Abstract
The gold standard for the diagnosis of sleep apnoea (SA) is polysomnography, consisting of overnight in-lab tests, which are expensive for both patients and healthcare systems. Airflow and pulse/oximetry signals contain most of the necessary information for detecting SA and widely simplify the data acquisition process, hence holding the promise to increase the availability of SA diagnosis and reduce waitlists. Deep learning has recently shown some interesting steps forward in analysing these signals in paediatric patients. Here we introduce a novel platform, REST, that is able to simultaneously detect the presence of apnoea, desaturation, and artefacts in input signals. To achieve this goal, we developed a novel 1D deep neural network architecture that leverages prior knowledge of the information distribution across signals, allowing for the concurrent detection and interpretation of target events. The platform was trained, validated, and tested on data from 86 paediatric patients. We show that our approach outperforms other three approaches from the literature, reaching 92.50% (1.10%), 98.30% (0.43%), and 97.59% (0.28%) balanced classification accuracies for apnoea, desaturation, and artefact, respectively (mean and standard deviation, in brackets). Notably, the REST platform also gives a confidence score as output, highlighting to the doctor the samples that need to be reviewed and further boosting the performances of the other samples. Lastly, based on gradient-weighted class activation mapping (grad-CAM) heatmaps, our platform allows the explanation of the decision process, pointing out the regions of the input signals in which events occur, increasing the reliability of the whole process for a human user.
Similar content being viewed by others
1 Introduction
Sleep apnoea (SA) is a syndrome characterised by a reduction and interruption of airflow either for enhanced flow resistance in the upper airway (obstructive sleep apnoea OSA) or for lack of drive to breathe (central sleep apnoea (CSA))1. Among the two, OSA is far more common (~ 95%) but can be equally dangerous. Indeed, SA can lead to various cardiovascular, neurocognitive, and metabolic comorbidities when left untreated. OSA has a prevalence in the range of 1–5% in the paediatric population. However, it is likely that there are undiagnosed cases due to a lack of resources and sleep laboratories2. Polysomnography (PSG) is the gold standard for diagnosing this disease, consisting of overnight in-lab tests in which multiple cardiorespiratory and sometimes even electrical encephalic characteristics are simultaneously measured1. Currently in the clinical practice, all these signals are visually inspected by specialised doctors to determine the number of flow interruptions (apnoea) or reductions (hypopnoea). The disease severity is then determined by the apnoea hypopnoea index (AHI) computed as the number of apnoea/hypopnoea events per hour of sleep.
Although PSG is the benchmark for SA diagnosis, different options, e.g., nocturnal pulse-oximetry and polygraphy, have recently been proposed to ease the measuring setup and lower the costs3,4. Indeed, it was demonstrated that airflow (AF) and oximetry signals contain most of the information necessary for the identification of respiratory events5.
In recent years, machine learning (ML) and deep learning (DL) have greatly impacted diverse biomedical fields, such as pain detection6recurrent pregnancy loss prediction7and cell phenotyping8,9,10,11,12. In the field of SA, ML, and DL show the potential for automating the analysis of sleep signals for apnoea detection with huge savings in terms of doctors’ and patients’ time.
The main approaches used in the literature for automatic OSA detection involve the use of feature-engineering strategies5,13followed by classical machine learning (ML) algorithms, i.e., logistic regression14,15,16,17neural networks18,19 and ensemble learning16,20. However, feature-engineering approaches require a laborious preliminary analysis of the data to obtain meaningful features. This limitation is solved by DL models, which, through the training process, automatically learn features from signals at the expense of reduced interpretability of the final results. Therefore DL based approaches have been widely used in the analysis of adult sleep data, mainly based on ECG, pulse/oximetry, and AF signals13,21,22,23. Convolutional neural networks (CNNs), recurrent neural networks (RNNs), and their combination have been mainly used in the analysis of electrocardiogram (ECG) signals24,25. Oximetry signals have been used to detect OSA using deep neural networks (DNNs) and CNNs26,27,28,29. Respiratory signals, i.e., oronasal AFs chest/abdominal band, were analysed through CNNs and RNNs to detect SA30,31,32,33,34.
Despite the success of DL in adult sleep scoring, these models are not guaranteed to generalise well to the paediatric population. In recent years, approaches have been proposed to use models pretrained on adult data by fine-tuning them on paediatric data35. Nonetheless, many studies have pointed out the necessity of developing models trained specifically on paediatric data20,36,37. The need for paediatric-specific models is motivated by physiological and developmental differences (such as different respiratory patterns38sleep architecture39), differences in diagnostic criteria (including event scoring criteria and thresholds40), and data quality (mainly sensors displacement due to children’s movement41). As an example, Nazih et al.42 showed that models trained on adult data performed worse on paediatric patients, especially those younger than 13 years. Baumert et al.43 found a performance drop of approximately 10% in the automatic sleep staging when training models on adults and testing them on paediatric data. Therefore, recent literature definitively points out the need for paediatric-specific models.
Recently, most of the works concentrated on DL based approaches for paediatric respiratory events detection, mainly based on the use of CNNs, RNNs, and their combination for OSA detection from AF and oximetry signals21.
