Abstract
Membrane channels play an important role in auditory processes. Connexins and pannexins are membrane channels that exist in the cochlea. Connexin26 and connexin30 have been previously shown to be differentially expressed in the developing cochlea of primates in a species-specific manner. However, whether other connexins and pannexins are also expressed in the developing primate cochlea has not been investigated. In this study, we sought to examine the expression patterns of connexin43, connexin31, and pannexin1 during cochlear development in a primate animal model, the common marmoset, to determine the differences observed between the marmoset and existing rodent models. Our observations revealed both interspecies differences in the expression patterns of these genes and similarities between rodents and primates. These results indicate that the spatiotemporal patterns of connexin and pannexin expression are complex and differ between rodents and primates. They also highlight the importance of carefully observing differences in developmental changes in connexins and pannexins between humans and rodents.
Similar content being viewed by others
Introduction
Hearing is achieved via finely controlled mechano-electro transduction in the peripheral sensory organ, the cochlea. Sound, air pressure waves, reach the cochlea via the tympanic membrane and ossicular chains. Pressure waves cause the mechanical stimulation of hair cells in the cochlea. In hair cells, this mechanical stimulation is converted into neuroelectric signaling, which reaches the brain via spiral ganglion neurons. The proper maintenance of electrolysis, cytoplasmic substances, small molecules, substrates, and metabolites is essential for mechanical-electrical conversion in hair cells. Finally, neuro-electrical signals reach the brain and are perceived as sound; this fine-tuned sequence of events results in hearing.
Membrane channels play an essential role in the hearing process1,2. They enable the transport of substances between adjacent cells, or between intra- and extra-cellular channels. Gap junctions are the particular type of membrane channels between adjacent cells3,4 that use connexin complexes, known as connexons, to connect cells. In the cochlea, gap junctions participate in hearing by directly exchanging electrolytes, such as potassium ions and calcium ions, as well as cytoplasmic substances, small molecules, substrates, and metabolites between the cochlear cells5,6,7. This substance exchange is essential for mechano-electro conversion in hair cells and required for other activities in the cochlea5,8. Thus far, several gap junction proteins in the cochlea have been reported, including connexin26 (CX26), CX30, CX31, and CX439,10,11. Pathogenic variants of genes encoding these connexins are known to cause hereditary hearing loss, suggesting that these connexins are essential for hearing in humans12,13.
Pannexins constitute another type of membrane channel14,15. Structurally, pannexins and connexins are very similar, consisting of four transmembrane domains, two extra-cellular and one intracellular loop, and intracellular N- and C-terminal tails; however, pannexins may not form intercellular junctions in the same way as connexins and form a single-membrane channel. This single-membrane channel facilitates communication with the extracellular environment and has been shown to allow the exchange of calcium and ATP16. Previous reports demonstrated that pannexins are also expressed in the cochlea17,18,19.
Our previous study demonstrated that the common marmoset, a non-human primate model animal, is a suitable surrogate for the human cochlea20,21. Recent studies have reported interspecies differences in cochlear gene expression patterns between rodents and primates22,23,24,25,26. Moreover, the common marmoset has a gestation period of about 150 days, and the cochlear development timeline in common marmosets is more closely aligned with that of humans than with rodents, as it takes three times longer than in rodents21. This longer cochlear developmental period has been noted as beneficial for observing the developmental expression patterns of several genes21,27.
Also, cochlear development in rodents may differ from that in primates. For example, in a previous study, we identified spatiotemporal changes in the CX26 and CX30 expression patterns in the developing cochlea of the common marmoset28suggesting the existence of primate-specific control of connexin expression patterns and specific functions of these gap junction proteins28. However, other connexins have not been investigated in this primate.
In the present study, we sought to determine the extent of interspecies differences observed between existing rodent models and the common marmoset. To do this, we examined the expression patterns of CX43 and CX31 in the common marmoset during cochlear development. We also investigated the expression patterns of pannexin1 (PANX1).
Results
CX43 expression in the cochlea of the common marmoset
First, we examined the CX43 expression pattern in the common marmoset. In the P0 cochlea of the common marmoset, CX43 expression was observed in basal cells of the stria vascularis and Type I lateral wall fibrocytes (Fig. 1). The CX43 expression domain did not overlap with and was expressed differently from the CX26 expression domain in this primate (Fig. 2).
