Abstract
Previous research suggests a link between inflammatory proteins and rotator cuff tears, but the specific protein remains unclear. This two-sample Mendelian randomization (MR) study used genetic variants from a genome-wide association study (GWAS) of 14,824 primarily European participants. We analyzed 91 circulating inflammatory proteins using inverse variance weighting (IVW) and validated results with MR-Egger, the weighted median approach, and sensitivity tests. MR analysis identified three inflammatory proteins with significant associations in the IVW analysis: Interleukin-20 (IL-20) levels (OR = 1.225, 95% CI = 1.036 − 1.450, P = 0.044), Leukemia inhibitory factor levels (LIF) (OR = 1.380, 95% CI = 1.008 − 1.889, P = 0.044), and Signaling lymphocytic activation molecule (SLAM) (OR = 1.183, 95% CI = 1.059 − 1.321, P = 0.015). Elevated IL-20, LIF and SLAM increased the risk of rotator cuff tears. This MR study reveals a potential association between IL-20, LIF and SLAM and the risk of rotator cuff tears, providing insights into their pathogenesis and potential therapeutic targets.
Similar content being viewed by others
Introduction
Rotator cuff tears, defined as tendon ruptures or detachments from the humeral head, represent a common etiology of shoulder pain and dysfuntion1,2. Notably, such tears are also frequently identified in asymptomatic individuals3,4. Epidemiological studies indicate that rotator cuff tears affect approximately 20–22% of the general population, with prevalence increasing with age. Asymptomatic tears are nearly twice as common as symptomatic ones3,5. The pathophysiology of rotator cuff tears is multifactorial, involving various intrinsic and extrinsic factors. Emerging evidence from basic and translational research highlights the pivotal role of inflammation in the onset and progression of rotator cuff tears. Although local inflammation within the tendon microenvironment is recognized as a key contributor to the pathophysiology of rotator cuff tears, some studies suggest that circulating inflammatory proteins may also play an active role in modulating tendon health. Systemic inflammation has been implicated in the initiation and progression of tendon disorders, potentially by facilitating immune cell trafficking and the infiltration of cytokines from the bloodstream into tendon tissue6,7. Studies have demonstrated that pro-inflammatory cytokines such as interleukin-6 (IL-6), interleukin-1β (IL-1β), and tumor necrosis factor-alpha (TNF-α) can cross endothelial barriers, thereby influencing local cellular responses—including tenocyte apoptosis, extracellular matrix (ECM) remodeling, and neoangiogenesis8,9. Elevated levels of these cytokines, along with cyclooxygenase-2 (COX-2), have been associated with tissue degradation, tendon atrophy, and the development of chronic pain symptoms10,11. Nevertheless, the precise molecular and cellular mechanisms through which systemic and local inflammation interact to drive rotator cuff degeneration remain incompletely understood12. A deeper understanding of these mechanisms is crucial for developing effective diagnostic and therapeutic strategies, which could significantly alleviate symptoms and enhance shoulder function. Previous animal experiments revealed that TNF-α IL-6, IL-1β, and IL-17 A is frequently expressed at elevated levels in the subacromial bursae tissue and worn tendon tissue of patients with rotator cuff tears13,14. However, the results of these studies are often susceptible to the influence of confounding factors.
This study uses a bidirectional MR approach to explore the potential associations between genetically predicted levels of circulating inflammatory proteins and the risk of rotator cuff tears. By using genetic variants as instrumental variables, this method helps mitigate confounding factors and clarify the directionality of these associations. Our aim is to delineate the genetic and inflammatory pathways that may predispose to or result from rotator cuff tears, potentially unveiling new therapeutic targets. The insights gained could lead to better prevention and treatment strategies, improving outcomes for those at risk or currently suffering from rotator cuff tears.
Materials and methods
Study design
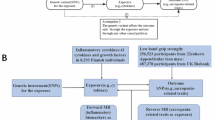
In this study, 91 circulating inflammatory proteins were selected as exposures based on a recent extensive study. A rigorous selection process was implemented to identify appropriate Instrumental variables (IVs) for proxying these exposures. The study utilized aggregated data from the IEU Open GWAS project focused on rotator cuff tears. We restricted our subject population to individuals of European ancestry to minimize demographic stratification biases. No further approval from the institutional review board (IRB) was required, as all data were sourced from previously approved studies with requisite ethical clearance and informed consent obtained from all participants. MR studies are predicated on three critical assumptions: (I) IVs, chosen as genetic variants, must be strongly associated with the exposure of interest. (II) IVs must be independent of any confounders that also affect the outcome. (III) The influence of IVs on the outcome should be exclusively mediated through the exposure. The structure of this bidirectional MR study aligns with established guidelines for conducting MR investigations, as outlined in Fig. 115,16.
