Abstract
Lupus nephritis (LN) is a common manifestation of systemic lupus erythematosus (SLE), affecting about half of the patients. This research uses genotype–phenotype associations to identify genetic markers, enhancing our understanding of LN pathogenesis. This study used genotype–phenotype association to explore genetic variants linked to LN in a Taiwanese cohort from the Taiwan Precision Medicine Initiative (TPMI). Genotyping was performed on 244 SLE and 63 LN patients using the TPM array. Genomic DNA underwent quality control via the Axiom Analysis Suite. Chi-squared tests identified significant variants, which were annotated and assessed for functional impacts using the 1000 Genomes Project, gnomAD, enabling comprehensive genetic analysis. This study on a Taiwanese cohort (LN, SLE, LN/SLE) identified four significant genetic variants (p-values < 10−6) associated with LN: rs1025129 in HGF, rs80282109 in BACH2, rs516119 in SOX1, and rs134545 in TTC28. These variants may affect LN pathogenesis through splicing junction changes and gene expression regulation, highlighting their potential as biomarkers. Our research identified four novel variants (rs134545, rs516119, rs1025129, rs80282109) in LN patients through a genotype–phenotype study on a Taiwanese population, suggesting their roles in LN pathogenesis and supporting their potential as new disease biomarkers.
Similar content being viewed by others
Introduction
Lupus nephritis (LN) is a common manifestation of systemic lupus erythematosus (SLE), affecting over 50% of patients and typically developing within the first five years after SLE diagnosis1,2,3. LN arises from an inflammatory response to immunogenic chromatin, like oxidized mitochondrial DNA and nucleic acids in exosomes. Accumulated chromatin activates DNA/RNA sensors (TLR7, TLR8, TLR9, RIG1/MDA5–MAVS, CGAS–STING), leading to increased type I interferon and cytokines4,5,6,7. Genetic polymorphisms (e.g., BANK1, BLK, TLR7) persistently activate B cells and autoantibodies production8,9. Complement proteins also play a role, with reduced circulating levels and intra-renal deposition leading to LN10.
Several studies in Taiwan have identified genetic variants that contribute to LN. A cross-sectional analysis pointed out that in the Taiwanese population, the presence of HLA-DRB10301 and DRB11501 alleles significantly increases the risk of SLE, while the presence of DRB1*1202 allele appears to be protective against LN11. Another study identified specific genetic variants in the IFN-λ3/4 genes, including notable variants—rs8099917, rs12979860, rs4803217, and rs469415590—that significantly increase the risk of developing LN in patients with SLE12.
Our study aims to showcase the utility of genotype–phenotype association study in predicting and understanding LN through data from participants in the Taiwan Precision Medicine Initiative (TPMI)13. We finally highlighted four variants—rs1025129, rs80282109, rs516119, and rs134545—that were associated with LN. Additionally, we characterized variants rs80282109 (in the BACH2 gene intron) and rs1025129 (in the upstream region of the HGF gene), as BACH2 and HGF are crucial for immune cell function. These findings provide significant genetic insights into LN risk variants within the Taiwan Han population.
Materials and methods
Ethics
This study utilized genome typing data from participants enrolled in accordance with the regulations of the Tri-Service General Hospital Institutional Review Board (IRB), under approval number 2-108-05-038. The organization and operation of the IRB comply with Good Clinical Practice (GCP) and the applicable laws and regulations. All participants provided written informed consent, allowing the collection and analysis of their data. Appropriate security measures will be implemented to safeguard the data, and all research team members are required to strictly uphold the confidentiality of identifiable information or establish relevant mechanisms. All investigators must report any Serious Adverse Events and Unanticipated Problems in accordance with governmental laws and regulatory requirements.
Participants enrolled in study
The study participants were recruited from Tri-Service General Hospital (TSGH) as part of the Taiwan Precision Medicine Initiative (TPMI), a collaborative effort between Academia Sinica and 16 leading medical centers in Taiwan. The primary objective of TPMI is to establish a comprehensive database containing detailed clinical information and genetic profiles of one million individuals from the Taiwanese Han population. In total, 27,529 participants were recruited through TPMI and underwent genotyping conducted by Academia Sinica. Participants were enrolled from various medical centers. Table 1 summarizes the age and sex distribution of TSGH participants included in this study, comprising individuals with lupus nephritis (LN), systemic lupus erythematosus (SLE), and controls. For the control group, participants were eligible if they had no autoimmune diseases, although other medical conditions were permitted. Exclusion criteria included a history of blood transfusion, leukemia, lymphoma, or prior chemotherapy.
