Abstract
Ovarian cancer is the most lethal gynecological malignancy, with platinum-based chemotherapy serving as the standard first-line treatment. However, the emergence of primary or acquired resistance significantly compromises patient survival. Among the mechanisms contributing to chemotherapy resistance, autophagy plays a crucial role. DnaJ heat shock protein family (Hsp40) member C5 (DNAJC5) is a key regulator of nervous system function. Recent studies suggest that DNAJC5 may also be involved in tumor progression. Based on this, we aimed to investigate the role of DNAJC5 in epithelial ovarian cancer (EOC) chemotherapy resistance. Survival analysis of the TCGA database revealed a strong association between elevated DNAJC5 expression and poor prognosis in ovarian cancer patients. Immunohistochemistry (IHC) and statistical analyses demonstrated that patients with high DNAJC5 expression frequently presented with advanced International Federation of Gynecology and Obstetrics (FIGO) stage and elevated CA125 levels. Functional experiments showed that overexpression of DNAJC5 in A2780 cells markedly increased resistance to cisplatin (DDP). Moreover, cisplatin-resistant EOC cell lines (A2780/DDP) exhibited high levels of DNAJC5 protein expression. Conversely, DNAJC5 knockdown enhanced cisplatin-induced apoptosis and inhibited autophagy in A2780/DDP cells, as confirmed by Annexin V/PI staining, CCK-8 assays, colony formation assays, and Western blot analysis. Further mechanistic investigations using RNA sequencing identified a strong correlation between DNAJC5 expression and endoplasmic reticulum (ER) stress. Western blot analysis confirmed that DNAJC5 overexpression upregulated BiP-IRE1α-XBP1 protein expression, which was significantly associated with both increased autophagy levels and enhanced DDP resistance. These findings were further supported by validation experiments using hydroxychloroquine(CQ) and 4µ8C, as well as in vivo animal studies. In conclusion, this study demonstrates that DNAJC5 overexpression promotes DDP resistance in EOC by modulating autophagy through the BiP-IRE1α-XBP1 signaling pathway.
Similar content being viewed by others
Introduction
Epithelial ovarian cancer (EOC) is the most prevalent subtype of ovarian cancer, accounting for approximately 80% of cases. Due to the absence of specific early symptoms, EOC is often diagnosed at an advanced stage, frequently with extensive abdominal metastasis1. As a result, it has the highest mortality rate among gynecological cancers2,3. The current standard treatment involves cytoreductive surgery followed by platinum-based chemotherapy, typically using cisplatin (DDP) or carboplatin4. Although this approach initially achieves favorable responses in many patients, nearly 75% experience relapse, primarily due to drug resistance, which significantly reduces the five-year survival rate to below 35%5. DDP resistance is particularly prevalent among EOC patients. Managing recurrent cases often necessitates increased DDP doses, leading to heightened toxicity and severe adverse effects6. Consequently, understanding the mechanisms underlying DDP resistance is essential for developing more effective therapeutic strategies and improving patient outcomes.
Autophagy, an intracellular self-digestion process, has recently been recognized as a major challenge in EOC chemotherapy, primarily due to its role in promoting DDP resistance under endoplasmic reticulum (ER) stress7. Autophagy is classified into three main types: macroautophagy, chaperone-mediated autophagy (CMA), and microautophagy8. Macroautophagy involves the formation of autophagosomes, which supply essential proteins and ATP to sustain cell survival under stress conditions9. CMA, in contrast, relies on Heat Shock Cognate Protein 70 (Hsc70) to bind specific cytosolic proteins, which are subsequently transported across the lysosomal membrane for degradation10. Microautophagy entails the direct invagination of the lysosomal membrane to engulf intracellular substrates11. Autophagy plays diverse roles in cellular function. In healthy cells, it preserves homeostasis by eliminating oncogenic substrates, toxic misfolded proteins, and damaged organelles, thereby preventing chronic cellular damage and malignant transformation. However, in cancerous cells, upregulated autophagy enhances survival and chemotherapy resistance. Notably, it is often detected at higher levels in malignant tissues than in adjacent normal tissues12.
ER stress, triggered by conditions such as hypoxia, glucose deprivation, oxidative stress, and certain drugs, can contribute to drug resistance in various cancers13. Typically, the heavy chain binding protein (BiP/HSPA5/GRP78), an ER-resident chaperone, binds to three ER-localized transmembrane signal transducers—inositol-requiring enzyme 1α (IRE1α), pancreatic ER kinase (PERK), and activating transcription factor 6 (ATF6)—to suppress their activity14. When ER stress occurs, BiP attaches to misfolded proteins and undergoes a conformational shift, which decreases its affinity for these transducers. This decreased affinity activates the transducers, initiating the unfolded protein response (UPR) to restore ER homeostasis. Among the UPR pathways, the IRE1α-X-box binding protein 1 (XBP1) pathway stands out as the most evolutionarily conserved and plays a pivotal role in this restoration process15.
DNAJC5, a member of the DnaJ Heat Shock Protein Family (Hsp40), is widely expressed and encodes a 198-amino-acid protein featuring two critical domains: the N-terminal J-domain and the cysteine-rich string domain16. The J-domain binds to Heat Shock Protein 70 (Hsp70) to activate its ATPase function, while the string domain, with its 12–15 cysteine residues, plays an essential role in DNAJC5’s functions in synaptic vesicle targeting and neurotransmitter release17,18. Research in Drosophila shows that mutations or knockouts of DNAJC5 lead to synaptic dysfunction, neurodegeneration, and reduced lifespans19. Additionally, DNAJC5 contributes to the clearance of neurotoxic proteins, such as misfolded Tau and α-Synuclein, in neurons20,21. Furthermore, DNAJC5 mutations are linked to cancer, with recent studies indicating that its overexpression promotes liver cancer progression22,23. Our preliminary clinical studies suggest a potential connection between DNAJC5 and EOC progression, highlighting its promise as a target for future research.
Our findings reveal that DNAJC5 promotes DDP resistance in EOC. We present the first evidence that DNAJC5 regulates ER stress by activating the IRE1α/XBP1 pathway, which increases autophagy and strengthens drug resistance in tumor cells. Silencing DNAJC5 and lowering microtubule-associated protein 1 light chain 3 (LC3) levels markedly reversed DDP resistance in resistant cell lines. Additionally, the IRE1α inhibitor 4µ8C amplified the antitumor effects of DDP in A2780/DNAJC5 cells, indicating that targeting DNAJC5 could be a promising approach to overcome DDP resistance in EOC.
Materials and methods
Cell culture and reagents
Human EOC cell lines A2780, SKOV3 and A2780/DDP were obtained from the Type Culture Collection of the Chinese Academy of Sciences (Shanghai, China). A2780, ID8, OVHM, and A2780/DDP cells were cultured in Dulbecco’s Modified Eagle’s Medium (DMEM) supplemented with 10% fetal bovine serum (FBS) and 1% penicillin/streptomycin (P/S). To maintain the drug-resistant phenotype of A2780/DDP cells, 2 µg/mL DDP was added weekly for two days to maintain their drug-resistant phenotype. SKOV3 cells were cultured in 5 A medium with 10% fetal bovine serum and 1% P/S. All cultures were maintained at 37 °C in a humidified atmosphere with 5% CO₂. For cell line authentication, all human cell lines underwent short tandem repeat (STR) analysis, and all experiments were performed with mycoplasma-free cells.
