (MAPK4), and ubiquitin associated and SH3 domain containing B (UBASH3B).
NRAD1 regulates the expression of hundreds of genes in TNBC (A) Log2 fold changes in gene expression following NRAD1 knockdown (GapmeR #4) compared to control in SUM149 (left) and MDA-MB-468 (right) cells, as determined by microarray (n = 3). A significance threshold pf p < 0.05 and fold change cutoffs of ± 1.4 are indicated by the dashed line. Genes of interest are labeled. The number of upregulated and downregulated genes meeting those cut-offs are indicated. (B) qPCR analysis of selected differentially expressed genes data represents relative expression in NRAD1 targeting GapmeR #4 and GapmeR #3 compared to GapmeR negative control in SUM149 and MDA-MB-468 (n = 5). Statistical significance was determined using one-way ANOVA followed by Dunnett’s multiple comparisons test. Error bars represent the standard deviation. Significant p-values are as follows: * < 0.05, ** p < 0.01, *** p < 0.001, **** p < 0.0001. (C) The total number of significant genes in each cell line were compared to one another and the overlapping upregulated and downregulated genes by NRAD1 knockdown by Gapmer #4 were identified by Venn diagrams. (D) Comparison between differentially expressed genes from microarrays (in A) in MDA-MB-468 (x-axis) and SUM149 (y-axis). Common genes that are also correlated with patient data from TCGA—Cell 2015 data set are highlighted. Genes upregulated with NRAD1 knockdown and negatively correlated with NRAD1 expression in patient data (light blue) and genes downregulated with NRAD1 knockdown and positively correlated with NRAD1 expression in patient data (Navy). Correlation between the two data sets was determined through spearman correlation analysis. (E) Pearson and Spearmen coefficients were calculated based on the co-expression of NRAD1 and all the genes in the TCGA Cell 2015 (n = 815); RNA-Seq RSEM log2. Genes are labeled based on their significance to NRAD1 microarray data (in A). Negatively correlated genes that are upregulated with NRAD1 knockdown in both cell lines are light blue, total number is indicated. Positively correlated genes that are downregulated with NRAD1 knockdown in both cell lines are Navy, total number is indicated. (F) Co-expression analysis between NRAD1 and DUSP5, SLC20A1, MAPK4 and UBASH3B in TCGA Cell 2015 dataset. Each point represents a patients expression profile, TNBC subtype patients are indicated in pink. Spearman correlation (R) and p-value significance are listed.
We validated the gene array data by performing qPCR on a few of these commonly regulated genes and included treatment with a second NRAD1-targeting antisense oligonucleotide, GapmeR #3, which also reduces NRAD1 levels, although to a lesser degree than GapmeR #4, Fig. 1B). A gene array performed with SUM149 cells treated with GapmeR # 3 (Supplementary Fig. S1A) revealed a gene expression change that positively correlated with the gene expression changes induced by GapmeR #4 (R = 0.384, Spearman, Supplementary Fig. S1B). However, the correlation is categorized as weak to moderate. While there is overlap in the genes affected by both GapmeR’s, specifically, with 129 commonly downregulated and 121 commonly upregulated genes, each also produces distinct gene expression changes (Supplementary Fig. S1B). The partial overlap is consistent with the GapmeRs targeting different regions of NRAD1 resulting in differing efficiency in knockdown (with GapmeR #4 inducing greater NRAD1 knockdown, Fig. 1B), and as described in detail later, GapmeR #3 targets additional NRAD1 transcripts.
We compared the overlap of the genes regulated by NRAD1 in two cell lines based on a fold change cutoff of 1.4 and p value < 0.05 and noted a partial overlap, with 165 genes commonly upregulated and 272 genes commonly downregulated in the two cell lines by NRAD1 knockdown (Fig. 1C, Supplementary File 4). Comparing the overall change in gene expression induced by NRAD1 knockdown in SUM149 versus MDA-MB-468 cells revealed a significant weak positive correlation (R = 0.227, Spearman). We identified the genes that were significantly up and down regulated in both cell lines in the correlation plot. Differing gene expression profiles could be attributed to the distinct origin of the two cell lines used. Although both SUM149 and MDA-MB-468 are classified as basal-like TNBC cell lines, SUM149 cells originate from a primary inflammatory breast cancer whereas MDA-MB-468 cells originate from a pleural effusion of metastatic breast adenocarcinoma39 The cell lines have distinct morphologies, molecular receptor expression, cytokine expression, and invasiveness profiles40,41,42. Together, this data suggests that NRAD1 is inducing both common mechanisms of gene regulation and gene regulation mechanism(s) that rely on cell line-specific factors.
The genes commonly regulated by NRAD1 in SUM149 and MDA-MB-468 cells from Fig. 1D were further analysed for co-expression with NRAD1 in breast cancer patient datasets from TCGA (Fig. 1E) and METABRIC (Supplementary Fig. S1C). Genes significantly upregulated (light blue dots) and downregulated (navy dots) by NRAD1 knockdown in the cell lines were associated with negative co-expression and positive co-expression, respectively, in the patient datasets. Selected genes from this analysis were visualized for co-expression with NRAD1 individually (Fig. 1F, TCGA, and Supplementary Fig. S1D, METABRIC). For example, MAPK4 is downregulated when NRAD1 is reduced by knockdown (Fig. 1A,B), and it is positively correlated for co-expression with NRAD1 (Fig. 1F, Supplementary Fig. S1D). Similarly, downregulated UBASH3B (Fig. 1A,B) is positively co-expressed with NRAD1 in patient tumors (Fig. 1F, Supplementary Fig. S1D). Inversely, genes that are upregulated upon NRAD1 knockdown in the cell lines, like dual specificity phosphatase 5 (DUSP5, Fig. 1A) and solute carrier family 20 member 1 (SLC20A1, Fig. 1A,B) are negatively co-expressed with NRAD1 (Fig. 1F, Supplementary Fig. S1D).
Many of these NRAD1 co-expressed genes are critical in breast cancer progression and associated with TNBC. For example, NRAD1-upregulated MAPK4 (Fig. 1A,B) promotes TNBC growth and prevents sensitivity to phosphatidylinositol 3-kinase (PI3K) blockade and is enriched in TNBC/basal-like breast cancers43. Similarly, NRAD1 upregulated UBASH3B (Fig. 1A,B), is a protein tyrosine phosphatase that overexpressed in TNBC and promotes metastasis44. Conversely, NRAD1-downregulated SLC20A1 and DUSP5 is lowly expressed in TNBC/basal-like breast cancers compared to other subtypes and DUSP5 induces tumor suppression and better prognosis in breast cancer45,46,47. We noted that based on co-expression with NRAD1, the NRAD1-regulated genes separated TNBC breast cancer patients (pink dots) from the other breast cancers (Fig. 1F). This suggests NRAD1, which is highly enriched in TNBC/basal-like breast cancers14, could play a role in regulating a subset of gene expression changes associated with TNBCs.
NRAD1 alters the miRNA landscape in TNBC cells, with cell line-specific effects
The results thus far indicated that NRAD1 regulates gene expression in TNBC; however, there are cell line and GapmeR-specific effects. In the transcriptome array data NRAD1 knockdown altered expression of some miRNA genes in a cell line-specific manner, such as MIR4521 in SUM149 cells and MIR361 in MDA-MB-468 cells, which encode the respective primary transcripts for miR-4521 and miR-361 (Fig. 1A). This suggests that NRAD1 effects on miRNAs could be a potential mechanism of gene expression regulation we observe in NRAD1 and could also explain some of the cell line-specific effects.
To explore this possibility of miRNA-regulation by NRAD1 in TNBC, we performed we performed an Affymetrix GeneChip miRNA 4.0 array on small RNA isolated from MDA-MB-468 cells and SUM149 cells treated with negative control or NRAD1-trageting GapmeRs (Fig. 2A, GapmeR #4; Supplementary Fig. S2A, GapmeR #3). The array detects levels of 2578 mature human miRNAs. We identified 82 and 72 upregulated mature miRNAs and 62 and 65 downregulated miRNAs in SUM149 and MDA-MB-468 (GapmeR #4), respectively (Fig. 2A, p-value < 0.05 and fold change cutoffs of ± 1.4.). Knockdown with GapmeR #3 similarly regulated miRNA levels in both cell lines (Supplementary Fig. S2A). Comparison of the changes in the miRNA levels induced by the two NRAD1-targeting GapmeRs revealed significant correlation and overlap with also distinct changes (Spearman correlation: SUM149, 0.473; MDA-MB-468, 0.471, Fig. 2B).
