Abstract
Cervical cancer is a leading cause of death from malignant tumors in women, and accurate evaluation of occult lymph node metastasis (OLNM) is crucial for optimal treatment. This study aimed to develop several predictive models—including Clinical model, Radiomics models (RD), Deep Learning models (DL), Radiomics–Deep Learning fusion models (RD-DL), and a Clinical–RD-DL combined model—for assessing the risk of OLNM in cervical cancer patients.The study included 130 patients from Center 1 (training set) and 55 from Center 2 (test set). Clinical data and imaging sequences (T1, T2, and DWI) were used to extract features for model construction. Model performance was assessed using the DeLong test, and SHAP analysis was used to examine feature contributions. Results showed that both the RD-combined (AUC = 0.803) and DL-combined (AUC = 0.818) models outperformed single-sequence models as well as the standalone Clinical model (AUC = 0.702). The RD-DL model yielded the highest performance, achieving an AUC of 0.981 in the training set and 0.903 in the test set. Notably, integrating clinical variables did not further improve predictive performance; the Clinical–RD-DL model performed comparably to the RD-DL model. SHAP analysis showed that deep learning features had the greatest impact on model predictions. Both RD and DL models effectively predict OLNM, with the RD-DL model offering superior performance. These findings provide a rapid, non-invasive clinical prediction method.
Similar content being viewed by others
Introduction
Cervical cancer ranks among the leading causes of death from malignant tumors in women, with its incidence rate surpassed only by breast, lung, and colon cancers. This disease poses significant threats to both the physical and mental health of women. Early detection, diagnosis, and treatment in clinical practice can effectively reduce cervical cancer mortality rates1. According to the 2019 Clinical Practice Guidelines for Cervical Cancer published by the National Comprehensive Cancer Network, surgery is the primary treatment for early-stage patients with cervical cancer2. In cases of lymph node metastasis (LNM), radiotherapy and chemotherapy are recommended as first-line treatments3,4. Therefore, accurate preoperative assessment of LNM in cervical cancer is crucial for treatment decision-making and prognosis. Currently, preoperative assessment of LNM in cervical cancer primarily relies on computed tomography (CT) or magnetic resonance imaging (MRI) to evaluate lymph node morphology. The most common method involves measuring the size of the lymph nodes (transverse diameter ≥ 10 mm, ) or assessing necrosis. This approach largely depends on the radiologist’s experience and exhibits relatively low accuracy. Additionally, many clinicians have reported instances where preoperative imaging fails to detect LNM, which is only confirmed postoperatively and is referred to as occult lymph node metastasis (OLNM)5. This issue necessitates treatment plan adjustments for 20–40% of patients after surgery, leading to a significant waste of medical resources. Therefore, there is an urgent need for a practical and efficient solution to improve the preoperative prediction of OLNM, aiming to optimize treatment plans for patients with cervical cancer.
Lambin6introduced the concept of radiomics, drawing from genomics and proteomics, proposing that medical images could be converted into high-dimensional data. This approach involves extracting high-throughput imaging features such as surface features, first-order statistics, and texture features to reflect tumor heterogeneity. With advancements in artificial intelligence, numerous radiomics applications have emerged in cervical cancer7,8,9,10. Radiomics has demonstrated unique value in quantifying tumor heterogeneity; however, its application still presents certain limitations. This method heavily relies on manual delineation of regions of interest by physicians, a process that is not only time-consuming and labor-intensive but also prone to introducing subjective biases, thereby limiting the reproducibility of the extracted features. In contrast, deep learning leverages large-scale pre-trained knowledge to operate efficiently even under small sample conditions. Deep transfer learning methods based on convolutional neural networks (CNNs) can automatically extract multi-level semantic features from regions of interest (ROIs), significantly reducing human intervention. These characteristics are particularly crucial in cross-center and multi-device scenarios11. Deep learning has been widely applied in cervical cancer research12,13,14,15.
To the best of our knowledge, only a few studies have integrated radiomics with 3D deep learning to predict OLNM in cervical cancer. This study aimed to construct and validate a prediction model based on radiomics and 3D deep learning to assess the risk of OLNM in cervical cancer patients. To further enhance the interpretability and clinical applicability of the model, we plan to integrate the SHAP method. Through this integration, the decision-making process of the model will be more clearly explained and visualized, making its application more intuitive and comprehensible, thereby further increasing its value in clinical practice.
Results
Patient characteristics
We collected clinical data from patients, including age, FIGO stage, and maximum tumor diameter measured by MRI (scored as 1 point for < 2 cm, 2 points for between 2 and 4 cm, and 3 points for > 4 cm), along with postoperative pathological data such as pathological type, lymph vascular space invasion (LVSI), deep stromal invasion, and histological grade (Table 1).
In both datasets, there were no statistically significant differences in average age, maximum tumor diameter, pathological type, and degree of differentiation between patients with and without OLNM (all P > 0.05). However, across all data, FIGO stage and deep stromal invasion were significantly associated with OLNM (P = 0.011, P < 0.001). This association was also observed in the training set (P = 0.003, P < 0.001). In contrast, in the test set, no significant differences were found in the FIGO stage and deep stromal invasion with respect to OLNM (P = 0.135, P = 0.067, respectively). Notably, there was a significant difference between positive LVSI and OLNM (P < 0.001).
