Abstract
The interaction between protein and DNA across three distinct conformations of protein chains is examined in the framework of band structures and density of states analysis, utilizing a tight-binding Hamiltonian model in conjunction with Green’s function technique. At a constant temperature and with a predetermined number of sub-sites on the DNA strand, the spectral diagrams reveal a flat energy dispersion curve for both the protein molecules and the DNA strand independently, demonstrating characteristics akin to those of semiconductors. An increase in the contact points between protein molecules and the DNA strand results in a transition from semiconducting to metallic properties, a change that is also affected by the hydrogen bonds contributed by the mutant protein at these contact points. The electronic characteristics of the protein-DNA system are modulated by the size of the DNA, leading to a conversion of localized states within the structures into less defined energy levels as the length of the DNA strand increases. The influence of temperature on the density of states causes variations in both the peak heights and their positions. The stretching effects of DNA influenced by the presence of protein molecules, result in modifications by redistributing spectral characteristics within the density of states. The interaction between protein and DNA is anticipated to have a direct impact on the electronic properties of DNA, which differ across various protein conformations, thus paving the way for new research opportunities with considerable biological significance.
Similar content being viewed by others
Introduction
Deoxyribonucleic acid (DNA) is a complex macromolecule1 composed of repetitive sequences of base pairs (bps), specifically adenine (A) paired with thymine (T) and guanine (G) paired with cytosine (C). These bps are interconnected through hydrogen bonds and are organized in a double-helix structure, which is reinforced by a sugar-phosphate backbone. The double-helix structure of the DNA molecule, distinguished by its \(\pi\)-electron cores due to base stacking, facilitates a one-dimensional (1D) pathway for electronic charge transport. The electronic characteristics of DNA are predominantly influenced by \(\pi\)-electrons, with each site possessing a single orbital, thereby rendering it appropriate for modeling as a tight-binding (TB) system characterized by defined nearest neighbor (nn) interactions2,3,4,5,6,7,8. Essential features such as self-assembly and self-recognition are crucial for the role of DNA molecules as carriers of genetic information, and these attributes may be leveraged in the design and construction of electronic circuits9. Additionally, DNA can be integrated with various materials and methodologies to produce an extensive range of biosensor designs10,11. In essence, nucleic acids serve a dual purpose; they not only transmit genetic information but also function as adaptable and potent biomolecules in the creation of electrochemical biosensors capable of detecting and identifying proteins12.
Proteins are intricate biological macromolecules composed of a specific sequence of amino acid residues13. The synthesis of proteins involves the linear linkage of amino acids via peptide bonds, which create a structural backbone for the protein, with diverse side chains branching off. In contrast to other biological macromolecules, proteins exhibit complex and irregular configurations. There are twenty distinct amino acids that serve as the essential building blocks of all proteins. The linear arrangement of these amino acids is known as the primary structure of the protein. Furthermore, under physiological conditions, each protein adopts a unique 3D conformation14,15, which is referred to as its native tertiary structure. This native configuration is crucial, as it dictates the functional characteristics of the protein. The relationship between a protein’s amino acid sequence and its native tertiary structure represents a significant challenge in biological research. While various computational techniques have been developed to forecast protein structures, only a few of these methods provide robust mathematical assurances. Lattice models have been particularly effective in exploring the intricacies of protein structures16. These models aid in deriving fundamental principles, making predictions, and improving our understanding of different protein properties17.
Interactions between proteins and DNA are fundamental, as they play a vital role in numerous cellular processes such as the regulation of gene expression, DNA replication, repair, recombination, and the organization of chromosomal DNA, thereby enabling cells to interpret and utilize the genetic information encoded in DNA for protein synthesis and essential functions18,19,20,21,22,23. A key concern regarding these interactions is the approach proteins utilize to search for specific targets on DNA. The ways in which proteins can detect relatively small cognate sequences, typically consisting of 15 to 20 bps for repressors and 4 to 6 bps for restriction enzymes within the millions of bps of non-specific chromosomal DNA, have largely gone unexplored by researchers for decades24. Studies indicate that protein molecules exhibit an exceptional proficiency in quickly finding their target within the DNA sequence25,26,27. To identify a specific DNA sequence, a protein requires multiple interactions with the bps found in the major groove of the DNA, achieved through the formation of hydrogen bonds28 or van der Waals interactions by the protein’s amino acid side chains with the bases. It has been established that the recognition of consensus sequences on DNA by proteins is contingent upon the entanglement of \(\pi\)-\(\pi\) electrons between DNA nucleotides and protein amino acid24.
DNA exhibits a helical configuration, and proteins have the capacity to modify this twist by either unwinding or overwinding the DNA29,30,31. The natural flexibility of DNA enables it to bend and twist in reaction to protein binding, with these deformations being pivotal in numerous cellular processes. Proteins can further induce twisting and stretching of DNA, thereby influencing its overall structure and functionality. The interactions between proteins and DNA represent a dynamic interplay of forces and conformations. These interactions are profoundly affected by the mechanical properties of DNA, particularly its capacity to twist and stretch. Indeed, proteins attach to DNA and can instigate alterations in its twist and stretch, while the mechanical state of DNA can, in turn, influence its interactions with proteins. Grasping this interplay is vital for comprehending many fundamental biological processes. This relationship between protein binding and DNA mechanics is crucial for gene regulation, DNA replication, repair, and other essential cellular functions. For instance, in the context of twist-stretch profiles of DNA chains, the relationship between twist and stretch for a collection of short DNA molecules, as modeled by a mesoscopic Hamiltonian, has been investigated29. This model also incorporates the bending and twisting fluctuations that occur between neighboring base pairs within the molecular stack. The results indicate that the chains typically exhibit over-twisting when subjected to an external stretching force, and this over-twisting is physically linked to a reduction in the average diameter of the helix, which corresponds to a damping effect on the fluctuations of the bps.
Artificial intelligence (AI) and deep learning (DL) techniques are transforming the investigation of DNA-protein interactions by offering robust tools for predicting binding sites, elucidating interaction mechanisms, and even designing innovative proteins32,33. These techniques utilize extensive datasets of established protein structures, DNA sequences, and interaction data to discern intricate patterns and relationships, resulting in predictions that are more precise and efficient compared to traditional methods. Recent developments in DNA-protein interaction research have been greatly accelerated by the implementation of AI and DL techniques34,35,36. These advancements are enabling not only more accurate forecasts of protein-DNA binding locations34,35, improving the understanding of protein-DNA interactions35, but also accelerating the process of drug discovery34. For example, AI and DL are advancing structure-based drug discovery by refining ligand binding site identification, estimating protein-ligand binding poses, and developing scoring functions34,35. Furthermore, AI-driven methods enable a comprehensive analysis of protein structures, which can be associated with the prediction of drug-target interactions. Additionally, AI and DL are being incorporated into computational models to scrutinize protein structures and forecast drug-target interactions.
