Abstract
Due to the scalability issues of transformers and the limitations of CNN’s lack of typical inductive bias, their applications in a wider range of fields are somewhat restricted. Therefore, the hybrid network architecture that combines the advantages of convolution and Transformer is gradually becoming a hot research and application direction. This article proposes an enhanced dual encoder network (EDE-Net) that integrates convolution and pyramid transformers for medical image segmentation. Specifically, we apply convolutional kernels and pyramid transformer structures in parallel in the encoder to extract features, ensuring that the network can capture local details and global semantic information. To efficiently fuse local details information and global features at each downsampling stage, we introduce the phase-based iterative feature fusion module (PIFF). The PIFF module first combines local details and global features and then assigns distinct weight coefficients to each, distinguishing their importance for foreground pixel classification. By effectively balancing the significance of local details and global features, the PIFF module enhances the network’s ability to delineate fine lesion edges. Experimental results on the GlaS and MoNuSeg datasets validate the effectiveness of this approach. On these two publicly available datasets, our EDE-Net significantly outperforms previous CNN-based (such as UNet) and transformer-based (such as Swin-UNet) algorithms.
Similar content being viewed by others
Introduction
Medical image segmentation plays a crucial role in image analysis, focusing on the identification and delineation of specific structures or areas of interest within medical images. By accurately distinguishing lesions from healthy tissue, this technology assists doctors in developing personalized treatment plans and enhances the precision of medical diagnosis and treatment.
In recent years, deep learning algorithms have achieved outstanding results in computer vision applications, demonstrating extraordinary performance1,2,3,4,5. At the same time, with the increasing aging of the global population, the demand for artificial intelligence in the medical field is growing day by day. Studies have shown that medical image segmentation techniques based on convolutional neural networks can significantly enhance lesion detection capabilities6,7,8,9. Currently, due to its ability to automatically extract and analyze effective features from lesion areas, fully convolutional networks (FCNs)10 and encoder-decoder11 architectures have surpassed traditional methods in terms of effectiveness.
Recently, the Transformer-based architecture12,13,14 has become an essential tool in the medical field and is widely applied in various medical imaging segmentation tasks. The self-attention mechanism in Transformer can effectively calculate the correlation between pixels in the entire image, which makes it have excellent global modeling performance. It is worth noting that the entire feature extractor of Swin-Unet15 is entirely composed of Transformers. In addition, this architecture introduces a novel shift window attention mechanism for medical image segmentation tasks by designing Swin Transformer blocks.
Although CNN-based methods significantly enhance both the accuracy and generalization of network models, the lack of global modeling capability in convolutional kernels limits further performance improvements. Conversely, purely Transformer-based architectures lack the inductive biases inherent to convolution, leading to limited local details feature extraction. To address this issue, many researchers have employed hybrid feature extraction modules16 that combine CNNs and Transformers to capture richer semantic information from images. The TransUNet17 network was the first medical segmentation model to integrate CNN and Transformer modules within the encoder. Since the initial feature map contains a substantial amount of interference, directly calculating pixel-wise correlations globally could disrupt the network’s learning trajectory. Thus, TransUNet leverages the local modeling strengths of CNNs to help the network capture useful foreground information at shallow encoding layers. As the network deepens, the foreground pixel content within the feature map increases. Ultimately, TransUNet applies the Transformer module at the encoder’s output to compute global pixel correlations, enabling the model to capture relationships between lesions of varying scales, thereby enhancing its capacity to accurately recognize complex structures.
The model that integrates CNN and Transformer can enhance the network’s capability to detect lesions, but neglecting the importance of local details and global features may reduce the model’s generalization ability15,17. To address this issue, we propose a hybrid network based on the advantages of convolution operation and self attention mechanism. Additionally, we propose the PIFF module, designed to automatically assess the significance of local details and global features in image analysis tasks. Specifically, the PIFF module first merges local details and global features. Then, by assigning appropriate weight coefficients to different types of features. Through this approach, PIFF can effectively distinguish the contribution of local and global information to foreground pixel classification. By utilizing a hybrid approach within the encoder and efficiently integrating long- and short-term dependencies via the Phase-based Iterative Feature Fusion Module (PIFF), we achieve notable advancements. The primary contributions of this work are as follows:
-
An integrated hybrid medical image segmentation framework, EDE-Net, based on a UNet-like encoder-decoder structure, is proposed. Unlike existing methods, we design an enhanced dual encoder based on a pyramidal Transformer and convolution. This encoder can extract local details and global features effectively.
-
To effectively judge the importance of local details information and global features obtained in each down-sampling stage, we also introduce a Phase-based Iterative Feature Fusion Module (PIFF). PIFF assigns corresponding weight coefficients to local details and global features, which can make the network focus on learning local details or global features that are beneficial to foreground pixel recognition.