CX43 expression in the P0 cochlea of the common marmoset. (A–C) CX43 expression in the basal cells of the stria vascularis and Type I lateral wall fibrocytes. In the stria vascularis, CX43 expression was not observed in MLANA-positive intermediate cells, but it was expressed in CLDN11-positive basal cells (arrowheads in C). (D) CX43 expression in the spiral ganglion neurons of P0 cochlea. No CX43 expression was observed in the spiral ganglion neurons. Nuclei were counterstained with Hoechst (blue). OC organ of Corti, StV stria vascularis, MC marginal cells, IC intermediate cells, BC border cells. Scale bar: 50 μm in (A, D), 20 μm in (B, C).
CX43 and CX26 are expressed in the P0 cochlea of the common marmoset. (A–C) Both CX43 and CX26 are expressed in the basal cells of the stria vascularis (arrowheads in B) and Type I lateral wall fibrocytes. CX43 does not colocalize with CX26. Nuclei were counterstained with Hoechst (blue). StV stria vascularis, MC marginal cells, IC intermediate cells, BC border cells. Scale bar: 20 μm in (A), 10 μm in (B), 5 μm in (C).
Next, we sought to identify developmental changes in the CX43 expression in the cochlea (Fig. 3). Until E101, CX43 was not observed to be expressed in the cochlear duct. In the E109 cochlea, CX43 expression was observed in the stria vascularis and spiral ligament fibrocytes. In the E120 cochlea, CX43 expression was evident in the developing marginal cells of the stria vascularis and type I fibrocytes, as observed in the P0 cochlea.
CX43 expression in the developing cochlea of the common marmoset. (A) CX43 expression pattern in the E101 cochlea. At E101, CX43 expression was observed in the developing Scala Vestibuli (arrowheads in A), but no expression was observed in the cochlear duct. (B, C) Expression patterns of CX43 in the E109 cochlea. In the E109 cochlea, low levels of CX43 were observed in the stria vascularis and lateral wall fibrocytes. (D–F) CX43 expression patterns in the E120 cochlea. In the E120 cochlea, CX43 expression was more evident than that in the E109 cochlea. CX43 expression was observed in the basal cells of the stria vascularis and lateral wall fibrocytes. Nuclei were counterstained with Hoechst (blue). CD cochlear duct, SpGN spiral ganglion neurons, StV stria vascularis, MC marginal cells, IC intermediate cells, BC border cells. Scale bar: 100 μm in (A), 50 μm in (B, D), 20 μm in (C, E, F).
CX31 expression in the cochlea of the common marmoset
Next, we investigated the CX31 expression pattern in the common marmoset cochlea. In the P0 cochlea, CX31 expression was observed in the stria vascularis and sensory epithelium of the cochlear duct (Fig. 4). In the stria vascularis, CX31 expression was observed in the basal cells. At this stage, CX31 expression was evident in the inner sulcus cells, inner border cells, Hensen’s cells, and spiral prominence cells. CX31 expression was not observed in lateral wall fibrocytes at this stage.
CX31 expression 1 in the P0 cochlea of the common marmoset. (A) CX31 expression in the stria vascularis and sensory epithelium of the cochlear duct. (B) In the stria vascularis, CX31 expression was observed in the basal cells (arrowheads in B). (C) CX31 was expressed in the inner sulcus cells, inner border cells, Hensen’s cells, and spiral prominence cells. Nuclei were counterstained with Hoechst (blue). StV stria vascularis, OC organ of Corti, MC marginal cells, IC intermediate cells, BC border cells, IHC inner hair cells, OHC outer hair cells, HC Hensen’s cells, ISC inner sulcus cells. Scale bar: 50 μm in (A, C), 20 μm in (B).
Next, we examined developmental changes in CX31 expression in the cochlea (Fig. 5). Until E109, CX31 expression was undetectable in the cochlea. At E120, CX31 was observed in the spiral prominence cells.
CX31 expression in the developing cochlea of the common marmoset. (A) CX31 expression pattern in the E109 cochlea. At this stage, no CX31 expression was observed in the cochlear duct. (B, C) CX31 expression pattern in the E120 cochlea of the common marmoset. Expression was evident in the spiral prominence cells (arrowhead in B). A higher-magnification image of the box is shown in (B). At this stage, low levels of expression were observed in the inner sulcus cells (arrow in C). No expression of CX31 in the stria vascularis was detected at this stage. Nuclei were counterstained with Hoechst (blue). StV stria vascularis, OC organ of Corti, IHC inner hair cells, OHC outer hair cells. Scale bar: 50 μm.
PANX 1 expression in the cochlea of the common marmoset
Finally, we investigated the PANX1 expression pattern in the common marmoset cochlea. In the P0 cochlea, PANX1 expression was observed in Type I, III, and V lateral wall fibrocytes (Fig. 6). PANX1 expression was also observed in the spiral ganglion, in the satellite cells surrounding the type I spiral ganglion neurons.
