Abstract
Plant diseases significantly harm crops, resulting in significant economic losses across the globe. In order to reduce the harm that these diseases produce, plant diseases must be diagnosed accurately and timely manner. In this work, a YOLO-LeafNet approach is proposed for detecting diseases from leaf images of four distinct species, namely, grape, bell pepper, corn, and potato. About 8850 leaf images have been acquired for this work from five different publicly available datasets on Kaggle. All the acquired images were pre-processed by applying four different image pre-processing operations. The number of images in the training dataset was tripled for better model performance by applying five different augmentation operations. The augmented dataset was then used to train YoloV5, YoloV8, and the proposed YOLO-LeafNet. The performance of all three models is evaluated in terms of recall, precision, and Mean Average Precision (mAP). The YoloV5 attained a precision of 0.861, recall of 0.868, mAP50 of 0.944, and 0.815 of mAP50-95, and YoloV8 attained 0.977 precision, 0.975 recall, 0.984 of mAP50, and mAP50-95 of 0.915, whereas the proposed the YOLO-LeafNet attained precision of 0.985, recall of 0.980, mAP50 of 0.990, and mAP50-95 of 0.940. The experimental results reveal that the proposed YOLO-LeafNet outperformed YOLOv5 and YOLOv8 in terms of all performance metrics.
Similar content being viewed by others
Introduction
In India, the agricultural industry plays a crucial role in employment, food security, and economic growth. Nearly half of India’s population is employed in agriculture, which contributes roughly 14% of the country’s GDP. According to the Economic Survey of India, in 2021–2022, 45.5% of the population was employed in agriculture1, which contributed 18.8% to the country’s GDP2. Plant diseases, however, have a major effect on agricultural productivity and quality3. Fungal infections, bacterial infections, environmental factors, parasitic nematodes, and insects are the reasons behind plant diseases. Early disease detection and disease control are essential for reducing this loss. Traditionally, plant disease detection has been done through visual inspection, which can be subjective and time-consuming, especially when working with multiple plants.
However, new approaches for detecting plant diseases, based on image processing, Machine Learning (ML), Deep Learning (DL) techniques, and hyperspectral imaging have been developed as a result of technological advancements. For instance, authors utilized multi-class SVM for detecting three distinct classes of apple diseases4. Authors utilized a feed-forward back propagation neural network for detecting mildew from grape leaves5. The authors utilized machine learning techniques for the prediction of crop yield6. DL-based approaches for detecting plant diseases have attracted growing interest in recent years due to their capacity to acquire complex representations from vast amounts of data. DL-based Convolutional Neural Networks (CNNs) are most widely utilized for detecting plant diseases. For instance, CNN was utilized for detecting three classes of apple diseases7. The authors utilized SqueezeNet and AlexNet to detect diseases from tomato leaf images8. Authors utilized AlexNet for detecting diseases from apple leaves9. The authors utilized a deep CNN to detect diseases from leaf images of guava10. Authors utilized AlexNet and GoogleNet to detect diseases from soybean leaves11. Authors utilized CNN to classify healthy/unhealthy leaves of apple and tomato12. Authors utilized VGG16 and GoogleNet to detect diseases from tomato leaves13. Authors utilized ResNet, DenseNet, and VGG to detect diseases from grape leaves14.
DL-based You Only Look Once (YOLO) can also be employed for the detection of plant diseases. For instance, YOLOv7 was utilized for detecting diseases from citrus leaves15. YOLO was utilized by Achyut Morbekar et al. to detect plant diseases16. Authors utilized YOLOv3 for classifying healthy and diseased tomato leaves17. Authors utilized YOLO-Dense for detecting diseases from tomato leaves18. YOLO can be viewed as a particular variety of CNN with an architecture specialized for object recognition tasks. A single forward pass of the network enables YOLO to detect numerous objects within a single image.
In19, the authors used SVM to identify diseases from images of grape leaves. A digital camera was utilized for capturing some grape leaf images, while others were downloaded from the internet. During segmentation, K-means clustering was employed. The obtained pictures were scaled down to 300 × 300. The suggested model achieved an accuracy of 88.89%. Furthermore, in20, the authors utilized a multi-class SVM for detecting diseases from leaf images of tomatoes. Images were acquired from a real environment. An inverse difference operation was utilized for segmenting the images. The proposed approach attained an accuracy of 93.75%. sA method based on deep CNN was put forth by Sumita Mishra et al.21 to identify diseases in maize leaves. Some parts of the dataset images were acquired from a real environment, while others came from the PlantVillage dataset. The training of the suggested model was done with 70% of the acquired images, validation using 20%, and testing using the remaining 10%. In the final layer of the suggested method, softmax was used as the activation function, while RELU was utilized in all other layers. The suggested model achieved 88.46%.
Authors in22 proposed a CNN with global average pooling for diagnosing six different classes of diseases, namely: Downy Mildew, Angular Leaf Spot, Anthracnose, Gray Mold, Black Spot, and Powdery Mildew from cucumber leaves. The data about 700 cucumber leaf images was gathered using a digital camera from the planting bases in China. The dataset’s image count was increased by applying geometric and intensity transformations during the augmentation stage. The suggested approach attained 94.65% accuracy. The authors in23 suggested a hybrid approach based on autoencoders and CNN to identify diseases in three different crops. Out of the 900 images obtained from the PlantVillage dataset, 600 were used for training the proposed model, while the remaining 300 images were utilized for its evaluation. The Adam optimizer was employed to improve accuracy. The shortcoming of the proposed model was its small dataset.
An approach based on DL for detecting diseases from soybean leaf images was suggested by the authors in24. Images of 1470 soybean leaves were collected from different farms. By using ten different augmentation operations to expand the dataset size, the proposed method achieved an accuracy of 94.29%. The authors in25 proposed a method based on DL to identify diseases from potato leaves. Potato leaf images were obtained from a publicly available dataset on Kaggle. Three different classes of potato leaves are recognized by the suggested approach. A 97.8% success rate is offered by the suggested method. The authors in26 compared existing ML (SVM, KNN, fully connected neural network) and DL algorithms created for detecting plant disease. The suggested study demonstrates that DL technology outperforms ML methods and offers classification accuracy of over 99%. Authors in27 suggested a CNN-based method for detecting and categorizing plant diseases. For the purpose of providing more accuracy than the existing models, the suggested model was constructed on the TensorFlow framework and achieved more than 95% accuracy. In addition, K-Means clustering was utilized to locate the disease-affected areas since it may be used to determine the precise quantity of fertilizer needed to safeguard plants.
In28, the authors suggested the KFHT-RLPBC model for identifying diseases from plant leaves, where time complexity is decreased during the stage of feature extraction. The three stages of the proposed model’s operation are pre-processing, feature extraction, and classification. Kuan Filter is utilized in the initial stage to improve image quality by reducing noise in it. In the second stage, the Hough transformation was used for extracting characteristics, namely: color, texture (contrast, correlation), and shape (area, perimeter, and length). The PlantVillage dataset was utilized to test the proposed model. The suggested model outperforms the existing model by attaining an accuracy of 88%. The authors in29 proposed an approach based on CNN for detecting diseases from leaf images of three plant species, namely: tomato, potato, and pepper. For this work, the dataset of about 20,636 leaf images related to 15 distinct classes was obtained from the PlantVillage, and the acquired images were resized to 128 × 128 during pre-processing, and then passed to the CNN model. The proposed approach attained an accuracy of 98.029%. The authors in30 proposed an approach based on faster R-CNN for diagnosing diseases from soybean leaf images. Images of soybean leaves affected by three distinct classes of diseases, namely bacterial spot, frogeye leaf spot, and virus disease, were gathered from farms in China. The stochastic gradient descent was employed for optimizing the parameters. The suggested method achieved an average mean precision of 83.34%. The authors in31 proposed MobileNet-V2 for detecting diseases in rice plants. 1100 rice plant images that were affected by twelve different diseases were gathered from the internet and the real-time environment. 80% of the dataset was utilized for training the proposed approach, and the remaining 20% was utilized to test it. The size of all the acquired images was changed to 224 × 224. Five different augmentation operations were applied to the acquired images to augment the number of images. The proposed approach achieved 99.67% accuracy.
