Abstract
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is relentlessly evolving, with emerging variants exhibiting heightened transmissibility and immune escape capabilities. Understanding the genetic diversity and seasonal transmission dynamics of SARS-CoV-2 in Thailand is crucial for implementing effective public health interventions. This study aims to elucidate the genetic diversity and seasonal transmission patterns of SARS-CoV-2 among patients with acute respiratory illness in Thailand in 2024. Among 8,096 Acute Respiratory Infection (ARI) cases analyzed, 1,152 samples tested positive for SARS-CoV-2, resulting in a positivity rate of 14.2%. The majority of infections occurred during a significant Outbreak from late summer to early rainy season, particularly between April and June, accounting for nearly 49% of all positive cases. The highest infection rate occurred in adults aged 31–40 years, with no significant association between gender and infection status (p = 0.583). Extensive genomic sequencing identified over seven distinct SARS-CoV-2 lineages, with the JN.1 lineage dominating early in the year. Recombinant variants, notably XEC, XDV.1, and XDY, emerged as significant contributors to the evolving landscape, reaching a prevalence of 57.1% by December. Phylogenetic analyses demonstrated a consistent evolutionary rate and identified critical emergence dates for new lineages, underscoring the virus’s ongoing evolution in Thailand. Our findings emphasize the necessity of continuous genomic surveillance for tracking variant transmission dynamics and managing public health responses effectively. Furthermore, the evolving genetic landscape of SARS-CoV-2 highlights the urgent need for adaptive vaccination strategies aligned with seasonal transmission patterns in Thailand.
Similar content being viewed by others
Introduction
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) is a single-stranded, positive-sense RNA virus classified within the Betacoronavirus genus of the Coronaviridae family1. As the causative agent of Coronavirus Disease 2019 (COVID-19), this virus is notorious for its high transmissibility and profound impact on global public health2. The SARS-CoV-2 genome is approximately 29,870 nucleotides long and features a single-stranded RNA structure that includes at least 13 identified open reading frames (ORFs). These ORFs are primarily arranged in a linear sequence from the 5′ to the 3′ end and are crucial for the virus’s biological functions3. Key structural proteins encoded by the SARS-CoV-2 genome include the spike (S), envelope (E), membrane (M), and nucleocapsid (N) proteins, all of which are essential for facilitating viral entry into host cells and for the assembly of new virions. Additionally, SARS-CoV-2 harbors multiple accessory and nonstructural proteins that help the virus evade the immune system, disrupt host cellular processes, and enhance both viral replication and transcription3,4,5,6.
As SARS-CoV-2 continues to evolve, new variants have emerged featuring mutations in the spike protein. Notably, sublineages of the omicron variant have become the predominant strains of the virus. These sublineages are characterized by a wide range of spike protein mutations that significantly increase their transmissibility and ability to evade immune responses7. The first omicron subvariant, BA.1, was detected in mid-November 2021 in South Africa and has been associated with enhanced viral replication and an elevated risk of reinfection8,9,10. After the emergence of BA.1, subsequent mutations led to the development of several significant subvariants, including BA.2, BA.2.75, and BA.4/5. By 2023, these subvariants, along with their descendants, were co-circulating globally, which has raised ongoing concerns among public health officials regarding the potential for increased transmission and impacts on vaccine efficacy11,12,13.
The increasing genetic diversity of SARS-CoV-2 has resulted in the concurrent emergence of multiple lineages, facilitating the identification of various recombinant forms. The Pango-designation project, established and released in early 2020, has meticulously cataloged over 200 distinct recombinant lineages of SARS-CoV-214,15. By late 2022, the Omicron XBB lineages, derived from the recombination of two co-circulating BA.2 sub-lineages (BJ.1 and BM.1.1.1), had become the dominant circulating variants16. However, by late 2023, the emergence of the JN.1 lineage in the United States rapidly supplanted the XBB lineages, establishing itself as the most prevalent Omicron variant globally by 202417. This transition highlights JN.1’s enhanced transmissibility and adaptability18,19. Ongoing genetic evolution has resulted in several JN.1 descendant, including JN.1.16, KP.2, and KP.3.1.117. Notably, in November 2024, the prevalence of a recombinant variant, XEC, a fusion of JN.1 lineages KS.1.1 and KP.3.3, significantly increased, further illustrating the virus’s ongoing genetic diversification17.
Since its emergence in late 2019, SARS-CoV-2 has been responsible for over 7 million confirmed deaths and nearly 700 million confirmed infections globally over a span of four years20. In Thailand, six distinct lineages of SARS-CoV-2 were identified in the first quarter of 2020, namely A, A.6, B, B.1, B.1.8, and B.5821. The B.1.36.16 variant was the primary driver of the major Outbreak during the second wave from December 2020 to January 2021. Following this, the Alpha variant emerged as the leading strain in the third wave from April to June 2021. The Delta variant characterized the fourth wave, which lasted from July to December 2021, while the Omicron variant significantly impacted the fifth wave from January to March 202222. In early 2023, the BA.2.75 variant was the most prevalent among COVID-19 cases; however, it was soon supplanted by the rising XBB.1.15 and XBB.1.16 variants, which gained dominance later in the year23.
Thailand’s commitment to genomic surveillance is exemplified by the submission of 25,178 SARS-CoV-2 sequences to GISAID in 202224. However, this number experienced a substantial decline of 78%, dropping to 5,550 sequences in 2023, and further decreased by 61% to just 2,131 sequences in 202424. Concurrently, scientists are examining the possibility of COVID-19 evolving into the same period as influenza or other respiratory diseases, spreading in the winter season of countries in the Northern and Southern Hemispheres25. In tropical countries such as Thailand, outbreaks can occur almost year-round, with peak transmission during the rainy season, from June to September each year26. As population immunity increases through vaccination and natural infection, and as the virus may mutate into less severe forms, the pattern of outbreaks could become more predictable.