The hypothesis principle of MR.
GWAS data sources for 91 inflammatory proteins
Updated GWAS summary statistics for 91 inflammatory proteins were derived from a study by Zhao et al.17involving 14,824 participants, predominantly of European descent, from 11 cohorts. These proteins were analyzed using genome-wide genetic data and plasma proteomics via the Olink Target Inflammation immunoassay panel. Each cohort’s GWAS was conducted using a linear regression-based additive genetic association model. The effect of each inflammatory protein is reported as a change in inverse-rank normalized protein level per allele dose. Adjustments for population substructure were made using genetic principal components. Age and sex were also included as covariates to mitigate potential confounding effects18. Comprehensive quality control measures and data sources are detailed in the original publication17.
Rotator cuff tears GWAS data sources
Data on rotator cuff tears were sourced from the IEU Open GWAS project (https://gwas.mrcieu.ac.uk/)19, with the specific GWAS ID for rotator cuff tears being ebi-a-GCST90044700. This dataset includes data from 5,701 cases and 406,310 controls, encompassing 16,110,542 single nucleotide polymorphisms (SNPs). Notably, all participants in both datasets were of European ancestry, which can help mitigate potential bias arising from demographic stratification18.
Selection of instrumental variables (IVs)
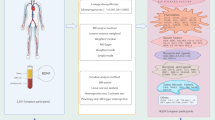
To ensure the validity of IVs in our MR analysis, we applied rigorous quality control measures for SNPs. SNPs were initially selected based on a significance threshold of P < 5 × 10⁻⁶20,21. To minimize the effects of linkage disequilibrium (LD), we applied a clumping criterion with an r² threshold of < 0.001 within a 10,000 kb window22,23. Additionally, the F statistic was utilized to confirm the strength of the IVs, with an inclusion threshold set at F > 10 to ensure robust associations23 (Fig. 2).
The research workflow of this study.
Statistical analysis
All analyses were conducted using R statistical software (version 4.2.1). MR was performed using the “TwoSampleMR” package in R. For multiplicity tests, the “MR_PRESSO” package was utilized. For each exposure, we conducted the primary MR analysis using the IVW method within a multiplicative random-effects framework24. This method combines Wald ratio estimates for individual SNPs to generate an overall causal estimate for each risk factor, with each SNP’s effect calculated as the ratio of its outcome association to its exposure association25. As IVW estimates can be influenced by pleiotropic instruments24we performed sensitivity analyses to account for pleiotropy. Cochran’s Q test evaluated heterogeneity among SNPs, and scatter plots of SNP-exposure and SNP-outcome associations were created to visualize the results26. A leave-one-out analysis assessed the influence of individual SNPs by sequentially excluding each SNP and recalculating the IVW estimates27. MR-PRESSO and MR-Egger regression were further applied to detect and correct horizontal pleiotropy, with MR-PRESSO identifying and removing significant outliers28 (Fig. 2, Supplementary Table S1). Odds ratios (ORs) with 95% confidence intervals (CIs) were calculated to represent the effects of a one-standard deviation increase in lipid-related traits. To investigate the bidirectional causal relationships between rotator cuff tears and inflammatory proteins, we designated rotator cuff tears as the “exposure” and inflammatory proteins associated with rotator cuff tears as the “outcome.” SNPs significantly associated with rotator cuff tears (P < 1 × 10− ⁵) were selected29,30. SNPs with F-statistic values below 10 were excluded to avoid weak instrument bias23. To ensure robustness, we applied the Benjamini-Hochberg false discovery rate (FDR) adjustment to P-values when analyzing associations between inflammatory proteins and rotator cuff tears. Associations were considered statistically significant if FDR-adjusted P-values were below 0.0531,32.
Results
Exploring the impact of 91 inflammatory proteins on the association of rotator cuff tears
In the MR analysis of 91 inflammatory proteins in relation to rotator cuff tears, genetically predicted levels of three proteins—IL-20, LIF, and SLAM—were significantly associated with rotator cuff tear risk based on the IVW analysis (Fig. 3, Supplementary Table S2). All reported P-values were adjusted for multiple comparisons using the FDR correction.
The result of MR analysis for 91 inflammatory proteins on rotator cuff tears.