For the experimental group, 244 individuals were diagnosed with SLE through diagnostic interviews conducted by eight clinical rheumatologists, following the 2019 European Alliance of Associations for Rheumatology (EULAR)/American College of Rheumatology (ACR) classification criteria. Among them, 63 individuals were identified as having lupus nephritis (LN), based on the presence of the renal disorder variable defined in the ACR criteria and/or biopsy confirmation according to the standards set by the International Society of Nephrology/Renal Pathology Society (ISN/RPS). All participants in the experimental group, similar to those in the control group, had no history of blood transfusion, leukemia, lymphoma, or chemotherapy.
Genotyping by TPM array
The process of DNA extraction and identification of nucleotide mutation in this study followed the following steps and these steps allowed researchers to analyze genetic variations and understand their association with various health conditions.: (1) DNA Extraction: Genomic DNA was extracted and purified from 3 mL of peripheral blood collected in EDTA vacutainers using the QIA symphony SP system by QIAGEN (Hilden, Germany). (2) Genotyping: Each participant’s purified DNA was loaded onto the Taiwan Precision Medicine (TPM) array chip. Genome type signals produced from the TPMI array were detected using the Axiom GeneTitan platform by Thermo Fisher Scientific (Sunnyvale, CA, USA). (3) Quality Control and Annotation: Genome SNP data quality control (QC), SNP calling, and sample annotations were performed using the Axiom Analysis Suite by Thermo Fisher Scientific (Sunnyvale, CA, USA).
Association study
Figure 1 outlines the steps involved in the genotype–phenotype association analysis. Initially, we analyzed genotyping data from the TPM array. SNPs (total 493,852 SNPs) obtained from TPM array with a low typing call rate (< 80%) were filtered out. The remaining SNP and phenotype data, comprising LN (n = 63), SLE (n = 244), and control (n = 27,529) groups, underwent association analysis using the chi-squared method with PLINK 1.9 software (https://zzz.bwh.harvard.edu/plink/)14. To find out different variants between SLE and LN, three traits were designed as LN (only LN patients together compared to control), SLE (only SLE patients together compared to control), and LN/SLE (LN patients compared to SLE patients without LN). We removed variants of low quality (minor allele frequency less than 0.05 and Hardy–Weinberg equilibrium less than 1 × 10–5) and selected variants with highly significant p-values (less than 1 × 10–6) for further investigation.
Genotype–phenotype association analysis pipeline in this study. Case groups (LN: 63 patients and SLE: 244 patients) and control group (other 27,529 participants) underwent genotyping identified by TPM array chip. Data from 493,852 SNPs were filtered, and 239,080 SNPs that passed were applied to chi-squared test for detecting risk factors.
Variant annotations and functional analysis
Significant variants related genes were annotated by utilizing the RefSeq Database (https://www.ncbi.nlm.nih.gov/refseq/), as described in wANNOVAR (https://wannovar.wglab.org/)15, a powerful tool for annotating functional consequences of genetic variations. To assess allele frequencies across diverse racial populations, we turned to publicly available databases:
-
1.
1000 Genomes Project16: This comprehensive resource catalogs common human genetic variation, drawing from openly consented samples of healthy individuals.
-
2.
Genome Aggregation Database (gnomAD)17: An international collaboration that harmonizes exome and genome sequencing data from various large-scale projects, making summary data accessible to the scientific community.
-
3.
Taiwan BioBank (https://taiwanview.twbiobank.org.tw/index): A locally curated human biobank in Taiwan integrates lifestyle, environmental factors, clinical medicine, and biological markers to create a foundation for personalized medicine and health in Taiwan18.