The authors in32 proposed deep ensemble neural networks (DENN) for autonomous plant disease identification. Data augmentation techniques like scaling, rotation, image enhancement, and translation were used to overcome overfitting, while hyperparameters were tweaked using transfer learning. The suggested model surpasses existing pre-trained models in terms of performance. The authors in33 used an EfficientNetV2 for identifying diseases from images of grape and cardamom leaves. Cardamom leaf images were obtained from the farm field using mobile phones, whereas grape leaf images were obtained from the PlantVillage dataset. U2-Net was used to remove the background of the acquired images. Cardamom disease detection using the recommended method was accurate to 98.26% and grape disease detection to 96.44%. Quan Huu Cap et al.34 presented LeafGAN for augmenting image data. The performance of the suggested LeafGAN and the existing vanilla CycleGAN was evaluated for categorizing cucumber leaf diseases. Five distinct classes of cucumber leaves were utilized to test the suggested model. The proposed method demonstrated a 7.4% improvement in classification performance and generated high-quality images post-augmentation, enabling it to outperform the vanilla CycleGAN in performance comparisons. A method based on YOLOv7 was suggested by Junyang Chen et al.15 for identifying diseases in citrus. The dataset for the proposed study, consisting of 1,266 images, was collected from a real-world environment. 80% of acquired images were utilized to train the proposed model, whereas the remaining 20% were for validation and testing of the proposed model (10% each). Five distinct data enhancement operations, including random cropping, mosaic enhancement, contrast enhancement, cutout, and scaling, were used, and the Mean Average Precision (mAP) was 97.29%. In35, a deep CNN was suggested to identify crop diseases from tomato, maize, potato, and apple leaf images. While the majority of the disease-infected leaves and healthy leaves were collected from the plantvillage dataset for forming the dataset, some of the images were captured with an HD camera. 100 epochs were used to train the suggested model. The suggested method achieved a precision of 93.78%, a recall of 94.91%, and an mAP of 40.6%. An automatic approach for the identification of diseases and plant leaf damage was presented by B. Sai Reddy36. The plantvillage dataset was utilized to obtain the images of strawberry, potato, apple, and grape leaves. The suggested method’s initial step used DenseNet to determine the type of disease from a plant leaf image. Images classified as healthy and different types of diseases were utilized for training the DenseNet model. Then, new leaf images were tested using this model. The suggested DenseNet model generated a classification accuracy of 100% while using fewer images during the training phase. Utilizing DL-based semantic segmentation, the second stage located the damage to the leaf. Every pair of RGB pixel values in the image was recovered, and the 1D CNN was used to train under supervision on the retrieved pixel values. The trained model was able to identify the pixel-level damage present in the leaves. The proposed method attained precision and recall values of 0.971 each, along with an accuracy of 97%. The authors in37 suggested an ML-based method for identifying diseases from tomato leaves. Before using histogram equalization to enhance the quality of the leaf images, the tomato leaf samples were resized to 256 × 256 pixels. The data space was partitioned into Voronoi cells through K-means clustering. Contour tracing was utilized for extracting the edges of the leaf sample. To extract the useful features from the leaf samples, many descriptors, including the discrete wavelet transform (DWT), principal component analysis (PCA), and grey-level co-occurrence matrix (GLCM), were applied. Finally, three distinct ML techniques, including SVM, CNN, and K-Nearest Neighbor (K-NN), were applied to classify the retrieved features. Using SVM, the proposed model’s accuracy was 88%, compared to 97% for K-NN and 99.6% for CNN.
The authors in38 proposed a DL-based approach for detecting five categories of rice plant diseases, namely: false smut, bacterial leaf blight, rice blast, sheath rot, brown leaf spot, and brown leaf spot. The images of rice crops were acquired from diverse rice fields across Kerala. The dataset’s volume was expanded through the implementation of diverse augmentation operations, namely: flipping, geometric alterations, cropping, rotation, random erasing, noise injection, and colour alteration. The suggested method achieved a higher accuracy of 91.4% using softmax combined with ReLU. The authors in39 used CNN for identifying four different classes of potato disease. For this work, around 5000 potato images were acquired from Iranian farms as well as the USDA and CFIA. Rotation and cropping operations on the images resulted in an increase in the dataset size. During the pre-processing stage, the acquired images were resized to 128 × 128. The proposed approach attained an accuracy between 98 and 100%. The authors in40 suggested a novel image processing approach and multi-class SVM to identify and categorize grape leaf diseases such as black rot, leaf blight, and black measles. The plantvillage dataset was utilized for acquiring images of grape leaves. K-means clustering was employed to automatically differentiate between the affected areas and the healthy sections of the leaf, followed by extracting features from three distinct color models: l*a*b, HSV, and RGB. In the proposed work, PCA was utilized to reduce the dimension of the features, and SVM was used as an effective classification approach. The relief feature selection process ultimately chose the most significant features. Although utilizing PCA for feature dimension reduction resulted in an accuracy of 98.97%, GLCM features produced a result of 98.71% accuracy. The authors in41 used five distinct YOLO models to identify three categories of maize diseases. Image acquisition was the initial step in the suggested method. Using a smartphone and varied weather conditions at various growth phases, images of maize leaves were acquired from research farms. The captured images were all downsized to 416 × 416 in the pre-processing phase, which was the second stage of the suggested methodology. The dataset was expanded by implementing different augmentation operations, including flipping, rotation, cropping, and scaling. After augmentation, images were passed for training and testing using different versions of YOLO.
Compared to conventional object detection algorithms, which employ sliding windows and make numerous passes through an image, this makes it quicker and computationally more effective. These techniques allow the automated and non-destructive evaluation of plants, enabling the quick screening of large plant populations. These methods can identify early disease signs before they become obvious to the human eye, facilitating timely intervention and possibly lowering crop losses.
The YOLO (You Only Look Once) framework is particularly significant for disease detection when compared to traditional machine learning (ML) and other deep learning (DL)-based approaches, due to its advantages in accuracy, speed, and simplicity. YOLO employs a single neural network to simultaneously predict bounding boxes and class probabilities, allowing it to achieve high accuracy, especially in detecting diseases within small and localized regions. As a single-stage object detection method, YOLO offers real-time detection capabilities, making it highly suitable for time-sensitive agricultural applications. Furthermore, its relatively simple architecture makes it easier to train and deploy compared to other complex object detection algorithms.
In this work, we propose a DL-based YOLO-LeafNet model for the detection of plant diseases from leaf images of four plant species: grape, bell pepper, potato, and corn. The key contributions of this study are as follows: First, the proposed YOLO-LeafNet model is capable of detecting eleven distinct classes of plant diseases, including healthy grape leaf, grape esca, grape black rot, grape leaf blight, healthy potato leaf, potato late blight, potato early blight, healthy corn leaves, corn common rust, healthy bell pepper leaf, and bacterial spot. This model is based on the latest YOLO architecture, offering both speed and efficiency in disease detection from leaf images.
Second, data augmentation techniques have been strategically utilized to increase the dataset size and introduce greater variability. This step is crucial for improving model generalization and reducing overfitting, thereby resulting in a more robust and effective training process. Third, the proposed work addresses the limitations of manual disease detection, which often requires extensive expertise, is time-consuming, and labor-intensive. By using YOLOv5, YOLOv8, and the proposed YOLO-LeafNet model, we demonstrate improved automation and accuracy in detecting diseases across the selected plant species, thereby minimizing the need for human intervention.
Finally, the automated detection approach contributes to reducing labor costs, pesticide usage, and crop losses. With timely and precise identification of plant diseases, farmers can implement targeted treatments, leading to significant cost savings and promoting sustainable agricultural practices.