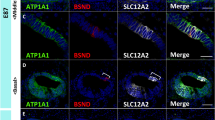
PANX1 expression in the P0 cochlea of the common marmoset. (A–C) PANX1 expression patterns in the P0 cochlea of the common marmoset. PANX1 expression was observed in type I, III, and V spiral ligament fibrocytes (A). No expression was observed in the ATP1A1-positive Type II and IV fibrocytes. PANX1 expression was also observed in the spiral ganglion (B), and in the satellite cells surrounding ATP1A3-positive Type I spiral ganglion neurons (C). Nuclei were counterstained with Hoechst (blue). StV stria vascularis, I type I spiral ligament fibrocytes, II type II spiral ligament fibrocytes, III type III spiral ligament fibrocytes, IV type IV spiral ligament fibrocytes, V type V spiral ligament fibrocytes. Scale bar: 50 μm in (A, B), 20 μm in (C).
Next, we examined developmental changes in PANX1 expression in the cochlea (Fig. 7). PANX 1 expression was evident in spiral ligament fibrocytes as early as in the E101 cochlea. As lateral wall fibrocytes developed, PANX1 expression increased. At E120, PANX1 expression was observed in type I, III, and V lateral wall fibrocytes, as observed in the P0 cochlea. PANX1 expression in the spiral ganglion was detected at E120.
PANX1 expression in the developing cochlea of the common marmoset. (A, B) PANX1 expression pattern in the E101 cochlea. At this stage, PANX1 expression was observed in the superolateral domain of developing lateral wall fibrocytes (arrowhead in A). (C) PANX1 expression pattern in the E109 cochlea in the lateral domain of the developing lateral wall fibrocytes (arrowheads in C). (D, E) PANX1 expression pattern in the E120 cochlea in the developing type I, III and V lateral wall fibrocytes. At this stage, low levels of PANX1 expression were observed in the Schwann cells of the spiral ganglion. Nuclei were counterstained with Hoechst (blue). CD cochlear duct, StV stria vascularis. Scale bar: 100 μm in (A), 50 μm in (B–D), 20 μm in (E).
Validation of the antibodies used in this study
To validate the specificities of the antibodies against CX43 and PANX1, used in this study, we performed immunostaining of the skin of the common marmoset. The antibodies against CX43 and PANX1 used in this study exhibited immunoreactivity in the hair follicles (Supplementary Fig. 1), consistent with the previous reports in humans29. The validation of the specificities of the antibody against CX31 was reported in the previous report20.
Discussion
In this study, we examined the CX43, CX31, and PANX1 expression patterns in the cochlea of the common marmoset. Our investigation revealed inter-species similarities and differences between primate and conventional rodent models. We also identified developmental changes in the expression patterns of these genes.
CX43 is encoded by the GJA1 gene on chromosome 6 in humans. Mutations in the GJA1 gene are known to cause oculodentodigital dysplasia (ODDD), which is accompanied by hearing loss30,31. GJA1 variants also cause non-syndromic autosomal recessive deafness32. In a rodent model, inhibition of CX43 was reported to lead to hearing loss, suggesting that CX43 is essential for normal hearing ability33,34. In addition, recent studies have suggested that CX43 expression changes in relation to age-related hearing loss35 and that CX43 is associated with sensorineural hearing loss induced by oxidative stress36. Several previous reports have investigated the expression patterns of CX43 in the cochlea in both rodents and humans11,37,38,39. Studies on rodents showed that CX43 was expressed in the stria vascularis and lateral wall fibrocytes of mice and rats11,37. However, a previous report showed a different CX43 expression pattern in mice, confirming that mutating Cx43 causes severe hearing loss, in which CX43 expression was reported in the cochlear nerve40. In the human cochlea, CX43 expression was reported in the basal cells of the stria vascularis and spiral ligament fibrocytes38.
CX43 expression in the cochlea of the common marmoset was observed in basal cells of the stria vascularis and type I spiral ligament fibrocytes. Our observations are in line with those of previous studies, suggesting that CX43 expression in these tissues is conserved between rodents and primates and essential for hearing. A previous study compared the expression pattern of CX43 with that of CX26 in a rodent cochlea and showed distinct differences in the localization of these CXs37. We also observed that CX43 expression in the lateral wall did not colocalize with CX26 in this primate. This specific expression pattern of CX43 is well preserved between rodents and primates. By contrast, CX43 expression in spiral ganglion neurons has not been detected in this primate, but has been reported in both rodents and humans11,38. Such inter-species differences would be beneficial to investigate in other primates.