It has been demonstrated that certain proteins may lose their capacity to bind to particular DNA sequences when exposed to low temperatures, whereas others may exhibit modified specificity37. At low temperatures, the interactions between proteins and DNA can undergo significant alterations. While some interactions may weaken or become disrupted due to diminished molecular motion, others may be enhanced or exhibit increased specificity. It is important to observe that while it is theoretically possible for protein-DNA interactions to occur at very low temperatures, the significant decrease in molecular motion greatly influences the dynamics necessary for stable and specific binding. However, some interactions may still emerge due to quantum mechanical effects or residual thermal energy. The specific outcomes are contingent upon the individual protein and DNA sequence involved, as well as the characteristics of their interaction38. Consequently, protein-DNA interactions can be profoundly affected by extremely low temperatures, potentially resulting in alterations in binding affinity, stability, and even the capacity of proteins to engage with DNA. In fact, lower temperatures typically enhance the stability of protein-DNA complexes. This phenomenon occurs because the reduction in thermal energy makes it more challenging for the complex to dissociate39. Furthermore, the changes in enthalpy and entropy associated with binding can differ markedly at low temperatures compared to physiological conditions. For example, certain protein-DNA interactions that are entropy-driven at room temperature may transition to being enthalpy-driven at low temperature40. Additionally, low temperatures can encourage the emergence of novel protein-DNA interactions that are not present at elevated temperatures41. Comprehending the interactions between proteins and DNA at low temperatures is crucial for grasping cellular responses. Numerous organisms face stress due to low temperatures, and their survival hinges on the effective operation of protein-DNA interactions42. By elucidating how proteins engage with DNA in cold environments, researchers can develop innovative materials and pharmaceuticals that operate under extreme conditions41. Investigations at low temperatures can yield valuable insights into the basic principles governing protein folding and the dynamics of protein-DNA interactions38.
Proteins, in contrast to other biological macromolecules, exhibit intricate and irregular structures. The geometric properties of proteins are related to the impact of their 3D conformation and the spatial arrangement of amino acids on the architecture, stability, and functionality of the protein. These geometric features include hydrogen bonding, van der Waals forces, and the overall folding configuration. Indeed, the models have a specific amino acid sequence, with differences in the distribution of hydrogen bonds, where thirteen hydrogen bonds are designed to simulate a mutation (disease) observed in a natural protein, as noted by43. The electronic properties of proteins arise from their amino acid composition, notably the presence of aromatic residues and their potential to form \(\pi\)-interactions, which play a crucial role in determining their folding, stability, and molecular interactions. Protein-DNA interaction research can leverage a TB model that portrays DNA as a system with \(\pi\)-electrons and specific nn interactions, allowing for the modeling of proteins as engaging in these interactions. A TB model for DNA-protein interactions employs the principles of TB theory to depict the relationship between a protein and DNA, conceptualizing the DNA molecule as a series of distinct sites for protein binding. The binding strength is influenced by the specific interactions between the amino acids of the protein and the bases of the DNA at each site, facilitating the computation of binding affinities and sequence specificity derived from the interaction energies at these locations24. In the TB framework concerning the interaction between 3D proteins and 1D DNA, proteins first move through 3D space. Upon meeting DNA, they transition to a 1D sliding motion along the DNA to locate their specific target site24. This research seeks to explore the interaction between protein and DNA by examining the band structure (BS) and density of states (DOS) within a system that comprises a double-stranded DNA configuration2,3,4,5 alongside three unique types of protein chains (Fig. 1a) designated as \(\mathrm {P_{1}}\), \(\mathrm {P_{2}}\), and \(\mathrm {P_{3}}\)28,43. In the context of the DNA model, each site corresponds to a nitrogenous base (A, T, C, and G), with electronic pathways illustrated as solid lines, representing the form of a ladder (Fig. 1b). The arrangement of protein molecules is dictated by a specific sequence of amino acids, with differences in the orientation of hydrogen bonds; a total of thirteen hydrogen bonds are utilized to model a mutation associated with a certain disease in a natural protein28,43. This examination is focused on protein chains that comprise thirty-six amino acids. The \(\mathrm {P_{1}}\) conformation shows a simple structural framework where amino acids are connected exclusively by covalent (peptide) bonds. In contrast, the \(\mathrm {P_{2}}\) and \(\mathrm {P_{3}}\) conformations exhibit a structural configuration in which amino acids are linked through both peptide bonds and non-covalent (hydrogen bond) interactions. This study builds on the protein models as presented in43. They utilized computer simulations within the lattice model framework to assess and validate the primary hypotheses of the protein engineering analysis regarding the transition state between the folded and unfolded states. In their model, a protein chain is represented as a self-avoiding walk on a cubic lattice, with covalently bonded residues occupying neighboring lattice sites. Their approach is based on the principle of conducting sequence design and folding simulations. This sequence was developed to achieve a low relative energetic stabilization, which makes the hydrogen bonds vital for their models. Besides, the double-stranded model offers a simplified yet effective representation of DNA’s double helix structure in computational studies, enabling researchers to simulate the behavior of DNA under various conditions. This model is significant for comprehending DNA’s role in biological processes and its potential applications in nanotechnology. It is utilized to investigate DNA’s properties, such as its structure, flexibility, and electronic transport. The model frequently incorporates elements like base pairing interactions, the flexibility of the sugar-phosphate backbone, and the helical twist to accurately simulate DNA’s behavior. Through this computational model, researchers can explore how DNA interacts with proteins, other molecules, or surfaces, which is essential for understanding gene regulation and the development of DNA-based technologies. This study utilized the TB Hamiltonian model alongside Green’s function methodology to examine the interactions between DNA and three conformations of protein models, considering a range of temperatures and varying hopping parameter values within the protein chains, as well as the dimerization effects of parallel hopping in the DNA and the effects of stretching of the DNA strand on the protein-DNA interaction. We have employed existence models for both protein chains and DNA strands to investigate the interaction between proteins and DNA. Due to the complex nature of these systems, we opted for a simplified model to make the problem more tractable. Such simplifications have been incorporated into the models utilized in this research. For example, the quantum transport of protein chains has been examined in6,28, while the electronic characteristics of DNA using a double-stranded model have been studied in3,4,5. Despite the simplifications, these models successfully capture the qualitative behavior of protein chains and DNA strands. Concerning the length of the DNA sequence, it is fundamentally important for the efficiency and stability of protein-DNA interactions. This length affects the search mechanism, binding affinity, and the kinetics of the interaction as a whole, thereby influencing numerous cellular processes that depend on these interactions, including gene regulation and DNA replication. The findings of this study indicate that the interaction between protein and DNA is influenced not only by the number of contact points, temperature, and the conformation of the protein, but also by the total length of the DNA segment under consideration.
Panel (a) presents a schematic illustration of the conformations of \(\textrm{P}_{1}\) (left), \(\textrm{P}_{2}\) (center), and \(\textrm{P}_{3}\) (right), with hydrogen bonds indicated by dashed lines. Panel (b) illustrates the double-stranded DNA, highlighting the connections between four sub-sites of a protein molecule and the DNA strand, along with the representation of the hopping terms.