NRAD1 alters the microRNA landscape in TNBC. (A) Log2 fold changes of miRNAs following NRAD1 targeting GapmeR #4 compared to GapmeR negative control in SUM149 and MDA-MB-468 cells, as determined by miRNA microarray (n = 3). Significance cut-offs of p-value < 0.05 and fold-change cutoffs of ± 1.4 and miRNAs meeting these cutoffs are labeled. (B) Correlation between the miRNA microarrays (in S2A and A) comparing NRAD1 targeting GapmeR #3 (x-axis) versus GapmeR #4 (y-axis) SUM149 or MDA-MB-468. Commonly regulated miRNAs are indicated by color (light blue downregulated and blue upregulated) and labeled. Spearman correlation and p-value are indicated. (C) Correlation between the miRNA microarrays comparing MDA-MB-468 (x-axis) and SUM149 (y-axis) in GapmeR #4 treated cells versus negative control. Commonly regulated genes are indicated by color (light blue downregulated and blue upregulated) and labeled. Bolded miRNA names have also been assessed by TaqMan qPCR assays (D). Spearman and p-value are listed. (D) Venn diagrams comparing the number of miRNAs upregulated or downregulated upon NRAD1 knockdown by GapmeR #4 in MDA-MB-468 and SUM149 (E) TaqMan qPCR of selected miRNAs identified in the miRNA microarray (in A). The data represents relative expression of NRAD1 targeting GapmeR #3 or GapmeR #4 compared to GapmeR negative control (n = 5). Statistical significance was determined using one-way ANOVA followed by Dunnett’s multiple comparisons test. Error bars represent the standard deviation. Significant p-values are as follows: * < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
This contrasted with the much weaker correlation and overlap in regulated miRNAs when the effects of the NRAD1-targeting GapmeRs was compared between cell lines (Fig. 2C, GapmeR #4; Supplementary Fig. S2B, GapmeR #3). The cell line and GapmeR-specific effects on some of the regulated miRNAs was further evidenced through individual quantification of the miRNAs with TaqMan qPCR assays (Fig. 2D). While some miRNAs like miR-4485-3p and miR-484 are commonly up and downregulated by both NRAD1-targeting GapmeRs and TNBC cell lines, other miRNAs were only regulated in one cell (e.g., miR-1303). The strong cell line-specific effects in terms of NRAD1 regulation of miRNAs may contribute to the observed cell line-specific gene regulation effects if the miRNAs target these genes (Fig. 1).
NRAD1-regulated miRNA-mRNA networks in TNBC cells are associated with cancer promoting biological processes
Having identified miRNAs regulated by NRAD1, we next evaluated their potential for mediating the gene expression effects induced by NRAD1. We used the MultiMiR tool to identify potential mRNA targets among the genes regulated by NRAD1 (Fig. 1) and the corresponding regulated miRNAs (Fig. 2)32. In SUM149 and MDA-MB-458 cells, 89% and 76% of the mRNAs regulated by NRAD1 are predicted targets of the miRNAs altered by the lncRNA (Fig. 3A, Supplementary File 3). These findings suggest that most gene expression changes induced by NRAD1 may be attributed to its effects on miRNAs.
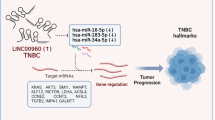
NRAD1-miRNA-mRNA networks and gene ontology analysis (A) Pie charts illustrating the percentage of NRAD1-regulated mRNAs predicted to be regulated by NRAD1-regulated miRNAs. (B) Biological processes identified through gene ontology enrichment of NRAD1 regulated miRNAs. (C) Cystoscope network plots depicting NRAD1-miRNA–mRNA relationships.
Many of the miRNAs regulated by NRAD1 have been associated with cancer promoting and tumor suppressive effects. For example, miR-4485-3p and miR-4458, which are upregulated upon NRAD1 knockdown in the TNBC cells (Fig. 2), have been implicated as having tumor suppressive effects in breast cancer48,49. While miR-1303 and miR-484 have cancer-promoting effects, which are downregulated upon NRAD1 knockdown, have cancer-promoting effects50,51. For example, in liver cancer miR-1303 induces proliferation, migration and invasion, and in breast cancer miR-484 is described as an OncomiR, suppressing the expression of apoptotic genes50,51. To obtain an overall analysis of the biological functions of the miRNAs regulated by NRAD1, we performed a gene ontology (GO) miRNA enrichment analysis, and this revealed significant enrichment of miRNAs associated with cancer-associated gene ontology biological processes (Supplementary File 3). In Fig. 3B we highlight some of these biological processes that were both in common to both cell lines (e.g., cell cycle, proliferation, mitochondrial and stress responses) or distinct to SUM149 (e.g., immune responses) or MDA-MB-468 cells (e.g., apoptosis and DNA damage responses). These functions are inclusive of effects that have been attributed to the miRNAs regulated by NRAD148,49,50,51, and consistent with our previous characterization of NRAD1 effects on the TNBC cells14. This suggests that NRAD1 orchestrates a regulatory network impacting both miRNAs and mRNAs. The resulting miRNA-mRNA networks were refined to highlight selected interactions enriched in GO processes and positions NRAD1 at the center of a miRNA-mRNA network that mediates cancer-promoting biological processes in TNBC (Fig. 3C, Supplementary File 3).
NRAD1-miRNA interactions are cell line-specific due to differences in miRNA and NRAD1 abundances and knockdown efficiencies
We next investigated the mechanism(s) by which NRAD1 alters miRNA levels. For a few miRNAs, such as downregulated miR-4521, its primary transcript, MIR4521 was also downregulated (Fig. 1). We analyzed the gene array data for miRNA host genes and identified a total of 1140, which would give rise to the majority of mature 5p and 3p miRNAs detected by the miRNA arrays. Among the 1140 miRNA host genes we noted that in MDA-MB-468 cells, NRAD1 knockdown resulted in 31 downregulated and 40 upregulated miRNA host genes and SUM149 cells, NRAD1 knockdown resulted in 16 downregulated and 34 upregulated miRNA host genes in SUM149 cells (Supplementary Fig. S3A). We next tracked what happened to the expression of the mature miRNAs that are generated from the down and upregulated miRNA host genes. This revealed that most down and upregulated miRNA host genes did not result in corresponding changes in mature miRNAs (Supplementary Fig. S3B). Therefore, transcriptional changes induced by NRAD1 of miRNA host genes likely are a minor contribution to the overall changes in mature miRNAs induced by NRAD1 (Fig. 2).
Investigation in other cancers have all identified individual ‘sponging’ ceRNA interactions between LINC00284 (i.e., NRAD1) and miRNAs, such as miR-205-3p in lung cancer and miR-27a in colorectal cancer21,22,23,24,25,26. To investigate potential ceRNA interactions NRAD1-regulated miRNAs (Fig. 2) we first identified all potential NRAD1 transcripts. The latest Ensembl release describes 14 NRAD1 transcripts located on chromosome 13: 43,908,669–44,042,929 with both shared, overlapping, and distinct exons (Fig. 4A). The longest transcripts NRAD1-201, 202, 204, and 206, are 3208, 2521, 2307, 2040 ribonucleotides long and share 649 ribonucleotides in common. NRAD1-201 and NRAD1-202 share a common region of 1705 ribonucleotides (from 1503b-3208b for NRAD1-201 and from 292b-1997b for NRAD1-202), which covers most of the NRAD1-202 transcript.
