Abstract
Accurate diagnosis of brain tumors is critical in understanding the prognosis in terms of the type, growth rate, location, removal strategy, and overall well-being of the patients. Among different modalities used for the detection and classification of brain tumors, a computed tomography (CT) scan is often performed as an early-stage procedure for minor symptoms like headaches. Automated procedures based on artificial intelligence (AI) and machine learning (ML) methods are used to detect and classify brain tumors in Computed Tomography (CT) scan images. However, the key challenges in achieving the desired outcome are associated with the model’s complexity and generalization. To address these issues, we propose a hybrid model that extracts features from CT images using classical machine learning. Additionally, although MRI is a common modality for brain tumor diagnosis, its high cost and longer acquisition time make CT scans a more practical choice for early-stage screening and widespread clinical use. The proposed framework has different stages, including image acquisition, pre-processing, feature extraction, feature selection, and classification. The hybrid architecture combines features from ResNet50, AlexNet, LBP, HOG, and median intensity, classified using a multilayer perceptron. The selection of the relevant features in our proposed hybrid model was extracted using the SelectKBest algorithm. Thus, it optimizes the proposed model performance. In addition, the proposed model incorporates data augmentation to handle the imbalanced datasets. We employed a scoring function to extract the features. The Classification is ensured using a multilayer perceptron neural network (MLP). Unlike most existing hybrid approaches, which primarily target MRI-based brain tumor classification, our method is specifically designed for CT scan images, addressing their unique noise patterns and lower soft-tissue contrast. To the best of our knowledge, this is the first work to integrate LBP, HOG, median intensity, and deep features from both ResNet50 and AlexNet in a structured fusion pipeline for CT brain tumor classification. The proposed hybrid model is tested on data from numerous sources and achieved an accuracy of 94.82%, precision of 94.52%, specificity of 98.35%, and sensitivity of 94.76% compared to state-of-the-art models. While MRI-based models often report higher accuracies, the proposed model achieves 94.82% on CT scans, within 3–4% of leading MRI-based approaches, demonstrating strong generalization despite the modality difference. The proposed hybrid model, combining hand-crafted and deep learning features, effectively improves brain tumor detection and classification accuracy in CT scans. It has the potential for clinical application, aiding in early and accurate diagnosis. Unlike MRI, which is often time-intensive and costly, CT scans are more accessible and faster to acquire, making them suitable for early-stage screening and emergency diagnostics. This reinforces the practical and clinical value of the proposed model in real-world healthcare settings.
Similar content being viewed by others
Introduction
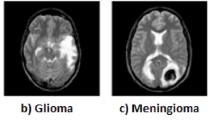
Abnormal cell growth, hormonal imbalance, and partial loss of vision are the symptoms of brain tumors. This tumor affects the body’s functions, including memory, emotions, vision, motor skills, responses, and breathing. Early diagnosis and disability can be averted, and surgical costs minimized. Computer-aided diagnosis, including MRI used due to advancements in medical imaging. The three main techniques (MRI, CT, and PET) for brain malignancies are frequently employed to analyze the anatomy of the brain1. Brain tumors are classified into three primary categories: gliomas, meningiomas, and pituitary tumors. Gliomas, accounting for approximately 30% of all brain tumors, are frequently malignant and have a median survival rate exceeding two years. Meningiomas are typically benign and characterized by slow growth. Although pituitary tumors are often benign, they can lead to significant health complications by disrupting endocrine function, resulting in hormonal imbalances and visual disturbances. The Brain tumor stage is an essential step in the process because proper diagnosis and prevention of the disease depend on a complete understanding for better survival; hence, early detection requires expert opinion in the evaluation of patients2.
Artificial intelligence (AI), machine learning (ML) algorithms, and Deep learning (DL) have made strong progress in medical imaging, and hybrid models are increasingly relevant3,4. Brain tumor medical treatment since then has been markedly optimized by a new technique that merges Deep Learning and Machine Learning. This pioneering way gets rid of human interpretation issues and becomes a better and more accurate detection tool for radiologists. The primary objective of this research is to contribute to the development of brain tumor detection and classification in CT scan images, providing a deep overview of the methodologies existing in the literature, starting from classic machine learning methods to state-of-the-art deep learning systems. Although MRI is frequently utilized for its contrast information in guiding treatment decisions, the manual evaluation of extensive MRI data is both time-intensive and expensive. Similarly, while CT scans play a critical role in the identification of brain tumors, they are also hindered by the challenges associated with manual interpretation. Beyond manual interpretation, automated analysis of CT scan images faces specific challenges, including lower soft-tissue contrast compared to MRI and variability in image quality due to differences in acquisition settings. These factors contribute to difficulty in extracting meaningful features, increasing the complexity of model design and limiting generalization across datasets4. Additionally, deep learning models often require large labeled datasets, which are scarce in the context of annotated CT brain tumor images. This study addresses these challenges by developing a hybrid model that reduces complexity through feature selection and improves generalization by integrating both hand-crafted and deep features from complementary sources. This systematically assesses the strengths and limitations of the latter approaches for the selection of the most efficient models that allow significant enhancement of the performance in brain tumor detection and classification in CT scan images.
The proposed hybrid model, combining hand-crafted and deep learning features, effectively improves brain tumor detection and classification accuracy in CT scans. It has the potential for clinical application, aiding in early and accurate diagnosis, and is vital for patient care. The selection of the relevant features in our proposed hybrid model was extracted using the Select KBest algorithm. Thus, it optimizes the proposed model performance. In addition, the proposed model incorporates data augmentation to handle the imbalanced datasets. We employed a scoring function to extract the features. The Classification is ensured using a multilayer perceptron neural network. The main objective of this study is to develop a novel hybrid model for brain tumor classification using CT scan images, which are more accessible and time-efficient than MRI. Unlike previous works that primarily rely on MRI data or a single deep learning model, our approach integrates both hand-crafted features and deep features, followed by feature selection using the SelectKBest algorithm and classification using an MLP neural network.
The key contributions of this paper include: (1) proposing a hybrid feature extraction strategy combining classical and deep features tailored for CT scans; (2) introducing a feature selection step to reduce complexity and enhance generalization; and (3) demonstrating the model’s superior performance on CT images compared to existing MRI-based approaches.
The research is presented here in four sections, beginning with the literature review section, where existing methods associated with the detection and classification of brain MRI and CT scans are discussed. In the next section, the proposed approach is explained, which begins with the image acquisition steps along with pre-processing, feature extraction, feature selection, and classification. The third section focuses on the experimental results, which evaluate the outcomes of applying the proposed approach to real data sets and provide an analysis of different models and their efficiency. Finally, the conclusion and future work are presented to summarize and encapsulate the findings and discoveries, along with future directions.