In the developing cochlea of the common marmoset, CX43 expression was obvious after E109, which is a relatively late stage of cochlear development in this animal21,41. Our previous investigation revealed that the formation of basal cells of the stria vascularis and lateral wall fibrocytes occurs in the late phase of cochlear development24,26. In this study, CX43 expression was observed during the maturation of basal cells or lateral wall fibrocytes, as previously observed for CX26 or CX3028.
CX31 is encoded by the GJB3 gene on chromosome 1 in humans. GJB3 pathogenic variants can cause non-syndromic hearing loss or erythrokeratodermia variabilis, a skin disorder42,43,44. Regarding the GJB3 gene, interspecies differences of the hearing loss phenotype have been identified between the rodent model and human patients45which were explained by differences in the CX31 expression patterns in the cochlea between the rodents and primates20. In the adult mouse cochlea, CX31 expression has been reported in the lateral wall fibrocytes10,46. By contrast, in the common marmoset cochlea, CX31 expression was also observed in the basal cells of the stria vascularis and supporting cells, as well as in spiral ligament fibrocytes20. Previous studies have investigated the developmental changes in the expression patterns of CX31 in the rodent cochlea46. A previous study revealed dynamic changes during cochlear development, with CX31 expression being transiently detected in hair and supporting cells in developing mice46. However, no previous study has examined the expression patterns in the developing primate cochlea.
Our observations revealed that CX31 expression was narrower than that in adults at birth; no expression was observed in spiral ligament fibrocytes, which had been reported in the adult cochlea of the common marmoset20. This indicated that the CX31 expression pattern does not become fully confined at birth but dynamically changes even after birth. In addition, distinct changes in the CX31 expression patterns, different than those reported in mice10have been observed. This observation of large inter-species diversity supports the idea that CX31 has a primate-specific role in hearing, as previously suggested20.
PANX1 is encoded by the PANX1 gene on chromosome 11 in humans. Pathogenic PANX1 gene variants have been reported to cause multisystem dysfunction accompanied by sensorineural hearing loss47. However, Panx1 is dispensable for hearing development and auditory function in mice48,49. This phenotypic discrepancy between human patients and rodent models implies the existence of inter-species differences, as reported for CX3120. Therefore, we investigated the PANX1 expression pattern in the primate cochlea.
Panx1 expression in the mouse has been reported broadly in the cochlea, including the sensory epithelium containing the organ of Corti and spiral ganglion neurons48. By contrast, our observation in the marmoset showed strong expression in some spiral ligament fibrocytes (Type I, III, and V) and satellite cells in the spiral ganglion. Previous functional analysis in rodents suggested that Panx1 is dispensable for hearing in rodents. However, since clinical observations indicate that PANX1 is probably indispensable in humans, the distinct primate-specific expression of PANX1 in the cochlea is likely to be important for hearing development. Inter-species differences between mice and humans regarding the cell-specific expression of PANX1 in the central nervous system have been suggested50. Our observation appears to align with this notion. To date, no study has reported the expression of PANX1 in the human cochlea. Investigating the human cochlea, which is difficult due to technical and ethical concerns, would be beneficial for unraveling differences in the PANX1 functional importance in hearing. Therefore, further studies are warranted.
This study examined the changes in the expression of CX43, CX31, and PANX1. Our observations revealed that CX43, CX31, and PANX1 are expressed during the late phase of cochlear development. A previous study examining other CXs showed that CX26 and CX30 expression in lateral wall fibrocytes starts being detected around E11528. Considering this previous report, the observation in this study suggested that the gap junction network in the spiral ligament fibrocytes and stria vascularis start being formed around the late phase of cochlear development. This observation is consistent with a previous study that examined the development of spiral-ligament fibrocytes in this primate24.
Nevertheless, the functional importance of CX43, CX31, and PANX1 in human hearing remains unclear. Previous studies describing the clinical features of patients bearing pathological variants of these genes have suggested that their normal function is essential for hearing32,42,43,47. Our observations revealed inter-species differences in the distribution of expression of these genes in the cochlea, suggesting the usefulness of examining the primate cochlea to investigate the mechanism of hearing loss caused by disruptive variants.
Conclusions
In this study, we investigated developmental changes in the expression of CX43, CX31, and PANX1 in the cochlea of a primate model animal, the common marmoset. Our observations revealed inter-species differences in the expression patterns of these genes, as well as similarities between rodents and primates. These results indicate that the spatiotemporal expression patterns of connexin and pannexin are complex and differ between rodents and primates. They also highlight the importance of carefully observing the differences in developmental changes in connexins and PANX1 between humans and rodents.