RD model
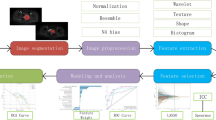
Radiomic features were extracted from T1-weighted sequences and subsequently processed through a series of analytical steps, including normalization, clustering analysis, Pearson correlation assessment, and feature selection using LASSO regression (Fig. 1). This process yielded four optimal radiomic features. Five machine learning models—GB, LGBM, LR, MLP, and RF were then developed and evaluated. Performance metrics included the area under the curve (AUC), sensitivity, specificity, F1-score, and their corresponding 95% confidence intervals (CI). The optimal model was selected based on the highest AUC on the testing set and favorable calibration performance. Among the models, the LGBM exhibited superior predictive accuracy, achieving an AUC of 0.693 on the test dataset.Similarly, the RD model developed from the T2-SPAIR sequence selected 3optimal radiomics features, yielding a test set AUC of 0.684. For the DWI sequence, a total of 7 optimal radiomics features were extracted, resulting in a test set AUC of 0.777. Combining data from all three sequences, the RD-combine model was constructed, integrating 13 optimal radiomics features, including 6 from the T1 sequence, 2 from the T2-SPAIR sequence, and 5 from the DWI sequence. This model achieved a test set AUC of 0.803 (Fig. 2).
(A) Case screening and division of trainingandtestset. (B) Theradiomics workflow. RD; radiomics model, DL, deep learning model, RD-DL radiomics -deep learning model development process.
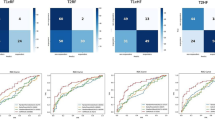
ROC and decision curves of radiomics models predicting cervical cancer OLNM. (A) ROC curves on training set (Red = Combined, Blue = DWI, Light Blue = T1, Orange = T2-SPAIR). (B) Decision curves on training set (Blue = Combined, Orange = DWI, Green = T1, Red = T2-SPAIR). (C) ROC curves on test set (Color scheme same as A). (D) Decision curves on test set (Color scheme same as A). AUC: area under the curve; RD-combine: fusion of T1/T2/DWI sequences.
DL model
The deep learning analysis of data derived from the T1 sequence ultimately yielded 7optimal features, resulting in a model with average predictive performance, as indicated by an AUC of 0.726. In contrast, the deep learning analysis based on T2-SPAIR obtained 8optimal features, yielding a model with lower prediction performance, characterized by an AUC of 0.621. Deep learning analysis of DWI data finally obtained 10 optimal features, characterized by an AUC of 0.768. Combining data from all three MRI sequences, the DL-combine model was created, integrating 17 relevant features, including 1 from the T1 sequence, 7 from the T2-SPAIR sequence, and 9 from the DWI sequence. This integrated model demonstrated enhanced predictive ability, achieving an AUC of 0.818 (Fig. 3).
ROC and decision curves of deep learning models predicting cervical cancer OLNM. (A) ROC curves on training set (Red = Combined, Blue = DWI, Light Blue = T1, Orange = T2-SPAIR). (B) Decision curves on training set (Blue = Combined, Orange = DWI, Green = T1, Red = T2-SPAIR). (C) ROC curves on test set (Color scheme same as A). (D) Decision curves on test set (Color scheme same as A). DL-combine: fusion of T1/T2/DWI deep features.
RD-DL model
We integrated radiomics features and deep learning features using both early fusion and transform fusion methods (transform fusion showed poorer performance, see Appendix 3). The early fusion method used LASSO to select 26 optimal features, including 12 radiomics features and 14 deep learning features. Subsequently, we developed the combined radiomics-deep learning (RD-DL) model. Following the integration of these features, the performance of each model improved to varying degrees, with the LGBM demonstrating the best performance (Fig. 4). The Delong test indicated significant differences in predictive performance between the RD-DL model, RD-combine model, and DL-combine model in the training set (Z = 3.802, 2.636, P < 0.05); however, no significant differences were observed in the test set (Z = 0.974, 1.349, P > 0.05). The performance metrics of all models on the test set are summarized in the table provided in Appendix 4.
ROC and decision curves of fusion models predicting cervical cancer OLNM. (A) ROC curves on training set (Red = RD-DL, Blue = DL-combine, Light Blue = RD-combine). (B) Decision curves on training set (Blue = RD-DL, Orange = DL-combine, Green = RD-combine). (C) ROC curves on test set (Color scheme same as A). (D) Decision curves on test set (Color scheme same as A). RD-DL: radiomics-deep learning fusion; DL-combine: deep learning multi-sequence fusion.
Clinical model and clinical-RD-DL model
In this study, recognizing that certain clinicopathological parameters are only obtainable postoperatively and thus unsuitable for preoperative modeling, we limited our predictors to non-invasive preoperative variables—age, FIGO stage, and tumor diameter. After normalizing these variables, we applied clustering analysis, Pearson correlation analysis, and LASSO regression for feature selection, which ultimately identified FIGO stage and tumor diameter as the most informative predictors. A LightGBM classifier built on these two features was evaluated via five-fold cross-validation, achieving an AUC of 0.722 in the training set and 0.702 in the testing set, indicating that preoperative clinical variables alone confer only modest predictive power (Fig. 5).We then attempted to integrate clinical features with radiomics and deep learning–derived features to construct a combined Clinical–Radiomics–Deep Learning (Clinical–RD–DL) model. However, during feature selection, the clinical variables were deemed redundant and thus excluded, resulting in the Clinical–RD–DL model exhibiting identical performance to the RD–DL model, with no observable gain or loss in predictive capability.