One of the main challenges in the automatic analysis and detection of paediatric respiratory events is the abundance of artefacts in the signals. These artefacts are mainly linked to detachment or incorrect positioning of the sensors, often caused by patients’ movement during sleep. Currently, to the authors knowledge, there are few works investigating the possibility to automatically detect artefacts in paediatric polygraphy44,45but none of them solve the problem in the real-world scenario. The automated detection of these artefacts is expected to significantly enhance the overall performance of such a system.
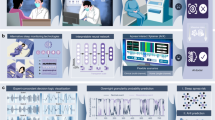
Here, we present REST, an acronym for “aRtificial intelligence based platform for the automatic and simultaneous Explainable detection of apnoea, oxygen deSaturation, and arTefacts in paediatric polygraphy exams”. Figure 1 illustrates the main workflow of the platform. In the first step, data on nasal flow, chest and abdominal band, pulse and saturation, are collected from paediatric patients (Fig. 1a). Signals are then preprocessed, i.e., auto-scaled and windowed (Fig. 1b). Signals are then used as input of a 1D CNN to detect apnoea, desaturation, and artefacts (Fig. 1c). Finally, the platform gives a gradient-based explanation of the obtained results, highlighting the most significative regions in the input signals that led the model to the final decision (Fig. 1d).
REST platform: the first step consists of the data acquisition, i.e., nasal airflow, chest and abdominal band, pulse, and oxygen saturation signals on paediatric patients (a). Then the signals are preprocessed, i.e., signal autoscaling and windowing (b). Signals are then fed in three monodimensional convolutional neural network for the simultaneous detection of apnoea, desaturations, and artefacts events (c). Lastly, an explanation of the decision process is given, highlighting the portions of the input signals that gave the main contribution to the final outcomes (d).
The main contributions of the present work are threefold:
-
1.
We developed the first platform for the simultaneous detection of apnoea, oxygen desaturation, and artefacts in paediatric AF and pulse/oximetry signals, explaining the decision process for each event.
-
2.
We designed and implemented a novel splitting architecture, i.e., dividing the analysis of AF and pulse from the oxygen saturation signals. By exploiting the prior knowledge on the distribution of information between signals, i.e., only saturation signals bring information about the desaturation events, the proposed 1D CNN architecture outperforms the approaches existing in literature.
-
3.
Lastly, the proposed platform provides a confidence score allowing specialised personnel, e.g., doctors, to revise unsure samples further boosting the accuracy of the whole process.
The splitting architecture place this work in the field of physics constrained neural networks. Indeed, the chosen architecture ultimately reduces the number of weights to be trained, by exploiting the prior on information’s distribution, i.e., the information on desaturation events is contained only in oxygen saturation signals. Therefore, the splitting architecture can be seen as a form of regularisation.
The platform was trained and tested on respiratory signals (oronasal AF, chest and abdominal band) pulse (heart rate) and oximetry (oxygen saturation) signals from paediatric patients from the Respiratory Unit of the Bambino Gesù Research Children’s Hospital in Rome. Based on the ability of a novel 1D CNN architecture, taking advantage of the prior information distribution between the signals, REST can detect the presence of apnoea, desaturation, and artefacts reaching 92.50% (1.10%), 98.30% (0.43%), and 97.59% (0.28%) test balanced classification accuracy, respectively (mean and standard deviation, in brackets, over 100 runs). Furthermore, we demonstrated the superiority of our approach with respect to a standard 1D CNN architecture21,46 and with respect to other approaches found in literature, i.e., bidimensional CNNs (2D CNNs)47 and 2D CNN combined with recurrent neural networks (RNNs)48.
In addition to the standard operating principle, REST also returns a confidence score, which can be used to estimate the model’s confidence in the classification of the sample. This characteristic allows the doctor to revise only a limited number of signals, improving the overall performance in terms of classification accuracies. Leveraging the power of recently developed explainability techniques49REST also gives an insight into the process that led the model to make a specific decision enhancing its reliability from a human user point of view.
Materials and methods
Subjects and signals
Eighty-six patients (51 male and 35 female; median age 7.5 years), referred to the Respiratory Unit of the Bambino Gesù Research Children’s Hospital in Rome, were enrolled for this study.
Data were acquired in the past years by the medical staff. Written informed consent was signed by the patients or by their legal tutor. All the data were provided to the authors anonymously, without any sensitive information about patients and their pathology. All methods were performed in accordance with the relevant guidelines and regulations (Declaration of Helsinki). The study received approval from the institutional review board of the Bioethical Committee of the Bambino Gesù Research Children’s Hospital in Rome (protocol no. 1934_OPBG_2019).
The sleep data, consisting of AF and pulse/oximetry signals, were randomly split into train (50%) and test (50%) (data from 43 patients for each set). The test partitioning was performed subject-wise to avoid the same subject being represented in both sets at the same time. The train set was further randomly split into train (2/3), i.e., data actually used for training the model, and validation (1/3), i.e., data used to assess the performance on an independent set and choose the optimal hyperparameters. The use of the validation set for the hyperparameters tuning and the selection of best-performing models, leaving the test set untouched during training and validation, is widely regarded as a best practice, allowing an unbiased estimate of the model performances on unseen data, especially when the test is completely independent50,51,52.
