Abstract
Cellulase is essential for cellulose hydrolysis and is used across industries such as food, feed, textiles, biofuel, and biomass pretreatment. However, its utility is restricted by high temperatures and salt concentrations. This study identified a novel cellulase gene (named c5-cel4) from Ebinur Salt Lake in Xinjiang, China using metagenomic technology. Its amino acid sequence has a 90.97% similarity to the GH5 family cellulase of Microbulbifer litoralis (WP_250463697.1). The gene was expressed in Escherichia coli, and the recombinant protein, C5-CEL4, was purified via Ni-NTA affinity chromatography. C5-CEL4, secreted extracellularly (0.886 U/mL), revealed a protein size 14 KDa smaller than predicted, with mass spectrometry indicating a truncated C-terminal of 135 amino acid residues. Optimal activity was observed at 50 °C and pH 7.0, maintaining over 80% activity at 40–65 °C and 45–50 °C for 2 h. The enzyme’s half-life was 60 min at 55–60 °C, retaining over 90% activity after 24 h in pH 5.0–12.0 buffers. C5-CEL4 showed activity against CMC-Na, bagasse xylan, and beech xylan, with Kcat values of 98.20 s− 1 and 12.32 s− 1 for CMC-Na and bagasse xylan, respectively. It also hydrolyzed wheat bran and maize stalks into reducing sugars. Remarkably, C5-CEL4 exhibited high salt tolerance, maintaining 100% activity in 0.5 M-5.0 M NaCl and after 9 months in 5.0 M NaCl. It retained over 90% activity in ionic liquids (BMIM-Ac, EMIM-Cl, BMIM-BF4) at 50% concentration and showed resistance to heavy metal ions (Co2+, Cu2+, Ag+, Mn2+, Pb2+, and Ni2+) and inhibitors (PMSF, DTT, Tween80, and SDS). In conclusion, C5-CEL4 is a robust cellulase with heat, alkali, salt, ionic liquid, and inhibitor resistance, alongside cellulase and xylanase activity, presenting significant potential for feed, food, and bioenergy applications.
Similar content being viewed by others
Introduction
Cellulose, a renewable organic macromolecular polysaccharide, is the most prevalent component in plant biomass1. The efficient degradation of cellulose by cellulolytic enzymes is a crucial step in the transformation of lignocellulosic materials. The complete hydrolysis of cellulose into glucose necessitates the synergistic action of endo-1,4-β-glucanase (EC 3.2.1.4), exo-1,4-β-glucanase (EC 3.2.1.91), and β-glucosidase (EC 3.2.1.21)2. Among these enzymes, endoglucanases target the amorphous regions within the cellulose molecule, initiating random cleavages that render the polymer more amenable to further hydrolysis by other cellulolytic enzymes, and are primarily responsible for cleaving internal glycosidic bonds3. Currently, cellulases find extensive applications across various industries, including pulp and paper, biofuels, food and beverage processing, textiles, animal feed, bio-nanomaterials, and phenolic compounds4. However, these enzymes are particularly vulnerable to denaturation under elevated temperatures and in saline or alkaline industrial environments, making thermal stability and salinity tolerance of paramount importance. Although numerous thermostable cellulases exhibit tolerance to salinity and polyextremophilic, only a few possess true halotolerance and the capability to maintain catalytic efficiency in high-salt conditions5,6. Consequently, the identification of salinity-tolerant cellulases is advantageous for cost reduction and efficiency enhancement in industrial processes.
Saline soil, characterized by its hypersaline nature, represents a significant reservoir for various salt-tolerant cellulases. Ionic liquids (ILs), defined as nonvolatile salts that are liquid below 100 °C, have demonstrated efficacy in solubilizing cellulose, hemicellulose, and lignin from plant biomass at moderate temperatures, thereby enhancing enzymatic hydrolysis7. These ILs exhibit minimal contamination, high thermal and chemical stability, high polarity, and very low toxicity. However, the activity of most commercial cellulases is compromised or inhibited by even small concentrations of ILs8. Consequently, the identification and engineering of new enzymes with improved IL tolerance are essential for advancing lignocellulosic bioprocesses that incorporate ILs.
As one of the planet’s most abundant renewable resources, cellulose’s effective utilization is pivotal for mitigating the energy crisis and reducing environmental pollution9. The direct utilization of cellulose is hindered by its crystalline structure and the protective role of lignin10. Cellulase, a potent biocatalyst, hydrolyzes cellulose into monosaccharides such as glucose, facilitating innovative resource utilization in sectors including energy, food, and chemicals. The enzymatic hydrolysis of lignocellulose by cellulases not only enables the efficient valorization of substantial amounts of agricultural waste but also mitigates the environmental impact associated with incineration and other conventional waste treatment methods. Despite the promising potential of cellulases, challenges remain in hydrolyzing complex substrates, including high production costs and suboptimal hydrolysis efficiency11. Furthermore, only a small fraction (0.1%-1%) of microorganisms can be cultured in laboratory conditions12,13. Metagenomic technologies, however, offer significant advantages in screening novel cellulases14. The integration of metagenomic and gene cloning techniques facilitates the discovery of extremophilic enzymes from harsh environments, which is anticipated to address the challenges of high production costs and enhance the hydrolysis efficiency of complex lignocellulosic substrates, thereby increasing sugar yields.
In this research, a novel cellulase gene, named c5-cel4, was extracted from metagenomic sequences sourced from the substrate of Ebinur Salt Lake in Xinjiang, China. The gene was cloned, a recombinant plasmid was engineered, and then introduced into Escherichia coli. The enzymatic activity of the recombinant cellulase was evaluated through heterologous expression and protein purification. Findings indicated that the enzyme exhibits thermostability, alkali resistance, halophilicity, and tolerance to ionic liquids, highlighting its potential utility in cellulase applications within the papermaking, textile, food, feed, and biofuel industries.
Materials and methods
Sample collection and metagenomic sequencing
Samples were collected from Ebinur Salt Lake in Xinjiang, China (45°09′35″ N, 83°53′21″ E), where the surface temperature was approximately 7.8 °C and the pH was 8.49. Bottom sediment samples were immediately frozen on dry ice for subsequent DNA extraction and metagenomic sequencing. DNA was extracted using the PowerSoil Kit (MOBIO, USA). Metagenomic sequencing was carried out by Suzhou GENWIZ using a HiSeq 2500 platform, and sequence data were analyzed via the IMG server (https://img.jgi.doe.gov/cgi-bin/mer/main.cgi).
Cellulase gene prediction and sequence analysis
A cellulase gene sequence, designated c5-cel4, was identified from the metagenomic database through functional prediction. The nucleotide sequence of the c5-cel4 gene was deposited in GenBank (accession NO.: PV014881). The amino acid sequence was translated using the Expasy-Translate tool, and the protein sequence was analyzed using the BLASTp program (http://blast.ncbi.nlm.nih.gov/Blast.cgi). Multiple sequence alignments and homology modeling were performed using RCSB PDB, SWISS-MODEL, AlphaFold Protein Structure Database, MEGA 7, and ESPript 3.0. Signal peptides were predicted using SignalP (http://www.cbs.dtu.dk/services/SignalP/), and protein domains were predicted using InterPro (ebi.ac.uk). Phylogenetic analysis was conducted using MEGA 7, with phylogenetic trees constructed via maximum likelihood and Poisson correction models.
Cloning and heterologous expression of C5-CEL4 gene
Primers for amplifying the full-length c5-cel4 gene were designed using Primer 5.0: TFH-c5-cel4-F (CATCATCATCATCATCATGAAATGATATTCGGCGCAGGTGCCCAG) and TFH-c5-cel4-R (GTGCTCGAGTGCGGCCGCAAGTCAGTTGGCGCTGCATTCCGGC). PCR amplification was performed with TransStarFastPfu Fly DNA polymerase (TransGen Biotech, China). The underlined sequences represent regions amplified by BamTech. The PCR protocol involved initial denaturation at 95 °C for 3 min, followed by 29 cycles of 98 °C for 20 s, 55 °C for 30 s, and 72 °C for 2 min, with a final extension at 72 °C for 10 min. PCR products were verified by 1.0% agarose gel electrophoresis (120 V, 25 min) and visualized under UV light. Bands matching theoretical sizes were sequenced, purified, and recovered using a gel extraction kit as per the manufacturer’s instructions. The PCR product was cloned into the pSHY211 vector15 using the pEASY-Uni Seamless Cloning and Assembly Kit (TransGen Biotech, China) to create the expression plasmid pSHY211-c5-cel4. Escherichia coli DH5α was employed for cloning and heterologous expression.