ROC and decision curves of Clinical model and Clinical- RD-DL Model predicting cervical cancer OLNM. (A) ROC curves on training set (Red = Clinical model, Blue = Clinical- RD-DL model). (B ) Decision curves on training set (Blue = Clinical model, Orange = Clinical- RD-DL model). (C) ROC curves on test set (Color scheme same as A). (D) Decision curves on test set (Color scheme same as A). RD-DL: radiomics-deep learning fusion; DL-combine: deep learning multi-sequence fusion.
Explanation of SHAP
To overcome the “black box” nature of machine learning models and improve interpretability, we visualized the final model using SHAP dependence plots, which explained how individual features affected the output of the predictive model. Figure 6A showed the distribution of SHAP values for the selected features, sorted by importance from top to bottom for each feature. The x-axis position represented the SHAP value for the feature in the same row. Points were colored differently to mark the contribution of all patients to the results, with red indicating high feature values and blue indicating low feature values. Figure 6B and C explained the assessment of individual patients. They visualized the SHAP values of features as a force predicting probability. The length of the arrow indicated the contribution of a specific feature to the SHAP value. Among them, patient B’s SHAP value was 1.8, higher than the baseline, indicating that the model predicted the patient had OLNM; patient C’s SHAP value was 0.4, lower than the baseline, indicating that the model predicted the patient did not have metastasis.
(A) Demonstrates the SHAP summary plot of our proposed model, elucidating the feature relevance and the contribution of combined features to the model’s predictive performance. (B,C) Together explain how the model predicts whether patients have occult LNM. In this context, patient B is predicted to have metastasis, while patient C is predicted not to have metastasis.
Discussion
Cervical cancer is one of the most prevalent gynecological tumors and a common cause of cancer-related death among women. Identifying the presence of LNM in patients with early-stage cervical cancer is crucial for the selection of appropriate treatment strategies. According to the 2019 NCCN guidelines, cervical cancer patients with imaging- or pathology-confirmed lymph node metastasis are classified as stage IIIC2. Although surgery is the standard for early-stage disease, detection of LNM shifts treatment to chemoradiotherapy. Preoperative imaging, however, fails to identify LNM reliably, leading to occult lymph node metastasis (OLNM) in 20–40% of patients and necessitating postoperative treatment adjustments16. To address this issue, we developed the Clinical model, RD model, DL model, combined RD-DL model, and Clinical- RD-DL model, for the preoperative assessment of OLNM in cervical cancer. The findings of this study demonstrated that the clinical model exhibited relatively poor predictive performance, primarily due to the limited amount of informative variables included. Similarly, models based on single MRI sequences consistently underperformed in comparison to those integrating multiple sequences. Notably, the RD and DL models yielded comparable results, whereas the RD–DL model achieved the highest predictive performance. This suggests that deep learning-derived features can provide complementary information to radiomics in the prediction of OLNM in cervical cancer, potentially enhancing preoperative risk stratification and supporting individualized treatment planning. Moreover, the combined Clinical–RD–DL model demonstrated identical performance to the RD–DL model, indicating that the incorporation of clinical variables did not confer additional predictive value.
In current clinical practice, the assessment of LNM in cervical cancer primarily involves CT and MRI imaging. These methods mainly evaluate whether the short diameter of the lymph node exceeds 10 mm or whether there is central necrosis. However, due to reactive hyperplasia and occult metastasis, misdiagnosis is highly likely16,17. PET-CT, while having very high specificity for diagnosing LNM, has limited clinical application due to its high cost18. Radiomics, as an emerging imaging analysis approach, has garnered substantial attention in cervical cancer research19,20,21,22. In this study, we developed radiomics (RD) models based on multi-sequence MRI—including T1-weighted, T2-weighted SPAIR, and diffusion-weighted imaging (DWI) sequences—which are routinely employed for cervical cancer evaluation. T1-weighted imaging provides detailed information on tissue morphology and size, while T2-SPAIR imaging enables visualization of tumor morphology and quantification of tumor volume and shape parameters. DWI, a functional imaging sequence, reflects the movement of water molecules constrained by the tissue microstructure, thereby facilitating the noninvasive detection of small metastatic lymph nodes23. Our results showed that the predictive performance of RD models based solely on T1 or T2-SPAIR sequences was relatively modest, with test set AUCs of 0.693 and 0.684, respectively. In contrast, the DWI-based model achieved a moderate performance, yielding a test AUC of 0.777. Notably, the combined use of all three sequences led to a substantial improvement in model performance, with the RD-combined model attaining an AUC of 0.803, which exceeded that of any single-sequence model. This finding is in line with Wang et al.9suggesting that functional imaging modalities such as DWI may more effectively capture the characteristics of OLNM compared to purely morphological sequences like T1 and T2-SPAIR. Furthermore, the complementary nature of tumor information derived from different MRI sequences likely contributes to the enhanced predictive capability of the combined model.