AF signals, i.e., nasal airflow, chest band, and abdominal band, were measured at a sampling frequency of 20 Hz, while heart rate and saturation were acquired at 1 Hz and then upsampled to 20 Hz, with a linear interpolation, to uniform the sampling rate between the different signals. Different signal interpolation methodologies were considered, namely cubic and nearest neighbour, but the linear method outperformed them in terms of balanced classification accuracies (Table S1).
Each polygraphy exam was divided into disjoint temporal segments (signal windowing) with a length of 512 samples (25.6 s). The windowed signals were then considered as samples. Three labels were then assigned by the medical staff to each sample to specify whether or not artefact, desaturation, and apnoea were present in the sample.
Based on training set statistics, data were auto-scaled, i.e., mean subtracted and normalised by the standard deviation53.
Model architecture and training
The neural network architecture is schematically represented in Fig. 2. REST model consists of two separated 1D CNN54 one for apnoea and the other for desaturation classification. The last feature layers, the output of the flattening layer, are then combined to detect the presence of artefacts. Apnoea and desaturation models share similar architectures except for input data dimensions, i.e. different number of signals, and for the number of filters learned during training. Specifically, apnoea model receives as input nasal flow, chest and abdominal band, and heart rate (pulse), while desaturation model receives as input only the oxygen saturation. Both the neural networks are constituted by a sequence of three blocks followed by a max pooling layer with input dimension of 2 × 1 and a stride of 2. The blocks are constituted concatenating three layers, i.e., 1D convolution, batch normalisation, and rectified linear units (ReLU) activation, for two times. Each convolution block learns an increasing number of filters (N, 2 N, 4 N for the apnoea network and M, 2 M, 4 M for the desaturation network). The output of the fourth block (learning 0.5 N or 0.5 M filters for apnoea or desaturation network respectively) is then flattened. The last layer consists in a fully connected layer with softmax activation function to ensure a probability-like output. Each sample for every task is then assigned to the class with highest posterior probability, from here on, confidence score. Details about CNN parameters are given in the Supplementary materials Tables S2-S4, including number, dimensions, and strides of the filters for the three models, i.e., apnoea, desaturation, and artefact classification. The loss functions used, one for each task, consist of the summation of the categorical cross-entropy and a L1 loss over the weights of the network, in formula:
where the \(\:Loss\) is the total loss, \(\:{N}_{s}\) is the total number of samples in the batch, \(\:{N}_{c}\) is the total number of classes, \(\:{y}_{ij}\) is the class label for the class \(\:j\) and sample \(\:i\:\)in one-hot-encoding, \(\:{p}_{ij}\) is the output probability of the network for the class \(\:j\) and sample \(\:i\), \(\underline{\varvec{\theta }}\) is the vector of network’s parameters, \(\lambda\) is the regularisation strength, \(\:M\) is the number of network’s parameters, and \(\theta _{k}\) is the kth parameter of the network.
It can be showed that minimising the first term (the categorical cross-entropy) on the right hand side of the Eq. (1), with respect to the networks parameters \(\underline{\varvec{\theta }}\) corresponds to minimising the Kullback-Leibler divergence between data classes and the predicted output probability55. The \(L1\) regularisation term consists of the sum of the absolute value of the weights of the network. This regularisation promotes sparse solutions with respect to network parameters enhancing the generalisation ability of the resulting model. The mutual importance between the two terms in the loss definition is regulated by the strength\(\lambda\), heuristically set to \(\lambda\,=\,{10}^{-3}\). The loss function was minimised using Adam optimiser56 with learning rate set to \(\:lr=\:{10}^{-4}\), and the moving average parameters set to \(\:{\beta\:}_{1}=0.9\) and \(\:{\beta\:}_{2}=0.999\).
REST model architecture. 1D CNN architecture for apnoea, desaturation, and artefact models.
The training set consisting of the AF and pulse/oximetry signals from 43 patients was randomly split into training (2/3) and validation (1/3). The training set was used to optimize the network parameters by minimizing the total loss function. The validation set served to heuristically guide the selection of hyperparameters, including the learning rate, regularization strength, and the number and dimensions of convolutional filters. The validation set was also used to select the best-performing network, i.e., selecting the model with the lowest validation loss, to evaluate the performances on the test set.
Comparative models
Three other CNN architectures were implemented to compare their performance with the proposed REST platform. The first comparative model is the standard 1D CNN. The difference with the REST architecture consists in the fact that all signals are fed in the same network (apnoea and desaturation networks coincide). The second comparative model is taken from47 and consists of a 2D CNN. Basically, all the signals together are interpreted as an image where the columns represent time and the rows represent the signals. The third comparative model is taken from48 and combines 2D CNN with LSTM. All these kinds of models from the literature do not exploit the prior knowledge about the information distribution in the signals, i.e., only oxygen saturation has information about desaturation events, and we believe that this is the main superiority of the proposed REST platform. All the models were trained with the same strategy and datasets proposed for the REST platform.
