Abstract
Host resistance evolution to natural enemies depends on genetic variation, symbiont associations, and trade-offs with host´s fitness traits. We selected isofemale lineages of Myzus persicae to investigate biological traits and molecular markers associated with aphid defensive responses to the parasitic wasp Diaeretiella rapae. The parasitization of M. persicae lineages by D. rapae ranged from 43 to 76%. Rickettsia was the predominant secondary symbiont. Six lineages were selected according to D. rapae parasitization rate (LP = low parasitization; HP = high parasitization) and Rickettsia association. Life history traits differed among M. persicae lineages. However, the association of adaptive costs with reduced susceptibility to D. rapae parasitization was not consistently observed. Rickettsia infection did not explain the enhanced aphid defense, but it did increase aphid fecundity. Comparative small structural variants analyses of M. persicae lineages indicated that approximately 6% of SNPs and 49% of InDels were unique to each aphid lineage or shared among LP or HP lineages. Only 1.7% of SNPs and 0.91% of InDels influenced genes involved with aphid immune response. Further exploration of non-coding variants and additional omics-related approaches are needed to fully characterize the mechanisms behind the differential parasitization success of M. persicae lineages by D. rapae.
Similar content being viewed by others
Introduction
Parasitoids use completely different life strategies when compared to true parasites as they kill their hosts upon completion of their development, putting a much stronger selection pressure for the evolution of defensive traits on their hosts1,2,3,4,5. The successful exploitation of a wide range of hosts by parasitoids is partially due to the diverse strategies parasitoids evolved for host exploitation6. But host defensive strategies also lead to parasitoid avoidance and parasitization escape7,8,9. Host populations that are maintained under strong selective pressure from parasitoids can face a bottleneck in their genetic composition due to the elimination of susceptible genotypes that are successfully parasitized from the total genetic pool available in the population,3,10,11,12,13 including those represented by genotypes associated with microbial symbionts14,15. Host-parasitoid population densities are known to fluctuate cyclically, keeping both populations in equilibrium and rarely leading to extinction events16,17,18. Thus, the survival of host genotypes that escape parasitization may drive selection for hosts that are resistant to parasitization19,20,21.
Diaeretiella rapae (MacIntosh) (Hymenoptera: Braconidae), a parasitoid of more than 60 species of aphids22, including Myzus persicae (Sulzer) (Hemiptera: Aphididae) is an example of a parasitoid exerting strong selective pressure on its hosts. Myzus persicae is a polyphagous pest of great economic importance, with excellent developmental capabilities on hundreds of plant species in more than 50 families23. Its economic importance is based on the ability to cause direct damage to plants in conjunction with its feeding habits, which result in host plant dehydration, loss of shoots and retardation of vegetative growth24. In addition, aphids can cause indirect damage by transmitting viruses and promoting black mold growth by excreting honeydew24. Aphid infestations of agricultural crops often require the implementation of pest control measures, and a set of new management strategies has been addressed to sum up to biocontrol methods to reduce the often-used synthetic chemical insecticides and their undesired side-effects25.
D. rapae is a key parasitoid in the biological control of aphids, including M. persicae, and exerts a strong selective pressure on aphid populations, driving the evolution of host defense mechanisms and attack strategies. Previous studies on M. persicae—D. rapae interactions have mainly focused on verifying the influence of the host plant on aphid suitability26 and on parasitoid attraction27, the effects of the secondary symbiont Rickettsiella viridis in the rate of parasitization28, and the side-effects of insecticides in this host-parasitoid interaction29,30. There have been also studies dedicated to compare D. rapae efficacy on aphid host species with different host plant ranges31. However, the genetic basis of the defensive mechanisms, the adaptive costs of resistance to parasitization and the role of secondary symbionts as defensive symbionts in this host-parasitoid interaction are seldom reported.
These gaps in knowledge highlight the complexity of the evolutionary arms race between aphids and their parasitoids. Co-evolution between hosts and parasitoids arises because of interactions throughout their evolutionary history, as hosts face selection pressure due to exposure to parasitoid-derived virulence factors whilst, at the same time, parasitoids face selection pressure from host defenses. As predicted by the red queen theory, each adaptation developed by one species faces a counteradaptation of the interacting species, such that the survival of both species depends on the continuous development of defense and attack strategies (arms race)11,32,33,34,35. Microorganisms can modulate their host phenotype36,37, and secondary symbionts have shown to modify their host response to parasitoid attack, with possible implications in the evolution of parasitoid virulence38. Defensive symbionts can affect host successful parasitization by resource limitation, such as lipids39, and by the production and release of metabolites toxic to natural enemies40,41.
Despite advances in the understanding of defense mechanisms against natural enemies, the physiological and behavioral strategies of hosts and their interactions with secondary symbionts are still poorly understood. Hamiltonella defensa and its associated bacteriophage are among the best-known defensive secondary symbionts. Phage-infected Hamiltonella defensa can produce toxins such as Shiga-like, CdtB and YD-repeat, which increase immature parasitoid mortality and strengthen host resistance42,43,44. Serratia symbiotica is another relevant secondary symbiont that affects successful parasitization by interfering with the production of plant volatiles involved in parasitoid attraction45. In aphids, other secondary symbionts (e.g., Rickettsia, Spiroplasma, Wolbachia) are also present in natural environments, but their role as defensive symbionts against natural enemies is still poorly understood, and an increase in the knowledge of parasitoid-host-symbiont interactions can open new perspectives for the development of more efficient biological control programs46.
Host-parasitoid interactions are frequency-dependent, which can be observed more strongly in laboratory than in field studies47. The frequency-dependent interaction combined with genetic variation allows interacting species to co-evolve2. Parasitoids are selected to evade defense mechanisms of common host genotypes, contributing to the selection of hosts with rare resistant genotypes48. This process is difficult to observe in field populations since the detection of resistant individuals depends on the host encounter and on the genotype of the natural enemy. Host genotypes with a lower ability to evade natural enemies’ encounters are more frequently attacked. However, parasitoid genotypes are subject to similar selection pressures, resulting in a selection process that occurs without the fixation of extreme host or parasitoid genotypes48,49. In laboratory experiments, genotypes with greater and/or lesser ability to avoid parasitoid attack face similar rates of parasitoid encounter under controlled abiotic and biotic conditions and similar patch structures, allowing the study of host/parasitoid genotypes with selected responses47.
The genetic variability that results in different host responses to parasitoid attack and parasitoid phenotypes with different levels of success in host parasitization can arise from mutation, increased gene flow, genetic drift, inbreeding depression, and selection50,51. The main and fastest way to generate genetic variability is through sexual reproduction, which allows for the perpetuation of mutations in the population, as well as genetic recombination between chromosomes52,53. Aphids developing in temperate regions exhibit seasonal reproductive polyphenism, alternating between asexual and sexual reproduction to produce eggs that will diapause in winter54,55,56. However, under tropical conditions, aphid species such as Myzus persicae reproduce exclusively by ameiotic (apomictic) parthenogenesis, in which chromosome division occurs by mitosis. In apomixis reproduction, the resulting offspring are genetically identical to their mothers, and the sources of genetic variation to promote phenotypic expression are now limited to rare genomic events, such as mutations, chromosomal rearrangements and mitotic recombination, and/or interaction with endosymbionts57,58,59.
Chromosomal rearrangements and interactions with symbionts have been shown to serve as sources of genetic variation leading to the manifestation of insecticide resistant, parasitoid defense and host plant adapted phenotypes14,60,61,62,63. Thus, both chromosomal and extra-chromosomal variation can influence the evolution of host defense mechanisms against natural enemies, which may invariably have associated adaptive costs64,65,66,67. Chromosomal rearrangements often result from the emergence of structural variants that can arise through various cellular mechanisms, including DNA replication and repair68. These variants can be broadly categorized into five classes: deletions, duplications, insertions, inversions, and translocations69. Structural variants introduce variability in gene copy number, position, orientation, and occasionally a combination of these effects70. As a result, structural variants drive genotypic change and facilitate the manifestation of diverse phenotypes.
In this context, we investigated the interaction between M. persicae and its parasitoid D. rapae, considering genetic and symbiotic factors that may influence the host response to parasitization. We sought to answer the following questions: (1) is there variation between isofemale lineages of M. persicae in their resistance to parasitism by D. rapae?; (2) does the presence of secondary symbionts influence the success of parasitism?; (3) are there biological differences between isofemale strains of M. persicae associated with the presence of secondary symbionts and their resistance to parasitism?; and (4) do small structural variants (SVs) in the M. persicae genome play a role in host adaptation and response to parasitism? Our central hypothesis is that the differential resilience to parasitism by D. rapae among M. persicae lineages is associated both with the presence of secondary symbionts and with structural variations in the genome that may modulate the host response to parasitoid attack.
Results
Secondary symbionts associated with Myzus persicae isofemale lineages
We evaluated the presence of secondary symbionts commonly associated with aphids in isolated infections and co-infections in 14 isofemale lineages of M. persicae. Diagnostic PCR analysis based on the 16S rRNA gene of the most common symbiotic bacteria harbored by aphids detected secondary symbionts associated with 11 (nearly 79%) out of the 14 M. persicae isofemale lineages subjected to parasitization by D. rapae. Spiroplasma (29%) and Rickettsia (71%) were the only secondary symbionts detected, occurring either in single- [Spiroplasma—9% (one lineage); Rickettsia—64% (seven lineages)] or double-infected aphids (27%—three lineages) (Fig. 1A).
Venn diagram showing the number of isofemale lineages of Myzus persicae with isolated infection or co-infection by the secondary symbionts Rickettsia and Spiroplasma (A) and mean parasitization (%) (±) of isofemale lineages of Myzus persicae by Diaeretiella rapae (Red dotted line = 75% percentile; black dotted line = 25% percentile) (Green stars indicate lineages selected for biological parameters and molecular markers analysis) (B).
Response of isofemale Myzus persicae lineages to parasitization by Diaeretiella rapae
We tested the response of 14 isofemale lineages of M. persicae to parasitization by D. rapae. The observed success of parasitization of M. persicae isofemale lineages by D. rapae varied between 43 and 76%, with five lineages with successful parasitization values within the fourth quartile (above 75% percentile) and another five within the first quartile (below 25% percentile) (Fig. 1B).
Rickettsia infections (r +) were detected in isofemale lineages with parasitization values within both quartiles (first quartile: Iso13 and Iso14; fourth quartile: Iso2 and Iso3), whereas Spiroplasma infections (s +) were detected in an isofemale lineage with a parasitism value within the first quartile (Iso12), and co-infections were detected in isofemale lineages with a parasitism rate only in the fourth quartile (Iso1 and Iso5).
Based on the scenario of infection by secondary symbionts in the two quartiles representing the lowest (first quartile) and highest (fourth quartile) values of successful parasitization, three isofemale lineages from the first [low parasitization (LP) group—Iso10LP/r-, Iso11LP/r- and Iso14LP/r+] and fourth [high parasitization (HP) group—Iso2HP/r+, Iso3HP/r+ and Iso4 HP/r-] quartiles were selected for further comparative analysis of life history traits and fertility life table determination to verify the association of fitness costs with the defensive response of M. persicae to parasitization by D. rapae (Fig. 1B).