Deep learning, through the convolution of the kernel with different regions of the image, can transform the image into deeper high-dimensional features. This enables a more comprehensive quantification of tumor heterogeneity and a more detailed description of tumor information1,24,25. In this study, we employed three ResNet models (ResNet50, ResNet101, and ResNet200) for 3D transfer learning to extract deep learning features. The results indicate that the ResNet200 model outperformed the others, with the performance of the remaining models detailed in Appendix 2. Although this might seem counterintuitive with a relatively small dataset, since deeper networks usually tend to overfit, the complex field of medical imaging is an exception. MRI images contain very detailed anatomical features that require deeper networks to capture effectively. As the number of layers increases, the model can learn more abstract representations of the images. For high-dimensional data like MRI images, deeper networks help capture more potential biomedical signals and disease patterns, thus improving the accuracy of classification or other tasks. Furthermore, in this study, we employed data augmentation techniques, including random translation, scaling, and rotation, which can help the model generalize better and alleviate overfitting issues. Based on the ResNet200 model, we extracted the top 50 relevant features for predictive model construction. Our findings indicate that the diagnostic performance of the DL models based on T1, and DWI sequences were average, with AUC values of 0.726 and 0.768, respectively. The predictive performance of the DL model based on the T2-SPAIR sequence was lower, with an AUC of 0.621. However, when all sequences were combined, the performance of the DL-combine model improved significantly, with an AUC reaching 0.818. The predictive performance of the DL models based on T1, T2-SPAIR, DWI, and the combination of the three sequences was roughly similar to that of the RD model, with no statistically significant difference (AUC 0.726 vs. 0.693, 0.621 vs. 0.682, 0.768 vs. 0.777, 0.818 vs. 0.803, P > 0.05).
Previous studies have demonstrated that radiomics and deep learning features provide complementary information, and integrating RD–DL features may further enhance predictive performance26,27,28,29. In this study, we implemented two feature integration strategies—early fusion and transform-based fusion (technical details provided in Appendix 3)—to construct predictive models for OLNM.
Our results indicated that the early fusion strategy outperformed the transform-based fusion approach in terms of predictive accuracy, whereas the latter exhibited signs of overfitting. We attribute this discrepancy primarily to fundamental differences in information preservation and modeling complexity between the two strategies. Specifically, early fusion directly concatenates raw features from multiple modalities at the model input level, thereby preserving the original information distribution and inter-modality complementarity. Moreover, its relatively simple architecture, reduced number of parameters, and lower computational overhead make the training process more stable and easier to optimize, particularly in small-sample scenarios.
In contrast, transform-based fusion relies on deep Transformer encoders to model inter-modality interactions. Although theoretically capable of capturing long-range dependencies, this approach introduces a larger model capacity and increased computational complexity. When trained on limited datasets, it is more prone to overfitting and its performance becomes highly sensitive to optimization strategies, regularization techniques, and parameter initialization.Further analysis suggests several specific factors contributing to the suboptimal performance of the transform-based fusion model in our study: Mismatch between model complexity and dataset size: The depth and parameter volume of the Transformer encoder are ill-suited for the moderate-sized dataset used, increasing the risk of overfitting.Undifferentiated weighting of modality features: All modalities are projected into a unified space and fused with equal weights, without introducing attention-based mechanisms to emphasize informative modalities and suppress noise.Absence of dynamic learning rate scheduling and regularization: The model does not incorporate strategies such as cosine annealing or attention-specific dropout, which may limit its generalizability and robustness.These findings align with recent research trends, wherein deep fusion architectures (e.g., Transformer-based models) are predominantly applied in large-scale, high-dimensional tasks such as histopathological image analysis. However, their application in MRI-based multimodal modeling remains challenging and requires further methodological refinement.
In the training cohort, the RD–DL model built via an early-fusion strategy achieved outstanding discrimination (AUC = 0.981). The Delong test indicated statistically significant differences in predictive performance between the RD–DL, RD–combine, and DL–combine models (P < 0.05). However, in the independent test set, the RD–DL model’s performance declined, achieving an AUC of 0.903, and the differences compared to the other models were not statistically significant (P > 0.05). The difference in AUC between the training and test sets (ΔAUC = 0.078) exceeded the commonly accepted generalization warning threshold (ΔAUC > 0.05), suggesting a potential risk of overfitting. This risk may be intensified by factors such as the limited sample size of 185 patients (130 in the training set and 55 in the independent test set), the use of different imaging equipment (Siemens scanners for training and Philips scanners for testing), and the reliance on N4 bias field correction, which may not fully mitigate equipment-related variations.Despite this performance drop, the RD–DL model still achieved a commendable AUC of 0.903 in the test set and maintained the highest net benefit within the threshold probability range of 0.2–0.8, underscoring its practical value in supporting clinical decision-making. To improve the model’s generalizability and clinical reliability, future research should focus on the following enhancements: Expanding to multicenter, large-sample cohorts to bolster model robustness; Implementing advanced harmonization techniques, such as ComBat correction or CycleGAN generative adversarial networks, to overcome the limitations of N4 correction and achieve cross-device feature alignment; Deploying an automated segmentation pipeline based on U-Net to minimize manual annotation bias.