Bioinformatic processing of the cancer genome atlas (TCGA) data
Transcriptomic and clinical data for EOC patients were retrieved from the TCGA database. Using the R package maxstat (version 0.7–25), the optimal cut-off for ENSG00000101152 (DNAJC5) was calculated, with a minimum group sample size set above 25% and a maximum below 75%. This cut-off divided patients into high and low expression groups. Prognostic differences between the groups were then analyzed using the survfit function in the R package survival. A log-rank test confirmed significant differences in prognosis between groups (p = 7.7e-3).
Tissue samples and immunohistochemistry (IHC) assay
Tissue samples were collected from Nanjing First Hospital. This investigation had received approval from the Ethics Committee of the Nanjing First Hospital (No.KY20240822-KS-06), and informed consent was obtained from all subjects and/or their legal guardians. All methods were performed in accordance with the relevant guidelines and regulations. Clinical samples were formalin-fixed, paraffin-embedded, and sectioned into 4-µm slices. EOC tumors were staged following the 2018 International Federation of Gynecology and Obstetrics (FIGO) stage, ensuring that no subjects had received immunotherapy or neoadjuvant chemotherapy prior to sample collection.
For IHC analysis, formalin-fixed, paraffin-embedded sections of EOC and adjacent tissues were mounted on slides, deparaffinized in xylene, rehydrated through a graded ethanol series, and rinsed with distilled water. Antigen retrieval was performed by microwaving slides in Ethylene Diamine Tetraacetic Acid (EDTA) buffer (pH 9.0) for 45 min after blocking endogenous peroxidase activity with 3% hydrogen peroxide. Primary antibodies were applied and incubated at 37 °C for over two h. After washing in buffer, secondary antibodies were added and incubated for 30 min, followed by gentle rinsing. Sections were subsequently treated with the Polink-2 Horseradish Peroxidase Diaminobenzidine (DAB) Detection Kit, counterstained with hematoxylin, dehydrated, and mounted with neutral balsam.
Protein expression was quantified using the Immunoreactive Score (IRS), calculated as staining intensity (0 = no staining, 1 = weak, 2 = moderate, 3 = strong) multiplied by the staining area (0 = no staining, 1 = 0–25%, 2 = 26–50%, 3 = 51–75%,4 = 76%-100%). Scores were categorized as follows: 0 = negative, 1–3 = low staining (+), 4–6 = moderate staining (++), and > 6 = high staining (+++). Staining intensity was independently assessed by two observers, and Chi-square tests were conducted to evaluate the correlation between DNAJC5 expression and clinical features such as age, FIGOstage, carbohydrate antigen 125(CA125) levels, lymph node status, and omental metastasis.
Transient transfection and lentivirus infection
Small interfering RNA (siRNA) targeting the DNAJC5 gene was procured from Tsingke (Beijing, China). A2780/DDP cells were seeded at a density of 1 × 10^5 cells per well in a 6-well plate and cultured until approximately 70% confluence was reached. The siRNA was combined with Lipo3000 reagent (#KGA9705-1.5, Keygen, China) and gently added to each well. After a 6-hour incubation, the transfection mixture was replaced with complete medium, and cells were collected 48 h post-transfection for subsequent analyses. The siRNA sequences targeting DNAJC5 were as follows (from 5’ to 3’):
-
Homo: CACUGACGGGUUCAACUAA
UUAGUUGAACCCGUCAGUG
-
Mouse: AGAAUAUCUUUAUGAAUUCCA
GAAUUCAUAAAGAUAUUCUGC
Lentiviral vectors for DNAJC5 overexpression (lentivirus-oeDNAJC5) and a negative control were designed by Genechem (Shanghai, China). A2780/DDP cells were exposed to viral supernatants containing 8 µg/mL polybrene (#40804ES76, YEASEN, China) for 16 h, then cultured in fresh medium supplemented with 1 µg/mL puromycin (without P/S) for one week. Western blot analysis verified successful transfection, and positively transfected cells were subsequently expanded for further experimentation.
RNA sequencing (RNA-seq) and bioinformatics analysis
Total RNA was extracted from cells using the RNA Isolater Total RNA Extraction Reagent (#R401-01, Vazyme, China). After confirming the quality of the samples, 2 µg of RNA from each sample was selected for mRNA enrichment using Oligo(dT) beads, which facilitated library preparation and sequencing. Following quality control, libraries were pooled according to their effective concentrations and the desired data output. Sequencing was conducted on the Illumina NovaSeq 6000 platform using the PE150 sequencing strategy. Differential gene expression analysis was performed using DESeq (v1.10.1), with differentially expressed genes identified based on criteria of |log2FoldChange| > 0.5 and p < 0.05. Gene Ontology (GO) enrichment analysis was carried out using GOseq (v1.22), and a volcano plot was generated with R software (v4.3.0).
Cell viability assay
Cell viability was assessed using the Cell Counting Kit-8 (CCK-8; #K1018, APE×BIO, USA). Cells were seeded in 96-well plates at a density of 4,000 cells per well and allowed to grow for 24 h. Following this incubation, the cells were treated with varying concentrations of DDP (HANSOH PHARMA, China). After 72 h, fresh medium containing 10% CCK-8 reagent was added, and the cells were incubated for an additional 1.5 h at 37℃. Absorbance at 450 nm was then measured using a microplate reader. To investigate the role of autophagy in DDP resistance, cells were pretreated with 10 µM hydroxychloroquine (CQ) for 8 h prior to DDP exposure, after which absorbance at 450 nm was measured again.
Plate colony for mation assay
For colony formation assays, cells were seeded in 12-well plates at a density of 300 cells per well and cultured for 10 days, with the medium replaced every 3 days. After the incubation period, colonies were fixed with 4% paraformaldehyde (#BL539A, Biosharp, China) and stained with crystal violet. Colony counts were then analyzed using ImageJ software.
Transwell invasion assay
For migration assays, a 24-well plate was prepared by adding 500 µL of DMEM containing 20% FBS to each well. Transwell inserts, pre-coated with a 1:8 Matrigel-DMEM mixture, were placed in each well. Cells, digested with 0.25% trypsin-EDTA (#2508964, Gibco, Canada), were resuspended in serum-free DMEM at a concentration of 8×104cells/mL. Subsequently, 100 µL of this cell suspension was added to each insert and incubated at 37 °C for 48 h. After incubation, cells on the lower surface of the inserts were fixed with 4% paraformaldehyde, stained with 0.1% crystal violet (#G1062, Solarbio, China), and images were captured from three randomly selected fields per condition under a microscope.
Wound healing assay
For wound healing assays, cells were seeded in 6-well plates at a density of 8×105 cells per well and grown to 80–90% confluence. A scratch was then created using a sterile pipette tip, and any detached cells were removed with Phosphate Buffered Saline (PBS). Serum-free medium was added, and images of the wound were captured at 0, 24, and 48 h. The wound areas were measured from three randomly selected fields per condition.
Cell apoptosis assay
Apoptosis was analyzed using the Annexin V-fluorescein isothiocyanate (FITC)/propidium iodide (PI) apoptosis detection kit (#KGA1102-100, KeyGEN BioTEC, China). Cells were seeded in 6-well plates and treated with DDP for 72 h. After treatment, the cells were harvested and stained with FITC and PI according to the manufacturer’s protocol. The stained cells were then analyzed using a FACSCanto II flow cytometer (BD Biosciences).