The consequences of the differential response to parasitization by Diaeretiella rapae and the association with Rickettsia on life history traits in isofemale lineages of Myzus persicae
We investigated whether isofemale lineages of M. persicae selected for high and low parasitization by D. rapae, as well as infection or not by the secondary symbiont Rickettsia impose adaptive costs by evaluating their life history traits and fecundity table. The survival of nymphal stage differed among the selected lineages of M. persicae (LR = 15.94, df = 5, p = 0.007). The lowest probability of reaching adulthood was observed for lineage Iso10LP/r- (probability of reaching adulthood = 0.776) and the highest probability of reaching adulthood was identified for lineage Iso11LP/r− (probability of reaching adulthood = 0.92) (Fig. 2A, Table 1). We also detected differences in the developmental time required for nymphs to become adults (χ2 = 63.10, df = 5, p < 0.001). The development time of Iso14LP/r+ was the longest observed among the lineages (7.4 ± 3.5 days), differing from all other lineages except Iso4HP/r-. The development time of Iso2HP/r+ and Iso11LP/r- was intermediate and differed from the development time of Iso14LP/r+ and Iso4HP/r-. Iso3HP/r+ and Iso10LP/r- were among those with the shortest developmental time (5.7 ± 2.5 and 5.8 ± 3.3 days, respectively), differing from that observed for Iso4HP/r- and Iso14LP/r+ (Fig. 2B, Table 1).
Biological parameters evaluated for selected lineages of Myzus persicae with high (Iso2HP/r+, Iso3HP/r+, Iso4HP/r-) and low (Iso10LP/r-, Iso11LP/r-, Iso14LP/r+) parasitization by Diaeretiella rapae. (A) Kaplan–Meier estimates of survival curves using Cox proportional hazards models, p < 0.05. (B) Immature development time (days ± se). Treatments followed by different letters are statistically different (p < 0.05). Green bars = indicate the lineages of the group with lower parasitism. Blue bars = represent the isolines with higher parasitism. (C) Daily female survival (%) and relative accumulated observed fecundity (%). (D) Heatmap of reproductive life table data and associated dendrograms obtained from hierarchical clustering based on Euclidean distance using Ward’s method. The color scale indicates the proximity of the mean of each isoline to the total mean of each measured parameter (white = low proximity; red = high proximity).
The tibia size of females, parameter used to determine the relative size of the adult aphid, also differed among the selected lineages (F5, 112 = 3.62; p = 0.0044). The tibia of adult females from Iso2HP/r+ were larger than those from Iso4HP/r- and Iso14LP/r+ which had the smallest size of the tibia. The tibia size in Iso2HP/r+, Iso4HP/r- and Iso14LP/r+ did not differ from the other lineages (Table 2). No differences in adult longevity were observed (F5, 174 = 0.6167; p = 0.6872), while differences in female fecundity were detected only between Iso10LP/r- and Iso11LP/r- (F5, 174 = 2.53; p = 0.027) (Table 2). All selected lineages appeared to have a similar rhythm of offspring production, with 80% of the total offspring being produced between days 13 and 15 of adulthood (Fig. 2C). Most lineages reached 80% of accumulated fecundity with almost 80% survival of females (Fig. 2C).
Analysis of fertility life table parameters of M. persicae lineages revealed differences in their net reproductive rate (Ro) (F5, 174 = 5.425; p < 0.001). Lineage Iso11LP/r- had the lowest Ro values, differing from Iso2HP/r+ (p = 0.048), Iso10LP/r- (p < 0.001) and Iso14LP/r+ (p = 0.038), but not from Iso3HP/r+ and Iso4HP/r-, which had intermediate values (Table 3). The intrinsic growth rate (rm) (F5, 174 = 7.1119; p < 0.001) and the finite growth rate (λ) (F5, 174 = 7.0172; p < 0.001) were higher for lineages Iso2HP/r+, Iso3HP/r+, Iso10LP/r-, and Iso14LP/r+ when compared to Iso11LP/r-. Rm and λ obtained for Iso3HP/r+ were also higher than those for Iso4HP/r- (Table 3). The time between generations (T) observed for isofemale lineage Iso3HP/r+ was shorter than those for Iso4HP/r- (p = 0.037), Iso10LP/r- (p < 0.001), and Iso14LP/r+ (p = 0.025), but like that obtained for Iso2HP/r+ and Iso11LP/r-. The doubling time (Dt) for Iso11LP/r- (3.5 d) was longer than that for all other lineages (p < 0.001) except Iso4HP/r- (Table 3). The doubling time (Dt) of Iso4HP/r- was also like that of lineages with intermediate Dt values, but different from that of Iso3HP/r+ (Table 3).
The results of the hierarchical clustering analysis based on Ro, Rm, T, TD and λ values revealed two major clusters, the first composed by the Iso4HP/r- and Iso11LP/ r- isofemale lineages and the second composed by the other lineages (Iso2HP/r+, Iso3HP/r+, Iso10LP/r- and Iso14LP/r+). Iso4HP/r- and Iso11LP/r- grouped in a well-defined cluster. Iso2HP/r+ and Iso14LP/r+ resolved in the innermost cluster of a large cluster also containing Iso3HP/r+ and Iso10LP/r- (Figs. 2D and S2). Iso3HP/r+ was the first to bifurcate from the cluster containing Iso10LP/r-, Iso2HP/r+, and Iso14LP/r+ because it had the highest values of Rm and λ, and the lowest values of Dt and T. Iso10LP/r- was the second to bifurcate out based on the highest values of Ro and T. Iso2HP/r+ and Iso14LP/r+ resolved together as they shared similar values for all fertility life table parameters (Fig. 2D).
The life history traits and fecundity table suggest that different mechanisms may be involved in the differential response to parasitism and infection by Rickettsia, considering that it was not possible to detect a pattern between lineages belonging to the HP or LP group and r + or r-.
Relation of small nucleotide variants in the genome of isofemale lineages of Myzus persicae selected for differential response to parasitization
We further investigated whether the difference in response to parasitization in selected isofemale lineages of M. persicae could be associated with small structural variants and SNPs in the genome of these isofemale lineages compared to a reference genome. Figure 3 summarizes the main results described so far (parasitism and fecundity) in relation to the detection of small structural variants (InDels) and SNPs.
Graphical summary results with parasitism, fecundity and number of variants (InDels and SNPs) in Myzus persicae isofemale lineages with high (Iso2HP/r+, Iso3HP/r+, Iso4HP/r-) and low (Iso10LP/r-, Iso11LP/r-, Iso14LP/r+) parasitization by Diaeretiella rapae.
DNA sequencing data
Illumina sequencing of the six selected M. persicae lineages resulted in a total of 140.6 million raw reads, ranging from 19.3 to 26.4 million reads per library with very high-quality scores (Table S1). The number of duplicated reads ranged from 19 to 25.5%, and only 10,806 to 23,238 bp reads per library failed the quality filters applied (Table S1). The number of paired reads ranged from 19 to 26 million/library, resulting in 138.5 million raw reads with 30% GC content (Table S1).
SNPs and InDels variant calling
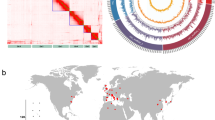
A total of 1,048,661 single nucleotide polymorphisms (SNPs) and 423,303 insertion-deletions (InDels) were detected, with 290,628 SNPs and 64,752 InDels surviving the filtering protocols used. Around 80% of all SNPs and InDels were associated with the intergenic and intron regions of the aphid genome, with only ~ 6% associated with coding regions (Figure S3). Most SNPs (85.1%) (Fig. 4A) and nearly half of InDels (60.8%) (Fig. 4B) were common to all lineages. The number of private SNPs detected in each lineage was quite variable, ranging from 737 in lineage Iso11LP/r- to 179 in lineage Iso3HP/r+ (Fig. 4A). A high number of specific SNPs was associated with each group of lineages, with 135 specific SNPs in the LP group and 117 in the HP group (Fig. 4A). A very high number of InDels was specifically detected in each lineage, ranging from 1,404 in lineage Iso14LP/r+ to 2,621 in lineage Iso4HP/r- (Fig. 4B). Lineages of the LP group shared 79 specific InDels, while those from the HP group 222 (Fig. 4B).
Overview of the SNPs (A) and InDels (B) that survived the filtering protocols, common to all the Myzus persicae isofemale lineages studied (Iso2HP/ r + , Iso3HP/r + , Iso4HP/r-, Iso10LP/r-, Iso11LP/r- and Iso14LP/r +), shared by two or more lineages, or unique to each group (high and low parasitism) or each lineage.
Nearly 1.7% SNPs and 0.91% InDels were classified as resulting in low, moderate, and high (Figure S4) according to their impact on gene function (Figures S5 and S6), and were found located with promoter 2 kb, 5’UTR, 3’UTR, cds, and intronic genomic regions, in which most SNPs and InDels are located in the promoter and codon regions (cds) (Fig. 5). Among the SNPs and InDels that were shared between lineages belonging to the LP and lineages belonging to the HP group, only seven SNPs and six InDels were classified as small structural variants with high or moderate impact in lineages of the HP group, while seven SNPs in those of the LP group (Table 4).
Overview of the moderate and high SNPs (pink color) and Indels (purple color) by genomic regions present in the genome of Myzus persicae lineages with high (Iso2HP/r+, Iso3HP/r+, Iso4HP/r-) and low (Iso10LP/r-, Iso11LP/r-, Iso14LP/r+) parasitization by Diaeretiella rapae.
Functional annotation led to the identification of GO and KO terms for only 7.5% and 13.2% of the detected structural variants and SNPs with a high or moderate impact, with the majority classified into replication, recombination and repair (L), carbohydrate transport and metabolism (G), transcription (K) and post-translational modification, protein turnover, chaperones (O) COG classes (Tables S2, S3, S4, S5, Fig. 6). In the HP group, only one high InDel and two moderate SNPs were detected, the InDel promoting the deletion of an adenine nucleotide being associated with the g2278.1 gene, which codes for the p52IPK protein, a suppressor of the p58IPK inhibitor of protein kinase RNA-activated (PKR), that may be associated with cellular responses to stress inducers. The SNPs resulting in the change of cytosine or guanine to thymine are related to the g15384 and g22415 genes, both of which code for a zinc-finger MYM-type protein, which may be related to immune response mechanisms. In the LP group, only one moderate SNP was found associated with the gene g19286, which encodes a nephrin-like protein, that does not appear to be associated with the phenotypes observed.
The functional COG classification of genes associated with high and moderate SNPs (A) and InDels (B) detected in Myzus persicae lineages selected for high (Iso2HP/ r + , Iso3HP/r + , Iso4HP/r-) and low (Iso10LP/r-, Iso11LP/r, Iso14LP/r +) parasitization by Diaeretiella rapae, or unique to each group (high and low parasitism) (C).
Enrichment analysis of GO terms for the high and moderate SNPs and InDels resulted in different top five terms in each category, with the most represented having catalytic or hydrolase activities (Fig. 7). KEGG pathway analysis identified InDels in 33 and SNPs in 20 pathways. Twelve genes with small structural variants specifically detected in LP lineages were associated with KEGG pathways, while no association was established for those of the HP group. KEGG pathway analysis also demonstrated that although small structural variants were associated specifically with each individual lineage or to each group (LP or HP) of lineages, all pathways identified for the LP group were also represented in at least one lineage of the HP group (Table S6).
Top five Gene Ontology (GO) functional analysis of genes associated with high and moderate SNPs (A) and InDels (B) unique to the high (Iso2HP/ r + , Iso3HP/r + , Iso4HP/r-) or low parasitization (Iso10LP/r-, Iso11LP/r, Iso14LP/r +) group or to each individual lineage of Myzus persicae selected for their differential response to parasitization by Diaeretiella rapae.