Literature review
This section reviews recent techniques for the detection and classification of brain tumors using MRI and CT scan images, detailing the methodologies, implementation strategies, datasets, and associated limitations, while providing recommendations for future advancements. Nabizadeh et al.5 presented an automatic system for detecting tumor slices and defining tumor areas, demonstrating high accuracy and low computational complexity in brain tumor tissue segmentation. It also compares statistical features over Gabor wavelet features for tumor segmentation applications. They are highly redundant and lead to high computational costs. Das et al.6 enhanced treatment planning, growth rate prediction, and clinical trial monitoring. It uses non-sub-sampled shearlet transformation, spatial fuzzy c-means clustering algorithm, and Gabor Filtering. Experimental results show high accuracy rates for various datasets of MRI, but the applicability of the proposed features has not yet been tested on real-time, diverse datasets. Deep learning, a hierarchical machine learning technique, uses multiple layers for intricate features and high-level inferences. In7, they focused on brain tumor classification using magnetic resonance images MRI. The proposed model uses CNNs and a multi-branch network with an inception block. The model was evaluated using the Br35H dataset and compared to other classification models. The CNN with a Multi-Branch Network and Inception block demonstrated exceptional performance. Another research proposed a novel parallel deep convolutional neural network (PDCNN) topology to extract global and local features from two parallel stages, addressing overfitting problems. The method uses regularized and batch normalization, resizing input images, and utilizing grayscale transformation. The method is effective in three MRI datasets, achieving accuracy of 97.33%, 97.60%, and 98.12%, compared to state-of-the-art techniques8. The challenge of non-Euclidean distances in image data and the incapacity of traditional models to infer pixel similarity based on proximity is what Ravinder et al.9 seek to overcome. A graph-based convolutional neural network (GCNN) model is proposed, which improves brain tumor detection and classification in MRI images. The model combines GNN and a 26-layered CNN, with Net-2 outperforming other networks. The model represents a critical alternative for the statistical detection of brain tumors in suspected patients, but requires a high computational demand.
To categorize brain malignancies into three groups: glioma, meningioma, and pituitary tumor, Zulfiqar et al.10 used CNN and Deep Learning using EfficientNets. A publicly available CE-MRI dataset is used to enhance five iterations of pre-trained models. The method involves loading ImageNet weights, adding top layers, and a fully connected layer. Experiments show robustness and data augmentation effects on model accuracy. Reyes et al.11 explored deep architectures like VGG, ResNet, EfficientNet, and ConvNeXt for a network based on two magnetic resonance imaging datasets. The results show several networks achieve high accuracy rates. The largest models are VGG, MobileNet, and EfficientNetB0, with VGG having over 171 million parameters. The fastest models are VGG and MobileNet, with ConvNext being the slowest. The majority of models need fine-tuning after transfer learning to attain the highest accuracy. Khushi et al.30 proposed a brain tumor classification method using MRI images and a pre-trained EfficientNetB4 model. They enhanced the Br35h dataset through data augmentation and employed transfer learning with adjustable learning rates and regularization. Their model achieved an accuracy of 99.87% on the augmented dataset. The study demonstrates the effectiveness of EfficientNetB4 in MRI-based tumor classification, highlighting the potential of deep learning models even on relatively small datasets. However, their model is limited to distinguishing tumor vs. non-tumor cases, lacking the capability for multi-class tumor classification, which is critical for clinical decision-making. Kar and Singh31 proposed MBTC-Net, a multimodal deep learning framework that classifies brain tumors using both CT and MRI scans. Their model leverages EfficientNetV2B0 for high-dimensional feature extraction and employs a multi-head attention mechanism to enhance spatial context understanding. Evaluated on three open-access datasets, including a multimodal CT + MRI dataset, MBTC-Net achieved classification accuracies of 97.54%, 97.97%, and 99.34% across 15-class, 6-class, and binary setups, respectively. Unlike traditional fusion methods, MBTC-Net processes MRI and CT images independently in the same framework, learning distinct representations that support cross-modal generalization. But the system was primarily evaluated in a binary classification setup, and performance on CT scans may be limited due to modality imbalance in the dataset. Future work is planned to address structured fusion and clinical validation. Table 1 provides a summary of the methods employed for tumor classification and detection based on a combination of deep learning methods and machine learning methods in brain images.
Based on the reviewed literature, most existing models either rely on deep learning architectures with millions of parameters—leading to high computational complexity or are limited in their ability to generalize due to modality constraints (MRI-only focus), limited datasets, or lack of feature optimization. Several studies employ powerful CNNs or attention-based models without adequate dimensionality reduction, which increases training time and hardware dependency. Others focus on binary classification only, failing to differentiate between tumor types. These limitations motivate the need for a more efficient and generalizable approach. The proposed hybrid model addresses these gaps by combining hand-crafted and deep features with a feature selection mechanism (SelectKBest), enabling reduced model complexity while maintaining strong classification performance across multiple tumor types in CT scan images. Unlike most hybrid feature-based approaches in the literature, which are predominantly developed for MRI-based brain tumor classification, our proposed method is tailored specifically for CT scan images, considering their distinct noise characteristics and lower soft-tissue contrast. To the best of our knowledge, no prior work has integrated LBP, HOG, and median intensity with deep features from both ResNet50 and AlexNet in a structured fusion pipeline for CT brain tumor classification. This combination, along with a feature selection stage, aims to reduce computational complexity while improving multi-class classification performance in a modality that is faster, more accessible, and more widely used in early clinical screening. This methodology provides a consistent response to the intricacies of the process of carrying out precise and expedient tumor detection.
Proposed framework
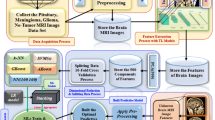
In this section, we explain our proposed framework. The proposed framework has different stages, including image acquisition, pre-processing, feature extraction, feature selection, and classification. The proposed methodology starts with the phase named Data Organization. In this process the collection of CT scan images from publicly accessible datasets as shown in Fig. 1. The improvement in image quality, considering noise, especially in the pre-processing step, relies on nonlinear filtering; further steps include feature selection and extraction using techniques from visual descriptors combined with deep features coming from pre-trained models. We choose appropriate methods to extract the relevant and useful characteristics related to the shape and textural changes of the tumors. In the classification phase, a new multi-layer perceptron model identifies and classifies different types of brain tumors based on their characteristic patterns. In addition, this process is lengthy because of the assurance of effective identification and classification of the unique attributes of brain tumors.
