Abstract
Generating novel molecular structures with desired pharmacological and physicochemical properties is challenging due to the vast chemical space, complex optimization requirements, predictive limitations of models, and data scarcity. This study focuses on investigating the problem of posterior collapse in variational autoencoders, a deep learning technique used for de novo molecular design. Various generative variational autoencoders were employed to map molecule structures to a continuous latent space and vice versa, evaluating their performance as structure generators. Most state-of-the-art approaches suffer from posterior collapse, limiting the diversity of generated molecules. To address this challenge, a novel approach termed PCF-VAE was introduced to mitigate the issue of posterior collapse, reduce the complexity of SMILES representations, and enhance diversity in molecule generation. In comparison to state-of-the-art models, PCF-VAE has been evaluated and compared in the MOSES benchmark at different diversity levels. Depending on the diversity level, PCF-VAE has a validity of 98.01% at D = 1, 97.10% at D = 2, and 95.01% at D = 3. It is important to note that PCF-VAE effectively generates molecules with a 100% unique structure. Both intDiv and intDiv2 are measures of internal diversity; intDiv2 ranges from 85.87 to 86.33% and intDiv ranges from 85.87 to 89.01%. Additionally, at D = 1, D = 2, and D = 3, PCF-VAE generates 93.77%, 94.71%, and 95.01% novel molecules, respectively. The results indicate that this research provides valuable insights into the challenges of molecular generation and contributes to the design of novel molecules with desired properties.
Similar content being viewed by others
Introduction
The primary objective of molecular design for novel materials and pharmaceuticals is to directly generate compounds possessing the intended properties. Apparently, this is a difficult task due to the vastness, discretness, and disorganization of a molecular space, which is composed of a variety of molecules. Specifically, a total of \(10^8\) molecules have been successfully generated, while it is approximated that there exist \(10^{23}\)–\(10^{60}\) molecules resembling drugs1,2. Although experimental techniques have made significant progress, it remains exceedingly challenging to identify molecules best suited for particular applications only through experimental methods.
In the field of pharmaceutical research, the task of identifying compounds with desirable characteristics is important. Various techniques, such as enumerated virtual libraries and de novo drug design, have been developed in order to identify promising compounds. The utilization of de novo drug design and other techniques broadens the scope of the search beyond the limitations of current physical compound libraries. This ultimately raises the probability of identifying drug candidates with particular and desired characteristics, as well as augmenting chemical diversity. Over the past decade, generative models have been developed specifically for the purpose of de novo drug design. The models undergo training that generates molecules from a predefined training set and are subsequently sampled to generate new molecules. The integration of reinforcement learning into generative models enables the generation of compounds that effectively accomplish specific objectives. The Simplified Molecular-Input Line-Entry System (SMILES) is widely used by many de novo generators to represent molecules. Linear molecular line notations such as SMILES are compatible with established natural language processing (NLP) generative models3. Previous research have shown that recurrent neural networks (RNNs), autoencoders, generative adversarial networks (GANs), and other generative models can be used for partial and quantitative de novo drug design4.
The generation of desirable molecules is a well recognized challenge in the design of pharmaceutical molecules. The properties of molecules exhibit a strong correlation, such that any alteration in one property leads to equally significant changes in the other properties. Furthermore, the generative models fail to effectively learn the molecular patterns in the SMILES representation of molecules, resulting in the production of undesired and invalid molecules. Additional, a prevalent problem encountered by the majority of VAE based models is their susceptibility to the posterior collapse effect, which results in the production of highly similar molecules.
Contributions
The PCF-VAE technique, which is an unique approach, has been implemented to generate novel molecules. This novel approach entails the reparameterization of the loss function of the VAE, specifically designed for the purpose of generating molecules. In order to improve the robustness of the input data for PCF-VAE, the SMILES strings are transformed into GenSMILES. This process simplifies the complexity of the SMILES strings and leads to the development of a larger range of feasible molecules.
Moreover, a novel methodology has been developed to incorporate diversity into the generated molecules. This entails integrating a diverse layer between the latent space and the decoder of PCF-VAE, which allows for precise manipulation of the variety and integrity of the generated molecules. PCF-VAE exhibits remarkable performance in comparison to other state-of-the-art (SOTA) methods. The key contributions of this proposed model are delineated as follows:
-
The molecules represented by the SMILES notations are transformed into GenSMILES. GenSMILES has been specifically designed to preserve the intrinsic semantic information of the molecules. This structural adaption enables the probabilistic model to understand long-term relationships inside the SMILES strings while simultaneously reducing their complexity.
-
Properties such as molecular weight, LogP (lipophilicity), and TPSA (topological polar surface area) are incorporated into GenSMILES representations of molecules. This integration makes it possible for PCF-VAE to generate molecules that meet specific requirements for acceptability.
-
A diversity parameter is used in PCF-VAE to control the diversity and validity of the molecules.