Discussion
Isofemale lineages of M. persicae exhibited high phenotypic variation in response to parasitization by D. rapae. Phenotypic plasticity in M. persicae has also been demonstrated for body color and size71, pathogen vectoring capacity72, and susceptibility to biotic and abiotic stressors71,73. However, we did not expect to observe such high variation within isofemale lineages under controlled parasitoid exposure conditions, as the offspring of parthenogenetic aphid females are clones of their mothers, once parthenogenesis occurs by apomixis64. Previous studies with clonal lines of the aphid Acyrthosiphon pisum have shown contrasting variability in resistance to parasitism by Aphidius ervi, resulting in the selection of lines that are completely resistant and others that are highly susceptible to parasitoid attack74,75, although the variability in response observed between individuals of the same line was not as high as the observed in our study.
The high variability observed in some isofemale lineages indicates the presence of factors that reduce genetic fidelity among clonal daughters. There are several examples in aphid species that demonstrate selection and evolution of clonal lines mainly in adaptation to host plants71, although several mechanisms have been reported to produce genetic variation in clonal aphids, such as DNA mutations76, chromosomal rearrangements, mitotic recombination, and interactions with symbionts58,59,64. Some of the events that generate genetic variability should not be as common in obligate asexual (anholocyclic) aphid lineages, lineages formed by asexual females that produce asexual females, as they are in holocyclic lineages, where sexual reproduction occurs during at least part of their life cycle71,77. Some studies have shown that the main source of clonal variation in response to parasitization is due to interaction with the secondary symbionts Hamiltonella defensa and Regiella insecticola. The host association with these symbionts significantly increased resistance of clonal host lines to parasitoid attack78,79.
In the isofemale lineages in our study, we did not detect infections with the most common defensive symbionts (H. defensa and R. insecticola), but we did examine the influence of Rickettsia on host fitness traits and host defense against parasitization. Our data was not conclusive on the role of Rickettsia as a defensive secondary symbiont, since Rickettsia infections were detected in aphid isofemale lineages belonging to the low and high parasitization groups. In addition, similar to a previous study with A. pisum, the most resistant isofemale lineages in our study exhibited levels of defense (~ 47—57% survival) like A. pisum lineages infected with defensive symbionts (~ 35–100%), leading us to question whether genetic variation in the strains is associated with the phenotypic variability observed80,81.
Our study showed that all isofemale lineages of M. persicae carry specific small genomic structural variants representing specific molecular markers from its interaction with the environment, which could explain the phenotypic differences observed. Structural variants, such as SNPs and InDels, have been previously linked to phenotypic plasticity and adaptive responses in aphids and other insects82,83,84,85. Despite the small number of structural variants that can induce high and moderate impact in gene functioning in LP and HP phenotypic groups of M. persicae, the functional annotation of one InDel and three SNPs revealed their association with stress signaling pathways and developmental processes that can be further explored to explain the observed phenotypic responses. Similar associations have been observed in studies investigating genomic adaptations in insects, where structural variants influence host resistance against parasitoids, wing plasticity and insecticide resistance82,83,84,85.
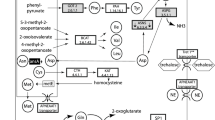
One of the InDels identified in lineages of HP group is associated with the g2278.1 gene encoding for the p52IPK protein, a suppressor of the p58IPK inhibitor of protein kinase RNA-activated (PKR). PKRs are key regulators of cellular response to stress inducers by activating other pathways of stress response86. Several of the activated stress responses by the PKR pathway are involved with humoral and cellular immune responses (e.g., JNK, p38, and NFkB). p38 is a mitogen activated protein kinase (MAPK) and regulates gene expression leading to antimicrobial peptide production87,88. MAPK also plays a key role in the cascade activation of the humoral and cellular immune response in insects leading to the encapsulation and melanization of wasp eggs in parasitized hosts89,90,91,92,93. Recent data has shown the efficacy of the antimicrobial peptides in activating the Toll/Imd immune pathways to protect silkworm larvae from dipteran parasitoids successful development94. Thus, we believe the InDel detected in the p52IPK gene could lower the capacity of the HP group of aphid lineages to build an immune response through the protein kinase cascade activation of cellular and humoral immune responses due to the reduced suppression of p58IPK, which in turn would result in a higher inhibition of PKRs by p58IPK.
SNPs detected in two genes (g15384.t1 and g22415.t1) in the HP group lineages encode zinc finger proteins of the MYM family (ZFP-MYM), which are also associated with the immune response. ZFP-MYMs are multifunction proteins with regulatory roles in SUMOylating by acting as SUMO-binding proteins, which directly interferes with the development, maturation, and activation of immune cells, precursors, and effectors95,96,97,98. Thus, alterations in the protein sequence of ZFP-MYMs would affect their binding capacity to target proteins would dysregulate the development and differentiation of immune cells and impair the cellular immune response against macroinvaders.
The only small variant within a host gene represented in all lineages of the LP group was a SNP in a gene encoding for a nephrin-like protein (g19286.t1 gene). Nephrins are related to cell adhesion and signaling, regulating the structure and function of nephrocytes99. Based on the information available, we were unable to establish a link between the alteration of this gene and the observed phenotype.
The SNPs and InDels identified in the HP lineage group offer some potential to explain the differential response of aphid lineages to the parasitization by D. rapae, but we believe that additional experiments on comparative differential gene expression could provide additional insights into the functional role of the large number of variants detected in intergenic/intronic regions. Intronic sequence variants can modulate genotype–phenotype relationship100 and they have been demonstrated to affect insect resistance to insecticides101,102, the development of human diseases103 and the response to drugs104. However, the occurrence of other mechanisms regulating gene expression in M. persicae should also be considered, since epigenetic changes (DNA methylation) also lead to the loss of M. persicae resistance to insecticides105,106.
The lack of association with host-defensive secondary symbionts and the control of other factors that could interfere with the aphid response to parasitization (nutritional quality of the host plant, age of the aphid and wasp, and genetic variation of the wasp)107,108,109 indicate that the selected isofemale lineages accumulated sufficient genetic diversity to generate the observed phenotypic variability. Similar patterns of intraspecific variation in resistance to parasitism have been observed in other aphid species, such as A. pisum, where genetic diversity among clones contributed to differential survival against parasitoids74,75. The environmental factors that produced the observed phenotypic variation require further physiological and molecular investigation, but our initial hypothesis is that structural variants associated mainly with immune genes and variants in intronic and intergenic regions in the genomes of the selected aphid lineages resulted in alterations in aphid response to parasitization.
The manifestation of defense strategies, regardless of their origin, most often results in changes in life history traits due to the energy costs involved. Changes that direct the allocation of larger amounts of energy to build a stronger immune response will often reduce the energy budget dedicated to sustaining other life history traits, such as fecundity or longevity66. The differences in life history traits observed among the isofemale lineages tested were similar to those observed by Gwynn et al.66 and Vorburger et al.67. However, the relationship between fitness costs and the ability of aphids to respond to D. rapae was not uniform among lineages of the LP group (Iso10LP/r-, Iso11LP/r-, and Iso14LP/r+). Only Iso11LP/r- (lower fecundity, Ro, Rm, and λ, and longer Dt) and Iso14LP/r+ (delayed development) showed significant negative changes in their fitness traits. While the observation of fitness costs in fecundity is commonly reported as a trade-off between female reproduction and immunity in insects110, other life history traits such as development time are rarely studied111.
The hypothesis that the reduced growth rate observed for Iso14LP/r+ is related to a fitness cost due to its greater ability to avoid parasitism is not supported by observations showing that the growth rate is genetically correlated positively with the larval cuticular immune response of the cotton leafworm Spodoptera littoralis83. The observed change in life history traits of Iso14LP/r+ may be related to a reduced allocation of energy to food acquisition to the detriment of other physiological processes, such as immune response and xenobiotic detoxification. Similar trade-offs between fitness and activities related to defense against pathogens and natural enemies have been reported in other insects. For instance, Freitak et al.112 found that immune activation in Pieris brassicae (L.) led to increased metabolic rates. Similarly, Kraaijeveld et al.33 demonstrated that the adoption of defense mechanisms against parasites and parasitoids resulted in reduced size and fecundity in Drosophila melanogaster. SNPs and InDels in Iso14LP/r+ were detected in genes encoding proteins involved in insect immunity (for example, zinc finger proteins of the MYM family and protein kinase-like inhibitor) and detoxification (g12062. t1 gene—multidrug resistance-associated protein belonging to the ABC transporter family)113,114. Reduced growth rates in aphids are generally associated with reduced metabolism to convert ingested food into energy115. Malnutrition has been reported to increase the susceptibility of insects to stress responses and to reduce their immune capacity116,117. Changes in the nutritional content of aphids may also result in aphids that do not provide the correct chemical stimuli to induce female wasps to lay their eggs while assessing for host suitability118,119.
The longer developmental time required for insects to complete their immature development due to malnutrition is often accompanied by changes in other life-history traits, such as fecundity. However, no reproductive changes were observed for Iso14LP/r+, like other studies in which no trade-offs were detected between reproduction and resistance to parasititization80,134,135. We believe that in our study this is due to its association with Rickettsia, given that this secondary symbiont positively affects fecundity in other host-symbiont associations120,121,122. Thus, Rickettsia may mitigate the costs associated with the reduced susceptibility of Iso14LP/r+ to D. rapae parasitization. Thus, the fitness traits observed in Rickettsia-infected isolines suggest that Rickettsia does not benefit the host response to parasitization by D. rapae, but the fecundity of Rickettsia-infected isofemale lineages was higher than that of uninfected isofemale lineages. The observed dissociation between the benefits of Rickettsia on aphid fecundity and its lack of effect on parasitoid resistance may be explained by several factors. First, the energetic trade-offs associated with immune activation may favor reproductive investment over parasitoid resistance, as resource allocation to immunity may come at the expense of other physiological functions, such as fecundity123. This trade-off has been observed in other insect systems, where increased reproductive output is often associated with a reduced ability to mount an effective immune response124. However, it is important to note that additional research (eg., analysis of antibiotic cured Rickettsia-infected lines) is still required to establish a causal relationship between Rickettsia infection and the aphid increased fecundity.
Furthermore, interactions with other symbionts could mediate the observed effects, as previous studies have shown that Rickettsia can alter the density of protective symbionts125 or reduce the number of primary aphid symbiont cells and the host fitness traits126,127. However, our data does not suggest that Rickettsia affects B. aphidicola density in a way that interferes with the provision of essential amino acids and vitamins to the host128,129,130, because we observed higher fecundity in Rickettsia-infected lineages. Finally, environmental conditions and host plant quality may influence both fecundity and immune function in aphids. Studies have shown that the benefits conferred by symbionts are highly context dependent, with changes in temperature and nutrient availability altering their effects131. Aphid symbionts, including Rickettsia, may confer benefits under specific environmental conditions that were not assessed in this study. Future studies should aim to explore these mechanisms in more detail and determine the specific conditions under which Rickettsia confer fitness advantages to aphids.
The observation of fitness costs depends on the environment, and the adaptive cost varies with environmental conditions132, suggesting the experimental conditions used did not allow for the expression of costs associated with the different abilities of the tested lineages to respond to parasitoid attack. It is also possible that the selected lineages evolved different mechanisms to cope with parasitoid attack, as aphid defense mechanisms may rely on physiological, morphological, or behavioral strategies133,134,135,136,137, which may or may not have associated costs. Aphid defense against parasitism may also be provided by the differential ability to metabolize and utilize host plant secondary compounds to influence the successful establishment and development of natural enemies138,139,140.
The results of this study have important practical implications for biological control strategies of aphids using parasitoids. The phenotypic variability observed in aphid lineages suggests the existence of multiple resistance mechanisms, which may pose a challenge to biological control. However, the identification of genetic variants (SNPs and InDels) associated with immune response genes, such as p52IPK and zinc finger proteins, provides molecular targets to potentially reduce aphid defenses. Furthermore, the role of secondary symbionts, such as Rickettsia, which increase fecundity without affecting parasitoid resistance, suggests that symbiont manipulation may be a promising strategy. Environmental factors, such as host plant quality and temperature, also influence the effectiveness of these interactions, highlighting the need for biological control strategies adapted to specific contexts. Future research should explore these mechanisms to develop more effective and sustainable control methods.