Related work
The traditional medical image segmentation method18,19,20 realizes the segmentation function by distinguishing the texture characteristics21, color characteristics or shape characteristics22 between the lesion and the normal tissue. These methods often fail to effectively extract features from the lesion area while mitigating the influences from different sources in medical images. For example, the morphological changes of different organs and tissues, the changes in lesion boundaries, and the inconsistency of imaging equipment will all produce great segmentation differences. In addition, threshold based methods are very common in medical image segmentation, among which Otsu method23 and Minimum Cross Entropy (MCE) method24 are two widely used techniques. The Otsu method is an algorithm that automatically determines the global threshold for image binarization. The core idea is to divide the grayscale histogram of an image into two parts, so that the variance between these two parts is maximized. The Otsu method assumes that the foreground and background of the image are globally consistent and sensitive to uneven lighting and noise. The minimum cross entropy method is a threshold selection method based on information theory. The core idea is to choose a threshold that minimizes the cross entropy between the foreground and background after segmentation. Although the minimum cross entropy method can better measure the separation between foreground and background, it has a high computational complexity and is also sensitive to noise.
Models based on CNNs
Conventional methods for medical image segmentation rely on the textural and geometric characteristics of images. These approaches often struggle to accurately localize the boundaries of lesion tissues, leading to a high incidence of mis-segmentation. With advancements in artificial intelligence, CNN-based segmentation techniques have become increasingly dominant in this field. Among these, FCN10 was the first model to leverage CNNs for image segmentation. While FCN demonstrates superior performance compared to traditional methods, its pooling operations result in a loss of crucial texture and edge information necessary for precise segmentation. Furthermore, UNet25 was the pioneering medical image segmentation model utilizing CNNs. It consists of an encoder, a decoder, and skip connections, where skip connections act as bridges to effectively supplement the encoder’s features to the decoder’s features. UNet++26 employs nested skip connections to mitigate the semantic gap inherent in UNet, but it still faces challenges in capturing global contextual features in feature maps. ResUNet++27 enhances the model’s ability to recognize foreground information by incorporating an attention module to filter out redundant noise. The MultiResUNet28 improves semantic information extraction by utilizing residual paths instead of skip connections for multi-scale analysis. M-Net29 enriches the extraction of semantic details by introducing multi-scale input features at various layers, which are then processed through multiple downsampling and upsampling stages. Additionally, noteworthy 3D medical image segmentation approaches, such as 3D UNet30, replace 2D convolutions with 3D convolutions, while V-net31 enhances UNet by leveraging 3D convolutions and proposing a Dice loss method to achieve improved segmentation results.
Models based on transformer
The CNN-based medical image segmentation model demonstrates effectiveness and feasibility. However, these approaches exhibit limitations in global modeling capabilities. To minimize mis-segmentation, it is essential to grasp the distant correlations between background and foreground pixels. Attempts to capture these long-range dependencies by constructing deeper networks or employing larger convolutional kernels often result in parameter explosion. To overcome these challenges, researchers have integrated Transformer into computer vision, achieving competitive performance across various tasks. Unlike convolutional blocks, Transformers leverage a Multi-Head Self-Attention (MHSA) mechanism for comprehensive global modeling. A prominent example of this in computer vision is the Vision Transformer (ViT)14, which effectively assesses the relationships among all pixels in an image. Following the success of ViT on large datasets, Transformer-based medical image segmentation technologies emerged. Chen et al.17 were the first to propose an encoder that combines Transformer and CNN for 2D medical image segmentation. TransClaw UNet32 developed a hybrid block of Transformer and CNN within the encoder for multi-organ segmentation tasks. The Multi-Compound Transformer33 achieved state-of-the-art results across six different benchmarks by integrating hierarchical scale semantic information into a cohesive framework. To enhance performance, Zhou et al.34 employed CNN and Transformer in parallel, allowing for the extraction of local details and global contextual information. To address overfitting and improve applicability to smaller medical image datasets, MedT35 enhanced the self-attention mechanism using gated axial attention modules. Furthermore, UCTransNet36 replaced traditional skip connections with channel Transformer modules to bridge semantic gaps, while Karimi et al.37 refined the MHSA mechanism for application between adjacent image patches.
The fixed size of the convolution kernel restricts its ability to extract features effectively, resulting in a generally weak global modeling capability for CNN-based medical image segmentation models. While Transformer architectures excel at capturing long-range dependencies, they often struggle to concentrate on specific regions during the feature extraction process. This lack of focused attention can hinder the Transformer model’s ability to precisely localize lesion boundaries, ultimately affecting the accuracy of segmentation results. In addition, the Segment Anything Model (SAM)38 achieves flexible and efficient segmentation performance due to its unique and inspiring segmentation mechanism, demonstrating enormous potential for application. However, SAM has a high dependence on training data, and current medical imaging datasets are usually small in scale, which to some extent limits the application of SAM in the field of medical image segmentation and requires further improvement. To address these challenges, we propose the EDE-Net network. EDE-Net employs an enhanced dual encoder to simultaneously extract local details information and global semantic features from images, and it assesses the significance of these local and global features through a PIFF Module.