Western blot
For protein extraction, cells were lysed on ice using lysis buffer (#P0013, Beyotime, China) for 30 min, with intermittent vortexing every 5 min. After lysis, high-speed centrifugation was performed to collect the supernatant. Protein concentrations were quantified using the Bicinchoninic Acid (BCA)protein assay kit (#BK0001-02, ACE, China). Samples were then mixed with 5×sodium dodecyl sulfate polyacrylamide gel electrophoresis (5×SDS-PAGE) loading buffer, heated at 95 °C for 10 min, and separated on FuturePAGE gels (#ET15010Gel, #ET15012Gel, ACE, China).
Proteins were transferred onto polyvinylidene fluoride(PVDF) membranes, blocked in Tris Buffered Saline with Tween 20(TBST) with 5% milk, and incubated overnight at 4 °C with primary antibodies. Afterward, Horseradish Peroxidase (HRP)-conjugated secondary antibodies were added and incubated for 2 h at room temperature. Protein bands were visualized using a chemiluminescence imaging system (Tanno5200) along with the SuperPico Enhanced chemiluminescence (ECL) Chemiluminescence Kit (#E422-C1-P1, Vazyme, China).
The antibodies used for Western blot analysis included β-actin (#66009-1-Ig, Proteintech, USA), glyceraldehyde-3-phosphate dehydrogenase(GAPDH,#60004-1-Ig, Proteintech, USA), DNAJC5 (#DF4011, Affinity, USA), BiP (#11587-1-AP, Proteintech, USA), IRE1α (#27528-1-AP, Proteintech, USA), phospho-IRE1α (Ser724)(p-IRE1α,#AF7150, Affinity, USA), XBP1u (#bs23973R, Bioss, China), the spliced form of X-box binding protein 1(XBP1s,#24868-1-AP, Proteintech, USA), PERK (#24390-1-AP, Proteintech, USA), activating transcription factor 4 (ATF4,#97038, Cell Signaling, USA), ATF6 (#AF6243, Beyotime, China), and LC3 (#14600-1-AP, Proteintech, USA), p62(#AF0279, Beyotime, China), Beclin-1(#11306-1-AP, Proteintech, USA).
Immunoprecipitation (IP)
The magnetic beads and antibodies were pretreated according to the specified protocol (#HY-K0202, MCE, USA). Subsequently, 400 µL of the diluted antibody was added to the magnetic beads, resuspended thoroughly, and incubated on a rotating mixer at room temperature for 30 min, followed by incubation at 4 °C for 2 h. The beads were then magnetically separated, collected, and washed thoroughly with buffer.
Next, 400 µL of the antigen sample was added, and the mixture was incubated under the same conditions. The beads were magnetically separated, and the supernatant was discarded. After resuspension, the beads underwent an additional magnetic separation, followed by the removal of the supernatant. Multiple washing steps were performed to ensure purity.Finally, 25–50 µL of 1× SDS-PAGE loading buffer was added to the beads, mixed thoroughly, and heated at 95 °C for 5 min. The beads were then magnetically separated, and the supernatant was collected for Western blot analysis.
RNA extraction and quantitative polymerase chain reaction(qPCR)
Total RNA was isolated using the RNA isolator reagent (#R401-01-AA, Vazyme, China), and its concentration and quality were assessed with a NanoDrop 2000 spectrophotometer (Thermo). 1 µg of total RNA was reverse-transcribed using the HiScript II One Step qRT-PCR SYBR Green Kit (#Q221, Vazyme, China) in a 10 µL reaction volume. Each experiment was conducted in triplicate, and the resulting data were analyzed using LightCycler 96 SW1.1 and Prism software. The primer sequences utilized for qPCR are provided in Supplementary Table 2.
Animal models
All experimental protocols were used and performed in strict accordance with the approval of the Animal Care and Use Committee of Nanjing First Hospital, Nanjing Medical University (No. DWSY-24143600), which are based on the recommendations of the National Institutes of Health and the ARRIVE Guidelines (https://arriveguidelines.org). For the subcutaneous xenograft model, eight-week-old female Balb/c nude mice were housed in High Efficiency Particulate Air (HEPA)-filtered barrier facilities. Tumor cells were prepared as a single-cell suspension at a concentration of 1 × 107 cells/mL, and each mouse received a subcutaneous injection of 4 × 106 cells to induce tumor growth. One week after tumor establishment, the treatment group received intraperitoneal injections of DDP at a dosage of 3 mg/kg, while the control group was administered saline three times per week. Tumor size and body weight were monitored weekly. The tumor inhibition rate was calculated using the following formula: Tumor Inhibition Rate = (Pre-treatment tumor volume - post-treatment tumor volume) / pre-treatment tumor volume×100.
Statistical analysis
Data were obtained from at least three independent experiments and are presented as mean ± standard deviation. Group differences were analyzed using chi-square tests and Student’s t-test, while Kaplan-Meier and log-rank tests were used to assess survival differences based on DNAJC5 expression. To investigate the relationships among DNAJC5 expression, FIGO stage, and CA125 levels, Mann–Whitney U analysis was performed to compare DNAJC5 expression across different FIGO stages, while Spearman correlation analysis was used to assess the strength and direction of associations between these variables. Statistical analyses were performed using SPSS (version 24.0, SPSS Inc., Chicago, IL, USA) and GraphPad Prism 8.0, with significance defined as P < 0.05(*), P<0.01(**), P<0.001(***).
Results
Increased expression of DNAJC5 in EOC and its association with disease progression
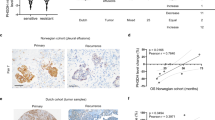
To investigate the role and underlying mechanisms of DNAJC5 in EOC, data from TCGA were analyzed. A significant correlation was identified between elevated DNAJC5 expression and reduced overall survival (Fig. 1a). To validate these findings, IHC analysis was performed on 67 EOC samples. Microscopic examination revealed that DNAJC5 was predominantly localized in the cytoplasm (Fig. 1b). Mann-Whitney U analysis of IRS demonstrated a significant association between DNAJC5 expression and advanced EOC stage (P < 0.001). Specifically, tumors at stages III/IV exhibited higher IRS than those at stages I/II, suggesting a potential oncogenic role of DNAJC5 (Fig. 1c).
DNAJC5 is highly expressed in ovarian cancer tissues and is associated with progression. (a) Analysis of DNAJC5 expression levels in ovarian cancer patients based on the TCGA dataset. Survival analysis was performed using the R package. (b) Representative IHC images of pathological sections from EOC patients, demonstrating varying levels of DNAJC5 expression (-, +, ++, +++). Optical magnification: 10 × 20.The scale bar represents 100 μm and is located in the lower-left corner of the images. (c) Mann-Whitney U analysis results between IHC IRS and FIGO staging of EOC patients; (d) Spearman Correlation Analysis results between the IHC staining intensity of DNAJC5 (-/+/++/+++), FIGO stage, and CA125 index value.
Based on these observations, clinical data were subsequently analyzed using chi-square testing and Spearman correlation analysis. A significant positive correlation was observed between the IHC staining intensity of DNAJC5 and serum CA125 levels (p = 0.016), a biomarker commonly used for EOC prognosis. These findings further highlight the prognostic significance of DNAJC5 in EOC (Table 1; Fig. 1d).