Proposed model.
Image acquisition
The first step is the data acquisition phase. We have used a brain tumor dataset. The dataset is obtained from a free online repository called Radiopaedia. The dataset provides pre-processed CT images from different patients. It can be accessed with the link: https://radiopaedia.org/search?scope=cases&sort=date_of_publication with 1388 computed tomography images representing the findings in each case. For model development, the dataset was randomly partitioned into 70% for training, 15% for validation, and 15% for testing. The split was stratified, ensuring that all four classes (glioma, meningioma, pituitary tumor, and normal brain) were proportionally represented in each subset. To prevent data leakage, care was taken to ensure that images from the same patient were not shared across training and testing sets. This division supports robust model evaluation and avoids overfitting, especially considering the class imbalance present in the original dataset. Table 2 shows the distribution of CT scan images across training, validation, and testing sets. Figure 2 represents sample images of each category in this dataset, including glioma, meningioma, pituitary tumor, and normal brain. The dataset includes variation in the quality of the images, due to a difference in scanning equipment and settings, and inter-patient variability, where the same type of tumors may differ in character. Another problem that this dataset faces is class imbalance, as some tumor types are less represented compared to others, which could result in biased performances from the models. These are some of the challenges that require highly developed pre-processing, feature extraction, and classification methods to handle the complexities inherent in the dataset.
Data augmentation
Data augmentation is a technique that artificially enlarges the number of datasets for detecting and classifying brain tumors. This approach will enhance the model’s capability to generalize, thus giving it the ability to operate even in the presence of errors in unseen and new diagnostic tests. However, it will simultaneously increase diagnostic accuracy and help in overcoming the sample size constraints and issues faced in image acquisition modalities. Performing augmentation increases the diversity of the dataset without any actual collection of new data, hence making the model more robust and avoiding the possibility of overfitting. The Image augmentation stage includes Rotation: Images are rotated within a range of ± 15 degrees. This helps the model become invariant to slight rotations in the input images. Zoom: Images are zoomed in or out within a range of ± 20% (0.2). This augmentation helps the model generalize better to images of varying scales. Contrast: The contrast of images can vary between 0.5 and 1.5 times the original contrast. Adjusting contrast aids in making the model robust to variations in image contrast across the dataset. Rescale: The pixel values of images are rescaled by a factor of 1/255. This normalization ensures that pixel values fall within the range of [0, 1], which is often beneficial for neural network training. These augmentations were applied only to the training set to prevent information leakage and ensure fair model evaluation. Figure 3 presents examples of enhanced CT sequences depicting brain tumors following the application of the previously described data augmentation techniques. Table 3 provides a detailed overview of the distribution of training data across various classes, both before and after the augmentation process was applied to the images within the training set of the dataset.
Sample images from each class.
Example of augmented data.
Pipeline of pre-processing steps.
Data pre-processing
Data pre-processing is a critical step in preparing the images for optimal feature extraction and subsequent analysis. The primary objective of pre-processing is to enhance the quality and consistency of the images, thereby improving the performance of the deep-learning models. Most raw CT images have variability in acquisition settings, noise, and inconsistent illumination that may obscure important features, leading to inaccuracies in the detection and classification of brain tumors. Effective pre-processing mitigates these challenges by standardizing the input data, reducing noise, and enhancing contrast. The pre-processing pipeline is shown in Fig. 4. First, cropping was done to eliminate the background of the brain area. This has the effect of reducing computation and allowing the model to focus only on the relevant data. The image is then resized at 256 × 256 to standardize the input data dimensions and to provide one consistent input size for feeding into the model, subsequently reducing the computational burden and enhancing learning efficiency. Gray scaling reduces the dimensionality while retaining important structural information, further simplifying data for robust feature extraction.
Non-linear filters such as Median filtering reduce noise and remove artifacts, which improves the signal-to-noise ratio and prevents the model from learning irrelevant patterns that operate on each pixel by replacing it with the median value of the intensities within a local neighborhood (kernel). A median filter works in a way that it takes the median of neighboring intensity values of each pixel one by one. The median filter, compared with linear filters like Gaussian blurring, suppresses impulse noise (like salt-and-pepper noise) effectively while preserving edges due to its median being less prone to outliers12,13.
Further, histogram equalization techniques are applied to enhance the contrast and keep the brightness of the images uniform for the proper distribution of intensities within an image. This helps to make the features of tumors more distinguishable. All these steps of pre-processing collectively optimize images for accurate and effective analysis during the training of deep learning models, improving both the detection and classification of brain tumors. This is achieved by mapping the original intensity values to new values based on their cumulative distribution function (CDF)14,15.
Feature extraction
Feature extraction is necessary to prepare the data for use by the classification models-especially complex tasks. It marries deep learning networks together with texture and shape descriptors, which are learning high-level features for tumor detection. Traditional methods are aimed at the extraction of high-level textural and structural features, which will promote precision and robustness in the detection and classification of brain tumors.
The hybrid nature of the proposed model lies in the combination of both handcrafted and deep learning-based feature extraction techniques. Specifically, we use Local Binary Pattern (LBP), Histogram of Oriented Gradients (HOG), and median intensity features as handcrafted descriptors. In addition to handcrafted descriptors, this study employs feature engineering with deep learning. Specifically, pre-trained CNNs (ResNet50 and AlexNet) are used in evaluation mode to extract deep semantic features, which are then engineered into the hybrid pipeline alongside LBP, HOG, and median intensity features. Unlike end-to-end fine-tuning, this approach treats CNNs as fixed feature extractors and explicitly combines their outputs with handcrafted features, thereby enhancing the diversity and discriminative power of the overall feature space. These diverse feature types are concatenated and passed through a feature selection process (SelectKBest algorithm) before classification using a multilayer perceptron (MLP). This fusion of classical texture/shape descriptors with high-level learned features enables the model to leverage complementary information, improving both accuracy and generalization. Local Binary Pattern (LBP), introduced by Ojala et al. in 1996, is a central tool in the classification of textures and the detection of tumors. The capability of LBP in picking up the fine textural details in CT images is possible because it compares pixels with one another and then checks the presence of texture.
Rotation and illumination are factors that could easily affect brain tumor classification; thus, feature extraction methods must be robust to these factors, which might happen for different positioning or conditions of the patient. The use of LBP satisfies this because it has a consistent and reliable texture descriptor that is invariant to such changes; hence, the features extracted are truly representative of the underlying tissue characteristics. The main challenge in the detection and classification of brain tumors is to accurately differentiate the tumor from normal tissue, especially when the tumor textures are subtle and varied. It is difficult for traditional feature extraction methods to overcome this challenge because they are susceptible to changes in image conditions. While focusing on the frequency of the local structure, LBP captures the local texture pattern at multiple scales; hence, this approach allows for the discrimination of normal versus abnormal tissues even under variable conditions.