Method
First, we present the overall structure of EDE-Net. Subsequently, we will provide a detailed explanation of the workflow of the Phase-based Iterative Feature Fusion Module (PIFF), the Channel-wise Cross Fusion Transformer Module (CCT), and the Channel-wise Cross Attention Module (CCA).
Overall network architecture.
Network architecture
Medical images are prone to uncontrolled elements in being acquired and often contain significant noise. Recent studies39,40 have shown that the ViT41,42 demonstrates superior global modeling abilities. Motivated by this finding, we construct the dual encoder utilizing the Pyramid Vision Transformer (PVT) architecture41, which computes structural representations through spatially reduced attention operations to decrease resource consumption. In particular, we adopt PVTv242, an enhanced iteration of PVT41, known for its improved feature extraction capabilities.
EDE-Net (Pseudo code for network processes).
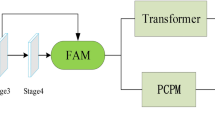
As illustrated in Fig. 1, our proposed EDE-Net primarily consists of an encoder, a CCT module, and a decoder. The encoder is designed as an enhanced dual encoder structure based on CNN integrated with a Pyramid Transformer, which serves to capture both short-term and long-term correlation dependencies within the image. We then fuse the feature information derived from various CNN and Transformer layers using a stage-based fusion module, with the combined output serving as the input feature map for the CCT. This approach effectively enhances the richness of the extracted feature information without introducing additional feature modules. To develop a robust model for medical image segmentation, we employ a combination of cross-entropy loss and Dice loss during the training process, continuously optimizing the model to achieve optimal performance.
Given the input samples \(X, P\in R^{H\times W\times 3}\), we input the features into two branches of the encoder, respectively. The left branch is the CNN layer, which obtains rich local details feature information from the input image through the computation of the convolution module, which can be denoted as \(X_i\in R^{\frac{H}{2^{(i+1)}} \times \frac{W}{2^{(i+1)}} \times C_i }\), \(C_i\in \{64,128,256,512,512\}\), \(i\in \{0,1,2,3,4\}\). The right branch is the pyramid Transformer layer, which acquires the global information from the input, which can be expressed as \(P_j\in R^{\frac{H}{2^{(j+1)}} \times \frac{W}{2^{(j+1)}} \times C_j }\), \(C_j\in \{128,256,512,512\}\), \(j\in \{1,2,3,4\}\). Then, we fuse \(X_1, X_2, X_3, X_4\) with the corresponding strata of \(P_1, P_2, P_3, P_4\) through the PIFF module to generate encoded outputs that are rich in both global semantic information and local detailed features. To eliminate semantic gaps and fuse multi-scale features, CCT and CCA fuse the features output by PIFF. Finally, the fused features are input into the decoder to obtain the final feature map. The specific process is shown in Algorithm 1.
Phase-based iterative feature fusion module(PIFF)
Detailed flow chart of the PIFF.
Due to the differing image processing approaches of Transformers and CNNs, we introduce the PIFF module to more effectively assess the significance of local detailed features obtained from the CNN encoder and the global semantic information derived from the Transformer, as illustrated in Fig. 2.
Firstly, the feature information maps output from each layer of CNN and Transformer are passed through per-element addition operation to further obtain the key information under different receptive fields.
Then, the feature map is input into different branches for feature extraction. For the upper branch, convolution operation and function mapping are applied separately to the feature map to output feature map \(F_3^{up}\).
For the lower branch, the feature map is first subjected to global average pooling(GAP) to preserve important foreground information. Then, convolution operation and function mapping are applied to the feature map respectively, and the output feature map \(F_3^{dn}\) is obtained.
Next, \(F_3^{up}\) and \(F_3^{dn}\) perform feature fusion to obtain a feature map \(G \in R^{C \times H \times W}\) that contains both local and global information.
Subsequently, we use the Sigmoid function to obtain the relevant weight coefficients \(\partial _1\) and \(\partial _2\).
To automatically evaluate the importance of global features and local information, we multiply the weight coefficients \(\partial _1\) and \(\partial _2\) with the feature maps X and P, respectively.
Finally, feature maps \(G_1\) and \(G_2\) are fused element by element and output as feature map T. At this point, the feature map T contains both global and local information that has undergone importance discrimination.
In the two stages of PIFF, we assign coefficients to global semantic information and local detail information, respectively. By automatically measuring the importance of local details and global semantic information, the network can better distinguish the edge information of the focal area.
Channel-wise cross fusion transformer module(CCT)
Due to the fact that different channel feature maps typically focus on different semantic patterns, adaptively fusing channel features is beneficial for the segmentation of complex medical images. In order to better explore the correlation between pixels in feature map channels, we use CCT modules instead of conventional skip connections.
Detailed flow figure of channel-wise cross attention module.
As shown in Fig. 3, the inputs \(F_1 \in R^{64 \times 224 \times 224}\), \(F_2 \in R^{128 \times 112 \times 112}\), \(F_3 \in R^{256 \times 56 \times 56}\), and \(F_4 \in R^{512 \times 28 \times 28}\) of the CCT module come from feature maps of different sizes in the encoder.