Model explainability using grad-CAM
We applied gradient-weighted class activation mapping (grad-CAM) to the trained models to get an explanation of the decision process. Grad-CAM is a model-specific explainability method based on gradients originally designed for CNNs aiming to generate feature attribution maps that identify the regions of the input signals that generate a certain output. The gradient-based attribution maps can be considered as heatmaps, in the sense that they highlight significative regions of the input signals, and can be obtained from all the convolution layers. The attribution maps of the positive class (apnoea, desaturation, or artefact) were computed following the algorithm of the original paper49. First, the gradient between the \(\:c\) class output and the feature maps resulting from the output of the last convolutional layer were calculated: in formula \(\frac{{\partial y^{c} }}{{\partial x_{j}^{i} }}\) where \({y}^{c}\) is the output of class c and \({x}_{j}^{i}\) is the activation of the neuron in position \(j\) of the feature map \(\:i\). These gradients are then averaged over the signal dimension to obtain the weights \({\alpha}_{i}^{c}\) as follows:
where N here is the dimension of each 1D feature map. The weights \({\alpha}_{i}^{c}\) can be thought as a partial linearisation of the nonlinear neural network from the convolutional layer chosen to the output and represents the importance of the single feature map with respect to the selected output class. Therefore, \({\alpha}_{i}^{c}\) can be seen as the average sensitivity of the class output \(c\) to that specific feature map \(i\). The heatmaps for class c are then obtained by a weighted combination of the feature maps \(\underline{x} ^{i}\) with \({\alpha}_{i}^{c}\) as weights followed by a ReLU activation.
Since each heatmap has the dimension of the feature maps of the convolutional layer chosen, we linearly upsampled it to the dimension of the input signals to allow proper visualisation. We selected the last convolutional layer to extract the heatmaps because it grants us the lowest possible approximation error between the actual neural network function and the grad-CAM linear approximation. The grad-CAM heatmaps have higher values in the regions of the input signals with high influence in determining the final output, i.e., regions containing the apnoea, desaturation, or artefact event.
The grad-CAM heatmaps were used to segment the apnoea, desaturation, and artefacts events. The segmentation was obtained simply by thresholding the explainability heatmap signal. The results were evaluated in terms of mean absolute error (Figure S1) defined as follows:
where \(\:{y}_{j}\) is the ground truth event signal, \(\:{\widehat{y}}_{j}\) is the predicted segmentation and \(\:N\) is the number of time points in the signal. Segmentation performances were evaluated also in terms of Apnoea Hypopnoea Index (real vs. predicted), defined here for each signal (512 samples) as the ratio between the apnoea and the total observation time (Figure S2).
Implementation details
All simulations including training, test, and explainability were conducted on a standard desktop computer equipped with intel i7 11th generation CPU and a 12 GB NVIDIA GeForce RTX 3060 dedicated GPU. All the models were developed in python 3.8.8. The main libraries used are tensorflow-gpu and numpy. The model weights require few memory space (49 KB for apnoea and desaturation, 534 KB for artefacts) to be saved. The model test sample prediction is relatively fast, processing 1 sample every 0.02 s on average.
Results
In this section, we present the results of the REST platform for three specific tasks: apnoea, desaturation, and artefact classification. The performances of the models are evaluated using confusion matrices, balanced classification accuracies, precision, recall, and F1-score metrics.
Each classification model was trained using the training dataset, composed of signals from 43 patients. After training, the models were validated on a separate validation set, as previously described. The model that demonstrated the best performance on the validation set was then selected for inference on the test set, which comprised signals from additional 43 independent subjects. This cross-validation strategy assures a fair test of the generalisation ability of the models employed.
Classification results for apnoea, desaturation, and artefacts
For the three tasks, i.e., apnoea, desaturation, and artefact detection, we obtained, on a specific randomly sampled run, the test set balanced classification accuracies of 92.2%, 99.0% and 97.6%, respectively. To better understand the discrimination abilities of the single models, we reported in Fig. 3 the confusion matrices for the three tasks. Notably, all classes show high accuracy values in three tasks. Table 1 reports precision, recall, and F1-score for each task (mean and standard deviation, in brackets, over 100 runs). The high values of all the performance metrics prove the ability of the REST platform to successfully classify independent test samples.
Balanced classification accuracy results. Confusion matrices were obtained from classification of the test set from the best model selected based on the validation set in a single run for the three tasks: apnoea, desaturation, and artefacts. The green cells contain correctly classified samples (on the first diagonal of the matrix). Pink cells include wrongly classified samples. In the last column, green values show single class accuracies, while the red values show the error rate. The cell on the bottom left reports average accuracy. Values in the last row report single-class precision in green and associated errors in red.
Comparison with other state-of-the-art methodologies
We investigated the effectiveness of the REST platform by comparing the results with the performances of the other three state-of-the-art methodologies. The first method used for comparison is a standard 1D CNN, i.e., without the separated apnoea and desaturation models. The second model, taken from47 is a 2D CNN where signals are concatenated as rows of an image. The last model used for comparison48 combines 2D CNN and LSTM (here referred to as 2D CNN + LSTM). All the models used for comparison do not employ the split architecture proposed in this paper; instead, they combine all the signals from the initial layers together. Table 2 reports the balanced classification accuracies obtained by the REST platform and by the other three comparative approaches (mean and standard deviation, in brackets, over 100 runs). The same signal preprocessing strategies, train-validation-test sets, hyperparameters, and model selection approaches were employed for the proposed approach and the comparative methods. The results show striking superiority in the performance of the proposed methodology in all the tasks, particularly in the desaturation classification. This is probably due to the fact that REST desaturation model receives in input the only signal carrying information about the desaturation. Instead, the other models also receive other signals enhancing model capacity without increasing the exploitable information, ultimately resulting in overfitting. In addition, the main difference between the three models used for comparison and the REST platform lies in the “split architecture”. Therefore, the authors impute these superior performances to the introduction in the model’s architecture of prior knowledge of the separated information between the tasks (two separated models for apnoea and desaturation), making the REST platform more data efficient and generalisable.