LBP extracts textural structures on various scales using a round neighborhood template. The sampling scale R (R > 0) with P (P > 0) sampled neighborhood pixels \({g_p}\)(P = 0,1,., P-1) uniformly dispersed around the circular neighborhood template16. In this research, we used (P, R) = (8, 1) to balance computational cost and feature richness.
In addition to LBP, the Histogram of Oriented Gradients (HOG) method is another critical feature extraction technique employed to enhance diagnostic performance. HOG employs the histogram of gradients to depict the distribution of directions that are essential for finding figures and contours. This includes computation, normalization, and creation, as well as the formation of the final feature vector. HOG is a powerful tool to grab edge and gradient information; therefore, it contributes to tumor detection and classification accuracy17.
In the field of medical imaging analysis, LBP and HOG are known to have brought a very valuable attribute known as robust features, which is achieved using statistical measures such as median intensity. The usefulness of this feature lies in its capability of pinpointing the tumor, thanks to its unique ability to stand out against the background noise and outliers. Here, the median is found to be a powerful force through which the image is mapped to its most representative value and thus can be used to not only reduce excitation, without decreasing performance, classification, and detection accuracy, but also can even improve it18,19.
In addition to traditional methods like LBP, HOG, and median intensity selection, deep learning features play a transformative role in brain tumor detection. Deep learning is a powerful technique in the detection of brain tumors, learning complex patterns with a higher level of abstraction within medical images. Contrary to classic machine learning, it allows the deep learner to draw finer details out of an image that could enable discrimination between abnormalities. This learning ability for the subtle features is highly important in medical image processing due to the possibility that the tumor may present shapes and sizes of all forms.
ResNet50 and AlexNet serve as baseline deep learning models, commonly used in the literature for medical imaging. They have been chosen because they are well-established architectures with proven performance in similar tasks. We have used ResNet50 as shown in Fig. 5, a deep convolutional neural network (CNN) architecture with 50 layers, for feature extraction from CT scan brain images. The ResNet50, in particular, is one of those models that is known for the introduction of residual connections that overcome the vanishing gradient problem in many networks during the extensive layers. ResNet50 is a model that combines depth and computational efficiency, which is the answer to the problem of training difficulties in deep networks, and is known to identify complex brain tumor imaging patterns and their deformities. ResNet50, with its residual structure, gives a new way to define features and thus can distinguish the tumor types and improve the accuracy of classification. That leads to more precise tumor detection, creating better characterizations21. If ResNet50 is employed for feature extraction, it usually only retains the convolutional layers serving as feature extractors by capturing hierarchical features from the input images upon removal of the last fully connected layers. The features extracted by ResNet50 amount to 2048 dimensions and can subsequently be fed into further machine-learning tasks or one or more fully connected layers. This study uses AlexNet for feature extraction. AlexNet is a deep learning architecture widely recognized for its landmark victory in the 2012 ImageNet Large Scale Visual Recognition Challenge (ILSVRC) and has since become one of the foundation models in computer vision (shown in Fig. 623. AlexNet’s deep convolution layers permit it to acquire complex spatial hierarchies, garnering detailed patterns and structures bound to brain cancer diagnostics. A concise architecture is accountable for the extraction of a lot of diversity of features compared to a more convoluted architecture such as VGGNet21. It attests that the AlexNet architecture can be quite helpful in at least many medical applications where either the computational resources are not strong enough or fast inference times are needed.
AlexNet solves problems related to medical imaging through the use of large convolutional layers that are then passed through nonlinear activations. The method itself improves the training process as well as the generalized data, especially in the case of small medical imaging datasets, by dealing with the data complexity and by overcoming the limitations of traditional feature extraction methods. AlexNet boosts feature extraction, thereby upgrading tumor classification, diagnostic precision, and dependable results. It gives the same importance to the accuracy of results and computational efficiency, and is therefore the most suitable for a brain tumor analysis, which consists of feature extraction and practical resource usage.
The Feature extraction part of AlexNet provides 9216 features that reduce error rates on the ImageNet dataset; thus, it is a suitable tool for the detection of intricate patterns in different imaging backgrounds. This work focuses on the convolutional layers, which are responsible for extracting features from input images hierarchically. The 9216 extracted features can further be used in subsequent machine learning tasks or for tumor classification in fully connected layers.
Architecture of the ResNet50 model.
Architecture of AlexNet model.
Feature selection
The key idea is to avoid overfitting and complexity, which is achieved by picking out data with the highest scores among others that are based on tests of statistical relevance.
To address this, we employed a scoring function \(~S~\left( {{f_i},y} \right)\) \({\text{~}}\left( {{\text{SelectKBest~algorithm}}} \right)\) \({\text{~to~evaluate~each~feature~}}{f_i} \in {F_{total}}~\). The function S measures the statistical dependence between features. \({f_i}\) The target variable \(y.A\) common approach is to use metrics such as Mutual Information (MI) or Chi-Square (\({x^2})\left[ {22} \right]\):
where \(p\left( {{f_i},y} \right)\) Is the joint probability distribution, and \(p\left( {{f_i}} \right)p\left( y \right)\) Are the marginal distributions?
To find relevant features useful in prediction and annotation, cross-validation is utilized; only good and useful ones are kept for further research. The primary goal of feature selection is to identify a subset of the most relevant features. \({F_{selected}}\)from a high-dimensional feature space \({F_{total}}=\left\{ {{f_1},{f_2},~ \ldots ,~{f_3}} \right\},~\) where n presents the total number of features. This step addresses issues such as redundancy, irrelevance, and noise, which can otherwise degrade model performance. The need for feature selection arises from challenges such as overfitting, increased computational cost, and reduced generalization ability. Each feature is ranked based on its score, and only the top k features are selected:
In this study, we use k = 10,000 to select the top features, combining Local Binary Pattern (LBP) features (\({n_{LBP}}=65,536\)) and Histogram of Oriented Gradients (HOG) features (\({n_{HOG}}=72,900{\text{~}}\)), resulting in a total initial feature space of (\({n_{total}}=138,436).\) The chosen value of \(k~\)was determined through cross-validation to balance the trade-off between model performance and computational efficiency.