Related work
Recent years have seen the usefulness of generative deep learning models in several domains such as remote sensing, image analysis, text generation, and molecule design5,6,7. Throughout the training phase, the model described in these papers exhibits the capacity to gain knowledge about the structural arrangement of pharmaceutical molecules. Consequently, the trained models have the ability to generate desired molecules8,9,10. Gómez-Bombarelli et al.11 use a RNN based VAE to design pharmacological compounds. Notably, the generated latent space is effectively optimized using Bayesian optimization techniques. The training dataset for the proposed model comprises drug molecules structured in SMILES format. The study conducted by Blaschke et al.12 employs AAE13 in combination with Bayesian optimization to produce potential drugs that specifically target the dopamine type 2 receptor. Comparing the molecules produced by VAE and AAE, Kadurin et al.14 also explores the application of NLP models for drug design. The input for these NLP models is the string representation of molecules SMILES15,16,17,18,19. Yuan et al.16 have focused on designing molecules for a specific target and conducting a comprehensive evaluation of the resulting molecules. The application of transfer learning is demonstrated by Gupta et al.18 and Segler et al.17 in the generation of optimal molecules, particularly in situations when the available training data is restricted. Reinforcement learning approaches are employed to adapt pre-trained generative models20,21,22. The objective of these modifications is to enforce several property constraints on the generated collection of molecules.
VAE has been widely used in the field of generative modeling and drug discovery. Some of the most related SOTA methods that uses VAE for the design of the drug are summarized below:
In their study, Zhang et al.23 propose D-VAE, an innovative VAE designed for directed acyclic graphs (DAGs). D-VAE incorporates an asynchronous message passing mechanism that takes into account computation dependencies. Through Bayesian optimization, this model efficiently encodes DAG computations into a continuous latent space, thereby enabling optimization tasks such as Bayesian network structure learning and neural architecture search.
Gómez et al. utilize an autoencoder to convert discrete molecular representations into continuous vectors, facilitating efficient exploration and optimization of chemical space. This technique, enhanced by gradient-based optimization, supports the creation of novel molecules and the forecasting of their characteristics11.
Lim et al. introduce the All SMILES VAE, leveraging RNNs across multiple SMILES strings to enable smooth and nearly bijective latent representations of molecular properties. This method significantly boosts the efficiency of property regression and optimization processes27.
Tempke et al. present an innovative VAE approach, generating over 7,000,000 reactions from 7000 starting reactions to autonomously produce chemical reactions, mitigating dataset biases. This expansion increases molecular diversity, posing challenges for experimental validation25.
Rigoni et al. propose the Conditional Constrained Graph Variational Autoencoder, which optimizes molecule generation using atom valence histograms. This model improves upon existing frameworks by better incorporating chemical principles, achieving enhanced performance on the QM9 and ZINC datasets26.
Liu et al.28 propose a dual-channel variational autoencoder (SmilesGEN) that jointly encodes molecular SMILES strings and phenotypic profiles to condition molecule generation on desired cellular responses. Their model integrates transcriptomic data into the latent space, enabling the generation of biologically relevant compounds tailored to specific gene expression signatures. While effective in aligning chemical and phenotypic spaces, the model does not explicitly incorporate pharmacophoric constraints or structural priors, which are critical for improving binding affinity and downstream activity.
Hu et al.29 introduce ACARL, a reinforcement learning framework designed to account for activity cliffs—regions in chemical space where small molecular changes lead to significant differences in biological activity. By incorporating an Activity Cliff Index and contrastive RL objectives, the model prioritizes regions with strong structure–activity relationships. Although their approach enhances SAR awareness in molecule generation, it is limited to post hoc learning without generative structure in latent space.
Table 1 presents the related works discussed earlier. This table includes the molecular representations, dataset sizes, as well as the contributions and research gaps identified in various research papers.
A prevalent challenge encountered in most of the VAE based approaches is the occurrence of the posterior collapse problem, wherein the model fails to effectively utilize samples from the latent space. In addressing this issue, the proposed PCF-VAE method demonstrates a reduction in the impact of posterior collapse, resulting in the generation of a greater variety of valid molecules on benchmark datasets. Furthermore, PCF-VAE proves successful in generating a more informative latent space, contributing to the generation of desirable molecules. The effectiveness of PCF-VAE in mitigating the posterior collapse problem and generating diverse and valid molecules showcases its potential as an improvement over existing VAE-based approaches.
Method
Molecule representation
The significance of data representation becomes critical in the domain of data-assisted methodologies, such as DL. A variety of representations, such as SMILES, graphs, fingerprints, and three-dimensional models, provide unique approaches to efficiently encapsulate the structural substance of molecules. Because of its string-like properties, SMILES has found widespread use in Data-assisted DL applications. SMILES was proposed by Weininger in 198830, long before the advent of modern DL methods, and it was not designed with generative DL methods in consideration. However, its widespread application in generative DL increases the importance of meticulous preprocessing to fit the proposed methodological framework.
SMILES represents molecular graphs as linear strings, utilizing a set of symbols designed to concisely capture molecular structure while maintaining human readability. For instance, c denotes aromatic carbon, C indicates aliphatic carbon, O represents oxygen, and N stands for nitrogen. Bond types are similarly encoded with distinct symbols: single (-), double (=), and triple (#). These encodings allow SMILES to represent complex molecular structures, including branches and rings, in a compact form.
However, SMILES relies heavily on paired symbols—such as parentheses for branches and numeric labels for ring closures—which introduces syntactic fragility. Experimental studies have identified these pair representations as a primary contributor to invalid molecule generation. In particular, as the depth and length of branches increase, correctly placing opening and closing parentheses becomes increasingly error-prone. Similarly, maintaining consistent numeric ring indices in large or nested ring systems presents challenges.
To address these issues, GenSMILES5 proposes an alternative representation that replaces paired symbols with single-token, position-aware representations. For instance, the SMILES string CC(CCCC)CC, which contains a branch of length four, is converted into the GenSMILES form CCCCCC)4CC, where the closing bracket ) is followed by a digit indicating the branch length.