Firstly, the input feature maps are processed through different convolution kernels to output feature maps of the same size.
Among them, \({Conv}_{(64,64,16,16)}\) represents that the number of input and output channels of the convolutional kernel is 64, the size of the convolutional kernel is 16 \(\times\) 16, and the stride is 16.
Next, we will perform spatial dimension transformation on \(Q_1\), \(Q_2\), \(Q_3\), and \(Q_4\).
We assemble feature maps \(Q_1\), \(Q_2\), \(Q_3\), and \(Q_4\) in the channel dimension to generate feature map Q.
Next, the feature maps \(Q_1\), \(Q_2\), \(Q_3\), \(Q_4\), and Q are inputted into the Multi head Cross Attention to calculate the correlation between channel pixels.
In the above formula, \(\otimes\) represents matrix multiplication. \(Q_i^T\) represents the conversion of pixel row and column positions in the feature map \(Q_i\).
Finally, the generated channel attention coefficients are multiplied with the pixels in Q.
The input feature map \(Y_i\) is fused with the feature map \(Q_i\) and subjected to dimensional transformation to output \(F_i^{\prime }\).
Finally, the feature map is upsampled to increase its spatial resolution.
Channel-wise cross attention module(CCA)
For the decoder, we do not simply piece together the feature maps, but use the CCA module to explore the correlations between pixels in the spatial dimension.
As shown in Fig. 3, feature map \(F_i^{\prime }\) and feature map \(D_i\) are subjected to pooling operation and full connection layer respectively, and then feature fusion is performed.
Next, we will spatially expand the generated attention coefficients.
Finally, the attention coefficient is multiplied with the feature map \(F_i^{\prime }\) for spatial pixel correlation calculation.
Loss function
Currently, the existing medical image datasets are relatively small in scale, and the target area to be segmented usually only accounts for a very small part of the entire image, which may lead to overfitting of the model during training.To address this challenge, we suggest adopting a composite loss function strategy that combines Dice loss function and cross entropy loss function to effectively alleviate overfitting problems and improve the model’s generalization ability. The main role of the Dice loss function is to solve the negative problem caused by the imbalance between foreground and background information in medical images, And it emphasizes foreground information more during the training process. The Dice loss is related to the Dice coefficient, which is used to evaluate the similarity between the label and the predicted value, the higher the Dice coefficient, the higher the similarity is proved. The Dice coefficient is shown as follows:
M and N represent two sets, respectively, where \({\left| {M \cap N} \right| }\) denotes the number of elements intersecting M and N, and \({\left| M \right| }\) and \({\left| N \right| }\) denote the number of elements in the sets M and N, respectively. The Dice loss is calculated as follows:
where M and N represent ground truth and predicted value, respectively. Meanwhile, using only the Dice loss function creates the problem of loss saturation. In contrast, cross entropy loss as multiclassification loss function, treats each pixel point equally when calculating pixel loss.
Where p(x) and q(x) represent ground truth and predicted values respectively. The ground truths are the labels in the datasets and the predicted values are the outputs predicted by the model. A small cross-entropy value indicates that the labeled and predicted values are similar and the model predicts better. To summarize, our loss function combines Dice loss and cross-entropy loss in the training process. In order to optimize both loss functions equally, these two loss functions are each given weight coefficients \({\mu _1}\) and \({\mu _2}\), both coefficients are fixed values of 0.5:
Experiments and results
Next, the relevant datasets, experimental details, and experimental evaluation metrics will be introduced. The relevant analysis results in comparative experiments and ablation studies will be provided.
Datasets
In this paper, we conduct extensive experiments on two public medical image datasets to validate the effectiveness of our proposed method.
GlaS43
This dataset is a dataset specifically designed for gland segmentation derived from images of colon tissue. It comprises a total of 165 images, with 85 allocated for training and 80 for testing. These images are slices of colorectal adenocarcinoma, processed under various laboratory conditions, with each slide originating from a different patient.
MoNuSeg44
This dataset is designed for nuclear segmentation across multiple organs. Its training set comprises 30 images, which are annotated with a total of 21,623 distinct kernels. The organs represented in these images include the stomach, prostate, liver, colon, bladder, kidney, and breast. The test set contains 14 images featuring organs such as the brain, breast, lung, prostate, bladder, colon, and kidney. Notably, lung and brain tissue images are exclusive to the test dataset, adding a level of difficulty for evaluation.
Implementation details
All experiments are training on an NVIDIA Tesla V100 with 24GB of memory, using Python 3.8 and PyTorch 1.11.0. The input resolution for all datasets is 224 \(\times\) 224, with a batch size of 4, a learning rate of 0.001, momentum of 0.9, and a weight decay rate of 1e-4 using the Adam optimizer. The loss function combines cross-entropy loss with Dice loss. All baseline methods in this paper utilize the same settings and loss functions.