Fusion of deep learning and hand-crafted features
The output from these pre-trained models, combined with selected hand-crafted features and median intensity, is merged into a hybrid feature vector. More precisely, it includes features from ResNet50 and AlexNet, with the chosen features from LBP and HOG, and one feature, median intensity, which is rather high-dimensional. This hybrid vector can represent an image more comprehensively compared to a normal feature vector and provide more insight. For that reason, the result of subsequent analysis tasks performed using this hybrid vector greatly improves their performance. Additionally, this combination enhances classification robustness and accuracy, forming the core of the proposed hybrid approach.
Classification using multilayer perceptron neural network (MLP)
The presence of dense layers in MLPs makes them very effective in the extraction of complex patterns and interactions in high-dimensional data, such as CT scan images; they are good at learning nonlinear relationships and finding intricate patterns, which is necessary to differentiate between tumor types. Although both CNNs and RNNs20 can handle the same task, the MLP performs an excellent trade-off between efficiency and model complexity. Overall, training and implementing MLPs are considerably easier than those of higher complex models, especially when such models are combined with solid and efficient feature extraction techniques. Besides, MLPs grant enough flexibility and adaptability in the majority of classification scenarios that can justify their application in our four-class classification task. The proposed MLP model consists of several layers of neurons, where an activation function has been introduced to bring non-linearity to the network. More specifically, the ReLU is an activation function since it avoids problems like the vanishing gradient problem in the hidden layers, allowing this model to learn complicated patterns in the data. It improves the efficiency of training since, for only positive neuron inputs, it gives an active output, which reduces the convergence of the network and computational complexity. It has batch normalization layers after the standardization of every input in a batch to keep the distribution of inputs constant. This helps in avoiding the problem of internal covariate shift, hence making training easier and more stable, besides better generalization. Then it uses dropout layers to avoid overfitting and has a final layer with Softmax activation that provides an output as a probability distribution for classification. It is designed for a four-class classification problem and provides the probability of each class. The model uses the top selected features as input to classify into four categories. The architecture of this model is presented in Fig. 7, and the hyperparameters of the proposed model are given in Table 4. The proposed hybrid model follows a modular training strategy. Handcrafted features (LBP, HOG, and median intensity) and deep features (from pre-trained ResNet50 and AlexNet) are extracted separately. Deep models are used in frozen mode without fine-tuning. All features are concatenated and reduced using SelectKBest. The final classification is performed using an MLP, trained with cross-entropy loss and the Adam optimizer. This multi-stage pipeline improves generalization and reduces model complexity.
Experimental results evaluation metrics
All experiments were conducted on a personal laptop equipped with an Intel Core i7 CPU, 32 GB RAM, and a 512 GB SSD, running Windows 11. The implementation was done in Python 3.9, utilizing libraries such as TensorFlow, Keras, PyTorch, scikit-learn, OpenCV, and NumPy. Feature extraction using pre-trained CNNs (ResNet50 and AlexNet) was performed in evaluation mode without fine-tuning. Feature extraction using pre-trained CNNs (ResNet50 and AlexNet) was performed in evaluation mode without fine-tuning. While this approach significantly reduces computational requirements and avoids the need for large annotated datasets, it may limit the ability of the networks to fully adapt to the domain-specific characteristics of CT brain images. Fine-tuning could potentially yield higher performance by allowing the pre-trained weights to adjust to modality-specific features, but it was avoided here to reduce complexity and prevent overfitting, given the limited dataset size. Data pre-processing, augmentation, and classification pipeline were developed and executed in this environment. We have used the key metrics to evaluate the performance of the model in its effectiveness in performing its intended work in classification. Accuracy gives the general correctness of the model’s performance; precision shows the percentage of positive instances predicted out of all positive predictions. Recall shows the model identifying all the actual positive instances correctly, while the F1-score gives a decent balance between precision and recall, which latter quite useful when the data is imbalanced. All these metrics are computed with the help of the confusion matrix, which categorizes the predictions into true positives, false positives, true negatives, and false negatives. This matrix is the very basis for each performance metric computation and allows a detailed look at strengths and weaknesses regarding the model’s performance in classification. To ensure robust performance evaluation and minimize bias due to dataset partitioning, a 10-fold cross-validation strategy was employed. The dataset was randomly divided into 10 equal subsets, where in each iteration, nine folds were used for training and one fold for testing. The reported metrics represent the average performance across all folds. This approach is particularly necessary for our study due to the relatively limited dataset size. By using multiple train–test splits, we reduce the risk that the model’s performance is overestimated or underestimated due to a single, potentially unrepresentative split. It also ensures that every sample in the dataset is used for both training and testing, providing a more reliable and generalizable performance estimate. Table 5 presents the prediction results of the proposed approach, which integrates hand-crafted features, ResNet50, and AlexNet with an MLP classifier, demonstrating strong performance across all evaluation metrics, and also shows the base model (hand-crafted features without pre-trained models) across the different types of tumors. Standard deviations (±) are included to indicate performance variability, estimated based on 10-fold cross-validation. Figure 8 illustrates the confusion matrices of the base model, the base model and ResNet50, the base model and AlexNet, and also the hybrid model (combination of base model, ResNet50, and AlexNet). We compare ResNet50, and AlexNet shows better performance in glioma, meningioma, normal tumor, and pituitary tumor. However, the combined model always has higher precision, recall, and overall accuracy compared to individual models. In our proposed model, ResNet50 and AlexNet, it is observed that the graph of the training curve lies above the graph of the validation curve. Also, its minimum validation loss is 0.22 while the minimum training loss is 0.02, as shown in Fig. 9. Figure 9 shows the curves of training and validation losses of the proposed hybrid model based on a pre-trained architecture and a base model. Figure 9 (a) shows a view of training and validation loss using the base model (hand-crafted features). Figure 9 (b) gives a view of a linear decrease of training loss using just the base model and ResNet50. Figure 9 (c) illustrates the same, however, with stronger oscillations for the validation loss, suggesting that the base model and AlexNet generalize a bit worse on this dataset. Hybrid architecture in Fig. 9 (d), already combining base model, ResNet50, and AlexNet with an MLP classifier (hybrid model), presents both low and stable validation losses while showing a smaller gap between training and validation loss that ensures good generalization and possibly robustness for the detection and classification of brain tumors.
The features c proposed model.
Base model, different pre-trained models and proposed hybrid model result in confusion matrices.
The effectiveness of the proposed approach is evaluated using a number of metrics across various tumour types and MLP classifiers.
Training and validation loss of base model, different pre-trained models and proposed hybrid model.