Ring representations are also simplified. For example, the SMILES string c1ccccc1, denoting benzene, is transformed into CCCCCC^6_1 in GenSMILES, where ^ signifies a ring, the digit 6 represents the number of atoms in the ring, and _1 encodes the number of bonds separating the first and last atoms.
GenSMILES is particularly advantageous in more complex scenarios, such as when branches occur within rings or when rings are embedded within branches. For example, the molecule C1C(CC)C1C represents a three-membered ring that contains a single-atom side branch. In GenSMILES, this is encoded as CCCC)2C^3_1, where the branch is denoted using a closing bracket followed by its relative depth, and the ring structure is captured compactly using the ^ symbol with numeric indicators of ring size and closure distance.
On the other hand, a molecule such as C(C1CCC1CC)C exhibits a ring embedded within a larger branched structure. The ring substructure C1CCC1 appears as part of a long side chain. In GenSMILES, this can be expressed as CCCCC^3_1CC)6C, where the ring is encoded linearly and merged into the main chain using single-token indicators. This linearized format eliminates the need for paired ring digits and nested parentheses, thus enhancing both syntactic robustness and model learnability.
These compact representations in GenSMILES reduce dependency on symbol pairing and nested structural context, leading to higher validity in generative tasks and more interpretable molecular encodings.
Following training, the generative model produces molecules in GenSMILES format. These can be smoothly translated back into valid SMILES strings using a set of deterministic derivation rules that preserve valency and structural integrity, ensuring chemically valid outputs while improving syntactic robustness during generation.
The SMILES notation of four FDA approved drugs with their molecule structures, SMILE representation and GenSMILES representation are shown in the Fig. 1. The structure of molecules are drawn with RDkit python package. The branches in the structure are represented by parentheses in SMILES whereas rings in the structure are represented with the pair of digits in SMILES notation. The elements of branch lies within the rings are not included in the length of the rings. All the cases such as branch within the ring, ring within the branch, branch within the branch and ring within the ring are shown in the diagram.
Drug molecules with their SMILES and GenSMILES representation.
As depicted in Fig. 2, the procedure of converting the SMILES representation to PCF-VAE input is illustrated. The conversion takes place specifically on the SMILES string corresponding to the chemical galidesivir, and the resultant GenSMILES is illustrated. Using the RDkit package, the initial SMILES representation of galidesivir, Nc1ncnc2c(C3NC(CO)C(O)C3O)c[nH]c12], is transformed into its kekulized form31. NC1N=CN=C2C(C3N-C(CO)C(O)C3O)=C[NH]C=12 is a kekulized form that maintains the structural characteristics of galidesivir. As a complex pharmaceutical compound, galidesivir consists of three branches and three rings, which are respectively denoted by parentheses and numerals. The respective counts of ring elements in the first, second, and third rings are five, five, and nine. 2, 1, and 1, respectively, are the bond values between the initial and final elements of the first, second, and third rings. Significantly, components contained within nested branches are omitted from the parent branch or ring, leaving the parent branch with a total of six elements. The young branches each consist of one and two elements. This structural information is presented simply in GenSMILES. Galidesivir is denoted by the GenSMILES notation NCN=CN=CCCNCCO)2CO)1C^5_1O)6=C[NH]C^9_2^-5_1. The numerical value appended to the ending bracket denotes the number of elements present in each branch. GenSMILES employs a tool called a tokenizer to count how many distinct characters there are in a string. In addition, an index is created for each character, and the resultant string is transformed into embedding matrices. The embedding matrices are subsequently utilized as input for the PCF-VAE model, thereby enhancing the model’s ability to conduct a thorough analysis and gain insight into the molecular structure of galidesivir.
Conversion of GenSMILES to embedding vector that act as input to PCF-VAE.
Dataset
The PCF-VAE has undergone training using the MOSES benchmark dataset32. The MOSES platform incorporates various renowned models and also offers evaluation metrics to assess the diversity and quality of the generated molecules. The molecules present in the MOSES dataset are derived from the ZINC dataset33. These molecules possess a molecular weight ranging from 250 to 350 Daltons. They have undergone refinement with specific constraints, ensuring that the number of rotatable bonds does not exceed 7 and the XLogP value is less than or equal to 3.5. The application of medicinal chemistry filters (MCFs) aids in the filtration of the molecules.
The MOSES dataset comprises a total of 1,936,962 molecular structures, which are subsequently divided into training, testing, and scaffold datasets, containing 1.6 million, 176,000, and 176,000 molecules respectively.
PCF-VAE architecture
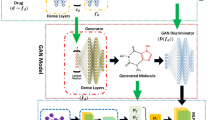
The PCF-VAE is constructed by using basic architecture of VAE. PCF-VAE contain two sub neural network: encoder and decoder. The encoder and decoder of PCF-VAE are constructed with GRU cells. The PCF-VAE encoder k convert the discrete GenSMILES representation of molecules into a fixed-size continuous vector and decoder convert samples form the fixed-size vector back to the real string. The objective of PCF-VAE is to learn the compressed bottleneck called latent space that hold relevant information in the data. The complete architecture of PCF-VAE is shown in the Fig. 3. A conditional properties vector is also combined with the continuous input vector to add more information in the latent space.
The architecture of PCF-VAE model.