Evaluation metrics
We utilize the Dice coefficient and intersection over union (IoU) ratio to assess model performance. TP is correct positive predictions, in which the predicted and actual results are positive. FP occurs when the prediction is positive, and the actual result is negative, meaning the model incorrectly identifies negative cases as positive. TN refers to correct negative predictions, where both predicted and actual results are negative. FN indicates a negative prediction when the actual result is positive, showing the model misclassifies positive cases as negative. The formulas for calculating the Dice coefficient and IoU are as follows:
HD95 considers the maximum difference between the predicted and the actual results as the measure of similarity. The calculation formula is presented in Formula (7):
Where A and B denote the predicted and the true result, respectively. h(A, B) represents the maximum value of the shortest distance from each point in A to B, i.e, Formula (13):
Where d(a, b) denotes the Euclidean distance between point a to point b.
Comparative experiment
From Table 1, it can be seen that the Dice and IoU of the EDE-Net proposed in this paper reach 90.65% and 83.79%, respectively. Compared to other existing segmentation models, EDE-Net achieved the highest scores in both key metrics. In addition, we present the qualitative comparison results in Fig. 4, and the relevant observations are as follows: (1) The UNet model often suffers from over segmentation or under segmentation during the segmentation process, as shown in the examples on lines 1, 2, and 4. (2) Introducing Transformer into UCTransNet helps alleviate these issues, demonstrating the powerful global context modeling capability of Transformer based hybrid models. However, due to its limited local modeling capability, the segmentation contours predicted by Swin-UNet appear relatively rough. (3) EDE-Net significantly outperforms other models in segmentation performance. Its unique enhanced dual encoder structure endows the model with stronger information capture capabilities, enabling precise identification of the edges of diseased tissues while obtaining richer details and semantic information. From Fig. 4, it can be seen that the image segmentation contour predicted by EDE Net is significantly smoother, for example, it stands out in the comparison between the first and fourth rows.
Segmentation maps visualized for comparison experiments on the GlaS.
Visualisation of segmentation maps for comparison experiments on the MoNuSeg.
Partial enlargement of the visualization results on the MoNuSeg.
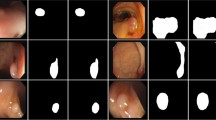
As shown in Table 2, we provide segmentation results for the comparison networks. The Dice and IoU scores of the proposed EDE-Net reach 79.73% and 66.42%, respectively. Figure 5 displays the predicted results from the comparison networks on the MoNuSeg dataset. To facilitate a closer examination of the actual segmentations produced by different networks, we present a partial enlargement of Fig. 5 in Fig. 6. From the first row of Figs. 5 and 6, it’s evident that UNet still misclassifies some background pixels as foreground pixels. In the second and third rows, the Transformer hybrid model shows improvements in some incorrect predictions. However, the Transformer’s weaker ability to differentiate detailed information can hinder kernel segmentation. In contrast, EDE-Net demonstrates a superior ability to capture cell nuclei compared to other models.
In EDE-Net, the parallel use of convolutional kernels and Transformers for feature extraction ensures that the network can simultaneously capture local detail information and global semantic information, thereby significantly improving the recognition accuracy of the network for lesion areas. In addition, the PIFF module effectively balances the importance of local detail features and global semantic features by assigning different weight coefficients, thereby enhancing the network’s ability to depict fine lesion edges.
Ablation study
To illustrate the effectiveness of each module, we designed relevant ablation experiments focusing on the dual encoder structure and the PIFF module, as presented in Tables 3 and 4. It’s important to note that in these tables, “EDE-Net (without PIFF)” indicates that the PIFF module was omitted from the model, and a simple element-wise addition was performed to fuse the features of the dual encoder. “EDE-Net (PIFF-one)” signifies that the PIFF module utilized only one stage for feature extraction. “EDE-Net (only CNN)” means that both branch encoders of EDE-Net consist solely of CNNs, while “EDE-Net (only Transformer)” indicates that both branch encoders are composed exclusively of Transformers.
Visualisation of ablation experiments on the GlaS Segmentation map.
Showing in Table 3, we provided ablation experiments on various components of EDE-Net. It can be seen that when EDE-Net does not use the PIFF module or only uses the PIFF module to calculate the importance of global features and local details in one stage, it will reduce the model’s ability to distinguish foreground information. In addition, the encoder’s ability to distinguish foreground information by using the mixed double branches of CNN and Transformer is better than that when the double branches are all composed of CNN or Transformer.
As illustrated in Fig. 7, our model demonstrates exceptional performance in segmenting the structures of diseased tissue, producing more complete predictions. This improvement can be attributed to the dual-encoder structure, which effectively enables the network to perform joint local details and global modeling, leading to a richer extraction of key information. For instance, in line 2 of Fig. 7, the dual encoder captures more detailed structural features of the lesion area, with larger segments aligning closely with the ground truth (GT) label map. The PIFF module further enhances this process by employing a specialized fusion technique to prevent information loss. As evidenced in row 1 of Fig. 7, the results achieved through the PIFF module are notably more comprehensive compared to the baseline model (UCTransNet). Consequently, EDE-Net exhibits a stronger learning capability.