This work consists of five distinct sections, with Sect. 1 being the introduction and summary of existing work is presented. In Sect. 2, the methodology for detecting plant diseases is proposed. The results of the proposed methodology and a comparison of both approaches, along with the proposed YOLO-LeafNet, have been discussed in Sect. 3. In Sect. 4, the discussion focuses on the results obtained from both versions of YOLO that were utilized, as well as the proposed YOLO-LeafNet model. It also includes a comparison of these results with those achieved by existing state-of-the-art approaches. Lastly, Sect. 5 provides a comprehensive conclusion to the proposed research work, offering insight and outlining future directions for advancements in plant disease detection. A brief summary of related work is presented in Table 1.
On the basis of the related work, the limitations of current plant disease detection models highlight several challenges that need to be addressed to enhance their real-world applicability and performance. These limitations are as follows:
-
1.
Many models are based on small or specialized datasets, which limits their ability to generalize to diverse and complex real-world agricultural conditions.
-
2.
The fixed image resizing and manual segmentation methods (e.g., K-means) may lead to the loss of important features.
-
3.
The ML-based methods, like SVM, depend on hand-crafted features, which may not capture complex patterns as effectively as deep learning.
-
4.
Most of the current studies do not adequately account for environmental factors like weather, lighting, limiting their robustness and practical usability in real-world agricultural fields.
-
5.
Some models with additional steps like feature extraction and transformation are computationally intensive, which makes them less suitable for deployment in real-time applications.
The proposed YOLO-LeafNet model aims to address the observed limitations in current plant disease detection models, ensuring improved performance and real-world applicability as follows:
-
1.
To overcome the limitation of using a small or specific dataset, the proposed model incorporates a comprehensive data collection from multiple sources/datasets. This enhances the model’s ability to generalize to diverse plant species and diseases.
-
2.
To overcome fixed image resizing and manual segmentation methods, the proposed model employs an auto-orientation operation, which automatically adjusts to the correct orientation, ensuring the important features are preserved.
-
3.
The proposed YOLO-LeafNet is superior then the traditional ML-based models by providing real-time, multi-class disease detection with faster processing and more efficient feature extraction through its backbone network.
-
4.
To handle variations in weather conditions, lighting, and other environmental factors, the model uses brightness adjustment and contrast enhancement during the data augmentation stage. This helps to make the proposed model more robust and applicable in real-world conditions.
-
5.
The proposed YOLO-LeafNet is based on the optimized YOLO architecture, enhanced with transfer learning and pre-trained weights, enabling fast, real-time object detection while minimizing computational resource requirements.
Proposed methodology
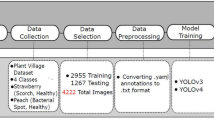
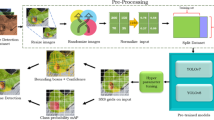
The proposed methodology has seven distinct stages. During the first stage, images of four different plant species’ leaves are gathered from publicly available datasets. In the second stage, image annotation, the acquired images are labelled into eleven distinct classes. In the third stage, image partitioning, these images are divided into three different sets. Next, pre-processing is performed on the acquired images to reduce the training time. In the fifth stage, pre-processed images are augmented by applying augmentation operations to increase the size of the training set. In the sixth stage, images are trained and tested using YoloV5, YoloV8, and the proposed YOLO-LeafNet. Moreover, in the last stage, the performance of YoloV5, YoloV8, and the proposed YOLO-LeafNet has been computed. Figure 1 depicts the flow of the suggested approach.
Proposed method’s framework.
Data acquisition
The proposed model’s first stage is data acquisition. Images of eleven different classes of four species of plants, namely: pepper, grape, corn, and potato, are acquired from five different publicly available datasets on Kaggle. The PlantDoc42,43, Potato disease leaf dataset (PLD)44,45, Corn or Maize leaf disease dataset43,46,47, PlantVillage48, and Grapevine disease images49 are the five different datasets from where images are acquired for this research work. The acquired images involve images of a single leaf and multiple leaves in a single frame. Table 2 shows the complete description of the data acquisition stage.
Data annotation
During the data annotation stage, acquired images are labelled into eleven different classes, namely: Healthy_Potato_Leaf (HPL), Healthy_Grape_Leaf (HGL), Healthy_Corn_Leaf (HCL), Healthy_Bell_Pepper (HBP), Potato_Early_Blight (PEB), Potato_Late_Blight (PLB), Bell_Pepper_Bacterial_Spot (BPBS), Grape_Esca (GE), Grape_Black_Rot (GBR), Grape_Late_Blight (GLB), and Corn_Common_Rust (CCR) using the Roboflow annotation tool50. The acquired images were imported into Roboflow, as it is an adaptable conversion tool designed specifically for computer vision datasets. It supports exporting data into a number of different alternative formats and can import a variety of annotation types with ease. The annotation tools utilized in this work made it easier to accurately define areas of interest within the images by generating bounding boxes. Specifically, in this work, a click-and-drag approach has been utilized to form a defined rectangle that encloses the objects of interest. Labels were methodically applied to the annotated items to indicate their classes or categories after the bounding box annotation. By offering important details about the recognized items inside the drawn bounding boxes, these labels function as essential metadata. After annotation, annotated images are split in an 8:1:1 ratio, out of which a maximum portion of images was utilized to train the proposed model, whereas the rest of the images are for testing and validation, respectively. Table 3 provides the details of the deployed datasets.
Image pre-processing
The third stage of the proposed approach is image pre-processing. The image pre-processing plays a vital role in the manipulation and enhancement of raw images before these are fed into a proposed DL model. The primary goals of the image pre-processing stage encompass enhancing the input image’s quality, optimizing feature extraction, and boosting the overall performance of the model. During this stage, four different image pre-processing operations, namely: resize, static crop, contrast adjustment, and auto-orient, are applied to acquire images for transforming them in order to reduce training time. Image resizing is responsible for standardizing input dimensions. During this operation, all the acquired images are resized to a fixed resolution, ensuring uniformity in size and aspect ratio of the images, which optimizes computational efficiency and enables batch processing during training of the proposed YOLO-LeafNet model. In this proposed work, all of the acquired images were resized to 416 × 416 to ensure that all input image samples have the same dimensions, which allows the model to process them uniformly during training and inference. This operation ensures that key features are retained while eliminating unnecessary background information. After resizing, the order of image pixels is standardized during the auto-orient operation. The auto-orient in the proposed YOLO-LeafNet is responsible for automatically correcting the orientation of the acquired input images to ensure that objects are consistently aligned in standard orientation, regardless of how the images were captured. After the auto-orient operation, contrast adjustment has been applied to the images. The contrast adjustment improves the visibility of features in an image. It highlights structures, features, and patterns that may be difficult to see in low-contrast images. Different lighting circumstances might result in images with differing contrast levels. By adjusting the contrast, images become more uniform and environment-adaptive while also helping to normalize these variations. In this work, the contrast of images is adjusted by applying contrast stretching. The objective of this operation is to make the most of the entire dynamic range of pixel values to improve the image’s visual appeal and suitability for further training or interpretation. Static crop is a fixed cropping technique that is responsible for extracting a specific region of interest on the basis of predetermined dimensions within the image. During static crop operation, acquired images are cropped vertically and horizontally in 10–90% of the range to highlight key characteristics and lessen the impact of irrelevant features by cropping an image to concentrate on a specific area. This may improve the efficacy and efficiency of later processing stages.