Figure 10 shows accuracy curves of feature extraction, pre-trained models, and the proposed MLP classifier. Training accuracy consistently exceeds validation accuracy, with a peak of 98.71 and 94.56 across epochs. Figure 10 compares the training and validation accuracy of the proposed hybrid model by using different pre-trained architectures. Figure 10 (a) shows training and validation accuracy based on base model. In Fig. 10 (b) (ResNet50 andbase model), training accuracy rapidly increases up to a high value, while the validation accuracy is stabilized around 90% with minor variations, hence the good performance with minor variability. Figure 10 (c) (AlexNet and base model) also shows high training accuracy, but its validation accuracy is highly variant and remains lower than the previous case, hence reducing generalization compared to ResNet50. It can be seen that Fig. 10 (d) stands for the hybrid model, which combines the base model, ResNet50, and AlexNet, and an MLP classifier. It has a high validation accuracy and shows stability in training and validation; moreover, the validation accuracy tends to follow the training curve, implying that the hybrid model has robustness with improved generalization for brain tumor classification. The ROC curve and AUC (Fig. 11) illustrate the model’s performance in distinguishing between the four brain tumor classes: glioma, meningioma, pituitary, and normal brain tissue for the base model, pre-trained models, and hybrid model. Each curve represents the true positive rate versus the false positive rate for a specific class using a one-vs-rest approach. The area under the curve (AUC) for each class exceeds 0.98, with meningioma and pituitary classes achieving perfect AUCs of 1.00 in the hybrid model. This indicates a high level of discriminative power and excellent classification performance across all categories. The near-ideal ROC profiles confirm the model’s robustness and effectiveness in handling multi-class medical image classification. Figure 12 presents performance comparisons of the base model, pre-trained model, and proposed hybrid model, using five metrics: accuracy, precision, sensitivity, specificity, and F1-score. Among the discussed methods, the proposed hybrid model (a combination of base model, AlexNet, and ResNet50 ) performed much better according to all five metrics. In other words, this suggests that the model has gained from the powers of both networks in a better way. More specifically, it achieved higher accuracy, which means better overall classification. Its improved precision and specificity reflect a reduced false positive rate. Besides, with this model, a little more sensitivity is shown, implying it can catch the true positives more reliably. Overall, its balanced performance across metrics makes the combination of AlexNet and ResNet50 much more robust and effective in this classification task. Figure 13 provides a quick comparison of the test accuracy for various classifiers. In this instance, the MLP classifier achieved the highest test accuracy at 94.82%, followed by SVM at 93.93%, KNN at 93.92%, and Random Forest at 87.67%. This visualization effectively highlights the classifier that performs best for the proposed approach. Figure 14 compares the proposed approach, which integrates ResNet50 and AlexNet with an MLP classifier and various optimization algorithms, showing that the Adam optimizer achieves the highest performance score of 94.82. The test accuracy of the proposed model significantly improves after incorporating base models, pre-trained models, with combinations of ResNet50 and AlexNet, yielding higher accuracy than those utilizing only one model or none at all, underscoring the effectiveness of deep learning models (Table 6). Additionally includes estimated standard deviations (±), derived from 10-fold cross-validation, to reflect the consistency of model performance. More importantly, no prior work has focused on CT scan images for the task at hand. Table 7: Comparative performance analysis for brain tumor classification with MRI modality. From this, it can be perceived that the proposed technique, applied to the CT scan modality, reflected the highest accuracy on all the performance metrics used for evaluation. All studies are based on MRI data, while our work focuses on CT scans. This comparison is included only for general context, not direct benchmarking.
Training and validation accuracy of the base model, different pre-trained models and proposed hybrid model.
Multi-Class ROC curve and AUC of the base model, pre-trained models and proposed hybrid model.
Efficiency of different metrics based on base model, different pre-trained models and proposed hybrid model.
Comparative analysis of the proposed hybrid model based on the various classifiers.
Proposed techniques analysis based on different optimizers.
Discussion
The proposed hybrid model outperformed the baseline and intermediate models in both accuracy and robustness. As illustrated in the confusion matrices (Fig. 8), the final hybrid model combining handcrafted features (LBP, HOG, median intensity) with deep features from ResNet50 and AlexNet achieved the highest true positive rates across all four classes, significantly reducing misclassification compared to individual models. Similarly, the training and validation accuracy curves (Fig. 10) show smoother convergence and less overfitting in the hybrid model, reflecting better generalization. This improvement is primarily attributed to the complementary nature of the features: handcrafted descriptors capture localized textural patterns relevant in CT images, while deep CNN features provide high-level semantic abstraction. Moreover, pre-processing techniques such as contrast enhancement and median filtering normalized image variability and reduced noise, contributing to consistent feature learning across varied CT inputs and reducing noise-induced variance, and facilitating better generalization. Data augmentation addresses class imbalance and data scarcity, further contributing to model robustness by exposing the classifier to a wider range of intra-class variations. Feature selection via SelectKBest further reduced dimensionality, minimizing model complexity while preserving discriminative power. The integration of features from multiple domains enriches the feature space, enabling the MLP classifier to learn more discriminative representations. Specifically, ResNet50 brings robust residual connections and deep abstraction capability, allowing the model to capture complex tumor characteristics. AlexNet, though shallower, contributes complementary patterns, often focusing on local features, which seem particularly beneficial for certain tumor classes such as meningioma and normal tissue (as observed in Fig. 11 ROC curves with AUC = 1.00). The hybrid model also demonstrates a smaller gap between training and validation curves (Figs. 9 and 10), suggesting reduced overfitting and improved generalization. Unlike the base model or individual CNN combinations, the hybrid model benefits from both local and global information. The MLP classifier, used as the final decision layer, is well-suited for integrating these heterogeneous feature types due to its flexibility in learning nonlinear relationships. Taken together, the hybrid architecture not only enhances classification accuracy but also achieves high AUC across all tumor classes, reflecting excellent discriminative power. Additionally, as shown in Table V, the proposed hybrid model achieves the highest test accuracy (94.82%), surpassing both the baseline and intermediate models, thus validating the effectiveness of the combined feature approach. These results demonstrate that the hybrid model effectively addresses the inherent complexity of brain tumor classification in CT images and provides a robust and generalizable solution. Meanwhile, deep features from pre-trained CNNs capture high-level spatial and contextual patterns that are learned from large-scale datasets, enhancing the model’s ability to generalize. Together, these components enabled the model to learn more robust and generalized representations, ultimately leading to improved performance. This superiority is quantitatively demonstrated in Fig. 12, where the hybrid model consistently outperforms all other models across key performance metrics, including accuracy, precision, sensitivity, specificity, and F1-score. The proposed hybrid model leverages complementary features and robust pre-processing to enhance generalization, reduce overfitting, and improve multi-class classification accuracy on CT brain tumor data. Its superior performance across key metrics validates the design choices and demonstrates its effectiveness in real-world clinical scenarios. Beyond its technical performance, the proposed automated hybrid model has strong potential for integration into clinical workflows. It can serve as a rapid, objective, and reproducible second-opinion system for brain tumor detection and classification from CT scans, helping radiologists prioritize urgent cases, reduce diagnostic delays, and minimize inter-observer variability. This is particularly valuable in resource-limited settings or emergency departments where MRI may not be feasible. The model’s ability to accurately differentiate glioma, meningioma, pituitary tumors, and normal brain tissue supports more informed treatment planning and monitoring, while its high sensitivity and specificity can facilitate earlier interventions and potentially improve patient outcomes. By automating routine screening and providing consistent decision support, the model could significantly reduce radiologists’ workload and enhance the efficiency of patient care.