Hamiltonian model
The TB Hamiltonian framework functions as a theoretical model for elucidating the dynamics of quantum systems. This model enables the examination of electronic characteristics across diverse materials by utilizing a Hamiltonian that integrates various factors. The TB Hamiltonian encompasses the complete system, which includes the DNA strand denoted by \(\hat{\mathcal{H}}_{\textrm{D}}\), the protein chain represented as \(\hat{\mathcal{H}}_{\textrm{P}}\), and their interactions or couplings symbolized by \(\hat{\mathcal{H}}_{\textrm{C}}\). The model is characterized by the following Hamiltonian:
each term, when articulated within the framework of second quantization, can be represented through its local and hopping components as follows:
and
in this context, \(\alpha\) refers to a and b strand indices of the DNA model, i (\(\ell\)) and j (\(\ell ^{\prime }\)) denote specific sub-sites within the DNA strand (protein chain). The notation \(\varepsilon _{i}\) (\(\epsilon _{\ell }\)) signifies the on-site energy associated with an electron positioned at the sub-site or nucleic acid (amino acid) i (\(\ell\)). The parameter \(t_{\textrm{D}ij}\) represents the hopping amplitudes for a \(\pi\)-electron moving from sub-site i to its nn j. It is important to observe that the parallel and perpendicular notations on \(t_{\textrm{D}ij}\) indicate hopping between nn sites within the same strand and between two distinct strands, respectively. It is worth mentioning that the perpendicular notation on \(t^{\perp }_{\textrm{D}i_{a}i_{b}}\) signifies hopping between two separate strands. Consequently, the electron may hop, for instance, from the i site on strand a to the corresponding site i on strand b. Thus, the perpendicular hopping pertains solely to the two bp mates on the complementary DNA strands, meaning \(i=j\). Also, the parameter \(t_{\textrm{P}\ell \ell ^{\prime }}\) (along with its counterpart \(t^{\prime }_{\textrm{P}\ell \ell ^{\prime }}\)) represents the nn hopping amplitude for a \(\pi\)-electron moving from sub-site \(\ell\) to sub-site \(\ell ^{\prime }\), which is associated with the energy of the peptide bond (hydrogen bond energy). Moreover, \(t_{\textrm{C}i\ell }\) indicates the overlap term between the DNA strand and the protein molecule and \(\mathrm {h.c.}\) denotes the Hermitian conjugate. Furthermore, the operators \(c^{\dag }_{i}\) (\(c^{\dag }_{\ell }\)) and \(c_{i}\) (\(c_{\ell }\)) are defined as the creation and annihilation operators for a \(\pi\)-electron at sub-site i (\(\ell\)). It is essential to acknowledge that variations in temperature can induce structural disorder and fluctuations within the protein28. Therefore, we investigate the effects of temperature changes on the hopping terms, particularly emphasizing the fluctuations in twist angles. We denote \(\theta _{\ell \ell ^{\prime }}\equiv \theta\) to represent the fluctuations in twist angles between adjacent sub-sites, which are comparable to the torsional angles found along a chemical bond. By assuming a Gaussian distribution for the variable \(\theta\), we find that \(\langle \theta \rangle =0\) and \(\langle \theta ^{2}\rangle =k_{\textrm{B}}T/(\mathcal{I}\omega ^{2})\), where \(\mathcal{I}\omega ^{2}\simeq 0.022\;\textrm{eV}\). Here, \(\mathcal{I}\) denotes the reduced moment of inertia related to the relative rotation of nn amino acids, while \(\omega\) represents the oscillator frequency associated with the vibrational mode28. These formulations enable the derivation of temperature-dependent hopping integrals, which can be approximated as \(\cos (\theta )\simeq 1-\theta ^{2}/2\). We operate within a framework where the physical constants are normalized to unity.
If we consider a DNA strand (protein chain) comprising \(N_{\textrm{D}}\) (\(N_{\textrm{P}}\)) sub-sites, the total Hamiltonian, denoted as \({\varvec{H}}\), can be represented in a matrix form as a \(N\times N\) matrix where \(N=N_{\textrm{D}}+N_{\textrm{P}}\). This matrix includes sub-matrices: \({\varvec{H}}_{\textrm{D}}\) for the DNA strand, \({\varvec{H}}_{\textrm{P}}\) for the protein molecule, and \({\varvec{H}}_{\textrm{C}}\) for the interaction or coupling component of the Hamiltonian:
In Eq. (5), the Hamiltonian sub-matrix of the DNA strand can be represented as:
in this equation, the parallel hopping amplitudes (\(t^{\parallel }_{\textrm{D}ij}\)) between nn sub-sites are represented by \(t_{1}\) and \(t_{2}\), whereas \(t_{3}\) denotes the term for perpendicular hopping (\(t^{\perp }_{\textrm{D}ij}\)). One would acknowledge that the distinct parallel hoppings lead to the introduction of dimerization parameters. According to the Hamiltonian in Eq. (6), we have selected a sequence of AT and GC within the DNA strand. In fact, a DNA strand characterized by repetitive AT and GC bps is a prevalent characteristic in genomes, exhibiting diverse distributions across various regions. These repeating motifs, particularly as simple sequence repeats, may be associated with numerous genomic functions and even diseases44. The protein Hamiltonian sub-matrix of Eq. (5) for the \(\mathrm {P_{1}}\) conformation can be presented by:
wherein \(\overline{t}_{\ell }\) is defined as \(t_{\textrm{P}\ell \ell +1}\cos (\theta )\). The Hamiltonian matrices for \(\mathrm {P_{2}}\) and \(\mathrm {P_{3}}\) can be calculated in a comparable fashion. The nonzero elements of the Hamiltonian sub-matrix presented in Eq. (5) related to the interaction component are contingent upon the number of coupling points between the protein molecule and the DNA strand. For instance, if the interaction between the protein and DNA occurs at a single point located at sub-site number 3 on one side of the DNA strand, the nonzero elements of \({\varvec{H}}_{\textrm{C}}\) would be represented as \(H^{\textrm{C}}_{3,N_{\textrm{D}}+1}=H^{\textrm{C}}_{N_{\textrm{D}}+1,3}=t_{\textrm{C}i\ell }\equiv t^{\prime \prime }_{1}\) (Fig. 1). As we wrap up this section, we emphasize that the characteristic equation presented below can serve to numerically identify the spectrum of the system through the total Hamiltonian matrix \({\varvec{H}}\):
where the symbol \(\mathcal{E}\) denotes the energy levels of \(\pi\)-electrons, and \({\varvec{I}}\) represents a unit matrix of size \(N\times N\). As a result, the system is defined by N eigenvalues.