Augmentation
Through augmentation, the training process becomes more data-efficient and allows for the generation of more training samples without the need for manual image data collection. Additionally, by introducing variances in the input data, augmentation helps in preventing overfitting and enhances the model’s capacity to handle conditions that vary. In this work, the volume of images in the dataset is increased from 8850 to 23,032 by applying five distinct augmentation operations, namely rotating 90°, flipping, zooming, brightness adjustment, shearing, and rotation. Figure 2 represents the augmentation operation results. During the flipping operation, pre-processed images were flipped both horizontally and vertically to simulate variations in image orientation, which helps the proposed YOLO-LeafNet become invariant to changes in the image’s spatial direction. The flipping operation ensures that the proposed YOLO-LeafNet does not become biased to the particular orientation of the objects and learns to detect features robustly. Rotation operation is another data augmenting technique that significantly enhances the generalization and robustness capabilities of the proposed YOLO-LeafNet model. In the proposed study, images were turned 90 degrees in the clockwise, counter-clockwise, and upside-down directions during rotation by 90 degrees. When the images are rotated at a certain angle, the corresponding bounding box coordinates must be recalculated to align with the new orientation. In the proposed YOLO-LeafNet, images were zoomed by 20% during the zooming operation to increase camera position variability. This augmentation step was designed to mimic variations in camera position and make the proposed model robust to scale. This operation emphasizes finer details of the leaves and plant structures, which could be important for accurately identifying subtle symptoms of disease. Shearing is a geometric transformation that skews the acquired images by shifting their pixels along one axis while keeping the other axis constant. For the purpose of training the suggested model from various angles, a shearing operation was applied to the acquired images. A 15° shear is done both horizontally and vertically in this work. This ensures the proposed model can detect disease patterns even when the images appear distorted due to natural conditions or camera positioning. In the proposed work, the brightness of images is adjusted by -20% (darken), and + 20% (Brighten) for the purpose of making the suggested method more robust to changes in lighting conditions. This operation ensures to simulate diverse illumination scenarios, for instance, overexposed or underexposed environments commonly encountered in field conditions. This operation enhances the model’s ability to detect disease features by training it on a broader range of lighting variations.
Sample from dataset after augmentation operations on CCR image (a) Pre-processed image, (b) Horizontal flip, (c) Vertical Flip, (d) Clockwise 90° rotation, (e) Counter clockwise 90° rotation, (f) Upside down 90° rotation, (g) 20% zoom-in, (h) Shear (15°, 15°), (i) Shear (15°, −15°), (j) Shear (−15°, 15°), (k) Shear (−15°, −15°), (l) Brightness adjustment (-20%), (m) Brightness adjustment (+ 20%), (n) Counter clockwise 45° rotation, (o) Clockwise 45° rotation.
Disease identification
For disease detection investigations, leaves are provided as input to DL-based YOLO Models after augmentation. During the disease identification stage, YoloV5, YoloV8, and the proposed YOLO-LeafNet model are trained to identify 11 distinct classes of leaves. The augmented images are passed as input to YoloV5, YoloV8, and the proposed YOLO-LeafNet for training all models. After completion of the training process, images from the validation set have been utilized for validating all three models. Lastly, the images from the testing set have been utilized for assessing the performance of YoloV5, YoloV8, and the proposed YOLO-LeafNet. The disease identification process using both models, along with the proposed YOLO-LeafNet model, is thoroughly discussed within Sect. 3.5.1, 3.5.2, and 3.5.3, respectively.
Disease identification using YoloV5
YoloV5 is an object detection model built on a CNN. YoloV5 consists of three main components: neck, head, and backbone. The first component of the model is the backbone, which is a pre-trained network utilized for extracting features from an image. This component increases the feature resolution of images by reducing spatial resolution. The neck component of the model allows for generalizing objects on different scales and sizes51. The final stage of the model is the head component, where detection is performed. Figure 3 depicts the architecture of YoloV5.
Architecture of YoloV5.
After augmentation, the leaf images are passed as input to the backbone component of YoloV5. Cross-Stage Partial (CSP) networks have a vital role in extracting valuable features from input images52,53. Spatial Pyramid Pooling (SPP) is a pooling layer that frees the network from its fixed-size restriction54. YoloV5 developed CSPDarknet, the foundation of Darknet, by integrating the CSP network. In order to ensure both accuracy and speed of inference as well as smaller model size, CSPNet reduces parameters and FLOPS by resolving issues with repetitive gradient details within extensive backbone structures and integrating gradient alterations into the feature map53. The neck component is a PANet for generating a feature pyramid. In addition, the feature grid, along with all feature levels, is connected by adaptive feature pooling, which makes the valuable data at each feature level propagate directly to the subnetwork that comes after18. The last component of YoloV5 is the head, which contains convolutional layers for detection. To achieve multi-scale prediction and enable the model to handle objects of varying sizes, the Yolo layer, serving as the head of YoloV5, generates feature maps in three different dimensions: 18 × 18, 36 × 36, and 72 × 7252. For this work, the YoloV5 model was trained with augmented leaf images for 25 epochs and involves 7,273,488 parameters.
Disease detection using YoloV8
YoloV8 is the latest version of the DL-based YOLO object detection model. A novel backbone network, an anchor-free split head, and new loss functions are being introduced in YoloV8 by ultralytics to improve performance metrics55. The CNN used by YoloV8 is composed of two primary sections: the head and the backbone. The core of YoloV8 is an improved version of the CSPDarknet53 architecture. To enhance information flow between the various layers, this design, which has 53 convolutional layers, uses CSP connections. A sequence of fully connected layers comes after a multiple convolutional layer that makes up the head of YoloV8. The bounding boxes, class probabilities, and objectness scores for the objects found in an image are predicted by these layers. One of the key features of YOLOv8 is the integration of a self-attention mechanism within the network’s head, enhancing its ability to capture long-range dependencies and improve feature representation. The self-attention mechanism allows the model to choose to focus on distinct regions of the image and modify the significance of various elements in accordance with their pertinence to the task. YoloV8’s capability to carry out multi-scaled object identification is another crucial feature. To identify items in an image that vary in size and scale, the model employs a feature pyramid network, which is composed of multiple layers56. The model consists of training, validation, and testing phases. During the training phase, the optimization of model parameters was carried out to improve object prediction, and after training, images were validated and tested55. In this work, YoloV8 has been trained for 25 epochs. A total of 11,129,841 parameters have been utilized, and stochastic gradient descent (SGD) was used for optimizing these parameters. Figure 4 shows the architecture of YoloV8.
Architecture of YoloV8.
Disease detection using proposed YOLO-LeafNet
The proposed YOLO-LeafNet architecture is based on the YOLOv8 architecture. YoloV8 itself builds upon the solid foundation of the CSPDarknet53 architecture. The key enhancement in YoloV8 lies in its utilization of cross-stage partial connections among the 53 convolutional layers, facilitating a more efficient flow of information across various layers56. In the proposed YOLO-LeafNet, which aims to further improve upon YOLOv8’s capabilities, two crucial enhancements are introduced in the backbone part of the architecture. Specifically, batch normalization and dropout layers are integrated into the backbone prior to passing data to the spatial pyramid pooling stage. The inclusion of batch normalization layers has been effective in stabilizing and accelerating the training process by normalizing activations across the batch dimension. This normalization technique helps in mitigating issues related to internal covariate shifts, thereby enhancing the overall convergence and performance of the model. Moreover, batch normalization fosters a more consistent flow of information through the network, ultimately leading to improved feature representations. Furthermore, the addition of dropout layers serves as a powerful regularization technique aimed at preventing overfitting. During the training phase fraction of neurons were deactivated, dropout layers discourage the model from too heavily relying on specific features. As a result, the network is driven to acquire more robust and generalizable representations of the input data, ultimately enhancing its ability to generalize well to unseen instances. After dropout layers, the features are passed as input to spatial pyramid pooling, rest of the architecture remains the same as YoloV8. The addition of batch normalization and a dropout layer improves the leave disease detection results. The proposed YOLO-LeafNet model consists of training, validation, and testing phases. During the training phase, the model’s parameters were optimized for improving object prediction, and after training, images were validated and tested55. In this work, the proposed YOLO-LeafNet model has been trained for 25 epochs. Figure 5 shows the architecture of the proposed YOLO-LeafNet.
Architecture of the proposed YOLO-LeafNet model.