DNAJC5 overexpression promotes DDP resistance in EOC cells in vitro
Clinical and database analyses indicate that elevated DNAJC5 expression in tumors is associated with poorer prognosis in EOC. To further elucidate its role in tumor growth, metastasis, invasion, and chemoresistance, DNAJC5 expression levels were quantified in four EOC cell lines (A2780, SKOV3, ID8, and OVHM) using Western blot analysis. Among these, SKOV3 cells exhibited the highest DNAJC5 expression, whereas A2780 cells showed the lowest levels (Fig. 2a).
Overexpression of DNAJC5 promotes DDP resistance in EOC cells in vitro. (a) Expression levels of DNAJC5 in different EOC cell lines. (b) Western blot analysis confirming DNAJC5 protein expression in A2780 cells infected with a lentivirus overexpressing DNAJC5. (c) A2780 and A2780/DNAJC5 cells were incubated with varying concentrations of DDP (50µM, 40µM, 30µM, 20µM, 10µM, 5 µM, 2.5µM, 1.25µM) for 72 h. Cells were then treated with CCK-8 reagent in the dark for 1.5 h, and the final IC50 values were determined. (d,e) Clonogenic assay of A2780/DNAJC5 cells. A total of 300 cells per well were seeded and treated with 3µM DDP after 24 h. After 10 days, colonies were stained with crystal violet and quantified using ImageJ software. The bar graph represents the effect of DDP on the viability of A2780 and A2780/DNAJC5 cells. (f,g) Apoptosis assay of A2780/DNAJC5 cells following treatment with 3µM DDP for 72 h. The bar graph illustrates the average apoptosis rate of A2780/DNAJC5 cells under DDP treatment compared to the Vector control.
To assess the functional role of DNAJC5, a stable A2780 cell line overexpressing DNAJC5 (A2780/DNAJC5) was established via lentiviral transduction, and successful overexpression was confirmed by Western blot analysis (Fig. 2b). CCK-8 assays revealed no significant difference in proliferation between A2780/DNAJC5 cells and vector controls (Supplementary Fig. 1a). Similarly, wound healing and transwell assays showed no substantial changes in migration or invasion capacity (Supplementary Fig. 1b–e). In contrast, following treatment with DDP, the half-maximal inhibitory concentration (IC50) of A2780/DNAJC5 cells increased to 19.51 µM, significantly higher than that of vector cells, indicating enhanced resistance to DDP (Fig. 2c). This chemoresistant phenotype was further supported by colony formation assays, which demonstrated a greater number of colonies in A2780/DNAJC5 cells after DDP exposure compared to controls (Figs. 2d–e). To examine the effect of DNAJC5 on apoptosis, A2780/DNAJC5 cells were treated with DDP and subjected to apoptosis assays. A significant reduction in apoptotic cell numbers was observed relative to vector controls, suggesting that DNAJC5 promotes cell survival under DDP treatment (Figs. 2f–g).
Knockdown of DNAJC5 reduces DDP resistance in A2780/DDP cells
Given that DNAJC5 overexpression enhanced DDP resistance in A2780 cells, the potential of reversing this effect through gene silencing was investigated in DDP-resistant A2780 cells (A2780/DDP). Western blot analysis confirmed that DNAJC5 expression was markedly elevated in A2780/DDP cells compared to parental A2780 cells (Fig. 3a). Following siRNA-mediated knockdown of DNAJC5 (si-DNAJC5), the IC50 of DDP in A2780/DDP cells decreased by 65% relative to control cells, indicating a strong association between DNAJC5 expression and chemoresistance (Figs. 3b–c). Consistently, si-DNAJC5 cells exhibited significantly reduced colony formation following DDP exposure (Figs. 3d–e), which was further corroborated by flow cytometry analysis showing increased apoptosis (Figs. 3f–g). These findings collectively demonstrate that DNAJC5 plays a crucial role in mediating DDP resistance in A2780 cells.
Silencing DNAJC5 expression reduces resistance in A2780/DDP cells. (a) Western blot analysis of DNAJC5 protein expression levels in A2780, A2780/DNAJC5, and A2780/DDP cells. (b) Western blot validation of the efficiency of siRNA-mediated knockdown of DNAJC5 in A2780/DDP cells. (c) Determination of the IC50 values for A2780/DDP and siDNAJC5-transfected A2780/DDP cells using the CCK-8 assay. Cells were incubated with a concentration gradient of DDP (50µM, 40µM, 30µM, 20µM, 10µM, 5 µM, 2.5µM, 1.25µM) for 72 h, after which the final IC50 values were obtained and plotted. (d,e) Clonogenic assay of A2780/DDP cells transfected with siDNAJC5. A total of 300 cells per well were seeded and treated with 3µM DDP. After 10 days, colonies were stained with crystal violet and quantified using ImageJ software. The bar graph represents the effect of DDP on the viability of A2780/DDP and siDNAJC5-transfected cells. The Vector group refers to A2780/DDP cells, while the siDNAJC5 group consists of A2780/DDP cells subjected to siRNA interference. (f,g) Apoptosis assay of A2780/DDP cells transfected with siDNAJC5 following treatment with 3µM DDP. The bar graph illustrates the average apoptosis rate of A2780/DDP cells under DDP treatment compared to the Vector control. The Vector group corresponds to A2780/DDP cells, whereas the siDNAJC5 group represents A2780/DDP cells with siRNA-mediated knockdown.
In previous experiments, Western blot analysis revealed higher DNAJC5 expression in SKOV3, ID8, and OVHM cells compared to A2780 cells, with varying expression levels (Fig. 2a). To determine whether DNAJC5-mediated DDP resistance is cell-type specific, siRNA knockdown of DNAJC5 was performed in these additional EOC cell lines. Efficient knockdown was verified by Western blot analysis (Supplementary Figs. 2a–c). Colony formation assays conducted under 3 µM DDP treatment for 10 days showed that, unlike in A2780/DDP cells, DNAJC5 silencing did not reduce colony formation in SKOV3, ID8, or OVHM cells (Supplementary Fig. 2d). Bar graph analyses supported this observation (Supplementary Figs. 2e–g). These results suggest that the role of DNAJC5 in mediating DDP resistance is likely cell-type dependent, underscoring the need for further investigation into its regulatory mechanisms in diverse cellular contexts.
DNAJC5 triggers ER stress-mediated autophagy and confers DDP resistance in EOC cells
To further investigate the regulatory role of DNAJC5 in EOC cells, RNA-seq analysis was performed on both A2780 andA2780/DNAJC5 cells, followed by bioinformatic analysis. The results revealed significant enrichment of the “response to endoplasmic reticulum stress” pathway in A2780/DNAJC5 cells (Fig. 4a), along with a marked upregulation of HSPA5 (BiP) expression (Fig. 4b), which was confirmed by qPCR (Supplementary Fig. 3a). These findings indicate a strong association between DNAJC5 and ER stress.