Methods
Preparation of Temporal bone samples for antibody staining
Cadaveric temporal bone samples from common marmoset fetuses at E101 (n = 3), E109 (n = 2), E120 (n = 3), and P0 (n = 3) were used in this study, as described in our previous reports21. Skin samples from common marmoset were used in this study. Adult animals were anesthetized via isoflurane inhalation (1.5–4%). Cesarean sections were performed as previously described21. Animal experiments were approved by the Animal Experiment Committee of Keio University (approval numbers: 11006, 08020) and performed following the ARRIVE guidelines and the guidelines of the National Institutes of Health and the Ministry of Education, Culture, Sports, Science, and Technology of Japan.
The embryos were deeply anesthetized on ice and dissected in phosphate-buffered saline (PBS). Temporal bones were dissected immediately after euthanasia and fixed overnight in 4% paraformaldehyde in PBS. The specimens were embedded in Tissue-Tek O.C.T. compound (Sakura Fine Technical Co., Ltd., Tokyo, Japan) for cross-sectioning. P0 specimens were decalcified in Decalcifying Solution B (Wako, Osaka, Japan) for one week and embedded in Tissue-Tek O.C.T. compound for cross-sectioning, as described in our previous reports21. For immunohistochemical analysis, we used 7-µm sections.
Immunohistochemistry
Immunohistochemical analysis was performed as follows: after a brief wash with PBS, the sections were heated (80° C) in 10 µM citrate buffer (pH 6) for 15 min. After another brief wash, the sections were pre-blocked in PBS containing 10% normal serum for 1 h at room temperature, incubated with the relevant primary antibodies overnight at 4 °C, and then incubated with Alexa Fluor-conjugated secondary antibodies for 60 min at room temperature. The nuclei were counterstained with Hoechst 33,258. Confocal imaging was performed using an LSM980 microscope with an AiryScan 2 microscope (Zeiss).
Primary and secondary antibodies used
The following primary antibodies were used: anti-ATP1A1 (Mouse IgG2a, a6F, DSHB, Iowa City, IA, USA, 1:500), anti-ATP1A3 (Mouse IgG1, MA3-915, Invitrogen, Carlsbad, CA, USA, 1:500), anti-TUBB3 (Mouse IgG2a, 801201, Biolegend, 1:500), anti-CALD1 (Mouse IgG2b, 66693-1, Proteintech, Rosemont, IL, USA, 1:500), anti-MLANA (Mouse IgG2a, 60348-1, Proteintech, 1:500), anti-POU4F3 (Mouse IgG1, Santa Cruz Biotechnology, Santa Cruz, CA, USA, 1:200), anti-CLDN11 (Mouse IgA, sc-271231, Santa Cruz Biotechnology, 1:100), anti-CX26 (Mouse IgG1, 13-8100, Invitrogen, 1:200), Anti-CX31 (Rabbit IgG, 12880-1-AP, Proteintech, 1:100), anti-CX43 (Rabbit IgG, ab11370, Abcam, Cambridge, UK, 1:500), anti-PANX1 (Rabbit IgG, ACC-234, Alomone labs, Jerusalem, Israel, 1:500), anti-MYO7A (Mouse IgG1, 138-1-s, DSHB, 1:500), anti-SOX2 (Goat IgG, AF2018, R&D, Minneapolis, MN, USA, 1:200).
The following secondary antibodies were used: goat anti-rabbit IgG, Alexa Fluor Plus 488 (A32731, Invitrogen, Waltham, MA, USA, 1:500), goat anti-rabbit IgG, Alexa Fluor Plus 555 (A32732, Invitrogen, 1:500), goat anti-mouse IgG, Alexa Fluor Plus 555 (A32727, Invitrogen, 1:500), goat anti-mouse IgG1, Alexa Fluor 488 (A21121, Invitrogen, 1:500), goat anti-mouse IgG1, Alexa Fluor 647 (A21240, Invitrogen, 1:500), goat anti-mouse IgG2a, Alexa Fluor 555 (A21137, Invitrogen, 1:500), goat anti-mouse IgG2a, Alexa Fluor 647 (A21241, Invitrogen, 1:500), goat anti-mouse IgG2b, Alexa Fluor 647 (A21242, Invitrogen, 1:500), goat anti-mouse IgG, IgM, IgA, Alexa Fluor 488 (A10667, 1:500, Invitrogen), donkey anti-rabbit IgG, Alexa Fluor Plus 488 (A32814, Invitrogen, 1:500), donkey anti-mouse IgG, Alexa Fluor Plus 555 (A32773, Invitrogen, 1:500), donkey anti-goat IgG, Alexa Fluor Plus 647 (A32849, Invitrogen, 1:500).