Due to the “black box” nature of machine learning, we used the SHAP method to analyze the contributions of different variables in order to gain a deeper understanding of the risk prediction model. SHAP analysis provides a general tool for assessing the feature importance of machine learning models. In interpretable machine learning models, we use SHAP values to explain model outputs by calculating the contribution of each input feature across all samples in the dataset. We visualized SHAP values in both global and local forms to study the effects and interactions between variables. If the SHAP value is positive, it indicates that the feature has a positive contribution to the model’s prediction, and vice versa. In our study, SHAP analysis showed that deep learning features extracted from the DWI sequence were the most informative features.
The main risk factors for OLNM in cervical cancer include deep stromal invasion, FIGO staging, LVSI, histological grading, and tumor diameter30. As the FIGO stage of cervical cancer increases and the depth of stromal invasion increases, the extent of cancer lesions expands, potentially increasing the risk of OLNM. Therefore, it is reasonable to believe that higher FIGO stages and deeper stromal invasion in early patients with cervical cancer may correlate with a poorer prognosis31,32,33,34,35,36. In this study, significant differences were observed between FIGO staging and deep stromal invasion with OLNM across the entire dataset (P = 0.011, P < 0.001), as well as within the training set (P = 0.003, P < 0.001). However, these differences were not statistically significant in the test set (P = 0.135, P = 0.067). These findings diverge from those reported in previous studies12,13,25. This discrepancy is attributed to the small sample size and potential selection bias inherent in the test set data sourced from Medical Center 2. This underscores the necessity to expand the data sample size in future studies.LVSI positivity can indicate an elevated risk of LNM and serves as an independent risk factor for LNM37,38. Consistent with these findings, our study also demonstrated a significant association between LVSI and OLNM, with P values < 0.001 in both the training and test sets. However, as LVSI status requires confirmation through postoperative pathology, it has not been incorporated into the construction of the preoperative prediction model. Currently, the correlation between tumor diameter, histological grading, and the occurrence of OLNM in cervical cancer remains controversial39,40,41,42,43. In our study, there was no significant association between tumor diameter, histological grading, and occult LNM (P = 0.139, P = 0.263).
Previous studies9,12,13,15 have shown that incorporating clinical factors can enhance predictive model performance. In our work, although a range of clinical variables was collected, key indicators such as LVSI and deep stromal invasion require postoperative pathological confirmation. Consequently, we developed a purely clinical model based on age, maximum tumor diameter, and FIGO stage, which yielded an AUC of 0.702 on the independent test set—indicative of only modest discrimination. We then fused these three clinical variables with the early-fusion RD–DL features to construct a ‘Clinical–RD–DL’ model. Contrary to expectations and prior reports9,12,13,15this fusion model performed equivalently to the standalone RD–DL model, showing no incremental benefit.We attribute these results to two main factors: Discrepant levels of information representation. RD–DL features, extracted via high-throughput methods such as gray-level co-occurrence matrices and wavelet transforms, capture microscopic tumor texture, morphology, and intratumoral heterogeneity—proxy markers of biological behaviors like local invasiveness and microvascular infiltration. In contrast, age, FIGO stage, and maximum tumor diameter convey only macroscopic anatomical and staging information and cannot reflect subtle heterogeneity or microenvironmental changes. Although FIGO stage delineates tumor spread, intrastage biological variability is more precisely quantified by imaging-derived features such as boundary sharpness and capsule integrity. Maximum tumor diameter measures only size, a parameter that shape metrics (e.g., volume, surface area) already capture more comprehensively. Age has a weak linkage to metastatic propensity and is not independently predictive.As a result, the clinical variables included here largely overlap with the information encoded in the RD–DL features and offer limited additional value, preventing the “clinical–RD–DL” model from outperforming the pure RD–DL model. Future studies should explore the incorporation of novel clinical biomarkers with stronger biological relevance (for example, SCC-Ag or serum biochemical markers) and validation on large, multicenter cohorts to maximize the predictive gains of multimodal fusion.
Conclusions
In summary, the RD and DL models developed in this study based on multiple MRI sequences demonstrate strong predictive performance and can be used as tools to predict OLNM in patients with cervical cancer. The RD-DL model demonstrates the highest predictive potential among the evaluated models, with an AUC of 0.903 in the test set, suggesting a trend towards better performance that warrants validation in larger, multicenter studies.
Materials & methods
We retrospectively collected clinical data from 2,314 cervical cancer patients who underwent MRI examinations from two medical centers. This study has obtained approval from the hospital ethics committee. As it is a retrospective study, the requirement for obtaining patient informed consent was waived. Our research was conducted in accordance with the principles of the Helsinki Declaration. This study has received ethical approval from the Ethics Review Committee of Guangyuan First People’s Hospital (Approval Number: 072LW2024012).
-
(I)
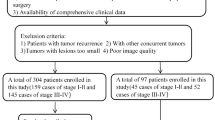
The inclusion criteria were as follows: (I) patients confirmed to have cervical cancer (either squamous cell carcinoma or adenocarcinoma) by pathological biopsy,
-
(II)
Patients clinically diagnosed with stage IB∼IIB disease (FIGO 2018),
-
(III)
Patients who underwent MRI examinations within 2 weeks before surgery, and whose preoperative imaging did not indicate lymph node metastasis, and did not undergo lymph node biopsy. (IV) patients who underwent total hysterectomy and pelvic lymph node dissection. The exclusion criteria included: (I) patients who received chemotherapy before surgery, (II) patients with poor image quality or motion artifacts, and (III) patients who did not undergo lymph node dissection or had clear LNM on MRI (Short-axis diameter ≥ 10 mm, or necrosis, irregular lymph node shape, blurred boundaries). Finally, 185 patients with cervical cancer were included in the study. Among these, 130 patients from Center 1 constituted the training set, while 55 patients from Center 2 comprised the test set.