For IL-20 levels, twelve SNPs were analyzed, all with F values > 10, indicating strong instrumental variables. The IVW method revealed a potential association between IL-20 levels and rotator cuff tears (OR = 1.225, 95% CI = 1.036–1.450, P = 0.044). Alternative MR methods, including MR Egger (OR = 1.053, 95% CI = 0.770–1.440, P = 1.000), weighted median (OR = 1.155, 95% CI = 0.920–1.449, P = 0.356), weighted mode (OR = 1.125, 95% CI = 0.806–1.570, P = 0.837), and simple mode (OR = 1.170, 95% CI = 0.799–1.711, P = 0.728), did not reach statistical significance. Nevertheless, the estimated effects across these methods were all in the same direction, lending some support to the IVW findings (Fig. 4).
The putative association of IL-20 levels on rotator cuff tears. (A) A forest plot delineated the associations of SNPs associated with IL-20 levels on rotator cuff tears, highlighting individual SNP contributions. (B) Analysis using a funnel plot indicated minimal heterogeneity within this MR study, suggesting a uniform distribution of the effects across studies. (C) The leave-one-out method revealed that the removal of individual SNPs does not significantly alter the overall conclusions, indicating robustness in our findings. (D) Scatter plots illustrating the genetic associations between IL-20 levels and rotator cuff tears were used to visually assess the correlation between genetic predispositions and disease outcomes.
For LIF levels, six SNPs were evaluated, all with F values > 10. The IVW method indicated a potential association with rotator cuff tears (OR = 1.380, 95% CI = 1.008–1.889, P = 0.044), and the weighted median method also produced a statistically significant result (OR = 1.529, 95% CI = 1.111–2.105, P = 0.046). Although the weighted mode (OR = 1.675, 95% CI = 1.082–2.593, P = 0.344) and simple mode (OR = 1.661, 95% CI = 1.059–2.607, P = 0.391) yielded comparable effect estimates, their results did not reach significance. Overall, the pattern of results was consistent with a potential positive association (Fig. 5).
The putative association of LIF levels on rotator cuff tears. (A) A forest plot was constructed to showcase the effects of SNPs linked to LIF levels on rotator cuff tears, presenting the influence of individual SNPs. (B) A funnel plot analysis indicated minimal heterogeneity across the MR analysis, supporting the consistency of the SNP effects. (C) Application of the leave-one-out method demonstrated that omitting individual SNPs does not critically alter the overarching conclusions, affirming the stability of our results. (D) Scatter plots were employed to depict the genetic associations between LIF levels and rotator cuff tears, facilitating a visual examination of the genetic linkage and potential association.
For SLAM levels, twenty-three SNPs were included, each with F values > 10. The IVW analysis showed a statistically significant association with rotator cuff tears (OR = 1.183, 95% CI = 1.059–1.321, P = 0.015). However, none of the alternative MR methods—MR Egger (OR = 1.139, 95% CI = 0.914–1.420, P = 1.000), weighted median (OR = 1.070, 95% CI = 0.913–1.254, P = 0.506), weighted mode (OR = 1.069, 95% CI = 0.877–1.303, P = 0.648), and simple mode (OR = 1.212, 95% CI = 0.937–1.568, P = 0.394)—achieved statistical significance. Despite this, the convergence of effect direction across methods enhances the credibility of the observed association (Fig. 6).
The putative association of SLAM levels on rotator cuff tears. (A) A forest plot illustrated the association of SNPs related to SLAM levels on rotator cuff tears, detailing the impact of each genetic variant. (B) A funnel plot analysis revealed minimal heterogeneity within this MR study, suggesting a consistent effect size across the included SNPs. (C) The leave-one-out method confirmed that removing individual SNPs did not significantly influence the overall conclusions, indicating robustness in the association inference. (D) Scatter plots were generated to visually represent the genetic associations between SLAM levels and rotator cuff tears, providing insight into the genetic correlations and their potential pathways.
Employing a range of MR methods, we found that the IVW approach yielded statistically significant associations for all three inflammatory proteins—IL-20, LIF, and SLAM. While alternative methods such as the weighted median, weighted mode, MR-Egger, and simple mode did not consistently reach statistical significance, their effect estimates were largely directionally consistent with those from the IVW approach (Supplementary Table S2). This consistency across methods, despite differences in statistical power and assumptions, strengthens the credibility of the observed associations. The IVW method is known for its higher statistical efficiency when the instrumental variables satisfy core assumptions—particularly the absence of horizontal pleiotropy and invalid instruments33. Therefore, in the context of no evidence for heterogeneity or directional pleiotropy, we primarily interpreted our findings based on the IVW estimates. Nevertheless, we acknowledge that alternative MR approaches offer robustness against certain violations of these assumptions, and their nonsignificant results warrant cautious interpretation of our findings rather than overreliance on any single method. All selected SNPs had F-statistics exceeding the conventional threshold of 10, suggesting sufficient instrument strength and a low likelihood of weak instrument bias. Furthermore, results from sensitivity analyses (Supplementary Table S3-4) supported the overall robustness of the associations34.