Despite the decreased submission rates, maintaining robust genomic monitoring remains essential for effectively tracking the virus’s evolution and its variants. Continuous surveillance and timely updates on vaccinations are vital for controlling and managing the virus’s impact as it may transition to a seasonal pattern. This study aims to examine the seasonal patterns and genetic diversity of SARS-CoV-2 among patients with acute respiratory illness in Thailand during the fifth year of the COVID-19 pandemic, offering crucial insights into the virus’s evolving dynamics and informing future public health strategies.
Results
Trends in COVID-19 incidence and demographics of infected patients in 2024
According to the Ministry of Public Health (MoPH), a total of 46,079 COVID-19 cases and 220 fatalities were reported between January and December 2024, reflecting a 6% reduction in cases and a 20% decrease in deaths compared to the previous year (Fig. 1a and b)27. Regional distribution of cases revealed that the Bangkok Metropolitan Region (BMR) accounted for 30.8% of the total cases, followed by the Northeastern region at 20.0% and the Southern region at 13.8%. The Western region recorded the lowest proportion, contributing only 3.4% to the overall total. The disproportionately high number of cases in the BMR may reflect its dense population, greater mobility, and broader access to testing services, which could have contributed to both increased transmission and case detection. Notably, the highest incidence of deaths occurred in May and June, accounting for 41% of total fatalities (Fig. 1b).
In this study, 8,096 Acute Respiratory Infection (ARI) cases submitted by Thai residents to partnering hospitals were analyzed. Of these, 1,152 samples tested positive for SARS-CoV-2 via real-time RT-PCR, resulting in a positivity rate of 14.2% (Fig. 1c). Data collection was conducted across Thailand’s seasonal divisions: summer (February to mid-May), rainy season (mid-May to October), and winter (November to February). The findings revealed an initial SARS-CoV-2 Outbreak wave in January, accounting for 13.3% of total positive samples (153/1,152), followed by a significant second wave from April to June, which represented 48.9% of total positives (563/1,152) during the summer months. Notably, positivity rates for SARS-CoV-2 declined below 10% from July to December, indicating a reduction in transmission during the latter part of the year.
Among the 1,152 SARS-CoV-2 infected patients, 539 (46.8%) were Male and 613 (53.2%) were female (Table 1). A Chi-square test showed no significant association between gender and infection status, χ²(1, N = 8,096) = 0.30, p = 0.583. The highest proportion of SARS-CoV-2 infections occurred in individuals aged 31–40 years (18.3%), followed by those aged 41–50 (17.7%) and 1 month–10 years (16.9%). The lowest was in the 11–20 years group (5.9%). Pairwise proportion tests with Bonferroni correction showed significantly higher infection rates in the 31–40 age group compared to the 0–10 (p < 0.001), 11–20 (p = 0.0011), and 51–60 (p < 0.001) groups. No significant differences were observed when compared to the 21–30, 41–50, or > 60 age groups (p > 0.05). These findings highlight a greater burden of infection among middle-aged adults.
SARS-CoV-2 variants distribution in Thailand
To investigate the distribution of SARS-CoV-2 lineages in Thailand from January to December 2024, a comprehensive analysis was conducted on 689 SARS-CoV-2 positive samples. Among these, 60 samples were randomly selected for complete genome sequencing, while the remaining 629 underwent partial spike sequencing. The study identified more than seven distinct lineages, each accompanied by its descendant variants [Fig. 2]. Notable lineages included JN.1, JN.1.16, KP.1 through KP.3, and LB.1, in addition to several recombinant lineages including XCU, XDQ, XDV, XDY, XEC, XEF, XEL, and XEN. The findings revealed that the frequency and distribution of these viral lineages demonstrated continuous variations over time.
(a) The distribution of SARS-CoV-2 lineages in Thailand from January to December 2024 includes each lineage along with its descendant lineages. (b) The variation frequencies over time are shown, where the size of each point indicates the weekly collection of sequences. These data points are assessed through logistic regression, with the shaded areas denoting the 95% confidence interval. Only the variants with the highest prevalence during the study period are displayed, emphasizing the most significant trends in variant distribution and their statistical reliability.
In alignment with the observed variant frequencies (Fig. 2a and b), the JN.1 lineage Maintained the highest prevalence from January to April, peaking at 90.2% (55/61) in February, followed by JN.1.16 at 11.1% (4/36) in March and KP.2 at 23.2% (16/69) in April. Following this period, the highest prevalence transitioned to KP.2 during May and June, reaching a peak of 28.6% (32/112) in June, while JN.1.16 emerged as the second most prevalent variant, accounting for 23.4% (32/137) and 25.9% (29/112) in those respective months. Additionally, LB.1 was first detected in April with a prevalence of 4.3% (3/69) and subsequently peaked at 35.0% (21/60) in July, establishing itself as a dominant variant before declining and ultimately disappearing by November.
From August to September, the predominant variant shifted once again to KP.3, which accounted for 29.0% of cases in both months (11/38 in August and 9/31 in September), primarily represented by sublineages KP.3.1.1, KP.3.2, and KP.3.3.1 (Supplement Fig. 1). This variant maintained its status as the second most prevalent variant through the end of December.