Results
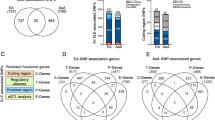
According to the genotype–phenotype association results, eight significant variants (p-value < 10–6) were identified including in the LN, SLE, and LN/SLE group when comparing each genotype–phenotype association patterns in the Fig. 2. The Q–Q plot depicted in Fig. S1A, notable variant points lie above the diagonal line, indicating a potential association with diseases. The genomic inflation factor, lambda, calculated from the median of observed and expected chi-square values, approaches 1, suggesting negligible inflation. This indicates the robustness of our test statistics, allowing for confident interpretation of the results. Additional quality control analyses, including principal component analysis (PCA) for assessing population stratification, are shown in Fig. S1B. The PCA plot demonstrates no distinct clustering patterns, suggesting the absence of population stratification among the study samples. To assess potential kinship relationships among samples, IBD analysis was performed. No sample exhibited a low IBS genetic distance (< 0.1) to any other, indicating that no significant relatedness or kinship was detected among the experimental samples. Finally, we tested individual genotype success rates at thresholds of 0.01, 0.02, 0.03, 0.04, 0.05, 0.1, 0.2, 0.3, 0.4, and 0.5. After applying a missing genotype rate threshold of 0.01, 95.8% of samples remained (Fig. S2). Consequently, we affirm that our genotype–phenotype association findings are minimally biased. After searching genotype–phenotype association catalog database, variants rs73366469 and rs117026326 showed records of associations with SLE19,20,21 and other variants did not have any association to SLE or immune disease in the genotype–phenotype association catalog.
Manhattan plot of genotype–phenotype association results in four traits. There were 239,080 variants detected in TSGH TPMI participants through a chi-squared test. Eight highly significant variants were selected according to the p-value < 10–6 (Bold points A-I on the plot) : rs12508616, rs2027856, rs117026326, rs73366469, rs76267797, rs80282109, rs516119, and rs134545. Variants with star symbol indicate records of associations with SLE in genotype–phenotype association catalog database. Circle from the inside to outside: LN, SLE, and LN/SLE.
Annotation analysis of these variants in Table 2 showed that four LN related variants, rs1025129, rs80282109, rs516119, and rs13454, were characterized in the different gene locations such as BACH2 (BTB Domain And CNC Homolog 2), CACNA2D1 (Calcium Voltage-Gated Channel Auxiliary Subunit Alpha2delta 1), HGF (Hepatocyte growth factor), SOX1(SRY-Box Transcription Factor 1), and TTC28 (Tetratricopeptide Repeat Domain 28). All variants demonstrated highly significant p-values, and the p-values adjusted using the Genomic Control (GC) method (Reference) also remained below 10⁻6. It is interesting that variants rs516119 presented negative effects (Odds Ratio < 1) to the LN, indicating that this particular genetic variant is less likely to develop the disease. Furthermore, when compared to each variant allele frequency in different populations in Table 3, SLE related variants (rs117026326, rs76267797, and rs73366469) showed high frequency not only in Taiwanese population (TPMI and Taiwan biobank) but also in Asian. When comparing LN-related variants, their allele frequency showed lower rate in the Taiwanese or Asian population compared to others, reflects the complex interplay of genetic diversity in Taiwan or Asia LN disease. Further investigation of the detailed allele frequency of variants in participants, as shown in Fig. 3, revealed that the LN-associated variants are specific to SLE and controls and may possibly be causal genetic factors in Taiwanese LN patients. Allele frequency of rs1025129, rs802809, and rs134545 in LN patients were higher than SLE and control. Moreover, a specific variant rs73366469 and rs117026326 in the SLE patients has similar frequency in both LN patients and these two variants could be risk factors to both LN and SLE.
Mutation frequency of significant variants on each group. Mutation frequency was presented as green to red color from low to high. LN and SLE group showed similar frequency in the SLE trait. Variants rs134545, rs516119, rs1025129, and rs80282109 with high frequency were observed in the LN group in LN and LN/SLE trait.
According to the genetic location and gene structure of the variants, LN-related variants support changes in genes through exon–intron splicing and gene expression regulation. Figure S3 shows that the genetic locations of two LN-related variants, rs80282109 and rs134545, are in the intronic region of BACH2 and TTC28, respectively. The other two variants, rs1025129 and rs516119, located in the upstream region of HGF and SOX1.
Discussion
LN presents significant clinical challenges due to its prevalence and complex treatment. Careful management is needed to mitigate long-term complications. Genetic factors greatly influence SLE and LN pathogenesis, with first-degree relatives of SLE patients having a significantly increased risk: 316 times higher for twins, 23 times for siblings, and 11 times for parents22. Over 100 susceptibility loci linked to SLE and LN involve genes related to lymphocyte function, immune signaling, DNA clearance, complement pathway, and renal injury23,24. Known mechanisms may indirectly influence LN development, despite the lack of direct proof connecting specific SNPs/genes to LN22.