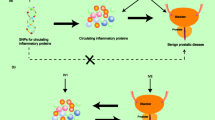
Exploring the impact of rotator cuff tears on the association of 91 inflammatory proteins
To investigate whether rotator cuff tears is associated with alterations in inflammatory protein levels, we conducted reverse MR analyses using twenty-two SNPs significantly linked to rotator cuff tears as instrumental variables. All selected SNPs had F-statistics > 10, indicating their strength and reliability as instruments. For IL-20 levels, the MR results showed no significant association across all methods: IVW (P = 0.462), MR Egger (P = 0.168), weighted median (P = 0.479), weighted mode (P = 0.938), and simple mode (P = 0.921). Likewise, analyses for LIF levels yielded no evidence of an association: IVW (P = 0.857), MR Egger (P = 0.441), weighted median (P = 0.958), weighted mode (P = 0.363), and simple mode (P = 0.503). Similarly, no significant association was observed for SLAM levels: IVW (P = 0.583), MR Egger (P = 0.466), weighted median (P = 0.731), weighted mode (P = 0.978), and simple mode (P = 0.922) (Table 1). Taken together, these findings suggest that rotator cuff tears is not linked to detectable changes in the circulating levels of IL-20, LIF, or SLAM. This indicates that rotator cuff tears may not exert a measurable effect on the systemic levels of these specific inflammatory proteins.
Discussion
In this MR study, we identified potential associations between genetically predicted levels of IL-20, LIF, and SLAM with increased risk of rotator cuff tears. These associations were statistically significant in the IVW analysis, and although other MR methods such as weighted median and MR Egger did not consistently reach significance, the consistent direction of effect estimates across methods reinforces the plausibility of our findings. By contrast, the reverse MR analyses did not support an effect of rotator cuff tears on these inflammatory protein levels, suggesting a unidirectional influence from inflammatory pathways toward tendon degeneration, rather than inflammation being a secondary consequence of structural changes in the rotator cuff.
The associations identified in this study between elevated levels of IL-20, LIF, and SLAM and the risk of rotator cuff tears support the hypothesis that inflammation may play an upstream role in tendon degeneration. IL-20, a member of the IL-10 cytokine family, plays a crucial role in promoting chronic inflammation and tissue remodeling. Studies have identified elevated IL-20 expression in degenerative musculoskeletal conditions, such as lumbar spinal stenosis and spondylolisthesis, where it contributes to inflammatory infiltration and extracellular matrix degradation35. Although, direct evidence of IL-20 receptor expression or functional effects in human tendon or tenocytes remains lacking, the role of IL-20 in related musculoskeletal tissues suggests a potential upstream role in tendon degeneration. LIF, a multifunctional cytokine involved in both inflammation and tissue regeneration, is increasingly recognized as a key modulator in musculoskeletal disease. Elevated LIF levels have been observed in conditions such as ankylosing spondylitis, where it exacerbates local inflammation and abnormal tissue remodeling36. While some study shown LIF is exert both reparative and pathological effects in joint tissues37its specific expression patterns and functional roles in tendon cells or tendon healing models have not been well characterized, and this represents an important direction for future studies.
The SLAM receptors, initially identified in immune cells, are also expressed in a variety of hematopoietic and non-hematopoietic cell types. These receptors play essential roles in immune cell activation and cytokine production, and have emerged as potential biomarkers in inflammatory and joint-related diseases38. However, direct evidence demonstrating the presence of SLAM receptors in tendon-resident cells or infiltrating immune cells within tendon tissue remains insufficient. Nevertheless, SLAM signaling plays a crucial role in coordinating macrophage and T cell function—two key immune cell populations involved in chronic tendon pathology39,40. Through the modulation of cytokine production and the resolution of inflammation, SLAM receptors may indirectly impact pathological features such as local inflammatory responses, muscle atrophy, and fibrosis observed in tendon degeneration. Furthermore, downstream pathways, including RhoA and ceRNA regulatory networks, have been proposed in other musculoskeletal diseases41,42; however, their specific involvement in tendon pathology has yet to be experimentally confirmed. These knowledge gaps underscore the necessity for further mechanistic investigations to clarify the roles of SLAM receptors and related signaling cascades in tendon disease processes.