In conclusion, we demonstrated variability among aphid isofemale lineages to parasitization and identified conserved small structure variants in all the HP lineage group that support the hypothesis of altered aphid immune function. Our comparative analysis of life history traits and fertility life table among selected lineages did not detect similar fitness costs in lineages of the LP group, suggesting they evolved different mechanisms to reduce the successful parasitization by D. rapae. Rickettsia improved aphid reproduction but did not interfere with the aphid response to the parasitoid. Since the effects of secondary symbionts on stress conditions may be dependent on the density of infection141,142,143, further studies are required to properly investigate the contribution of Rickettsia to M. persicae. Further studies are also needed to characterize the defense mechanisms that isofemale lineages of M. persicae evolved to avoid the successful parasitization by D. rapae, particularly those involving transcriptomics and epigenomics. These findings highlight the complexity of aphid-parasitoid-symbiont interactions and provide valuable insights into biological control strategies. However, limitations remain, such as the need to further explore the role of genetic and epigenetic variants, as well as the influence of environmental factors, to develop more precise and adaptive control methods.
Methods
Insects and rearing
Aphid isofemale lineages were established from adult aphids collected from cabbage (Brassica oleracea var. acephala) plants in experimental plots at Esalq/USP (22° 42′ 46.1″ S; 47° 37′ 37.8″ W) and in home vegetable gardens in Piracicaba (22° 42′ 38.041″ S; 47° 38′ 30.221″ W) and Americana (22° 44′ 20.216″ S; 47° 18′ 19.325″ W), state of São Paulo, and from canola plants (Brassica napus) in Coxilha (28° 13′ 51.809″ S; 52° 24′ 13.752″ W), state of Rio Grande do Sul. Female aphids were isolated in plastic containers containing a leaf of cabbage var. Georgia (Brassica oleracea var. acephala) as a substrate for aphid feeding and reproduction. A total of 43 isofemale lineages were established. Insects were reared under controlled laboratory conditions (20 ± 2 °C; 70 ± 10% RH; 14L:10D) and cabbage leaves were replaced weekly.
The parasitoid strain was obtained from mummified aphids collected in Piracicaba, state of São Paulo (22° 42′ 46.1″ S; 47° 37′ 37.8″ W). Aphid mummies were individualized in glass tubes (8 × 1 cm) containing a droplet of honey to feed the emerging wasp. After emergence, couples were formed and allowed to mate for 24 h. After mating, females were placed in plastic cages (20 × 15 × 10 cm) containing cabbage leaves infested with aphids for parasitization for 48 h. Wasps were then removed and nymphs were kept in cages until aphid mummification. Mummies were collected and transferred to clean dishes lined with filter paper. A droplet of honey was applied to the inside of the lid as a food source for the emerging wasps. The wasps were allowed to mate, and mated females were used to parasitize new aphids. After seven generations under laboratory conditions, the isofemale lineage demonstrating the highest rate of mummification was selected for the experiments. The selected isoline was maintained under controlled conditions (20 ± 2 °C; 60 ± 10% RH; 14L:10D) and continuously reared on 2nd and 3rd instars of M. persicae as previously described.
Plants
Cabbage plants were obtained from seedTr (TopSeed®) sown in seedling trays of 200 cells. Seedlings with a height of 10 cm and three to four true leaves were transferred to 550 mL containers with Tropstrato HT® substrate for cultivation in a greenhouse. Plants were sprayed every two weeks with a foliar fertilizer (Home, Maxx Garden—composition: 150 g/L N, 80 g/L P, 80 g/L K, 400 mg/L B, 100 mg/L Co, 600 mg/L Cu, 500 mg/L Mn, 100 mg/L Mo, 1 g/L Zn, and 60 g/L chelating agent) to stimulate sprouting. Seedlings were also treated every other week with a hydroponic solution containing micro (20 mg/L ZnSO4.H2O, 30 mg/L CuSO4.5H2O, and 90 mg/L FeNaEDTA) and macronutrients (140 mg/L NH4H2PO4, 830 mg/L MgSO4, 360 mg/L KNO3, 80 mg/L NH4NO3, and 905 mg/L CaN2O6) adapted from Hoagland and Snyder144.
Assessing the parasitization of Myzus persicae isofemale lineages by Diaeretiella rapae
A total of 14 isofemale lineages of M. persicae were subjected to parasitization by D. rapae. Third instars M. persicae from each lineage were individually placed on 6 cm cabbage leaf discs placed onto a layer of 2% agar supplemented with 0.02% methyl parahydroxybenzoate (Nipagin®) in a Petri dish and allowed to settle for one hour before individual exposure to a host-fed, 24-h-old, mated D. rapae female for assisted observation of parasitism. Each female wasp was used to parasitize up to seven aphids randomly selected from each isofemale lineage. Nymphs were removed shortly after insertion of the parasitoid ovipositor, which was confirmed by the visualization of the oviposition hole on the cuticle surface, and transferred to a new leaf disc. The leaf discs were replaced every three days to allow the aphid full development. Aphids were maintained on the discs for 7–10 days for mummification (successful parasitization observed) or adult development (failed parasitization or successful aphid defense). Mummified aphids were collected and transferred to glass tubes for adult parasitoid emergence. The experiment was set up in a completely randomized design with 30 replicates (one replicate = one individual host aphid).
Parasitism data were statistically analyzed using Bernoulli generalized linear models145, including the effects of female isoline in the linear predictor. The significance of the isofemale lineage effect was assessed by analysis of deviance. The selection of isofemale lineages for further biological and genomic analyses was based on their distribution in the first (above the 25% percentile) and fourth (below the 75% percentile) quartiles to represent isolines with low and high parasitization rates, respectively.
Assessing symbionts in aphids
Specimens of each M. persicae isofemale lineage used in parasitization assays with D. rapae were subjected to DNA extraction by a salting-out protocol64 (Supplementary Material and Methods). Aphid gDNA was used to detect associated symbionts using diagnostic PCR primarily based on the analysis of the 16S rRNA gene of the most common symbiotic bacteria harbored by aphids (Supplementary Material and Methods; Table S7)146,147,148,149. We selected representatives infected (Iso2HP/r+, Iso3HP/r+ and Iso14LP/r+) and uninfected (Iso4 HP/r-, Iso10LP/r- and Iso11LP/r-) by the Rickettsia among the lineages distributed in the first and fourth quartiles according to their resilience to parasitization for further biological and genomic analyses. Rickettsia was selected because it was the only secondary symbiont to be represented in single infection in aphid isofemale lineages represented in the first and fourth quartiles. Moreover, Rickettsia has also been reported to play nutritional and defensive roles in association with white flies150.
Assessment of life history traits and fertility life tables of Myzus persicae isofemale lineages with different levels of parasitization by Diaeretiella rapae
The existence of adaptive costs associated with the differential response of M. persicae to parasitism by D. rapae was assessed on a selected set of isofemale lineages by evaluating life history traits at the immature and adult stages of the host. Six selected isofemale lineages were subjected to comparative biology experiments, three belonging to the high parasitism (HP) group (Iso2HP/r+, Iso4 HP/r-, and Iso3HP/r+) and three belonging to the low parasitism (LP) group (Iso10LP/r-, Iso14LP/r+, and Iso11LP/r-). LP and HP groups contained Rickettsia-infected (r +) and uninfected (r-) lineages. We evaluated the immature aphid development time (days) and survival (%) by setting up 16 replicates for each selected isofemale lineage. Each replicate was represented by a pool of 10 neonates infesting a potted cabbage seedling, for a total of 160 nymphs per isofemale lineage.
Thirty newly emerged females were randomly selected for each isofemale lineage and individually placed on a cabbage plant. Females were monitored daily for survival, and daily and total fecundity (number of nymphs produced) was recorded. We fitted an inverse Gaussian generalized linear model to the longevity data (since there was no censoring), including the effects of isofemale lineage in the linear predictor. We assessed the significance of the isoline effect using F-tests. We fitted a negative binomial model to the fecundity data, including the effects of isofemale lineage in the linear predictor, and assessed the significance of the isofemale lineage effect using F-tests (since the dispersion parameter was estimated). We performed multiple comparisons by obtaining the 95% confidence intervals for the true linear predictors.
The fecundity and longevity data obtained were used to construct fertility life tables to assess the reproductive success of each selected lineage by calculating and comparing the mean interval between generations (T), finite growth rate (λ), intrinsic growth rate (Rm), net reproduction rate (Ro), and time to duplication (TD)151,152. Plants and nymphs were maintained as before, and nymph mortality and adult development were checked daily.
Twenty adult females obtained from each isofemale lineage were also sampled and used to estimate aphid size by measuring the length of slide-mounted left metathoracic tibia. Measurements were performed using a stereomicroscope connected to a digital image analysis system (Motic Images Plus 2.0).
Aphid immature development time was analyzed using Cox proportional hazards models, including the effects of isofemale lineage in the linear predictor. We assessed the significance of the lineage effect using likelihood ratio (LR) tests for nested models and performed multiple comparisons by obtaining 95% confidence intervals for the true linear predictors. Survival curves were estimated using the Kaplan–Meier estimator to obtain probabilities of M. persicae nymphs reaching adulthood (p < 0.05).
Uncertainty in life table parameters was estimated by Jackknife resampling according to Maia and Luiz153, using the SAS-based routine of Maia et al.154. The obtained means were compared using the Tukey test (p < 0.05). Multivariate analyses were also used to evaluate the life table data using multivariate linear models, including the effects of isofemale lineage as a linear predictor. The significance of the isofemale lineage effect was assessed using Pillai’s trace test, and a heatmap was created to visualize clusters of isofemale lineages based on life table parameters using hierarchical clustering based on Euclidean distances and Ward’s method. A gamma generalized linear model was fitted to the tibia size data, including the effects of isofemale lineage in the linear predictor. We assessed the significance of the isofemale lineage effect using F-tests and performed multiple comparisons by obtaining the 95% confidence intervals for the true linear predictors.
All analyses were performed in R (R Core Team, 2020). Package survival155 was used to fit the Cox proportional hazards models, ggplot2156 was used to generate the plots, and hnp157 was used to assess the goodness-of-fit of the model.
Evaluation and identification of small structural variants (SNPs and InDels) in the selected Myzus persicae isofemale lineages
Genomic sequence data from the six M. persicae isofemale lineages selected from the parasitization assays (Iso2HP/r+, Iso3HP/r+, Iso4 HP/r-, Iso10LP/r-, Iso11LP/r, and Iso14LP/r+) were obtained by extracting and sequencing genomic DNA from a pool of five (5) adult aphids from each isofemale lineage. Insects were subjected to DNA extraction using the Universal AllPrep DNA/RNA/miRNA kit (Qiagen®), according to the manufacturer’s recommendations. Samples with an absorbance of approximately 1.8 were sent to genomic sequencing on the Illumina Novaseq6000 platform using the paired-end strategy (150 bp) in a sequencing service provider.
The reads were subjected to quality control using the FastQC tool v.0.11.8158. Quality trimming and adapter clipping were performed using Trimmomatic v.038 (parameters LEADING:15 TRAILING:15 SLIDINGWINDOW:5:15 MINLEN:50)159 and sequences shorter than 50 bp were removed. Only high quality paired-reads from the libraries of each isofemale lineage were aligned against the reference genome of M. persicae (G006 v.3.0)160 (https://bipaa.genouest.org/sp/myzus_persicae_g006/analysis/3) using Bowtie2 v.2.5.3 aligner software161. SAMtools v.1.21 was used to convert SAM files into sorted BAM files162 (Table S8).