Experimental results and comparison
The VGG19. ResNet-50, YoloV5, YoloV8, and proposed YOLO-LeafNet have been implemented for detecting diseases from the leaves of four different plant species. The models are trained and tested on Google Colab utilizing Tesla T4 GPU, using torch 1.13.1 + cu116, and Python version 3.9.16. SGD has been utilized with a value of 0.001 as the learning rate. The models were trained with a learning rate of 0.001, a batch size of 16, and a momentum set to 0.937. The intersection over union (IoU) threshold was set to 0.4, and the confidence threshold was set to 0.5. The computational complexity of the proposed YOLO-LeafNet model was measured at 28.5 Giga Floating Point Operations Per Second (GFLOPS), indicating its lightweight design and suitability for resource-constrained environments. The proposed YOLO-LeafNet achieved an average inference speed per frame of 0.0180 s, which is equivalent to 55.6 frames per second. The performance of all models is evaluated in terms of three different performance metrics, namely: precision, recall, and mAP (mAP50, and mAP50-95). Figures 6, 7, 8, 9, 10, 11 show disease prediction results using YoloV5, YoloV8, and the proposed YOLO-LeafNet model, respectively.
(a): Original image, (b) YoloV5 prediction, (c) YoloV8 prediction, (d) the proposed YOLO-LeafNet prediction.
(a) Original image, (b) YoloV5 prediction, (c) YoloV8 prediction, (d) the proposed YOLO-LeafNet prediction.
Figure 6a shows an image of the original leaf affected by Potato_Early_Blight disease. Figure 6b shows the prediction made by YoloV5, which predicts leaf is affected by Potato_Early_Blight with a confidence score of 60%, and Fig. 6c shows the YoloV8 prediction. YoloV8 predicts the same label with a 65% confidence score, whereas the proposed YOLO-LeafNet model predicts the same label with a confidence score of 86%, shown in Fig. 6d.
(a) Original image, (b) YoloV5 prediction, (c) YoloV8 prediction, (d) the proposed YOLO-LeafNet prediction.
Figure 7a shows multiple leaves in the image affected by Potato_Late_Blight disease. Figure 7b shows the prediction made by YoloV5, which predicts only one instance of Potato_Late_Blight with a confidence score of 30%, Fig. 7c shows the prediction made by YoloV8, which predicts leaves are affected by Potato_Late_Blight with the highest confidence score of 74% for one instance in an image, whereas the proposed YOLO-LeafNet model predicts the same label with the highest confidence score of 84%, shown in Fig. 7d.
(a) Original image, (b) YoloV5 prediction, (c) YoloV8 prediction, (d) the proposed YOLO-LeafNet prediction.
Figure 8a shows a leaf image affected by Grape_Esca disease. Figure 8b shows the prediction made by YoloV5, which predicts leaf is affected by Grape_Esca with 76% confidence score, and the YoloV8 predicts the same label with the confidence score of 97%, shown in Fig. 8c, whereas proposed YOLO-LeafNet predicts the same label with 99% confidence, shown in Fig. 8d.
(a) Original image, (b) YoloV5 prediction, (c) YoloV8 prediction, (d) The proposed YOLO-LeafNet prediction.
Figure 9a shows a leaf image affected by Grape_Leaf_Blight disease. Figure 9b shows the prediction made by YoloV5, which predicts leaf is affected by Grape_Leaf_Blight with 76% confidence score, and YoloV8 model predicts the same label with the confidence score of 97%, shown in Fig. 9c, whereas the proposed YOLO-LeafNet predicts the same label with 99% confidence, as shown in Fig. 9d.
Figure 10a shows multiple leaves in the image affected by Grape_Black_Rot disease. Figure 10b shows the prediction made by YoloV5, which predicts Grape_Black_Rot only for one instance in an image with confidence score of 77%, for one instance in an image, whereas YoloV8 and the proposed YOLO-LeafNet model predicts the same label for three instances in same image with the highest confidence score of 96%, shown in Fig. 10c and d respectively.
(a) Original image, (b) YoloV5 prediction, (c) YoloV8 prediction, (d) the proposed YOLO-LeafNet prediction.
Figure 11a shows a leaf image affected by Corn_Common_Rust disease. Figure 11b shows the prediction made by YoloV5, which predicts leaf is affected by Corn_Common_Rust with 65% confidence score, and Fig. 11c shows the prediction made by YoloV8, which predicts leaf is affected by Corn_Common_Rust with 95% confidence score, whereas Fig. 11d shows the proposed model prediction. The proposed YOLO-LeafNet predicts the same label with a confidence score of 97%.
Performance metrics
The performance metrics evaluate how well VGG19, ResNet-50, YoloV5, YoloV8, and the proposed YOLO-LeafNet perform for detecting all eleven classes of plant leaves, which have been measured in terms of recall, precision, and mAP. The precision and recall metrics are computed based on the confusion matrix, while the area under the precision-recall curve contributes to the calculation of mAP.
Confusion matrix
The effectiveness of yolo based models is assessed by the confusion matrix. It is a matrix where the rows represent the actual classes, the columns represent the predicted classes, and each cell indicates the number of instances corresponding to a specific combination of actual and predicted classes. The confusion metrics for YoloV5, YoloV8, and the proposed YOLO-LeafNet models are shown in Figs. 12 and 13, and 14, respectively.
Confusion matrix for YOLOv5 model.
Confusion matrix for YoloV8 model.
Confusion matrix for the proposed YOLO-LeafNet model.
Precision
The proportion of accurately predicted positive instances among all positive instances that were predicted is known as precision. It is calculated as:
where FP stands for False Positive and TP stands for True Positive.
Recall
Recall counts how many of the actual positive instances were accurately predicted out of all the positive instances. It is computed as:
where FN stands for False Negative.
mAP (mean average precision)
The precision-recall curve is first computed for each class by plotting the precision and recall numbers at various IoU thresholds. The average precision (AP) for that class is then evaluated using the area under the precision-recall curve (AUC-PR). In our study, mAP50 and mAP50-95 are calculated.
-
a.
mAP50
It calculates the AP of the utilized model at a fixed IoU threshold of 50%. The IoU calculates the degree of overlap between the anticipated and actual bounding boxes. In simple words, a predicted object is considered correct if it overlaps with the ground truth object by at least 50%. It reflects the model’s ability to localize objects with moderate precision.
-
b.
mAP50-95
It is an estimate of AP over various IoU levels between 0.5 and 0.95. The model’s effectiveness is evaluated using recall and precision metrics with a step size of 0.05 at IoU thresholds varying from 0.5 to 0.95 for calculating mAP50-95. This metric captures the model’s performance across varying levels of localization accuracy, from coarse to very precise.
Precision, Recall, mAP50, and mAP50-95 are computed for all models. Figures 15, 16, 17, 18 and 19 show experimental results of YoloV5, VGG19, ResNet-50, YoloV8, and YOLO-LeafNet models, respectively, class-wise.
Precision, Recall, mAP50, and mAP50-95 attained by YoloV5.
Precision, recall, mAP50, mAP50-95 attained by VGG-19.
Precision, Recall, mAP50, mAP50-95 attained by ResNet-50.
Precision, recall, mAP50, mAP50-95 attained by YoloV8.
Precision, recall, mAP50, mAP50-95 attained by the proposed YOLO-LeafNet.
To show the effectiveness and comparison of all models, experimental findings are summarized in Fig. 17. Among all models, YoloV5 has the least performance. VGG19 and ResNet-50 show improvement due to residual learning. YoloV8 and the proposed model have shown significant improvements. The proposed model shows the best balance between precision and recall.
Comparison of precision, recall, F1-Score, mAP50, and mAP50-95.
Figure 20 shows the comparison of five models in terms of precision, recall, F1-score, mAP50, and mAP50-95. The graph highlights that the proposed YOLO-LeafNet model outperforms all models, depicting its effectiveness in real-time plant leaf disease detection. Higher mAP indicates the ability of the most accurate detection.
Comparison with existing approaches
The performance of the proposed approach is compared with various state-of-the-art approaches on the basis of the dataset from which images were acquired, the model utilized for detecting diseases, how many classes of diseases were detected by the respective model, and what value of performance metrics was attained by the models. A comparison of the proposed approach with existing state-of-the-art methods is presented in Table 4.