Our findings are in agreement with prior histological and molecular studies that report elevated levels of inflammatory cytokines—including IL-1β, IL-6, and TNF-α—in degenerated or torn rotator cuff tissues43. For example, IL-6 has been shown to mediate fibroblast activation and promote matrix remodeling in early tendinopathy44while IL-17 A contributes to the initiation of tendon inflammation via the activation of MMPs and chemokine pathways45. However, most of these prior investigations were observational or conducted in animal models, making it challenging to infer association or establish the direction of effect. By employing MR, our study mitigates the influence of confounding and reverse causality, providing stronger evidence for a unidirectional relationship from elevated inflammatory proteins to tendon degeneration. Notably, reverse MR analyses did not support an effect of rotator cuff tears on circulating inflammatory protein levels, reinforcing the idea that inflammation precedes structural changes, rather than being a reactive consequence of tendon injury.
Collectively, these results suggest that IL-20, LIF, and SLAM may represent mechanistic links between immune dysregulation and tendon degeneration. These inflammatory proteins may represent potential biomarkers for the early identification of rotator cuff pathology and could be explored as prospective targets for therapeutic intervention aimed at modulating upstream inflammatory pathways. However, their precise roles in tendon biology and disease progression require further experimental validation. Clinically, the MR-based approach enhances causal inference and provides a strong rationale for future trials of anti-inflammatory or immunomodulatory therapies in tendinopathies, particularly among individuals with chronic inflammation or systemic immune dysregulation.
However, This study also has several limitations that should be acknowledged. First, although we systematically investigated 91 circulating inflammatory proteins, the underlying GWAS data were primarily derived from individuals of European ancestry. This population-specific dataset may limit the generalizability of our findings to other ethnic groups. Future replication studies in more diverse populations are necessary to confirm the observed associations. Second, while we identified potential associations between genetically predicted inflammatory protein levels and the risk of rotator cuff tears, MR reflects the impact of long-term, lifelong exposures based on genetic predisposition. Therefore, our findings may not fully capture transient or stage-specific fluctuations in inflammatory protein levels that occur during the progression of tendon pathology. Further studies incorporating time-resolved proteomic or transcriptomic data are warranted to elucidate these dynamic processes46.Third, although the IVW method produced statistically significant results in our primary analyses, other complementary MR methods—including the weighted median, weighted mode, MR-Egger, and simple mode—did not consistently reach statistical significance. Although the general concordance in the direction of effect across these methods supports the plausibility of our findings, the absence of consistent statistical significance highlights potential sensitivity to the choice of analytical method. These alternative approaches are more robust to violations of key MR assumptions, such as horizontal pleiotropy or the presence of invalid instruments. Consequently, our results should be interpreted with caution, and further validation using larger, independent datasets and orthogonal analytical frameworks is needed to strengthen causal inference. Fourth, this study lacks direct experimental validation regarding the roles of IL-20, LIF, and SLAM in tendon tissue. The expression of these protein receptors in tenocytes or tendon-resident progenitor cells has not been conclusively demonstrated, and histological evidence for SLAM-positive immune cell infiltration in human rotator cuff tendons remains unavailable. Additionally, while signaling pathways such as RhoA and ceRNA networks have been implicated in other degenerative musculoskeletal conditions, their functional involvement in tendon pathology has not yet been established. These limitations emphasize the exploratory and hypothesis-generating nature of our work. Although IL-20, LIF, and SLAM appear to be potential mediators linking systemic immune dysregulation with tendon degeneration, their precise mechanistic roles in tendon biology remain to be clarified. Future research should focus on mechanistic studies involving receptor expression analysis, functional assays in tendon-derived cells, and in vivo modeling to elucidate their potential as biomarkers or therapeutic targets in rotator cuff disease.
Conclusion
In summary, our findings indicate a potential association between genetically proxied levels of IL-20, LIF, and SLAM and the risk of rotator cuff tears, as supported by IVW analysis and consistent directions in sensitivity analyses. After adjusting for multiple comparisons using the FDR, these associations remained statistically significant, reinforcing the role of dysregulated immune signaling in tendon degeneration rather than as a secondary response to structural disruption. By leveraging MR methodology, this study minimizes confounding and reverse causation, providing stronger evidence for a unidirectional effect from inflammation to tendon pathology. IL-20, LIF, and SLAM may serve as candidate biomarkers or therapeutic targets, highlighting the potential of targeting upstream inflammatory pathways for the early prevention and clinical management of rotator cuff disease.