Abundant studies in Taiwan have investigated various genetic loci among SLE patients. A study analyzed 2429 SLE patients and 48,580 controls in Taiwan using GWAS and polygenic risk scores (PRS). Multiple PRS models combining SLE, ANA, anti-dsDNA, and anti-Sm markers improved SLE prediction (AUC 0.64). Novel genes and HLA haplotypes (HLA-DQA101:01, HLA-DQB105:01) were also identified, aiding early diagnosis and precision medicine25. Another study investigated six SNPs in TLR3, TLR7, and TLR8 genes in 795 Taiwanese SLE patients and 1162 controls. TLR7 rs3853839-G was significantly associated with SLE susceptibility in females and several clinical features. TLR8 rs3764880-G was linked to oral ulcers and pericardial effusion. TLR3 rs3775296-T was associated with photosensitivity and anemia. Certain TLR7/TLR8 haplotypes influenced SLE risk, suggesting TLR variations contribute to SLE susceptibility and phenotypic diversity26.
The variant rs1025129, located in the upstream regulatory region of the HGF gene, is crucial due to the role of HGF in tissue regeneration and anti-inflammatory responses. HGF interacts with the c-Met receptor, activating pathways like PI3-kinase and STAT-327,28,29. HGF also promotes podocyte migration and epithelial-to-mesenchymal transformation, which is essential for renal injury repair in LN30. This variant may modulate HGF gene expression, influencing cell migration. In-vivo studies show that peak podocyte apoptosis, aligned with TGF-β1 expression and initial albuminuria, occurs before mesangial expansion in glomerulosclerosis models, highlighting TGF-β1’s role in fibrosis and inflammation31,32. The balance between HGF and TGF-β is critical for LN prognosis, emphasizing TGF-β’s dual role in renal disease progression and nephrotoxicity33,34.
Notably, the variant rs80282109, located in the intronic region of the BACH2 gene, illustrates the intricate link between genetic variations and immune function regulation. BACH2 is critical in the development and function of B and T lymphocytes, playing a pivotal role in maintaining immune homeostasis. Within B cells, BACH2 is essential for balancing differentiation into antibody-producing plasma cells and preserving the capacity of B cells to mature and react to antigens35. For T cells, BACH2 controls the differentiation of CD4 + T cells into various subtypes, such as regulatory T cells (Tregs) and effector T cells36. Elevated levels of BACH2 suppress “myeloid genes” in pre- and pro-B cells, steering them away from the lymphoid lineage37. A study found that LN patients who experienced more frequent relapses exhibited higher microRNA-148a (miR-148a) expression in naive and memory B cells, alongside lower BACH2 expression in B lymphocytes38. MiR-148a is prevalent in B cells and plasma cells and regulates critical B cell transcription factors such as BACH2, enhancing plasma cell differentiation and maintaining B cell tolerance39,40. The regulation of miR-148a over BACH2 suppresses the maturation and homeostasis of B lymphocytes41. Thus, the presence of variant rs80282109 may influence the splicing and subsequent expression of BACH2, potentially altering its function and impacting the immune regulatory pathways crucial in LN pathogenesis.
Conversely, rs516119, positioned upstream of the SOX1 gene, demonstrated negative effects on LN, suggesting a reduced likelihood of disease development associated with this genetic variant. The SOX gene family predominantly regulates sex determination and development, with certain subgroups playing critical roles in neuronal development. In the SOX gene family, the SOX1 gene belongs to the SOXB1 subclass, which are expressed in neuronal tissues and are believed to play a role in regulating cell differentiation42. More reports indicate that SOX4, a member of a different subgroup within the SOX gene family, is associated with the pathogenesis of LN, as it is essential for the survival of pro-B cells in mice43 and plays a role in TGF-β signaling, suppressing the Th2 immune response by regulating GATA344.