The strain E. coli DH5α-pSHY211-c5-cel4 was initiated in 15 mL LB broth with 100 µg/mL kanamycin at 37 °C and 180 RPM overnight to prepare the seed culture. A 5% inoculum of this seed culture was then transferred into a mixed medium consisting of 200 mL Glycerol-based salt medium (Glycerol 20 g/L, (NH4)2SO4 4 g/L, MgSO4·(7H2O) 1.2 g/L, CaCl2 0.3 g/L, K2HPO4 1 g/L, KH2PO4 1 g/L, NaNO3 1 g/L, pH 7.0) and 100 mL LB (yeast extract 5 g/L, peptone 10 g/L, NaCl 10 g/L, pH 7.0) with 100 µg/mL kanamycin, incubated at 37 °C and 180 RPM for 36 h with continuous shaking. The culture was centrifuged at 4,000 RPM for 30 min to harvest the supernate, which served as the extracellular crude enzyme solution.
Purification of intracellular and extracellular Recombinant cellulase C5-CEL4
The biomass was isolated, resuspended in 25 mL Phosphate Buffered Saline (pH 7.6) with 10 mM imidazole, and sonicated in an ice-water bath for cell disruption. The lysate was centrifuged at 4 °C and 12,000 RPM for 20 min, and the supernate was collected as the intracellular crude enzyme solution. Both intracellular and extracellular enzyme solutions were filtered and applied to an equilibrated Ni-NTA affinity chromatography column (Histrap, TransGen Biotech, China).
The Ni-NTA affinity chromatography column was resuspended in a 500 mL conical flask and agitated at ≤ 15 °C at 150 RPM for 1.5 h to facilitate protein binding. Intracellular and extracellular proteins were purified and characterized for recombinant cellulase as described by Yin et al. (2017)16. Protein concentrations were quantified with a Bradford assay kit (Order NO. C503031, Sangon Biotech, China), using bovine serum protein as the standard. Purified C5-CEL4 enzyme was resolved on a 12% SDS-PAGE, and protein bands were visualized with Coomassie Brilliant Blue R-250. Cellulase activity in the gel was determined via an enzyme profiling method. Specifically, samples were loaded onto a 12% SDS-acrylamide gel containing 0.5% (w/v) CMC-Na. Post-electrophoresis, the gel was treated with 2.5% (v/v) Triton X-100 (pH 7.0) for 30 min, soaked in 200 mM phosphate buffer for 15 min, and incubated in phosphate buffer (pH 7.0) at 45 °C for 1 h. The gel was then stained with 0.2% (w/v) Congo red for 10 min at 25 °C, and excess dye was removed with 1 M NaCl until clear active bands appeared against the gel background.
Analysis of extracellular Recombinant cellulase C5-CEL4 by mass spectrometry
The protein bands were excised into 1 mm³ pieces and placed into 1.5 mL Eppendorf tubes. They were decolorized with a 50% ACN (acetonitrile)-50% 50 mM NH4HCO3 (ammonium bicarbonate) solution for 10–30 min, followed by aspiration and disposal of the solution. This step was repeated until the pellets became colorless. Subsequently, 1000 µL of 100% ACN was added, incubated for 30 min until the pellets turned white and compact, then the ACN was discarded, and the pellets were air-dried.
For reduction and alkylation, 100 µL of 10 mM DTT (dithiothreitol) was added to the sample and incubated in a 56℃ water bath for 1 h. The supernate was removed and discarded. Then, 100 µL of 20 mM IAM (iodoacetamide) was added, and the sample was incubated in the dark at room temperature for 1 h, followed by removal and disposal of the supernate. The sample was then treated with 500 µL of decolorizing solution, vortexed, and the solution was aspirated and discarded. Another 1000 µL of 100% ACN was added, and the pellets were incubated until they turned white and compact, then the ACN was discarded, and the pellets were air-dried.
Next, 50 µL of trypsin (25 ng/µL in 50 mM NH4HCO3) was added, and the sample was sealed and incubated at 37℃ for 16 h. For peptide extraction, 100 µL of extraction solution (5% trifluoroacetic acid (TFA)-50% ACN-45% water) was added, and the sample was incubated at 37℃ for 1 h. The sample was sonicated for 5 min, centrifuged for 5 min, and the supernate was transferred to a new Eppendorf tube. This extraction process was repeated once, and the combined extracts were dried via vacuum centrifugation. Post-digestion, peptides were desalted using a desalting column and dried in a vacuum centrifugal concentrator at 45℃.
Data acquisition was performed using an Easy-nLC 1200 coupled with a Q Exactive™ Hybrid Quadrupole-Orbitrap™ Mass Spectrometer. Chromatographic separation involved mobile phase A (0.1% formic acid in water) and mobile phase B (0.1% formic acid, 80% ACN in water) with a gradient: 0–2 min, 4–8% B. 2–35 min, 8–28% B. 35–55 min, 28–40% B. 55–56 min, 95% B. and 56–66 min, 95% B. The flow rate was set at 0.6 µL/min through a column of 150 μm i.d. × 170 mm, packed with Reprosil-Pur 120 C18-AQ 1.9 μm particles.
Mass spectrometry parameters for full MS were: resolution 70,000, AGC target 4e8, maximum IT 20 ms, scan range 300–1800 m/z, and data type profile. For dd-MS2: resolution 17,500, AGC target 1e5, maximum IT 50 ms, TopN 20, and stepped NCE 30. Data analysis was conducted using Byonic 4.2.2 (https://www.proteinmetrics.com/products/byonic) software.
Molecular cloning and functional comparison of GH5 family DNA fragments of C5-CEL4
Primer 5.0 was employed to design concatenated primers: c5-cel4-gh5-F2 (CATCATCATCATCATCATGAAATGATATTCGGCGCAGGTGC) and c5-cel4-gh5-R2 (GTGCTCGAGTGCGGCCGCAAGTCAGCTGGAGCTCGAAGAGGAA) to amplify the c5-cel4-gh5 gene. The c5-cel4-gh5 gene was cloned, expressed, and purified analogously to c5-cel4. Monoclonal strains of C5-CEL4 and C5-CEL4-GH5 were cultured in a mixed liquid medium comprising 6 mL of basal salts and 3 mL of LB broth with 100 µg/mL kanamycin. Each strain was inoculated into three tubes as replicates and incubated at 37℃ with shaking at 180 RPM for 36 h. Post-incubation, cultures were chilled at 4℃, centrifuged at 12,000 RPM for 20 min, and the supernate was harvested as the extracellular crude enzyme solution. Escherichia coli biomass was collected, resuspended in 5 mL of Phosphate Buffered Saline (pH 7.6), disrupted by ultrasonic fragmentation in an ice-water bath, and centrifuged at 4℃, 12,000 RPM for 20 min to obtain the intracellular crude enzyme solution. The enzymatic activities of C5-CEL4 and C5-CEL4-GH5 in both intracellular and extracellular fractions were compared.
Enzyme activity assay
Activity against CMC was determined by measuring the release of reducing sugar, with 1% (w/v) CMC as substrate, by the 3,5-dinitrosalicylic acid (DNS) assay. One unit (U) of CMCase activity was defined as the amount of enzyme to release 1µmol glucose-equivalent reducing sugars per minute17. The assay involved adding 10 µL of purified enzyme to 90 µL of an optimal pH buffer containing 1% CMC-Na, incubating at the optimal temperature for 30 min, and terminating the reaction with 150 µL of DNS. The mixture was heated at 90 °C for 10 min for color development, cooled, and 150 µL was transferred to a 96-well plate for absorbance measurement at 540 nm using a microplate reader. Each experiment was conducted in triplicate with one control.