Green’s function and DOS
Green’s functions are essential in various applications of quantum mechanics, especially in the study of transport phenomena, as they enable the investigation of particle dynamics within complex systems. The Green’s function is fundamentally connected to the Hamiltonian operator of the system, and its properties are closely related to the system’s energy spectrum. The matrix representation of the Green’s function45 associated with the Hamiltonian can be formulated as:
wherein \(E=\mathcal{E}+\textrm{i}0^{+}\). The imaginary component of the Green’s function at a specific energy level reflects the probability of finding a state at that energy. In the context of quantum mechanics, the DOS at a given energy can be directly obtained from the imaginary part of the trace of the Green’s function associated with that energy. To determine the DOS for the system, it is necessary to calculate the trace over all quantum numbers that characterize the Hamiltonian:
The trace presented in Eq. (10) can be calculated in the following manner:
where both site indices i and \(\ell\) are shown by m. To obtain the DOS for either DNA or protein independently, one may incorporate the relevant sub-matrices from Eqs. (6) and (7) into the Green’s function as defined in Eq. (9).
Results and discussion
The formalism established in the preceding sections is employed to examine the coupling of various protein conformations, designated as \(\mathrm {P_{1}}\), \(\mathrm {P_{2}}\), and \(\mathrm {P_{3}}\), with the double-strand DNA, utilizing the energy spectrum of the system. This study investigates the effects of different numbers of contact points (CPs) between protein and DNA, the consequences of altering the length of the DNA strand, and the impact of temperature on the BS and DOS of the system. Initially, we outline the key parameters necessary for obtaining numerical results. The DNA model features three distinct hopping parameters, where \(t^{\parallel }_{\textrm{D}\ell \ell ^{\prime }}\equiv \{t_{1}, t_{2}\}\) represent the parallel hopping terms between bases, and \(t^{\perp }_{\textrm{D}\ell \ell ^{\prime }}\equiv t_{3}\) denotes the perpendicular hopping amplitudes (Fig. 1), with values of \(t_{1}=-0.35\;\textrm{eV}\) and \(t_{3}=-0.005\;\textrm{eV}\)46, and the dimerization parameter is defined as \(t_{2}=0.75t_{1}\). Furthermore, the hopping term within the protein is established based on the values referenced in28, specifically \(t_{\textrm{P}\ell \ell ^{\prime }}\equiv t_{p}=-0.3\;\textrm{eV}\). For the assessment, a configuration of \(t_{p}\) is generated across thirty-six sub-sites, with \(t_{p}\) values varying within the range of \(-0.3\le t_{p}\le -0.26\). Consequently, as outlined following Eq. (7), we derive \(\overline{t}_{\ell }=t_{p}\cos (\theta )\) for \(\ell =1,2,3,\cdots ,35\), utilizing thirty-five distinct values of \(t_{p}\). Additionally, the energy associated with hydrogen bonds indicates relatively weak intermolecular interactions compared to peptide bonds, as illustrated in Fig. 1a by dashed lines, thus we set \(t^{\prime }_{p}=-0.03\;\textrm{eV}\). Despite the relative weakness of hydrogen bonds, their abundance and the specific arrangement of these bonds among amino acids in the chain can considerably stabilize and influence the spatial configuration of proteins. It is crucial to recognize that protein sequences offer a more intricate alignment than DNA, as DNA is made up of merely four nucleotides, while proteins are formed from twenty distinct amino acids. In this regard, we explore thirteen connections that play a vital role in stabilizing these conformational structures28,43. We posit that all on-site energies within the system are normalized to zero, thereby creating a baseline for the energy levels. Although this may seem like an oversimplification, this basic model has successfully captured the semiconducting characteristics of DNA double-strands by utilizing diverse values for the hopping parameters or dimerization effects2,3,4,5. In general, when the on-site energy is set to zero, the Hamiltonian simplifies, facilitating an analysis of the correlation between the hopping parameters and the lattice geometry. To design a semiconducting material by manipulating the on-site energy and hopping parameters, one can employ a tight-binding model to depict the electronic structure of the material. By fixing the on-site energy at zero and adjusting the parallel hopping parameters between adjacent sites, it becomes feasible to create an energy band gap, which is a characteristic feature of semiconductors2,4,5. As a result, rather than maintaining a uniform overlap among sub-sites in each segment of the system, we suggest that the hopping parameters fluctuate across protein chains from one site to another, while still being close to the value indicated in28 and taking into account the dimerization within the DNA component of the system. In essence, to engineer a semiconducting material through the manipulation of on-site energy and hopping parameters, one can utilize a TB model to represent the electronic structure of the material. By establishing the on-site energy at zero and altering the hopping parameters among adjacent sites, it is possible to induce the emergence of an energy band gap, a defining feature of semiconductors. Furthermore, with the on-site energy fixed at zero, the Hamiltonian simplifies, enabling a clearer analysis of the interplay between hopping parameters and lattice geometry. It is significant to highlight that while a protein may interact with DNA at a singular site, effective recognition and robust binding typically necessitate interactions at multiple sites along a short sequence. This is achieved through various non-covalent interactions, including hydrogen bonds, ionic interactions, and van der Waals forces, which collectively enhance the stability of the protein-DNA complex47. The primary atomic interactions occur between the nitrogen and oxygen atoms of the nucleotide bases, which are accessible in the major and minor grooves of the DNA structure, facilitating the formation of hydrogen bonds with specific amino acid side chains of the protein48. Furthermore, it is noteworthy that the distance between adjacent nucleotides (or bps) in a DNA strand is approximately \(0.34\;\textrm{nm}\), while the inter-atomic distance between amino acids in a protein chain is generally around \(0.38\;\textrm{nm}\), although this can fluctuate based on the specific amino acids involved and the overall conformation of the protein. The typical range for interaction distances between side chains is estimated to be between 4 and 6 \(\textrm{nm}\)49. In the context of DNA bases such as G, C, T, and A, the term ’hopping electrons’ refers to the transfer of electrons from one base to another along the DNA strand. G is particularly notable for its propensity to accept electrons due to its relatively low ionization potential, which facilitates its ability to gain electrons. This phenomenon is termed electron hopping, as electrons transition between adjacent bases in the DNA sequence, predominantly occurring between stacked GC pairs. Among the DNA bases, G exhibits the lowest ionization energy, followed by A, C, and T, which has the highest ionization energy. This indicates that G is the most readily ionizable base, while T is the least. The energy required for electron transfer, which is the process of hopping between an amino acid and a nucleic acid, typically ranges from approximately \(0.5\;\textrm{eV}\) to \(2\;\textrm{eV}\), contingent upon the specific amino acid and nucleic acid base involved, as well as the distance separating them. The most probable electron hopping events occur between aromatic amino acids, such as tyrosine and tryptophan, and the nucleobases G or A24,50. We analyze the interactions of \(\pi\)-electrons among protein chains and DNA strand, considering one to four CPs between amino acids and nucleic acids. Four unique interactions between protein molecules and the DNA strand are analyzed separately. These interactions are classified into four distinct modes of interaction: (i) sub-site 1 of the protein molecules, (ii) sub-sites 19 and 20 of the protein molecules, (iii) sub-sites 29, 30, and 31 of the protein molecules, and (iv) sub-sites 1, 2, 3, and 4 of the protein molecules (see Fig. 1). Each mode delineates a unique interaction pattern between the protein molecules and the DNA strand. The coupling terms (\(t_{\textrm{C}i\ell }\)) are quantified as \(t^{\prime \prime }_{1}=-1.0\;\textrm{eV}\), \(t^{\prime \prime }_{2}=-1.25\;\textrm{eV}\), \(t^{\prime \prime }_{3}=-0.75\;\textrm{eV}\), and \(t^{\prime \prime }_{4}=-0.5\;\textrm{eV}\), which arise from the interaction of various sub-sites of DNA with protein molecules in each mode of interaction. In order to investigate the interaction between the proteins and the DNA strand, we analyze a DNA model that incorporates \(N_{\textrm{D}}\) values of 40 and 160 sub-sites.