Data availability
The datasets analysed during the current study are available in the EBI GWAS Catalog (accession numbers from GCST90274758 to GCST90274848) (https://www.ebi.ac.uk/gwas/) and IEU Open GWAS (https://gwas.mrcieu.ac.uk/).
References
Schmidt, C. C., Jarrett, C. D. & Brown, B. T. Management of rotator cuff tears. J. Hand Surg. Am. 40 (2), 399–408. https://doi.org/10.1016/j.jhsa.2014.06.122 (2015).
Grusky, A. Z., Song, A. & Kim, P. Factors associated with symptomatic rotator cuff tears: the rotator cuff outcomes workgroup cohort study. Am. J. Phys. Med. Rehabil. 100 (4), 331–336. https://doi.org/10.1097/PHM.0000000000001684 (2021).
Yamamoto, A. et al. Prevalence and risk factors of a rotator cuff tear in the general population. J. Shoulder Elb. Surg. 19 (1), 116–120. https://doi.org/10.1016/j.jse.2009.04.006 (2010).
Moosmayer, S. et al. Prevalence and characteristics of asymptomatic tears of the rotator cuff: an ultrasonographic and clinical study. J. Bone Joint Surg. Br. 91 (2), 196–200. https://doi.org/10.1302/0301-620X.91B2.21069 (2009).
Minagawa, H. et al. Prevalence of symptomatic and asymptomatic rotator cuff tears in the general population: from mass-screening in one village. J. Orthop. 10 (1), 8–12. https://doi.org/10.1016/j.jor.2013.01.008 (2013).
Millar, N. L., Murrell, G. A. & McInnes, I. B. Inflammatory mechanisms in tendinopathy - towards translation. Nat. Rev. Rheumatol. 13 (2), 110–122. https://doi.org/10.1038/nrrheum.2016.213 (2017).
Dakin, S. G. et al. Inflammation activation and resolution in human tendon disease. Sci. Transl Med. 7 (311), 311ra173. https://doi.org/10.1126/scitranslmed.aac4269 (2015).
Schulze-Tanzil, G. et al. The role of pro-inflammatory and immunoregulatory cytokines in tendon healing and rupture: new insights. Scand. J. Med. Sci. Sports. 21 (3), 337–351. https://doi.org/10.1111/j.1600-0838.2010.01265 (2011).
Fu, C. et al. Cellular and molecular modulation of rotator cuff muscle pathophysiology. J. Orthop. Res. 39 (11), 2310–2322. https://doi.org/10.1002/jor.25179 (2021).
Struzik, S. et al. Selected clinical features fail to predict inflammatory gene expressions for TNF-α, TNFR1, NSMAF, Casp3 and IL-8 in tendons of patients with rotator cuff tendinopathy. Arch. Immunol. Ther. Exp. (Warsz). 69 (1), 6. https://doi.org/10.1007/s00005-021-00610-z (2021).
Cipollaro, L. et al. Immunohistochemical features of rotator cuff tendinopathy. Br. Med. Bull. 130 (1), 105–123. https://doi.org/10.1093/bmb/ldz016 (2019).
Yuan, T. et al. Proteomic analysis reveals rotator cuff tears caused by oxidative stress. Ther. Adv. Chronic Dis. 12, 2040622320987057. https://doi.org/10.1177/2040622320987057 (2021).
Millar, N. L. et al. IL-17A mediates inflammatory and tissue remodelling events in early human tendinopathy. Sci. Rep. 6, 27149. https://doi.org/10.1038/srep27149 (2016).
Stengaard, K. et al. Early-stage inflammation changes in supraspinatus muscle after rotator cuff tears. J. Shoulder Elb. Surg. 31, 1344–1356. https://doi.org/10.1016/j.jse.2021.12.046 (2022).
Skrivankova, V. W. et al. Strengthening the reporting of observational studies in epidemiology using Mendelian randomization: the STROBE-MR statement. JAMA 326, 1614–1621. https://doi.org/10.1001/jama.2021.18236 (2021).
Burgess, S. et al. Guidelines for performing Mendelian randomization investigations: update for summer 2023. Wellcome Open. Res. 4, 186. https://doi.org/10.12688/wellcomeopenres.15555.3 (2019).