Suppose the input of PCF-VAE is s and Z be the latent dimension. Assume that the learning parameters for encoder is denoted by \(\theta\) and c be the condition vector. The objective of PCF-VAE is to find PCF-VAE the encoder distribution \(P_\theta (Z/s, c)\). Samples z \(\sim\) \(P_\theta (Z/s )\) are used by VAE decoder \(P_\phi (s/Z, c)\) to generate new samples \(\hat{s}\). Consider \(\phi\) as decoder’s learn-able parameters. Since the real \(P_\theta (Z/s)\) is intractable, it is necessary to approximate it using a Gaussian distribution \(Q_\theta (Z/s, c)\). The Loss function (ELBO) used in PCF-VAE is
The first term in the loss function, \(- \mathbb {E}_{z}[\log P_\phi (s \mid z, c)]\), represents the reconstruction error. This term ensures that the model accurately reconstructs the input molecules from the latent representation. The second term, \(D_{\text {KL}}[Q_\theta (z \mid s, c) \parallel P_\phi (z \mid c)]\), denotes the Kullback–Leibler (KL) divergence, which regularizes the encoder by encouraging the approximate posterior distribution to align closely with the prior Gaussian distribution, typically defined as \(\mathcal {N}(0, I)\) with \(\mu = 0\) and \(\sigma = 1\).
During training, it is observed that the PCF-VAE tends to generate molecules that closely resemble those in the training set. This behavior is primarily attributed to the phenomenon of posterior collapse, where the decoder becomes overly dependent on the conditioning variable c, thereby ignoring the latent variable z.
Posterior collapse
The reconstruction loss plays a pivotal role in assessing the degree of similarity between the input data and the output data generated by the decoder in the context of a VAE. It measures how effectively the decoder can reconstruct the original input, acting as a driving force for the model to produce accurate and faithful representations. On the other hand, the kl divergence, short for Kullback–Leibler divergence, serves as a measure of dissimilarity between the prior and posterior distributions of the latent space. It quantifies the extent to which the latent space distribution deviates from the prior distribution, reflecting the model’s ability to capture meaningful variations in the data.
By combining these two components, the loss function employed in VAE training provides a comprehensive guidance mechanism. It encourages the model to generate molecules that not only exhibit a high level of fidelity to the input data but also possess a diverse range of molecular structures. This dual objective of maintaining validity and diversity is crucial in applications such as drug discovery, where the generation of novel and unique molecular candidates is of utmost importance.
However, in the initial stages of training, the RNN-based VAE architecture may encounter a phenomenon known as the posterior collapse effect. This effect refers to the situation where the model fails to effectively utilize the full range of the latent space, resulting in an inadequate exploration of its potential. Consequently, the generated molecules may exhibit a striking resemblance to the molecules present in the training set, lacking the desired diversity and novelty.
The posterior collapse effect poses a challenge to the VAE training process, as it restricts the model’s ability to explore and generate genuinely novel molecules. Instead, it tends to produce a limited number of molecular structures that closely resemble those seen during training. The consequence is a diminished diversity in the generated molecules, which can hinder the model’s capacity to discover new chemical entities with unique properties.
To address the problem of posterior collapse, several strategies have been proposed in the literature, focusing primarily on the use of regularization techniques and the incorporation of auxiliary objectives during training. These methods are designed to promote effective utilization of the latent space and encourage the generation of diverse and novel molecular structures. However, directly applying such techniques to datasets represented in the SMILES format remains challenging, owing to the structural and syntactic differences between SMILES strings and natural language sequences.
To mitigate posterior collapse in RNN-based VAE architectures, we propose a novel strategy that introduces a dynamic weighting factor, denoted as \(\alpha\), applied to the KL divergence term in the ELBO loss. At the initial stages of training, \(\alpha\) is set to a value close to zero, thereby minimizing the influence of the KL term and allowing the model to prioritize reconstruction accuracy. As training progresses, \(\alpha\) is gradually increased, allowing the KL divergence to progressively regularize the latent distribution towards the standard Gaussian prior. This annealing process helps balance reconstruction fidelity and latent space regularization, effectively reducing the risk of posterior collapse.
The modified ELBO formulation employed in the PCF-VAE model is expressed as:
Training of model
The training and validation of the PCf-VAE are evaluated through the analysis of multiple metrics depicted in Fig. 4, across 150 epochs on the MOSES benchmark dataset. The KL loss initially rises, reaching its maximum near the 20th epoch, subsequently declining consistently. This behavior demonstrates the model’s progressive alignment of the approximate posterior with the prior distributions, signifying enhanced latent space representation. Likewise, the mutual Information varies across the epochs, exhibiting no distinct upward or downward trajectory. This variability indicates that the information encoded by the latent representation regarding the input data fluctuates dynamically throughout the training process.
In PCF-VAE validation, we meticulously monitor (a) Kullback–Leibler loss, (b) mutual information, (c) reconstruction loss, (d) reconstruction accuracy, and (e) test validation.
The Reconstruction Loss exhibits a sharp drop in the early epochs, subsequently stabilizing as training progresses. This signifies that the model progressively enhances its ability to reconstruct the input data, with errors diminishing markedly over time. The reconstruction accuracy steadily increases from an initial value of approximately 0.55 to nearly 0.80 by the end of training. This consistent enhancement underscores the model’s increasing capacity to preserve critical attributes of the input data in its acquired representations.