Discussion
Evaluating the performance of the YoloV5 model
The YoloV5 attained an overall precision of 86.1%, a recall of 86.8%, mAP50 of 94.4%, and mAP50-95 of 81.5% for detecting all eleven classes of plant leaves, including healthy and diseased ones. The Fig. 17 shows the performance metrics achieved for each class individually using YOLOv5.
Diseases like GBR, CCR, PEB, PLB, GLB, GE, and BPBS are particularly well detected, with all three metrics, recall, precision, and mAP, consistently maintaining a high level. Despite having fewer training images or instances of the class in the provided dataset, the HCL has good overall performance scores. The HGL class exhibits average performance across all metrics and is expected to improve with a high instance of the class in the provided dataset. HPL, as compared to the other ten classes, has performed poorly in terms of recall.
Evaluating the performance of the YoloV8 model
The YoloV8 attained an overall precision of 97.7%, a recall of 97.5%, mAP50 of 98.4%, and mAP50-95 of 91.5% for detecting all eleven classes of plant leaves, including healthy and diseased ones. Performance metrics attained for each class individually using YoloV8 are shown in Fig. 18.
Diseases like CCR, PLB, PEB, GBR, GLB, GE, and BPBS are particularly well detected, with all three metrics, precision, recall, and mAP, being high and stable. Despite having fewer training images or instances of the class in the provided dataset, healthy leaves like HPL, HCL, HBPL, and HGL have good overall performance scores. In comparison to the other ten classes, CCR has average performance despite having high training images or instances of the class in the provided dataset.
Evaluating the performance of the proposed YOLO-LeafNet model
The proposed YOLO-LeafNet attained an overall precision of 98.5%, a recall of 98.0%, mAP50 of 99%, and mAP50-95 of 94% for detecting all eleven classes of plant leaves, including healthy and diseased ones. The proposed YOLO-LeafNet provides efficient prediction with a high confidence score for all the classes. The performance metrics attained for each class individually using YoloV8 are shown in Fig. 19.
Evaluating YoloV5, YoloV8, and the proposed YOLO-LeafNet model
While comparing the disease detection performance of YOLOv5, YOLOv8, and proposed YOLO-LeafNet across distinct classes, several notable patterns emerge. The YoloV5 utilized in this work shows decent performance across most classes, with notable strengths with high recall for classes like PEB and PLB, although its precision is very significant, leading to higher false positives in certain categories, like healthy potato leaf. The YoloV8 utilized in this work exhibits improvement over the YoloV5, showcasing higher precision and recall for many classes, indicating more accurate disease detection overall. However, the proposed YOLO-LeafNet consistently outperforms both YoloV5 and YoloV8, by attaining the highest precision of 98.5%, 98% of recall, and mAP of 94%. Overall, YoloV8 presents advancements over YoloV5. Proposed YOLO-LeafNet stands out as the most effective model, showcasing superior performance across all the classes by attaining the highest precision, recall, and mAP. The overall comparison of both versions of YOLO models utilized in this work, along with the proposed YOLO-LeafNet results in terms of all performance metrics, is shown in Fig. 19.
Discussion on comparison of the proposed methodology with existing approaches
The effectiveness of both the models, along with the proposed YOLO-LeafNet, has been assessed by comparing their performance against six existing state-of-the-art approaches. Table 4 illustrates the comparison between the proposed approach with the existing approaches.
The proposed YOLO-LeafNet outperformed VGG16 in aspects of recall and precision for detecting potato diseases25, whereas YoloV5, utilized in this work, falls behind by a slight difference in values of performance metrics. While making the performance comparison, both of the utilized models and the proposed YOLO-LeafNet model outperformed (Mathew & Mahesh, 2022)57 YoloV5 model when the performance for detecting bacterial bell pepper spot was compared. The YoloV5 utilized in the proposed work outperformed the YoloV5 utilized by57 for two main reasons: First, before being fed into a proposed DL model, the pre-processing approaches utilized in this work improve the raw images and standardized image pixels. In order to guarantee that every input image sample has the same dimensions and that the model can process them consistently during training and inference, 416 × 416 images were used as input for the proposed method. Furthermore, by emphasizing structures, details, and patterns that could be hard to notice in low-contrast images, the contrast adjustment applied at this stage enhances the visibility of features in an image. Images can be made more uniform and environment-adaptive by altering the contrast, which also helps to normalize these differences. The cropping process has also been applied to sample images in this work in order to emphasize important elements and reduce the impact of unimportant features. This technique allows an image to focus on a particular area that was not included in the work that was suggested in57. Second, the augmentation stage, which was likewise absent from the work proposed in57, helps minimize overfitting by introducing variances in the input data and enhancing the model’s capacity to manage variation in conditions. In58, the authors applied ten different models for detecting corn diseases, but the performance of all the applied models was outperformed by the suggested YOLO-LeafNet. DCNN35 attained only an overall 40.6% mAP for detecting diseases from apple, potato, tomato, and corn, whereas both the model along with proposed YOLO-LeafNet in this work attained mAP around 95%. The authors in41 compared the performance of five different versions of YOLO models for detecting three classes of maize leaves. Both the model utilized in this work, along with the proposed YOLO-LeafNet, outperformed YOLOv3-tiny, YOLOv5s41 in aspects of all three performance metrics. The YoloV5 utilized in this work outperformed YOLOv441 in aspects of precision, whereas YOLO8 and the proposed YOLO-LeafNet performed better than YOLOv441 in all aspects of the performance metrics. The YoloV5 used in this work outperformed YOLO7s41 in aspects of mAP. In contrast, YoloV8 utilized in this work and proposed YOLO-LeafNet performed better than YOLO7s41 in terms of recall and mAP. For detecting diseases from corn/maize leaves, both YoloV5, YoloV8, along with the proposed YOLO-LeafNet model, outperformed YoloV8n41 in aspects of recall, precision, and mAP. Several recent studies support the advancement of YOLO-based models in diverse detection tasks, which align with the objectives of YOLO-LeafNet. For instance, authos in59 highlight the effectiveness of deep learning in extracting spatial patterns from complex environments, which parallels our model’s ability to identify disease features across varied leaf textures. Authors in60 propose a lightweight YOLO-PDGT model for unripe pomegranate detection, emphasizing model efficiency and speed key considerations also addressed in our framework for real-time plant disease diagnosis. Furthermore, in61 (2025) enhance YOLO with attention mechanisms for poultry tracking, a concept we integrate into YOLO-LeafNet to boost detection accuracy across multiple plant species and disease conditions.
Conclusion and future scope
The effect of plant diseases on crop quality and productivity can be significantly reduced by early detection of plant diseases utilizing DL-based technologies. In this work, the diseases have been detected from the leaf images of grape, potato, corn, and bell pepper using two existing versions of YOLO, namely: YoloV5, YoloV8, and a new approach named YOLO-LeafNet has been proposed. The results reveal that YoloV8 offers precision, recall, and mAP of more than 97% and outperforms YoloV5 in terms of all the performance measures. Moreover, YOLOv8 outperformed the performance of various existing approaches; the proposed YOLO-LeafNet outperformed YOLOv8 in terms of all the performance measures. To produce the best possible disease detection results, pre-processing and data augmentation operations must be carefully chosen. In future work, more plant species can be added to the model, as well as the proposed YOLO-LeafNet approach can be integrated with drones for real-time field monitoring and IoT sensors for environmental data collection. This integration could facilitate efficient, large-scale monitoring of crops and early detection of disease, enhancing decision-making in agriculture. Furthermore, integrating the YOLO-LeafNet model with a mobile application would provide farmers with easy access to identifying plant diseases at an early stage.
Data availability
The datasets used in this study are publically available at https://www.kaggle.com/datasets/raman31/leafnet-dataset. Acquired from: (1) https://www.kaggle.com/datasets/emmarex/plantdisease. (2) https://www.kaggle.com/datasets/abdulhasibuddin/plant-doc-dataset. (3) https://www.kaggle.com/datasets/rizwan123456789/potato-disease-leaf-datasetpld. (4) https://www.kaggle.com/datasets/smaranjitghose/corn-or-maize-leaf-disease-dataset. (5) https://www.kaggle.com/datasets/piyushmishra1999/plantvillage-grape.