As shown in Table 4, the Dice and IoU scores of EDE-Net surpass those of EDE-Net (only Transformer) and EDE-Net (only CNN), indicating that relying solely on the Transformer for feature extraction makes it challenging for the encoder to capture the fine tissue details of the lesion. Conversely, using only CNN for feature extraction limits the encoder’s ability to learn the overall structural information of the lesion. Additionally, not employing PIFF to assess the importance of local details and global features diminishes the network’s capacity to differentiate foreground information effectively.
Visualisation of ablation experiments on the MoNuSeg Segmentation map.
As shown in Fig. 8, the nuclei boundaries of most cells are closely connected, complicating the segmentation task for our model. While the segmentation results from our model show improvement over the baseline (UCTransNet), they are still not entirely complete compared to the labeled images. In the fourth row of Fig. 8, the network utilizing a dual encoder structure achieves a more comprehensive segmentation map relative to the baseline (UCTransNet). This enhancement is due to the dual encoder’s ability to capture both local detail and global features, thereby minimizing the risk of losing crucial information. To accurately identify the boundaries of the nuclei, the network must effectively integrate local details and global features. Thus, the PIFF module aids in acquiring the structure and boundary information of cell nuclei more precisely, facilitating a more thorough joint modeling process within the network. Overall, our network demonstrates a distinct advantage in cell nuclei segmentation.
As shown in Table 5, the algorithm proposed in this paper has a low number of parameters and moderate calculation speed. This article aims to improve the segmentation performance of existing segmentation models without increasing network complexity, which further demonstrates the effectiveness of the proposed approach.
Conclusions
To better leverage the advantages of convolutional and self-attention mechanisms, we propose a model called EDE-Net, based on CNN and pyramid Transformer. In the dual-branch encoder, we use a CNN branch to capture local image details and a Transformer branch to calculate long-range dependencies of input features. This parallel structure enables joint modeling of both local and global features. To better assess the importance of features extracted from each branch, we introduce the PIFF module, which performs importance calculations on local and global features to achieve precise localization of pathological tissues. Without any pre-processing, EDE-Net was extensively tested on two publicly available datasets. Experimental results show that our method demonstrates a stronger learning capability and better segmentation performance than current approaches. However, EDE-Net has certain limitations, particularly regarding segmentation performance for complex cell nucleus boundaries. The challenge lies in the difficulty of localizing nuclei with tightly connected boundaries, which increases the risk of misclassification as background. We aim to address this limitation in future work.
Data availability
Gland Segmentation Dataset (GlaS): The dataset is titled “MICCAI 2015 Gland Segmentation Challenge” and is accessible under the Synapse ID: d7d041a88c9f5ac3535a5f088913e8a2 or doi: https://doi.org/10.1016/j.media.2016.08.008. Gland Segmentation Dataset is available on https://opendatalab.org.cn/OpenDataLab/GlaS. Multi-organ Nucleus Segmentation Dataset (MoNuSeg): The dataset is titled “Multi-organ Nucleus Segmentation Dataset” and is accessible under the collection ID: 7e4897888610f1bf15389fc48866c228 or doi: 10.1109/TMI.2017.2677499. Multi-organ Nucleus Segmentation Dataset is available on https://opendatalab.org.cn/OpenDataLab/MoNuSeg.
References
Rauf, Z. et al. Attention-guided multi-scale deep object detection framework for lymphocyte analysis in IHC histological images. Microscopy 72(1), 27–42 (2023).
Ali, M. L., Rauf, Z., Khan, A. R. & Khan, A. Channel boosting based detection and segmentation for cancer analysis in histopathological images. In 2022 19th International Bhurban Conference on applied sciences and technology (IBCAST), 1–6 (2022).
Khan, S. H. et al. Segmentation of shoulder muscle MRI using a new region and edge based deep auto-encoder. Multimed. Tools Appl. 82(10), 14963–14984 (2023).
Zhang, B. et al. Multi-scale segmentation squeeze-and-excitation UNet with conditional random field for segmenting lung tumor from CT images. Comput. Methods Programs Biomed. 222, 106946 (2022).
Liu, Y. et al. MESTrans: Multi-scale embedding spatial transformer for medical image segmentation. Comput. Methods Programs Biomed. 233, 107493 (2023).
Ghosal, P. et al. Compound attention embedded dual channel encoder–decoder for MS lesion segmentation from brain MRI. Multimed. Tools Appli. 1–33 (2024).
Zhang, L. et al. Generalizing deep learning for medical image segmentation to unseen domains via deep stacked transformation. IEEE Trans. Med. Imaging 39(7), 2531–2540 (2020).
Chen, B., Liu, Y., Zhang, Z., Lu, G. & Kong, A. W. K. Transattunet: Multi-level attention-guided u-net with transformer for medical image segmentation. IEEE Trans. Emerg. Top. Comput. Intell. 8(1), 55–68 (2023).
