Abstract
The growing crisis of antimicrobial resistance (AMR) underscores the critical need for innovative antimicrobial discoveries. Novel antibiotics targeting the bacterial cell wall remain an attractive area of research, due to their conservation and essentiality in bacteria and their absence in eukaryotic cells. Antibiotics targeting lipid II are of special interest due to the reduced potential for target modification of lipid components and their surface accessibility to inhibitors. In this study, we identified 3-phenyl-4-phenoxypyrazole analogues named PYO12 and PYO12a with bactericidal activity against gram-positive bacteria and low cytotoxicity for different types of mammalian cells. Gram-negative bacteria were resistant to PYO12 activity through extrusion of this compound via efflux pumps. Exposure to PYO12 induces expression of genes involved in resistance to antimicrobials targeting the cell wall, suggesting that PYO12 acts via binding to lipid II or other lipid intermediates involved in peptidoglycan or teichoic acid biosynthesis. Antagonism of PYO12 antibacterial activity by undecaprenyl-pyrophosphate supports the idea that PYO12 may bind to the lipid moiety of lipid II blocking the shuttling of peptidoglycan precursors across the cytoplasmic membrane. These findings open opportunities to further develop these compounds as antibiotics targeting bacterial cell wall synthesis.
Similar content being viewed by others
Introduction
Antimicrobial resistance (AMR) has recently become one of the leading causes of death worldwide1 highlighting the need for new antibiotics with low potential for resistance development. Bacterial cell wall synthesis is still one of the most desirable targets for antibiotic development due to its essentiality for bacterial growth and the lack of similar structures in mammalian cells. Antibiotics targeting the bacterial cell wall mainly act through inhibition of cell wall synthesis enzymes (e.g. ampicillin) or binding to cell wall building blocks such as lipid II (e.g. vancomycin or nisin)2.
Lipid II is an essential precursor in peptidoglycan biosynthesis in all bacteria. It is initially synthesized on the inner leaflet of the cytoplasmic membrane, where its undecaprenyl-pyrophosphate (C55-PP) hydrocarbon chain acts as a lipid anchor. During peptidoglycan synthesis, lipid II shuttles across the membrane to incorporate its disaccharide MurNAc-GlNAc-pentapeptide ʻbuilding blockʼ into the growing peptidoglycan polymer. The lipid intermediate C55-PP is then dephosphorylated and translocated back to the cytosolic side by a yet unknown mechanism where the formation of lipid II starts again3. Lipid II formation is a limiting step in cell wall synthesis due to the fixed number of undecaprenyl-phosphate (C55-P) molecules per cell (2 × 105 molecules)4,5. The essentiality of lipid II, its high conservation in bacteria and its accessibility to inhibitors on the outside of the cytoplasm makes it a highly attractive target for antibiotic development. One of the main disadvantages of antibiotics targeting lipid II is that they are usually not effective against Gram-negative bacteria, mainly due to the low permeability of the LPS populated outer membrane and the presence of multiple efflux pumps6.
The main lipid II binders used in the clinic are glycopeptides, the most well-known being vancomycin that binds to lipid II through the carboxyl end of the pentapeptide D-Ala-D-Ala7 however resistance has risen due to the substitution of D-Ala-D-Ala by D-Ala-D-Lactate8. Other lipid II binding antibiotics, such, nisin, and cyclic depsipeptides9 recognize the C55-PP moiety of the lipid II, while other antibiotics such as bacitracin10 directly bind to the lipid intermediate C55-PP preventing lipid II formation. An advantage of targeting C55-PP is that this molecule is also used for teichoic acid11 and capsule biosynthesis12 so various pathways can be inhibited simultaneously. Resistance to these antibiotics has been reported by the induction of ABC transporters VraDEH and BraDE, which play a role in antibiotic transport and detoxification13 but resistance due to modification of the C55-PP lipid carrier has not been reported presumably due to the difficulty for bacteria to sufficiently modify the lipid moiety and retain functionality14.
The discovery of small molecules able to bind to the lipid moiety of the lipid II to surpass resistance emergence in Gram-positive bacteria has drawn a lot of interest in recent years. Different small molecules have been found to bind to the undecaprenyl moiety of lipid II15,16,17,18 however, cytotoxicity has remained an issue for these molecules (with selectivity indexes, i.e., ratio between the compound’s antibacterial activity and its cytotoxicity, between 1.216 and 417 for small molecules targeting the lipid moiety of lipid II).
Our previous work showed 3,4-diphenylpyrazoles (DPP) to be candidate antimicrobials with a mechanism of action related to or caused by loss of inner membrane integrity in a similar manner to nisin19. Here, we further explored the DPP scaffold and found a 3-phenyl-4-phenoxypyrazole (PYO) derivative referred to as compound PYO12 and the related PYO12a, with low cytotoxicity in vitro and bactericidal activity against a panel of Gram-positive bacteria. We demonstrated that exposure to PYO12 leads to loss of bacterial membrane integrity and hypothesize that it acts in a similar manner to nisin by binding to lipid II or other lipid intermediates. To corroborate the mechanism of action, we quantified the expression of cell wall stress genes vraX and cwrA which are known to be significantly upregulated by agents that target the bacterial cell wall or membrane, such as bacitracin, vancomycin and antimicrobial peptides (AMPs)18,20,21. We also analysed the expression of the regulons controlled by two-component systems (GraRS, BraRS, and VraRS), which have been linked to decreased susceptibility to antibiotics targeting lipid II and the bacterial cell wall22. PYO12 induced expression of cell wall stress genes and our studies showing antagonism of antibacterial activity by C55-PP indicate that PYO12 binds to the C55-PP moiety of lipid II and is a promising lead for the development of a novel antibiotic.