Biochemical characterization
The optimal pH of purified C5-CEL4 was determined across a pH range of 3.0–10.0 using citrate-disodium phosphate buffer for pH 3.0–8.0 and glycine-sodium hydroxide buffer for pH 8.0–10.0. The optimal temperature was identified by measuring enzyme activity at temperatures from 20 to 75 °C under the optimal pH conditions. Thermal and pH stability were assessed by measuring residual enzyme activity after incubation at various temperatures (40, 45, 50, 55, and 60 °C) for time intervals (0, 20, 40, 60, 80, 100, and 120 min), and across a pH gradient (3.0–13.0) for exposure durations of 4 °C, 12 h, and 24 h.
Influence of salts, ionic liquids, metal ions, and chemical reagents on C5-CEL4
The optimal concentration of NaCl (ranging from 0 to 5.0 M) was established at 50 °C and pH 7.0. The purified enzyme solution was incubated at 4 °C in varying salt concentrations (0–5.0 M) for nine months, after which the residual enzyme activity was assessed without a salt removal procedure. The surface electrostatic potential of C5-CEL4 was predicted using AlphaFold Protein Structure Database and PyMOL 2.6 software. The influence of different ionic liquids (1-butyl-3-methylimidazole tetrachloroborate BMIM-BF4, 1-butyl-3-methylimidazole acetate BMIM-Ac, and ethyl-3-methylimidazolium chloride EMIM-Cl) in varying concentrations (1%, 2.5%, 5%, 10%, 15%, 20%, 25%, 30%, 35%, 40%, 45%, and 50% w/v) on enzyme activity was investigated at 50 °C and pH 7.0. The purified enzyme solution was incubated at 4 °C for 24 h in the aforementioned ionic liquid solutions (1%, 5%, 10%, 20%, 30%, 40%, 45%, and 50% w/v) at pH 7.0, and the residual enzyme activity was measured.
The effects of metal ions, chemical reagents, and ionic liquids on C5-CEL4 activity were studied using 1 mM and 10 mM concentrations of various metal ions (Na+, K+, Mg2+, Fe3+, Ca2+, Zn2+, Co2+, Cu2+, Ag+, Mn2+, Pb2+, and Ni2+), and 0.1% and 1% concentrations of different chemical reagents (EDTA, PMSF, DTT, Tween 80, SDS, CTAB—corresponding to ethylenediaminetetraacetic acid, phenylmethylsulfonyl fluoride, dithiothreitol, polysorbate-80, sodium dodecyl sulfate, and cetyltrimethylammonium bromide, respectively). Additionally, 1% and 10% concentrations of various alcohols (methanol, ethanol, isopropanol, and β-mercaptoethanol) were incorporated into the enzyme solution reaction system. Enzyme activity was compared to the control (untreated enzyme solution, set as 100%).
Substrate specificity and kinetic constants of C5-CEL4
Experiments were conducted under controlled conditions identical to those described above, excluding any additives in the reaction mixture. Substrate specificity of C5-CEL4 intracellular and extracellular enzymes was examined using CMC-Na, bagasse xylan, beechwood xylan, corn cob xylan, Avicel, and cellobiose (each at 1% w/v). Kinetic constants of C5-CEL4 intracellular and extracellular enzymes were determined using varying concentrations of CMC-Na and bagasse xylan (2–20 mg/mL), incubated for 5/10 minutes at 50 °C and pH 7.0. The Michaelis-Menten constant (Km) and the maximum reaction velocity (Vmax) were calculated.
TLC analysis
A reaction mixture containing 1% CMC-Na and 10 µg of purified enzyme was incubated under optimal pH and temperature conditions for 2 h. Hydrolysis products were analyzed using thin-layer chromatography (TLC) on silica gel 60 plates (Merck, Darmstadt, Germany). The solvent system used was 1-butanol/acetic acid/water (2:1:1 v/v/v). Sugars were detected post-treatment at 120 °C for 10 min with a freshly prepared 5% (v/v) H2SO4 ethanol spray. Sugar standards included glucose (G1), cellulosic disaccharide (G2), cellulosic trisaccharide (G3), and cellulosic tetrasaccharide (G4).
Utilization of C5-CEL4 in the enzymatic degradation of agricultural straw
The raw substrates of wheat bran and corn stalks obtained from the market were washed with tap water to eliminate impurities and then air-dried. The dried substrates were ground and sieved. A 5 g portion of the substrate was mixed with 100 mL of distilled water and subjected to a water bath at 90 °C for 4 h. Post-treatment, the substrate was rinsed with distilled water until the rinse water was clear. The substrate was then dried in an oven at 50 °C for subsequent use. The dried substrate was divided into four 0.2 g portions, placed into 10 mL tubes, and combined with 4 mL of a pH 7.0 citric acid-disodium hydrogen phosphate buffer. The mixture was stirred thoroughly and left to soak overnight. The supernate was centrifuged at 12,000 RPM for 15 min, and the supernate was discarded. To three experimental groups, 0.05 mg/mL of C5-CEL4 was added to the substrate, while the control group received no enzyme solution. All groups were adjusted to a final volume of 5 mL with the same buffer. The mixtures were shaken well and incubated at the optimal temperature, with reducing sugar yield monitored every 2 h.
Statistical analysis
Experiments were performed in triplicate unless otherwise noted, and the results were expressed as mean values. Statistical analyses were conducted using SPSS 20.0, with results presented as mean ± SEM. One-way ANOVA followed by Tukey’s test was used to compare multiple groups (0.01 < p-value < 0.05).
Results
Metagenomic sequencing and sequence analysis of C5-CEL4
Metagenomic sequencing of DNA extracted from the sediment of Ebinur Salt Lake yielded a comprehensive dataset comprising 4.2 Gbp and including 32,394 contigs longer than 500 bp. This dataset enabled the identification of 135,361 unique genes, among which 15 were predicted to be cellulase genes. Based on gene integrity and sequence novelty, a new cellulase gene, named c5-cel4, was identified. The gene was sequenced, and the absence of a stop codon within the ORF (Open Reading Frame) was confirmed. The c5-cel4 gene is 1,347 bp long, encoding 448 amino acids, and lacks a signal peptide, indicating cytoplasmic localization. The deduced protein, without a signal peptide, comprises 448 amino acids with a calculated molecular weight of 47.54 KDa and a theoretical pI of 4.51.
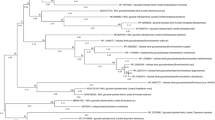
Comparative analysis with NCBI data showed a 90.97% similarity between C5-CEL4 and GH5 family cellulases from Microbulbifer litoralis, based on the expressed amino acid sequences. Phylogenetic analysis indicated that C5-CEL4 clusters with a cellulase from Microbulbifer halophilus (GenBank: WP 265722981.1) with 85% support, identifying it within the glycosyl hydrolase family (Supplementary Figure S1). This protein shares 70% similarity with GH5 family endoglucanases from Cellvibrio japonicus (PDB: 8BQA, 8BQC). Multiple sequence alignments demonstrated high homology of C5-CEL4 with GH5 family cellulases 8BQA, 8BQC, 8C10, 4M1R, and endoglucanase 1EGZ (Supplementary Figure S2). Structural analysis confirmed that C5-CEL4 contains a catalytic domain typical of GH5 cellulases (Fig. 1a). Homology modeling via SWISS-MODEL yielded a GMQE value of 0.94 with GH5 endoglucanases from Microbulbifer rhizosphaerae. The protein exhibits a conserved (β/α)8 TIM barrel fold with eight loops around the active site, contributing to its catalytic versatility, substrate specificity, thermal stability, and pH stability (Fig. 1b).
Protein structure analysis of C5-CEL4. (a), structural domain prediction of the protein sequence of C5-CEL4 using InterPro. The structure describes a cellulase catalytic structural domain (CD). (b), GH5 family cellulase C5-CEL4, homology modeled on the SWISS-MODEL server (https://swissmodel.expasy.org/interactive) using a Microbulbifer rhizosphaerae-derived GH5 family endoglucanase as a template (UniProt accession no: A0A7W4WCK3).