Zhao, J. H. et al. Genetics of Circulating inflammatory proteins identifies drivers of immune-mediated disease risk and therapeutic targets. Nat. Immunol. 24, 1540–1551. https://doi.org/10.1038/s41590-023-01588-w (2023).
Liang, Y. C. et al. Association of Circulating inflammatory proteins with type 2 diabetes mellitus and its complications: a bidirectional Mendelian randomization study. Front. Endocrinol. (Lausanne). 15, 1358311. https://doi.org/10.3389/fendo.2024.1358311 (2024).
Sollis, E. et al. The NHGRI-EBI GWAS catalog: knowledgebase and deposition resource. Nucleic Acids Res. 51, D977–D985. https://doi.org/10.1093/nar/gkac1010 (2023).
Ji, D., Chen, W. Z., Zhang, L., Zhang, Z. H. & Chen, L. J. Gut microbiota, Circulating cytokines and dementia: a Mendelian randomization study. J. Neuroinflammation. 21 (1), 2. https://doi.org/10.1186/s12974-023-02999-0 (2024).
Yu, M., Li, Y., Li, B. & Ge, Q. Inflammatory biomarkers and delirium: a Mendelian randomization study. Front. Aging Neurosci. 15, 1221272. https://doi.org/10.3389/fnagi.2023.1221272 (2023).
Myers, T. A., Chanock, S. J. & Machiela, M. J. LDlinkR: an R package for rapidly calculating linkage disequilibrium statistics in diverse populations. Front. Genet. 11, 157. https://doi.org/10.3389/fgene.2020.00157 (2020).
Burgess, S., Small, D. S. & Thompson, S. G. A review of instrumental variable estimators for Mendelian randomization. Stat. Methods Med. Res. 26 (5), 2333–2355. https://doi.org/10.1177/0962280215597579 (2017).
Burgess, S., Butterworth, A. & Thompson, S. G. Mendelian randomization analysis with multiple genetic variants using summarized data. Genet. Epidemiol. 37 (7), 658–665. https://doi.org/10.1002/gepi.21758 (2013).
Palmer, T. M. et al. Instrumental variable Estimation of causal risk ratios and causal odds ratios in Mendelian randomization analyses. Am. J. Epidemiol. 173 (12), 1392–1403. https://doi.org/10.1093/aje/kwr026 (2011).
Cohen, J. F. et al. Cochran’s Q test was useful to assess heterogeneity in likelihood ratios in studies of diagnostic accuracy. J. Clin. Epidemiol. 68 (3), 299–306. https://doi.org/10.1016/j.jclinepi.2014.09.005 (2015).
Burgess, S. & Thompson, S. G. Interpreting findings from Mendelian randomization using the MR-Egger method. Eur. J. Epidemiol. 32 (5), 377–389. https://doi.org/10.1007/s10654-017-0276-5 (2017).
Verbanck, M. et al. Detection of widespread horizontal Pleiotropy in causal relationships inferred from Mendelian randomization between complex traits and diseases. Nat. Genet. 50 (5), 693–698. https://doi.org/10.1038/s41588-018-0099-7 (2018).
Chen, J. et al. Antioxidants, minerals and vitamins in relation to crohn’s disease and ulcerative colitis: A Mendelian randomization study. Aliment. Pharmacol. Ther. 57 (4), 399–408. https://doi.org/10.1111/apt.17392 (2023).
Xie, J. et al. The associations between modifiable risk factors and nonalcoholic fatty liver disease: A comprehensive Mendelian randomization study. Hepatology 77 (3), 949–964. https://doi.org/10.1002/hep.32728 (2023).
Che, B. et al. Causal inference between pernicious anemia and cancers: a bidirectional two-sample Mendelian randomization analysis. BMC Cancer. 24 (1), 586. https://doi.org/10.1186/s12885-024-12354-y (2024).
Li, W. & Wang, W. Causal effects of exposure to ambient air pollution on cancer risk: insights from genetic evidence. Sci. Total Environ. 912, 168843. https://doi.org/10.1016/j.scitotenv.2023.168843 (2024).
Fei, Y., Yu, H., Wu, Y. & Gong, S. The association between immune cells and ankylosing spondylitis: a bidirectional Mendelian randomization study. Arthritis Res. Ther. 26, 24. https://doi.org/10.1186/s13075-024-03266-0 (2024).
Ouyang, Y. & Dai, M. Associations between systemic inflammatory cytokines and adhesive capsulitis: a bidirectional Mendelian randomization study. Front. Immunol. 15, 1380889. https://doi.org/10.3389/fimmu.2024.1380889 (2024).