The recombinant lineages were first identified in May, beginning with an initial prevalence of 8.8% (12/137). They subsequently became predominant during the last three months of the year, reaching 38.2% (13/34) in October and peaking at 57.1% (4/7) in December. For example, XDV.1 began to significantly contribute in mid-year, first detected in May at 5.1% (7/137) and increasing to 10.5% (4/38) in August (Supplement Fig. 1). Meanwhile, XDY peaked in September at 9.7% (3/31). The variant XEC emerged as the most prevalent within the recombinant lineages, first identified in September at 3.2% (1/31), and then rising to 15.8% (3/19) in November and 14.3% (1/7) in December. Additionally, in October, other recombinant variants detected included XEF at 2.9% (1/34), and both XEL and XEN at 8.8% each (3/34). This dynamic landscape of variant prevalence highlights the ongoing evolution and transmission patterns of SARS-CoV-2 in Thailand, underscoring the necessity for continuous genomic surveillance to adapt public health responses effectively.
Maximum likelihood and time‑scaled phylogenetic reconstruction of sars‑cov‑2 variants
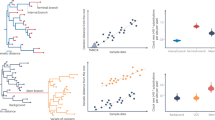
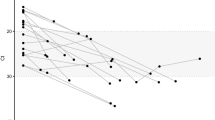
To establish initial trees for Bayesian phylodynamic analyses, time-scaled phylogenies were constructed based on the maximum likelihood (ML) tree topologies and their corresponding collection dates (Fig. 3a and b). The ML phylogenetic tree, derived from complete genome sequences, illustrates the molecular evolution of SARS-CoV-2 isolates both from Thailand and globally. Molecular clock analysis of the dataset provided robust evidence for the time-stable, clock-like evolution of these lineages, demonstrating an R² value of 0.5 and a correlation coefficient of 0.7, along with a substitution rate of 1.38 × 10–3 substitutions per site per year (Fig. 3c). This analysis employed the optimal rooting approach, which minimizes the mean squared residuals, to effectively investigate the relationships between genetic divergence and sampling dates.
(a) A global maximum likelihood (ML) tree was constructed from a dataset of complete genome sequences (N = 423) using TreeTime, employing the oldest method for phylogenetic analysis. (b) A time-scaled ML tree was generated by integrating collection dates into the ML tree from panel (a), thus serving as the primary reference for further analysis. (c) A regression analysis of root-to-tip genetic distances against sampling dates for this comprehensive dataset, estimated using TempEst, revealed a significant positive molecular clock signal.
The Bayesian phylogenetic tree, constructed from an Additional set of 60 complete SARS-CoV-2 genomes (Supplement Table 1) alongside global sequences collected between December 2023 and December 2024 (Fig. 4), indicates that the most recent common ancestor of this dataset is estimated to have emerged in August 2022. Overall, the Thai and global SARS-CoV-2 samples did Not exhibit distinct phylogenetic clustering, suggesting similar patterns of viral circulation and shared lineages during the study period. However, after September 2024, a divergence was observed, with Thai samples not clustering within the sublineage clusters KP3.1.1, KP3.2, and KP3.3.1. The absence of these newly emerging KP3 sublineages in Thailand during the study period May reflect differences in transmission dynamics, introduction events, or the temporal scope of sampling. The molecular evolutionary rate for the dataset was estimated at 0.63 × 10–3 nucleotide substitutions per site per year (sub/site/year), with a 95% highest posterior density interval (HPDI) ranging from 0.57 × 10–3 to 0.70 × 10–3 sub/site/year.
Maximum Clade Credibility (MCC) tree of SARS-CoV-2 variant populations circulating in Thailand (n = 176) and globally (n = 247), inferred using molecular clock analysis of samples collected between December 2023 and December 2024. The scale bar indicates nucleotide substitutions per site.
Table 2 details the sequence change rates and the time to the most recent common ancestors (TMRCA) as determined by Markov chain Monte Carlo (MCMC) methods. Among all variants, XDV showed the highest substitution rate (0.89 × 10–3) and the greatest nucleotide divergence (4.56 × 10–4), with a tMRCA estimated around late November 2023. In contrast, JN.1 had the lowest substitution rate (0.45 × 10–3) and an earlier tMRCA in early March 2023. Variants KP.1 to KP.3 and LB.1 shared comparable nucleotide divergence values (3.66 × 10–4 and 4.09 × 10–4, respectively), with substitution rates of 0.56 × 10–3 and 0.75 × 10–3. However, KP.1–3 emerged later, with a tMRCA in mid-November 2023, while LB.1 diverged slightly after, in early January 2024. XEC and XDY both emerged more recently, with tMRCAs in May 2024. While their substitution rates were similar (0.55 and 0.56 × 10–3), XDY showed notably lower nucleotide divergence (1.72 × 10–4) compared to XEC (3.33 × 10–4), possibly reflecting differences in evolutionary dynamics or sampling time frames.
Discussion
The COVID-19 Outbreak, first identified in late 2019, arrived in Thailand in early 2020, leading to a series of ongoing transmission events that prompted national concern28. Following nearly three years of extensive outbreaks and significant public health challenges, the Thai Ministry of Public Health reclassified COVID-19 from a ‘dangerous infectious disease’ to a ‘disease under surveillance’ and announced a revised reporting protocol in October 2022 that focuses exclusively on case counts among hospitalized patients29. By 2023, the severity of COVID-19 had significantly diminished, evidenced by a marked reduction in fatalities27. In 2024, the reported death toll decreased to just 220 cases, representing a stark contrast to the high mortality rates observed in the first three years of the pandemic, which included 21,614 deaths in 2021, 11,971 in 2022, and 848 in 202327. This reduction in severity can be attributed to several factors, including increased vaccination coverage, the emergence of less virulent variants, and improved treatment modalities. As a result, COVID-19 is now recognized as a seasonal respiratory illness, similar to other common respiratory infections.