Discussion
We developed a novel platform, REST, able to detect the presence of apnoea, desaturation, and artefacts from AF and pulse-oximetry signals with a temporal length of around 25 s in paediatric patients. Our approach is built on the recent advancements in DL technology, which have demonstrated the ability of CNNs to extract significant information in strictly related tasks21,57. Specifically, our approach exploited a diverse 1D CNN architecture and incorporates prior knowledge on the distribution of information between the signals useful for the various tasks, i.e., apnoea and desaturation. This architecture acts as a form of regularisation, decreasing the number of parameters to be trained while maintaining the capacity to discriminate the classes. This choice leads to realising two distinct models for desaturation and apnoea. Results show that our model is able to reach optimal classification results (Fig. 3; Table 1) and outperforms other literature approaches (Table 2). Another advantage of the proposed methodology for the artefact detection lies in the data-driven training, rather than relying on hard coded rules. This gives flexibility to the proposed strategy allowing easy generalisation to other datasets with different types of artefacts.
In this study we considered all the types of apnoea, i.e., obstructive, central, and mixed apnoea, joined in the same class. This decision was due to insufficient data in the two less-represented categories, central and mixed apnoea. The intraclass variability likely contributed to the lower performance of our approach in apnoea detection compared to desaturation and artefact detection.
The proposed platform has the great advantage of giving as output not just whether or not a certain sample belongs to a class but also a confidence score, a number between zero and one, which can be interpreted as the sample’s posterior probability of belonging to a class. Here, we show (Fig. 4) that by selecting only samples with a confidence score higher than a threshold the platform achieves higher performances. After measuring the performances with the threshold varying between 0.6 and 0.95 (with step 0.05), we selected 0.85 as a compromise between the samples retained and the accuracy achieved (Fig. 4) reaching 94.7% (1.1%), 99.4% (0.3%), 99.6% (0.1%) for apnoea, desaturation, and artefacts respectively (mean and standard deviation, in brackets, over 100 runs). These results are obtained by retaining the 82.6% (1.2%), 93.8% (0.8%), and 92.7% (0.4%) of data available in the test set from apnoea, desaturation, and artefact data, respectively (mean and standard deviation, in brackets, over 100 runs). As a collateral result, we can observe that most of the samples with confidence scores below the threshold are incorrectly classified. Therefore, this analysis shows a hidden yet striking potential of our approach: the REST platform could be used to reduce the amount of work to be performed by the doctors (17.4% (1.2%), 6.2% (0.8%), and 7.3% (0.4%) for the three tasks), highlighting the “uncertain” samples that need to be reviewed while improving the overall performances on the test set.
Sensitivity analysis of balanced classification accuracy (a) and ratio of samples selected (b) to the threshold used for samples selection. The accuracies were obtained after the selection of samples with probability out of the softmax layer higher than a threshold spanning between 0.6 and 0.95 (with step 0.05) (a). Ratio of samples retained for classification by the model (b). The samples with a probability lower than the threshold can be reported to specialised doctors for an accurate reanalysis.
Leveraging recent advancements in the field49 the REST platform gives an explanation of the decision process that led to the final output. Figure 5 shows grad-CAM mono-dimensional heatmaps obtained from the last convolutional layer of the network, as described in the methodology section, along with the input signals. Although it involves only a few derivatives and linear combinations, grad-CAM allows us to understand which portions of the input signals have contributed to the final class attribution. Therefore, grad-CAM equips the REST platform with a human-friendly explanation of the decision process. Figure 5 shows a case example in which the network is able to correctly identify discriminative regions of the input signals in which the event happened (apnoea, desaturation, and artefact in Fig. 5).
Figure 5a shows the input signals used by the apnoea detection system, which include nasal airflow, heart rate, and signals from the abdominal and chest bands, represented in green, cyan, blue, and purple, respectively. The corresponding one-dimensional grad-CAM heatmap is also illustrated in red, highlighting the regions of interest in the sensor signals during the experiment. Figure 5b shows the oxygen saturation signal (in blue) used as input for the desaturation detection model, alongside its corresponding grad-CAM heatmap (in red). Figure 5c shows the nasal flow, abdominal and chest band, heart rate signals represented in green, blue, purple, and cyan respectively. The corresponding one-dimensional grad-CAM heatmap, illustrated in red, highlights the region containing artefacts in the signals.
Grad-CAM explanation of the REST’s decision process. Plots of input apnoea’s and desaturation’s model signals and associated grad-CAM mono-dimensional heatmap for explainability for a case example from the test set. The heatmap shows higher values where the events occur (apnoea (a), desaturation (b), and artefacts (c)). The local maxima in the grad-CAM heatmaps highlight portions of signals that the model identifies as relevant events. Signals are rescaled between − 1 and 1 to allow better visualisation.
In all figures, the peaks of the grad-CAM heatmap signals align with the areas in the input sensor signals where the apnoea, desaturation, and artefacts events, respectively, are detected in Fig. 5a–c. These maxima highlight the portions of the signals that the models identify as critical events.