In the end, the test Validity curve indicates the model’s generalization performance on unique data. Commencing from an initial value of approximately 0.92, it demonstrates a consistent increase, nearing 0.98 by the 150th epoch. This upward trend validates the model’s ability to generalize effectively, ensuring strong validity and robustness during the training process. These metrics collectively demonstrate the model’s efficacy in acquiring meaningful representations, reconstructing input data, and generalizing to novel samples.
Molecule generation
The de novo generation of molecules through PCF-VAE involves abstaining from the utilization of explicit chemical rules. Instead, molecules are engendered by employing latent space samples as input to the well-trained PCF-VAE model, wherein conditioned properties play a pivotal role in generating desirable molecules. The efficacy of PCF-VAE in the production of desirable molecules is evident. The commencement symbol ’X’ is introduced as input, alongside target properties and Gaussian samples. The symbol generated in correspondence to ’X’ serves as input for the subsequent symbol generation. This iterative process persists until the generated sequence attains its maximum length.
It has been noted that the generation of molecules by PCF-VAE in the vicinity of the specified target yields highly similar molecular structures. To regulate the diversity within the generated molecules, a controllable diversity parameter is employed. The ensuing equation is utilized to imbue diversity into the generated molecular entities.
The latent representations of PCF-VAE, denoted by \(\mu\) and \(\sigma\), depict the hidden samples. The variable ’g’ represents Gaussian samples characterized by a zero mean and a standard deviation denoted by \(D_i\). The variable \(D_i\) can assume values of 1, 2, and 3.
Result and discussion
PCF-VAE extend the straightforward structure of a conditional VAE. The model exhibits a reduced time complexity attributed to its simplicity, resulting in a less number of learnable parameters. While the conditional VAE has been previously documented in the literature34, its applicability in drug design is constrained by inherent drawbacks such as sub-optimal performance on standard SMILES representations, the posterior collapse problem, and inefficient exploration of latent space. To overcome these limitations, several research studies35,36 propose intricate and expansive architectures. Although these architectures attain a SOTA level of validity and diversity in generated molecules, the intricate nature of the process leads to an extension in the time required for completion.
A method has been developed to mitigate the specific limitations associated with conditional VAEs, which is used in the context of PCF-VAE. By using GenSMILES, the complexity associated with SMILES representation is reduced.
In the relevant literature discussed in Section 2, SMILES preprocessing, posterior collapse, and directed search have received insufficient consideration. Currently, as far as we know, unbounded generative neural networks comprise the majority of implementations. Furthermore, to meet particular goals, optimization methods are applied, including techniques such as post-processing approaches (e.g., transfer learning and reinforcement learning) and Bayesian optimization11.
The PCF-VAE underwent assessment using the MOSES platform, as detailed by32. The MOSES platform offers the following evaluation metrics:
-
Validity: The validation metric measures the total number of valid molecules, which are defined as having realistic molecular structures, out of all the molecules that were generated. The RDkit Python package conducts a thorough evaluation of generated molecules’ conformity to predefined structural criteria in order to facilitate the determination of molecular validity (# in the metrics indicates “Total number of”).
$$validity=\frac{\text {\# valid molecules}}{\text {\# generated molecules }}$$ -
Uniqueness: The evaluation encompasses two distinct uniqueness metrics. Firstly, the Unique@1k metric quantifies the uniqueness within the cohort of 1000 valid molecules. Similarly, the Unique@10k metric extends this assessment to the broader scope of 10,000 generated molecules, measuring the total count of unique molecular structures within this larger set.
$$Uniqueness =\frac{set(\text {\# generated molecules )}}{\text {\# generated molecules }}$$ -
Internal Diversity: The evaluation incorporates the Internal Diversity metric to quantify the diversity within the generated set of molecules. In deep learning models, a prevalent challenge known as mode collapse can arise. Internal Diversity serves as a measure to assess the extent of mode collapse, addressing scenarios where the model produces molecules closely resembling each other, concentrating predominantly within specific regions of chemical space. The metric values, denoted as \(IntDiv_1(G)\) and \(IntDiv_2\), are experimentally determined. These metric values are constrained within the range of 0–1, providing a proportional indication of the level of diversity exhibited by the generated molecules. The equation of the metric is:
$$\begin{aligned} intDiv_p(G) = 1 - \sqrt{\frac{1}{\left| G^2 \right| } \sum _{m_1, m_2 \epsilon G} } T (m_1, m_2)^p \end{aligned}$$(5) -
Filters: Filters refer to predetermined restrictions that are enforced throughout the process of molecule generation. Potentially undesirable fragments could be included in the generated molecules. The function of filters is to remove these undesired fragments.
-
Novelty: A quantitative measure of the percentage of molecules missing from the training set is provided. A reduced degree of novelty signifies that the model is prone to overfitting.
$$Novelty= 1 - \frac{\text {\# generated molecules} \; \cap \text {\# Test set molecules } }{\text {\# Test set molecules }}$$ -
Frechet ChemNet Distance (FCD): The FCD is a metric introduced by37 to facilitate the comparison of the distributions of two sets of molecules: the generated set (\(G_r\)) and the training set (\(P_r\)). Let \(m_r\) and \(C_r\) denote, respectively, the covariance and mean of \(P_r\) as a Gaussian distribution. The Gaussian mean and covariance of \(G_r\) are represented by the symbols m and C, respectively. The computation of the FCD metric is possible by employing the equation provided, as suggested by Préuer et al., where d represents the FCD.