This study investigated the seasonal trends and genetic diversity of SARS-CoV-2 among patients with acute respiratory illness in Thailand during the fifth year of the COVID-19 pandemic, providing a comprehensive analysis of acute respiratory infection cases throughout 2024, which includes both inpatient and outpatient data across all seasons. The results demonstrate that COVID-19 transmission follows a distinct seasonal pattern, marked by a significant increase in cases from late summer to the early rainy season, particularly between April and June.
In the early years of COVID-19’s seasonal transition, its incidence peaked earlier than other respiratory infections, such as influenza and respiratory syncytial virus30,31. Although Thailand’s official rainy season begins in mid-May, the interplay of school reopenings and the Songkran festival in mid-April, characterized by extensive travel and gatherings, contributes significantly to an early rise in infection rates.
Following this peak, case numbers gradually declined from September to the end of the year, before experiencing another surge in early winter (January to mid-March), primarily due to lower temperatures. Unlike temperate regions where COVID-19 surges predominantly occur during winter32,33,34. Thailand’s tropical climate lacks a distinct cold season. While there is evidence of increased cases during cooler months, the most pronounced peak reliably occurs at the onset of the rainy season each year. This highlights the unique seasonal dynamics of COVID-19 in Thailand, characterized by year-round transmission with a marked surge during early monsoon months. Such insights are crucial for informing public health strategies tailored to local seasonal patterns.
This study observed the highest prevalence of COVID-19 infection among individuals aged 31 to 40 years, with statistically significant differences compared to several other age groups, including children (0–10 years), adolescents (11–20 years), and older adults aged 51–60 years. These findings are consistent with previous research reporting a greater burden of SARS-CoV-2 infection among middle-aged adults, possibly due to increased social mobility and occupational exposure during the post-pandemic reopening phase35. Although a slightly higher proportion of female cases was observed, statistical analysis showed no significant association between gender and infection status. This aligns with earlier studies that reported comparable infection rates between males and females35. However, consistent with earlier findings, male patients have been reported to experience more severe disease outcomes and increased mortality36.
The current study revealed that the predominance of variants shifted throughout the year, with JN.1 dominating in the earlier part of the year, followed by transitions to KP.2 and LB.1. After August, KP.3 became the predominant variant, primarily represented by the sublineages KP.3.1.1, and KP.3.3.1. The present study aligns with a report from South Korea, which found that the proportions of various sub-lineages, including JN.1, KP.2, LB.1, and KP.3, exhibited the highest prevalence during the period from April to August 202437. That report also indicated that the KP.3 sub-lineages with the highest proportions were identified as KP.3.3.1, KP.3.3, and KP.3.1.1. The ongoing genetic evolution of SARS-CoV-2 poses significant challenges for vaccine development, as newly emerging variants may diminish the efficacy of existing vaccines. The variants identified in Thailand closely align with those circulating globally, highlighting the interconnected nature of SARS-CoV-2 evolution and underscoring the need for continuous genomic surveillance and vaccine adaptation to effectively combat the evolving pandemic. Notably, while the overall variant dynamics in Thailand closely mirrored global trends, the absence of the newly emerging KP.3 sublineages (KP.3.1.1, KP.3.2, and KP.3.3.1) in Thai samples after September 2024 suggests potential regional differences in viral evolution and spread. This highlights the importance of timely genomic surveillance to detect emerging sublineages that may initially appear in global datasets but remain undetected locally.
The current study estimated a nucleotide evolutionary rate of approximately 0.63 × 10–3 substitutions per site per year (sub/site/year), consistent with reported during the XBB wave in the Republic of Korea (late 2022 to late 2023), where substitution rates ranged from 0.56 × 10–3 to 0.91 × 10–3 sub/site/year38. Similarly, a study conducted in the United States reported a whole-genome evolutionary rate of 0.67 × 10–3 (sub/site/year), further supporting the comparability of our findings39. When compared to other respiratory viruses, this rate is similar to that of human rhinovirus (0.66 × 10–3) and human metapneumovirus (0.71 × 10–3), while it is lower than that of respiratory syncytial virus (0.76 × 10–3) and influenza A, which ranges from 2.21 to 3.37 × 10–340,41,42,43.
The present study provides important insights into the molecular epidemiology and evolutionary dynamics of SARS-CoV-2 circulating in Thailand and globally. By integrating genomic surveillance with phylogenetic analysis, the findings contribute to the early detection of emerging variants and the identification of transmission patterns. The ability to monitor viral evolution in near real-time supports early warning systems and enables the timely adjustment of public health measures. Moreover, the phylogenetic similarities observed between Thai and global samples emphasize the importance of international surveillance and data sharing to anticipate local outbreaks influenced by global viral movements. These insights are instrumental for guiding vaccine policy, resource allocation, and national preparedness planning, particularly in the context of rapidly evolving variants such as JN.1 and KP.1-KP.3.
During the study period, vaccination strategies globally were updated to address the Omicron JN.1 subvariant, with reformulated vaccines being deployed. In Thailand, however, these JN.1-targeted vaccines were introduced later, toward the end of 2024, and their uptake was limited due to high cost and voluntary administration. This may have influenced the transmission dynamics observed in the region.