Nucleotide variants were detected using the HaplotypeCaller v.4.3.0 available in the Genome Analysis Toolkit (GATK, https://gatk.broadinstitute.org/)163 with the default parameters. The GenomicsDBImport and GenotypeGVFs tools (GATK) were used to merge the gVCF files into a single VCF and to genotype the nucleotide variants. The preliminary list of identified variants was filtered using the SelectVariants tool (GATK), using the -select-type-to-include argument to obtain a subset containing only SNPs, while another subset only with InDels (insertions and deletions). The SNPs-only subset was filtered by quality metrics using the VariantFiltration tool (GATK) (DP < 3 || QD < 20.0 || MQ < 20.0 || FS > 10.0 || MQRankSum < -2.0 || MQRankSum > 2.0 || ReadPosRankSum < -2.0 || ReadPosRankSum > 2.0 || BaseQRankSum < -2.0 || BaseQRankSum > 2.0 || SOR > 3.0). Similarly, the subset containing only InDels was filtered by quality metrics (size = 1 to 50 bp || QD < 20.0 || FS > 30.0 || ReadPosRankSum < -2.0 || ReadPosRankSum > 2.0 || MQRankSum < 30.0 || BaseQRankSum < -2.0 || BaseQRankSum > 2.0).
The classification of filtered SNPs and InDels by genomic region (Figure S1) was performed using BEDTools Intersect v.2.31.1164, vcf2bed v.2.4.41165, and gencode_regions (https://github.com/saketkc/gencode_regions) tools. For these analyses, the reference genome (G006 v.3.0)160 (https://bipaa.genouest.org/sp/myzus_persicae_g006/analysis/3) structural annotation file was used as the input file for the gencode_regions program, which allows the extraction of 3’UTR, 5'UTR, CDSs, promoters, genes, and introns from GTF/GFF files, resulting in annotation files in bed format. The bed files were then used in the BEDTools intersect v.2.31.1 program together with the bed file of filtered SNPs and InDels previously converted from VCF format to bed using the vcf2bed v.2.4.41 tool. To ensure that SNPs and InDels were counted only once, the intersection was performed in the following hierarchy: promoter 2 kb > 5'UTR > 3'UTR > cds > introns, with the bed containing all filtered SNPs and InDels being used for the first region, and the input file containing SNPs and InDels that did not match the previous region being used for subsequent regions. SNPs and InDels that did not match any of the above genomic regions were classified as belonging to intergenic regions.
Genomic variant annotation and functional effect prediction of SNPs and InDels were performed using the SnpEff software v.5.2166. Subsequently, SNPs and InDels were specifically selected using the filtering tool provided by the SnpSift software v.5.2167 according to the effect in genomic sequences: high—significant functional consequences; moderate—SNPs that result in missense substitutions and InDels resulting in conservative or disruptive changes; low—SNPs that did not result in amino acid changes, and InDels observed in the splice region and at initiation and termination sites; and modifier—effects predicted in intronic, intergenic, promoter and UTR regions. Gene sequences harboring high and moderate effects, present exclusively in isofemale lineages with low (LP) or high parasitism rate (HP) groups or specific isofemale lineages were queried against the NCBI non-redundant protein database (nr-NCBI) using both the BLASTx algorithm and the Diamond high-throughput sequence aligner168, with an e-value cutoff of 10–3. These sequences were also subjected to functional annotation using the Eggnog mapper v.2.0.1 tool to obtain the gene ontology (GO) terms and the clusters of orthologous genes (COGs) classification to analyze the biological processes and pathways in which the high and moderate SNPs and InDels were involved. The mapping analysis was performed using KEGG mapper tool (https://www.genome.jp/kegg/mapper/reconstruct.html) against the Kyoto Encyclopedia of Genes and Genomes169,170,171.
Data availability
The short read DNA sequencing of the six isofemale lineages of Myzus persicae and the molecular sequence of COI used for Diaeretiella rapae identification are fully available at NCBi (BioProject PRJNA1169216 and GeneBank ON243747) (Table S9). The scripts used are available at https://github.com/MarianePossignolo/Repository-Variation-in-isolines-of-Myzus-persicae and https://hackmd.io/@QF2W6Z2XT6qOMO2EYuxCRQ/H1S_fG31kl. Data will become publicly available upon manuscript acceptance. Data access can also be obtained by contacting the corresponding author or the first author, Mariane P. Gomes (email: mariane.gomes@usp.br).
References
Eggleton, P. & Gaston, K. J. “Parasitoid” species and assemblages: Convenient definitions or misleading compromises?. Oikos 59, 417–421. https://doi.org/10.2307/3545155 (1990).
Henter, H. J. & Via, S. The potential for coevolution in a host-parasitoid system: Genetic variation within an aphid population in susceptibility to parasitic wasp. Evolution 49, 427–438. https://doi.org/10.2307/2410267 (1995).
Kraaijeveld, A. R., Alphen, J. J. M. V. & Godfray, H. C. J. The coevolution of host resistance and parasitoid virulence. Parasitology 116, S29–S45. https://doi.org/10.1017/s0031182000084924 (1998).
Lafferty, K. D. & Kuris, A. M. Trophic strategies for animal diversity and body size. Trends Ecol. Evol. 17, 507–513. https://doi.org/10.1016/s0169-5347(02)02615-0 (2002).
Weinersmith, K. L. What’s gotten into you? A review of recent research on parasitoid manipulation of host behavior. Curr. Opin. Insect Sci. 33, 37–42. https://doi.org/10.1016/j.cois.2018.11.011 (2019).
Pennacchio, F. & Strand, M. R. Evolution of developmental strategies in parasitic Hymenoptera. Annu. Rev. Entomol. 51, 233–258. https://doi.org/10.1146/annurev.ento.51.110104.151029 (2006).
Carton, Y. & Nappi, A. J. Drosophila cellular immunity against parasitoids. Parasitol. Today 13, 218–227. https://doi.org/10.1016/S0169-4758(97)01058-2 (1997).
Schmid-Hempel, P. Evolutionary ecology of insect immune Defenses. Annu. Rev. Entomol. 50, 529–551. https://doi.org/10.1146/annurev.ento.50.071803.130420 (2005).
Carton, Y., Poirié, M. & Nappi, A. J. Insect immune resistance to parasitoids. Insect Sci. 15, 67–87. https://doi.org/10.1111/j.1744-7917.2008.00188.x (2008).
Godfray, H. C. J. Parasitoids: Behavioral and Evolutionary Ecology (Princeton University Press, 1994).
Kraaijeveld, A. R. & Godfray, H. C. J. Evolution of host resistance and parasitoid counter-resistance. Adv. Parasitol. 70, 257–280. https://doi.org/10.1016/S0065-308X(09)70010-7 (2009).
Koltz, A. M., Culler, L. E., Bowden, J. J., Post, E. & Høye, T. T. Dominant arctic predator is free of major parasitoid at northern edge of its range. Front. Ecol. Evol. 7, 250. https://doi.org/10.3389/fevo.2019.00250 (2019).
Moore, M. E., Hill, C. A. & Kingsolver, J. G. Differing thermal sensitivities in a host-parasitoid interaction: High, fluctuating developmental temperatures produce dead wasps and giant caterpillars. Funct. Ecol. 35, 675–685. https://doi.org/10.1111/1365-2435.13748 (2021).
Oliver, K. M., Campos, J., Moran, N. A. & Hunter, M. S. Population dynamics of defensive symbionts in aphids. Proc. R. Soc. B 275, 293–299. https://doi.org/10.1098/rspb.2007.1192 (2007).
Chaplinska, M., Gerritsma, S., Dini-Andreote, F., Falcao Salles, J. & Wertheim, B. Bacterial communities differ among Drosophila melanogaster populations and affect host resistance against parasitoids. PLoS ONE 11, e0167726. https://doi.org/10.1371/journal.pone.0167726 (2016).
Reeve, J. D. Stability, variability, and persistence in host-parasitoid systems. Ecology 71, 422–426. https://doi.org/10.2307/1940295 (1990).
Comins, H. N., Hassell, M. P. & May, R. M. The spatial dynamics of host-parasitoid systems. J. Anim. Ecol. 61, 735–748. https://doi.org/10.2307/5627 (1992).
Singh, A. Stochasticity in host-parasitoid models informs mechanisms regulating population dynamics. Sci. Rep. 11, 16749. https://doi.org/10.1038/s41598-021-96212-y (2021).
Benassi, V., Frey, F. & Carton, Y. A new specific gene for wasp cellular immune resistance in Drosophila. Heredity 80, 347–352. https://doi.org/10.1046/j.1365-2540.1998.00303.x (1998).
Kacsoh, B. Z. & Schlenke, T. A. High hemocyte load is associated with increased resistance against parasitoids in Drosophila suzukii, a relative of D. melanogaster. PLoS ONE 7, e34721. https://doi.org/10.1371/journal.pone.0034721 (2012).
Fors, L., Markus, R., Theopold, U. & Hambäck, P. A. Differences in cellular immune competence explain parasitoid resistance for two coleopteran species. PLoS ONE 9, e108795. https://doi.org/10.1371/journal.pone.0108795 (2014).
Stark, J. D. & Acheampong, S. A demographic and modeling approach to determine the suitability of two hosts, Brevicoryne brassicae (Linnaeus) and Myzus persicae (Sulzer) (Heteroptera: Aphididae) of the aphid parasitoid, Diaeretiella rapae (McIntosh) (Hymenoptera: Aphidiidae). Pan-Pac. Entomol. 83, 75–79. https://doi.org/10.3956/0031-0603-83.1.75 (2007).
Weber, G. Genetic variability in host plant adaptation of the green peach aphid, Myzus persicae. Entomol. Exp. Appl. 38, 49–56. https://doi.org/10.1111/j.1570-7458.1985.tb03497.x (1985).
Dedryver, C. A., Le Ralec, A. & Fabre, F. The conflicting relationships between aphids and men: A review of aphid damage and control strategies. C. R. Biol. 333, 539–553. https://doi.org/10.1016/j.crvi.2010.03.009 (2010).
Javed, K. et al. Exploring innovative strategies to control aphids: Meta-analysis and a critical view on what we have and what the future is. J. Pest Sci. 98, 51–57. https://doi.org/10.1007/s10340-024-01852-4 (2025).
Guigo, P. L., Maingeneau, A. & Corff, J. L. Performance of an aphid Myzus persicae and its parasitoid Diaeretiella rapae on wild and cultivated Brassicaceae. J. Plant Interac. 7, 326–332. https://doi.org/10.1080/17429145.2011.628417 (2012).
Girling, R. D., Hassall, M., Turner, J. G. & Poppy, G. M. Behavioural responses of the aphid parasitoid Diaeretiella rapae to volatiles from Arabidopsis thaliana induced by Myzus persicae. Entomol. Exp. Appl. 120, 1–9. https://doi.org/10.1111/j.1570-7458.2006.00423.x (2006).
Soleimannejad, S., Ross, P. A. & Hoffmann, A. A. Effect of Rickettsiella viridis endosymbionts introduced into Myzus persicae aphids on parasitism by Diaeretiella rapae: A combined strategy for aphid control?. Biol. Control 187, 105377. https://doi.org/10.1016/j.biocontrol.2023.105377 (2023).
Desneux, N. et al. Diaeretiella rapae limits Myzus persicae populations after applications of deltamethrin in oilseed rape. J. Econ. Entomol. 98, 9–17. https://doi.org/10.1093/jee/98.1.9 (2005).