$$\begin{aligned} d^2((m, c), (m_r, c_r)) = \mid \mid m-m_r\mid \mid ^{2}_{2} + Tr(C +C_r - 2(CC_r^{\frac{1}{2}}) \end{aligned}$$(6) -
Similarity to the Nearest Neighbour (SNN): The metric is calculated by averaging the Tanimoto similarity between the fingerprints of molecules generated (\(m_G\)) and their nearest neighbor molecules (\(m_R\)) in the reference set (R). This average similarity is denoted as T(\(m_G\), \(m_R\)).
$$\begin{aligned} T(m_G, m_R) = \frac{1}{\left| G \right| } \sum _{m_G\epsilon G} max_{m_R \epsilon R } T (m_G, m_R) \end{aligned}$$(7)The RDKit package is employed to calculate the similarity to the nearest neighbor. A low metric value indicates a considerable distance between the generated set and the reference molecules, while a high metric value suggests proximity. The metric range is confined within 1 and 0.
-
Fragment Similarity (Frag): This metrics are employed to evaluate and compare the distribution of BRICS fragments within the generated set (G) in contrast to the reference set (R). It serve as analytical tools to assess the resemblance or dissimilarity between the generated and reference distributions of fragments, providing valuable insights into the structural composition of the molecular sets under consideration. The metric is defined as :
$$\begin{aligned} Frag (G, R) = \frac{\sum _{f\epsilon F} [c_f (G). c_f(R)]}{\sqrt{\sum _{f \epsilon F} c^2f(G)} \sqrt{\sum _{f \epsilon F} c^2 f(R)}} \end{aligned}$$(8) -
Scaffold Sililarity (Scaf) : This metric exhibits resemblances to fragment similarity—as described by Bemis and Murcko38, the comparison is performed utilizing Bemis-Murcko scaffolds. By concentrating on the shared Bemis-Murcko scaffolds, the Scaf metric evaluates the structural similarity between the generated set and the reference set. This method of analysis contributes to an exhaustive comprehension of the structural similarities and distinctions between the two sets under consideration by facilitating a nuanced evaluation of the molecular scaffolds. The scaffold similarity metric is defined as
$$\begin{aligned} Scaf (G, R) = \frac{\sum _{s\epsilon S} [c_s (G). c_s(R)]}{\sqrt{\sum _{s \epsilon S} C_s^{2}(G)} \sqrt{\sum _{s \epsilon S} C_s^{2}} (R)} \end{aligned}$$(9)
Additionally, MOSES provides SOTA methods for comparing proposed models. The following description outlines the models under consideration:
-
CharRNN17 : The model works by calculating the likelihood of the following SMILE symbol based on the probability of the previous symbol in the sequence. The guiding principle of the model’s operation is to maximize the log-likelihood, which signifies that the observed sequence of SMILE symbols should have the highest possible probability. By following this methodology, the model is consistently trained to forecast the subsequent symbol that is most likely to occur at each iteration, which is consistent with the primary goal of increasing the probability of the generated sequences as a whole.
-
VAE11 : There are two separate neural networks in the model, called an encoder and a decoder. SMILES representations are converted by the encoder network to a hidden space, and samples from the hidden space are converted back into SMILES format by the decoder. The model’s loss function is comprised of two distinct components: KL divergence and reconstruction loss.
-
Adversarial Autoencoder (AAE)13 : The KL divergence term in VAE loss function is replaced with adversarial objectivein AAE. The auxillary network called discriminator is trained distinguished samples from hidden space and the prior distribution. The AAE training depends upon discriminator and encoder-decoder network. AAE is also trained on SMILES representation of molecules.
-
JTN-VAE39 : The JT-VAE model operates in two separate phases. During the initial phase, the JT-VAE constructs a tree structure which symbolizes the framework of subgraph elements. The valid subgraphs obtained from the training set are comprised of these components. These subgraphs are subsequently compiled into molecular structures (graphs).
-
LatentGAN40: An innovative model is presented, referred to as LatentGAN, which amalgamates components from the Generative Adversarial Network (GAN) and Variational Autoencoder (VAE) architectures. The conversion of SMILES representations to latent spaces is the responsibility of the LatentGAN VAE. Following this, hidden samples are generated by the GAN component for utilization by the decoder of the LatentGAN VAE.
-
Nguyen et al.41 propose the TGVAE, a novel model that integrates transformer architecture with graph neural networks and variational autoencoders to enhance molecular generation. TGVAE operates on molecular graph representations, capturing complex structural features.
Other non-neural models NGram, HMM and combinatorial generator is also used for comparison.
The PCF-VAE model demonstrates superior performance when compared to SOTA models (discussed earlier). Across Tables 2 and 3, the PCF-VAE consistently outperforms the VAE model. Specifically, in Table 2, the PCF-VAE achieves a validity score of 0.9789 at diversity level 1, surpassing all other models. Furthermore, the PCF-VAE successfully attains 100% uniqueness at both the 1 k and 100 k levels.
Regarding additional metrics, the PCF-VAE showcases favorable results at different level of diversity. It achieves values of 0.8587, 0.8527, 0.990, and 0.9377 for intDiv, intDiv2, filters, and novelty, respectively, surpassing most SOTA methods.
In Table 3, a comparison of models using FCD, SNN, Frag, and Scaf metrics with Test and TestSF molecules is provided. The Frag metric value of the PCF-VAE surpasses that of all other SOTA methods. While the PCF-VAE does not consistently outperform other models across all metrics, it strikes a balance with diversity among the various metrics, providing competitive performance overall.