Heterologous expression, and purification of C5-CEL4
The c5-cel4 was cloned into the pSHY211 C-His vector (with 6 His tags) and confirmed by sequencing. Recombinant C5-CEL4 was secreted extracellularly, with optimal fermentation conditions at 37 °C for 36 h, yielding an enzyme specific activity of 0.886 U/mL(Fig. 3a). Both intracellular and extracellular crude enzyme solutions were purified using a Ni-NTA affinity chromatography column, and SDS-PAGE analysis showed a ~ 33 KDa band for both, which differed from the theoretical size (47.54 KDa) by about 14 KDa. (Fig. 2, The original figure is in Supplementary Figure S3.). Sequence analysis indicated the presence of a His tag at the N-terminus but not at the C-terminus, which lacked a stop codon. Consequently, the C-terminus was selected for protein sequencing analysis.
SDS-PAGE analysis and enzyme activity profiling of recombinant C5-CEL4 intracellular and extracellular purified enzyme. (a), extracellular purified enzyme. (b), intracellular purified enzyme. Lane 1, protein molecular weight marker. Lane 2, E. coli DH5α/pSHY211-c5-cel4 lysate. Lane 3, purified C5-CEL4. Lane 4, zymogram of purified enzyme.
Monitoring of extracellular crude enzyme activity of C5-CEL4 heterologous expression in Escherichia coli (a), and comparison of intracellular and extracellular cellulase activities of C5-CEL4 and C5-CEL4-GH5 (b). The error line represents ± SEM (n = 3). Different letters (a, b, c, d) indicate significant differences between groups (p < 0.05, it was derived through multiple comparative analyses conducted using the Duncan method in SPSS.). The same letters indicate no significant differences between groups.
The LC-MS/MS-based C-terminal sequencing18 of the C5-CEL4 protein revealed its terminal sequence as IVRGWDGGGGSSSSSSSSSS, ending at amino acid 314 (Supplementary Figure S4). Consequently, the theoretical molecular weight and isoelectric point of the expressed protein were 34.12 KDa and 4.69, respectively. This premature termination led to a truncated protein, with its actual molecular mass matching the theoretical value (34.12 KDa), confirming the successful expression and purification. Zymography detected cellulase activity, displayed as a clear band on a red background (Fig. 2). Due to the premature termination, the C-terminal region (residues 315 to 448) containing the CBM domain was absent. Primers were designed to amplify the catalytic domain (C5-CEL4-GH5), to obtain the recombinant target protein C5-CEL4-GH5 (Supplementary Figure S5). Activity assays revealed similar intracellular activities for C5-CEL4 and C5-CEL4-GH5, though their extracellular activities differed significantly. The extracellular activity of C5-CEL4-GH5 was notably low, likely due to the absence of the sequence necessary for secretion in Escherichia coli(Fig. 3b).
The influence of temperature and pH on C5-CEL4 activity
The optimal temperature for both intracellular and extracellular pure enzyme activities is 50 °C, maintaining above 80% activity between 35 °C and 60 °C (Fig. 4a). The optimal pH for both activities is 7.0, with over 70% of maximal activity preserved within a pH range of 6.0 to 8.0 (Fig. 4b). Thermal stability analysis indicates that C5-CEL4 retains 100% activity after 2 h at 40 °C and 74% after the same period at 50 °C, with a half-life of 70 min at 55 °C and 60 °C (Fig. 4c). pH stability analysis shows the enzyme maintains over 50% of its initial activity across a pH range of 3.0 to 13.0. Following incubation at 4 °C for 12 and 24 h, the enzyme’s activity remains above 80% within the pH range of 4.0 to 12.0 (Fig. 4d).
Effect of temperature and pH on the activity and stability of recombinant cellulase C5-CEL4. (a), effect of temperature on the activity of C5-CEL4. (b), effect of pH on the activity of C5-CEL4. (c), effect of temperature on the stability of intracellular enzyme C5-CEL4. (d), effect of pH on the stability of intracellular enzyme C5-CEL4. Initial activity was 100%, and each value in the graphs represents the mean ± SEM (n = 3),100% of extracellular enzyme = 24.36 ±0.23 U/mg and 100% of intracellular enzyme = 19.6 ± 3.47 U/mg.
Impact of salts and ionic liquids on C5-CEL4
As depicted in Fig. 5a, the optimal NaCl concentration for C5-CEL4 enzymatic activity ranged from 2.5 to 3.0 M, with activity levels exceeding 100% even at a near-saturated concentration of 5.0 M. The enzyme maintained its activity across a broad NaCl concentration spectrum from 0.5 to 5.0 M. Salinity tolerance analysis revealed that C5-CEL4 preserved approximately 100% or higher activity at NaCl concentrations between 0.5 and 5.0 M after prolonged exposure of 9 months at 4 °C (Fig. 5b). Furthermore, the enzyme exhibited near 100% activity when 1 mM or 10 mM of NaCl, NaNO3, or Na2SO3 were incorporated into the reaction mixture, suggesting that these sodium salts did not significantly impact enzymatic function (Supplementary Table S1). As shown in Fig. 6a, b, the higher prevalence of acidic amino acids on the enzyme’s surface contributes to a negative overall electrostatic potential, which enhances hydrophilicity and reduces hydrophobicity. In contrast, the GH5-family cellulase GH5_419 (PDB code: 6XSU) from Ruminococcus flavefaciens has a lower amount of acidic amino acids on its surface (Fig. 6c, d). The GH5-family cellulase is structurally similar to our enzyme. This modification improves the enzyme’s water-binding capacity and prevents aggregation in high-salt environments20. C5-CEL4’s activity increased to over 110% in the presence of 1%-10% EMIM-Cl ionic liquids (Fig. 5c) and maintained over 60% relative activity at 20% ionic liquid concentration. Additionally, the enzyme retained about 90% or greater activity and was marginally activated following 24 h of incubation in 40% ionic liquids (Fig. 5d). These results confirm that C5-CEL4 is resilient to both salts and ionic liquids.
Impact of NaCl and ionic liquids on C5-CEL4 activity. (a), influence of NaCl on the catalytic performance of intracellular and extracellular C5-CEL4 enzymes. (b), stability assessment of intracellular C5-CEL4 enzyme under varying NaCl concentrations after a 9-month incubation period. (c), effect of ionic liquid solutions on the enzymatic activity of intracellular C5-CEL4. (d), stability evaluation of intracellular C5-CEL4 enzyme in ionic liquid solutions following 24 h of incubation at 4 °C. The initial enzymatic activity was standardized to 100%, with each graphical data point representing the mean ± SEM, where 100% corresponds to 19.6 ± 3.47 U/mg.
Predicted surface electrostatic potentials of C5-CEL4 and GH5-family cellulase. (a) Surface electrostatic potential of C5-CEL4. (b) Surface electrostatic potential obtained by flipping figure a by 180° in Pymol 2.6. (c) Surface electrostatic potential of TfCel5A. (d) Surface electrostatic potential obtained by flipping figure c by 180° in Pymol 2.6. Negative and positive electrostatic potentials are shown in red and blue, respectively. Note: Fig. 6a was plotted in Pymol 2.6 from the structure prediction map of Fig. 1b. Figure 6c was plotted in Pymol 2.6 based on the structure of GH5_4 downloaded from the Protein Data Bank(https://www.rcsb.org/). GH5_4 (PDB code: 6XSU) is a GH5 family cellulase from Ruminococcus flavefaciens.