Results
Antimicrobial activity of PYO derivatives against S. aureus
Starting with the DPP scaffold previously published19 we identified a set of commercially available analogues (PYO1-PYO22, Table 1) with a 4-phenoxy group instead of the original 4-phenyl group. We then assessed their antibacterial activity against S. aureus str. Newman. The most potent derivatives reached a minimal inhibitory concentration (MIC) of 1 µg/ml (PYO1 and PYO12, Table 1). In the DPP series, replacing the R1 methyl group with a trifluoromethyl group improved antibacterial activity. To explore if the DPP’s SAR could be transferred to the novel series, we synthesized PYO23 (Figure SF2), the trifluoromethyl analogue of one of the most potent compounds of the series PYO1. In contrast to the original series, this transformation resulted in a decrease in antibacterial potency.
The activity of PYO14 suggests that the replacement of the R2-hydroxyl of ring A, a resorcinol, with a methoxy group is tolerated. The improved activity of the R4-propyloxy derivative PYO20 and the R4-benzyloxy PYO11 compared to their hydroxy analogues (PYO18 and PYO13, respectively) indicate that O-alkylation at this position can be beneficial in terms of antibacterial activity. The matched molecular pair PYO1/PYO4 shows that the R5-ethyl substitution causes a marked improvement in activity. Methylation is tolerated at the R3-position (PYO19), whilst the R6-methyl derivative is only weakly active (PYO22).
Favourable substitutions on the C-ring are a chlorine or isopropyl in position R7 (PYO1, PYO17, PYO18, PYO20, PYO23/PYO2), a phenyl in position R8 (PYO12) and a bromine in position R9 (PYO14). We also investigated the impact of exchanging the -O-linker with a -S-linker by synthesizing PYO24 (Figure SF3). Its antimicrobial potency was similar to the parent analogue PYO1, indicating that the replacement is bioisosteric.
Cytotoxicity of antimicrobial PYOs
The cytotoxicity of the PYOs (Table 1) was tested using the human kidney cell line HEK293 and calculated as LC50 (lethal concentration 50, i.e. the concentration required to kill 50% of the cell population). All the compounds with antibacterial activity against S. aureus str. Newman were toxic to mammalian cell lines at low concentrations with only three compounds (PYO1, PYO12 and PYO24) showing a selectivity index (drug MIC/LC50) higher than 10. We decided to further investigate the SAR of PYO12 and its antimicrobial mechanism because it had a different structure to other compounds of the series due to the presence of a phenyl group at position R8. The cytotoxicity of PYO12 for the liver cell line HepG2, commonly used in toxicology studies, gave LC50 value of 11.5 ± 1.33 µg/ml, and selectivity index of 11.5. To allow the comparison between PYO derivatives and commonly used antimicrobials, the toxicity of nisin, vancomycin and novobiocin was also calculated, as well as their selectivity indexes.
Finally, we tested the potential damaging effects of PYO12 on biological membranes using the sensitive red blood cell haemolysis assay. Incubation of 150 µg/ml of PYO12 for 30 min with sheep red blood cells in PBS resulted in complete haemolysis. However, haemolysis dropped below 10% at 75 µg/ml and to 0% at 37.5 µg/ml. The HC50 (concentration that causes 50% haemolysis in RBCs) was calculated to be 121 µg/ml, resulting in a selectivity index of 60.5 (Supplementary Figure SF1). The three mentioned antimicrobials did not show haemolysis in the highest concentrations tested of 250 µg/ml.
Since PYO structures are derived from known Hsp90 inhibitors (mammalian chaperone) we tested the inhibition of Hsp90 by PYO12, however no inhibition was detected. This was expected due to the bulky substitution of PYO12 probably leading to reduced affinity to the ATP domain19.
Spectrum of activity
The selectivity index of compound PYO12 prompted us to test its antimicrobial activity against Bacillus subtilis and a panel of common opportunistic human pathogens, methicillin-resistant Staphylococcus aureus (MRSA), vancomycin susceptible and resistant Enterococcus faecium and Enterococcus faecalis, Streptococcus pneumoniae, Escherichia coli and Pseudomonas aeruginosa, and Streptococcus suis a common pig pathogen (Table 2). Compound PYO12 showed inhibitory activity against all Gram-positive bacteria tested with MICs ranging from 1 to 16 µg/ml with E. faecalis being the least susceptible Gram-positive species. PYO12 had no inhibitory effect against Gram-negative E. coli and P. aeruginosa.
The TolC-dependent efflux pumps are mainly responsible for the lack of efficacy of PYO12 in Gram-negative bacteria
We hypothesized that the lack of activity against Gram-negative bacteria was due to reduced permeability caused by the outer membrane and/or effective efflux of the compounds. We investigated the antibacterial activity of the compounds against E. coli (D21f2), a strain with truncated a lipopolysaccharide (LPS) barrier, and a tolC mutant of E. coli (JW5503), a strain defective in compound efflux.
Whilst none of the compounds inhibited WT E. coli ATCC25922 growth, some derivatives showed improved activity against E. coli D21f2, with MIC as low as 16 µg/ml for PYO17 and PYO18. However, the MIC remains high (32–128 µg/ml) for most compounds (Table 3), indicating that the LPS layer has a limited impact on compound penetrability. The MIC for E. coli ΔtolC was lower or the same as for S. aureus for 12 of 21 active compounds (Table 3) indicating a key role for efflux pumps in the efflux of PYO12 and related compounds in E. coli. In the cases where the MIC was significantly higher for E. coli ΔtolC than for S. aureus, an ethyl or isopropyl group substitution was present, suggesting these groups reduce penetrability through the outer membrane (compounds PYO1, PYO2, PYO19, PYO20, Table 1).