Agarwal, R., Ghosal, P., Sadhu, A. . K., Murmu, N. & Nandi, D. Multi-scale dual-channel feature embedding decoder for biomedical image segmentation. Comput. Methods Programs Biomed. 257, 108464 (2024).
Long, J., Shelhamer, E. & Darrell, T. Fully convolutional networks for semantic segmentation. In Proceedings of the IEEE conference on computer vision and pattern recognition (CVPR 2015). Boston, MA, USA 3431–3440 (2015).
Oktay, O. et al. Attention u-net: Learning where to look for the pancreas. Preprint at arxiv:1804.03999 (2018).
Sharma, A. L., Sharma, K., Srivastava, U. P., Ghosal, P. February. U-SwinTrans: Automated Skin Lesion Segmentation Using Swin Transformer. In 2025 3rd International Conference on Intelligent Systems, Advanced Computing and Communication (ISACC) 25–30 (IEEE, 2025).
Zhang, Y., Liu, H. & Hu, Q.: Transfuse: Fusing transformers and CNNs for medical image segmentation. In Medical image computing and computer assisted intervention–MICCAI 2021: 24th international conference, Strasbourg, France, September 27–October 1, 2021, proceedings, Part I 24 14–24. (Springer International Publishing, 2021).
Dosovitskiy, A. et al. An image is worth 16x16 words: Transformers for image recognition at scale. Preprint at arXiv:2010.11929 (2020).
Cao, H. et al. Swin-unet: Unet-like pure transformer for medical image segmentation. In Proceedings of the Computer Vision-ECCV 2022 Workshops, Part III, Tel Aviv , Israel, 23–27 Volume 13803, 205–218 (2022).
Muksimova, S. et al. A lightweight attention-driven YOLOv5m model for improved brain tumor detection. Comput. Biol. Med. 188, 109893 (2025).
Chen, J. N. et al. Transunet: Transformers make strong encoders for medical image segmentation. Preprint at arXiv: 2102.04306 (2021).
Mamonov, A. V., Figueiredo, I. N., Figueiredo, P. N. & Tsai, Y. H. R. Automated polyp detection in colon capsule endoscopy. IEEE Trans. Med. Imaging 33(7), 1488–1502 (2014).
Tajbakhsh, N., Gurudu, S. R. & Liang, J. Automated polyp detection in colonoscopy videos using shape and context information. IEEE Trans. Med. Imaging 35(2), 630–644 (2015).
Bernal, J. et al. WM-DOVA maps for accurate polyp highlighting in colonoscopy: Validation vs. saliency maps from physicians. Comput. Med. Imaging Graph. 43, 99–111 (2015).
Fiori, M., Musé, P. & Sapiro, G. A complete system for candidate polyps detection in virtual colonoscopy. Int. J. Pattern Recognit. Artif. Intell. 28(7), 1460014 (2014).
Maghsoudi, O. H. Superpixel based segmentation and classification of polyps in wireless capsule endoscopy. In 2017 IEEE Signal Processing in Medicine and Biology Symposium (SPMB 2017), 1–4.
Jumiawi, W. A. H. & El-Zaart, A. Otsu Thresholding model using heterogeneous mean filters for precise images segmentation. In 2022 international conference of advanced Technology in Electronic and Electrical Engineering (ICATEEE), 1–6. (IEEE, 2022).
Jumiawi, W. A. H. & El-Zaart, A. A boosted minimum cross entropy thresholding for medical images segmentation based on heterogeneous mean filters approaches. J. Imaging 8(2), 43 (2022).
Ronneberger, O., Fischer, P. & Brox, T. U-Net: Convolutional networks for biomedical image segmentation. In Proceedings of the 18th International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI 2015), Part III, Munich, Germany , 5–9, Volume 9351, 234–241 (2015).
Zhou, Z., Rahman Siddiquee, M.M., Tajbakhsh, N. & Liang, J. Unet++: A nested u-net architecture for medical image segmentation. In Proceedings of the 4th International Workshop and 8th International Workshop on Deep Learning in Medical Image Analysis-and-Multimodal Learning for Clinical Decision Support (DLMIA 2018 and ML-CDS 2018), Held in Conjunction with MICCAI 2018, Granada, Spain, 20 September 2018; Volume 11045, 3–11.
Jha, D. et al. Resunet++: An advanced architecture for medical image segmentation. In 2019 IEEE international symposium on multimedia (ISM 2019), 225–2255.
Ibtehaz, N. & Rahman, M. . S. MultiResUNet: Rethinking the U-Net architecture for multimodal biomedical image segmentation. Neural Netw. 121, 74–87 (2020).
Fu, H. et al. Joint optic disc and cup segmentation based on multi-label deep network and polar transformation. IEEE Trans. Med. Imaging 37(7), 1597–1605 (2018).
Çiçek, Ö., Abdulkadir, A., Lienkamp, S. S., Brox, T. & Ronneberger, O. 3D U-Net: Learning dense volumetric segmentation from sparse annotation. In Medical Image Computing and Computer-Assisted Intervention–MICCAI 2016: 19th International Conference, Athens, Greece, October 17–21, 2016. Proceedings, Part II 19, 424–432 (2016).