MDA-MB-468 and SUM149 have differing NRAD1 transcript abundance and GapmeR knockdown efficiencies which affects potential ceRNA interactions with miRNAs (A) NRAD1 chromosomal locations and transcript maps as described in ensembl. The binding locations of the GapmeRs and primers are indicated on transcripts. (B) NRAD1 transcript abundance in SUM149 and MDA-MB-468 cells using primers specific to NRAD1-201, NRAD1-202, or NRAD1-201,-202, -204, and -206 (n = 7). Expression is relative to reference genes. Error bars indicate standard deviation. Statistical significance was determined by two-way Anova followed by multiple comparisons post-test. Significant p values are indicated as follows: **p < 0.01, ****p < 0.0001. (C) Relative abundance of NRAD1-201, NRAD1-202, or NRAD1-201, -202, -204, and -206 in SUM149 and MDA-MB-468 cells treated with NRAD1-targeting GapmeR #3 or GapmeR #4 compared to negative control GapmeR. Values are relative to reference genes and negative control treated samples. Error bars indicate standard deviation. Statistical significance was determined using one-way ANOVA followed by Dunnett’s multiple comparisons test, ns = not significant, ****p < 0.0001. (D) The number of unique miRNAs that were predicted to bind NRAD1 transcript variants are plotted against the transcript’s identifier and a Venn diagram identifies the number of overlapping and distinct miRNAs predicted to bind the transcripts. (E) A Euler plot identifies the overlap of regulated miRNAs that are predicted to bind the NRAD1 transcript identified through lncBook. The dots indicate which NRAD1 transcript(s) is predicted to be bound by the regulated miRNAs in the bars above. The bargraphs above indicate the miRNAs regulated by the NRAD1-targeting GapmeRs (top) and in each cell line (middle). (F) The miRNAs predicted to bind NRAD1 through lncBook, are divided into groups; those that are not regulated by NRAD1 and those that regulated by NRAD1 knockdown by GapmeR #3 or GapmeR #4 and the no treatment abundance of the miRNA from miRNA microarrays are compared. Significance was determined by unpaired t-test.
We completed QPCR analysis with primers specific to NRAD1-201 and NRAD1-202, or primers that detect NRAD1-201, 202, 204, and 206 together (Fig. 4A). Overall, MDA-MB-468 has higher levels of NRAD1 compared to SUM149 cells, with over two-fold more NRAD1-201 and NRAD1-202 than in SUM149 cells. NRAD1-201 is especially more abundant in MDA-MB-468 cells (Fig. 4B). Comparing the levels of NRAD1 transcripts with primers that detect NRAD1-201, 202, 204, and 206 versus primers specific to NRAD1-201 or NRAD1-202 suggests that NRAD1-201 and NRAD1-202 are the predominate transcript isoforms expressed in the TNBC cells. We further confirmed the expression of the transcript in the cells by cloning the full length NRAD1-201 and NRAD1-202 sequences from RNA isolated from MDA-MB-468 cells (Supplementary File 3).
We next compared the level of transcript knockdown achieved with the GapmeRs (Fig. 4C). We noted that GapmeR #4 only targets transcripts NRAD1-201, 202, 204 and 206, while GapmeR #3 targets these four transcripts, plus 205, 207, 208, 209 and 212 (Fig. 4A). Given the sufficiency of GapmeR #4 in inducing phenotypic cellular effects14, and associated gene expression changes in the TNBC cancer cells (Fig. 1), it suggests that transcripts 201, 202, 204 and 206 inclusive (and their shared common sequence) are the critical mediators of NRAD1 effects. Quantification of knockdown with the three primer sets suggest more efficient knockdown with GapmeR #4 for NRAD1-201, 202, 204 and 206 in comparison to GapmeR #3 (Fig. 4C). In evaluating the knockdown of NRAD1-201 and NRAD1-202 specifically, NRAD1-202 was more efficiently knocked down in both cell lines, and better NRAD1-202 knockdown was achieved in SUM49 cells. In sharp contrast NRAD1-201 was not significantly knocked down in SUM149 cells. The transcript-specific knockdown efficiencies with respect to NRAD1-201 and 202 in MDA-MB-468 versus SUM149 cells provide a potential mechanism for both the similar and differing miRNA and mRNA regulation effects we observe (Figs. 1, 2 and 3).
Having characterized the relative expression of NRAD1 transcripts in the TNBC cells, we accessed the Lncbook 2.0 tool, which provides comprehensive analysis of validated and predicted canonical miRNA-lncRNA interactions34. These interactions typically depend on sequence complementarity in the 5′ seed region of miRNAs and target RNAs10,52,53. The seed region, typically spans between 6–8 nucleotides, starting at position 2 from the 5′ end of the mature miRNAs. This revealed hundreds of predicted miRNA interactions with the NRAD1 transcripts and that NRAD1-201 and NRAD1-202 have the greatest potential for miRNA interactions (over 2.5 × more than any other NRAD1 transcript, Fig. 4D, Supplementary File 3). We compared the number of unique miRNAs in each NRAD1 transcript and noted a shared core of 103 predicted miRNAs between and NRAD1-201, NRAD1-202, NRAD1-204, and NRAD1-206 (Supplementary Fig. S4, Supplementary File 3). NRAD1-201 and NRAD1-202 also have the most predicted unique miRNA interactions; 101 and 87, respectively (Fig. 4E, Supplementary File 3).
Having identified predicted miRNA interactions for each NRAD1 transcript, (Supplementary File 3), we then cross-referenced these miRNAs with the list of miRNAs regulated by NRAD1 in each cell line upon knockdown by GapmeR #3 and GapmeR #4 (Fig. 2) Approximately 16% of the miRNAs predicted to interact with at least one NRAD1 transcript is also regulated by GapmeRs #3 and/or #4 (Fig. 4E). We compared the relative abundance of all miRNAs predicted to interact with NRAD1 and grouped them as either regulated by NRAD1 (i.e. altered levels upon GapmeR treatment) or not regulated by NRAD1 (no change in miRNA levels upon GapmeR treatment). This revealed that regulated miRNAs are more abundant and have a narrower standard deviation than non-regulated miRNAs (Fig. 4F), with a greater fold increase in MDA-MB-468 cells (fourfold) compared to SUM149 cells (twofold). The greater abundance of regulated miRNAs (especially in MDA-MB-468 cells) and narrower range of the regulated miRNAs versus non-regulated miRNAs is consistent the reliance of ceRNA interactions on equimolar stochiometric relationships54. This concentration dependence on ceRNA interactions is supported by NRAD1 transcripts being over twofold greater in MDA-MB-468 cells than in SUM149 cells (Fig. 4B), which is the reflected in the abundance of miRNAs that are regulated in the respective cell lines (Fig. 4F). The differing abundance of miRNAs and the NRAD1 transcripts results in cell line specific differences in terms of the miRNAs regulated by the lncRNA through ceRNA interactions.
Predicted binding to NRAD1-201 and NRAD1-202 accounts for most the GapmeR-mediated regulations of miRNAs through ceRNA interactions with NRAD1 transcripts (Fig. 4F). NRAD1-201 and NRAD1-202 are predicted to bind 33 of the regulated miRNAs in commonly (second column), and individually they are predicted to bind 35 (first column) and 15 (fourth column) of the regulated miRNAs, respectively (Fig. 4E, Supplementary File 3). We also identified regulated miRNAs specific to that cell line or GapmeR treatment (Fig. 4E). Consistent with the poor NRAD1-201 knockdown in SUM149 cells, regulated miRNAs predicted to bind NRAD1-201 are more abundant in the MDA-MB-468 dataset and comparably lowly represented in the SUM149 dataset. In contrast, miRNAs bound to NRAD1-202 were more abundant in the SUM149 dataset versus MDA-MB-468 dataset (9 versus 5), and consistent with greater knockdown efficiency of NRAD1-202 in SUM149 cells. The analysis was also consistent GapmeR #4 inducing greater effects on the regulated miRNAs predicted to bind the transcripts. Together these analyses suggest that miRNA interactions with specific NRAD1 transcripts contributes to cell line and GapmeR-specific differences in the miRNAs regulated by NRAD1 in the breast cancer cells.