Structure-activity relationship of PYO12 derivatives
To further study the SAR of PYO12, we purchased analogues from commercial vendors (PYO12e-f, PYO12i and PYO12l, Molport, Riga) or performed targeted synthesis (PYO12a-d, PYO12g-h, PYO12j-k, PYO121m-n, O2H group, Oxford, supplementary data, Figure SF4-SF13). With respect to the SAR (Fig. 1), elimination of the R1-methyl group (general scaffold in Table 1) in the pyrazole ring (B ring) improves antibacterial activity when combined with the R5-ethyl group (PYO12c/PYO12d and PYO12j/PYO12k), whilst the MIC in unchanged in the R5-unsubstituted matched molecular pair PYO12a/PYO12. In all three cases removing the R1-methyl increases the selectivity index over cytotoxicity in the two listed cell lines. N-methylation of the pyrazole increases cytotoxicity, but it does not alter the MIC, indicating that the hydrogen bond donor in the ring B is not essential for the antibacterial activity (compare PYO12 to PYO12n and PYO12m).
Structure of fourteen PYO12 analogues found using similarity search. A, B and C rings are indicated in PYO12 structure. Minimal inhibitory concentration (μg/ml) against S. aureus and LC50 (μg/ml) for HEK293 and HepG2 cells is included.
A distinctive feature of PYO12 is the presence of a phenyl in position R8. The addition of a chlorine to the para position of this phenyl does not have a significant effect on the activity (compare PYO12 vs. PYO12b). Removal of the R8-phenyl is detrimental for the antimicrobial activity (see pair PYO12d/PYO12e). Moving the phenyl to the R7 position results in a loss of antibacterial activity (see pairs PYO12/PYO12g and PYO12a/PYO12h). However, potency is recovered when substituting the R5 position with an ethyl (PYO12c/PYO12j and PYO12d/PYO12k). The potency of PYO12l suggests that the O-spacer is not required for activity.
In this set of compounds, PYO12j stands out for the excellent MIC 0.25 µg/ml. PYO12j cytotoxicity in HepG2 and HEK cells is 11.9 ± 0.6 µg/ml and 18.1 ± 2.8 µg/ml, respectively, giving selectivity indexes of 47.6 and 72.4, which are reasonable for optimization. PYO12a is the derivative with best profile, as it maintains the antibacterial potency of PYO12 and shows low cytotoxicity in both cell lines.
Compound PYO12 is bactericidal and increases bacterial membrane permeability
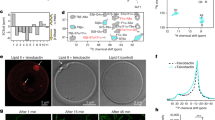
Previous studies showed that pyrazole derivatives compromise S. aureus membrane integrity19. To investigate if this is the putative mechanism of action of compound PYO12 we performed assays using SYTOX Green, a dye which only enters the bacteria and stains nucleic acids when membrane permeability is compromised. Like nisin which was used as a positive control, addition of 1xMIC (1 µg/ml) of PYO12 to S. aureus for 5 min increased membrane permeability (Fig. 2A).
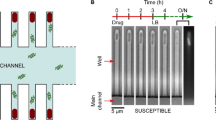
To rule out the possibility that we were measuring dead bacteria after exposure to PYO12, we performed a time-kill curve assay with different concentrations of PYO12 (Fig. 2B). At 1xMIC (1 µg/ml), PYO12 has a bacteriostatic effect (Fig. 2B). At 2xMIC (2 µg/ml) PYO12 is bactericidal after 30 min exposure, and this effect is more pronounced with 4xMIC. From these results we concluded that at 1xMIC, PYO12 increases membrane permeability prior to any loss of viability.
Finally, to show that the mechanism of action in Gram-negative bacteria is also related to disruption of the inner membrane, we performed the membrane damage assay using E. coli ΔtolC (Fig. 2C). Even though the Gram-negative E. coli ΔtolC strain needs a higher exposure time to show membrane damage, we can see that after 30 min, 1xMIC exposure to PYO12 (2 µg/ml), showed similar membrane damage as the colistin control (1xMIC = 4 µg/ml).
Membrane permeability and time-kill curve in S. aureus and E. coli ΔtolC by PYO12. (A) Membrane integrity assessed by SYTOX Green uptake and analysed by flow cytometry. Overlay histograms represent fluorescence intensity of stained bacterial cells (event count vs FITC fluorescence). Increased FITC fluorescence (488 nm) of bacterial population occurs after 5 min exposure to 0.5, 1 and 2xMIC of PYO12 compared to the non-treated control and negative control treated with novobiocin (1xMIC). Similar shifts in fluorescence are observed with the positive control nisin (1xMIC), a known membrane permeabilizing antimicrobial. (B) Time-kill curve of S. aureus incubated with PYO12 at 0.5, 1, 2 and 4xMIC (0.5-4 μg/ml). Graphs show CFU/ml vs time after addition of PYO12. Error bars show standard deviation (SD) of three independent experiments. (C) Membrane integrity after 10 min and 30 min exposure to 0.5, 1 and 2xMIC of PYO12 to E. coli ΔtolC compared to the non-treated control. Negative control (novobiocin, blue) and positive control (colistin, yellow) are included.
Putative mechanism of action of PYO12 involves binding to lipid II or other cell-wall lipid intermediates
To provide further evidence that PYO12 works by targeting cell-wall or bacterial membrane, expression of genes involved in cell wall stress was quantified by PCR. Exposure of S. aureus for 10 min to 1xMIC of PYO12 (1 µg/ml) led to strong overexpression of vraX and cwrA genes (fold-change 629.64 and 736.84, respectively, Fig. 3), genes known to be activated by lipid II binding18 or cell-wall targeting antibiotics respectively20,21 further suggesting that the mechanism of action for PYO12 reduces bacterial cell envelope integrity.
Expression data (Fig. 3) of vraD, dltA and murZ, pbp2 and sgtB, suggests that both BraRS and VraRS TCS are also activated by exposure to 1xMIC of PYO1213,23. As expected, expression of the GraRS-regulated dltA24 was not significantly changed (fold-change 0.89), most likely due to the neutral charge of PYO12. Genes murZ, pbp2 and sgtB were moderately upregulated (28.48, 3.28, 2.06-fold change, respectively), indicating VraRS activation due to cell-wall stress. Interestingly, vraD was highly upregulated (599-fold), suggesting BraRS is activated. Based on these results, we hypothesized that PYO12 acts by binding to lipid II or intermediates such as undecaprenyl-pyrophosphate.