Milletari, F., Navab, N. & Ahmadi, S. A. V-net: Fully convolutional neural networks for volumetric medical image segmentation. In Proceedings of the Fourth International Conference on 3D Vision (3DV 2016), Stanford, CA, USA, 25–28, 565–571 (2016).
Yao, C., Hu, M., Li, Q., Zhai, G. & Zhang, X. -P. Transclaw U-Net: Claw U-Net With Transformers for Medical Image Segmentation. In 2022 5th International Conference on Information Communication and Signal Processing (ICICSP), Shenzhen, China, 280–284 (2022).
Ji, Y. et al. Multi-compound transformer for accurate biomedical image segmentation. In 24th International Conference, Strasbourg, France, September 27–October 1, 2021, Proceedings, Part I 24, 326–336 (2021).
Zhou, H. Y. et al. nnformer: Interleaved transformer for volumetric segmentation. Preprint at arxiv:2109.03201 (2021).
Valanarasu, J. M. J., Oza, P., Hacihaliloglu, I. & Patel, V. M. Medical transformer: Gated axial-attention for medical image segmentation. In Medical image computing and computer assisted intervention–MICCAI 2021: 24th international conference, Strasbourg, France, September 27–October 1, 2021, proceedings, part I 24, 36–46.
Wang, H., Chao, P., Wang, J., Zaiane, O. R. & Uctransnet: Rethinking the skip connections in u-net from a channel-wise perspective with transformer. In AAAI Conf. Artif. Intell. 36(3), 2441–2449 (2022).
Karimi, D., Vasylechko, S. D. & Gholipour, A. Convolution-free medical image segmentation using transformers. In Medical image computing and computer assisted intervention–MICCAI 2021: 24th international conference, Strasbourg, France, September 27–October 1, 2021, proceedings, part I 24, 78–88.
Wu, J. et al. Medical SAM adapter: Adapting segment anything model for medical image segmentation. Med. Image Anal. 102, 103547 (2025).
Bhojanapalli, S. et al. Understanding robustness of transformers for image classification. In Proceedings of the IEEE/CVF international conference on computer vision, 10231–10241.
Xie, E. et al. SegFormer: Simple and efficient design for semantic segmentation with transformers. Adv. Neural. Inf. Process. Syst. 34, 12077–12090 (2021).
Wang, W. et al. Pyramid vision transformer: A versatile backbone for dense prediction without convolutions. In Proceedings of the IEEE/CVF international conference on computer vision, 568–578 (2021).
Wang, W. et al. Pvt v2: Improved baselines with pyramid vision transformer. Comput. Vis. Media 8(3), 415–424 (2022).
Sirinukunwattana, K. et al. Gland segmentation in colon histology images: The glas challenge contest. Med. Image Anal. 35, 489–502 (2017).
Kumar, N. et al. A multi-organ nucleus segmentation challenge. IEEE Trans. Med. Imaging 39(5), 1380–1391 (2019).
Ates, G. C., Mohan, P. & Celik, E. Dual cross-attention for medical image segmentation. Eng. Appl. Artif. Intell. 126, 107139 (2023).
Gao, Y., Zhou, M. & Metaxas, D. N. UTNet: A hybrid transformer architecture for medical image segmentation. In Medical Image Computing and Computer Assisted Intervention–MICCAI,. 24th International Conference, Strasbourg, France, September 27–October 1, 2021, Proceedings, Part III 2, 61–71 (2021).
Li, X. et al. Attransunet: An enhanced hybrid transformer architecture for ultrasound and histopathology image segmentation. Comput. Biol. Med. 152, 106365 (2023).
Liu, A. et al. MLAGG-Net: Multi-level aggregation and global guidance network for pancreatic lesion segmentation in histopathological images. Biomed. Signal Process. Control. 86, 105303 (2023).
Fu, Y., Liu, J. & Shi, J. TSCA-Net: Transformer based spatial-channel attention segmentation network for medical images. Comput. Biol. Med. 170, 107938 (2024).
Yin, Y. et al. AMSUnet: A neural network using atrous multi-scale convolution for medical image segmentation. Comput. Biol. Med. 162, 107120 (2023).
Xu, Q., Ma, Z., Na, H. E. & Duan, W. DCSAU-Net: A deeper and more compact split-attention U-Net for medical image segmentation. Comput. Biol. Med. 154, 106626 (2023).
Funding
Open access funding provided by China Mobile Research Institute (CMRI).
Author information
Authors and Affiliations
Contributions
Depeng Wang and Yibo Sun wrote the main manuscript text and Hong Chen and Xiaolei Zhao prepared figures. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, D., Sun, Y., Chen, H. et al. Image segmentation network based on enhanced dual encoder. Sci Rep 15, 35983 (2025). https://doi.org/10.1038/s41598-025-15274-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-15274-4