Bacci, L. et al. Concentration-mortality responses of Myzus persicae and natural enemies to selected insecticides. J. Environ. Sci. Health 47, 1930–1937. https://doi.org/10.1080/03601234.2012.676494 (2012).
Blande, J. D., Pickett, J. A. & Poppy, G. M. Attack rate and success of the parasitoid Diaeretiella rapae on specialist and generalist feeding aphids. J. Chem. Ecol. 30, 1781–1795. https://doi.org/10.1023/B:JOEC.0000042401.52088.54 (2004).
van Valen, L. The Red Queen. Am. Nat. 111, 809–810. https://doi.org/10.1086/283213 (1977).
Kraaijeveld, A. R., Ferrari, J. & Godfray, H. C. J. Costs of resistance in insect-parasite and insect-parasitoid interactions. Parasitology 125, S71–S82. https://doi.org/10.1017/s0031182002001750 (2003).
Vienne, D. M. et al. Cospeciation vs host-shift speciation: Methods for testing, evidence from natural associations and relation to coevolution. New Phytol. 198, 347–385. https://doi.org/10.1111/nph.12150 (2013).
Brockhurst, M. A. et al. Running with the Red Queen: The role of biotic conflicts in evolution. Proc. R. Soc. B 281, 20141383. https://doi.org/10.1098/rspb.2014.1382 (2014).
Moran, N. A. Symbiosis as an adaptive process and source of phenotypic complexity. Proc. Natl. Acad. Sci. 104, 8627–8633. https://doi.org/10.1073/pnas.0611659104 (2007).
Feldhaar, H. Bacterial symbionts as mediators of ecologically important traits of insect hosts. Ecol. Entomol. 36, 533–543. https://doi.org/10.1111/j.1365-2311.2011.01318.x (2011).
Vorburger, C. & Perlman, S. J. The role of defensive symbionts in host–parasite coevolution. Biol. Rev. 93(1747–1764), 2018. https://doi.org/10.1111/brv.12417 (2018).
Paredes, J. C., Herren, J. K., Schüpfer, F. & Lemaitre, B. The role of lipid competition for endosymbiont-mediated protection against parasitoid wasps in Drosophila. MBio https://doi.org/10.1128/mbio.01006-16 (2016).
Ballinger, M. J., Gawryluk, R. M. R. & Perlman, S. J. Toxin and genome evolution in a Drosophila defensive symbiosis. Genome Biol. Evol. 11, 253–262. https://doi.org/10.1093/gbe/evy272 (2019).
Oliver, K. M. & Perlman, S. J. Toxin-mediated protection against natural enemies by insect defensive symbionts. Adv. Insect Physiol. 58, 277–316. https://doi.org/10.1016/bs.aiip.2020.03.005 (2020).
Oliver, K. M., Russel, J. A., Moran, N. A. & Hunter, M. S. Facultative bacterial symbionts in aphids confer resistance to parasitic wasps. Proc. Natl. Acad. Sci. 100, 1803–1807. https://doi.org/10.1073/pnas.0335320100 (2003).
Oliver, K. M., Degnan, P. H., Hunter, M. S. & Moran, N. A. Bacteriophages encode factors required for protection in a symbiotic mutualism. Science 325, 992–994. https://doi.org/10.1126/science.1174463 (2009).
Rouïl, J., Jousselin, E., Coeurdcier, A., Cruaud, C. & Manzano-Marín, A. The protector within: comparative genomics of APSE phages across aphids reveals rampant recombination and diverse toxin arsenals. Genome Biol. Evol. 12, 878–889. https://doi.org/10.1093/gbe/evaa089 (2020).
Frago, E., Mala, M. & Weldegergis, B. T. Symbionts protect aphids from parasitic wasps by attenuating herbivore-induced plant volatiles. Nat. Commun. 8, 1860. https://doi.org/10.1038/s41467-017-01935-0 (2017).
Sentis, A., Hemptinne, J. L., Magro, A. & Outreman, Y. Biological control needs evolutionary perspectives of ecological interactions. Evol. Appl. 15, 1537–1554. https://doi.org/10.1111/eva.13457 (2022).
Stiling, P. D. The frequency of density dependence in insect host-parasitoid systems. Ecology 68, 844–856. https://doi.org/10.2307/1938356 (1987).
Carius, H. J., Little, T. J. & Ebert, D. Genetic variation in a host-parasite association: Potential for coevolution and frequency-dependent selection. Evolution 55, 1136–1145. https://doi.org/10.1111/j.0014-3820.2001.tb00633.x (2001).
Barrett, J. A. Frequency dependent selection in plant fungal interactions. Philos. Trans. R. Soc. B 319, 473–483. https://doi.org/10.1098/rstb.1988.0060 (1988).
Amos, W. Factors affecting levels of genetic diversity in natural populations. Philos. Trans. R. Soc. B 353, 177–186. https://doi.org/10.1098/rstb.1998.0200 (1998).
Vellend, M. & Geber, M. A. Connections between species diversity and genetic diversity. Ecol. Lett. 8, 767–781. https://doi.org/10.1111/j.1461-0248.2005.00775.x (2005).
Crow, J. F. Advantages of sexual reproduction. Dev. Genet. 15, 205–213. https://doi.org/10.1002/dvg.1020150303 (1994).
Gerber, N. & Kokko, H. Sexual conflict and the evolution of asexuality at low population densities. Proc. R. Soc. B 283, 20161280. https://doi.org/10.1098/rspb.2016.1280 (2016).
Moran, N. A. The evolution of aphid life cycles. Annu. Rev. Entomol. 37, 321–348. https://doi.org/10.1146/annurev.en.37.010192.001541 (1992).
Vorburger, C., Lancaster, M. & Sunnucks, P. Environmentally related patterns of reproductive modes in the aphid Myzus persicae and the predominance of two “superclones” in Victoria, Australia. Mol. Ecol. 12, 3493–3504. https://doi.org/10.1046/j.1365-294X.2003.01998.x (2003).
Shah, M. A. et al. Population ecology of aphid pests infesting potato. Sustain. Agric. Rev. 28, 153–181. https://doi.org/10.1007/978-3-319-90309-5_5 (2018).
Sunnucks, P., England, P. R., Taylor, A. C. & Hales, D. F. Microsatellite and chromosome evolution of parthenogenetic Sitobion aphids in Australia. Genetics 144, 747–756. https://doi.org/10.1093/genetics/144.2.747 (1996).
Oliver, K. M., Moran, N. A. & Hunter, M. S. Costs and benefits of a superinfection of facultative symbionts in aphids. Proc. R. Soc. B. 273, 1273–1280. https://doi.org/10.1098/rspb.2005.3436 (2006).
Guerrero, R., Margulis, L. & Berlanga, M. Symbiogenesis: The holobiont as a unit of evolution. Int. J. Microbiol. 16, 133–143. https://doi.org/10.2436/20.1501.01.188 (2013).
Brown, P. A. & Blackman, R. L. Karyotype variation in the corn leaf aphid, Rhopalosiphum maidis (Fitch), species complex (Hemiptera: Aphididae) in relation to host-plant and morphology. Bull. Entomol. Res. 78, 351–363. https://doi.org/10.1017/S0007485300013110 (1988).
Blackman, L., Spencel, J. M., Field, M. & Devonshire, A. L. Chromosomal location of the amplified esterase genes conferring resistance to insecticides in Myzus persicae (Homoptera: Aphididae). Heredity 75, 297–302. https://doi.org/10.1038/hdy.1995.138 (1995).
Russell, J. A. & Moran, N. A. Costs and benefits of symbiont infection in aphids: Variation among symbionts and across temperatures. Proc. R. Soc. B 273, 603–610. https://doi.org/10.1098/rspb.2005.3348 (2006).
Kikuchi, Y. et al. Symbiont-mediated insecticide resistance. Proc. Natl. Acad. Sci. U.S.A. 109, 8618–8622. https://doi.org/10.1073/pnas.1200231109 (2012).
Sunnucks, P. & Hales, D. F. Numerous transposed sequences of mitochondrial cytochrome oxidases I-II in aphids of the genus Sitobion (Hemiptera: Aphididae). Mol. Biol. Evol. 13, 510–524. https://doi.org/10.1093/oxfordjournals.molbev.a025612 (1996).
Rigby, M. C., Hechinger, R. F. & Stevens, L. Why should parasite resistance be costly?. Trends Parasitol. 18, 116–120. https://doi.org/10.1016/S1471-4922(01)02203-6 (2002).
Gwynn, D. M., Callaghan, A., Gorham, J., Walters, K. F. A. & Fellowes, M. D. E. Resistance is costly: Trade-offs between immunity, fecundity, and survival in the pea aphid. Proc. R. Soc. B 272, 1803–1808. https://doi.org/10.1098/rspb.2005.3089 (2005).
Vorburger, C., Gouskov, A. & Von Burg, S. Genetic covariation between effectiveness and cost of Defense in aphids. Biol. Lett. 4, 674–676. https://doi.org/10.1098/rsbl.2008.0382 (2008).
Hastings, P. J., Lupski, J. R., Rosenberg, S. M. & Ira, G. Mechanisms of change in gene copy number. Nat. Rev. Genet. 10, 551–564. https://doi.org/10.1038/nrg2593 (2009).
Hastings, P. J., Ira, G. & Lupski, J. R. A microhomology-mediated break-induced replication model for the origin of human copy number variation. PLoS Genet. 5, e1000327. https://doi.org/10.1371/journal.pgen.1000327 (2009).
Freeman, J. L. et al. Copy number variation: New insights in genome diversity. Genome Res. 16, 949–961. https://doi.org/10.1101/gr.3677206 (2006).
Loxdale, H. D. The nature and reality of the aphid clone: Genetic variation, adaptation, and evolution. Agr. Forest. Entomol. 10, 81–90. https://doi.org/10.1111/j.1461-9563.2008.00364.x (2008).
Terradot, L. et al. Molecular characterization of clones of the Myzus persicae complex (Hemiptera: Aphididae) differing in their ability to transmit the potato Leafroll luteovirus (PLRV). Bull. Entomol. Res. 89, 355–363. https://doi.org/10.1017/S0007485399000498 (1999).
Losey, J., Harmon, J., Ballantyne, F. & Brown, C. A polymorphism maintained by opposite patterns of parasitism and predation. Nature 388, 269–272. https://doi.org/10.1038/40849 (1997).
Henter, H. J. & Via, S. The potential for coevolution in a host-parasitoid system I. Genetic variation within an aphid population in susceptibility to a parasitic wasp. Evolution 49, 427–438. https://doi.org/10.1111/j.1558-5646.1995.tb02275.x (1995).
Ferrari, J., Müller, C. B., Kraaijeveld, A. R. & Godfray, H. C. J. Clonal variation and covariation in aphid resistance to parasitoids and a pathogen. Evolution 55, 1805. https://doi.org/10.1554/0014-3820(2001)055[1805:cvacia]2.0.co (2001).
Loxdale, H. D. Rapid genetic changes in natural insect populations. Ecol. Entomol. 35, 155–164. https://doi.org/10.1111/j.1365-2311.2009.01141.x (2010).
Loxdale, H. D. & Lushai, G. Maintenance of aphid clonal lineages: Images of immortality?. Infect. Genet. Evol. 3, 259–269. https://doi.org/10.1016/S1567-1348(03)00091-1 (2003).
Oliver, K. M., Moran, N. A. & Hunter, M. S. Variation in resistance to parasitism in aphids is due to symbionts not host genotype. Proc. Natl. Acad. Sci. 102, 12795–12800. https://doi.org/10.1073/pnas.0506131102 (2005).