Finally, to find further supporting evidence for binding to lipid II, we exposed B. subtilis to 1xMIC of PYO12 (1 µg/ml) for 10 min, and measured expression of liaI in comparison to the DMSO control. Exposure to PYO12 caused a 16-fold induction of liaI, which is in the regulon controlled by the LiaRS TCS regulon supporting the lipid II binding theory25.
Differential expression of genes involved in response to cell-wall targeting antibiotics in S. aureus exposed to 1 μg/ml of PYO12. In red cwrA and vraX are genes generally involved in the cell wall stress stimulon (CWSS), in green vraD is regulated by the TCS BraRS, in blue dltA regulated by the TCS GraRS, and in violet sgtB, pbp2 and murZ which are regulated by the TCS VraRS. Differential expression is expressed as log2 of the fold change as calculated by qPCR and the 2−ΔΔCt method.
Binding of PYO12 to undecaprenyl-pyrophosphate
To check if PYO12 binds to undecaprenyl-pyrophosphate(C55-PP), we performed a competition assay in which PYO12 was incubated for 10 min with 0.5-4-fold molar excess of C55-PP, before determining its growth inhibitory activity against S. aureus. When S. aureus is exposed to PYO12 in the presence of 2-fold molar ratio of the lipid, the MIC increases significantly, suggesting that PYO12 binds to C55-PP and is therefore not available for binding to the bacterial lipid intermediate (Fig. 4). Incubation of PYO12 with 4-fold molar ratio of the monophosphate C55−P no difference in MIC was detected, indicating that the pyrophosphate moiety is necessary for binding. This also indicates that an unspecific binding to lipid is unlikely.
Antagonism of C55-PP on antimicrobial activity of PYO12. Heat map represents growth of S. aureus (OD600) in a 96-well plate after exposure to different concentrations of PYO12 (0.125-8 8 µg/ml) in presence of different molar ratios of C55-PP (0 to 4) and C55-P (4). Red indicates bacterial growth while green indicates no-growth. Experiments were independently repeated in triplicate and graph shows the average growth per well in these experiments.
PYO12 shows low potential for resistance development
No spontaneous resistant mutants were found when inoculating 1011 CFU of S. aureus Newman onto MHB plates containing 5x and 10x MIC of PYO12 (5 and 10 µg/ml, respectively), indicating a low potential for spontaneous resistance development. Next, we tested the emergence of resistance mutants when the bacteria were exposed to sub-inhibitory concentrations of PYO12. From four experiments, only one resulted in the emergence of a stable resistant mutant with MIC 32 µg/ml after 3 days of exposure to sub-inhibitory concentrations to PYO12. Genome sequence analysis revealed the following mutations in the resistant strain: an internal deletion SaeR (1-228 D48-53) and a truncation in YbjH (1-138) (VCF accession number PRJEB86156, EMBL-EBI-EVA), both genes have been related to β-lactam resistance.
Discussion
Our previous research showed that diphenylpyrazoles (DPP) exert antimicrobial activity by disrupting membrane integrity19. However, their toxicity toward mammalian cells, due to disruption of biological membranes and/or targeting of the Hsp90 chaperone, limited optimization. To address this limitation, we modified the DPP scaffold and identified PYO12, a phenoxy-phenyl-pyrazol derivative with low haemolytic activity against red blood cells. Further optimization led to PYO12a which showed no cytotoxicity against HEK293 cells and only low cytotoxicity against HepG2 cells.
The increased permeability observed in the SYTOX Green assay suggested that PYO12 causes membrane damage to S. aureus. To obtain further evidence that the mechanism of action was related to cell wall or membrane disruption, we quantified expression of genes involved in responses to cell-wall targeting antibiotics. Expression analysis revealed activation of both BraRS and VraRS TCS by PYO12. Moderate upregulation of the VraRS regulon suggests that PYO12 causes cell-wall stress, while strong upregulation of vraDEH, similar to that reported for nisin treatment26 indicates activation of BraRS, a sensor of lipid II binders. These data suggest that PYO12 interacts with lipid II or other lipid intermediate22. This hypothesis is reinforced by a 16-fold over-expression of liaI in B. subtilis, a gene specifically induced by the LiaRS membrane stress regulator in presence of lipid II binders25. Although the LiaRS regulon is also induced in by organic solvents25 the low haemolytic activity of PYO12 suggests that it does not disrupt biological membranes in a similar fashion. It is noteworthy that previous studies in S. aureus treated with novobiocin (bacterial gyrase inhibitor) did not lead to over-expression of any of the aforementioned genes27.
Furthermore, competition with 2 M excess of free undecaprenyl pyrophosphate (C55-PP) significantly increased the MIC of PYO12 for S. aureus, suggesting that part of PYO12 binds to C55-PP. Binding to lipid C55-P was not found, indicating that the pyrophosphate moiety is critical for binding. Similar results were reported for the lipid II targeting small molecules THCz and DCAP16,17. These findings suggests that PYO12 binds to C55-PP either directly as a lipid intermediate or as part of lipid II, although direct binding assays are required to confirm lipid II interaction.
PYO12 displayed potent activity against a panel of Gram-positive pathogens (E. faecalis vanA+, E. faecium vanA+, vanB+ and vanAvanB+ and MRSA), showing no cross-resistance with lipid II binders vancomycin or methicillin. PYO derivatives had no effect on Gram-negative E. coli and P. aeruginosa. Improved susceptibility in E. coli ΔtolC mutant indicated that efflux pumps play a major role in the lack of activity in Gram-negative bacteria. However, we cannot discard increased sensitivity may be due to other of the pleiotropic effects that can be caused by deletion of TolC. Damage to the inner membrane was also seen in E. coli ΔtolC, strengthening the hypothesis that PYO12 specifically disrupts the inner membrane, possibly by binding to lipid II or its intermediates. Time-kill curves showed a dose-dependent bactericidal effect in S. aureus, adding to the attractive properties of this compound.