Intronic variant rs134545 located in the middle region of the TTC28 gene, could potentially influence gene splicing or regulatory elements, thereby affecting gene expression. Given the involvement of TTC28 in the mitotic cycle, this variant might have a broad impact on cell division and proliferation within immune cells. However, the link between TTC28 and LN remains underexplored in existing research. Previous studies indicate that TTC28 regulates the mitotic cell cycle and localizes in the midbody45. In the vascular endothelial cells of the blood–brain barrier (BBB), TTC28 interacts with ICAM1, a membrane receptor involved in multiple sclerosis (MS) pathogenesis. This interaction facilitates the transcellular migration of inflammatory T-cells into the CNS46,47, a key mechanism in MS development.
However, our study has limitations, including reliance on specific genotyping technologies like the TPM array chip and bioinformatics tools, affecting sensitivity and specificity. Additionally, the cross-sectional design limits understanding of temporal gene expression changes and disease progression, necessitating longitudinal studies for comprehensive assessment.
Conclusion
This genetic screening study among Taiwanese LN patients identified four significant variants: rs1025129 (upstream of HGF), rs80282109 (intronic region in BACH2), rs516119 (upstream of SOX1), and rs134545 (intronic region in TTC28). Variant rs1025129 may influence HGF expression, crucial for tissue regeneration and inflammation mitigation. Variant rs80282109 might alter the role of BACH2 in immune regulation. Variant rs516119 shows a potential protective effect against LN. Variant rs134545 in TTC28 could impact cell division and proliferation. These findings highlight the need for functional assays to explore their effects on gene expression and protein function, aiding in understanding LN pathogenesis. This genetic survey marks a significant step toward personalized medical approaches for diagnosing and managing LN, leading to targeted therapies and enhanced predictive models of disease susceptibility.
Limitations
The present study was limited by a small sample size, which precluded adequate assessment of statistical power in the GWAS analysis. Therefore, this investigation should be considered a preliminary exploration of genetic variants associated with LN and SLE. Future studies will focus on expanding participant recruitment to achieve a sufficient sample size, thereby ensuring robust statistical power and enabling more definitive conclusions.
Data availability
The data supporting the findings of this study are not publicly available due to project and IRB policies, as well as the sensitive nature of the participant information. However, some data that do not contain any identifiable personal records may be made available upon reasonable request, in accordance with the consent agreements, by contacting the author. If necessary, please contacted Feng-Cheng Liu to request the data from this study.
References
Hanly, J. G. et al. The frequency and outcome of lupus nephritis: results from an international inception cohort study. Rheumatology 55(2), 252–262 (2016).
Cervera, R. et al. Morbidity and mortality in systemic lupus erythematosus during a 10-year period: a comparison of early and late manifestations in a cohort of 1000 patients. Medicine 82(5), 299–308 (2003).
Singh, S., Saxena, R. & Palmer, B. F. Lupus nephritis. Am. J. Med. Sci. 337(6), 451–460 (2009).
Lyn-Cook, B. D. et al. Increased expression of Toll-like receptors (TLRs) 7 and 9 and other cytokines in systemic lupus erythematosus (SLE) patients: ethnic differences and potential new targets for therapeutic drugs. Mol. Immunol. 61(1), 38–43 (2014).
Devarapu, S. K. & Anders, H.-J. Toll-like receptors in lupus nephritis. J. Biomed. Sci. 25, 1–11 (2018).
Wang, J. et al. Association of abnormal elevations in IFIT 3 with overactive cyclic GMP-AMP synthase/stimulator of interferon genes signaling in human systemic lupus erythematosus monocytes. Arthritis Rheumatol. 70(12), 2036–2045 (2018).
Kato, Y. et al. Apoptosis-derived membrane vesicles drive the cGAS–STING pathway and enhance type I IFN production in systemic lupus erythematosus. Ann. Rheum. Dis. 77(10), 1507–1515 (2018).
Hom, G. et al. Association of systemic lupus erythematosus with C8orf13–BLK and ITGAM–ITGAX. N. Engl. J. Med. 358(9), 900–909 (2008).
Souyris, M. et al. TLR7 escapes X chromosome inactivation in immune cells. Sci. Immunol. 3(19), eaap8855 (2018).
Song, D. et al. Complement alternative pathway’s activation in patients with lupus nephritis. Am. J. Med. Sci. 353(3), 247–257 (2017).
Pan, C. et al. Molecular analysis of HLA-DRB1 allelic associations with systemic lupus erythematous and lupus nephritis in Taiwan. Lupus 18(8), 698–704 (2009).