In order to give a statistical evaluation of the performances of the explanation, we leveraged grad-CAM heatmaps to segment the apnoea, desaturation, and artefacts events, simply by thresholding the heatmaps signals. Notably, we found out that, even if the model was not trained for the event segmentation task, it was able to reach low mean absolute errors in the event segmentation tasks and good Apnoea Hypopnoea Index (AHI) prediction, both in test (Figure S1-2). These results suggest that the split-architecture proposed not only increased the event detection accuracy, but it also enabled the model to spontaneously learn the information of event localisation in the data.
Conclusions
This paper introduces a platform (REST) for the automatic detection of apnoea, desaturation, and artefacts in paediatric AF and pulse/oximetry data. The novelty of the proposed approach resides in several fundamental aspects.
REST is the first platform for the automatic and simultaneous detection of apnoea, desaturation, and artefacts from AF and pulse/oximetry data in paediatric patients. The concurrent detection of these three events is pivotal to ensure the possibility of a detailed diagnosis from specialised doctors. On the other hand, the use of the AF and pulse/oximetry signals allows this platform to be used on data taken directly from patients’ homes (telemedicine). Therefore, the proposed approach could reduce the pressure on hospitals, which is a crucial aspect, as highlighted by the pandemic experience.
The proposed platform introduces a novel architecture splitting the models between desaturation and apnoea incorporating prior knowledge about the distribution of information between signals. This choice enables REST to outperform, on this dataset, other approaches found in literature, reaching 92.50% (1.10%), 98.30% (0.43%), and 97.59% (0.28%) balanced accuracy for apnoea, desaturation, and artefact classification, respectively (mean and standard deviation, in brackets, over 100 runs).
Our approach introduces a confidence score, allowing for the selection of samples that need to be revised (confidence score below the threshold) by the doctor while boosting the performances on the remaining samples (confidence score over the threshold). Lastly, our platform allows an explanation of the results outputting the grad-CAM mono-dimensional heatmaps highlighting the regions of the input signals where the events occur.
As already stated, in this work, we merged all three types of apnoea (obstructive, central, and mixed) into one class. Nonetheless, apnoea detection represents a step forward in clinical practice, allowing specialised doctors to distinguish between the three types of apnoea and significantly reducing the amount of work to be performed. However, future studies will be directed toward this objective.
The proposed model can generalise well on a completely independent set of data, i.e., different patients, even if the dimension of the training set is relatively small. This is also due to the innovative architecture proposed that acts as a form of regularisation, reducing the risk of overfitting. The dimension of the dataset available in this study gave us the opportunity to highlight the robustness of the proposed platform when compared to other approaches present in the literature. We acknowledge the need of future studies validating the proposed methodology on large dataset, eventually investigating the possibility of training the models on artificially generated data.
The platform outputs a confidence score, allowing the selection of uncertain samples that need revision from specialised doctors while boosting performances on the remaining data. Lastly, the proposed approach gives an explanation of the decision process, enhancing the information extracted and the reliability of the whole platform for human users.
Furthermore, the small memory consumption for the weight storage (hundreds of KBs) and the rapid processing time (around 0.02 s per sample) (see “Materials and Methods”, Section “Implementation details”) make our approach suitable for implementation and real-world application even on standard computers.
The high performances in detecting apnoea and desaturation, combined with the automatic detection of experimental artefacts, obtained by the REST platform, prefigure its usage in a real-world scenario.
Data availability
Data supporting the findings of this study will be provided by the corresponding author under reasonable request.
References
Li, Z., Celestin, J. & Lockey, R. F. Pediatric sleep apnea syndrome: an update. J. Allergy Clin. Immunol. Pract. 4, 852–861 (2016).
Marcus, C. L. et al. Diagnosis and management of childhood obstructive sleep apnea syndrome. Pediatrics 130, e714–e755 (2012).
Tan, H. L., Gozal, D., Ramirez, H. M., Bandla, H. P. R. & Kheirandish-Gozal, L. Overnight polysomnography versus respiratory polygraphy in the diagnosis of pediatric obstructive sleep apnea. Sleep 37, 255–260 (2014).
Tan, H. L., Kheirandish-Gozal, L. & Gozal, D. Pediatric home sleep apnea testing slowly getting there! Chest 148, 1382–1395 (2015).
Bertoni, D. & Isaiah, A. Towards patient-centered diagnosis of pediatric obstructive sleep apnea—a review of biomedical engineering strategies. Expert Rev. Med. Devices. 16, 617–629 (2019).
Casti, P. et al. Metrological characterization of a pain detection system based on transfer entropy of facial landmarks. IEEE Trans. Instrum. Meas. 70, 1–8 (2021).
Bruno, V. et al. Machine learning (ML) based-method applied in recurrent pregnancy loss (RPL) patients diagnostic work-up: a potential innovation in common clinical practice. Sci. Rep. 10, 1–12 (2020).
Mencattini, A. et al. Deep-Manager: a versatile tool for optimal feature selection in live-cell imaging analysis. Commun. Biol. 6, 241 (2023).
Comes, M. C. et al. A camera sensors-based system to study drug effects on in vitro motility: the case of PC-3 prostate cancer cells. Sensors 20, 1531 (2020).
Mencattini, A. et al. NeuriTES. Monitoring neurite changes through transfer entropy and semantic segmentation in bright-field time-lapse microscopy. Patterns 2, (2021).