Resistant mutants revealed a deletion in saeR that would prevent phosphorylation of this regulator due to the absence of the conserved aspartic acid D5128. SaeR regulon is activated in presence of β-lactam antibiotics29 and regulate expression of haemolysin and biofilm formation30. A truncation was found in ybjH, an adaptor protein that promotes proteolysis of Spx regulator31. Spx is known to play an important role in β-lactam and vancomycin resistance32,33. A non-functional YbjH would lead to an overabundance of Spx and upregulation of genes involved in resistance to cell-wall and membrane damaging antibiotics, consistent with PYO12 interacting with bacterial membrane. The importance of both genes in the normal virulence and metabolism of S. aureus also suggest that in vivo these mutants would be compromised during an infection.
Among the 14 analogues synthesized, only PYO12a had improved biological properties, with very low cytotoxicity. The selectivity index of PYO12a (S.I. =36) is still below that of clinically relevant antimicrobials such as novobiocin (S.I. = 115, Table 1) so further optimization is necessary. Nevertheless, PYO12a shows a substantially improved selectivity index compared to previously reported lipid II targeting small molecules (S.I. 1.2 in DCAP17 and 4 THCz16. Modifications on the aromatic rings of PYO12 led to large variations in MIC and cytotoxicity, making the SAR difficult to analyse. As most analogues showed some cytotoxicity, further toxicity studies on different cell types or organoids are needed for PYO12a prior to progressing to in vivo experimental models. As these structures are analogues of published DPPs19 inhibitors of Hsp90, it is possible that inhibition of Hsp90 is still retained by some of the presented structures. However, the parent molecule PYO12 did not lead to inhibition of Hsp90 in our assays.
In conclusion, we hypothesize that the small molecule PYO12 binds to undecaprenyl pyrophosphate containing molecules on the bacterial membrane, leading to membrane disruption and bacterial death. Whether PYO12 binding to lipid intermediates leads to membrane disruption through pore formation or by intercalation in the membrane is still unknown. The limited activity of PYO12 against Gram-negative bacteria likely results from efflux from the periplasm before reaching the inner membrane. PYO12a, a derivative with similar structure to PYO12, exhibited low cytotoxicity for mammalian cells. The discovery of small molecules such as PYO12a, which may bind to lipid II or other lipid intermediates while maintaining low mammalian cytotoxic, represents an exciting finding that can open new paths for antibiotic development.
Materials and methods
Chemical compounds
Chemicals PYO1-22 and PYO12e, PYO12f, PYO12i and PYO12l were purchased from MolPort (MolPort, Riga, Latvia) in powder form and were > 95% pure by HPLC. Synthesis schemes, purity and characterization of synthetized PYO23, PYO24, PYO12, PYO12a, PYO12b, PYO12c, PYO12d, PYO12g, PYO12h, PYO12j, PYO12k, PYO12m, PYO12n can be found in Supplementary Data (Supplementary Figures SF2-SF13). Synthesis was performed by O2H group (Oxford). All compounds were resuspended in dimethyl-sulfoxide (DMSO) to a final concentration of 50 mg/ml. Antibiotics vancomycin, novobiocin and nisin were obtained from Sigma Aldrich.
Bacterial strains and growth conditions
Staphylococcus aureus str. Newman, S. aureus USA300 (Methicillin Resistant S. aureus, MRSA), Enterococcus faecalis (strain E2440), E. faecalis vanA+ (strain E1654), Enterococcus faecium (strain E1162), E. faecium vanA+ (strain E0155), E. faecium vanB+ (strain E7422), E. faecium vanAvanB+ (strain E8426), Bacillus subtilis str. 168 1A700, Escherichia coli ATCC 25,922, E. coli D21f2, E. coli JW550334 and Pseudomonas aeruginosa ATCC 27,853 were grown in Mueller-Hinton Broth (MHB) (Oxoid, Basingstoke, UK) and incubated at 37 ˚C. Streptococcus suis P1/7 and Streptococcus pneumoniae A66 were cultured in THY media (Todd-Hewitt Broth (Oxoid, Basingstoke, UK) + 0.2% yeast extract (BD Biosciences)) at 37 ˚C and 5% CO2.
Minimal inhibitory concentration (MIC)
The MIC was determined using the microdilution method according to the recommendations of the European committee on antimicrobial susceptibility testing (EUCAST)35. Compounds were diluted in 96-well plates (two-fold dilution range 50–0.39 µg/ml) in MHB (THY for S. suis and S. pneumoniae) in a final volume of 100 µl per well. Bacteria were added as a 1:100 dilution of an overnight culture, 100 µl per well to a final volume of 200 µl per well. Controls were added for bacteria with and without 1% DMSO (vehicle control), and medium alone was added to control for contamination. Plates were incubated for 18 h at 37 ˚C or 37 ˚C and 5% CO2 for S. suis and S. pneumoniae. The lowest concentration where no bacterial growth was detected by optical density at 600 nm (OD600) was recorded as MIC.
Cytotoxicity using alamar blue assay
Human embryonic kidney cells (HEK293, ATCC-CRL-1573) and human hepatocellular carcinoma cells (HepG2, ATCC-HB-8065) were obtained from the American Type Culture Collection (ATCC). Cells were seeded in 96-well plates at a density of 50.000 cells/well in Dulbecco’s Modified Eagle Medium (DMEM) containing glutaMax and phenol red (Gibco) supplemented with 1% penicillin/streptomycin (Gibco) and 10% foetal bovine serum (FBS) (Gibco). Cells were incubated at 37 °C and 5% CO2. Exterior wells were filled with PBS to avoid evaporation issues. Once cells reached a confluency of 80–90%, culture media was removed and replaced with 200 µl of exposure media (DMEM without phenol red and antibiotics (Gibco)) containing a 2-fold dilution series of the test compounds (range 50–1.56 µg/ml). FBS was omitted from the culture medium to avoid interaction with the tested compounds. Controls containing only exposure medium, exposure medium with 1% DMSO (vehicle control), exposure medium with 20% DMSO and wells containing no cells were included. After 24-hour exposure at 37 °C and 5% CO2, the exposure medium was replaced by 100 µl 10% Alamar Blue (Invitrogen) in exposure medium. Fluorescence (λex = 541 nm, λem = 590 nm) was measured after 45-minutes exposure using Spectramax M5 (Molecular Devices LLC, San Jose, CA, USA). Cell viability compared to the vehicle control was calculated and inhibition curves for each compound were fitted (non-linear curve fit, four variables, bottom constrained = 0) using Prism 9 (Graphpad Software, San Diego, USA). The IC50 value for 50% cell viability was calculated based on the fitted curves.