Vorburger, C., Gehrer, L. & Rodriguez, P. A strain of the bacterial symbiont Regiella insecticola protects aphids against parasitoids. Biol. Lett. 6, 109–111. https://doi.org/10.1098/rsbl.2009.0642 (2010).
Martinez, A. J., Weldon, S. R. & Oliver, K. M. Effects of parasitism on aphid nutritional and protective symbioses. Molec. Ecol. 23, 1594–1607. https://doi.org/10.1111/mec.12550 (2013).
Martinez, A. J. et al. Aphid-encoded variability in susceptibility to a parasitoid. BMC Evol. Biol. 14, 127. https://doi.org/10.1186/1471-2148-14-127 (2014).
Hita, M. T. et al. Genetic localization of a Drosophila melanogaster resistance gene to a parasitoid wasp and physical mapping of the region. Genome Res. 9, 471–481 (1999).
Fellowes, M. & Godfray, H. The evolutionary ecology of resistance to parasitoids by Drosophila. Heredity 84, 1–8. https://doi.org/10.1046/j.1365-2540.2000.00685.x (2000).
Singh, K. S. et al. Global patterns in genomic diversity underpinning the evolution of insecticide resistance in the aphid crop pest Myzus persicae. Commun. Biol. 4, 847. https://doi.org/10.1038/s42003-021-02373-x (2021).
Gregory, L. E., Driscoll, R. M. H., Parker, B. J. & Brisson, J. A. Impacts of body colour, symbionts and genomic regions on the pea aphid wing plasticity variation. Molec. Ecol. 34, e17660. https://doi.org/10.1111/mec.17660 (2025).
Gale, M. et al. P52r IPK regulates the molecular cochaperone P58 IPK to mediate control of the RNA-dependent protein kinase in response to cytoplasmic stress. Biochemistry 41, 11878–11887. https://doi.org/10.1021/bi020397e (2002).
Han, Z. S. et al. A conserved p38 mitogen-activated protein kinase pathway regulates Drosophila immunity gene expression. Mol. Cell. Biol. 18, 3527–3539. https://doi.org/10.1128/MCB.18.6.3527 (1998).
Martín-Blanco, E. p38 MAPK signaling cascades: Ancient roles and new functions. BioEssays 22, 637–645. https://doi.org/10.1002/1521-1878(200007)22:7%3c637::AID-BIES6%3e3.0.CO;2-E (2000).
Cytrynska, M. Studies on the role of protein kinase A in humoral immune response of Galleria mellonella larvae. J. Insect Physiol. 52, 744–753. https://doi.org/10.1016/j.jinsphys.2006.04.002 (2006).
Tsakas, S. & Marmaras, V. J. Insect immunity and its signalling: An overview. Invertebr. Surviv. J. 7, 228–238 (2010).
Eleftherianos, I. et al. Haemocyte-mediated immunity in insects: Cells, processes and associated components in the fight against pathogens and parasites. Immunology 164, 401–432. https://doi.org/10.1111/imm.13390 (2021).
Trainor, J. E. & Mortimer, N. T. Immune cell production is targeted by parasitoid wasp virulence in a Drosophila–parasitoid wasp interaction. Pathogens 10, 49. https://doi.org/10.3390/pathogens10010049 (2021).
Mahanta, D. K. et al. Insect-pathogen crosstalk and the cellular-molecular mechanisms of insect immunity: Uncovering the underlying signaling pathways and immune regulatory function of non-coding RNAs. Front Immunol. 14, 1169152. https://doi.org/10.3389/fimmu.2023.1169152 (2023).
Yang, J. et al. The Toll/IMD pathways mediate host protection against dipteran parasitoids. J. Insect Physiol. 153, 104614. https://doi.org/10.1016/j.jinsphys.2024.104614 (2024).
Guzzo, C. M. et al. SUMO-binding proteins as effectors and regulators of SUMO modification. Cancer Res. 70, 2184–2184. https://doi.org/10.1158/0008-5472.FBCR09-B69 (2009).
Paddibhatla, I., Lee, M. J., Kalamarz, M. E., Ferrarese, R. & Govind, S. Role for sumoylation in systemic inflammation and immune homeostasis in Drosophila larvae. PLoS Pathog. 6, e1001234. https://doi.org/10.1371/journal.ppat.1001234 (2010).
Hegde, S., Soory, X. A., Kaduskar, B. & Ratnaparkhi, G. SUMO conjugation regulates immune signaling. Fly 14, 62–79. https://doi.org/10.1080/19336934.2020.1808402 (2020).
Sajeev, T. K. et al. SUMO and SUMOylation pathway at the forefront of host immune response. Front. Cell Dev. Biol. 9, 681057. https://doi.org/10.3389/fcell.2021.681057 (2021).
Zhuang, S. et al. Sns and Kirre, the Drosophila orthologs of Nephrin and Neph1, direct adhesion, fusion and formation of a slit diaphragm-like structure in insect nephrocytes. Development 136, 2335–2344. https://doi.org/10.1242/dev.031609 (2009).
Cooper, D. N. Functional intronic polymorphisms: Buried treasure awaiting discovery within our genes. Hum. Genom. 4, 284–288. https://doi.org/10.1186/1479-7364-4-5-284 (2010).
Baxter, S. W. et al. Mis-spliced transcripts of nicotinic acetylcholine receptor α6 are associated with field evolved spinosad resistance in Plutella xylostella (L.). PLoS Genet. 6, e1000802. https://doi.org/10.1371/journal.pgen.1000802 (2010).
Vaulin, O. V., Karagodin, D. A., Novgorodova, T. A. & Glupov, V. V. Analysis of Anopheles messeae sl intron gene polymorphism associated with imidacloprid resistance. J. Vector Ecol. 45, 220–232. https://doi.org/10.1111/jvec.12393 (2020).
Salih, M. H., Al-Azzawie, A. F. & Al-Assie, A. H. A. Intronic SNPs and genetic diseases: A review. IJRASB 8, 267–274. https://doi.org/10.31033/ijrasb.8.2.36 (2021).
Wang, D., Guo, Y., Wrighton, S. A., Cooke, G. E. & Sadee, W. Intronic polymorphism in CYP3A4 affects hepatic expression and response to statin drugs. Pharmacogenom. J. 11, 274–286. https://doi.org/10.1038/tpj.2010.28 (2011).
Field, L. M., Devonshire, A. L., Ffrench-Constant, R. H. & Forde, B. G. Changes in DNA methylation are associated with loss of insecticide resistance in the peach-potato aphid Myzus persicae (Sulz.). FEBS Lett. 243, 323–327. https://doi.org/10.1016/0014-5793(89)80154-1 (1989).
Field, L. M. Methylation and expression of amplified esterase genes in the aphid Myzus persicae (Sulzer). Biochem. J. 349, 863–868. https://doi.org/10.1042/bj3490863 (2000).
Hufbauer, R. A. Pea aphid-parasitoid interactions: Have parasitoids adapted to differential resistance?. Ecology 82, 717–725. https://doi.org/10.2307/2680191 (2001).
Wang, S. Y., Chi, H. & Liu, T. X. Demography and parasitic effectiveness of Aphelinus asychis reared from Sitobion avenae as a biological control agent of Myzus persicae reared on chili pepper and cabbage. Biol. Control 92, 111–119. https://doi.org/10.1016/j.biocontrol.2015.10.010 (2016).
Kumar, S., Kashyap, S. & Soni, S. The foraging behavior of Aphelinus asychis Walker (Hymenoptera: Aphelinidae) and Aphidius ervi (Haliday) (Hymenoptera: Braconidae) on Myzus persicae (Sulzer) (Hemiptera: Aphididae). Phytoparasitica 47, 351–360. https://doi.org/10.1007/s12600-019-00735-0 (2019).
Schwenke, R. A., Lazzaro, B. P. & Wolfner, M. F. Reproduction–immunity trade-offs in insects. Annu. Rev. Entomol. 61, 239–256. https://doi.org/10.1146/annurev-ento-010715-023924 (2016).
Cotter, S. C., Kruuk, L. E. B. & Wilson, K. Costs of resistance: Genetic correlations and potential trade-offs in an insect immune system. J. Evol. Biol. 17, 421–429. https://doi.org/10.1046/j.1420-9101.2003.00655.x (2004).
Freitak, D., Ots, I., Vanatoa, A. & Hörak, P. Immune response is energetically costly in white cabbage butterfly pupae. Proc. R. Soc. Lond. 270, S220–S222. https://doi.org/10.1098/rsbl.2003.0069 (2003).
Kang, X. L., Zhang, M., Wang, K., Qiao, X. F. & Chen, M. H. Molecular cloning, expression pattern of multidrug resistance associated protein 1 (mrp1, abcc1) gene, and the synergistic effects of verapamil on toxicity of two insecticides in the bird cherry-oat aphid. Arch. Insect Biochem. Physiol. 92, 65–84. https://doi.org/10.1002/arch.21334 (2016).
Huang, X., Liu, D., Zhang, R. & Shi, X. Transcriptional responses in defense-related genes of Sitobion avenae (Hemiptera: Aphididae) feeding on wheat and barley. Mol. Entomol. 112, 382–395. https://doi.org/10.1093/jee/toy329 (2019).
Calow, P. Ecology, evolution and energetics: A study in metabolic adaptation. Adv. Ecol. Res. 10, 1–62. https://doi.org/10.1016/S0065-2504(08)60233-0 (1977).
Ojala, K., Julkunen-Tiitto, R., Lindström, L. & Mappes, J. Diet affects the immune defence and life-history traits of an Arctiid moth Parasemia plantaginis. Evol. Ecol. Res. 7, 1153–1170 (2005).
Singer, M. S., Mason, P. A. & Smilanich, A. M. Ecological immunology mediated by diet in herbivorous insects. Integr. Comp. Biol. 54, 913–921. https://doi.org/10.1093/icb/icu089 (2014).
Vinson, S. B. & Iwantsch, G. F. Host regulation by insect parasitoids. Q. Rev. Biol. 55, 143–165. https://doi.org/10.1086/411731 (1980).
Sequeira, R. & Mackauer, M. Covariance of adult size and development time in the parasitoid wasp Aphidius ervi in relation to the size of its host, Acyrthosiphon pisum. Evol. Ecol. 6, 34–44. https://doi.org/10.1007/BF02285332 (1992).
Himler, A. G. et al. Rapid spread of a bacterial symbiont in an invasive whitefly is driven by fitness benefits and female bias. Science 332, 254–256. https://doi.org/10.1126/science.1199410 (2011).
Cass, B. N. et al. Conditional fitness benefits of the Rickettsia bacterial symbiont in an insect pest. Oecologia 180, 169–179. https://doi.org/10.1007/s00442-015-3436-x (2015).
Kliot, A., Kontsedalov, S., Lebedev, G., Czosnek, H. & Ghanim, M. Combined infection with Tomato yellow leaf curl virus and Rickettsia influences fecundity, attraction to infected plants and expression of immunity-related genes in the whitefly Bemisia tabaci. J. Gen. Virol. 100, 721–731. https://doi.org/10.1099/jgv.0.001233 (2019).
Bashir-Tanoli, S. & Tinsley, M. C. Immune response costs are associated with changes in resource acquisition and not resource reallocation. Funct. Ecol. 28, 1011–1019. https://doi.org/10.1111/1365-2435.12236 (2014).
Nystrand, M. & Dowling, D. K. Transgenerational interactions involving parental age and immune status affect female reproductive success in Drosophila melanogaster. Proc. R. Soc. B 281, 20141242. https://doi.org/10.1098/rspb.2014.1242 (2014).