Influence of metal ions and chemical reagents on C5-CEL4
As presented in Supplementary Table S1, C5-CEL4 activity was significantly enhanced by 1 mM Co2+ (114.4 ± 1.1%), 1 mM Mn2+ (135.6 ± 8.7%), and 10 mM Mn2+ (212.4 ± 0.4%). Conversely, the enzyme was moderately inhibited by 1 mM and 10 mM concentrations of K+, Mg2+, Fe3+, Ca2+, Zn2+, Cu2+, Ag+, and Ni2+, and exhibited about a 9% reduction in activity with 10 mM Pb2+. Specifically, Co2+ increased activity by approximately 14%, whereas Mn2+ significantly boosted activity by roughly 2.1-fold. The inhibitors EDTA, PMSF, and DTT at 0.1% concentration had negligible effects on C5-CEL4. However, EDTA and PMSF at 1.0% concentration mildly inhibited the enzyme. At 0.1%, Tween 80 enhanced enzymatic activity by about 37% and showed slight activation at concentrations up to 1%. In contrast, SDS and CTAB at both 0.1% and 1.0% concentrations slightly inhibited enzyme activity. Methanol, ethanol, and isopropanol at 1% and 10% concentrations had no significant effect on enzyme activity. The enzyme retained 76% activity in 1% β-ME but was completely inactivated at 10% concentration.
Substrate specificity and kinetic constants of C5-CEL4
The substrate specificity and kinetic constants of C5-CEL4 were detailed in Supplementary Table S2 and Table 1. Both extracellular and intracellular C5-CEL4 demonstrated activities for CMC-Na (24.36 ± 0.23 U/mg and 19.6 ± 3.47 U/mg), bagasse xylan (13.24 ± 0.61 U/mg and 2.5 ± 0.77 U/mg), and beechwood xylan (14.06 ± 0.53 U/mg and 1.48 ± 2.01 U/mg), respectively, while showing no activity for corn cob xylan, Avicel, and cellobiose. The Vmax, Km, Kcat, and Kcat/Km values for extracellular C5-CEL4 with CMC-Na were 158.73 µmol/min/mg, 45.03 mg/mL, 98.20 S− 1, and 2.18, respectively. For intracellular C5-CEL4 with CMC-Na, these values were 357.14 µmol/min/mg, 101.71 mg/mL, 220.95 S− 1, and 2.17, respectively. The kinetic parameters Vmax, Km, Kcat, and Kcat/Km for extracellular C5-CEL4 with bagasse xylan were 15.55 µmol/min/mg, 16.05 mg/mL, 12.32 S− 1, and 0.77, respectively. However, these parameters were not determined for intracellular C5-CEL4 with bagasse xylan.
Thin-layer chromatography (TLC) analysis of CMC-Na hydrolyzed by C5-CEL4
The hydrolysis products of CMC-Na by C5-CEL4 were analyzed using TLC, as illustrated in Fig. 7. Results indicated that C5-CEL4 cleaved cellulose oligosaccharide chains randomly, producing cellulotriose (G3) and other fiber oligosaccharides.
TLC analysis of CMC-Na hydrolyzed by C5-CEL4. Lane 1, standards: glucose (G1), cellobiose (G2), cellotriose (G3), and cellotetraose (G4). Lane 2, CMC-Na without enzyme solution. Lane 3, hydrolysis of CMC-Na by purified C5-CEL4.
Analysis of sugar production from agricultural straw hydrolyzed by C5-CEL4
As depicted in Fig. 8, C5-CEL4 hydrolyzed wheat bran and cornstalk crude substrates. The hydrolysis of wheat bran was more effective, with reducing sugar concentration increasing to approximately 1.2 µmol/mL at 2 h and stabilizing at 3 µmol/mL after 10 h. In contrast, the hydrolysis of cornstalks was less efficient, with a maximum sugar concentration of about 0.2 µmol/mL at 4 h before ceasing entirely. Hydrolysis of wheat bran and cornstalk by C5-CEL4 showed its ability to degrade both cellulose and hemicellulose polymers. It means that C5-CEL4 may be an excellent candidate for hydrolysis of agricultural residues.
Comparative analysis of sugar yield from C5-CEL4 hydrolyzed wheat bran (a) and corn stalks (b). Data points on the graph represent the mean values ± SEM.
Discussion
Endoglucanases from the GH5 family, integral to hybrid enzymes for biomass conversion, necessitate considerable thermal stability21. Current research identifies optimal reaction temperatures for most GH5 enzymes at 50 °C and above (Table 2). Notably, C5-CEL4 exhibits robust enzymatic activity across a broad temperature spectrum, with an optimal reaction temperature of 50 °C, indicating its potential as a thermostable cellulase. This higher thermal stability is crucial for industrial, animal husbandry, and biomass conversion applications. Enhancing the economic feasibility of industrial processes hinges on the optimal temperature and thermal stability of potent endoglucanases22. Our findings reveal that, unlike most GH5 endoglucanases which exhibit low alkaline tolerance and pH stability, C5-CEL4 maintains high activity at a higher optimal pH range of 5.0–9.6 (Table 2). The pH stability of C5-CEL4 is on par with the salt-tolerant AgCMCase from Aspergillus glaucus CCHA, both retaining over 40% residual activity from pH 4.0 to 10.0. Consequently, C5-CEL4 is an excellent candidate for industries in alkaline conditions, such as textiles, paper, and detergents. In summary, the moderate thermal stability and extensive pH tolerance of C5-CEL4 present significant potential in biomass conversion applications.
C5-CEL4 also exhibits superior salt tolerance relative to other GH5 family endoglucanases (Table 2). Structural analyses suggest an abundance of acidic amino acids on the protein surface, imparting a negative charge that enhances solubility in high-salt solutions by forming hydrated ion networks with cations and preventing protein aggregation through surface electrostatic repulsion. The salt tolerance mechanisms of most of the salt-tolerant cellulases reported so far are associated with a higher distribution of negative charges on their protein surfaces(Table S3)27,28,29,30. Lignocellulosic biomass in saline soils is challenging for microbial degradation due to high salt concentrations. However, salt-tolerant cellulases maintain high activity and stable catalytic performance in such environments31. The application of salt-tolerant cellulases can facilitate the degradation of saline lignocellulosic biomass. Acid or alkali pretreatment of lignocellulosic biomass generates significant salt during pH adjustment, impairing the efficiency of salt-intolerant cellulases32.
Therefore, salt-tolerant cellulases are preferable for degrading pretreated lignocellulosic biomass, offering greater application potential than conventional cellulases. The high salt tolerance of C5-CEL4 suggests its utility in the feed, textile, and biofuel industries. It has been postulated that common structural motifs within the GH5 family contribute to the relative resistance of these enzymes to ionic liquids33. C5-CEL4 retained over 90% activity after 24 h in 50% BMIM-Ac, EMIM-Cl, and BMIM-BF4. In contrast, GH5 family Cel5A activity dropped to 50% after 5 h in 2.5 M BMIM-Cl at room temperature34. Cellulase from Trichoderma reesei showed a decline to 31% activity after 24 h in 30% EMIM-Ac35. Additionally, Aspergillus fumigatus endoglucanase retained only about 80% activity after 12 h in 10% EMIM-Ac36. Compared to these studies, C5-CEL4 demonstrates exceptional ionic liquid tolerance, underscoring its potential for biomass conversion applications.
Some metal ions protect enzymes from thermal denaturation and can be relied upon to promote their activity and stability at higher temperatures37. Therefore, it is clear that Mn2+ and Co2+ protected C5-CEL4 from thermal denaturation during the reaction at 50 °C, instead promoting its catalytic activity. It has been shown that Mn2+ has a protective effect against thermal inactivation of the enzyme isolated from Escherichia coli38,39. Previous studies have demonstrated that Co2+ and Mn2+ augment the enzymatic activity of various enzymes, including GH5 family endoglucanases such as EG5C40TrepCel3/TrepCel441, CEL-5A42Cel776Sc43and CelRH544. This highlights the activating influence of Co2+ and Mn2+ on most GH5 family endoglucanases. Conversely, 0.1% EDTA, PMSF, and DTT did not impact C5-CEL4 enzyme activity, whereas SDS exhibited a slight inhibitory effect. C5-CEL4 was marginally inactivated by 1% EDTA, PMSF, and SDS, but showed slight activation with 1% DTT. Enzyme activity increased by 1.3-fold with 0.1% Tween 80, known to enhance cellulase stability45. The enzyme’s activity remained unaffected by methanol, ethanol, and isopropanol, with moderate ethanol concentrations shown to promote cellulase activity46. C5-CEL4’s performance in the presence of divalent cations, inhibitors, and alcohols common in industrial detergents indicates its robust potential for industrial applications.