NRAD1 regulates most miRNAs through non-ceRNA mechanisms, including top hit miR-4485-3p
We next compared the proportion miRNAs regulated by NRAD1 (Supplementary File 1) that are predicted to have NRAD1 binding interactions (Supplementary File 3). The proportion of regulated miRNA that have predicted ceRNA interactions range from 19 to 38%. Inversely, this suggests that between 60–80% of all miRNAs regulated by NRAD1 are independent of ceRNA interactions (Fig. 5A, Supplementary File 3). The number of regulated miRNAs that do not interact with NRAD1 may be even greater if the strength of predicted miRNA interactions with NRAD1 is also considered. The Lncbook tool also provides the binding energy of the interactions, which is measured in kilocalories per mole (kcal/mol)34,54,55. Stronger interactions correspond to increasingly negative values, with highly stable complexes usually having energies less than -20 kcal/mol. We visualized the regulated miRNAs and their predicted binding energies with NRAD1- 201 and 202 transcripts (Fig. 5B). Computational analyses frequently use the − 20 kcal/mol as a cutoff to distinguish highly likely functional miRNA–mRNA interactions from weaker ones. Hence most of the regulated miRNAs have weak potential interactions with NRAD1, suggesting a lower likelihood of functionality. Notably, only a fraction of the miRNAs (14 in SUM149 cells and 15 in MDA-MB-468 cells) predicted to bind the NRAD1 transcripts had thermal binding energies lower than the − 20 kcal/mol. These include miR-1229-5p, a miRNA that is upregulated in both cell lines upon NRAD1 knockdown. In SUM149 cells, miR-370-3p is predicted to bind all four transcripts and has the lowest thermal binding energy of − 31.43 kcal/mol; however, it is not regulated in MDA-MB-468 cells. Similarly, miR-760 regulation is specific to MDA-MB-468 cells and has a predicted energy of − 28.48 kcal/mol to NRAD1. These cell line-specific differences can be attributed to differing NRAD1 transcript and miRNA abundances in the cell lines (Fig. 4B,F), which prevent even highly favorable binding interactions if the RNAs are not present in similar equimolar concentrations54.
The majority of miRNA regulation by NRAD1 is through ceRNA-independent mechanisms. (A) Venn diagrams summarize the number of miRNAs predicted to bind NRAD1, the miRNAs regulated by NRAD1, and those that are both regulated by and predicted to bind NRAD1 in SUM149 and MDA-MB-468 cells. (B) The predicted bindings sites and binding energy of miRNAs (from lncBook) that are regulated in the miRNA microarray in SUM149 or MDA-MB-468 cells upon targeting of NRAD1 transcripts 201, 202, 204 and 206 by GapmeR #4. Heatmaps indicate predicted binding energy of miRNAs along the NRAD1 transcript (top plots). The common region between all four transcript is indicated by a navy bar. (C) Potential non-canonical binding site of miR-4485-3p to NRAD1 transcripts, lines indicate sequence binding complementarity. (D) Relative luciferase activity generated by MDA-MB-468 cells transfected with pmirGLO dual-luciferase miRNA target expression vector bearing the potential non-canonical NRAD1 target sequence for miR-4485-3p (wildtype) or mutated version and treated with mimic-miR-4485-3p or mimic negative control (n = 4). Luciferase activity is normalized to the cells treated with the mimic negative control and wildtype vector. Significance was determined by paired t-test, ns not significant.
Given that at least 60–80% of miRNA regulation by NRAD1 are independent of the canonical ceRNA interactions identified through the Lncbook tools we considered alternative mechanisms of miRNA–RNA interactions. The predicted miRNA interactions we identified are isolated to those containing the canonical 5′end seed region complementarity; however, other miRNA–RNA interactions have been reported, including interactions with 3′ end of mature miRNAs56,57,58. Non-canonical miRNA 3-RNA interactions presents a plausible alternative mechanism for regulation of miRNAs by NRAD1.
One of the most upregulated miRNAs upon NRAD1 knockdown by GapmeR #3 and GapmeR #4 in both MDA-MB-468 and SUM149 cells, miR-4485-3p (Fig. 2), is among the list of miRNAs not predicted to interact with NRAD1 by the Lncbook analysis (Fig. 5A, Supplementary File 3). We therefore scanned the NRAD1 transcripts for alternative sequence complementarity with miR-4485-3p that would not be detected by the LncBook tool but could be a non-canonical region of interaction. This revealed one region of high complementarity in the 3′ end of miR-4485-3p with a sequence present in the NRAD1-201 and 202 transcripts, which are targeted by GapmeR #3 and #4 (Fig. 5C). To further interrogate the impact of increased levels of miR-4485-3p on gene expression we assessed NRAD1 regulated mRNAs for possible miR-4485-3p targeting and found that several mRNAs downregulated upon NRAD1 knockdown are predicted to be regulated by miR-4485-3p (Fig. 3C, Supplementary File 3). By performing qPCR, we confirmed that NRAD1 knockdown with GapmeR #3 and #4 downregulated these mRNA targets, consistent with the observed upregulation of miR-4485-3p. The targeting of the mRNAs by miR-4485-3p was further confirmed by treating the cells with miR-4485-3p mimic, which similarly reduced the mRNAs levels in the cells (Supplementary Fig. S5). This data further confirms that miR-4485-3p plays a meaningful part in NRAD1 regulated gene expression through mRNA targeting.
To investigate this potential non-canonical interaction between miR-4485-3p and NRAD1 we performed a miRNA luciferase reporter assay, which is commonly used to confirm miRNA interactions with target RNA sequences59. We cloned the putative target sequence, or a mutated version that should not interact with miRNA, into the reporter plasmid and transfected the plasmids into MDA-MB-468 cells. We then transfected the cells with mimic miR-4485-3p or negative control and quantified the resulting a luciferase. A significant decrease in luciferase activity indicates a positive sequence-specific interaction. Instead, we observed a lack of inhibition of luciferase activity, suggesting that the identified putative region of interaction between the miR-4485-3p and NRAD1 transcript region is not functional (Fig. 5D). We recognize that the negative result in the luciferase assay does not refute the possibility of other ceRNA interactions between NRAD1 and miR-4485-3p. Together with the negative Lncbook analysis that revealed no predicted canonical binding interaction and the lack of other non-canonical sequences that we could find, it is probable that top hit miR-4485-3p is being regulated by NRAD1 in the TNBCs by non-ceRNA mechanisms.
NRAD1 regulates key miRNA biogenesis protein DICER in TNBC cells
The generation of miRNAs is a multi-step process and alterations at any step could change the levels of mature miRNAs generated at the final step. Canonical miRNA biogenesis originates from the nuclear transcription of miRNA genetic loci and the generation of the longer primary-miRNAs (pri-miRNA)10. These pri-miRNAs are cleaved by the microprocessor complex to form precursor-miRNAs (pre-miRNA), which are exported out of the nucleus by exportin5 and RAN-GTP. Cytoplasmic pre-miRNAs undergo the final steps in the maturation processes where the hairpin loop is cleaved by Dicer to form duplex miRNA and then further processed by argonaut (AGO) proteins to form mature miRNA. The sense (5p) and anti-sense (3p) mature miRNAs are formed from the respective 5′ and 3′ arms of the hairpin pre-miRNA. There are variations of this pathway that occur naturally in the body and specifically are disrupted in disease. As in some diseases like cancer, miRNA alteration often stems from disruption in key pathways responsible for their production (Fig. 6A)60.
NRAD1 knockdown reduces protein levels of key miRNA biogenesis player, DICER. (A) Summary of the microRNA biogenesis pathway and where the players act in the pathway. (B) QPCR assessment of effects of NRAD1 knockdown on the miRNA biogenesis pathway regulators in SUM149 and MDA-MB-468 cells. Data represents relative expression in NRAD1 knockdown (GapmeR #3 and GapmeR #4) compared to GapmeR negative control (n = 5). Statistical significance was determined using one-way ANOVA followed by Dunnett’s multiple comparisons test. Significant p-values are as follows: *< 0.05, **p < 0.01. Error bars represent the standard deviation. (C) DICER protein western blot analysis in SUM149 (n = 4) and MDA-MB-468 (n = 6). The bar graphs represent the quantified fold change in the DICER band induced by GapmeR #3 or GapmeR #4 compared to the negative control GapmeR and normalized to the total protein for each sample. Representative images of one of the multiple independent experiments are shown. Statistical significance was determined using one-way ANOVA followed by Dunnett’s multiple comparisons test. Significant p-values are as follows: *< 0.05, **p < 0.01, ***p < 0.001 (D) NRAD1 ChIRP-seq peaks at the DICER1 locus. (E) A network plot of the miRNAs regulated by NRAD1 that are predicted/validated to target DICER1 mRNA.