DNAJC5 triggers ER stress-mediated autophagy and confers DDP resistance in EOC cells. (a) The bar graph presents the top 30 enriched pathways identified by KEGG analysis. Notably, the “Response to endoplasmic reticulum stress” pathway was highly enriched in A2780/DNAJC5 cells. (b) Volcano plot illustrating differentially expressed genes in transcriptome sequencing of A2780 vs. A2780/DNAJC5 cells. A total of 25 genes were upregulated and 63 genes were downregulated, with significance criteria of p adj < 0.05 and |log₂FoldChange| > 0.5. The differential expression of HSPA5 (BiP) was particularly significant (p = 2.25 × 10⁻²⁰⁶). (c,d) Western blot analysis of BiP and the three branches—PERK, IRE1α, and ATF6—in A2780 and A2780/DNAJC5 cells. (e) Western blot verification of BiP-IRE1α-XBP1 branch protein expression in siDNAJC5-transfected A2780/DDP cells compared to untreated A2780/DDP cells. (f–h) IP analysis detected the interaction between DNAJC5 and BiP in A2780/DNAJC5 cells, A2780/DNAJC5 cells treated with 3.5 μM DDP for 12 h, and A2780/DDP cells; (i,j) Western blot analysis of autophagy-related proteins expression in A2780/DNAJC5 and siDNAJC5-transfected A2780/DDP cells. (k) Determination of IC50 values in A2780/DNAJC5 cells following autophagy inhibition with chloroquine using CCK-8 assay. Cells were treated with 10µM chloroquine for 8 h, followed by incubation with different concentrations of DDP for 72 h before absorbance measurement.
ER stress, typically triggered by adverse cellular conditions, can facilitate chemoresistance by inducing autophagy. This response involves the activation of three major sensors: PERK, IRE1α, and ATF6. Under stress, BiP dissociates from these sensors upon binding misfolded proteins, leading to the activation of downstream pathways. The contribution of each signaling branch varies depending on cellular context and environmental stimuli. To validate the role of DNAJC5 in ER stress-mediated autophagy, further experiments were conducted. Western blot analysis showed that the BiP-IRE1α-XBP1 pathway was selectively activated in DNAJC5-overexpressing A2780 cells, as indicated by increased levels of phosphorylated IRE1α and spliced XBP1 (XBP1s) (Fig. 4c), while the PERK and ATF6 branches remained largely unaffected (Fig. 4d). In contrast, knockdown of DNAJC5 in A2780/DDP cells significantly suppressed the BiP-IRE1α-XBP1 signaling axis (Fig. 4e).
To elucidate the mechanistic relationship between DNAJC5 and BiP-IRE1α-XBP1 signaling, IP assays were conducted. Under basal conditions, no direct interaction between DNAJC5 and BiP was detected in A2780/DNAJC5 cells (Fig. 4f). However, upon exposure to 3.5 µM DDP for 12 h, a binding interaction was observed (Fig. 4g). A similar interaction was detected in A2780/DDP cells, suggesting that DDP treatment may facilitate or enhance the interaction between DNAJC5 and BiP. These results imply that DNAJC5 may contribute to chemoresistance by modulating BiP-dependent ER stress signaling (Fig. 4h).
In addition, Western blot assays demonstrated that DNAJC5 regulates autophagy in EOC cells. Overexpression of DNAJC5 in A2780 cells promoted autophagic activity, as indicated by increased LC3-II conversion, elevated Beclin-1 expression, and reduced P62 levels (Fig. 4i). Conversely, silencing DNAJC5 in A2780/DDP cells led to partial suppression of autophagy (Fig. 4j). To further evaluate the role of autophagy in DDP resistance, A2780/DNAJC5 cells were pretreated with 10 µM CQ to pharmacologically inhibit autophagy, followed by 3 µM DDP treatment. After 72 h, CCK-8 assays revealed that autophagy inhibition led to a decrease in DDP resistance, as evidenced by a reduced IC50 (Fig. 4k). Collectively, these findings suggest that DNAJC5 enhances DDP resistance in EOC cells, potentially through the promotion of autophagy via activation of the BiP-IRE1α-XBP1 signaling pathway.
Inhibition of the Bip-IRE1α-XBP1-LC3 pathway reduces DNAJC5-mediated DDP resistance
To elucidate the role of ER stress in DNAJC5-mediated DDP resistance, the impact of IRE1α pathway inhibition on drug sensitivity was examined in EOC cells. A2780/DNAJC5 cells were treated with the IRE1α RNase inhibitor 4µ8C for 72 h. Western blot analysis subsequently revealed a marked reduction in the expression levels of BiP-IRE1α-XBP1 pathway components and LC3 following 4µ8C treatment (Fig. 5a).
Combined inhibition with IRE1α RNase inhibitor 4µ8C mitigates DDP resistance. (a) Western blot analysis of BiP-IRE1α-XBP1 branch proteins and LC3 protein expression in A2780/DNAJC5 cells following treatment with 4µ8C (IRE1 Inhibitor III). Cells were treated with 30µM 4µ8C, with one medium change and drug replenishment during the incubation period. After 72 h, proteins were extracted for analysis. (b) Determination of IC50 values in A2780/DNAJC5 cells following 4µ8C treatment using the CCK-8 assay. (c,d) Clonogenic assay of A2780/DNAJC5 cells following 4µ8C treatment. A total of 300 cells per well were seeded and treated with 3µM DDP. After 10 days, colonies were stained with crystal violet and quantified using ImageJ software. The bar graph represents the effect of IRE1αpathway inhibition on the viability of A2780 and A2780/DNAJC5 cells under DDP treatment. (e,f) Apoptosis assay of A2780/DNAJC5 cells treated with 4µ8C, followed by 3µM DDP treatment. The bar graph illustrates the average apoptosis rate of A2780/DNAJC5 cells after 4µ8C treatment compared to Vector.
Functional assays further supported these findings. Cell viability, assessed using the CCK-8 assay, was significantly lower in A2780/DNAJC5 cells treated with both 4µ8C and DDP compared to those treated with DDP alone (Fig. 5b). Similarly, colony formation assays demonstrated a pronounced decrease in colony numbers under the combined treatment (Figs. 5c–d). Apoptosis assays also indicated a higher proportion of apoptotic cells following dual treatment with 4µ8C and DDP, relative to DDP treatment alone (Figs. 5e–f).
DNAJC5 impedes DDP efficacy in an EOC mouse model
To validate the in vitro findings, the role of DNAJC5 in DDP resistance was assessed using an in vivo EOC model in nude mice bearing A2780/DNAJC5 tumors. The mice were divided into four groups based on DNAJC5 expression and treatment regimen: A2780 cells with saline (Vector), A2780 cells with DDP (Vector + DDP), A2780/DNAJC5 cells with saline (DNAJC5), and A2780/DNAJC5 cells with DDP (DNAJC5 + DDP).Once tumor volumes reached approximately 100 mm³, DDP or saline was administered intraperitoneally three times per week, and body weight was monitored throughout the treatment period. After three weeks, tumors were harvested for further analysis.Notably, in the saline-treated groups, tumors in the DNAJC5 group grew more rapidly than those in the Vector group, and this disparity became more pronounced following DDP treatment. Additionally, the tumor inhibition rate was significantly higher in the Vector + DDP group compared to the DNAJC5 + DDP group (Figs. 6a–c), further emphasizing the role of DNAJC5 in conferring DDP resistance.