Haemolysis
Haemolysis protocol was adapted from Evans et al.36. Briefly, sheep red blood cells (RBC) 10% washed pool cells in PBS (Rockland Immunochemicals, Limerick, PA, USA) were diluted in PBS to a final concentration of 2% RBCs, and 200 µl were exposed to 500, 250 and 125 µM of compound PYO12 for 30 min. Non-treated control, 1% DMSO (vehicle control) and complete cell lysis control (Triton-X 2%) were included. Cells were centrifuged 1000 g for 15 min at 4 °C, then 100 µl transferred to a 96-well plate and absorption at 540 nm measured using Spectramax M5. Percentage of haemolysis was calculated:
Membrane permeability using SYTOX-green
Five hundred microliters of S. aureus str. Newman in exponential phase in MHB (OD600 = 0.4) were incubated for 5 min with 2, 1 and 0.5 µg/ml of PYO12. Nisin was included as a positive control (1x MIC = 6.25 µg/ml) and novobiocin (1xMIC = 0.5 µg/ml), which does not damage the membrane, as a negative control. Non-treated bacteria were also included as negative control. For E. coli ΔtolC colistin (1xMIC = 3 µg/ml) was used as positive control. After treatment, cells were centrifuged 4.000 r.p.m. for 5 min, resuspended in filtered PBS and dyed for 5 min in the dark with 0.5 µM of SYTOX® Green (Molecular Probes, Invitrogen). Cells were centrifuged and resuspended with PBS to remove unbound dye. Three hundred microliters of sample were transferred to a 96-well clear-bottom plate (Corning Incorporated). Fluorescence was measured using a CytoFlex® flow cytometer (Beckman Coulter, Brea, CA, USA) using the green (FITC) channel with the excitation wavelength at 488 nm. Overlay histograms were created with CytExpert Software (Bechman Coulter, Brea, CA, USA).
Time-kill curve
S. aureus str. Newman was grown to exponential phase (OD600 = 0.25, read in 96-well plate) in MHB media. Five millilitres of exponential culture were treated with 8, 4, 2–1 µg/ml of PYO12, a non-treated control was added. At 0, 10, 30 min, 1 h, 2 h,3 h, 4 h and 6 h post-treatment, 20 µl were transferred to 180 µl sterile PBS and 1:10 serial dilutions were made in PBS (dilution factors: 1:10 to 1:106). Five microliters of each dilution were plated in MHB plates. Colonies were counted following overnight incubation at 37 ˚C. Three independent experiments were performed.
Relative expression using qPCR
S. aureus str. Newman or Bacillus subtilis str. 168 1A700 were grown to exponential phase (OD600 = 0.4) in 50 ml MHB media at 37 °C. Bacteria were divided into four 15 ml Falcon tubes (Corning Incorporated) with 6 ml exponential culture per tube. Two tubes were supplemented with 1% DMSO (vehicle control) and two with 1 µg/ml of PYO12 for both S. aureus and B. subtilis. After 10 min at 37 °C cells were centrifuged 4.000 r.p.m. for 10 min at 4 °C, the supernatant was discarded, and the pellet immediately frozen in liquid nitrogen and stored at -80 °C until RNA was extracted.
RNA from the pellet was extracted using RNeasy Mini Kit (QIAGEN) using manufacturer protocol with slight adaptations. Frozen pellet was resuspended in 700 µl of RLT buffer containing 0.1% β-mercaptoethanol. The resuspended pellet was transferred to lysing matrix B 2 ml tubes (MP Biomedicals) and lysis was carried out in a Fast-Prep 24 5G bead beating grinder and lysis system (MP Biomedicals) using the following parameters: All-MetalQuickprep, 2 cycles, 40 s, 6.0 m/sec. Lysing matrix B tubes were centrifuged at 10.000 r.p.m. for 30 s, supernatant transferred to a clean 1.5 ml Eppendorf tube and manufacturer protocol was followed. RNA concentration was determined using Qubit RNA broad range (BR) kit (Invitrogen) in Qubit 4 fluorometer (Invitrogen). cDNA was synthesized from 1 µg RNA using QuantiTect reverse transcription kit (QIAGEN).
Quantitative PCR for relative gene expression was performed in 96-well clear PCR plates (Bio-Rad) and GoTaq qPCR Master Mix (Promega) in a CFX96 Real-Time system (Bio-Rad). Primers (Supplementary Table ST1) were designed using Benchling to amplify 100–200 bp amplicons of the gene of interest. Relative gene expression was calculated using the 2−ΔΔCt method37 and data for the gyrA gene for normalisation. The experiment, including replicates (n = 2) was independently repeated twice.
Undecaprenyl pyrophosphate antagonist assay
Antagonist assays were performed as reported previously17 with small adaptations. Briefly, undecaprenyl-pyrophosphate (C55-PP) and monophosphate (Larodan, Solna, Sweden) was mixed with compound PYO12 in molar ratios of 1:0.5, 1:1, 1:2 and 1:4 (PYO12:C55-PP) and incubated for 30 min at room-temperature. An MIC test was then performed using the mixed PYO12:C55-PP as starting compound, with concentrations of PYO12 ranging from 8 to 0.125 µg/ml. A control including only C55-PP was included to ensure that the lipid has no anti-bacterial effect. An extra control was added with C55-PP solvent, chloroform: methanol 2:1. Experiments were independently performed in triplicate.