The Fig. 5 compares the distributions of Molecular Weight, LogP, and TPSA properties between actual (red) and PCF-VAE-generated (blue) molecules at three diversity levels (D = 1, D = 2, D = 3). At D = 1, the generated distributions closely align with the actual data, indicating high fidelity. As diversity increases (D = 2 and D = 3), the generated distributions deviate more, showing increased variation. Molecular Weight and LogP distributions broaden and shift, while TPSA shows increased multimodality and divergence. This trend reflects a trade-off: higher diversity yields more novel molecules but with less resemblance to real molecular properties. The PCF-VAE model effectively adjusts molecular diversity while controlling structural realism.
Property distribution comparison between actual and PCF-VAE-generated molecules across different diversity levels (D = 1, 2, 3).
Conclusion
The efficacy of the PCF-VAE model arises from its use of a reparameterized loss function. By adding reparameterized loss, the model can find the best balance between the reconstruction and KL divergence parts, which improves performance. The PCF-VAE constructs an informative latent space, that outperformed baseline models like HMM, NGram, CharRNN, and LatentGAN in terms of key measures such as validity, novelty, and diversity. Notably, PCF-VAE achieved a validity score of up to 0.9801, Novelty reaching 0.9501, and consistently high values for Fragmentation (Frag) and Scaffold (Scaf) metrics. The enriched and diverse latent representation enhances the model’s generative capabilities, empowering it to generate molecules that are valid, diverse, and distinctive qualities that hold significant importance in the generative process. However, PCF-VAE excels in generating molecules with high diversity and novelty, it shows marginal variations in Scaffold metrics, particularly in the TestSF. Additionally, PCF-VAE also struggle to maintain chemical realism in extreme diversity settings, occasionally generating unstable or synthetically infeasible molecules.
Future prospects indicate substantial opportunities for the advancement and enhancement of the PCF-VAE model. One promising approach entails enhancing the PCF-VAE to facilitate fine-tuning, focusing on the attainment of specific objectives or the optimization of different features of the generated molecules. This method enables a more customized and advanced generation process, yielding molecules with enhanced properties that meet specific requirements.
Data availability
In the proposed methodology, the dataset utilized for training is publicly accessible via the following link: https://github.com/molecularsets/moses.
References
Polishchuk, P. G., Madzhidov, T. I. & Varnek, A. Estimation of the size of drug-like chemical space based on gdb-17 data. J. Comput.-Aided Mol. Design 27(8), 675–679 (2013).
Bhadwal, A.S., & Kumar, K. Nc-vae: Normalised conditional diverse variational autoencoder guided de novo molecule generation. J. Supercomput. 1–22 (2024).
Kumari, M. & Kaul, A. Recent advances in the application of vision transformers to remote sensing image scene classification. Remote Sensing Lett. 14(7), 722–732 (2023).
Kumari, M., & Kaul, A. Efficient classification of remote sensing images using df-dnlstm: a deep feature densenet bidirectional long short term memory model. Int. J. Syst. Assurance Eng. Manag. 1–18 (2024).
Bhadwal, A. S., Kumar, K. & Kumar, N. Gensmiles: An enhanced validity conscious representation for inverse design of molecules. Knowl.-Based Syst. 268, 110429 (2023).
Kaul, A. & Kumari, M. A literature review on remote sensing scene categorization based on convolutional neural networks. Int. J. Remote Sensing 44(8), 2611–2642 (2023).
Bhadwal, A. S., Kumar, K. & Kumar, N. Nrc-vabs: Normalized reparameterized conditional variational autoencoder with applied beam search in latent space for drug molecule design. Expert Syst. Appl. 240, 122396 (2024).
White, D. & Wilson, R. C. Generative models for chemical structures. J. Chem. Inform. Model. 50(7), 1257–1274 (2010).
Bhadwal, A.S., & Kumar, K. Gva: Gated variational autoencoder for de novo molecule generation. In: 2022 IEEE 9th Uttar Pradesh Section International Conference on Electrical, Electronics and Computer Engineering (UPCON), pp. 1–5 (2022). IEEE.
Singh Bhadwal, A., & Kumar, K. Direct de novo molecule generation using probabilistic diverse variational autoencoder. In: Computer Vision and Machine Intelligence: Proceedings of CVMI 2022, pp. 13–22. Springer, (2023).
Gómez-Bombarelli, R. et al. Automatic chemical design using a data-driven continuous representation of molecules. ACS Central Sci. 4(2), 268–276 (2018).
Blaschke, T., Olivecrona, M., Engkvist, O., Bajorath, J. & Chen, H. Application of generative autoencoder in de novo molecular design. Mol. Inform. 37(1–2), 1700123 (2018).
Makhzani, A., Shlens, J., Jaitly, N., Goodfellow, I., & Frey, B. Adversarial autoencoders. arXiv preprint arXiv:1511.05644 (2015).
Kadurin, A., Nikolenko, S., Khrabrov, K., Aliper, A. & Zhavoronkov, A. drugan: An advanced generative adversarial autoencoder model for de novo generation of new molecules with desired molecular properties in silico. Mol. Pharm. 14(9), 3098–3104 (2017).
Bjerrum, E.J., & Threlfall, R. Molecular generation with recurrent neural networks (rnns). arXiv preprint arXiv:1705.04612 (2017).