This study provides a comprehensive year-long analysis of the seasonal distribution of COVID-19 in Thailand, utilizing a substantial dataset collected over the entire year with a large number. However, several limitations must be acknowledged. Since the analysis focuses exclusively on a single year, multi-year investigations are necessary to fully understand the seasonal fluctuations of COVID-19 outbreaks. Regarding variant analysis, despite efforts to conduct sequencing throughout the year, resource constraints, including budget limitations, limited the ability to sequence all cases. The limited number and temporal distribution of fully sequenced genomes may affect the representativeness of lineage dynamics, so caution is warranted when interpreting results based on this subset. Additionally, in instances where patients exhibited low viral loads (as indicated by high Ct values), variant identification was not feasible. For evolutionary analysis, the inclusion of more robust metadata would enhance the accuracy and effectiveness of datasets used to estimate the nucleotide substitution rate and the time to the most recent common ancestor (tMRCA).
In conclusion, this study provides a comprehensive analysis of COVID-19 incidence and variant dynamics in Thailand throughout 2024. It highlights a notable seasonal trend in infection rates and emphasizes the predominance of specific SARS-CoV-2 lineages, including KP.3 and its sublineages, particularly in the latter part of the year. The findings underscore the importance of continuous genomic surveillance to monitor the evolving landscape of variants and inform public health strategies effectively. Additionally, the estimated molecular evolutionary rate and the emergence timing of key lineages contribute valuable insights to the understanding of SARS-CoV-2 evolution and its impact on the pandemic landscape in Thailand.
Materials and methods
Institutional review board statement
The research was carried out following the Good Clinical Practice (GCP) under the principles outlined in the Declaration of Helsinki and received approval from the Institutional Review Board of the Faculty of Medicine, Chulalongkorn University, Thailand (approval number IRB0933/67). To ensure patient confidentiality, all data and identifiers were anonymized. Since this was a retrospective study, the Institutional Review Board waived the requirement for informed consent.
Specimens collection
In this study, 8,096 specimens were collected using throat swabs or nasopharyngeal swabs with Flock swabs (Copan Diagnostics, Murrieta, CA) or Modono sterile swabs (Modono, New Delhi, India), and were subsequently placed in viral transport media (VTM) or universal transport media (UTM). The samples were obtained from patients presenting with acute respiratory illness characterized by symptoms such as fever, sore throat, rhinorrhea, cough, or dyspnea within seven days of symptom onset. The study included both outpatients and inpatients seeking medical care at various hospitals in Bangkok as well as at Chum Phae Hospital in Khon Kaen province. Collected samples were tested for multiple respiratory viruses, including influenza viruses (A and B), SARS-CoV-2, respiratory syncytial virus (RSV), adenovirus, parainfluenza viruses, human metapneumovirus, seasonal coronaviruses, and rhinovirus. Residual specimens were then sent to the Center of Excellence in Clinical Virology at the Faculty of Medicine, Chulalongkorn University, where they were stored at − 20 °C until further analysis for this study.
RNA extraction and molecular testing
RNA extraction and molecular testing were conducted as previously outlined23. In brief, RNA was extracted from a 200-µL aliquot of the supernatant using the MagLEAD 12gC instrument (Precision System Science, Chiba, Japan), following the manufacturer’s protocols meticulously. Real-time reverse transcription-PCR (RT-PCR) was performed using the Roche LightCycler® 480 (LC480) instrument, employing specific primers and probes targeting the nucleocapsid gene (N1 and N2) in accordance with the guidelines established by the Centers for Disease Control and Prevention (CDC)44. The amplification of the housekeeping gene Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) was included as an endogenous control for each sample, utilizing primers and probes as outlined in prior studies45. The thermocycling conditions included an initial 10-minute incubation at 45 °C, followed by a 5-minute denaturation step at 95 °C, and then 45 amplification cycles at 95 °C for 15 s and 60 °C for 30 s. Each testing run included positive and negative (nontemplate) controls, with a sample classified as positive for SARS-CoV-2 if the cycle threshold (CT) value was ≤ 38. Additionally, a selection of positive SARS-CoV-2 samples was randomly chosen for variant classification via partial spike sequencing and complete genome sequencing.
Amplification and sequencing of the partial Spike gene
From January to December 2024, a total of 629 SARS-CoV-2 RNA samples were processed, with partial amplification of the spike gene performed on individuals in Thailand confirmed to be infected with COVID-19. The amplification utilized primers detailed in Supplementary Table 2. Briefly, RT-PCR was performed in a total volume of 25 µL, consisting of 2–3 µL of total RNA (ranging from 100 ng to 1 µg), 0.5 µM of each primer, 12.5 µL of 2X Reaction Mix (which contains 0.4 mM of each dNTP and 3.2 mM MgSO4), 1 µL of the SSIII RT/Platinum Taq Mix, and nuclease-free water. The Superscript III One-Step RT-PCR system with Platinum Taq High Fidelity was utilized according to the manufacturer’s guidelines (Invitrogen, Carlsbad, CA, USA). The PCR protocol involved an initial incubation at 45 °C for 30 min, followed by 40 cycles that included denaturation at 95 °C for 30 s, annealing at 50 °C for 30 s, and extension at 68 °C for 1 min and 45 s. A final extension step was carried Out at 68 °C for 5 min. Both forward and reverse primers were utilized concurrently for sequencing and product amplification, conducted at First BASE Laboratories Sdn Bhd (Selangor Darul Ehsan, Malaysia), ensuring comprehensive coverage and accuracy in the analysis.