Resistance development assays
We assessed the emergence of spontaneous resistance by plating 1011 CFU of S. aureus Newman onto MHB plates containing 5x and 10x MIC concentrations of PYO12 (5 and 10 µg/ml, respectively) and incubated overnight at 37 °C. Emergence of resistant mutants exposed to sub-inhibitory concentrations of PYO12 was assessed as follows: tubes containing 2 ml of MHB media supplemented with 0.5, 1 and 2x MIC of PYO12 were inoculated 1:100 with an overnight culture of S. aureus and incubated overnight at 37 °C. The culture that grew at the highest concentration of PYO12 was then subcultured into tubes containing 0.5, 1, 2x MICPreviousday. This was repeated every day until resistance arose. Whole genome sequencing of these resistant cultures was provided by MicrobesNG (https://www.microbesng.com) using Illumina-sequencing. Variant analysis was performed using Galaxy (https://usegalaxy.org/) VarScan tool38. VCF files were deposited in the European Variation Archive (EVA) at EMBL-EBI (https://www.ebi.ac.uk/eva/) under accession number PRJEB86156.
Data availability
The dataset analysed during the current study are available in the EMBL-EBI-EVA (European Variation Archive) repository, under accession number PRJEB86156.
References
Antimicrobial Resistance, C. Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis. Lancet 399, 629–655. https://doi.org/10.1016/S0140-6736(21)02724-0 (2022).
Bugg, T. D., Braddick, D., Dowson, C. G. & Roper, D. I. Bacterial cell wall assembly: still an attractive antibacterial target. Trends Biotechnol. 29, 167–173. https://doi.org/10.1016/j.tibtech.2010.12.006 (2011).
Bouhss, A., Trunkfield, A. E., Bugg, T. D. & Mengin-Lecreulx, D. The biosynthesis of peptidoglycan lipid-linked intermediates. FEMS Microbiol. Rev. 32, 208–233. https://doi.org/10.1111/j.1574-6976.2007.00089.x (2008).
Storm, D. R. & Strominger, J. L. Binding of bacitracin to cells and protoplasts of micrococcus lysodeikticus. J. Biol. Chem. 249, 1823–1827. https://doi.org/10.1016/s0021-9258(19)42861-5 (1974).
Kramer, N. E. et al. Resistance of Gram-positive bacteria to Nisin is not determined by lipid II levels. FEMS Microbiol. Lett. 239, 157–161. https://doi.org/10.1016/j.femsle.2004.08.033 (2004).
Breukink, E. & de Kruijff, B. Lipid II as a target for antibiotics. Nat. Rev. Drug Discov. 5, 321–332. https://doi.org/10.1038/nrd2004 (2006).
Perkins, H. R. Specificity of combination between mucopeptide precursors and Vancomycin or Ristocetin. Biochem. J. 111, 195–205. https://doi.org/10.1042/bj1110195 (1969).
Arthur, M. & Courvalin, P. Genetics and mechanisms of glycopeptide resistance in enterococci. Antimicrob. Agents Chemother. 37, 1563–1571. https://doi.org/10.1128/AAC.37.8.1563 (1993).
Grein, F., Schneider, T. & Sahl, H. G. Docking on lipid II-A widespread mechanism for potent bactericidal activities of antibiotic peptides. J. Mol. Biol. 431, 3520–3530. https://doi.org/10.1016/j.jmb.2019.05.014 (2019).
Storm, D. R. & Strominger, J. L. Complex formation between bacitracin peptides and isoprenyl pyrophosphates. J. Biol. Chem. 248, 3940–3945. https://doi.org/10.1016/s0021-9258(19)43823-4 (1973).
Watkinson, R. J., Hussey, H. & Baddiley, J. Shared lipid phosphate carrier in the biosynthesis of teichoic acid and peptidoglycan. Nat. New. Biol. 229, 57–59. https://doi.org/10.1038/newbio229057a0 (1971).
Troy, F. A., Frerman, F. E. & Heath, E. C. The biosynthesis of capsular polysaccharide in aerobacter aerogenes. J. Biol. Chem. 246, 118–133. https://doi.org/10.1016/s0021-9258(18)62540-2 (1971).
Hiron, A., Falord, M., Valle, J., Debarbouille, M. & Msadek, T. Bacitracin and Nisin resistance in Staphylococcus aureus: a novel pathway involving the bras/brar two-component system (SA2417/SA2418) and both the brad/brae and vrad/vrae ABC transporters. Mol. Microbiol. 81, 602–622. https://doi.org/10.1111/j.1365-2958.2011.07735.x (2011).
Muller, A., Klockner, A. & Schneider, T. Targeting a cell wall biosynthesis hot spot. Nat. Prod. Rep. 34, 909–932. https://doi.org/10.1039/c7np00012j (2017).
Chauhan, J., Kwasny, S. M., Fletcher, S. & Opperman, T. J. Leeuw, E. P. H. Optimization of a small-molecule lipid II binder. Bioorg. Med. Chem. Lett. 29, 1849–1853. https://doi.org/10.1016/j.bmcl.2019.04.046 (2019). de.
Ludwig, K. C. et al. The dual mode of antibacterial action of the synthetic small molecule DCAP involves lipid II binding. J. Am. Chem. Soc. 146, 24855–24862. https://doi.org/10.1021/jacs.4c05138 (2024).
Reithuber, E. et al. THCz: small molecules with antimicrobial activity that block cell wall lipid intermediates. Proc. Natl. Acad. Sci. USA 118 https://doi.org/10.1073/pnas.2108244118 (2021).
Malin, J., Shetty, A. C., Daugherty, S. C. & de Leeuw, E. P. Effect of a small molecule lipid II binder on bacterial cell wall stress. Infect. Drug Resist. 10, 69–73. https://doi.org/10.2147/IDR.S126254 (2017).