We wondered if NRAD1 affects levels of the miRNA biogenesis pathway players and assessed their expression levels by qPCR post NRAD1 knockdown with GapmeR #3 and #4 (Fig. 6B). This revealed significant expression effects in various genes. This includes expression changes adenosine deaminase RNA specific (ADAR), which plays a role in miRNA biogenesis and function, including converting adenosines to inosines of pri-miRNA, which may block miRNA maturation61. In SUM149 cells ADAR was highly downregulated by GapmeR #3 but upregulated by GapmeR #4 and in MDA-MB-468 ADAR was similarly downregulated by GapmeR #3 but not changed by GapmeR#4. This GapmeR-specific effect on an enzyme which could block miRNA maturation and function and contribute to some of the GapmeR specific effects we observed in Fig. 2. However, given that GapmeR #4 is the more efficient knockdown of NRAD1 reduction and that ADAR expression was not altered in MDA-MB-468 cells treated with GapmeR #4, we did not follow up on ADAR. Only a few genes were consistently changed in the same direction by both GapmeRs and in both cell lines. Among the more consistent effects, was the downregulation of critical generator of mature miRNAs, DICER1. To interrogate these effects further, we performed western blots on protein extracts and quantified DICER protein levels in the cells post NRAD1 knockdown (Fig. 6C; uncropped blots, Supplementary Fig. S6).
The qPCR and western blot analyses suggest regulation of DICER at the mRNA (DICER1) and at the protein level by NRAD1. In our previous work, we identified NRAD1 interactions with chromatin by performing chromatin isolation by RNA Purification (ChIRP) sequencing14. The analysis revealed greater enrichment of NRAD1 binding at genetic loci among the genes regulated by NRAD1. Genetic loci with ≥ 3 NRAD1 peaks correlated with gene regulation by NRAD114. We accessed the dataset to assess for NRAD1 binding to the DICER1 genetic region (Supplementary File 3)14. This revealed five NRAD1 peaks directly bound to the DICER1 gene and another 24 NRAD1 peaks associated with the DICER1 locus (Fig. 6D). Recent studies have demonstrated that lncRNAs can regulate gene expression from considerable genomic distances, often through mechanisms such as chromatin looping and enhancer-like activity. Binding of lncRNAs at sites over 100kb away from a target gene, as observed in my data at the DICER1 locus, can influence transcription by facilitating long-range interactions between regulatory elements and promoters62,63. We also considered the possibility of post-transcriptional modulation through miRNA targeting. MulitmiR analysis of the miRNAs regulated NRAD1 (Supplementary File 3), revealed multiple NRAD1-regulated miRNAs that target the DICER1 mRNA (Fig. 6E). Therefore, the downregulation of DICER by NRAD1 is likely due to a combination of transcriptional (e.g., chromatin effects) and post-transcriptional (e.g., modulation of miRNAs that target DICER1) mechanisms. Further studies are needed to decipher the exact mechanism of DICER1 regulation by NRAD1.
NRAD1 regulates the location and biogenesis of mitochondrial miRNAs, including miR-4485-3p
As an essential effector of mature miRNA production, DICER downregulation by NRAD1 knockdown is a likely contributor for downregulation of miRNAs upon NRAD1 knockdown (Fig. 6). It does not explain the upregulation of miRNAs like miR-4485-3p, for which we also ruled out ceRNA interactions (Fig. 5). Previous research identified that miR-4485-3p is generated from anti sense non-coding mitochondrial RNA 2 (ASncmtRNA2), which through endonuclease cleavage produces the mitochondrial-origin miR-4485-3p38,64,65. We therefore wondered if NRAD1 is affecting this process, leading to changes in miR-4485-3p.
To test this hypothesis, we first wondered if NRAD1 transcripts are in the mitochondria. We had previously assessed nuclear and cytoplasmic fractions for NRAD1 abundance and found that NRAD1 is a predominantly nuclear-localized14; however, transcript-specific localization and mitochondrial localization was not assessed. So, we performed two isolation methods, one to isolate the nuclear and cytoplasmic fractions and the second to isolate the mitochondria from the cytoplasmic fraction. We first confirmed that the NRAD1 transcripts are predominately nuclear over cytoplasmic by qPCR analysis of cytoplasm and nuclear fractions (Fig. 7A). As positive controls we included quantification of the nuclear-localized nuclear paraspeckle assembly transcript 1 (NEAT1) and cytoplasmic differentiation antagonizing non-protein coding RNA (DANCR), which confirmed successful fractionation (Fig. 7A). We then performed mitochondrial isolation from the cytoplasmic fraction and found mitochondrial enrichment of the NRAD1 transcripts in the mitochondria (especially of NRAD1-201 in MDA-MB-468 cells), along with mitochondrial-localized RNA controls, cytochrome c oxidase 1 (COX1) and mitochondrial 12S rRNA (MT-12S) (Fig. 7B). The presence of NRAD1 in the mitochondria, especially the more abundant NRAD1-201 transcript in MDA-MB-468 cells that has extensive ceRNA potential (Fig. 4), provides potential for function of NRAD1 in the mitochondria in terms of miRNA regulation.
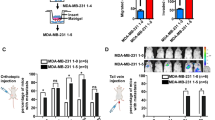
Subcellular localization of NRAD1 and the effect of NRAD1 knockdown on miRNA localization and biogenesis in the mitochondria. (A) The relative percentage of COX1, MT-12S, DANCR, and NRAD1 transcripts in nuclear and cytoplasmic fractions of SUM149 and MDA-MB-468 (n = 4) as quantified by qPCR. (B) The relative percentages of NRAD1, COX1, MT-12S, and DANCR transcripts in the mitochondria and mitochondria-depleted cytoplasmic fractions of SUM149 and MDA-MB-468 cells (n = 4) as quantified by pPCR. (C) The relative percentage of miR-4485-3p, miR-1303, miR-4521, miR-484, RNU48, and miR-25-3p in nuclear and cytoplasmic fractions of SUM149 and MDA-MB-468 (n = 4) as quantified by TaqMan qPCR assay. (B) The relative percentage of miR-4485-3p, miR-1303, miR-4521, miR-484, RNU48, and miR-25-3p in the mitochondria and mitochondria-depleted cytoplasmic fractions of SUM149 and MDA-MB-468 cells (n = 4) as quantified by TaqMan qPCR assays. (E) The mitochondrial origin of miR-4485-3p is derived from the degradation of antisense mitochondrial RNA-2 (ASncmtRNA-2) transcript. (F) The levels of ASncmtRNA-2 are quantified by qPCR in SUM149 and MDA-MB468 cells treated with NRAD1-targetting GapmeR #3 or GapmeR #4 relative to levels in the GapmeR negative control samples (n = 3). Statistical significance was determined using one-way ANOVA followed by Dunnett’s multiple comparisons test. Error bars represent the standard deviation. P value indicated as follows: * < 0.05, ns = not significant. (G) The effect of NRAD1 knockdown (by GapmeR #3 or GapmeR #4) compared to negative control GapmeR on the percentages of miR-4485-3p, miR-1303, miR-4521, miR-484, RNU48, and miR-25-3p in mitochondrial fraction. Data represents the percentage of whole cytoplasmic RNA fraction that is cytoplasmic. Statistical significance was determined using one-way ANOVA followed by Dunnett’s multiple comparisons test. Error bars represent the standard deviation. Significant p-values are as follows: *< 0.05, **p < 0.01, ns not significant.
We expanded our sub-cellular localization analysis to miR-4485-3p and other miRNA examples regulated by NRAD1 (Fig. 7C,D). We included additional miRNA controls with reported nuclear (RNU48) and cytoplasmic (miR-25-3p) location66,67. Comparing the nuclear versus cytoplasmic fractions of MDA-MB-468 cells revealed that all miRNAs were predominately located to the cytoplasm, except for RNU48, as expected (Fig. 7C). Further fractionation of the mitochondria from the remainder of the cytoplasm revealed the specific high enrichment of miR-4485-3p in the mitochondria of MDA-MB-468 cells, consistent with prior studies that showed its mitochondrial origin. This was in contrast with other miRNAs (i.e., miR-1303, miR-4521, miR-484, and miR-25-3p) that are predominately cytoplasmic localized. (Fig. 7D). We noted that RNU48 was split between the mitochondrial and cytoplasmic fractions.