Data availability
The datasets used and analyzed during the current study are available from the corresponding author on reasonable request.
References
Lang, F., Vallon, V., Knipper, M. & Wangemann, P. Functional significance of channels and transporters expressed in the inner ear and kidney. Am. J. Physiol. Cell. Physiol. 293, C1187–1208. https://doi.org/10.1152/ajpcell.00024.2007 (2007).
Mittal, R. et al. Indispensable role of ion channels and transporters in the auditory system. J. Cell. Physiol. 232, 743–758. https://doi.org/10.1002/jcp.25631 (2017).
Kumar, N. M. & Gilula, N. B. The gap junction communication channel. Cell 84, 381–388. https://doi.org/10.1016/s0092-8674(00)81282-9 (1996).
Nielsen, M. S. et al. Gap junctions. Compr. Physiol. 2, 1981–2035 (2012).
Jagger, D. J. & Forge, A. Connexins and gap junctions in the inner ear–it’s not just about K + recycling. Cell Tissue Res. 360, 633–644 (2015).
Zhu, Y., Zong, L., Mei, L. & Zhao, H. B. Connexin26 gap junction mediates MiRNA intercellular genetic communication in the cochlea and is required for inner ear development. Sci. Rep. 5, 15647. https://doi.org/10.1038/srep15647 (2015).
Anselmi, F. et al. ATP release through connexin hemichannels and gap junction transfer of second messengers propagate Ca signals across the inner ear. P Natl. Acad. Sci. USA. 105, 18770–18775. https://doi.org/10.1073/pnas.0800793105 (2008).
Zhao, H. B., Kikuchi, T., Ngezahayo, A. & White, T. W. Gap junctions and cochlear homeostasis. J. Membr. Biol. 209, 177–186. https://doi.org/10.1007/s00232-005-0832-x (2006).
Forge, A. et al. Gap junctions in the inner ear: comparison of distribution patterns in different vertebrates and assessement of connexin composition in mammals. J. Comp. Neurol. 467, 207–231 (2003).
Xia, A. P. et al. Expression of connexin 31 in the developing mouse cochlea. Neuroreport 11, 2449–2453. https://doi.org/10.1097/00001756-200008030-00022 (2000).
Cohen-Salmon, M. et al. Expression of the connexin43-and connexin45-encoding genes in the developing and mature mouse inner ear. Cell Tissue Res. 316, 15–22. https://doi.org/10.1007/s00441-004-0861-2 (2004).
Martinez, A. D., Acuna, R., Figueroa, V., Maripillan, J. & Nicholson, B. Gap-junction channels dysfunction in deafness and hearing loss. Antioxid. Redox Signal. 11, 309–322. https://doi.org/10.1089/ars.2008.2138 (2009).
Rabionet, R., Gasparini, P. & Estivill, X. Molecular genetics of hearing impairment due to mutations in gap junction genes encoding beta connexins. Hum. Mutat. 16, 190–202 (2000). doi:Doi 10.1002/1098 – 1004(200009)16:3 < 190::Aid-Humu2 > 3.0.Co;2-I
Penuela, S., Gehi, R. & Laird, D. W. The biochemistry and function of pannexin channels. Biochim. Biophys. Acta. 1828, 15–22. https://doi.org/10.1016/j.bbamem.2012.01.017 (2013).
MacVicar, B. A. & Thompson, R. J. Non-junction functions of pannexin-1 channels. Trends Neurosci. 33, 93–102. https://doi.org/10.1016/j.tins.2009.11.007 (2010).
Chen, J., Zhu, Y., Liang, C., Chen, J. & Zhao, H. B. Pannexin1 channels dominate ATP release in the cochlea ensuring endocochlear potential and auditory receptor potential generation and hearing. Sci. Rep. 5, 10762. https://doi.org/10.1038/srep10762 (2015).
Wang, X. H., Streeter, M., Liu, Y. P. & Zhao, H. B. Identification and characterization of pannexin expression in the mammalian cochlea. J. Comp. Neurol. 512, 336–346. https://doi.org/10.1002/cne.21898 (2009).
Abitbol, J. M. et al. Differential effects of pannexins on noise-induced hearing loss. Biochem. J. 473, 4665–4680. https://doi.org/10.1042/BCJ20160668 (2016).