D’Orazio, M. et al. Machine learning phenomics (MLP) combining deep learning with time-lapse-microscopy for monitoring colorectal adenocarcinoma cells gene expression and drug-response. Sci Rep 12, (2022).
D’Orazio, M. et al. Deciphering cancer cell behavior from motility and shape features: peer prediction and dynamic selection to support cancer diagnosis and therapy. Front. Oncol. 10, 1–11 (2020).
Gutiérrez-Tobal, G. C. et al. Reliability of machine learning to diagnose pediatric obstructive sleep apnea: systematic review and meta-analysis. Pediatr. Pulmonol. 57, 1931–1943 (2022).
Alvarez, D. et al. Symbolic dynamics to enhance diagnostic ability of portable oximetry from the phone oximeter in the detection of paediatric sleep Apnoea. Physiol. Meas. 39, 104002 (2018).
Wu, D., Li, X., Guo, X., Qin, J. & Li, S. A simple diagnostic scale based on the analysis and screening of clinical parameters in paediatric obstructive sleep Apnoea hypopnea syndrome. J. Laryngol Otol. 131, 363–367 (2017).
Calderón, J. M., Álvarez-Pitti, J., Cuenca, I., Ponce, F. & Redon, P. Development of a minimally invasive screening tool to identify obese pediatric population at risk of obstructive sleep apnea/hypopnea syndrome. Bioengineering 7, 1–13 (2020).
Barroso-García, V. et al. Bispectral analysis of overnight airflow to improve the pediatric sleep apnea diagnosis. Comput. Biol. Med. 129, 104167 (2021).
Hornero, R. et al. Nocturnal oximetry-based evaluation of habitually snoring children. Am. J. Respir Crit. Care Med. 196, 1591–1598 (2017).
Vaquerizo-Villar, F. et al. Detrended fluctuation analysis of the oximetry signal to assist in paediatric sleep apnoea-hypopnoea syndrome diagnosis. Physiol. Meas. 39, 114006 (2018).
Jiménez-García, J. et al. Assessment of airflow and oximetry signals to detect pediatric sleep Apnea-Hypopnea syndrome using adaboost. Entropy 22, 670 (2020).
Vaquerizo-villar, F. et al. A convolutional neural network to classify sleep stages in pediatric sleep apnea from pulse oximetry signals. 108–113 (2022). https://doi.org/10.1109/MELECON53508.2022.9842917
Mostafa, S. S., Mendonça, F. & Ravelo-García, A. G. Morgado-Dias, F. A systematic review of detecting sleep apnea using deep learning. Sens. (Switzerland). 19, 1–26 (2019).
Peng, D. et al. A bimodal feature fusion convolutional neural network for detecting obstructive sleep apnea/hypopnea from nasal airflow and oximetry signals. Artif. Intell. Med. 150, 102808 (2024).
Zarei, A., Beheshti, H. & Asl, B. M. Detection of sleep apnea using deep neural networks and single-lead ECG signals. Biomed. Signal. Process. Control. 71, 103125 (2022).
Dey, D., Chaudhuri, S. & Munshi, S. Obstructive sleep Apnoea detection using convolutional neural network based deep learning framework. Biomed. Eng. Lett. 8, 95–100 (2018).
Nikkonen, S., Afara, I. O., Leppänen, T. & Töyräs, J. Artificial neural network analysis of the oxygen saturation signal enables accurate diagnostics of sleep apnea. 1–9 (2019).
Leino, A. et al. Neural network analysis of nocturnal SpO2 signal enables easy screening of sleep apnea in patients with acute cerebrovascular disease. Sleep. Med. 79, 71–78 (2021).
Mostafa, S. S., Mendonca, F., Ravelo-Garcia, A. G., Julia-Serda, G. & Morgado-Dias, F. Multi-objective hyperparameter optimization of convolutional neural network for obstructive sleep apnea detection. IEEE Access. 8, 129586–129599 (2020).
Mostafa, S. S., Baptista, D., Ravelo-García, A. G., Juliá-Serdá, G. & Morgado-Dias, F. Greedy based convolutional neural network optimization for detecting apnea. Comput. Methods Programs Biomed. 197, 105640 (2020).
Haidar, R., Koprinska, I. & Jeffries, B. Sleep apnea event detection from nasal airflow using convolutional neural networks. Lect Notes Comput. Sci. (including Subser. Lect Notes Artif. Intell. Lect Notes Bioinformatics). 10638 LNCS, 819–827 (2017).
Choi, S. H. et al. Real-time apnea-hypopnea event detection during sleep by convolutional neural networks. Comput. Biol. Med. 100, 123–131 (2018).
Yue, H. et al. Deep learning for diagnosis and classification of obstructive sleep apnea: A nasal airflow-based multi-resolution residual network. Nat. Sci. Sleep. 13, 361–373 (2021).
Elmoaqet, H., Eid, M., Glos, M., Ryalat, M. & Penzel, T. Deep recurrent neural networks for automatic detection of sleep apnea from single channel respiration signals. Sens. (Switzerland). 20, 1–19 (2020).
Kemidi, M., Marur, D. R. & Reddy, C. V. K. Obstructive sleep apnea detection using optimized Bi-LSTM with random forest based exhaustive feature selector. Multimed Tools Appl. 1–23. https://doi.org/10.1007/s11042-024-18837-1 (2024).