Having confirmed that miR-4485-3p is mitochondrial, we next considered if its upregulation upon NRAD1 knockdown is due to the cleavage/degradation of its pre-cursor ASncmtRNA-2 (Fig. 7E). Consistent with this hypothesis, while NRAD1 knockdown increases miR-4485-3p (Fig. 2), it inversely decreases the levels of its precursor, ASncmtRNA-2, consistent with miRNA biogenesis dynamics (Fig. 7F). Mechanistically, others have shown that the stability of ASncmtRNA-2 and its generation of miR-4485-3p is affected by ceRNA interactions in the hairpin loop region38. Previous ChIRP-seq analysis of NRAD1 revealed that NRAD1 genomic binding regions were highly over-represented in T-rich motifs14. We note that the predicted binding region of ASncmtRNA-2 is 67% U-rich (Supplementary Fig. S7)—which would classify as a U-rich binding regions. Although a U-rich binding region in RNA is not strictly equivalent to a T-rich binding region in DNA, they are functionally analogous due to the chemical similarity between uracil (U) in RNA and thymine (T) in DNA. There are examples of RNA-binding proteins that recognize U-rich RNA sequences also bind T-rich DNA sequences with similar specificity and affinity68. Therefore, the identified putative predicted binding region between NRAD1 and ASncmtRNA-2 (Supplementary Fig. S7) is consistent with the nucleic acid regions primarily bound by NRAD1 as demonstrated by ChIRP-seq14. The reduction of NRAD1 upon knockdown could reduce the stability ASncmtRNA-2 leading to the increased production miR-4485-3p.
Finally, we wondered how NRAD1 affected the mitochondrial-localization of NRAD1-regulated miRNAs (Fig. 7G). The obvious changes were observed in the typically cytoplasmic miR-1303 and miR-484, where upon NRAD1 knockdown, their enrichment was increased in the mitochondria. This was a sharp shift from their overall reduction upon NRAD1 knockdown (Fig. 2). The altered levels of miRNAs in the mitochondria appeared to be a miRNA-specific effect and not a general phenomenon of miRNAs regulated by NRAD1. For example, miR-4521 is a downregulated miRNA, which is mainly cytoplasmic, and it’s still lowly detected in the mitochondrial fraction (Fig. 7G). Changes in miRNA localization in sub-cellular compartments will also affect the potential function since their interactions with target RNA is dependent on direct interactions.
Discussion
In this study we investigated the role of TNBC-enriched NRAD114, in regulating gene expression in breast cancer and found that the lncRNA influences cancer-promoting gene expression through miRNA regulating effects. Based on the general understanding miRNA–lncRNA interactions and how lncRNAs regulate miRNAs, the effects of NRAD1 on miRNAs were surprising in several ways. First, NRAD1 is localized primarily in the nucleus and mitochondria of the TNBC cells and yet has regulatory effects on miRNAs, which is generally believed to be cytoplasmic phenomenon among functionally studied lncRNAs. LncRNAs with predominate nuclear subcellular localization are comparably infrequently reported as having effects on miRNA regulation. This is possibly due to diminished opportunities for direct miRNA–lncRNA interactions, and with respect to mechanisms of gene regulation by lncRNAs, miRNA sponging is the most investigated mechanism of gene regulation by lncRNAs8,16,34.
We do find evidence of miRNA–ceRNA interactions/sponging between NRAD1 and regulated miRNAs; however, miRNA–ceRNA interactions/sponging explain only a fraction of the miRNA regulation by NRAD1 in the TNBC cells, contributing to at most 40% of the miRNAs regulated by NRAD1. We noted that miRNA regulation by NRAD1 through binding interactions only occurred if the miRNAs were in a narrow concentration range in the cells, and lowly or very highly expressed miRNAs are not regulated by NRAD1 despite having predicted binding interactions. This likely reflects the equimolar concentration dependency of physiologically functional ceRNA interactions54. Direct interactions of NRAD1 with miRNAs leading to altered levels was also transcript-dependent and partly explained the differences in miRNAs regulated by NRAD1 in the two cell lines. This finding underscores the complexity of lncRNA–miRNA interactions and suggests that alternative splicing or differential expression of NRAD1 transcripts could fine-tune its regulatory functions in different cellular contexts or cancer types69.
We find that majority of miRNA regulation by NRAD1 (at least 60%, depending on the cell line and condition) is independent of ceRNA mechanisms and NRAD1 therefore does not require direct interaction with miRNAs to alter their levels. This is consistent with the primary nuclear (and mitochondrial) localization of the NRAD1 transcripts, since ceRNA interactions with most cytoplasmic miRNAs would occur in the cytoplasm. Instead, we find alternative mechanisms where NRAD1 knockdown reduces levels of key miRNA biogenesis protein DICER. This suggests that NRAD1 may have a broader impact on miRNA populations by modulating the miRNA processing machinery. Our analysis suggests that NRAD1 affects the transcriptional and post-transcriptional regulation of DICER1. Additional experiments, including assay for transposase accessible chromatin with sequencing (ATAC-seq) would provide critical information about the effects of NRAD1 on chromatin openness, which analyzed alongside ChIRP-seq and RNA-seq data could then be used to interrogate the mechanism of transcriptional regulation by NRAD1. After the consequence of NRAD1 on chromatin structure is mapped, additional experiments such as mutagenesis of specific NRAD1 binding sites associated with altered chromatin structure would be helpful for identifying the most functionally impactful NRAD1 binding sites in terms of subsequent transcriptional effects.
Further investigation into NRAD1’s regulation of the other components of the miRNA biogenesis pathway could provide insights into the global effects of NRAD1 on miRNA homeostasis in cancer cells.
The sub-cellular origin and localization of the miRNAs is another mechanism of miRNA regulation by NRAD1. One of the most highly NRAD1-regulated miRNAs, miR-4485-3p, is of mitochondrial origin and our analysis is consistent with NRAD1 affecting its biogenesis from its mitochondrial precursor ASncmtRNA-2. We also observed that NRAD1 affects the subcellular mitochondrial distribution of some of its regulated miRNAs (e.g. miR-1303 and miR-484). This novel finding implies that NRAD1 may influence miRNA function by altering their accessibility to target mRNAs or other regulatory complexes in different cellular compartments. It is unclear whether NRAD1 acts directly on these miRNAs, miRNA precursors (i.e. ASncmtRNA-2) or indirectly through regulation of other factors such as RNA-binding proteins. Additional functional studies are needed to determine the mechanism by which NRAD1 achieves this redistribution of miRNAs and future findings may reveal new principles of lncRNA-mediated gene regulation.
The widespread effects of NRAD1 on gene expression highlight its potential as a master regulator of breast cancer gene expression programs. As suggested by our co-expression analysis NRAD1 gene regulation is reflected in breast cancer patient tumor data and it is a factor influencing the gene expression that is unique to TNBCs. NRAD1 heighted expression in TNBC contributes to the gene expression that is associated with TNBC (e.g., high MAPK4 expression in TNBC). These gene expression effects are linked to the miRNAs regulated by NRAD1. NRAD1 acts as a hub in miRNA–mRNA regulatory networks, influencing the expression of numerous genes associated with cancer-promoting biological pathways. These gene expression changes are consistent with previous work demonstrating how NRAD1 promotes tumor growth, cancer stem cell maintenance, and cancer cell survival. Our previous ChIRP-seq analysis revealed that up to 60% of genes regulated by NRAD1 have enriched NRAD1 binding in their loci; however, 40% of NRAD1-regulated genes have no binding of NRAD1 to their loci14. This implies post-transcriptional mechanism of gene regulation. This study has now identified miRNA regulation as a major post-transcriptional mechanism of gene regulation by NRAD1 in TNBC.
Conclusion
Our study reveals NRAD1 as a multifaceted regulator of miRNA–mRNA networks in TNBC, employing diverse mechanisms to influence gene expression. Its effects on miRNAs are also dependent on its primary nuclear and mitochondrial localization, which is a departure from the traditional idea that lncRNA regulate miRNA primarily due to ceRNA interactions in the cytoplasm. These findings not only expand our understanding of lncRNA–miRNA interactions in cancer but also highlight NRAD1 as a potential therapeutic target for reprogramming the TNBC transcriptome. Future research building on these discoveries may lead to novel RNA-based therapies and improved treatment strategies for TNBC and potentially other aggressive cancers.