MRI acquisition
All MRI examinations were performed using a 3.0T MRI scanner (Siemens VIDA3.0T, Philips Ingenia3.0T). Patients were instructed to fast for a minimum of 4 h before the examination. During the procedure, patients were positioned supine on the examination bed with their heads oriented toward the front, using an abdominal coil for image acquisition. The scanning sequence included axial T1-weighted imaging (T1WI), axial T2-weighted imaging with spectral attenuated inversion recovery (T2WI-SPAIR), axial diffusion-weighted imaging (DWI) with B values of 0 and 800 s/mm², coronal T2WI, and dynamic enhanced T1WI. All MRI data were stored in the hospital’s picture archiving and communication system in Digital Imaging and Communications in Medicine (DICOM) format before any further processing.
Radiomics feature extraction
After normalizing and performing N4 bias correction on all images17a radiologist with 5 years of experience in pelvic imaging diagnostics used 3D Slicer 5.4.0 software to manually delineate the ROI along the lesion’s edge on axial T1, axial T2-SPAIR, and axial DWI images (Fig. 1). The ROI was saved in NII format and subsequently imported into the Onekey platform 3.1.8 for radiomics feature extraction. Given the relatively small size of the dataset in this study, the authors applied data augmentation techniques—namely random translation, scaling, and rotation—to enhance model performance and reduce the risk of overfitting. Each sequence yielded 1198 features, categorized into three types: surface features, first-order statistics, and texture features. To evaluate the consistency between two observers, thirty cases were randomly selected from the training set, and another radiologist with 15 years of experience in pelvic imaging redrew the ROI. The intra-class correlation coefficient (ICC) was used to evaluate consistency, and features with good consistency (ICC > 0.9) were retained.
Deep learning feature extraction
In this study, the authors opted to utilize a pre-trained Convolutional Neural Network model, for 3D deep transfer learning due to its ability to operate effectively with fewer training samples and save computational resources44. The original images from each patient were cropped and resampled to dimensions of 90 mm × 90 mm × 90 mm. Subsequently, image signal intensities were normalized to a range between 0 and 1, and all images were uniformly saved in NII format. These processed images were then input into the Onekey platform 3.1.8 for model training. Given the limited data size in this study, the authors employed data augmentation techniques, including random translation, scaling, and rotation, to enhance model performance and mitigate the risk of overfitting. Finally, the authors extracted deep learning features using the ResNet50, ResNet101, and ResNet200 models, with the ResNet200 model showing the best performance (the performance of ResNet50 and ResNet101 is detailed in Appendix 1). When employing the ResNet200 model, 2048 deep learning features were extracted for each sequence. Due to the high feature dimensionality extracted by deep learning models such as ResNet200, we use the Minimum Redundancy Maximum Relevance (mRMR) algorithm for feature compression. This approach reduces model complexity, prevents overfitting, and improves computational efficiency and interpretability. The method maximizes the correlation between features and labels while minimizing redundancy between features, ensuring that the final selected feature set is both representative and non-redundant. This, in turn, helps build a more robust predictive model. Ultimately, the top 50 deep learning features most strongly correlated with the target variable were retained.
Feature selection and modeling
To eliminate the influence of dimensional differences among features, the original variables were first standardized using Z-score normalization (formula: Xnorm\(\:=\frac{\text{x}-\varvec{\upmu\:}\text{}}{{\upsigma\:}}\)), transforming them into a standard normal distribution with a mean of 0 and a standard deviation of 1. Spearman rank correlation analysis was then performed to evaluate feature redundancy, with a correlation coefficient threshold of ( |ρ| > 0.9 ) used to identify and remove highly correlated features, thereby retaining those most representative of OLNM. During the feature selection phase, the least absolute shrinkage and selection operator (LASSO) regression was applied, and fivefold cross-validation was used to determine the optimal regularization parameter (α) by minimizing the log loss of the binary classification task. Features with non-zero coefficients were retained, and a coefficient path plot was generated to visualize the shrinkage process. Based on the selected features, five classification models were established: Gradient Boosting (GB), Light Gradient Boosting Machine (LGBM), Logistic Regression (LR), Multilayer Perceptron (MLP), and Random Forest (RF). The performance of each model was systematically evaluated using metrics including area under the curve (AUC), sensitivity, specificity, F1-score, and their respective 95% confidence intervals (CIs).The optimal model was selected based on its highest AUC value in the test set and a well-fitted calibration curve, indicating superior predictive accuracy and calibration, with a Brier score below 0.1.Following this workflow, five types of models were constructed to comprehensively evaluate the predictive value of various data sources for assessing OLNM in cervical cancer: Clinical model, radiomics model, deep learning model, radiomics–deep learning fusion model, clinical model, and integrated radiomics–deep learning–clinical model.(Fig. 1).