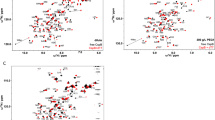
The DOS associated with an individual DNA strand and three distinct conformations of the protein chains at a constant temperature of \(T=0.001\) (\(3.48\textrm{K}\)), along with \(N_{\textrm{D}}=40\) sub-sites of DNA, demonstrates semiconducting properties and localization, with the DNA showing a more pronounced energy gap.
Figure 2 illustrates the DOS of DNA in comparison to three distinct conformations of protein molecules at a constant temperature of \(T=0.001\) or \(3.48\;\textrm{K}\) and for \(N_{\textrm{D}}=40\) sub-sites of DNA. The results indicate that all examined molecules display semiconducting properties, with DNA demonstrating a more substantial energy gap. The DOS for these finite systems is characterized by localization due to the quantization of energy levels inherent to the system. In such finite systems, the energy states are discrete, possessing a limited number of quantum states, in contrast to an infinite system where energy levels are continuous. This discreteness results in a localized DOS, wherein the likelihood of locating an electron at a specific energy level is concentrated in defined spatial regions rather than being uniformly distributed. The boundaries of these molecules confine particle movement, leading to quantized energy levels, which means that only certain discrete energy values are permissible for the particles within the system. Consequently, the DOS, which quantifies the number of available energy states per unit energy, becomes concentrated at these discrete energy levels. This phenomenon engenders a localized DOS, where the probability of finding an electron at a given energy level is focused in particular spatial locations. The implications of a localized DOS in finite systems are profound, influencing the behavior of particles within the system and potentially altering the physical properties of the system.
On the left side, the BS is displayed, whereas the right side demonstrates the DOS for DNA interacting with each of the three conformations of the protein molecules, with each panel corresponding to a particular number of CPs. The BS is shown solely for the \(\mathrm {P_{1}}\) conformation of the protein, with the temperature held constant at \(T=0.001\) (\(3.48\;\textrm{K}\)) and \(N_{\textrm{D}}=40\).
Figure 3 depicts the DOS and BS of the protein-DNA system, focusing on the interactions between three unique conformations of protein molecules and a DNA strand. Each panel within this figure presents distinct CP numbers for DNA-protein interactions, with parameters set at \(N_{\textrm{D}}=40\) and \(T=0.001\) or \(3.48\;\textrm{K}\). The BS is represented solely for the \(\mathrm {P_{1}}\) conformation of the protein to enhance the clarity of the diagram. The BS is depicted in terms of site index, which encompasses 76 sub-sites within the protein-DNA system. The BS diagrams are positioned on the left, while the DOS for all three conformations is shown on the right, with the DOS diagrams corresponding to the BS represented in matching color. It is crucial to acknowledge that while a traditional BS plot, predicated on the assumption of a periodic lattice, is not directly applicable to finite systems, we can still explore the energy levels of protein-DNA complexes in a manner that emulates the BS. This exploration can be conducted by plotting the energy levels as a function of a relevant parameter, such as the site index, which provides a representation of the band-like features inherent in this system. Figure 3 illustrates discrete energy levels that can be graphed against the selected parameter to depict the energy distribution. Essentially, within this finite system, the BS, when expressed in terms of the site index, represents the energy levels plotted against the site number or the position of each block within the system. This effectively showcases how an electron’s energy varies as it moves from one sub-site to another within the protein-DNA framework, with each distinct energy level corresponding to a band in the system, and the site index acting as a discrete variable along the x-axis of the graph. The confinement of electrons to limited dimensions leads to the quantization of energy levels. As a result, instead of forming a continuous band, the allowed energy states are converted into discrete values, resembling distinct steps or individual levels. This phenomenon, referred to as quantum confinement, arises when a system of constrained dimensions leads to the quantization of its energy levels. Additionally, the flat band in the BS represents a region where the energy of electrons remains constant regardless of their momentum. This results in a flat energy dispersion curve on a graph, indicating that the charge carriers within this band possess nearly zero group velocity and are highly localized. Furthermore, this figure illustrates that three conformations exhibit distinct energy levels or sub-band characteristics, implying the absence of states between these levels, particularly near the zero energy point. When a system, such as a molecule, engages with its environment, like a DNA strand, energy transfer may occur between the two. This engagement can result in the emergence of novel energy states or modifications to the density of pre-existing states. If the interaction between the system and its environment generates new energy levels or substantially modifies the density of existing ones, it may be observed as peaks in the DOS. In Fig. 3, the influence of protein-DNA interactions is evident, as indicated by the presence of peaks in the BS and DOS, with the quantity of additional peaks. The observed peaks in the DOS, or the additional bands in the BS, are a consequence of the interactions between protein molecules and DNA strands. Such interactions may give rise to the emergence of novel energy levels or modifications in the density of pre-existing levels, thereby producing peaks in the DOS. The precise positioning and morphology of these peaks are contingent upon the characteristics of the interaction between the protein and the DNA. The peaks resulting from protein-DNA interactions resemble those caused by impurities within a system. In fact, the presence of impurities in a material can generate localized states that profoundly influence the electronic structure and characteristics of the material. These localized states may occur around the impurity atom itself. The characteristics of these peaks, including their position, height, and width, are contingent upon various factors such as the nature of the impurity, its concentration, and its interactions with the surrounding host material. The presence of impurities within a material can result in the emergence of peaks in the DOS that are situated away from zero energy. These peaks, commonly known as impurity bands, originate from the localized energy levels established by the impurity atoms, which contrast with the band structure of the unadulterated material.