Yuan, W. et al. Chemical space mimicry for drug discovery. J. Chem. Inform. Model. 57(4), 875–882 (2017).
Segler, M. H., Kogej, T., Tyrchan, C. & Waller, M. P. Generating focused molecule libraries for drug discovery with recurrent neural networks. ACS Central Sci. 4(1), 120–131 (2018).
Gupta, A. et al. Generative recurrent networks for de novo drug design. Mol. Inform. 37(1–2), 1700111 (2018).
Bhadwal, A.S., Kumar, K., & Kumar, N. Gmg-ncdvae: Guided de novo molecule generation using nlp techniques and constrained diverse variational autoencoder. ACM Trans. Asian Low-Resource Lang. Inform. Process. (2023).
Guimaraes, G.L., Sanchez-Lengeling, B., Outeiral, C., Farias, P.L.C., & Aspuru-Guzik, A. Objective-reinforced generative adversarial networks (organ) for sequence generation models. arXiv preprint arXiv:1705.10843 (2017).
Jaques, N., Gu, S., Bahdanau, D., Hernández-Lobato, J.M., Turner, R.E., & Eck, D. Sequence tutor: Conservative fine-tuning of sequence generation models with kl-control. In: International Conference on Machine Learning, pp. 1645–1654 (2017). PMLR
Olivecrona, M., Blaschke, T., Engkvist, O. & Chen, H. Molecular de-novo design through deep reinforcement learning. J. Cheminform. 9(1), 1–14 (2017).
Zhang, M., Jiang, S., Cui, Z., Garnett, R., & Chen, Y. D-vae: A variational autoencoder for directed acyclic graphs. Adv. Neural Inform. Process. Syst. 32 (2019).
Alperstein, Z., Cherkasov, A., & Rolfe, J.T. All smiles variational autoencoder. arXiv preprint arXiv:1905.13343 (2019).
Tempke, R. & Musho, T. Autonomous design of new chemical reactions using a variational autoencoder. Commun. Chem. 5(1), 40 (2022).
Rigoni, D., Navarin, N., & Sperduti, A. Conditional constrained graph variational autoencoders for molecule design. In: 2020 IEEE Symposium Series on Computational Intelligence (SSCI), pp. 729–736 (2020). IEEE
Lim, J., Ryu, S., Kim, J. W. & Kim, W. Y. Molecular generative model based on conditional variational autoencoder for de novo molecular design. J. Cheminform. 10, 1–9 (2018).
Liu, H., Tian, S., & Liu, X. Phenotypic profile-informed generation of drug-like molecules via dual-channel variational autoencoders. arXiv preprint arXiv:2506.02051 (2025).
Hu, X., Liu, G., Zhao, Y. & Zhang, H. Activity cliff-aware reinforcement learning for de novo drug design. J. Cheminform. 17(1), 54 (2025).
Weininger, D. Smiles, a chemical language and information system. 1. Introduction to methodology and encoding rules. J. Chem. Inform. Comput. Sci. 28(1), 31–36 (1988).
Landrum, G. RDKit: Open-source Cheminformatics. https://www.rdkit.org/
Polykovskiy, D. et al. Molecular sets (moses): A benchmarking platform for molecular generation models. Front. Pharmacol. 11, 565644 (2020).
Irwin, J. J., Sterling, T., Mysinger, M. M., Bolstad, E. S. & Coleman, R. G. Zinc: a free tool to discover chemistry for biology. J. Chem. Inform. Model. 52(7), 1757–1768 (2012).
Lim, J., Ryu, S., Kim, J. W. & Kim, W. Y. Molecular generative model based on conditional variational autoencoder for de novo molecular design. J. Cheminform. 10(1), 1–9 (2018).
Liao, Z., Xie, L., Mamitsuka, H. & Zhu, S. Sc2mol: A scaffold-based two-step molecule generator with variational autoencoder and transformer. Bioinformatics 39(1), 814 (2023).
Skalic, M., Jiménez, J., Sabbadin, D. & De Fabritiis, G. Shape-based generative modeling for de novo drug design. J. Chem. Inform. Model. 59(3), 1205–1214 (2019).
Preuer, K., Renz, P., Unterthiner, T., Hochreiter, S. & Klambauer, G. Fréchet chemnet distance: A metric for generative models for molecules in drug discovery. J. Chem. Inform. Model. 58(9), 1736–1741 (2018).
Bemis, G. W. & Murcko, M. A. The properties of known drugs. 1. Molecular frameworks. J. Med. Chem. 39(15), 2887–2893 (1996).
Jin, W., Barzilay, R., & Jaakkola, T. Junction tree variational autoencoder for molecular graph generation. In: International Conference on Machine Learning, pp. 2323–2332 (2018). PMLR
Prykhodko, O. et al. A de novo molecular generation method using latent vector based generative adversarial network. J. Cheminform. 11(1), 1–13 (2019).
Nguyen, T., & Karolak, A. Transformer graph variational autoencoder for generative molecular design. Biophys. J. (2025).
Author information
Authors and Affiliations
Contributions
A.B. and M.K. wrote the main manuscript text and A.K. worked upon model evaluation and supervised it . All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Bhadwal, A.S., Kumari, M. & Kumar, A. PCF-VAE: posterior collapse free variational autoencoder for de novo drug design. Sci Rep 15, 34152 (2025). https://doi.org/10.1038/s41598-025-14285-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-14285-5