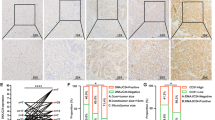
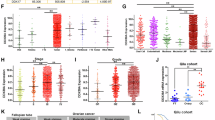
DNAJC5 impedes EOC response to DDP in vivo. (a) Representative images of tumor tissues from nude mice in the A2780/DNAJC5 (DNAJC5) and A2780 (Vector) groups following treatment with either DDP (3 mg/kg) or saline. Each group consisted of 5 mice for repeated validation to reduce errors (b,c) Tumor volume and weight measurements of excised tumors from the different treatment groups. (d) IHC staining and representative images of tumor tissue sections for Ki-67, DNAJC5, and proteins involved in the BiP-IRE1α-XBP1-LC3 pathway. Optical magnification: 10 × 20 .The scale bar represents 100 μm and is located in the lower-right corner of the images. (e–l) The bar graph presents statistical analyses of the IHC score. Each antibody was validated by repeated staining on tumor sections from five mice in the same group, and the resulting data were analyzed using a t-test.
To explore the underlying mechanisms, IHC analysis was performed on tumor sections (Fig. 6d). Ki67 staining revealed a higher proliferative index in the DNAJC5 group than in the Vector group, suggesting that DNAJC5 enhances tumor cell proliferation in vivo. This observation contrasts with previous in vitro findings and may be attributed to the influence of the complex tumor microenvironment, which is not adequately recapitulated in two-dimensional cell culture systems (Fig. 6e).
Moreover, in the Vector + DDP group, intraperitoneal DDP administration led to a relative increase in DNAJC5 expression, accompanied by moderate activation of the BiP-IRE1α-XBP1-LC3 signaling axis. These results suggest that DDP exposure may induce DNAJC5 upregulation in A2780 cells. However, the magnitude of this upregulation was insufficient to confer a resistance phenotype comparable to that observed in the DNAJC5 + DDP group, implying that the level of DNAJC5 expression is a critical determinant of DDP resistance in EOC (Figs. 6f–l).
In summary, these findings indicate that targeting DNAJC5 may alleviate ER stress, reduce DDP resistance, and impede tumor progression in epithelial ovarian cancer.
Discussion
DNAJC5, a member of the HSP40 family, is ubiquitously expressed across various tissues and is implicated in both neural and cancer biology24. This study identifies DNAJC5 as a potential prognostic biomarker in EOC. Clinical and experimental analyses demonstrated that DNAJC5 enhances DDP resistance by suppressing DDP-induced apoptosis in EOC cells. RNA-seq and subsequent assays further revealed that DNAJC5 facilitates macroautophagy through activation of the BiP–IRE1α–XBP1 signaling pathway, thereby indirectly promoting DDP resistance. Importantly, co-treatment with DDP and an IRE1α inhibitor effectively reversed DNAJC5-mediated resistance, highlighting IRE1α as a promising therapeutic target to mitigate platinum resistance in EOC (Fig. 7).
DNAJC5-mediated ER Stress activation induces autophagy and enhances DDP resistance in EOC. In response to DDPtreatment in EOC patients, intracellular DNAJC5 expression is upregulated. This facilitates the binding of BiP to misfolded proteins within the ER, leading to its dissociation from IRE1α and subsequent phosphorylation (p-IRE1α). p-IRE1α promotes the splicing of XBP1u, increasing XBP1s expression. As a result, Beclin-1 and LC3II protein levels rise, while P62 undergoes degradation, leading to enhanced autophagic flux and the development of DDP resistance.
TCGA database indicated a strong association between high DNAJC5 expression and poor prognosis. IHC and statistical evaluation demonstrated that DNAJC5 expression positively correlates with FIGO stage in EOC patients, and elevated expression is frequently accompanied by increased CA125 levels. These findings suggest that DNAJC5 is associated with tumor progression, in line with observations in hepatocellular carcinoma (HCC)22. Functional assays further confirmed that DNAJC5 enhances the capacity of A2780 cells to evade DDP-induced apoptosis and confers resistance to cisplatin. Collectively, these results support the notion that targeting DNAJC5 may offer a viable strategy for overcoming DDP resistance in EOC.
In addition, the downstream mechanisms of DNAJC5 in EOC were investigated. RNA-seq enrichment analysis revealed a significant association between high DNAJC5 expression and ER stress. ER stress has been implicated in chemotherapy resistance due to its regulatory role in essential cellular processes under stress conditions25,26. Previous studies have shown that the IRE1α-XBP1 signaling axis promotes IL-6 expression through direct promoter binding, with IL-6 modulating autophagy and chemoresistance by influencing the phosphorylation of BCL2-interacting protein (BECN1)15,27. In this study, the association between DNAJC5 and ER stress was further validated via Western blot analysis. Notably, DNAJC5 selectively activated the BiP-IRE1α-XBP1 branch of ER stress, with minimal effects on the BiP-PERK-ATF4 and BiP-ATF6 pathways. IP assays confirmed that DNAJC5 binds to BiP upon DDP treatment. This interaction was also observed in A2780/DDP cells, suggesting that DDP enhances the association between DNAJC5 and BiP, potentially by increasing binding affinity or the frequency of substrate delivery. This mechanism may contribute to the development of chemoresistance.
Building on these findings, the mechanism by which DNAJC5-mediated ER stress induces DDP resistance in EOC was further explored. Western blot analysis showed that DNAJC5 overexpression elevated the LC3-II/LC3-I ratio and Beclin-1 levels while reducing P62 expression. These results support the hypothesis that DNAJC5 activates autophagy through ER stress regulation. ER stress is a well-established inducer of autophagy. For example, in a study of colon cancer, Xiaohan Jiang et al. demonstrated that the ER membrane protein B-cell receptor-associated protein 31 (BAP31) promotes apoptosis and autophagy by mediating ER-mitochondrial communication. BAP31 significantly increased LC3-II accumulation, indicating autophagy activation, whereas PERK knockdown suppressed this effect, suggesting that BAP31 induces autophagy via the PERK-ATF4 pathway28. Similarly, in tongue squamous cell carcinoma (TSCC), melatonin (MT) treatment upregulated ATF4, ATF6, BiP, BAP31, and CHOP expression in CRL-1623 cells, leading to ER stress-mediated autophagy and apoptosis29.
To further investigate this association, ER stress and autophagy levels were evaluated in A2780/DDP cells. Elevated DNAJC5 expression was confirmed in these cells. Silencing DNAJC5 led to reduced BiP-IRE1α-XBP1 expression, a decreased LC3-II/LC3-I ratio, lower Beclin-1 protein levels, and increased P62 expression. Notably, partial reversal of DDP resistance was observed following DNAJC5 knockdown. The contribution of autophagy to chemoresistance was further substantiated by treatment with CQ30. To elucidate the mechanistic relationship between DNAJC5, ER stress, autophagy, and DDP resistance, A2780/DNAJC5 cells were exposed to the IRE1α inhibitor 4µ8C31. The results show that inhibition of IRE1α effectively suppressed activation of the IRE1α/XBP1 signaling axis, decreased the LC3-II/LC3-I ratio, and reduced DDP tolerance in A2780/DNAJC5 cells. These findings support the hypothesis that DNAJC5 indirectly promotes DDP resistance by inducing ER stress, which in turn enhances autophagy.
In vivo studies further confirmed these findings, demonstrating that DNAJC5 contributes to DDP resistance in EOC by activating ER stress-mediated autophagy. Accordingly, targeting DNAJC5 or its downstream pathways may offer a promising strategy for overcoming platinum-based resistance in EOC.