As previously mentioned in relation to Fig. 2, the individual components of the system, namely protein and DNA, exhibit semiconducting properties. However, as illustrated in all panels of Fig. 3, the interaction between these two components results in a narrower band gap in panel (a) when compared to the DOS of the separate protein or DNA shown in Fig. 2. Furthermore, panels (b) to (d) of Fig. 3 reveal a metallic behavior, characterized by the emergence of a peak at the zero point energies and diminish of the gap around the Fermi level. Thus, it can be concluded that the interaction between protein and DNA significantly modifies the electrical characteristics of the system, effectively enhancing its metallic properties. This transition from semiconducting to metallic behavior is attributed to the increased availability of electron channels in both components of the system. In order to concentrate more on this phenomenon, the band energies of the system are illustrated in Fig. 4, depicting only the one lower and one upper bands relative to zero energy for various CP numbers. These correspond to a specific conformation of the protein chain at a constant temperature of \(T=0.001\) or \(3.48\;\textrm{K}\) with \(N_{\textrm{D}}=40\) in all panels. It is observed that as the number of CPs increases, the gap around the Fermi level vanishes, resulting in metallic behavior. This phenomenon can be ascribed to the rising quantity of electron channels that enhance mobility within the system, which is further elucidated by the heightened interaction of impurities with DNA resulting from the increasing number of charge carriers. Furthermore, the presence of non-covalent (hydrogen) bonds in the \(\mathrm {P_{2}}\) and \(\mathrm {P_{3}}\) conformations creates additional channels when compared to the \(\mathrm {P_{1}}\) conformation. This additional bonding becomes more evident when the \(\mathrm {P_{3}}\) conformation of the protein molecule interacts with DNA in three CPs, which incorporate hydrogen bonds in the CPs, resulting in a greater reduction of energy level intervals approaching the zero point energy. Enhancing the quantity of bonding interactions between two components of the system typically results in a reduction of the gap energy. This phenomenon occurs because intensified bonding interactions facilitate greater overlap of atomic orbitals, which in turn leads to a broader distribution of energy levels within the energy bands, consequently resulting in a diminished gap between them.
The length of a DNA molecule can greatly influence the manner in which proteins interact with it. The sensitivity to the length of the DNA strand is also observable in the mechanical response of DNA to external forces51. Extended DNA molecules frequently demonstrate a greater affinity for protein binding, attributed to the heightened likelihood of protein-DNA interactions, a phenomenon referred to as the antenna effect. This effect occurs because longer DNA offers more chances for a protein to locate its specific binding site through diffusion along the DNA strand52. In fact, proteins are capable of traversing the DNA molecule (1D diffusion) to identify their specific binding sites. Lengthier DNA facilitates more effective diffusion, thereby increasing the probability of locating the target52. Nevertheless, our focus here is on examining the effects of DNA length concerning four distinct CPs between protein and DNA in their interaction. Notably, the positions of these CPs remain constant, while the length of the DNA strand is variable. To examine the impact of DNA length on protein-DNA interactions, the DOS for the system is illustrated for three distinct values of \(N_{\textrm{D}}=40,160,800\) at a fixed temperature of \(T=0.001\) or \(3.48\;\textrm{K}\) on Fig. 5. This analysis incorporates four CPs between the protein and DNA, along with a defined conformation of the protein chain represented in each panel of Fig. 5. This illustration demonstrates that the electronic properties of the protein-DNA system are affected by the size of the DNA, leading to a transformation of localized states within the structures into less distinct energy levels or sub-bands. Additionally, regardless of the length of DNA, the illustration reveals that the curves display symmetry about the zero-energy point, which is a result of the particle-hole symmetry present in the Hamiltonian framework. The figure indicates that the amplitude of peaks in the DOS diminishes with the increasing length of the DNA strand. This phenomenon occurs because the discrete energy levels characteristic of smaller systems, which are represented as sharp peaks in the DOS, tend to broaden and become less distinct as the system expands, with the spacing between these levels decreasing. Specifically, smaller systems demonstrate quantization, where energy is confined to particular levels, resulting in clearly defined peaks in the DOS. Conversely, as the system size increases, the energy levels converge, leading to a reduction in the visibility of quantization effects. Consequently, the total number of accessible energy states rises, contributing to a denser DOS. This implies that while the individual peaks may exhibit reduced height, the cumulative area beneath the DOS curve, indicative of the total number of states, increases. Ultimately, as the system enlarges, these discrete energy levels broaden and merge, causing the peaks to become less pronounced and resulting in a more continuous and smooth DOS. In all scenarios depicted across the three panels, it is evident that proteins generate identical peaks and alter the electronic properties of DNA, transitioning its behavior from semiconducting to metallic, irrespective of the DNA strand’s length. As the DNA size increases, the peak heights related to protein states in the DOS typically diminish. This occurs because, in larger DNA, the protein states tend to broaden and blend with the conduction band or other energy bands, resulting in a reduction of the localized DOS at particular energy levels. Conversely, in shorter DNA, the protein molecule establishes localized states at a specific energy level. Consequently, this leads to a pronounced peak in the DOS at that energy.
The BS of the protein-DNA system tends towards zero energy across multiple conformations of the protein molecule and various CPs. The figure illustrates a single lower and a single upper band in relation to zero energy for different quantities of CPs. These bands are associated with a specific conformation of the protein chain maintained at a constant temperature of \(T=0.001\) or \(3.48\;\textrm{K}\) with \(N_{\textrm{D}}=40\).
The influence of DNA length on protein-DNA interactions is demonstrated by the DOS for the system at two specific values of \(N_{\textrm{D}}=40,160\) at a constant temperature of \(T=0.001\) or \(3.48\;\textrm{K}\), where DNA strands engage with the various conformations of the protein molecule depicted in each panel.
The influence of temperature on the DOS curves related to protein-DNA interactions is illustrated in panels (a), (b), and (c) for three unique conformations of protein chains at two varying temperatures, \(T=0.001\) (\(3.48\;\textrm{K}\)) and \(T=0.010\) (\(34.8\;\textrm{K}\)).
Figure 6 illustrates the influence of temperature on the DOS curves associated with protein-DNA interactions, as shown in panels (a), (b), and (c) for three distinct conformations of protein chains at two different temperatures, \(T=0.001\) (\(3.48\;\textrm{K}\)) and \(T=0.010\) (\(34.8\;\textrm{K}\)). An increase in temperature results in modifications to both the peak heights and positions across the three protein conformations interacting with DNA. This phenomenon arises from the alterations in the hopping terms within the protein molecules, which are temperature-dependent and reflected in the Hamiltonian of the protein chain. Consequently, changes in the hopping terms of the protein molecules modify the system’s spectrum, leading to variations in both the position and height of the peaks. It is evident that the DOS, which quantifies the number of accessible electronic states at a specific energy level, is intrinsically linked to the energy spectrum of the material. Any shifts or alterations in the energy spectrum will correspondingly affect the DOS. For instance, the emergence of a new energy level in the spectrum will manifest as an additional peak in the DOS, while a shift in an existing energy level will result in a corresponding displacement of the peak in the DOS. Furthermore, the peak height, indicative of the number of states, may also be influenced by changes in the energy spectrum. Furthermore, this phenomenon can be understood as a standard broadening of the fermionic distribution at elevated temperatures; the energy levels of electrons also increase, facilitating their movement throughout the system. As temperature rises, atomic vibrations intensify, leading to an increased average separation between atoms, a process referred to as thermal expansion. This increased interatomic distance reduces the potential barriers faced by electrons, consequently decreasing the energy required to transition electrons across the energy gap between the peaks. This leads to changes in both the heights and positions of the peaks.