Whole-genome sequencing
Residual SARS-CoV-2 PCR-positive respiratory specimens (N = 60) were utilized for viral sequencing. Samples with a Ct value of 25 or lower were specifically selected for whole-genome sequencing, which was carried out by adapting a previously established protocol to enhance accuracy and efficiency in the genomic analysis23. The sequencing and identification of complete SARS-CoV-2 genomes were conducted using the Celemics Comprehensive Respiratory Virus Panel (Celemics Inc., Incheon, Republic of Korea), facilitating accurate and efficient genomic analysis. Briefly, RNA extraction involved combining 25 ng of isolated RNA with an RNA fragmentation buffer, followed by first-strand cDNA synthesis using a specialized master mix. The cDNA was converted into double-stranded form through incubation with a second-strand synthesis mix, after which it was cleaned, repaired, and modified with poly(A) tail oligomers. The A-tailed DNA was then ligated to adapters and purified using CeleMag cleanup beads before amplification to create an adapter-ligated library with CLM polymerase and unique dual-index primers. The quality of the library was assessed using automated capillary gel electrophoresis, ensuring DNA fragments were in the 200 to 400 bp range. Next-Generation Sequencing (NGS) was conducted on the Illumina NextSeq 500 system, and the resulting FASTQ data were trimmed, assembled, and analyzed through the Celemics Virus Verifier pipeline to generate consensus sequences.
Maximum likelihood phylogenetic analysis and molecular clock assessment
In this study, a comprehensive dataset of 423 genomic sequences was compiled, which included 60 newly collected sequences from this research and 116 sequences from other provinces in Thailand (Supplement Table 3). This dataset was further enriched by integrating 247 globally representative SARS-CoV-2 genomes retrieved from the GISAID database within the timeframe of the study, from December 2023 to December 2024, thereby enhancing its robustness. The dataset was aligned utilizing MAFFT v.746, and the alignments were subsequently partitioned by codon position. TreeTime47 was used to construct a maximum likelihood tree, which provided the framework for the Bayesian phylodynamic analyses based on the tree topologies and corresponding collection dates. The resulting maximum likelihood phylogeny and time-scaled tree were visualized using the ggtree package48 in R. v.4.4.249. Additionally, a regression analysis of root-to-tip genetic distance against sampling time was conducted using TempEST v1.5.350.
Bayesian phylogenetic analysis and genetic distance estimation
A complete genome dataset was utilized to reconstruct time-scaled phylogenies through Bayesian inference, employing Markov Chain Monte Carlo (MCMC) techniques via the BEAST software (v.2.4.8)51. To determine the most appropriate evolutionary model, combinations of three coalescent tree priors (constant population size, exponential growth, and Bayesian skyline) and two molecular clock models (strict and uncorrelated lognormal relaxed clocks) were independently evaluated. Model fit was assessed through marginal likelihood estimation (MLE) via path sampling and stepping-stone sampling52. Log Bayes Factors (logBF), calculated as the difference in MLE between competing models, were interpreted following Kass and Raftery’s criteria53, with logBF > 5 indicating very strong support. Among all tested models, the strict molecular clock with a constant population size prior exhibited the highest marginal likelihood and was decisively supported over alternative models (Supplement Table 4). Final phylogenetic inference was therefore conducted under the selected best-fit model. Two independent MCMC chains, each consisting of 200 million steps, were run and combined using the BEAGLE library54 to enhance computational performance. Parameters and trees were sampled every 20,000 steps, with the initial 20% discarded as burn-in. Convergence and adequate sampling were confirmed in Tracer v1.7.155, with effective sample sizes (ESS) exceeding 200 for all key parameters. The resulting posterior tree distributions were summarized using TreeAnnotator v1.8.4 to generate a maximum clade credibility (MCC) tree, and phylogenies were visualized with FigTree (https://github.com/rambaut/figtree/releases).
Average genetic distances were calculated using MEGA-X version [10.2.6], employing the Kimura 2-parameter model with variance estimation enabled56.
Statistical analysis
Associations between the categorical variable gender and infection status were examined using Pearson’s chi-square test of independence. Pairwise comparisons of infection proportions among age groups were conducted with Bonferroni correction to adjust for multiple testing. Statistical significance was set at p < 0.05. All analyses were performed using R version 4.4.249.
Data availability
The datasets generated and/or analyzed in this study are available in the National Center for Biotechnology Information (NCBI) repository, with accession numbers PV290368-PV290427 for complete genome data and PV291042-PV291670 for partial spike sequences.
References
Coronaviridae Study Group of the International Committee on Taxonomy of Viruses. The species severe acute respiratory syndrome-related coronavirus: classifying 2019-nCoV and naming it SARS-CoV-2. Nat. Microbiol. 5, 536–544 (2020).
Hu, B., Guo, H., Zhou, P. & Shi, Z. L. Characteristics of SARS-CoV-2 and COVID-19. Nat. Rev. Microbiol. 19, 141–154 (2021).
Malone, B., Urakova, N., Snijder, E. J. & Campbell, E. A. Structures and functions of coronavirus replication-transcription complexes and their relevance for SARS-CoV-2 drug design. Nat. Rev. Mol. Cell. Biol. 23, 21–39 (2022).
Yang, H. & Rao, Z. Structural biology of SARS-CoV-2 and implications for therapeutic development. Nat. Rev. Microbiol. 19, 685–700 (2021).
Rashid, F. et al. Roles and functions of SARS-CoV-2 proteins in host immune evasion. Front. Immunol. 13, 940756. https://doi.org/10.3389/fimmu.2022.940756 (2022).
Mariano, G., Farthing, R. J., Lale-Farjat, S. L. M. & Bergeron, J. R. C. Structural characterization of SARS-CoV-2: where we are, and where we need to be. Front. Mol. Biosci. 7, 605236. https://doi.org/10.3389/fmolb.2020.605236 (2020).