Chemotherapy resistance remains a major clinical challenge in the treatment of EOC, and few molecular markers are currently available for predicting drug sensitivity or identifying resistance at an early stage32.This study demonstrated that DNAJC5 expression was elevated to varying degrees across all FIGO stages in tumor tissues from patients with EOC, with the most pronounced increase observed in intermediate and advanced stages. This elevated expression was significantly associated with shorter overall survival. In addition, high expression of DNAJC5 was significantly correlated with increased CA125 values. These associations underscore the potential of DNAJC5 as a prognostic biomarker for tumor malignancy and therapeutic response. Mechanistically, DNAJC5 was found to promote DDP resistance in EOC cells by activating autophagy through ER stress. These findings collectively suggest that therapeutic strategies targeting DNAJC5—such as gene therapy or the development of small-molecule inhibitors—may reduce ER stress and autophagic activity, thereby preventing or reversing DDP resistance in patients with EOC.
Despite these findings, certain limitations should be acknowledged. DNAJC5-mediated DDP resistance was validated only in A2780 cells, with no significant effects observed in other EOC cell lines such as SKOV3, ID8, and OVHM. Although these lines exhibited varying levels of DNAJC5 expression, siRNA-mediated knockdown did not significantly alter their inherent DDP resistance. These observations imply that the contribution of DNAJC5 to DDP resistance may be subtype-specific. Further studies involving a broader panel of EOC cell lines are warranted to comprehensively evaluate its therapeutic potential.
Conclusion
In summary, this study demonstrates that DNAJC5 contributes to DDP resistance in EOC by promoting autophagy via activation of the BiP-IRE1α-XBP1 signaling pathway. These findings offer important insights into potential molecular targets for overcoming chemoresistance in EOC. Nevertheless, the functional role of DNAJC5 across various ovarian cancer subtypes remains insufficiently characterized, largely due to the limited scope of the current study. Consequently, further comprehensive and systematic investigations are warranted to fully elucidate its biological significance and therapeutic potential.
Data availability
Sequence data that support the findings of this study have been deposited in the National Center for Biotechnology Information with the BioProject ID PRJNA1244766.
Change history
28 January 2026
A Correction to this paper has been published: https://doi.org/10.1038/s41598-025-34686-w
Abbreviations
- DDP:
-
Cisplatin
- EOC:
-
Epithelial ovarian cancer
- DNAJC5:
-
DnaJ Heat Shock Protein Family (Hsp40) Member C5
- Hsp40:
-
Heat Shock Protein Family
- HCC:
-
Hepatocellular carcinoma
- IHC:
-
Immunohistochemistry
- CCK-8:
-
Cell Counting Kit-8
- BiP:
-
Heavy chain binding protein
- IRE1α:
-
Inositol-requiring enzyme 1α
- XBP1:
-
X-box binding protein 1 gene
- XBP1u:
-
The non-splicing form of X-box binding protein 1
- XBP1s:
-
The spliced form of X-box binding protein 1
- LC3:
-
Microtubule-associated protein 1 light chain 3
- ER stress:
-
Endoplasmic reticulum stress
- CMA:
-
Chaperone-mediated autophagy
- Hsc70:
-
Heat shock cognate protein 70
- PERK:
-
Pancreatic ER kinase
- ATF6:
-
Activating transcription factor 6
- UPR:
-
Unfolded protein response
- Hsp70:
-
Heat Shock Protein 70
- DMEM:
-
Dulbecco’s Modified Eagle’s Medium
- STR:
-
Short tandem repeat
- EDTA:
-
Ethylene diamine tetraacetic acid
- DAB:
-
Diaminobenzidine
- IRS:
-
Immunoreactive score
- FIGO:
-
International Federation of Gynecology and Obstetrics
- CA125:
-
Carbohydrate antigen 125
- siRNA:
-
Small interfering RNA
- GO:
-
Gene ontology
- PBS:
-
Phosphate buffered saline
- FBS:
-
Fetal bovine serum
- FITC:
-
Fluorescein isothiocyanate
- PI:
-
Propidium iodide
- BCA:
-
Bicinchoninic acid
- 5×SDS-PAGE:
-
5×sodium dodecyl sulfate polyacrylamide gel electropheresis
- PVDF:
-
Polyvinylidene fluoride
- TBST:
-
Tris Buffered Saline with Tween 20
- HRP:
-
Horseradish peroxidase
- ECL:
-
Enhanced chemiluminescence
- GAPDH:
-
Glyceraldehyde-3-phosphate dehydrogenase
- p-IRE1α:
-
Phospho-IRE1α (Ser724)
- ATF4:
-
Activating transcription factor 4
- qPCR:
-
Quantitative polymerase chain reaction
- NIH:
-
National Institutes of Health
- HEPA:
-
High Efficiency Particulate Air
- TCGA:
-
The Cancer Genome Atlas
- IC50 :
-
Half maximal inhibitory concentration
- TDP-43:
-
TAR DNA binding protein-43
- BECN1:
-
Myosin-like BCL2-interacting protein
- RNA-seq:
-
RNA sequencing
- CQ:
-
Hydroxychloroquine
References
Armbruster, S., Coleman, R. L. & Rauh-Hain, J. A. Management and treatment of recurrent epithelial ovarian cancer. Hematol. Oncol. Clin. N. Am. 32, 965–982. https://doi.org/10.1016/j.hoc.2018.07.005 (2018).
Kossaï, M., Leary, A., Scoazec, J. Y. & Genestie, C. Ovarian cancer: A heterogeneous disease. Pathobiol. J. ImmunoPathol Mol. Cell. Biol. 85, 41–49. https://doi.org/10.1159/000479006 (2018).
Shen, S. et al. A predictive model for initial platinum-based chemotherapy efficacy in patients with postoperative epithelial ovarian cancer using tissue-derived small extracellular vesicles. J. Extracell. Vesicles. 13, e12486. https://doi.org/10.1002/jev2.12486 (2024).
Karakashev, S. & Aird, K. M. Ovarian cancer: how can resistance to chemotherapy be tackled? Future Oncol. (London England). 13, 2737–2739. https://doi.org/10.2217/fon-2017-0235 (2017).
Li, X. et al. Inhibition of VEGFA increases the sensitivity of ovarian cancer cells to chemotherapy by suppressing VEGFA-mediated autophagy. OncoTargets Therapy. 13, 8161–8171. https://doi.org/10.2147/ott.S250392 (2020).
Gu, J. et al. Murine double minute 2 SiRNA and wild-type p53 gene therapy enhances sensitivity of the SKOV3/DDP ovarian cancer cell line to cisplatin chemotherapy in vitro and in vivo. Cancer Lett. 343, 200–209. https://doi.org/10.1016/j.canlet.2013.10.011 (2014).
Mahoney, E. et al. ER stress and autophagy: new discoveries in the mechanism of action and drug resistance of the cyclin-dependent kinase inhibitor flavopiridol. Blood 120, 1262–1273. https://doi.org/10.1182/blood-2011-12-400184 (2012).
Liu, S. et al. Regulator of cell death. Cell Death Dis. 14, 648. https://doi.org/10.1038/s41419-023-06154-8 (2023).
Griffey, C. J. & Yamamoto, A. Macroautophagy in CNS health and disease. Nat. Rev. Neurosci. 23, 411–427. https://doi.org/10.1038/s41583-022-00588-3 (2022).
Lo Dico, A. et al. CMA mediates resistance in breast cancer models. Cancer Cell Int. 23, 133. https://doi.org/10.1186/s12935-023-02969-9 (2023).