The stretching of the DNA strand refers to the modification of its structure, where the atoms are either separated or compressed. This phenomenon can arise through various mechanisms, including the introduction of impurities or defects, as well as the interaction of proteins with the DNA strand, which can lead to distortions or stretching of its configuration. These alterations may be reflected in changes to the electronic DOS, potentially resulting in a shift of spectral features within the DOS. Taking into account the effect of stretching on the DOS, we can manipulate it within the TB model by varying the hopping parameters in DNA and the coupling strength between protein molecules and the DNA strand. For this purpose, we selected two additional sets of hopping parameters: one set (set 2) consists of 0.8 of the values of the hopping parameters used to create previous figures (set 1), while the other set has been increased by a factor of 1.2 (set 3) compared to set 1. Figure 7 demonstrates the effect of reducing the hopping parameters in DNA and the coupling strength between protein molecules and the DNA strand (set 2) relative to set 1 by a factor of 0.8, as well as the effect of increasing the same hopping parameters (set 3) by a factor of 1.2 on the DOS curves related to the protein-DNA system for three conformations of protein molecules across three panels at fixed temperature \(T=0.001\) or \(3.48\;\textrm{K}\) with \(N_{\textrm{D}}=40\). The variations in the hopping terms, whether they are increasing or decreasing, result in alterations in the DOS by shifting and redistributing spectral features within the DOS. These changes can stem from local structural distortions, such as stretching, which introduce localized energy levels into the energy band. Consequently, this leads to an increase or decrease in the DOS at specific energy levels, resulting in a shift of the peaks in the DOS. This shift alters the overall shape of the DOS when comparing the unstretched DNA sequence to the stretched DNA strand. Therefore, the interaction between protein molecules and the unstretched DNA sequence, as opposed to the stretched DNA strand, can be distinctly influenced by the DOS within the TB model. This has biological implications, including gene regulation, DNA repair, and the efficiency of protein-DNA interactions53.
The impact of various hopping parameters resulting from the elongation of DNA sequences on the DOS of the protein-DNA system is analyzed for three unique conformations of protein chains, with each conformation illustrated in an individual panel. The temperature is maintained at \(T=0.001\) or \(3.48\;\textrm{K}\) with \(N_{\textrm{D}}=40\).
Conclusion
In summary, the TB Hamiltonian model was employed alongside the Green’s function methodology to explore the protein-DNA interaction across three unique protein chain conformations, referred to as \(\mathrm {P_{1}}\), \(\mathrm {P_{2}}\), and \(\mathrm {P_{3}}\), within the context of BS and DOS analysis. Under constant temperature and a fixed number of sub-sites within the DNA strand, the BS and DOS exhibited the flat energy dispersion curves for both the protein molecules and the DNA strand independently, which can be attributed to the existence of localized states within these structures. Indeed, the confines of these molecules restrict particle mobility, resulting in quantized energy levels. This indicates that only specific discrete energy values are allowed for the particles within the system. Furthermore, these finite structures exhibited characteristics akin to those of semiconductors, which are associated with the existence of a forbidden zone near zero energy, with DNA showing a more pronounced energy gap. The interaction between the two components of the system introduces novel effects in the protein-DNA system that are contingent upon the number of CPs between the protein and DNA. An increase in CPs leads to a transition from semiconducting to metallic characteristics, which is also influenced by the contribution of hydrogen bonds from the mutant protein at these CPs. Such interactions may lead to the formation of new energy levels or alterations in the density of existing levels, consequently generating peaks in the DOS. The electronic properties of the protein-DNA system are influenced by the size of the DNA, resulting in a transformation of localized states within the structures into less defined energy levels as the length of the DNA strand increases. This phenomenon arises because the discrete energy levels typical of smaller systems, depicted as sharp peaks in the DOS, tend to broaden and become less defined as the system enlarges, with the spacing between these levels diminishing. The effect of temperature on the BS and DOS, irrespective of the type of configuration, results in changes in both the peak heights and positions due to modifications in the hopping terms within the protein molecules and alterations in the system’s spectrum. This occurrence can be interpreted as a conventional broadening of the fermionic distribution at higher temperatures; the energy levels of electrons also rise, promoting their movement throughout the system. The fluctuations in the hopping parameters, whether they are on the rise or decline, caused by the stretching effects of DNA due to the presence of protein molecules, lead to changes by shifting and redistributing spectral characteristics within the DOS. The interaction between protein and DNA is expected to directly affect the electronic properties of DNA, which vary across different protein conformations, thereby opening new research avenues with significant biological implications.
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
Watson, J. D. & Crick, F. H. C. Molecular structure of nucleic acids: A structure for deoxyribose nucleic acid. Nature 171, 737 (1953).
Mousavi, H., Khodadadi, J. & Grabowski, M. Electronic properties of long DNA nanowires in dry and wet conditions. Solid State Commun. 222, 42 (2015).
Mousavi, H. & Grabowski, M. Nonlinear electron transport across short DNA segment between graphene leads. Solid State Commun. 279, 30 (2018).
Mousavi, H., Jalilvand, S., Sani, S. S., Hartman, J. A. L. & Grabowski, M. Electronic properties of different configurations of double-strand DNA-like nanowires. Solid State Commun. 319, 113974 (2020).
Jalilvand, S., Sepahvand, R. & Mousavi, H. Electronic behavior of randomly dislocated RNA and DNA nanowires: A multi-model approach. Euro. Phys. J. Plus 137, 928 (2022).
Mousavi, H. Electron transport through protein chains interfacing nanoribbon and nanotube leads. Solid State Commun. 404, 116057 (2025).
Mousavi, H. Quantum transport in protein chains. Sci. Rep. 15, 26703 (2025).
Mousavi, H. & Emami, R. Protein chains in tight-binding framework. J. Mol. Model. 31, 223 (2025).
Keren, K., Berman, R. S., Buchstab, E., Sivan, U. & Braun, E. DNA-templated carbon nanotube field-effect transistor. Science 302, 1380 (2003).
Cao, X., Chen, C. & Zhu, Q. Biosensors based on functional nucleic acids and isothermal amplification techniques. Talanta 253, 123977 (2023).
Fu, Z., Lu, Y.-C. & Lai, J. J. Recent advances in biosensors for nucleic acid and exosome detection. Chonnam. Med. J. 55, 86 (2019).
Gunasekaran, B. M., Srinivasan, S., Ezhilan, M. & Nesakumar, N. Nucleic acid-based electrochemical biosensors. Clin. Chim. Acta 559, 119715 (2024).
Bhopatkar, A. A., Uversky, V. N. & Rangachari, V. Disorder and cysteines in proteins: A design for orchestration of conformational see-saw and modulatory functions. Prog. Mol. Biol. Transl. Sci. 174, 331 (2020).
Douglas, S. M. et al. Self-assembly of DNA into nanoscale three-dimensional shapes. Nature 459, 414 (2009).
Rajendran, A., Endo, M. & Sugiyama, H. Single-molecule analysis using DNA origami. Angew. Chem. Int. Ed. Engl. 51, 874 (2012).
Hart, W.E. & Newman, A. Handbook of Computational Molecular Biology, Ch. 30, Edited by S. Aluru, 1st Ed. (2005).
Dill, K. A. et al. Principles of protein folding: A perspective from simple exact models. Prot. Sci. 4, 561 (1995).