Leclair, M. et al. Consequences of coinfection with protective symbionts on the host phenotype and symbiont titres in the pea aphid system. Insect Sci. 24, 798–808. https://doi.org/10.1111/1744-7917.12380 (2017).
Chen, D. Q., Montllor, C. B. & Purcell, A. H. Fitness effects of two facultative endosymbiotic bacteria on the pea aphid, Acyrthosiphon pisum, and the blue alfalfa aphid. A. Kondoi. Entomol. Exp. Appl. 95, 315–323. https://doi.org/10.1046/j.1570-7458.2000.00670.x (2000).
Sakurai, M., Koga, R., Tsuchida, T., Meng, X.-Y. & Fukatsu, T. Rickettsia symbiont in the pea aphid Acyrthosiphon pisum: Novel cellular tropism, effect on host fitness, and interaction with the essential symbiont Buchnera. Appl. Environ. Microbiol. 71, 4069–4075. https://doi.org/10.1128/AEM.71.7.4069-4075.2005 (2005).
Baumann, P., Moran, N. A. & Baumann, L. The evolution and genetics of aphids endosymbionts. Bioscience 47, 12–20. https://doi.org/10.2307/1313002 (1997).
Silva, F. J., Ham, R. C. H. J., Sabater, B. & Latorre, A. Structure and evolution of the leucine plasmids carried by the endosymbiont (Buchnera aphidicola) from aphids of the family Aphididae. Microbiol. Lett. 168, 43–49. https://doi.org/10.1111/j.1574-6968.1998.tb13253.x (1998).
Viñuelas, J. et al. Conservation of the links between gene transcription and chromosomal organization in the highly reduced genome of Buchnera aphidicola. BMC Genom. 8, 143. https://doi.org/10.1186/1471-2164-8-143 (2007).
Corbin, C. et al. Heritable symbionts in a world of varying temperature. Heredity 118, 10–20. https://doi.org/10.1038/hdy.2016.71 (2017).
Hunt, J., Bussière, L. F., Jennions, M. D. & Brooks, R. What is genetic quality?. Trends Ecol. Evol. 19, 329–333. https://doi.org/10.1016/j.tree.2004.03.035 (2004).
Minchella, D. J. Host life-history variation in response to parasitism. Parasitology 90, 205. https://doi.org/10.1017/S0031182000049143 (1985).
Ferrari, J., Muller, C. B., Kraaijeveld, A. R. & Godfray, H. C. J. Clonal variation and covariation in aphid resistance to parasitoids and a pathogen. Evolution 55, 1805–1814. https://doi.org/10.1554/0014-3820(2001)055[1805:CVACIA]2.0.CO;2 (2001).
von Burg, S., Ferrari, J., Müller, C. B. & Vorburger, C. Genetic variation and covariation of susceptibility to parasitoids in the aphid Myzus persicae—No evidence for trade-offs. Proc. R. Soc. B 275, 1089–1094. https://doi.org/10.1098/rspb.2008.0018 (2008).
Sandrock, C., Gouskov, A. & Vorburger, C. Ample genetic variation but no evidence for genotype specificity in an all-parthenogenetic host-parasitoid interaction. J. Evol. Biol. 23, 578–585. https://doi.org/10.1111/j.1420-9101.2009.01925.x (2010).
Vilcinskas, A. The role of epigenetics in host–parasite coevolution: Lessons from the model host insects Galleria mellonella and Tribolium castaneum. Zoology 119, 273–280. https://doi.org/10.1016/j.zool.2016.05.004 (2016).
Turlings, T. C. J. & Benrey, B. Effects of plant metabolites on the behavior and development of parasitic wasps. Écoscience 5, 321–333. https://doi.org/10.1080/11956860.1998.11682472 (1998).
Desneux, N., Barta, R. J., Hoelmer, K. A., Hopper, K. R. & Heimpel, G. E. Multifaceted determinants of host specificity in an aphid parasitoid. Oecologia 160, 387–398. https://doi.org/10.1007/s00442-009-1289-x (2009).
Harvey, J. A. & Gols, R. Effects of plant-mediated differences in host quality on the development of two related endoparasitoids with different host-utilization strategies. J. Insect Physiol. 107, 110–115. https://doi.org/10.1016/j.jinsphys.2018.03.006 (2018).
Brown, B. L., Creed, R. P., Skelton, J., Rollins, M. A. & Farrell, K. J. The fine line between mutualism and parasitism: Complex effects in a cleaning symbiosis demonstrated by multiple fields experiments. Oecologia 170, 199–207. https://doi.org/10.1007/s00442-012-2280-5 (2012).
Martinez, J. et al. Should symbionts be nice or selfish? Antiviral effects of Wolbachia are costly but reproductive parasitism is not. PLoS Pathog. 11, e1005021. https://doi.org/10.1371/journal.ppat.1005021 (2015).
Hopkins, S. R., Wojdak, J. M. & Belden, L. K. Defensive symbionts mediate host–parasite interactions at multiple scales. Trends Parasitol. 33, 53–64. https://doi.org/10.1016/j.pt.2016.10.003 (2017).
Hoagland, D. R. & Snyder, W. C. Nutrition of strawberry plant under controlled conditions: (a) effects of deficiencies of boron and certain other elements: (b) susceptibility to injury from sodium salts. PASHS 30, 288–294 (1934).
McCullagh, P. & Nelder, J. A. Generalized Linear Models 2nd edn. (Chapman & Hall, 1989).
Haine, E. R. Symbiont-mediated protection. Proc. R. Soc. 275, 353–361. https://doi.org/10.1098/rspb.2007.1211 (2008).
Moran, N. A., McCutcheon, J. P. & Nakabachi, A. Genomics and evolution of heritable bacterial symbionts. Annu. Rev. Genet. 42, 165–190. https://doi.org/10.1146/annurev.genet.41.110306.130119 (2008).
Oliver, K. M., Degnan, P. H., Burke, G. R. & Moran, N. A. Facultative symbionts of aphids and the horizontal transfer of ecologically important traits. Annu. Rev. Entomol. 55, 247–266. https://doi.org/10.1146/annurev-ento-112408-085305 (2010).
Vorburger, C. Symbiont-conferred resistance to parasitoids in aphids—Challenges for biological control. Biol. Control 116, 17–26. https://doi.org/10.1016/j.biocontrol.2017.02.004 (2018).
Fan, Z. Y. et al. Rickettsia infection benefits its whitefly hosts by manipulating their nutrition and defense. Insects 13, 1161. https://doi.org/10.3390/insects13121161 (2022).
Krips, O. E., Witul, A., Willems, P. E. L. & Dicke, M. Intrinsic rate of population increase of the spider mite Tetranychus urticae on the ornamental crop gerbera: Intraspecific variation in host plant and herbivore. Entomol. Exp. Appl. 89, 159–168. https://doi.org/10.1046/j.1570-7458.1998.00395.x (1998).
Ricklefs, R. E. A economia da natureza 5th edn. (Guanabara Koogan, 2003).
Maia, A. H. N. & Luiz, A. J. B. Programa SAS para análise de tabelas de vida e fertilidade de artrópodes: o método jackknife. (Comunicado Técnico, 33), Embrapa: Jaguariúna, p. 11 (2006).
Maia, A. H. N., Luiz, A. J. B. & Campanhola, C. Statistical inference on associated fertility life table parameters using Jackknife technique: Computational aspects. J. Econ. Entomol. 93, 511–518. https://doi.org/10.1603/0022-0493-93.2.511 (2000).
Therneau, T. A package for survival analysis in R. R package version 3.2-7. Available online at: https://CRAN.R-project.org/package=survival (2020).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer, 2016).
Moral, R. A., Hinde, J. & Demétrio, C. G. B. Half-normal plots and overdispersed models in R: The hnp Package. J. Stat. Softw. 81, 1–23. https://doi.org/10.18637/jss.v081.i10 (2017).
Andrews, S. FastQC: A quality control tool for high throughput sequence data [Online]. Available online at: http://www.bioinformatics.babraham.ac.uk/projects/fastqc (2010).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120. https://doi.org/10.1093/bioinformatics/btu170 (2014).
Singh, K. S. et al. Global patterns in genomic diversity underpinning the evolution of insecticide resistance in the aphid crop pest Myzus persicae. Commun. Biol. 4, 847. https://doi.org/10.1038/s42003-021-02373-x (2021).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25. https://doi.org/10.1186/gb-2009-10-3-r25 (2009).
Li, H. et al. 1000 Genome project data processing subgroup, the sequence alignment/map format and SAMtools. Bioinformatics 25(16), 2078–2079. https://doi.org/10.1093/bioinformatics/btp352 (2009).
Poplin, R. et al. Scaling accurate genetic variant discovery to tens of thousands of samples. bioRxiv 201178. https://doi.org/10.1101/201178 (2017).
Quinlan, A. R. & Hall, I. M. BEDTools: A flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842. https://doi.org/10.1093/bioinformatics/btq033 (2010).
Neph, S. et al. BEDOPS: High-performance genomic feature operations. Bioinformatics 28, 1919–1920. https://doi.org/10.1093/bioinformatics/bts277 (2012).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms. SnpEff. Fly 6, 80–92. https://doi.org/10.4161/fly.19695 (2012).
Cingolani, P. et al. Using Drosophila melanogaster as a model for genotoxic chemical mutational studies with a new program. SnpSift. Front. Genet. 15, 3–35. https://doi.org/10.3389/fgene.2012.00035 (2012).
Buchfink, B., Xie, C. & Huson, D. Fast and sensitive protein alignment using DIAMOND. Nat. Methods 12, 59–60. https://doi.org/10.1038/nmeth.3176 (2015).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30. https://doi.org/10.1093/nar/28.1.27 (2000).
Kanehisa, M. Toward understanding the origin and evolution of cellular organisms. Protein Sci. 28, 1947–1951. https://doi.org/10.1002/pro.3715 (2019).
Kanehisa, M., Furumichi, M., Sato, Y., Matsuura, Y. & Ishiguro-Watanabe, M. KEGG: Biological systems database as a model of the real world. Nucleic Acids Res. 53, D672–D677. https://doi.org/10.1093/nar/gkae909 (2025).
Acknowledgements
We thank Dr. Silvana Savaris, Dr. Eduardo Shimbori (ESALQ/USP) and Dr. Ana Paula Wengrat for the identification of the insect species used in this work. To Dr. Roberto A. Zucchi (ESALQ/USP) for allowing the use of his laboratories for the development of part of the experiments reported. We also thank the São Paulo Research Foundation (FAPESP) for providing the grant to the first author (Grant: 2019/06876-1, 2020/07896-3), Brazilian National Research Council (CNPq) (grant: 2022/164221) and the Coordination of Superior Level Staff Improvement (CAPES) (Finance Code 001) for additional funds.
Funding
Funds were provided by the São Paulo Research Foundation (FAPESP) (grant: 2019/06876-1, 2020/07896–3), the Brazilian National Research Council (CNPq) (grant: 2022/164221) and the Coordination of Superior Level Staff Improvement (CAPES) (Finance Code 001).
Author information
Authors and Affiliations
Contributions
Conceptualization and methodology: MPG and FLC. Investigation and data curation: MPG. Formal analysis: MPG, RAM, IARL, TB, RR and ECL. Supervision and funding acquisition: FLC. Writing—original draft: MPG. Writing—editing: MPG and FLC. Review: All authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Gomes, M.P., Lira, E.C., de Lara, I.A.R. et al. Genomic structure and life history variation of isofemale lineages of Myzus persicae with different levels of parasitization by Diaeretiella rapae. Sci Rep 15, 35515 (2025). https://doi.org/10.1038/s41598-025-19494-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-19494-6