Tang, W., Ahmad, S., Shestopalov, V. I. & Lin, X. Pannexins are new molecular candidates for assembling gap junctions in the cochlea. Neuroreport 19, 1253–1257. https://doi.org/10.1097/WNR.0b013e32830891f5 (2008).
Hosoya, M., Fujioka, M., Ogawa, K. & Okano, H. Distinct expression patterns of causative genes responsible for hereditary progressive hearing loss in Non-Human primate cochlea. Sci. Rep. 6, 22250. https://doi.org/10.1038/srep22250 (2016).
Hosoya, M., Fujioka, M., Murayama, A. Y., Okano, H. & Ogawa, K. The common marmoset as suitable nonhuman alternative for the analysis of primate cochlear development. FEBS J. 288, 325–353. https://doi.org/10.1111/febs.15341 (2021).
Hosoya, M. et al. A primate model animal revealed the inter-species differences and similarities in the subtype specifications of the spiral ganglion neurons. Sci. Rep. 14, 25166. https://doi.org/10.1038/s41598-024-76892-y (2024).
Hosoya, M. et al. Distribution of macrophages in the developing cochlea of the common marmoset, a primate model animal. Front. Immunol. 14, 1229414. https://doi.org/10.3389/fimmu.2023.1229414 (2023).
Hosoya, M. et al. Development of cochlear spiral ligament fibrocytes of the common marmoset, a nonhuman model animal. Sci. Rep. 13, 11789. https://doi.org/10.1038/s41598-023-39003-x (2023).
Hosoya, M., Fujioka, M., Okano, H. & Ozawa, H. Mapping of Notch signaling in the developing organ of Corti in common marmosets. Front. Neuroanat. 17, 1188886. https://doi.org/10.3389/fnana.2023.1188886 (2023).
Hosoya, M. et al. Development of the stria vascularis in the common marmoset, a primate model. Sci. Rep. 12, 19811. https://doi.org/10.1038/s41598-022-24380-6 (2022).
Hosoya, M. et al. Neuronal development in the cochlea of a nonhuman primate model, the common marmoset. Dev. Neurobiol. 81, 905–938. https://doi.org/10.1002/dneu.22850 (2021).
Hosoya, M. et al. Dynamic Spatiotemporal Expression Changes in Connexins of the Developing Primate’s Cochlea. Genes (Basel). 12 https://doi.org/10.3390/genes12071082 (2021).
Cowan, K. N., Langlois, S., Penuela, S., Cowan, B. J. & Laird, D. W. Pannexin1 and Pannexin3 exhibit distinct localization patterns in human skin appendages and are regulated during keratinocyte differentiation and carcinogenesis. Cell. Commun. Adhes. 19, 45–53. https://doi.org/10.3109/15419061.2012.712575 (2012).
Loddenkemper, T., Grote, K., Evers, S., Oelerich, M. & Stogbauer, F. Neurological manifestations of the oculodentodigital dysplasia syndrome. J. Neurol. 249, 584–595. https://doi.org/10.1007/s004150200068 (2002).
Paznekas, W. A. et al. GJA1 mutations, variants, and connexin 43 dysfunction as it relates to the oculodentodigital dysplasia phenotype. Hum. Mutat. 30, 724–733. https://doi.org/10.1002/humu.20958 (2009).
Liu, X. Z. et al. Mutations in GJA1 (connexin 43) are associated with non-syndromic autosomal recessive deafness. Hum. Mol. Genet. 10, 2945–2951. https://doi.org/10.1093/hmg/10.25.2945 (2001).
Zhang, J. et al. Suppression of connexin 43 leads to strial vascular Hyper-Permeability, decrease in endocochlear potential, and mild hearing loss. Front. Physiol. 11, 974. https://doi.org/10.3389/fphys.2020.00974 (2020).
Wang, J. & Song, Q. Inhibition of connexin 43 induces hearing loss in postnatal mice. Physiol. Int. 108, 43–53. https://doi.org/10.1556/2060.2021.00008 (2021).
Pineros, J., Zhu, X., Ding, B. & Frisina, R. D. Connexins 30 and 43 expression changes in relation to age-related hearing loss. Hear. Res. 444, 108971. https://doi.org/10.1016/j.heares.2024.108971 (2024).
Ding, L. & Wang, J. MiR-106a facilitates the sensorineural hearing loss induced by oxidative stress by targeting connexin-43. Bioengineered 13, 14080–14093. https://doi.org/10.1080/21655979.2022.2071021 (2022).
Suzuki, T., Takamatsu, T. & Oyamada, M. Expression of gap junction protein connexin43 in the adult rat cochlea: comparison with connexin26. J. Histochem. Cytochem. 51, 903–912. https://doi.org/10.1177/002215540305100705 (2003).