The extracellular cellulase activity of C5-CEL4 secreted by Escherichia coli was measured at 0.886 U/mL (Fig. 3a). It has been reported that signal peptides enhance the secretion and expression of cellulase in Escherichia coli. When compared to the crude enzyme activity in the fermentation broth, which ranges from 0.81 to 1.33 U/mL, this activity is slightly lower47. Unlike some GH5 family endoglucanases, the C5-CEL4 extracellular enzyme exhibited a low Km and high Vmax, indicating a higher affinity for CMC-Na and requiring less substrate to reach Vmax (Table 1). The Vmax of C5-CEL4 was significantly higher than those of CelC307 (62.58 U/mg)48, Cel-5 M (27.1 U/mg)49, StCel5A (194 U/mg)50, and BaGH5-WT (6.0 U/mg)51, suggesting C5-CEL4’s superior catalytic capability. Additionally, Vmax is directly proportional to enzyme concentration, thus increasing enzyme concentration enhances reaction rates under sufficient substrate conditions. Furthermore, C5-CEL4 displayed activity toward CMC-Na and xylan from various sources, indicating its multifunctional nature. Multifunctional cellulases are vital for advancing the renewable bioeconomy by aiding in the conversion of agricultural residues into feed for ruminants. The synergistic action of multifunctional enzymes can pre-treat the complex structure of fibers, improving the utilization of agricultural residues by ruminants41. Consequently, C5-CEL4 may serve as an effective enzyme in feed additives.
Fibre trisaccharides (G3) are regarded as potent prebiotic oligosaccharides52. Numerous studies53,54,55have shown that oligosaccharides are crucial for promoting probiotic growth and maintaining gut flora stability. This implies the potential application of C5-CEL4 in prebiotic production.
Cellulases hydrolyze the β-1,4-glycosidic bonds of cellulose to form soluble oligosaccharides, cellobiose, and glucose. However, cross-linking of the xylan and cellulose regions can limit the attachment of cellulases, whereas the breakdown of the β-1,4-xyloside bonds by xylanases increases the accessibility of cellulose to hydrolyzing cellulases, thereby increasing the efficiency of the enzyme mixture in disrupting the cellulose structure56. Therefore, a mixture of cellulases and xylanases has been shown to be a very efficient method for achieving hydrolysis of lignocellulosic material57. However, it is costly if individual cellulase and xylanase blends are used for hydrolysis during application. If multifunctional enzymes are applied in biomass hydrolysis, the cost of applying multiple enzymes can be reduced and the efficiency of hydrolysis can be improved. The cellulase and xylanase activities of C5-CEL4 improved its efficiency in hydrolyzing wheat bran and cornstalks, and it has a large potential for agricultural applications. Wheat bran and cornstalks, both rich in cellulose, are common agricultural waste materials. Hydrothermal pretreatment of these substrates offers several benefits over acid-base pretreatment, including significantly reduced environmental impact, lower capital investment, the avoidance of chemical use, and diminished by-product formation58. This method enhances lignocellulose saccharification by disrupting the cell wall matrix, thereby improving cellulase accessibility to cellulose microfibrils59. The enzyme C5-CEL4 has demonstrated the ability to hydrolyze hot water-treated wheat bran and maize stalks to produce reducing sugars, indicating that hydrothermal pretreatment enhances cellulase hydrolysis efficiency60,61. These observations suggest that C5-CEL4 is promising for applications in biofuel production, biochemicals, the textile industry, the food industry, and environmental treatment11.
Conclusion
In this study, a novel cellulase gene, c5-cel4, was identified from the metagenome of substrates in Ebinur Salt Lake, Xinjiang, cloned, and expressed in Escherichia coli. Structural analysis revealed that the C5-CEL4 protein has a high surface concentration of acidic amino acids, resulting in a negative electrostatic potential. This feature enhances the enzyme’s surface hydrophilicity and water-binding capacity, thereby conferring salt resistance. C5-CEL4 exhibited hydrolytic activity against both CMC-Na and bagasse xylan, confirming its multifunctional nature and resistance to heat, alkali, salt, and ionic liquids. Additionally, the enzyme demonstrated tolerance to various metal ions, inhibitors, and alcohols, and was effective in hydrolyzing hot water-pretreated wheat bran and corn stalks to produce reducing sugars. These findings position C5-CEL4 as an ideal candidate for biomass conversion and industrial applications in washing and textiles, with significant potential in the feed, food, and bioenergy industries.
Data availability
Original contributions presented in the study are included in the article/Supplementary Materials. The nucleotide sequence of the C5-CEL4 gene was submitted to GenBank (https://www.ncbi.nlm.nih.gov/nuccore/PV014881). Further inquiries can be directed to the corresponding author(yinyirui@dali.edu.cn).
References
Yin, Y. R. et al. Characterization of an alkali-tolerant, thermostable, and multifunctional GH5 family endoglucanase from Thermoactinospora rubra YIM 77501T for prebiotic production. Biomass Convers. Biorefinery. 12 (8), 1–10. https://doi.org/10.1007/s13399-020-00848-1 (2020).
Yin, Y. R. et al. Expression and characterization of a cold-adapted, salt- and glucose-tolerant GH1 β-glucosidase obtained from Thermobifida halotolerans and its use in sugarcane Bagasse hydrolysis. Biomass Convers. Biorefinery. 11, 1-9. https://doi.org/10.1007/s13399-019-00556-5 (2020).
Ibrahim, A. M., Hamouda, R. A., El-Naggar, N. E. & Al-Shakankery, F. M. Bioprocess development for enhanced endoglucanase production by newly isolated bacteria, purification, characterization and in-vitro efficacy as anti-biofilm of Pseudomonas aeruginosa. Sci. Rep. 11 (1), 9754. https://doi.org/10.1038/S41598-021-87901-9 (2021).
Sangita, C., Rajkumar, A., Singh, P. T., Ashiq, M. & Sanjay, S. Extremophilic Fungal Cellulases: Screening, Purification, Catalysis, and Applications in Extremophilic Fungi (ed. Sahay). 347-365. https://doi.org/10.1007/978-981-16-4907-3_16 (Springer, 2022).
Patel, S. & Saraf, M. Perspectives and application of halophilic enzymes. In Sustainable Development and Biodiversity (eds. Maheshwari, D. & Saraf, M.). 403-419. https://doi.org/10.1007/978-3-319-14595-215 (Springer, 2015).
Zarafeta, D. et al. Discovery and characterization of a thermostable and highly halotolerant GH5 cellulase from an Icelandic hot spring isolate. PLoS One. 11 (1), e0146454. https://doi.org/10.1371/journal.pone.0146454 (2016).
Zhang, T., Supratim, D. & Jerry, E. Identification of a haloalkaliphilic and thermostable cellulase with improved ionic liquid tolerance. Green Chem. 13 (8), 2083–2090. https://doi.org/10.1039/C1GC15193B (2011).
Hebal, H. et al. Enhanced activity of hyperthermostable Pyrococcus Horikoshii endoglucanase in Superbase ionic liquids. Biotechnol. Lett. 44 (8), 961–974. https://doi.org/10.1007/s10529-022-03268-5 (2022).
Vasić, K., Knez, Ž. & Leitgeb, M. Bioethanol production by enzymatic hydrolysis from different lignocellulosic sources. Molecules 26 (3), 753. https://doi.org/10.3390/molecules26030753 (2021).
Sun, W., Li, X. Z., Zhao, J. & Qin, Y. Q. Pretreatment strategies to enhance enzymatic hydrolysis and cellulosic ethanol production for biorefinery of corn Stover. Int. J. Mol. Sci. 23 (21), 13163. https://doi.org/10.3390/ijms232113163 (2022).
Xu, C. Z., Tong, S. S., Sun, L. Q. & Gu, X. L. Cellulase immobilization to enhance enzymatic hydrolysis of lignocellulosic biomass: an all-inclusive review. Carbohydr. Polym. 321, 121319–121319. https://doi.org/10.1016/J.CARBPOL.2023.121319 (2023).