Fan, Y. et al. SARS-CoV-2 Omicron variant: recent progress and future perspectives. Signal. Transduct. Target. Ther. 7. https://doi.org/10.1038/s41392-022-00997-x (2022).
Fischer, C. et al. Emergence and spread of the SARS-CoV-2 Omicron (BA.1) variant across africa: an observational study. Lancet Glob Health. 13, e256–e267 (2025).
Viana, R. et al. Rapid epidemic expansion of the SARS-CoV-2 Omicron variant in Southern Africa. Nature 603, 679–686 (2022).
Karim, S. S. A., Karim, Q. A. & Omicron SARS-CoV-2 variant: a new chapter in the COVID-19 pandemic. Lancet 398, 2126–2128 (2021).
Tan, C. W. et al. Comparative neutralisation profile of SARS-CoV-2 omicron subvariants BA.2.75 and BA.5. Lance. Microbe. 3, e898. https://doi.org/10.1016/S2666-5247(22)00220-8 (2022).
Cao, Y. et al. Omicron escapes the majority of existing SARS-CoV-2 neutralizing antibodies. Nature 602, 657–663 (2022).
Dejnirattisai, W. et al. SARS-CoV-2 omicron-B.1.1.529 leads to widespread escape from neutralizing antibody responses. Cell 185, 467 (2022).
Rambaut, A. et al. Addendum: A dynamic nomenclature proposal for SARS-CoV-2 lineages to assist genomic epidemiology. Nat. Microbiol. 6, 415 (2020).
O’Toole, Á. et al. Assignment of epidemiological lineages in an emerging pandemic using the Pangolin tool. Virus Evol. 7, veab064. https://doi.org/10.1093/ve/veab064 (2021).
Tamura, T. et al. Virological characteristics of the SARS-CoV-2 XBB variant derived from recombination of two Omicron subvariants. Nat. Commun. 14, 2800 (2023).
Ma, K. C. et al. Genomic surveillance for SARS-CoV-2 variants: circulation of Omicron XBB and JN.1 Lineages - United states, May 2023-September 2024. MMWR Morb Mortal. Wkly. Rep. 73, 938–945 (2024).
Lu, Y., Ao, D., He, X. & Wei, X. The rising SARS-CoV-2 JN.1 variant: evolution, infectivity, immune escape, and response strategies. MedComm (2020) 5, e675. https://doi.org/10.1002/mco2.675 (2024).
Ou, G. et al. Evolving immune evasion and transmissibility of SARS-CoV-2: the emergence of JN.1 variant and its global impact. Drug Discov Ther. 18, 67–70 (2024).
World Health Organization. WHO COVID-19 Dashboard. https://data.who.int/dashboards/covid19/cases (2025).
Buathong, R. et al. Multiple clades of SARS-CoV-2 were introduced to Thailand during the first quarter of 2020. Microbiol. Immunol. 65, 405–409 (2021).
Puenpa, J. et al. Investigation of the molecular epidemiology and evolution of Circulating severe acute respiratory syndrome coronavirus 2 in Thailand from 2020 to 2022 via Next-Generation sequencing. Viruses 15, 1394 (2023).
Puenpa, J. et al. Genomic epidemiology and evolutionary analysis during XBB.1.16-predominant periods of SARS-CoV-2 Omicron variant in bangkok, thailand: December 2022-August 2023. Sci. Rep. 14, 645 (2024).
Global Initiative on Sharing All Influenza Data (GISAID). Pandemic coronavirus causing COVID-19. https://gisaid.org/ (2025).
Zanobini, P. et al. Global patterns of seasonal influenza activity, duration of activity and virus (sub)type circulation from 2010 to 2020. Influenza Other Respir Viruses. 16, 696–706 (2022).
Suntronwong, N. et al. Climate factors influence seasonal influenza activity in bangkok, Thailand. PLoS One. 15, e0239729. https://doi.org/10.1371/journal.pone.0239729 (2020).
The Ministry of Public Health. COVID-19 Situation, Thailand. https://ddc.moph.go.th/covid19-dashboard/ (2025).
Hinjoy, S. et al. Self-assessment of the Thai department of disease control’s communication for international response to COVID-19 in the early phase. Int. J. Infect. Dis. 96, 205–210 (2020).
World Health Organization Thailand. Weekly Situation Update no. 249. https://cdn.who.int/media/docs/default-source/searo/thailand/2022_10_12_tha-sitrep-249-covid-19.pdf?sfvrsn=148e6a0_1 (2022).
Suntronwong, N. et al. Characterizing genetic and antigenic divergence from vaccine strain of influenza A and B viruses Circulating in Thailand, 2017–2020. Sci. Rep. 11, 735 (2021).
Thongpan, I., Vongpunsawad, S. & Poovorawan, Y. Respiratory syncytial virus infection trend is associated with meteorological factors. Sci. Rep. 10, 10931 (2020).
Centers for Disease Control and Prevention. COVID-19 can surge throughout the year. https://www.cdc.gov/ncird/whats-new/covid-19-can-surge-throughout-the-year.html (2024).
Patterson, K. et al. Characterising COVID-19 school and childcare outbreaks in Canada in 2021: a surveillance study. BMJ Public. Health. 2, e000248. https://doi.org/10.1136/bmjph-2023-000248 (2024).
Shamsa, E. H., Shamsa, A. & Zhang, K. Seasonality of COVID-19 incidence in the united States. Front. Public. Health. 11, 1298593 (2023).
Zaher, K., Basingab, F., Alrahimi, J., Basahel, K. & Aldahlawi, A. Gender differences in response to COVID-19 infection and vaccination. Biomedicines 11, 1677 (2023).