Data availability
The datasets generated during the current study are available in the GEO repository, with the accession codes GSE294997 (gene array data) and GSE295075 (miRNA array data).
References
Arnold, M. et al. Current and future burden of breast cancer: Global statistics for 2020 and 2040. Breast 66, 15 (2022).
Waks, A. G. & Winer, E. P. Breast cancer treatment: A review. JAMA J. Am. Med. Assoc. 321, 288–300 (2019).
Bianchini, G., De Angelis, C., Licata, L. & Gianni, L. Treatment landscape of triple-negative breast cancer—expanded options, evolving needs. Nat. Rev. Clin. Oncol. 19, 91–113 (2021).
Cejalvo, J. M. et al. Intrinsic subtypes and gene expression profiles in primary and metastatic breast cancer. Cancer Res. 77, 2213–2221 (2017).
Bernard, P. S. et al. Supervised risk predictor of breast cancer based on intrinsic subtypes. J. Clin. Oncol. 27, 1160 (2009).
Koboldt, D. C. et al. Comprehensive molecular portraits of human breast tumours. Nature 490, 61–70 (2012).
Okines, A. & Turner, N. Developing therapies for triple-negative breast cancer subtypes. Lancet Oncol. 25, 149–151 (2024).
Venkatesh, J., Wasson, M.-C.D., Brown, J. M., Fernando, W. & Marcato, P. LncRNA-miRNA axes in breast cancer: Novel points of interaction for strategic attack. Cancer Lett. 509, 81–88 (2021).
GENCODE-Human Release Statistics.
O’Brien, J., Hayder, H., Zayed, Y. & Peng, C. Overview of microRNA biogenesis, mechanisms of actions, and circulation. Front. Endocrinol. (Lausanne) 9, 388354 (2018).
Arun, R. P., Cahill, H. F. & Marcato, P. Breast cancer subtype-specific miRNAs: Networks, impacts, and the potential for intervention. Biomedicines. 10 (2022).
Wasson, M. C. D., Venkatesh, J., Cahill, H. F., McLean, M. E., Dean, C. A. & Marcato, P. LncRNAs exhibit subtype-specific expression, survival associations, and cancer-promoting effects in breast cancer. Gene. 901 (2024).
Zhang, Y. et al. Long noncoding RNA LINP1 regulates repair of DNA double-strand breaks in triple-negative breast cancer. Nat. Struct. Mol. Biol. 23, 522–530 (2016).
Vidovic, D. et al. ALDH1A3-regulated long non-coding RNA NRAD1 is a potential novel target for triple-negative breast tumors and cancer stem cells. Cell Death Differ. 27, 363–378 (2020).
Statello, L., Guo, C. J., Chen, L. L. & Huarte, M. Gene regulation by long non-coding RNAs and its biological functions. Nat. Rev. Mol. Cell Biol. 22, 96–118 (2020).
Karagkouni, D. et al. DIANA-LncBase v3: indexing experimentally supported miRNA targets on non-coding transcripts. Nucleic Acids Res. 48, D101–D110 (2020).
Bridges, M. C., Daulagala, A. C. & Kourtidis, A. LNCcation: lncRNA localization and function. J. Cell Biol. 220, e202009045 (2021).
Zhang, X. et al. Mechanisms and functions of long non-coding RNAs at multiple regulatory levels. Int. J. Mol. Sci. 20, 5573 (2019).
Ratti, M. et al. MicroRNAs (miRNAs) and long non-coding RNAs (lncRNAs) as new tools for cancer therapy: First steps from bench to bedside. Target Oncol. 15, 261 (2020).
Li, Y., Li, W., Hoffman, A. R., Cui, J. & Hu, J. F. The nucleus/mitochondria-shuttling LncRNAs function as new epigenetic regulators of mitophagy in cancer. Front. Cell Dev. Biol. 9, 699621 (2021).
Sheng, W. et al. Upregulation of Linc00284 promotes lung cancer progression by regulating the miR-205-3p/c-Met axis. Front. Genet. 12, 694571 (2021).
You, J. et al. Oncogenic long intervening noncoding RNA Linc00284 promotes c-Met expression by sponging miR-27a in colorectal cancer. Oncogene 40, 4151–4166 (2021).
Zhu, M., Yan, X., Zhao, Y., Xue, H., Wang, Z., Wu, B., Li, X. & Shen, Y. lncRNA LINC00284 promotes nucleus pulposus cell proliferation and ECM synthesis via regulation of the miR-205-3p/Wnt/β-catenin axis. Mol. Med. Rep. 25 (2022).
Ke, C. et al. ALDH1A3-Linc00284 axis mediates the invasion of colorectal cancer by targeting TGFβ signaling via sponging miR-361-5p. Int. J. Genom. 2022, 6561047 (2022).
Yan, D., Wu, F., Peng, C. & Wang, M. Silencing of LINC00284 inhibits cell proliferation and migration in oral squamous cell carcinoma by the miR-211-3p/MAFG axis and FUS/KAZN axis. Cancer Biol. Ther. 22, 149–163 (2021).
Ruan, Z. & Zhao, D. Long intergenic noncoding RNA LINC00284 knockdown reduces angiogenesis in ovarian cancer cells via up-regulation of MEST through NF-κB1. FASEB J. 33, 12047–12059 (2019).
Hagedorn, P. H. et al. Locked nucleic acid: modality, diversity, and drug discovery. Drug Discov. Today 23, 101–114 (2018).
Cerami, E. et al. The cBio cancer genomics portal: An open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2 (2012).
Gao, J. et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal 6, pl1 (2013).
Curtis, C. et al. The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature 486, 346 (2012).
Kern, F. et al. miEAA 2.0: integrating multi-species microRNA enrichment analysis and workflow management systems. Nucleic Acids Res. 48, W521–W528 (2020).
Ru, Y. et al. The multiMiR R package and database: integration of microRNA–target interactions along with their disease and drug associations. Nucleic Acids Res. 42, e133 (2014).
Shannon, P. et al. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Li, Z. et al. LncBook 2.0: integrating human long non-coding RNAs with multi-omics annotations. Nucleic Acids Res. 51, D186–D191 (2023).
Krüger, J. & Rehmsmeier, M. RNAhybrid: microRNA target prediction easy, fast and flexible. Nucleic Acids Res. 34, W451 (2006).
Lewis, B. P., Shih, I. H., Jones-Rhoades, M. W., Bartel, D. P. & Burge, C. B. Prediction of mammalian microRNA targets. Cell 115, 787–798 (2003).
Betel, D., Koppal, A., Agius, P., Sander, C. & Leslie, C. Comprehensive modeling of microRNA targets predicts functional non-conserved and non-canonical sites. Genome Biol. 11 (2010).
Fitzpatrick, C., et al. Mitochondrial ncRNA targeting induces cell cycle arrest and tumor growth inhibition of MDA-MB-231 breast cancer cells through reduction of key cell cycle progression factors. Cell Death Dis. 10 (2019).
Holliday, D. L. & Speirs, V. Choosing the right cell line for breast cancer research. Breast Cancer Res. 13, 215 (2011).
Chelouche-Lev, D. et al. Different signalling pathways regulate VEGF and IL-8 expression in breast cancer: implications for therapy. Eur. J. Cancer 40, 2509–2518 (2004).
Fujii, Y., Krishnamurthy, S. & El-Zein, R. Ultrastructural analysis of inflammatory breast cancer cell clusters in an ex vivo environment mechanically mimicking the lymph vascular system. Breast Cancer (Auckl) 15, 11782234211056134 (2021).
Crespo, B. et al. Androgen and estrogen β receptor expression enhances efficacy of antihormonal treatments in triple-negative breast cancer cell lines. Int. J. Mol. Sci. 25, 1471 (2024).
Wang, W. et al. MAPK4 promotes triple negative breast cancer growth and reduces tumor sensitivity to PI3K blockade. Nat. Commun. 13, 1–14 (2022).