Garg, R. et al. Biochemical and structural characterization of a novel halotolerant cellulase from soil metagenome. Sci. Rep. 6 (1), 39634. https://doi.org/10.1038/srep39634 (2016).
Yin, Y. R. et al. Characterization of a metagenome-derived thermostable Xylanase from Tengchong hot spring. Biomass Convers. Biorefnery. 14 (9), 10027-10034. https://doi.org/10.1007/s13399-022-03296-1 (2022).
Handelsman, J., Liles, M., Mann, D., Riesenfeld, C. & Goodman, R. M. Cloning the metagenome: Culture-independent access to thediversity and functions of the uncultivated microbial world. Methods Microbiol. 33 (02), 241–255. https://doi.org/10.1016/S0580-9517 (2002).
Yin, Y. R. et al. Characterization of a GH10 extremely thermophilic Xylanase from the metagenome of hot spring for prebiotic production. Sci. Rep. 13 (1), 16053. https://doi.org/10.1038/s41598-023-42920-6 (2023).
Yin, Y. R. et al. The hybrid strategy of Thermoactinospora rubra YIM 77501T for utilizing cellulose as a carbon source at different temperatures. Front. Microbiol. 8, 942. https://doi.org/10.3389/fmicb.2017.00942 (2017).
Miller, G. L. Use of Dinitrosalicylic acid reagent for determination of reducing sugar. Anal. Chem. 31 (3), 426–428. https://doi.org/10.1021/ac60147a030 (2002).
Wilkins, J. A., Kaasik, K., Chalkley, R. J. & Burlingame, A. L. Characterization of prenylated C-terminal peptides using a thiopropyl-based capture technique and LC-MS/MS. Mol. Cell Proteom. 19(6), 1005–1016. https://doi.10.1074/mcp.RA120.001944 (2020).
Glasgow, E. M. et al. A structural and kinetic survey of GH5_4 endoglucanases reveals determinants of broad substrate specificity and opportunities for biomass hydrolysis. J. Biol. Chem. 295 (51), 17752–17769. https://doi.org/10.1074/jbc.RA120.015328 (2020).
Teo, S. C. et al. Characterizing a Halo-Tolerant GH10 Xylanase from roseithermus Sacchariphilus strain RA and its CBM-Truncated variant. Int. J. Mol. Sci. 20 (9), 2284–2284. https://doi.org/10.3390/ijms20092284 (2019).
Zheng, F. et al. Activity and thermostability of GH5 endoglucanase chimeras from mesophilic and thermophilic parents. Appl. Environ. Microbiol. 85 (5), e02079–e02018. https://doi.org/10.1128/AEM.02079-18 (2019).
Patel, A. & Shah, A. Purification and characterization of novel, thermostable and non-processive GH5 family endoglucanase from Fomitopsis meliae CFA 2. Int. J. Biol. Macromol. 182, 1161–1169. https://doi.org/10.1016/j.ijbiomac.2021.04.110 (2021).
Yang, C. et al. Discovery of new cellulases from the metagenome by a metagenomics-guided strategy. Biotechnol. Biofuels. 9, 138. https://doi.org/10.1186/s13068-016-0557-3 (2016).
Li, Z. Q. et al. The unique GH5 cellulase member in the extreme halotolerant fungus Aspergillus glaucus CCHA is an endoglucanase with multiple tolerance to salt, alkali and heat: prospects for straw degradation applications. Extremophiles 22 (4), 675–685. https://doi.org/10.1007/s00792-018-1028-5 (2018).
Khan, A. et al. Isolation and characterization of an acidic, Salt-Tolerant endoglucanase Cel5A from a bacterial strain Martelella endophytica YC6887 genome. Mol. Biotechnol. 63 (4), 305-315. https://doi.org/10.1007/s12033-020-00295-3 (2021).
Cheng, J. B. et al. Isolation and characterization of a non-specific endoglucanase from a metagenomic library of goat rumen. World J. Microbiol. Biotechnol. 32 (1), 12. https://doi.org/10.1007/s11274-015-1957-4 (2016).
Kern, M. et al. Structural characterization of a unique marine animal family 7 cellobiohydrolase suggests a mechanism of cellulase salt tolerance. Proc. Natl. Acad. Sci. U S A. 110 (25), 10189–10194. https://doi.org/10.1073/pnas.1301502110 (2013).
Leadbeater, D. R. & Bruce, N. C. Functional characterisation of a new halotolerant seawater active glycoside hydrolase family 6 cellobiohydrolase from a salt marsh. Sci. Rep. 14(1), 3205–3205. https://doi.10.1038/S41598-024-53886-4 (2024).
Zhang, H. D. et al. Structural insight of a trimodular halophilic cellulase with a family 46 carbohydrate-binding module. Public Lib. Sci. One 10(11), e0142107. https://doi.10.1371/journal.pone.0142107 (2015).
Cai, L. N., Xu, S. N., Lu, T., Lin, D. Q. & Yao, S. J. Salt-tolerant and thermostable mechanisms of an endoglucanase from marine Aspergillus niger. Bioresour. Bioprocess. 9(1), 44. https://doi.10.1186/s40643-022-00533-3 (2022).
Cai, L. N., Xu, S., Lu, T., Lin, D. Q. & Yao, S. J. Salt-tolerant mechanism of marine Aspergillus Niger cellulase cocktail and improvement of its activity. Chin. J. Chem. Eng. 28 (4), 1120-1128. https://doi.org/10.1016/j.cjche.2019.11.012 (2022).
Ariaeenejad, S. et al. A novel high performance in-silico screened metagenome-derived alkali-thermostable endo-β–1,4-glucanase for lignocellulosic biomass hydrolysis in the harsh conditions. BMC Biotechnol. 20 (1), 56. https://doi.org/10.1186/s12896-020-00647-6 (2020).
Pottkämper, J. et al. Applying metagenom Ics for the identification of bacterial cellulases that are stable in ionic liquids. Green. Chem. 11, 957–965. https://doi.org/10.1039/b820157a (2009).
Liang, C. N. et al. Cloning and characterization of a thermostable and halo-tolerant endoglucanase from Thermoanaerobacter tengcongensis MB4. Applied. Microbiol. Biotechnol. 89 (2), 315–326. https://doi.org/10.1007/s00253-010-2842-6 (2011).
Wang, Y., Mark, R., Douglas, H. & Labbé, N. Compatible ionic liquid-cellulases system for hydrolysis of lignocellulosic biomass. Biotechnol. Bioeng. 108 (5), 1042–1048. https://doi.org/10.1002/bit.23045 (2011).
Singer, S. W. et al. Enrichment, isolation and characterization of fungi tolerant to 1-ethyl-3-methylimidazolium acetate. J. Appl. Microbiol. 110 (4), 1023–1031. https://doi.org/10.1111/j.1365-2672.2011.04959.x (2011).
Cruz, E., Moraes, L. P. D., Costa, E. A., Martins, M. L. L. & Barbosa, J.B. Optimization of food-waste based culture medium for cellulase production by Thermophilic bacillus sp SMIA–2 and effect of divalent metal ions on activity and stability of the enzyme at higher temperatures. Int. J. Adv. Eng. Res. Sci. 6(7), 331–337 (2019). https://doi.org/10.22161/ijaers.6741
Adalberto, P. R. et al. Effect of divalent metal ions on the activity and stability of β-galactosidase isolated from Kluyveromyces lactis. Revista De Ciências Farmacêuticas Básica E Aplicada. 31 (3), 143–150 (2010). https://rcfba.fcfar.unesp.br/index.php/ojs/about
Vasconcellos, V. M., Tardioli, P. W., Giordano, R. L. C. & Farinas, C. S. Addition of metal ions to a (hemi)cellulolytic enzymatic cocktail produced in-house improves its activity, thermostability, and efficiency in the saccharification of pretreated sugarcane Bagasse. New Biotechnol. 33 (3), 331–337. https://doi.org/10.1016/j.nbt.2015.12.002 (2016).