Redding, S. & Greene, E. C. How do proteins locate specific targets in DNA?. Chem. Phys. Lett. 570, 1 (2013).
Rice, P. A. & Correll, C. C. Protein-Nucleic Acid Interactions: Structural Biology (The Royal Society of Chemistry, Cambridge, 2008).
Murugan, R. DNA-protein interactions under random jump conditions. Phys. Rev. E 69, 011911 (2004).
Murugan, R. Generalized theory of site-specific DNA-protein interactions. Phys. Rev. E 76, 011901 (2007).
Murugan, R. Packaging effects on site-specific DNA-protein interactions. Phys. Rev. E 79, 061920 (2009).
Vasumathi, V. & Daniel, M. Base-pair opening and bubble transport in a DNA double helix induced by a protein molecule in a viscous medium. Phys. Rev. E 80, 061904 (2009).
D’Acunto, M. Protein-DNA target search relies on quantum walk. Biosyst. 201, 104340 (2021).
Kamagata, K., Itoh, Y. & Subekti, D. G. R. How p53 molecules solve the target DNA search problem: A review. Int. J. Mol. Sci. 21, 1031 (2020).
Monico, C., Capitanio, M., Belcastro, G., Vanzi, F. & Pavone, F. S. Optical methods to study protein-DNA interactions in vitro and in living cells at the single-molecule level. Int. J. Mol. Sci. 14, 3961 (2013).
Bonnet, I. et al. Sliding and jumping of single EcoRV restriction enzymes on noncognate DNA. Nucleic Acids Res. 36, 4118 (2008).
Sarmentoa, R. G., Frazaob, N. F. & Macedo-Filhoc, A. Electronic transport on the spatial structure of the protein: Three-dimensional lattice model. Phys. Lett. A 381, 276 (2017).
Zoli, M. Twist-stretch profiles of DNA chains. J. Phys. Condens. Matter 29, 225101 (2017).
Xu, W., Dunlap, D. & Finzi, L. Energetics of twisted DNA topologies. Biophys. J. 120, 3242 (2021).
Strick, T., Allemand, J.-F., Croquette, V. & Bensimon, D. Twisting and stretching single DNA molecules. Prog. Biophys. Mol. Biol. 74, 115 (2000).
Liu, X., Yu, E., Zhao, Q., Han, H. & Li, Q. Enzymes as green and sustainable tools for DNA data storage. Chem. Commun. 61, 2891 (2025).
Mann, M., Kumar, C., Zeng, W.-F. & Strauss, M. T. Artificial intelligence for proteomics and biomarker discovery. Cell Syst. 12, 759 (2021).
Sim, J., Kim, D., Kim, B., Choi, J. & Lee, J. Recent advances in AI-driven protein-ligand interaction predictions. Curr. Opin. Struct. Biol. 92, 103020 (2025).
Suh, D., Lee, J. W., Choi, S. & Lee, Y. Recent applications of deep learning methods on evolution- and contact-based protein structure prediction. Int. J. Mol. Sci. 22, 6032 (2021).
Qu, C. & Du, H. Advance technologies for DNA-protein interactions and future research prospect. Crop Des. 4, 100082 (2025).
Kimijima, J., Inagawa, A., Miyagawa, A., Nasuno, E. & Uehara, N. Probing the interaction between biomolecules under sub-zero temperature conditions by electrophoresis in ice grain boundaries. Anal. Chim. Acta. 1311, 342713 (2024).
Foguel, D. & Silva, J. L. Cold denaturation of a repressor-operator complex: the role of entropy in protein-DNA recognition. PNAS 91, 8244 (1994).
Zheng, J., Guo, N., Huang, Y., Guo, X. & Wagner, A. High temperature delays and low temperature accelerates evolution of a new protein phenotype. Nat. Commun. 15, 2495 (2024).
Privalov, P. L. et al. What drives proteins into the major or minor grooves of DNA?. J. Mol. Biol. 365, 1 (2007).
Hauschild, K. E. et al. Temperature-sensitive protein-DNA dimerizers. PNAS 102, 5008 (2005).
Muthusamy, M., Kim, J.-H., Kim, J. A. & Lee, S.-I. Plant RNA binding proteins as critical modulators in drought, high salinity, heat, and cold stress responses: An updated overview. Int. J. Mol. Sci. 22, 6731 (2021).
Gutin, A. M., Abkevich, V. & Shakhnovich, E. A protein engineering analysis of the transition state for protein folding: Simulation in the lattice model. Fold. Des. 3, 183 (1998).
Liao, X. et al. Repetitive DNA sequence detection and its role in the human genome. Commun. Biol. 6, 954 (2023).
Economou, E. N. Green’s Functions in Quantum Physics 3rd edn. (Springer-Verlag, 2006).
Paez, C. J., Schulz, P. A., Wilson, N. R. & Römer, R. A. Robust signatures in the current-voltage characteristics of DNA molecules oriented between two graphene nanoribbon electrodes. N. J. Phys. 14, 093049 (2012).
Pandey, P., Hasnain, S. & Ahmad, S. Protein-DNA interactions, encyclopedia of bioinformatics and computational. Biology 2, 142 (2019).
Luscombe, N. M., Laskowski, R. A. & Thornton, J. M. Amino acid-base interactions: A three-dimensional analysis of protein-DNA interactions at an atomic level. Nucleic Acids Res. 29, 2860 (2001).
Rico, F., Rigato, A., Picas, L. & Scheuring, S. Mechanics of proteins with a focus on atomic force microscopy. J. Nanobiotechnol. 11, S3 (2013).
Beratan, D. N. Why are DNA and protein electron transfer so different?. Annu. Rev. Phys. Chem. 70, 71 (2019).
Singh, A. & Singh, N. Pulling DNA: The effect of chain length on the mechanical stability of DNA chain. Macromol. Symp. 357, 64 (2015).
Shimamoto, N. et al. Dependence of DNA length on binding affinity between TrpR and trpO of DNA. Sci. Rep. 10, 15624 (2020).
King, G. A., Peterman, E. J. G. & Wuite, G. J. L. Unravelling the structural plasticity of stretched DNA under torsional constraint. Nat. Commun. 7, 11810 (2016).
Funding
This study is in no way funded by any organization and is an independent investigation carried out exclusively by the author of the study.
Author information
Authors and Affiliations
Contributions
The authors have contributed, in equal measure, to the study’s conception and findings. Material preparation, data collection and analysis were performed by Hamze Mousavi and Ronak Emami. The first draft of the manuscript was written by Hamze Mouavi and other authors commented on previous versions of the manuscript. All authors read and approved the final manuscript. Formulation and formal analysis; coding; results interpretation and writing the original draft were performed by Hamze Mousavi. Coding; results interpretation; and editing were performed by Ronak Emami.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Mousavi, H., Emami, R. Protein-DNA interaction in tight-binding paradigm. Sci Rep 15, 30565 (2025). https://doi.org/10.1038/s41598-025-15972-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-15972-z
Keywords
This article is cited by
-
Quantum charge transport in DNA and RNA systems coupled to nanoribbon electrodes
Discover Nano (2026)