Chen, J.-Y. et al. Interferon-λ3/4 genetic variants and interferon-λ3 serum levels are biomarkers of lupus nephritis and disease activity in Taiwanese. Arthritis Res. Ther. 20, 1–11 (2018).
Wei, C.-Y., Yang, J.-H., Yeh, E.-C. Genetic profiles of 103,106 individuals in the Taiwan Biobank provide insights into the. NPJ Genom. Med. 6(1) (2021).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81(3), 559–575 (2007).
Yang, H. & Wang, K. Genomic variant annotation and prioritization with ANNOVAR and wANNOVAR. Nat. Protoc. 10(10), 1556–1566 (2015).
Consortium GP. A global reference for human genetic variation. Nature 526(7571), 68 (2015).
Gudmundsson, S. et al. Variant interpretation using population databases: Lessons from gnomAD. Hum. Mutat. 43(8), 1012–1030 (2022).
Feng, Y.-C.A., Chen, C.-Y., Chen, T.-T., Kuo, P.-H., Hsu, Y.-H., Yang, H.-I., et al. Taiwan Biobank: A rich biomedical research database of the Taiwanese population. Cell Genom. 2(11) (2022).
Tangtanatakul, P. et al. Meta-analysis of genome-wide association study identifies FBN2 as a novel locus associated with systemic lupus erythematosus in Thai population. Arthritis Res. Ther. 22, 1–13 (2020).
Langefeld, C. D. et al. Transancestral mapping and genetic load in systemic lupus erythematosus. Nat. Commun. 8(1), 16021 (2017).
Song, Q. et al. Genome-wide association study on Northern Chinese identifies KLF2, DOT1L and STAB2 associated with systemic lupus erythematosus. Rheumatology 60(9), 4407–4417 (2021).
Kuo, C.-F. et al. Familial aggregation of systemic lupus erythematosus and coaggregation of autoimmune diseases in affected families. JAMA Intern. Med. 175(9), 1518–1526 (2015).
Mohan, C. & Putterman, C. Genetics and pathogenesis of systemic lupus erythematosus and lupus nephritis. Nat. Rev. Nephrol. 11(6), 329–341 (2015).
Yin, X. et al. Meta-analysis of 208370 East Asians identifies 113 susceptibility loci for systemic lupus erythematosus. Ann. Rheum. Dis. 80(5), 632–640 (2021).
Chen, Y.-C. et al. Multiple polygenic risk scores can improve the prediction of systemic lupus erythematosus in Taiwan. Lupus Sci. Med. 11(1), e001035.2 (2024).
Wang, C.-M. et al. Genetic variations in Toll-like receptors (TLRs 3/7/8) are associated with systemic lupus erythematosus in a Taiwanese population. Sci. Rep. 4(1), 3792 (2014).
Liu, Y. & Yang, J. Hepatocyte growth factor: new arsenal in the fights against renal fibrosis?. Kidney Int. 70(2), 238–240 (2006).
Herrero-Fresneda, I. et al. HGF gene therapy attenuates renal allograft scarring by preventing the profibrotic inflammatory-induced mechanisms. Kidney Int. 70(2), 265–274 (2006).
Stracke, S. et al. Differentiating and proliferative effects of HGF in renal proximal tubular cells are mediated via different signalling pathways. Nephrol. Dial. Transplant. 13(6), 1398–1405 (1998).
Rampino, T. et al. Hepatocyte growth factor and its receptor Met are induced in crescentic glomerulonephritis. Nephrol. Dial. Transplant. 20(6), 1066–1074 (2005).
Schiffer, M. et al. Apoptosis in podocytes induced by TGF-β and Smad7. J. Clin. Investig. 108(6), 807–816 (2001).
Diwakar, R., Pearson, A. L., Colville-Nash, P., Brunskill, N. J. & Dockrell, M. E. The role played by endocytosis in albumin-induced secretion of TGF-β1 by proximal tubular epithelial cells. Am. J. Physiol.-Ren. Physiol. 292(5), F1464–F1470 (2007).
Ohtsuka, K., Gray, J. D., Stimmler, M. M., Toro, B. & Horwitz, D. A. Decreased production of TGF-β by lymphocytes from patients with systemic lupus erythematosus. J. Immunol. 160(5), 2539–2545 (1998).