Conclusion
This study presented a novel hybrid learning framework for the detection and classification of brain tumors from CT scan images, focusing on four classes: glioma, meningioma, pituitary tumor, and normal brain. The proposed approach integrates handcrafted features (LBP, HOG, median intensity) with deep features extracted from ResNet50 and AlexNet, followed by feature selection using SelectKBest and classification via a Multilayer Perceptron. By combining localized textural descriptors with high-level semantic representations, the model achieved superior performance compared to individual baseline and intermediate models, reaching a maximum test accuracy of 94.82% and achieving high AUC values across all classes. Robust pre-processing steps, including contrast enhancement, median filtering, and data augmentation, contributed to the model’s ability to handle class imbalance, reduce noise-induced variability, and generalize to unseen cases. The hybrid architecture effectively addressed challenges of model complexity and generalization by leveraging complementary feature domains, enabling the classifier to capture both local and global tumor characteristics. The findings highlight the clinical potential of the proposed model as a decision-support tool for radiologists, providing fast, accurate, and automated classification of CT brain images. By enhancing diagnostic confidence and reducing manual workload, this framework can serve as a valuable component in modern computer-aided diagnosis pipelines. With further validation and optimization, it could be deployed in real-world clinical environments to improve diagnostic efficiency and patient outcomes.
Limitations and future work
The proposed hybrid model demonstrated high classification accuracy and robustness, and several limitations remain. First, the dataset used in this study, although diverse in terms of tumor types (glioma, meningioma, pituitary, and normal), was collected from a single online source (Radiopaedia) and may not fully represent the variability seen in multi-institutional clinical settings. This could limit the generalizability of the results. Second, the dataset size is relatively modest, and although data augmentation mitigated class imbalance to some extent, performance on rare tumor types may still be biased. Third, the study relied solely on CT imaging; while this modality is widely accessible, the absence of complementary imaging modalities such as MRI may have limited the ability to capture certain tumor characteristics. Finally, model explainability was not the primary focus of this study, which could hinder its direct clinical adoption. Future work will address these limitations through several directions. First, additional hybrid architectures will be explored, incorporating advanced deep learning backbones (e.g., DenseNet, EfficientNet, Vision Transformers) and a wider range of handcrafted descriptors to evaluate their impact on accuracy and efficiency. Second, multimodal imaging integration (CT + MRI) will be investigated to leverage complementary anatomical and soft-tissue information. Third, explainable AI (XAI) techniques will be developed to provide interpretable decision-making outputs, enhancing trust among clinicians. Fourth, large-scale clinical validation will be conducted across multiple healthcare institutions to assess real-world performance under varied acquisition protocols and patient demographics. Finally, optimization strategies aimed at reducing computational demands will be pursued to enable fast, resource-efficient deployment in routine clinical workflows.
Data availability
Data is provided within the manuscript.
References
Sh. Anantharajan, Gunasekaran, S., Gunasekaran, T., Subramanian, R. & Venkatesh MRI brain tumor detection using deep learning and machine learning approaches. Measure. Sens. 31, 101026 (2024).
Priya, A. & Vasudevan, V. Brain tumor classification and detection via hybrid alexnet-gru based on deep learning. Biomed. Signal Process. Control. 89, 105716. https://doi.org/10.1016/j.bspc.2023.105716 (2024).
Raghavendra, U. et al. Brain tumor detection and screening using artificial. https://doi.org/10.1016/j.compbiomed.2023.107063 (2025).
Nazir, M., Shakil, S. & Khurshid, K. Role of deep learning in brain tumor detection and classification (2015 to 2020): a review. Comput. Med. Imaging Graph. 91, 101940. https://doi.org/10.1016/j.compmedimag.2021.101940 (2021).
Nabizadeh, N. & Kubat, M. Brain tumors detection and segmentation in MR images: Gabor wavelet vs. statistical features. Comput. Electr. Eng. 45, 286–301. https://doi.org/10.1016/j.compeleceng.2015.02.007 (2015).
Das, P. & Das, A. Multi-scale cross spectral coherence and phase spectral distribution based measurement in non-subsampled shearlet domain for classification of brain tumors. Expert Syst. Appl. 247, 123329. https://doi.org/10.1016/j.eswa.2024.123329 (2024).
Rastogi, D., Johri, P., Tiwari, V. & Elngar, A. Multi-class classification of brain tumor magnetic resonance images using multi-branch network with inception block and five-fold cross validation deep learning framework. Biomed. Signal Process. Control. 88, 105602. https://doi.org/10.1016/j.bspc.2023.105602 (2024).
Rahman, T. I slam Brain tumor detection and classification using parallel deep convolutional neural networks. Measurement: Sens. 26, 100694. https://doi.org/10.1016/j.measen.2023.100694 (2023).
Ravinder, M. et al. Enhanced brain tumor classification using graph convolutional neural network architecture. Nat. Portfolio. https://doi.org/10.1038/s41598-023-41407-8 (2025).
Zulfiqar, F., Bajwa, U. & Mehmood, Y. Multi-class classification of brain tumor types from MR images using efficientnets. Biomed. Signal Process. Control. 84, 104777. https://doi.org/10.1016/j.bspc.2023.104777 (2023).
Reyes, D. & Sánchez, J. Performance of convolutional neural networks for the classification of brain tumors using magnetic resonance imaging. Heliyon 10, e25468. https://doi.org/10.1016/j.heliyon.2024.e25468 (2024).
Otair, M. Exclusive median filter (EMF) for enhancing image quality. AIP Conf. Proc. (2023).