Statistical methods
All statistical analyses were conducted using SPSS 26.0 software and the Onekey platform 3.1.8. All data conforming to a normal distribution are presented asx̄ ± s. The χ² test was used for discrete variables, while the independent sample t-test was used for continuous variables. Consistency between the two physicians in extracting radiomics features was determined using the ICC, with an ICC exceeding 0.9 indicating good consistency.
The diagnostic performance of the model was evaluated based on the area under the curve (AUC) value. A model with an AUC greater than 0.9 was considered to have excellent predictive performance, while an AUC exceeding 0.8 indicated good diagnostic performance. AUC between 0.7 and 0.8 denoted average diagnostic performance, while values between 0.6 and 0.7 suggested low diagnostic performance. The DeLong test was used to evaluate differences in AUC values. Decision curve analysis (DCA) was employed to assess the clinical utility of the model. A p-value less than 0.05 was considered statistically significant.
Data availability
The code source used is available at the following link: https://github.com/OnekeyAI-Platform/onekey.
References
Bray, F. et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 68, 394–424 (2018).
Bhatla, N. et al. Revised FIGO staging for carcinoma of the cervix Uteri. Int. J. Gynecol. Obstet. 145, 129–135 (2019).
Shi, T. Y. et al. RAD52 variants predict platinum resistance and prognosis of cervical cancer. PLoS One. 7, e50461 (2012).
Wang, T. et al. Preoperative prediction of pelvic lymph nodes metastasis in early-stage cervical cancer using radiomics nomogram developed based on T2-weighted MRI and diffusion-weighted imaging. Eur. J. Radiol. 114, 128–135 (2019).
Liu, B., Gao, S. & Li, S. A comprehensive comparison of CT, MRI, positron emission tomography or positron emission tomography/ct, and diffusion weighted imaging-MRI for detecting the lymph nodes metastases in patients with cervical cancer: a meta-analysis based on 67 studies. Gynecol. Obstet. Invest. 82, 209–222 (2017).
Lambin, P. et al. Radiomics: extracting more information from medical images using advanced feature analysis. Eur. J. Cancer. 48, 441–446 (2012).
Wang, T. et al. Preoperative prediction of parametrial invasion in early-stage cervical cancer with MRI-based radiomics nomogram. Eur. Radiol. 30, 3585–3593 (2020).
Li, Z. et al. MR-based radiomics nomogram of cervical cancer in prediction of the lymph‐vascular space invasion preoperatively. J. Magn. Reson. Imaging. 49, 1420–1426 (2019).
Wang, T. et al. A MRI radiomics-based model for prediction of pelvic lymph node metastasis in cervical cancer. World J. Surg. Oncol. 22, 55 (2024).
Xin, Z. et al. An MRI-based machine learning radiomics can predict short‐term response to neoadjuvant chemotherapy in patients with cervical squamous cell carcinoma: A multicenter study. Cancer Med. 12, 19383–19393 (2023).
Landhuis, E. Deep learning takes on tumours. Nature 580, 551–554 (2020).
Qian, W. et al. RESOLVE-DWI-based deep learning nomogram for prediction of normal-sized lymph node metastasis in cervical cancer: a preliminary study. BMC Med. Imaging. 22, 221 (2022).
Li, P. et al. Deep learning nomogram for predicting lymph node metastasis using computed tomography image in cervical cancer. Acta Radiol. 64, 360–369 (2023).
Liu, Y. et al. Development of a deep learning-based nomogram for predicting lymph node metastasis in cervical cancer: a multicenter study. Clin. Transl Med. 12, e938 (2022).
Jeong, S. et al. Comparing deep learning and handcrafted radiomics to predict chemoradiotherapy response for locally advanced cervical cancer using pretreatment MRI. Sci. Rep. 14, 1180 (2024).
Zhu, M. et al. Imaging evaluation of para-aortic lymph nodes in cervical cancer. Acta Radiol. 64, 2611–2617 (2023).
Dovrou, A. et al. A segmentation-based method improving the performance of N4 bias field correction on T2weighted MR imaging data of the prostate. Magn. Reson. Imaging. 101, 1–12 (2023).
Olthof, E. P. et al. Diagnostic accuracy of MRI, CT, and [18F] FDG-PET-CT in detecting lymph node metastases in clinically early-stage cervical cancer—a nationwide Dutch cohort study. Insights Imaging. 15, 36 (2024).
Liu, Q. et al. Identification of lymph node metastasis in pre-operation cervical cancer patients by weakly supervised deep learning from histopathological whole‐slide biopsy images. Cancer Med. 12, 17952–17966 (2023).
Yan, H. et al. Machine learning-based multiparametric magnetic resonance imaging radiomics model for preoperative predicting the deep stromal invasion in patients with early cervical cancer. J. Imaging Inf. Med. 37, 230–246 (2024).
Bizzarri, N. et al. Radiomics systematic review in cervical cancer: gynecological oncologists’ perspective. Int. J. Gynecol. Cancer. 33, 1522–1541 (2023).
Wu, Q. et al. Radiomics analysis of multiparametric MRI evaluates the pathological features of cervical squamous cell carcinoma. J. Magn. Reson. Imaging. 49, 1141–1148 (2019).