Fernandez-Ciruelos, B. et al. Repurposing Hsp90 inhibitors as antimicrobials targeting two-component systems identifies compounds leading to loss of bacterial membrane integrity. Microbiol. Spectr. e0014624 https://doi.org/10.1128/spectrum.00146-24 (2024).
Dengler, V., Meier, P. S., Heusser, R., Berger-Bachi, B. & McCallum, N. Induction kinetics of the Staphylococcus aureus cell wall stress stimulon in response to different cell wall active antibiotics. BMC Microbiol. 11, 16. https://doi.org/10.1186/1471-2180-11-16 (2011).
Balibar, C. J. et al. cwrA, a gene that specifically responds to cell wall damage in Staphylococcus aureus. Microbiol. (Reading). 156, 1372–1383. https://doi.org/10.1099/mic.0.036129-0 (2010).
Kawada-Matsuo, M. et al. Three distinct two-component systems are involved in resistance to the class I bacteriocins, Nukacin ISK-1 and Nisin A, in Staphylococcus aureus. PLoS One. 8, e69455. https://doi.org/10.1371/journal.pone.0069455 (2013).
Kuroda, M. et al. Two-component system VraSR positively modulates the regulation of cell-wall biosynthesis pathway in Staphylococcus aureus. Mol. Microbiol. 49, 807–821. https://doi.org/10.1046/j.1365-2958.2003.03599.x (2003).
Meehl, M., Herbert, S., Gotz, F. & Cheung, A. Interaction of the GraRS two-component system with the VraFG ABC transporter to support vancomycin-intermediate resistance in Staphylococcus aureus. Antimicrob. Agents Chemother. 51, 2679–2689. https://doi.org/10.1128/AAC.00209-07 (2007).
Mascher, T., Zimmer, S. L., Smith, T. A. & Helmann, J. D. Antibiotic-inducible promoter regulated by the cell envelope stress-sensing two-component system liars of Bacillus subtilis. Antimicrob. Agents Chemother. 48, 2888–2896. https://doi.org/10.1128/AAC.48.8.2888-2896.2004 (2004).
Zhao, X. et al. Analysis of the gene expression profile of Staphylococcus aureus treated with Nisin. Food Control. 59, 499–506. https://doi.org/10.1016/j.foodcont.2015.06.018 (2016).
Schroder, W. et al. Altering gene expression by aminocoumarins: the role of DNA supercoiling in Staphylococcus aureus. BMC Genom. 15, 291. https://doi.org/10.1186/1471-2164-15-291 (2014).
Mainiero, M. et al. Differential target gene activation by the Staphylococcus aureus two-component system SaeRS. J. Bacteriol. 192, 613–623. https://doi.org/10.1128/JB.01242-09 (2010).
Kuroda, H., Kuroda, M., Cui, L. & Hiramatsu, K. Subinhibitory concentrations of beta-lactam induce haemolytic activity in Staphylococcus aureus through the SaeRS two-component system. FEMS Microbiol. Lett. 268, 98–105. https://doi.org/10.1111/j.1574-6968.2006.00568.x (2007).
Gao, P. et al. SaeR as a novel target for antivirulence therapy against Staphylococcus aureus. Emerg. Microbes Infect. 12, 2254415. https://doi.org/10.1080/22221751.2023.2254415 (2023).
Engman, J., Rogstam, A., Frees, D. & Ingmer, H. Wachenfeldt, C. The YjbH adaptor protein enhances proteolysis of the transcriptional regulator Spx in Staphylococcus aureus. J. Bacteriol. 194, 1186–1194. https://doi.org/10.1128/JB.06414-11 (2012). von.
Nielsen, T. K. et al. The Spx stress regulator confers high-level beta-lactam resistance and decreases susceptibility to last-line antibiotics in methicillin-resistant Staphylococcus aureus. Antimicrob Agents Chemother, e0033524 https://doi.org/10.1128/aac.00335-24 (2024).
Jousselin, A., Kelley, W. L., Barras, C., Lew, D. P. & Renzoni, A. The Staphylococcus aureus thiol/oxidative stress global regulator Spx controls trfa, a gene implicated in cell wall antibiotic resistance. Antimicrob. Agents Chemother. 57, 3283–3292. https://doi.org/10.1128/AAC.00220-13 (2013).
Baba, T. et al. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol. Syst. Biol. 2, 0008 https://doi.org/10.1038/msb4100050 (2006).
EUCAST. https://www.eucast.org/ast_of_bacteria/mic_determination (2022).
Evans, B. C. et al. Ex vivo red blood cell hemolysis assay for the evaluation of pH-responsive endosomolytic agents for cytosolic delivery of biomacromolecular drugs. J. Vis. Exp. e50166 https://doi.org/10.3791/50166 (2013).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 25, 402–408 https://doi.org/10.1006/meth.2001.1262 (2001).
Koboldt, D. C. et al. VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 22, 568–576. https://doi.org/10.1101/gr.129684.111 (2012).
Acknowledgements
We would like to thank the Utrecht Medical Centre (UMC) group from Prof. Rob Willems for providing the enterococcal isolates. We would also like to thank to O2H Group, specially David Davies, Ashish Patel, Hemal Soni, Vinod Bhoge and Claudio Dagostin for the design and synthesis of compounds PYO12a-PYO12n.
Author information
Authors and Affiliations
Contributions
P.F. and J.M. acquired funding, conceptualized the studies, supervised and provided needed resources, B.F.C. and M.A. conceived, conducted and analyzed the experiments, F.T. supervised and help in conceptualization, B.F.C wrote the first draft and all authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fernandez-Ciruelos, B., Albanese, M., Taverne, F. et al. Novel antimicrobial 3-phenyl-4-phenoxypyrazole derivatives target cell wall lipid intermediates with low mammalian cytotoxicity. Sci Rep 15, 35646 (2025). https://doi.org/10.1038/s41598-025-19561-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-19561-y