Dhar, H., Kasana, R. C., Dutt, S. & Gulati, A. Cloning and expression of low temperature active endoglucanase EG5C from Paenibacillus sp. IHB B 3084. Int. J. Biol. Macromol. 81, 259–266. https://doi.org/10.1016/j.ijbiomac.2015.07.060 (2015).
Meng, Z. X., Yang, C. J., Leng, J., Zhu, W. Y. & Cheng, Y. F. Production, purification, characterization and application of two novel endoglucanases from Buffalo rumen metagenome. J. Anim. Sci. Biotechnol. 14 (1), 16. https://doi.org/10.1186/S40104-022-00814-z (2023).
Ma, L. L. et al. Cloning and characterization of low-temperature adapted GH5-CBM3 endo-cellulase from Bacillus subtilis 1AJ3 and their application in the saccharification of Switchgrass and coffee grounds. AMB Express. 10 (1), 42. https://doi.org/10.1186/s13568-020-00975-y (2020).
Escuder-Rodríguez, J. J. et al. Characterization of a novel thermophilic metagenomic GH5 endoglucanase heterologously expressed in Escherichia coli and Saccharomyces cerevisiae. Biotechnol. Biofuels Bioprod. 15 (1), 76. https://doi.org/10.1186/S13068-022-02172-4 (2022).
Wierzbicka-Woś, A. et al. Biochemical characterization of a novel monospecific Endo-β–1,4-Glucanase belonging to GH family 5 from a rhizosphere metagenomic library. Front. Microbiol. 10, 1342. https://doi.org/10.3389/fmicb.2019.01342 (2019).
Okino, S., Ikeo, M., Ueno, Y. & Taneda, D. Effects of tween 80 on cellulase stability under agitated conditions. Bioresour Technol. 142, 535–539. https://doi.org/10.1016/j.biortech.2013.05.078 (2013).
Liang, C. Y. et al. Synergistic enhancement effect of compound additive of organic alcohols and biosurfactant on enzymatic hydrolysis of lignocellulose. Fermentation 8 (12), 725–725. https://doi.org/10.3390/FERMENTATION8120725 (2022).
Cui, Y. et al. Efficient secretory expression of Recombinant proteins in Escherichia coli with a novel actinomycete signal peptide. Protein Expr Purif. 129, 69–74. https://doi.org/10.1016/j.pep.2016.09.011 (2017).
Mohammadi, S. et al. Expression, characterization, and activity optimization of a novel cellulase from the thermophilic bacteria Cohnella sp. A01. Sci. Rep. 12 (1), 10301. https://doi.org/10.1038/S41598-022-14651-7 (2022).
Patel, M., Patel, H. M. & Dave, S. Determination of bioethanol production potential from lignocellulosic biomass using novel Cel–5m isolated from cow rumen metagenome. Int. J. Biol. Macromol. 153, 1099–1106. https://doi.org/10.1016/j.ijbiomac.2019.10.240 (2020).
Tambor, J. H. et al. Recombinant expression, activity screening and functional characterization identifies three novel endo–1,4-β-glucanases that efficiently hydrolyse cellulosic substrates. Appl. Microbiol. Biotechnol. 93 (1), 203–214. https://doi.org/10.1007/s00253-011-3419-8 (2012).
Singh, S., Kumar, K., Nath, P. & Goyal, A. Role of Glycine 256 residue in improving the catalytic efficiency of mutant endoglucanase of family 5 glycoside hydrolase from Bacillus amyloliquefaciens SS35. Biotechnol. Bioeng. 117 (9), 2668–2682. https://doi.org/10.1002/bit.27448 (2020).
Ubiparip, Z., Moreno, D. S., Beerens, K. & Desmet, T. Engineering of cellobiose phosphorylase for the defined synthesis of cellotriose. Appl. Microbiol. Biotechnol. 104 (19), 8327–8337. https://doi.org/10.1007/s00253-020-10820-8 (2020).
Kyu, M. T., Dar, B., Aye, S. S. & Matsuda, T. Prebiotic oligosaccharides prepared by enzymatic degradation of dietary fibers in rice grains. J. Nutr. Sci. Vitaminol. 65, S143–S147. https://doi.org/10.3177/jnsv.65.S143 (2019).
Saini, R. et al. Recent advancements in prebiotic oligomers synthesis via enzymatic hydrolysis of lignocellulosic biomass. Bioengineered 13 (2), 2139–2172. https://doi.org/10.1080/21655979.2021.2023801 (2022).
Karnaouri, A., Matsakas, L., Krikigianni, E., Rova, U. & Christakopoulos, P. Valorization of waste forest biomass toward the production of cello-oligosaccharides with potential prebiotic activity by utilizing customized enzyme cocktails. Biotechnol. Biofuels. 12, 285. https://doi.org/10.1186/s13068-019-1628-z (2019).
Verma, D. & Satyanarayana, T. Cloning, expression and applicability of thermo-alkali-stable Xylanase of Geobacillus thermoleovorans in generating xylooligosaccharides from agro-residues. Bioresour Technol. 107, 333–338. https://doi.org/10.1016/j.biortech.2011.12.055 (2012).
Meng, Z. X. et al. Characterization of a novel bifunctional enzyme from Buffalo rumen metagenome and its effect on in vitro ruminal fermentation and microbial community composition. Anim. Nutr. 13, 137–149. https://doi.org/10.1016/j.aninu.2023.01.004 (2023).
Yang, B. & Wyman, C. E. Pretreatment: the key to unlocking low-cost cellulosic ethanol. Biofuel Bioprod. Bior. 2, 26–40. https://doi.org/10.1002/bbb.49 (2008).
Kumar, P., Barrett, D. M. & Delwiche, M. J. Methods for pretreatment of lignocellulosic biomass for efficient hydrolysis and biofuel production. Ind. Eng. Chem. Res. 48, 3713–3729. https://doi.org/10.1021/ie801542g (2009).
Merali, Z. et al. Characterization of cell wall components of wheat Bran following hydrothermal pretreatment and fractionation. Biotechnol. Biofuels. 8, 23. https://doi.org/10.1186/s13068-015-0207-1 (2015).
Lu, X. Q., Zheng, X. J., Li, X. Z. & Zhao, J. Adsorption and mechanism of cellulase enzymes onto lignin isolated from corn Stover pretreated with liquid hot water. Biotechnol. Biofuels. 9, 118. https://doi.org/10.1186/s13068-016-0531-0 (2016).
Acknowledgements
We extend our gratitude to Dr. Yonghong Liu (Xinjiang Institute of Ecology and Geography, Chinese Academy of Sciences) and Professor Wenjun Li (Sun Yat-sen University) for their invaluable assistance in sample collection from Ebinur Salt Lake. Additionally, I would like to express my appreciation to the molecular ecology team at Dali University, including all faculty and students, for their support and contributions to this study.
Funding
This research was supported by the Yunnan Applied Basic Research Project (No. 202101AU070138 and 202401AT071166), Science and Technology Projects of Xizang Autonomous Region, China (XZ202501ZY0019), and Xingdian Talent Support Program of Yunnan Province (No. 230212528080).
Author information
Authors and Affiliations
Contributions
L.J.W., L.Q.Y., and Y.R.Y. conceived the study. L.J.W. constructed the constitutive expression plasmid. M.H. and C.Q.H. cloned the gene and cultured strains. N.Y. and D.Z. constructed the three-dimensional structure of the protein. R.W. and M.L. purified the recombinant proteins. X.C.G. and Z.H.L. measured enzymatic activity. K.Q.X. and Z.F.Y. performed the data analysis and mapping. The manuscript was authored by L.J.W., M.H., L.Q.Y., and Y.R.Y. All authors contributed to the discussion of results and provided comments on the manuscript. All authors reviewed and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, LJ., Huang, M., Hu, CQ. et al. Characterization of a novel halophilic and thermostable multifunctional cellulase from Ebinur Salt Lake. Sci Rep 15, 32877 (2025). https://doi.org/10.1038/s41598-025-16035-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-16035-z