Haimov, S., Tabakhov, A., Tauman, R. & Behar, J. A. Deep learning for pediatric sleep staging from photoplethysmography: A transfer learning approach from adults to children. IEEE Trans. Biomed. Eng. 72, 760–767 (2024).
Crowson, M. G. et al. Paediatric sleep apnea event prediction using nasal air pressure and machine learning. J. Sleep. Res. 32, 1–7 (2023).
Vaquerizo-Villar, F. et al. Automatic assessment of pediatric sleep apnea severity using overnight oximetry and convolutional neural networks. Proc. Annu. Int. Conf. IEEE Eng. Med. Biol. Soc. EMBS, 633–636 (2020).
Fleming, S. et al. Normal ranges of heart rate and respiratory rate in children from birth to 18 years of age: A systematic review of observational studies. Lancet 377, 1011–1018 (2011).
Altevogt, B. M. & Harvey, R. C. Sleep disorders and sleep deprivation: An unmet public health problem. (2006).
Berry, R. B. et al. Rules for scoring respiratory events in sleep: update of the 2007 AASM manual for the scoring of sleep and associated events. J. Clin. Sleep. Med. 8, 597–619 (2012).
Marcus, C. L. et al. Feasibility of comprehensive, unattended ambulatory. J. Clin. Sleep. Med. 10, 913–918 (2014).
Nazih, W., Shahin, M., Eldesouki, M. I. & Ahmed, B. Influence of channel selection and subject’s age on the performance of the single channel EEG-based automatic sleep staging algorithms. Sensors 23, (2023).
Baumert, M., Hartmann, S. & Phan, H. Automatic sleep staging for the young and the old–Evaluating age bias in deep learning. Sleep. Med. 107, 18–25 (2023).
Poets, C. F. & Stebbens, V. A. Detection of movement artifact in recorded pulse oximeter saturation. Eur. J. Pediatr. 156, 808–811 (1997).
Lynn Kern, K., Katherine, M., Pumpki, B., Mohini, S., Stephen, C. & V. & Intervention for non-verbal and minimally-verbal individuals with autism: A systematic review. Int. J. Pediatr. Res. 5, 1–7 (2019).
Erdenebayar, U., Ji, Y., Park, J., Yeon, E. & Lee, K. Deep learning approaches for automatic detection of sleep apnea events from an electrocardiogram. Comput. Methods Programs Biomed. 180, 105001 (2019).
Jiménez-García, J. et al. A 2D convolutional neural network to detect sleep apnea in children using airflow and oximetry. Comput. Biol. Med. 147, 105784 (2022).
Jiménez-García, J. et al. An explainable deep-learning architecture for pediatric sleep apnea identification from overnight airflow and oximetry signals. Biomed. Signal. Process. Control. 87, 105490 (2024).
Selvaraju, R. R. et al. Grad-CAM: Visual explanations from deep networks via gradient-based localization. In IEEE international conference on computer vision (2017).
Xu, Y. & Goodacre, R. On splitting training and validation set: A comparative study of cross-validation, bootstrap and systematic sampling for estimating the generalization performance of supervised learning. J. Anal. Test. 2, 249–262 (2018).
Westphal, M. & Brannath, W. Evaluation of multiple prediction models: A novel view on model selection and performance assessment. Stat. Methods Med. Res. 29, 1728–1745 (2020).
Raschka, S. Model evaluation, model selection, and algorithm selection in machine learning. (2018).
Despagne, F. Neural networks in multivariate calibration. Analyst 123, 157R–178R (1998).
Kiranyaz, S. et al. 1D convolutional neural networks and applications: A survey. Mech. Syst. Signal. Process. 151, 107398 (2021).
Butcher, J. C. Numerical Methods for Ordinary Differential Equations (Wiley, 2003).
Kingma, D. P., Ba, J. & Adam A method for stochastic optimization. in International Conference on Learning Representations (2015).
Vaquerizo-villar, F. et al. A convolutional neural network architecture to enhance oximetry ability to diagnose pediatric obstructive sleep apnea. IEEE J. Biomed. Health Inf. 25, 2906–2916 (2021).
Acknowledgements
R.C. acknowledges funding from Italian Ministry of Health in the project 5 × 1000, code 5 M-2019-23680191.
Author information
Authors and Affiliations
Contributions
M.D., A.M, and E.M. wrote the original draft. M.D., E.V., R.C., E.M. conceptualization. M.D., E.V., J.F., G.A., G.C., P.C., A.M., data curation. M.D., E.M. formal analysis and visualization. M.D., E.V., G.A., G.C., P.C., investigation. M.D., J.F., A.M., E.M., methodology. M.D. software and validation. E.V., M.P., R.C., E.M. resources. R.C., E.M. supervision. M.R., R.C., E.M. funding acquisition. E.V., M.P., R.C. data collection and treatment. E.M. project administration. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
D’Orazio, M., Verrillo, E., Filippi, J. et al. Artificial intelligence based platform for the automatic and simultaneous explainable detection of apnoea, oxygen desaturation, and artefacts in paediatric polygraphy exams (REST). Sci Rep 15, 34024 (2025). https://doi.org/10.1038/s41598-025-13630-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-13630-y