References
Share of Agriculture in Employment Rose. Manufacturing Declined in 2021-22: PLFS. https://thewire.in/economy/share-of-agriculture-in-employment-rose-manufacturing-declined-in-2021-22-plfs (Accessed 29 Mar 2023).
Page 262 - economic_survey_2021–2022. https://www.indiabudget.gov.in/economicsurvey/ebook_es2022/files/basic-html/page262.html (Accessed 29 Mar 2023).
Ramanjot & Mittal, U. A Review of various plant disease detection algorithms. In 2022 10th Int. Conf. Reliab. Infocom Technol. Optim. (Trends Futur. Dir. ICRITO 2022, no. Ml), https://doi.org/10.1109/ICRITO56286.2022.9964538 (2022).
Dubey, S. R. & Jalal, A. S. Detection and classification of apple fruit diseases using complete local binary patterns. In 2012 3rd Int. Conf. Comput. Commun. Technol., vol. 2012, 346–351 https://doi.org/10.1109/ICCCT.2012.76 (2012).
Sannakki, S. S., Rajpurohit, V. S., Nargund, V. B. & Kulkarni, P. Diagnosis and classification of grape leaf diseases using neural networks. In 2013 4th Int. Conf. Comput. Commun. Netw. Technol. (ICCCNT), vol. 2013, 1–5 https://doi.org/10.1109/ICCCNT.2013.6726616 (2013).
Joshua, S. V. et al. Crop yield prediction using machine learning approaches on a wide spectrum. Computers Mater. Continua. 72(3), 5663–5679. https://doi.org/10.32604/cmc2022.027178 (2022).
Malathy, S., Karthiga, R., Swetha, K. & Preethi, G. Disease detection in fruits using image processing. In 2021 6th International Conference on Inventive Computation Technologies (ICICT), 747–752 https://doi.org/10.1109/ICICT50816.2021.9358541 (2021).
Durmus, H., Gunes, E. O. & Kirci, M. Disease detection on the leaves of the tomato plants by using deep learning. In 2017 6th Int. Conf. Agro-Geoinformatics, 1–5 https://doi.org/10.1109/Agro-Geoinformatics.2017.8047016 (2017).
Liu, B., Zhang, Y., He, D. & Li, Y. Identification of apple leaf diseases based on deep convolutional neural networks. Symmetry 10(1), 11. https://doi.org/10.3390/sym10010011 (2017).
Howlader, M. R., Habiba, U., Faisal, R. H. & Rahman, M. M. Automatic recognition of guava leaf diseases using deep convolution neural network. In 2019 International Conference on Electrical, Computer and Communication Engineering (ECCE), 1–5. https://doi.org/10.1109/ECACE.2019.8679421 (2019).
Jadhav, S. B., Udupi, V. R. & Patil, S. B. Identification of plant diseases using convolutional neural networks. Int. J. Inf. Technol. 13(6), 2461–2470. https://doi.org/10.1007/s41870-020-00437-5 (2021).
Francis, M. & Deisy, C. Disease detection and classification in agricultural plants using convolutional neural networks - a visual understanding. In 2019 6th Int. Conf. Signal Process. Integr. Networks, 1063–1068 https://doi.org/10.1109/SPIN.2019.8711701 (2019).
Kibriya, H., Rafique, R., Ahmad, W. & Adnan, S. M. Tomato leaf disease detection using convolution neural network. In 2021 Int. Bhurban Conf. Appl. Sci. Technol. 346–351. https://doi.org/10.1109/IBCAST51254.2021.9393311 (2021).
Akshai, K. P. & Anitha, J. Plant disease classification using deep learning. In 2021 3rd Int. Conf. Signal Process. Commun., 407–411 https://doi.org/10.1109/ICSPC51351.2021.9451696 (2021).
Chen, J. et al. A multiscale lightweight and efficient model based on YOLOv7: applied to citrus orchard. Plants https://doi.org/10.3390/plants11233260 (2022).
Morbekar, A., Parihar, A. & Jadhav, R. Crop disease detection using YOLO. In 2020 Int. Conf. Emerg. Technol. INCET 2020, 1–5 https://doi.org/10.1109/INCET49848.2020.9153986 (2020)
Ponnusamy, V., Coumaran, A., Shunmugam, A. S., Rajaram, K. & Senthilvelavan, S. Smart glass: Real-time leaf disease detection using YOLO transfer learning. In 2020 IEEE Int. Conf. Commun. Signal. Process. 1150–1154. https://doi.org/10.1109/ICCSP48568.2020.9182146 (2020).
Wang, X. & Liu, J. Tomato anomalies detection in greenhouse scenarios based on YOLO-Dense. Front. Plant. Sci. 12, 1–11. https://doi.org/10.3389/fpls.2021.634103 (2021).
Padol, P. B. & Yadav, A. A. SVM classifier based grape leaf disease detection. In 2016 Conference on Advances in Signal Processing (CASP), 175–179 https://doi.org/10.1109/CASP.2016.7746160 (2016).
Vetal, S. Tomato plant disease detection using image processing. Ijarcce 6(6), 293–297. https://doi.org/10.17148/ijarcce.2017.6651 (2017).
Mishra, S., Sachan, R. & Rajpal, D. Deep convolutional neural network based detection system for Real-time corn plant disease recognition. Procedia Comput. Sci. 167, 2003–2010. https://doi.org/10.1016/j.procs.2020.03.236 (2020).
Zhang, S., Zhang, S., Zhang, C., Wang, X. & Shi, Y. Cucumber leaf disease identification with global pooling dilated convolutional neural network. Comput. Electron. Agric. 162, 422–430. https://doi.org/10.1016/j.compag.2019.03.012 (2019).
Khamparia, A. et al. Seasonal crops disease prediction and classification using deep convolutional encoder network. Circuits Syst. Signal. Process. 39(2), 818–836. https://doi.org/10.1007/s00034-019-01041-0 (2020).
Wu, Q., Zhang, K. & Meng, J. Identification of soybean leaf diseases via deep learning. J. Inst. Eng. Ser. A. 100(4), 659–666. https://doi.org/10.1007/s40030-019-00390-y (2019).
Tiwari, D. et al. Potato leaf diseases detection using deep learning. In Proc. Int. Conf. Intell. Comput. Control Syst. ICICCS 2020, no. January 2021, 461–466 https://doi.org/10.1109/ICICCS48265.2020.9121067 (2020).
Bharali, P., Bhuyan, C. & Boruah, A. Plant disease detection by leaf image classification using convolutional neural network. Commun. Comput. Inform. Sci. 1025 CCIS, 194–205. https://doi.org/10.1007/978-981-15-1384-8_16 (2019).
Yadhav, S. Y., Senthilkumar, T., Jayanthy, S. & Kovilpillai, J. J. A. Plant disease detection and classification using CNN model with optimized activation function. In 2020 International Conference on Electronics and Sustainable Communication Systems (ICESC), 564–569 https://doi.org/10.1109/ICESC48915.2020.9155815 (2020).
Deepa, N. R. & Nagarajan, N. Kuan noise filter with Hough transformation based reweighted linear program boost classification for plant leaf disease detection. J. Ambient Intell. Humaniz. Comput. 12(6), 5979–5992. https://doi.org/10.1007/s12652-020-02149-x (2021).
Jasim, M. A. & Al-Tuwaijari, J. M. Plant leaf diseases detection and classification using image processing and deep learning techniques. In Proc. 2020 Int. Conf. Comput. Sci. Softw. Eng. CSASE 2020 259–265 https://doi.org/10.1109/CSASE48920.2020.9142097 (2020).
Zhang, K., Wu, Q. & Chen, Y. Detecting soybean leaf disease from synthetic image using multi-feature fusion faster R-CNN. Comput. Electron. Agric. 183, 106064. https://doi.org/10.1016/j.compag.2021.106064 (2021).
Chen, J., Zhang, D., Zeb, A. & Nanehkaran, Y. A. Identification of rice plant diseases using lightweight attention networks. Expert Syst. Appl. 169, 114514. https://doi.org/10.1016/j.eswa.2020.114514 (2021).