Thoeny, H. C. et al. Metastases in normal-sized pelvic lymph nodes: detection with diffusion-weighted MR imaging. Radiology 273, 125–135 (2014).
Sun, R. J. et al. CT-based deep learning radiomics analysis for evaluation of Serosa invasion in advanced gastric cancer. Eur. J. Radiol. 132, 109277 (2020).
Zhang, X. F. et al. Deep-learning-based radiomics of intratumoral and peritumoral MRI images to predict the pathological features of adjuvant radiotherapy in early-stage cervical squamous cell carcinoma. BMC Women’s Health. 24, 182 (2024).
Ran, J. et al. Development and validation of a nomogram for preoperative prediction of lymph node metastasis in lung adenocarcinoma based on radiomics signature and deep learning signature. Front. Oncol. 11, 585942 (2021).
Li, X., Yang, L. & Jiao, X. Comparison of traditional radiomics, deep learning radiomics and fusion methods for axillary lymph node metastasis prediction in breast cancer. Acad. Radiol. 30, 1281–1287 (2023).
Jiang, Y. et al. Non-invasive tumor microenvironment evaluation and treatment response prediction in gastric cancer using deep learning radiomics. Cell. Rep. Med. 4, 101146 (2023).
Gu, W. et al. Development and validation of CT-based radiomics deep learning signatures to predict lymph node metastasis in non-functional pancreatic neuroendocrine tumors: a multicohort study. eClinicalMedicine 65, 102269 (2023).
Ferrandina, G. et al. Can we define the risk of lymph node metastasis in early-stage cervical cancer patients? A large-scale, retrospective study. Ann. Surg. Oncol. 24, 2311–2318 (2017).
Epstein, E. et al. Early-stage cervical cancer: tumor delineation by magnetic resonance imaging and ultrasound—a European multicenter trial. Gynecol. Oncol. 128, 449–453 (2013).
Rizzo, S. et al. Pre-operative MR evaluation of features that indicate the need of adjuvant therapies in early stage cervical cancer patients. A single-centre experience. Eur. J. Radiol. 83, 858–864 (2014).
Song, J. et al. Value of diffusion-weighted and dynamic contrast-enhanced MR in predicting parametrial invasion in cervical stromal ring focally disrupted stage IB–IIA cervical cancers. Abdom. Radiol. 44, 3166–3174 (2019).
Manganaro, L. et al. Staging, recurrence and follow-up of uterine cervical cancer using MRI: updated guidelines of the European society of urogenital radiology after revised FIGO staging 2018. Eur. Radiol. 31, 7802–7816 (2021).
Laliscia, C. et al. MRI-based radiomics: promise for locally advanced cervical cancer treated with a tailored integrated therapeutic approach. Tumori J. 108, 376–385 (2022).
Plotti, F. et al. Tailoring parametrectomy for early cervical cancer (Stage IA-IIA FIGO): A review on surgical, oncologic outcome and sexual function. Minerva Obstet. Gynecol. 73, 149–159 (2021).
Becker, A. S. et al. MRI texture features May predict differentiation and nodal stage of cervical cancer: a pilot study. Acta Radiol. Open. 6, 2058460117729574 (2017).
Jiang, X. et al. MRI based radiomics approach with deep learning for prediction of vessel invasion in early-stage cervical cancer. IEEE/ACM Trans. Comput. Biol. Bioinform. 18, 995–1002 (2020).
He, Z. et al. The value of HPV genotypes combined with clinical indicators in the classification of cervical squamous cell carcinoma and adenocarcinoma. BMC Cancer. 22, 776 (2022).
Horn, L. C. et al. Tumor size is of prognostic value in surgically treated FIGO stage II cervical cancer. Gynecol. Oncol. 107, 310–315 (2007).
Turan, T. et al. Prognostic effect of different cut-off values (20 mm, 30 mm and 40 mm) for clinical tumor size in FIGO stage IB cervical cancer. Surg. Oncol. 19, 106–113 (2010).
Bai, H. et al. The potential for less radical surgery in women with stage IA2–IB1 cervical cancer. Int. J. Gynecol. Obstet. 130, 235–240 (2015).
Beavis, A. L. et al. Sentinel lymph node detection rates using indocyanine green in women with early-stage cervical cancer. Gynecol. Oncol. 143, 302–306 (2016).
He, K. et al. Identity Mappings in Deep Residual Networks. Comput Vis-ECCV. 9908, 630-645 (2016).
Acknowledgements
We thank the research team and the OnekeyAI team for their support in this study.
Author information
Authors and Affiliations
Contributions
G.T. led the project conception and design and contributed to reviewing and providing feedback. S.L. developed and validated the methods and wrote and edited the manuscript. Y.G. conducted experiments, collected data, and contributed to data analysis and interpretation. Y.Y. performed data analysis and interpretation and created visualizations and charts. Q.M. provided resources and support and contributed to reviewing and providing feedback. W.H. provided resources and support and managed and coordinated the project. All authors contributed to reviewing and editing the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Luo, S., Guo, Y., Ye, Y. et al. Prediction of cervical cancer lymph node metastasis based on multisequence magnetic resonance imaging radiomics and deep learning features: a dual-center study. Sci Rep 15, 29259 (2025). https://doi.org/10.1038/s41598-025-13781-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-13781-y