Liu, W., Boström, M., Kinnefors, A., Edin, F. & Rask-Andersen, H. Connexin 43 expression in the human cochlea: an immunohistochemistry study. J. Hear. Sci. 1, 21–29 (2011).
Liu, W. et al. Possible role of gap junction intercellular channels and connexin 43 in satellite glial cells (SGCs) for preservation of human spiral ganglion neurons: A comparative study with clinical implications. Cell. Tissue Res. 355, 267–278. https://doi.org/10.1007/s00441-013-1735-2 (2014).
Abitbol, J. M., Kelly, J. J., Barr, K. J., Allman, B. L. & Laird, D. W. Mice harbouring an oculodentodigital dysplasia-linked Cx43 G60S mutation have severe hearing loss. J. Cell Sci. 131, jcs214635. https://doi.org/10.1242/jcs.214635 (2018).
Hosoya, M. et al. Early development of the cochlea of the common marmoset, a non-human primate model. Neural Dev. 17 https://doi.org/10.1186/s13064-022-00162-8 (2022).
Liu, C. et al. Mutations in the gene encoding gap junction protein β-3 associated with autosomal dominant hearing impairment. Nat. Genet. 20, 370–373 (1998).
López-Bigas, N. et al. Connexin 31 (GJB3) is expressed in the peripheral and auditory nerves and causes neuropathy and hearing impairment. Hum. Mol. Genet. 10, 947–952 (2001).
Gottfried, I. et al. A mutation in GJB3 is associated with recessive erythrokeratodermia variabilis (EKV) and leads to defective trafficking of the connexin 31 protein. Hum. Mol. Genet. 11, 1311–1316. https://doi.org/10.1093/hmg/11.11.1311 (2002).
Plum, A. et al. Connexin31-deficiency in mice causes transient placental dysmorphogenesis but does not impair hearing and skin differentiation. Dev. Biol. 231, 334–347 (2001).
Lopez-Bigas, N., Arbones, M. L., Estivill, X. & Simonneau, L. Expression profiles of the connexin genes, Gjb1 and Gjb3, in the developing mouse cochlea. Mech. Dev. 119 (Suppl 1), 111–115. https://doi.org/10.1016/s0925-4773(03)00102-3 (2002).
Shao, Q. et al. A germline variant in the PANX1 gene has reduced channel function and is associated with multisystem dysfunction. J. Biol. Chem. 291, 12432–12443. https://doi.org/10.1074/jbc.M116.717934 (2016).
Zorzi, V. et al. Mouse Panx1 is dispensable for hearing acquisition and auditory function. Front. Mol. Neurosci. 10, 379. https://doi.org/10.3389/fnmol.2017.00379 (2017).
Abitbol, J. M. et al. Double deletion of Panx1 and Panx3 affects skin and bone but not hearing. J. Mol. Med. (Berl). 97, 723–736. https://doi.org/10.1007/s00109-019-01779-9 (2019).
Pietrowski, M. J. et al. Glial purinergic signaling in neurodegeneration. Front. Neurol. 12 https://doi.org/10.3389/fneur.2021.654850 (2021).
Acknowledgements
We thank Saki Ninomiya for providing technical support. We thank Ayako Y. Murayama, Sho Yoshimatsu, Junko Okahara and Hideyuki Okano for providing materials.
Funding
MH was supported by a grant from the Japanese government, MEXT KAKENHI (Grant-in-Aid for Scientific Research (C) 24K12727), the Keio Medical Association, and Keio University Medical Science Fund, Keio Gijuku Academic Development Funds, and Grant for Research Encouragement Award of the Japanese Society of Otorhinolaryngology-Head and Neck Surgery. MS was supported by a grant from Japanese government, MEXT KAKENHI (23K08972). MU was supported by a grant from Japanese government, MEXT KAKENHI (24K12706).
Author information
Authors and Affiliations
Contributions
M.H., S.S., M.U., M.S., T.N., N.O., H.O., conceived and designed the experiments. M.H. wrote the manuscript. M.H. performed most of the experiments. M.H., M.U., and M.S. analyzed the data. All authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
MH was founding scientist of Otolink Inc. Other authors have no conflicts of interest directly relevant to the content of this article.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Hosoya, M., Suda, S., Ueno, M. et al. Distinct Connexin43, Connexin31, and Pannexin1 expression patterns in the cochlea of a non-human primate. Sci Rep 15, 29527 (2025). https://doi.org/10.1038/s41598-025-15685-3
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-15685-3