Vallabhajosyula, S., Sistla, V. & Kolli, V. K. K. Transfer learning-based deep ensemble neural network for plant leaf disease detection. J. Plant. Dis. Prot. 129(3), 545–558. https://doi.org/10.1007/s41348-021-00465-8 (2022).
Sunil, C. K., Jaidhar, C. D. & Patil, N. Cardamom plant disease detection approach using EfficientNetV2. IEEE Access. 10, 789–804. https://doi.org/10.1109/ACCESS.2021.3138920 (2022).
Cap, Q. H., Uga, H., Kagiwada, S. & Iyatomi, H. LeafGAN: An effective data augmentation method for practical plant disease diagnosis. IEEE Trans. Autom. Sci. Eng., 19(2), 1258–1267. https://doi.org/10.1109/TASE.2020.3041499.
Gajjar, R., Gajjar, N., Thakor, V. J., Patel, N. P. & Ruparelia, S. Real-time detection and identification of plant leaf diseases using convolutional neural networks on an embedded platform. Vis. Comput. 38(8), 2923–2938. https://doi.org/10.1007/s00371-021-02164-9 (2022).
Sai Reddy, B. & Neeraja, S. Plant leaf disease classification and damage detection system using deep learning models. Multimed Tools Appl. 81(17), 24021–24040. https://doi.org/10.1007/s11042-022-12147-0 (2022).
Harakannanavar, S. S., Rudagi, J. M., Puranikmath, V. I., Siddiqua, A. & Pramodhini, R. Plant leaf disease detection using computer vision and machine learning algorithms. Glob. Trans. Proc. 3(1), 305–310. https://doi.org/10.1016/j.gltp.2022.03.016 (2022).
Haridasan, A., Thomas, J. & Raj, E. D. Deep learning system for paddy plant disease detection and classification. Environ. Monit. Assess. https://doi.org/10.1007/s10661-022-10656-x (2023).
Arshaghi, A., Ashourian, M. & Ghabeli, L. Potato diseases detection and classification using deep learning methods. Multimed Tools Appl. 82(4), 5725–5742. https://doi.org/10.1007/s11042-022-13390-1 (2023).
Javidan, S. M., Banakar, A., Vakilian, K. A. & Ampatzidis, Y. Diagnosis of grape leaf diseases using automatic K-means clustering and machine learning. Smart Agric. Technol. 3, 100081. https://doi.org/10.1016/j.atech.2022.100081 (2023).
Khan, F. et al. A mobile-based system for maize plant leaf disease detection and classification using deep learning. Front. Plant. Sci. 14, 1–18. https://doi.org/10.3389/fpls.2023.1079366 (2023).
Plant Doc Dataset | Kaggle. https://www.kaggle.com/datasets/abdulhasibuddin/plant-doc-dataset (Accessed 01 Apr 2024).
Singh, D. et al. PlantDoc: A dataset for visual plant disease detection. In ACM Int. Conf. Proceeding Ser., 249–253 https://doi.org/10.1145/3371158.3371196 (2020).
Potato Disease Leaf Dataset(PLD). | kaggle.https://www.kaggle.com/datasets/rizwan123456789/potato-disease-leaf-datasetpld (Accessed 01 Apr 2024).
Rashid, J. et al. Multi-level deep learning model for potato leaf disease recognition. Electronics. 10(17), 2064 https://doi.org/10.3390/electronics10172064 (2021).
Corn or Maize Leaf Disease Dataset | Kaggle. https://www.kaggle.com/datasets/smaranjitghose/corn-or-maize-leaf-disease-dataset (Accessed 01 Apr 2024)
Geetharamani, G. Identification of plant leaf diseases using a nine-layer deep convolutional neural network. Comput. Electr. Eng. 76, 323–338. https://doi.org/10.1016/j.compeleceng.2019.04.011 (2019).
PlantVillage Dataset | Kaggle. https://www.kaggle.com/datasets/emmarex/plantdisease (Accessed 01 Apr 2024).
Grapevine Disease Images | Kaggle. https://www.kaggle.com/datasets/piyushmishra1999/plantvillage-grape (Accessed 01 Apr 2024).
Inc, R. Roboflow Annotate: Label Faster Than Ever, (2023). https://roboflow.com/annotate (Accessed 17 Nov 2024).
YOLO v5 model architecture [Explained]. https://iq.opengenus.org/yolov5/ (Accessed 30 Mar 2024).
YOLO V5 — Explained and Demystified – Towards AI. https://towardsai.net/p/computer-vision/yolo-v5-explained-and-demystified (Accessed 31 Mar 2024).
Wang, C. Y. et al. CSPNet: A new backbone that can enhance learning capability of CNN. In 2020 IEEE/CVF Conference on Computer Vision and Pattern Recognition Workshops (CVPRW), vol. 2020, 1571–1580. https://doi.org/10.1109/CVPRW50498.2020.00203 (2020).
Spatial Pyramid Pooling Explained | Papers With Code. https://paperswithcode.com/method/spatial-pyramid-pooling (Accessed 11 Apr 2024).
YOLOv8 Docs. https://docs.ultralytics.com (Accessed 30 Mar 2024).
Understanding. YOLOv8 architecture, applications & features. https://www.labellerr.com/blog/understanding-yolov8-architecture-applications-features/ (Accessed 18 Nov 2024).
Mathew, M. P. & Mahesh, T. Y. Leaf-based disease detection in bell pepper plant using YOLO v5. Signal. Image Video Process. 16(3), 841–847. https://doi.org/10.1007/s11760-021-02024-y (2022).
Fraiwan, M., Faouri, E. & Khasawneh, N. Classification of corn diseases from leaf images using deep transfer learning. Plants 11(20), 1–14. https://doi.org/10.3390/plants11202668 (2022).
Zhou, L., Cai, J. & Ding, S. The identification of ice Floes and calculation of sea ice concentration based on a deep learning method. Remote Sens. 15(10), 2663. https://doi.org/10.3390/rs15102663 (2023).
Fan, Z. et al. YOLO-PDGT: A lightweight and efficient algorithm for unripe pomegranate detection and counting. Measurement 254, 117852. https://doi.org/10.1016/j.measurement.2025.117852 (2025).
Jiang, D., Wang, H., Li, T., Gouda, M. A. & Zhou, B. Real-time tracker of chicken for poultry based on attention mechanism-enhanced YOLO-Chicken algorithm. Comput. Electron. Agric. 237, 110640. https://doi.org/10.1016/j.compag.2025.110640 (2025).
Acknowledgements
This work was supported by King Saud University, Riyadh, Saudi Arabia, through Ongoing Research Funding program - Research Chairs, (ORF-RC-2025-2800).
Author information
Authors and Affiliations
Contributions
Ramanjot: Conceptualization, Data Curation, Writing – Original Draft.Usha Mittal: Methodology, Formal Analysis, Writing – Review & Editing.Ankita Wadhawan: Software Development, Validation, Visualization.Ahmad Almogren: Methodology, Validation, Resources, Project Administration, Writing – Review & Editing.Jimmy Singla: Investigation, Writing – Review & Editing.Salil Bharany: Conceptualization, Writing – Review & Editing, Supervision, Project Administration.Seada Hussen: Data Curation, Writing – Review & Editing, Supervision.Ateeq Ur Rehman: Conceptualization, Methodology, Writing – Review & EditingAsma A. Al-Huqail: Conceptualization, Supervision, Resources, Writing – Review & Editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Kaur, R., Mittal, U., Wadhawan, A. et al. YOLO-LeafNet: a robust deep learning framework for multispecies plant disease detection with data augmentation. Sci Rep 15, 28513 (2025). https://doi.org/10.1038/s41598-025-14021-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-14021-z
This article is cited by
-
A novel approach for disease and pests detection in potato production system based on deep learning
Scientific Reports (2026)
-
Lightweight scalable deep learning framework for real time detection of potato leaf diseases
Scientific Reports (2026)