Jin, J. M. et al. Gender differences in patients with COVID-19: focus on severity and mortality. Front. Public. Health. 8, 152 (2020).
Kim, J. M. et al. Prevalence of recent COVID-19 variants and Cell-based Omicron KP.3 infectivity analysis. PHWR 17, 1671–1681 (2024).
No, J. S. et al. Dynamics of SARS-CoV-2 variants during the XBB wave in the Republic of Korea. Virus Res. 350, 199471 (2024).
Wang, S. et al. Molecular evolutionary characteristics of SARS-CoV-2 emerging in the united States. J. Med. Virol. 94, 310–317 (2022).
Phyu, W. W. et al. Evolutionary dynamics of Whole-Genome influenza A/H3N2 viruses isolated in Myanmar from 2015 to 2019. Viruses 14, 2414 (2022).
Bhattacharjee, U., Chakrabarti, A. K., Kanungo, S. & Dutta, S. Evolutionary dynamics of influenza A/H1N1 virus Circulating in India from 2011 to 2021. Infect. Genet. Evol. 110, 105424 (2023).
Briese, T. et al. Global distribution of novel rhinovirus genotype. Emerg. Infect. Dis. 14, 944–947 (2008).
Yang, C. F. et al. Genetic diversity and evolution of human metapneumovirus fusion protein over Twenty years. Virol. J. 6, 138 (2009).
Centers for Disease Control and Prevention. The CDC 2019 novel coronavirus (2019 nCoV) real time Rt-PCR diagnostics. https://www.fda.gov/media/134922/download.pdf (2019).
Puenpa, J., Suwannakarn, K., Chansaenroj, J., Vongpunsawad, S. & Poovorawan, Y. Development of single-step multiplex real-time RT-PCR assays for rapid diagnosis of enterovirus 71, coxsackievirus A6, and A16 in patients with hand, foot, and mouth disease. J. Virol. Methods. 248, 92–99 (2017).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Sagulenko, P., Puller, V., Neher, R. A. & TreeTime Maximum-likelihood phylodynamic analysis. Virus Evol. 4, vex042. https://doi.org/10.1093/ve/vex042 (2018).
Yu, G., Smith, D. K., Zhu, H., Guan, Y. & Lam, T. T. Y. Ggtree: an R package for visualization and annotation of phylogenetic trees with their covariates and other associated data. Methods Ecol. Evol. 8, 28–36 (2017).
R Core Team. R: A Language and Environment for Statistical Computing.R Foundation for Statistical Computing, Vienna, Austria (2014).
Rambaut, A., Lam, T. T., Carvalho, M., Pybus, O. G. & L. & Exploring the Temporal structure of heterochronous sequences using tempest (formerly Path-O-Gen). Virus Evol. 2, vew007 (2016).
Bouckaert, R. et al. BEAST 2: a software platform for bayesian evolutionary analysis. PLoS Comput. Biol. 10, e1003537. https://doi.org/10.1371/journal.pcbi.1003537 (2014).
Lartillot, N. & Philippe, H. Computing Bayes factors using thermodynamic integration. Syst. Biol. 55, 195–207 (2006).
Kass, R. E. & Raftery, A. E. Bayes factors. J. Am. Stat. Assoc. 90, 773–795 (1995).
Ayres, D. L. et al. BEAGLE: an application programming interface and high-performance computing library for statistical phylogenetics. Syst. Biol. 61, 170–173 (2012).
Rambaut, A., Drummond, A. J., Xie, D., Baele, G. & Suchard, M. A. Posterior summarization in bayesian phylogenetics using tracer 1.7. Syst. Biol. 67, 901 (2018).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 35, 1547–1549 (2018).
Acknowledgements
We thank all staff from the Center of Excellence in Clinical Virology, Faculty of Medicine, Chulalongkorn University, for their help with the laboratory testing.
Funding
This work was supported by the Health Systems Research Institute (HSRI No. 68 − 026), the Center of Excellence in Clinical Virology, Chulalongkorn University, King Chulalongkorn Memorial Hospital, the MK Restaurant Group and Aunt Thongkam Foundation, the Education and Public Welfare Foundation, and the BJC Big C Foundation. Jiratchaya Puenpa reports that financial support was also provided by the Second Century Fund Fellowship of Chulalongkorn University.
Author information
Authors and Affiliations
Contributions
J.P., Y.P. drafted the manuscript; J.P., Y.P. analyzed the data, prepared figures, and interpreted the results; L.W., S.K. collected specimens and data; P.V., R.A. performed the laboratory tests; J.W., N.W., Y.P. designed the study and revised the manuscript. All authors reviewed the manuscript, provided critical feedback, and approved the final draft.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Informed consent
Due to the retrospective nature of the study, the Institutional Review Board of the Faculty of Medicine, Chulalongkorn University, Thailand (approval number IRB0933/67) waived the need to obtain informed consent.
Generative AI and AI-assisted technologies in the writing process
We hereby declare that generative AI (ChatGPT-4.0) was employed solely for the refinement of language and enhancement of readability within this manuscript, including the translation from Thai to English. All content, concepts, and findings were wholly conceived and developed by the authors. Following the language editing process, the authors conducted a thorough review of the final version to ensure its accuracy, integrity, and adherence to academic standards.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Puenpa, J., Vichaiwattana, P., Aeemjinda, R. et al. Seasonal trends and molecular evolution of SARS-CoV-2 in acute respiratory illness patients during the fifth year of COVID-19, Thailand. Sci Rep 15, 35703 (2025). https://doi.org/10.1038/s41598-025-19609-z
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-19609-z