Yang, F., Xu, K., Zhou, Y. G. & Ren, T. Insight into autophagy in platinum resistance of cancer. Int. J. Clin. Oncol. 28, 354–362. https://doi.org/10.1007/s10147-023-02301-5 (2023).
Ghosh, B. et al. Simvastatin exerts neuroprotective effects post-stroke by ameliorating endoplasmic reticulum stress and regulating autophagy/apoptosis balance through pAMPK/LC3B/ LAMP2 axis. Exp. Neurol. 381, 114940. https://doi.org/10.1016/j.expneurol.2024.114940 (2024).
Clarke, H. J., Chambers, J. E., Liniker, E. & Marciniak, S. J. Endoplasmic reticulum stress in malignancy. Cancer Cell. 25, 563–573. https://doi.org/10.1016/j.ccr.2014.03.015 (2014).
Chen, X. & Cubillos-Ruiz, J. R. Endoplasmic reticulum stress signals in the tumour and its microenvironment. Nat. Rev. Cancer. 21, 71–88. https://doi.org/10.1038/s41568-020-00312-2 (2021).
Fang, P. et al. IRE1α-XBP1 signaling pathway regulates IL-6 expression and promotes progression of hepatocellular carcinoma. Oncol. Lett. 16, 4729–4736. https://doi.org/10.3892/ol.2018.9176 (2018).
Tobaben, S. et al. A trimeric protein complex functions as a synaptic chaperone machine. Neuron 31, 987–999. https://doi.org/10.1016/s0896-6273(01)00427-5 (2001).
Zhang, H. et al. Cysteine string protein interacts with and modulates the maturation of the cystic fibrosis transmembrane conductance regulator. J. Biol. Chem. 277, 28948–28958. https://doi.org/10.1074/jbc.M111706200 (2002).
Lee, J., Xu, Y. & Ye, Y. Safeguarding lysosomal homeostasis by DNAJC5/CSPα-mediated unconventional protein secretion and endosomal microautophagy. Front. Cell. Dev. Biol. 10, 906453. https://doi.org/10.3389/fcell.2022.906453 (2022).
Burgoyne, R. D. & Morgan, A. Cysteine string protein (CSP) and its role in preventing neurodegeneration. Semin. Cell Dev. Biol. 40, 153–159. https://doi.org/10.1016/j.semcdb.2015.03.008 (2015).
Johnson, J. N., Ahrendt, E. & Braun, J. E. CSPalpha: the neuroprotective J protein. Biochem. Cell. biology = Biochimie Et Biol. Cell.laire. 88, 157–165. https://doi.org/10.1139/o09-124 (2010).
Fontaine, S. N. et al. DnaJ/Hsc70 chaperone complexes control the extracellular release of neurodegenerative-associated proteins. EMBO J. 35, 1537–1549. https://doi.org/10.15252/embj.201593489 (2016).
Wang, H. et al. DNAJC5 promotes hepatocellular carcinoma cells proliferation though regulating SKP2 mediated p27 degradation. Biochim. et Biophys. Acta Mol. Cell. Res. 1868, 118994. https://doi.org/10.1016/j.bbamcr.2021.118994 (2021).
Xu, H. et al. LINC00624 affects hepatocellular carcinoma proliferation and apoptosis through the miR-342-3p/DNAJC5 axis. J. Biochem. Mol. Toxicol. 38, e23650. https://doi.org/10.1002/jbt.23650 (2024).
Wu, S. et al. Unconventional secretion of α-synuclein mediated by palmitoylated DNAJC5 oligomers. eLife 12 https://doi.org/10.7554/eLife.85837 (2023).
Xu, Y. et al. IRE1α-XBP1s axis regulates SREBP1-dependent MRP1 expression to promote chemoresistance in non-small cell lung cancer cells. Thorac. Cancer. 15, 2116–2127. https://doi.org/10.1111/1759-7714.15442 (2024).
Pongking, T. et al. Cannabidiol exhibits potent anti-cancer activity against gemcitabine-resistant cholangiocarcinoma via ER-stress induction in vitro and in vivo. BMC Complement. Med. Ther. 24, 325. https://doi.org/10.1186/s12906-024-04610-2 (2024).
Hu, F. et al. IL-6 regulates autophagy and chemotherapy resistance by promoting BECN1 phosphorylation. Nat. Commun. 12, 3651. https://doi.org/10.1038/s41467-021-23923-1 (2021).
Jiang, X. et al. p20BAP31 induces autophagy in colorectal cancer cells by promoting PERK-Mediated ER stress. Int. J. Mol. Sci. 25 https://doi.org/10.3390/ijms25105101 (2024).
Liu, H. et al. Melatonin inhibits tongue squamous cell carcinoma: interplay of ER stress-induced apoptosis and autophagy with cell migration. Heliyon 10, e29291. https://doi.org/10.1016/j.heliyon.2024.e29291 (2024).
Carew, J. S. et al. REDD1 is a determinant of the sensitivity of renal cell carcinoma cells to autophagy Inhibition that can be therapeutically exploited by targeting PIM activity. Cancer Lett. 613, 217496. https://doi.org/10.1016/j.canlet.2025.217496 (2025).
Arena, A. et al. The Inhibition of IRE1alpha/XBP1 axis prevents EBV-driven lymphomagenesis in NSG mice. Microbiol. Spectr. 11, e0263623. https://doi.org/10.1128/spectrum.02636-23 (2023).
Siahestalkhi, E. K. et al. Characterization of LncRNAs contributing to drug resistance in epithelial ovarian cancer. Med. Oncol. (Northwood Lond. Engl.). 42(84). https://doi.org/10.1007/s12032-025-02628-1 (2025).
Acknowledgements
We acknowledge the supported from the National Natural Science Foundation of China (grant number: 82173205, 82273196), Health Science and Technology Development Key Program of Nanjing (zkx22029), and the biological image resources provided by Figdraw (ID: PORUW1f400).
Author information
Authors and Affiliations
Contributions
S.Z.: Writing – review & editing, Data curation, Methodology, Project administration, Resources, Supervision, Validation, Funding acquisition. W.Z.: Conceptualization, Formal analysis, Investigation, Methodology. S.H.: Writing – original draft, Writing – review & editing, Data curation, Software, Validation, Visualization. L.C.: Writing – original draft, Data curation, Software, Visualization. Y.C., S.L., B.Y., Y.C.: Data curation, software.
Corresponding author
Ethics declarations
Ethics declarations
The use of human tissues in this study was approved by the Ethics Committee of Nanjing First Hospital, Nanjing Medical University (No.KY20240822-KS-06). Written informed consent was obtained from the individual(s) for the publication of any data included in this article. The animal experiments were approved by the Animal Care and Use Committee of Nanjing First Hospital, Nanjing Medical University (No. DWSY-24143600). All of the methods were performed in accordance with relevant guidelines and regulations. All of the methods are reported in accordance with the ARRIVE guidelines.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this Article was revised: In the original version of this Article, Shijia Huang and Ling Chen were omitted as equally contributing authors. In addition, the Supplementary Material files 1 and 3 published with this Article, contained errors. Full information regarding the corrections made can be found in the correction for this Article.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Huang, S., Chen, L., Chen, Y. et al. DNAJC5 promotes cisplatin resistance in epithelial ovarian cancer by autophagy induced by the BiP/IRE1α/XBP1 endoplasmic reticulum stress pathway. Sci Rep 15, 34323 (2025). https://doi.org/10.1038/s41598-025-16625-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-16625-x