Chalghoumi, S. & Smiti, A. Median filter for denoising MRI: literature review. In presented at the International Conference on Decision Aid Sciences and Applications (DASA) (2022).
Behera, D. Enhancement of images using various histogram equalization Techniques. IOSR J. Electron. Commun. Eng. (IOSR-JECE) 8, 38–41 (2013).
Yelmanov, S., Hranovska, O. & Romanyshyn, Y. A new approach to the implementation of histogram equalization in image processing. In 3rd International Conference on Advanced Information and Communications Technologies (AICT) (2019).
Sh. Lan, X. et al. Amulti-channel framework based local binary pattern with two novel local feature descriptors for texture classification. Digit. Signal Proc. 140, 104124 . https://doi.org/10.1016/j.dsp.2023.104124 (2023).
Sharma, A. et al. HOG transformation based feature extraction framework in modified Resnet50 model for brain tumor detection. Biomed. Signal Process. Control. 84, 104737. https://doi.org/10.1016/j.bspc.2023.104737 (2023).
Gou, F., Wu, J. & Zhang, Y. A tumor MRI image segmentation framework based on Class-Correlation pattern aggregation in medical Decision-Making system. Mathematics 11 (5), 1187. https://doi.org/10.3390/math11051187 (2023).
Gonzalez, R. C. & Woods, R. E. Digital Image Processing 4th edn (Pearson Education, 2018).
Ji, Z. et al. Generalized rough fuzzy c-means algorithm for brain MR image segmentation. Comput. Methods ProgramsBiomed. 108, 644–655. https://doi.org/10.1016/j.cmpb.2011.10.010 (2012).
Lavanya, N. & Nagasundaram, S. A survey of brain tumor detection and classification based on Machine, Deep Learning and CNN based Transfer Learning techniques. In Proceedings of the 5th International Conference on Inventive Research in Computing Applications (ICIRCA IEEE Xplore Part Number: CFP23N67-ART; ISBN: 979-8-3503-2142-5 (2023).
Rodriguez, A. & Laio, A. Feature selection via mutual information: new theoretical insights. arXiv preprint arXiv:1907.07384. Retrieved from arXiv.org (2019).
Krizhevsky, A., Sutskever, A. & Hinton, I. ImageNet classification with deep convolutional neural networks. Adv Neural Inf. Proces Syst 2012, 25 (2012). https://papers.nips.cc/paper/2012/hash/c399862d3b9d6b76c8436e924a68c45b-Abstract.html.
Sarkar, A., Md. Maniruzzaman, M., Ashik, M. & Ahmad An effective and novel approach for brain tumor classification using AlexNet CNN feature extractor and multiple eminent machine learning classifiers in MRIs. J. Sens. https://doi.org/10.1155/2023/122461919, 1224619 (2023).
Khairandish, M., Sharma, M., Jain, V., Chatterjeed, J. & Jhanjhi, N. Z. A hybrid CNN-SVM threshold segmentation approach for tumor detection and classification of MRI brain images. IRBM 43 (Issue 4), 290–299. https://doi.org/10.1016/j.irbm.2021.06.003 (2022).
Zheng, N., Zhang, G., Zhang, Y. & Rashid, F. Brain tumor diagnosis based on Zernike moments and support vector machine optimized by chaotic arithmetic optimization algorithm. Biomed. Signal Process. Control. 82, 104543. https://doi.org/10.1016/j.bspc.2022.104543 (2023).
Reddy, K. & Dhuli, R. Segmentation and classification of brain tumors from MRI images based on adaptive mechanisms and ELDP feature descriptor. Biomed. Signal Process. Control. 76, 103704. https://doi.org/10.1016/j.bspc.2022.103704 (2022).
Kumar, K. S., Prasad, A. Y. & Metan, J. A hybrid deep CNN-Cov-19-Res-Net transfer learning architype for an enhanced brain tumor detection and classification scheme in medical image processing. Biomed. Signal Process. Control. 76, 103631. https://doi.org/10.1016/j.bspc.2022.103631 (2022).
Raghuram, B. & Hanumanthu, B. Brain tumor image identification and classification on the internet of medical things using deep learning. Measure. Sens. 30, 100905 (2023).
Hafiz, M., Arfan, J., Tehreem, M. & Sheeraz, A. A novel approach to classify brain tumor with an effective transfer learning based deep learning model. Braz. Archiv. Biol. Technol. 67, e24231137 (2025).
Kar. Satrajit, K. & Pawan, M. B. T. C. N. Multimodal brain tumor classification from CT and MRI scans using deep neural network with multi-head attention mechanism. Med. Novel Technol. Devices. 27, 100382 (2025).
Alamin, T. et al. An efficient deep learning model to categorize brain tumor using reconstruction and fine-tuning, reconstruction and fine-tuning. Expert Syst. Appl. 230, 120534 (2023).
Takwan, R. & Saiful, I. MRI brain tumor detection and classification using parallel deep convolutional neural networks. Measure. Sens. 26, 100694 (2023).
Sh. Kumar, N. et al. Brain tumor classification modified ResNet50 model based on transfer Learning, Biomed. Signal Process. Control 86, 105299 (2023).
Fatima, Z., Usama, B. & Yaser, M. Multi-class classification of brain tumor types from MR images using efficientnets. Biomed. Signal. Process. Control. 84, 104777 (2023).
Muhammad, A. et al. Brain tumor classification utilizing deep features derived from high-quality regions in MRI images. Biomed. Signal Process. Control. 85, 104988 (2023).
Loganayagi, T., Meesala, S., Balajee, M. & Venkata, R. T. Hybrid deep Maxout-VGG-16 model for brain tumour detection and classification using MRI images. J. Biotechnol. 405, 124–138 (2025).
Amreen, B. & Yung, B. A lightweight multi-path convolutional neural network architecture using optimal features selection for multiclass classification of brain tumor using magnetic resonance images. Results Eng.. 25, 104327 (2025).
Acknowledgements
This research is funded by the European University of Atlantic.
Author information
Authors and Affiliations
Contributions
Roja Ghasemi .Naveed Islam. Farhan Amin, Samin Bayat and Muhammad Shabir wrote the main manuscript text, and Shahid Rahman . Isabel de la Torre. Ángel Kuc Castilla. Debora Libertad Ramírez Vargas García prepared figures 1-3. All authors reviewed the manuscript.”
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ghasemi, R., Islam, N., Bayat, S. et al. Detection and classification of brain tumor using a hybrid learning model in CT scan images. Sci Rep 15, 35085 (2025). https://doi.org/10.1038/s41598-025-18979-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-18979-8