Lee, S. T. et al. Protein tyrosine phosphatase UBASH3B is overexpressed in triple-negative breast cancer and promotes invasion and metastasis. Proc. Natl. Acad. Sci. U. S. A. 110, 11121–11126 (2013).
Reddi, K. K. et al. ASAH1 facilitates TNBC by DUSP5 suppression-driven activation of MAP kinase pathway and represents a therapeutic vulnerability. Cell Death Dis. 15, 1–17 (2024).
Liu, T. et al. The suppression of DUSP5 expression correlates with paclitaxel resistance and poor prognosis in basal-like breast cancer. Int. J. Med. Sci. 15, 738 (2018).
Onaga, C. et al. High SLC20A1 expression is associated with poor prognoses in claudin-low and basal-like breast cancers. Anticancer Res. 41, 43–54 (2021).
Fitzpatrick, C. et al. Mitochondrial ncRNA targeting induces cell cycle arrest and tumor growth inhibition of MDA-MB-231 breast cancer cells through reduction of key cell cycle progression factors. Cell Death Dis. 10 (2019).
Liu, X., Wang, J. & Zhang, G. miR-4458 regulates cell proliferation and apoptosis through targeting SOCS1 in triple-negative breast cancer. J. Cell Biochem. 120, 12943–12948 (2019).
Tahtasakal, R. et al. miR-484 as an ‘OncomiR’ in breast cancer promotes tumorigenesis by suppressing apoptosis genes. Ann. Surg. Oncol. 32 (2025).
Huang, J. et al. Micro RNA miR-1303 promotion of proliferation, migration and invasion of human liver cancer cells is enhanced by low Talin 1 (TLN1) expression. Anticancer Res. 42, 4715–4725 (2022).
Kehl, T. et al. About miRNAs, miRNA seeds, target genes and target pathways. Oncotarget 8, 107167 (2017).
O’Brien, J., Hayder, H., Zayed, Y. & Peng, C. Overview of microRNA biogenesis, mechanisms of actions, and circulation. Front. Endocrinol. (Lausanne) 9, 402 (2018).
Diener, C., Keller, A. & Meese, E. The miRNA–target interactions: An underestimated intricacy. Nucleic Acids Res. 52, 1544–1557 (2024).
Ghoshal, A., Shankar, R., Bagchi, S., Grama, A. & Chaterji, S. MicroRNA target prediction using thermodynamic and sequence curves. BMC Genom. 16, 999 (2015).
Chen, J. S., Zeller, R. W., Chen, J. S. & Zeller, R. W. Using engineered microRNAs as vectors for animal RNA interference: Promises and challenges. Adv. Biosci. Biotechnol. 5, 301–310 (2014).
Flamand, M. N., Gan, H. H., Mayya, V. K., Gunsalus, K. C. & Duchaine, T. F. A non-canonical site reveals the cooperative mechanisms of microRNA-mediated silencing. Nucleic Acids Res. 45, 7212 (2017).
Kosek, D. M., Banijamali, E., Becker, W., Petzold, K. & Andersson, E. R. Efficient 3′-pairing renders microRNA targeting less sensitive to mRNA seed accessibility. Nucleic Acids Res. 51, 11162–11177 (2023).
Clément, T., Salone, V. & Rederstorff, M. Dual luciferase gene reporter assays to study miRNA function. Methods Mol. Biol. 1296, 187–198 (2015).
Annese, T., Tamma, R., De Giorgis, M. & Ribatti, D. microRNAs biogenesis, functions and role in tumor angiogenesis. Front. Oncol. 10, 581007 (2020).
Tomaselli, S. et al. ADAR enzyme and miRNA story: A nucleotide that can make the difference. Int. J. Mol. Sci. 14, 22796 (2013).
Gil, N. & Ulitsky, I. Regulation of gene expression by cis-acting long non-coding RNAs. Nat. Rev. Genet. 21, 102–117 (2019).
Ferrer, J. & Dimitrova, N. Transcription regulation by long non-coding RNAs: mechanisms and disease relevance. Nat. Rev. Mol. Cell Biol. 25, 396 (2024).
Bianchessi, V. et al. The mitochondrial lncRNA ASncmtRNA-2 is induced in aging and replicative senescence in endothelial cells. J. Mol. Cell Cardiol. 81, 62–70 (2015).
Farfán, N., Sanhueza, N., Briones, M., Burzio, L. O. & Burzio, V. A. Antisense noncoding mitochondrial RNA-2 gives rise to miR-4485-3p by Dicer processing in vitro. Biol. Res. 54 (2021).
Bai, B., Liu, H. & Laiho, M. Small RNA expression and deep sequencing analyses of the nucleolus reveal the presence of nucleolus-associated microRNAs. FEBS Open Bio 4, 441–449 (2014).
Zhang, J. et al. Excessive miR-25-3p maturation via N6-methyladenosine stimulated by cigarette smoke promotes pancreatic cancer progression. Nat. Commun. 10, 1–15 (2019).
Kim, H. S. et al. Distinct binding properties of TIAR RRMs and linker region. RNA Biol. 10, 579 (2013).
Khan, M. R., Wellinger, R. J. & Laurent, B. Exploring the alternative splicing of long noncoding RNAs. Trends Genet. 37, 695–698 (2021).
Funding
The article was funded by grant support to P.M. from the Canadian Institutes of Health Research (CIHR, PJT 162313 and 191871). H.F.C. is funded by a Research Nova Scotia graduate studentship and a Nova Scotia Graduate Scholarship. J.M.B was partly supported by Genomics in Medicine scholarships from the Dalhousie Medical Research Foundation (DMRF), Cancer Research Training Program (CRTP) scholarships funded through the Beatrice Hunter Cancer Research Institute (BHCRI) and the Terry Fox Research Institute, and Scotia Scholars Awards funded through Research Nova Scotia. M.L.T was funded by a BHCRI Summer Studentship with funds provided from the Canadian Cancer Society’s Carol Ann Cole Comfort Heart Summer Studentship for Breast Cancer Research, and by a National Sciences and Engineering Research Council of Canada (NSERC) undergraduate student research award (NSERC-USRA). J.V. is a trainee of the CRTP of the BHCRI and was supported by funds provided by GIVETOLIVE. M.-C.D.W. is funded by funds provided to PM from Canadian Breast Cancer Foundation-Atlantic Region Endowed Chair in Breast Cancer Research and donation made to Dalhousie University to support breast cancer research conducted by PM. M.C.D.W was funded by a doctoral award from the Canadian Institutes of Health Research (CIHR), a Killam Predoctoral Scholarship, Research Nova Scotia graduate studentship, and is a trainee of the CRTP BHCRI. R.P.A is funded by a I3V Dr. David H. Hubel Postdoctoral Fellowship, is a trainee of the CRTP-BHCRI and was funded by funds generously provided by the Canadian Cancer Society’s JD Irving, Limited—Excellence in Cancer Research Fund. M.E.M was funded by a Faculty of Medicine Scholarship and a Nova Scotia Graduate Scholarship. DV was supported by a Master’s award from the CIHR.
Author information
Authors and Affiliations
Contributions
H.F.C. conceptualized and designed the study and performed the experiments, data analysis, and data interpretation, and wrote the manuscript. J.M.B., J.V., M.C.D.W, R.P.A, M. E. M., and D.V. performed experiments and interpreted the data. PM conceptualized, designed and supervised the study, acquired funding, performed data interpretation and wrote the manuscript. All authors edited and revised the manuscript and were involved in the final approval of the manuscript.
Corresponding author
Ethics declarations
Competing interests
J.V., M.C.D.W, and P.M. are co-founders of Oncolinc Therapeutics Inc., a startup aimed at developing targeted therapies against lncRNAs as novel immunotherapies. P.M. is also a co-founder of Theranib Inc. a startup aimed at developing small molecule drugs targeting ALDH1A3 for the treatment of cancer. H.F.C., J.M.B., M.L.T.,R.P.A.,M.E.M.,and D.V have no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Cahill, H.F., Brown, J.M., Leslie-Toogood, M. et al. LncRNA NRAD1 regulates the triple-negative breast cancer transcriptome by miRNA biogenesis, localization, and predominately non-ceRNA interactions. Sci Rep 15, 26708 (2025). https://doi.org/10.1038/s41598-025-12415-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-12415-7