Huang, K. Y. et al. The roles of IL-19 and IL-20 in the inflammation of degenerative lumbar spondylolisthesis. J. Inflamm. 15, 19. https://doi.org/10.1186/s12950-018-0195-6 (2018).
Kong, W. et al. LIF is dysregulated in ankylosing spondylitis and contributes to bone formation. Int. J. Rheum. Dis. 25, 592–600. https://doi.org/10.1111/1756-185X.14312 (2022).
Ortiz-Muñoz, G. et al. Cystic fibrosis transmembrane conductance regulator dysfunction in platelets drives lung hyperinflammation. J. Clin. Invest. 130 (4), 2041–2053. https://doi.org/10.1172/JCI129635 (2020).
Zheng, Y. et al. Role of SLAM of receptors in the pathogenesis of rheumatoid arthritis: insights and application. Front. Pharmacol. 14, 1306584. https://doi.org/10.3389/fphar.2023.1306584 (2023).
Millar, N. L. et al. Inflammation is present in early human tendinopathy. Am. J. Sports Med. 38 (10), 2085–2091. https://doi.org/10.1177/0363546510372613 (2010).
Malmgaard-Clausen, N., Moelkjaer, Kjaer, M. & Dakin, Stephanie, G. Pathological Tendon Histology in Early and Chronic Human Patellar Tendinopathy, Translational Sports Medicine, 2799665, 9 pages, 2022. (2022). https://doi.org/10.1155/2022/2799665
Davies, M. R. et al. Lysophosphatidic acid-induced RhoA signaling and prolonged macrophage infiltration worsens fibrosis and fatty infiltration following rotator cuff tears. J. Orthop. Res. 35 (7), 1539–1547. https://doi.org/10.1002/jor.23384 (2017).
Huang, B. Z., Jing-Jing, Y., Dong, X. M., Zhuan Zhong, X. N. & Liu Analysis of the lncRNA-Associated competing endogenous RNA (ceRNA) network for tendinopathy. Genet. Res. (Camb). 2022, 9792913. https://doi.org/10.1155/2022/9792913 (2022).
Jomaa, G. et al. A systematic review of inflammatory cells and markers in human tendinopathy. BMC Musculoskelet. Disord. 21, 78. https://doi.org/10.1186/s12891-020-3094-y (2020).
Stauber, T. et al. Il-6 signaling exacerbates hallmarks of chronic tendon disease by stimulating reparative fibroblasts. Elife 12, RP87092. https://doi.org/10.7554/eLife.87092 (2025). Published 2025 Feb 7.
Arvind, V. & Huang, A. H. Reparative and maladaptive inflammation in tendon healing. Front. Bioeng. Biotechnol. 9, 719047. https://doi.org/10.3389/fbioe.2021.719047 (2021). Published 2021 Jul 19.
Nagase, H., Visse, R. & Murphy, G. Structure and function of matrix metalloproteinases and timps. Cardiovasc. Res. 69, 562–573. https://doi.org/10.1016/j.cardiores.2005.12.002 (2006).
Funding
National Natural Science Foundation of China (82405214), Postdoctoral Fellowship Program of CPSF (GZC20240344), China Postdoctoral Science Foundation (2024M760668), University-Hospital Joint Fund Project of Guangzhou University of Chinese Medicine (GZYZS2024U02), National Demonstration Pilot Project for the Inheritance and Innovative Development of Traditional Chinese Medicine (YN2024B001), Guangdong Provincial Bureau of Traditional Chinese Medicine (20251434). Hainan Provincial Natural Science Foundation of China (823RC596).
Author information
Authors and Affiliations
Contributions
The study was conceptualized and designed by Li Zhou. Data collection, organization, and analysis were managed by Yao Wang, Binbin Xu, and Chenhao Ma. Donghai Huang contributed substantially to the creation of figures, thereby enhancing the manuscript’s analytical depth and visual presentation. The initial draft of the manuscript was written by Yao Wang, with critical revisions provided by Binbin Xu. Funding was secured by Yao Wang and Li Zhou.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethical approval
All datasets utilized in this study are publicly available and were derived from research that had previously secured patient consent and ethical approval. Consequently, no additional ethical approval or informed consent was necessary for this analysis.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, Y., Xu, B., Ma, C. et al. Association between inflammatory proteins and rotator cuff tears: a bidirectional Mendelian randomization study. Sci Rep 15, 26987 (2025). https://doi.org/10.1038/s41598-025-12785-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-12785-y