Yamamoto, K. & Loskutoff, D. J. Expression of transforming growth factor-β and tumor necrosis factor-α in the plasma and tissues of mice with lupus nephritis. Lab. Investig. 80(10), 1561–1570 (2000).
Igarashi, K., Ochiai, K., Itoh-Nakadai, A. & Muto, A. Orchestration of plasma cell differentiation by Bach2 and its gene regulatory network. Immunol. Rev. 261(1), 116–125 (2014).
Yang, L., Chen, S., Zhao, Q., Sun, Y., Nie, H. The critical role of Bach2 in shaping the balance between CD4+ T cell subsets in immune-mediated diseases. Mediat. Inflamm. 2019 (2019).
Itoh-Nakadai, A. et al. The transcription repressors Bach2 and Bach1 promote B cell development by repressing the myeloid program. Nat. Immunol. 15(12), 1171–1180 (2014).
Yap, D. Y. et al. B cell subsets and cellular signatures and disease relapse in lupus nephritis. Front. Immunol. 11, 1732 (2020).
Porstner, M. et al. miR-148a promotes plasma cell differentiation and targets the germinal center transcription factors Mitf and Bach2. Eur. J. Immunol. 45(4), 1206–1215 (2015).
Gonzalez-Martin, A. et al. The microRNA miR-148a functions as a critical regulator of B cell tolerance and autoimmunity. Nat. Immunol. 17(4), 433–440 (2016).
Muto, A. et al. Bach2 represses plasma cell gene regulatory network in B cells to promote antibody class switch. EMBO J. 29(23), 4048–4061 (2010).
Miyagi, S., Kato, H. & Okuda, A. Role of SoxB1 transcription factors in development. Cell. Mol. Life Sci. 66, 3675–3684 (2009).
Sun, B. et al. Sox4 is required for the survival of pro-B cells. J. Immunol. 190(5), 2080–2089 (2013).
Kuwahara, M. et al. The transcription factor Sox4 is a downstream target of signaling by the cytokine TGF-β and suppresses TH2 differentiation. Nat. Immunol. 13(8), 778–786 (2012).
Buzgariu, W., Wenger, Y., Tcaciuc, N., Catunda-Lemos, A.-P. & Galliot, B. Impact of cycling cells and cell cycle regulation on Hydra regeneration. Dev. Biol. 433(2), 240–253 (2018).
Huttlin, E. L. et al. Architecture of the human interactome defines protein communities and disease networks. Nature 545(7655), 505–509 (2017).
Abadier, M. et al. Cell surface levels of endothelial ICAM-1 influence the transcellular or paracellular T-cell diapedesis across the blood–brain barrier. Eur. J. Immunol. 45(4), 1043–1058 (2015).
Acknowledgements
We extend our gratitude to all the participants and investigators involved in the TPMI. This study was supported by Academia Sinica 40-05-GMM and AS-GC-110-MD02. We are also appreciative of the Center for Precision Medicine and Genomic (CPMG) at Tri-Service General Hospital, National Defense Medical University, for their invaluable assistance in gene information support (Founding number: TSGH-B-112020, TSGH-B-113022).
Funding
This work was partially supported by grants (Tri-Service General Hospital, National Defense Medical University; Funding number: TSGH-B-112020, TSGH-B-113022 and NSTC: (1) 114-2314-B-281-009-MY3. (2) 114-2314-B-281-001. (3) 113-2314-B-016-048. (4) 114-2314-B-016-002).
Author information
Authors and Affiliations
Contributions
J.C.H.L. and F.C.L. conducted the analysis and wrote the manuscript. K.S.H. and J.H.L. performed the computational work for the genotype–phenotype association analysis. J.C.H.L. and K.S.H. interpreted the results and wrote the manuscript portions concerning the genotype–phenotype association analysis. H.C.C., S.Y.K., S.J.C., C.C.L., Y.C.L., F.C.Y. and F.C.L. provided original data. Y.J.H. and F.C.L. organized and supervised this project. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Approval statement
This study is approved by Tri-Service General Hospital Institutional Review Board under approval number 2-108-05-038.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Lee, J.CH., Hung, KS., Lyu, JH. et al. Identification of genetic variants associated with lupus nephritis in a Taiwanese cohort through systematic genetic screening. Sci Rep 15, 35534 (2025). https://doi.org/10.1038/s41598-025-19569-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-19569-4