Abstract
Long non-coding RNA (lncRNA) LINC01354 is involved in the progression of some cancer types, but the expression profile and prognostic value of this lncRNA in most tumor types, as well as its role and mechanism in esophageal cancer (ESCA), remain poorly characterized. Here, we identify LINC01354 as a tumor-suppressive lncRNA that inhibits ESCA progression by disrupting cell division signaling. Clinically, a pan-cancer analysis revealed widespread downregulation of LINC01354 in tumors compared to normal specimens, including ESCA and breast cancer (BRCA). Low tumor LINC01354 expression correlated with poor overall survival (OS) in ESCA and BRCA patients. In addition, the low expression of LINC01354 in tumors was validated again by two ESCA cohorts, and LINC01354 showed promising diagnostic potential. Moreover, tissue microarray (TMA) staining based on fluorescence in situ hybridization (FISH) once again confirmed the down-regulation of LINC01354 expression in ESCA. Functionally, enrichment analysis of LINC01354’s co-expressed genes indicated its critical role in ESCA cell cycle regulation, and its tumor suppressive effect was verified by an in vivo tumor-bearing mouse model. The inhibition of OE-LINC01354 on ESCA cell-cycle progression was further demonstrated by flow cytometry. Mechanistically, high-depth proteomic profiling revealed that overexpression of LINC01354 alters key cell division regulators, including AURKA, AURKB, BRCA2, CCND1, CDC20, and SOX9, etc. Thus, these findings establish the LINC01354-cell division axis as a novel tumor-suppressive mechanism in ESCA and highlight its therapeutic potential.
Similar content being viewed by others
Introduction
Esophageal cancer (ESCA), a leading cause of cancer-related mortality, claims over 440,000 lives annually worldwide1. The geographical variation in the incidence of ESCA varies greatly between the two histological subtypes (adenocarcinoma and squamous cell carcinoma), of which the latter has a high incidence in China1,2,3. Early ESCA detected by screening can be effectively treated under endoscopy, while the survival rate of inoperable advanced patients remains dismal4,5. Identifying the mechanisms that influence the malignant progression of ESCA is expected to improve clinical treatment strategies. Therefore, it is urgent to explore effective molecular targets and markers for the diagnosis and treatment of ESCA.
The biological functions of endogenous non-coding RNAs in physiological and pathological processes have been widely recognized, earning the Nobel Prize in Physiology or Medicine. Among these, long non-coding RNAs (lncRNAs) are increasingly acknowledged for their critical roles in human diseases, spanning processes from cell division to apoptosis, and from development to tumorigenesis6,7,8. Over the past decade, numerous ESCA-associated lncRNAs, such as CASC9, LINC00022, TTN-AS1, and TSPEAR-AS29,10,11,12, have been identified. These lncRNAs exhibit oncogenic functions in ESCA and demonstrate diagnostic or prognostic values, highlighting their potential for clinical applications. However, current understanding of tumor-suppressive lncRNAs in ESCA remains limited. LINC01354 was initially identified as a lncRNA in colon cancer, with studies reporting its involvement in the proliferation and metastasis of colon and gastric cancers13,14,15. Nevertheless, the expression status, clinical relevance, and functional mechanism of LINC01354 in most human cancers remain unclear.
By mining multiple bioinformatics databases, we identified that LINC01354 is generally down-regulated in human pan-cancer. In six cancer types ( BRCA, ESCA, KIRC, UVM, COAD, and STAD), its expression level predicted patients’ overall survival. Moreover, low level of LINC01354 showed strong diagnostic performance for ESCA. Preclinically, overexpression of LINC01354 significantly suppressed the growth of ESCA tumors in nude mice and inhibited cell-cycle process in vitro. Mechanically, using high-depth proteomics technology, we uncovered that LINC01354 regulated the levels of proteins associated with cell division signaling. In conclusion, our findings advance the understanding of tumor-suppressive lncRNAs whose downregulation promotes ESCA and provide a candidate biomarker for its diagnosis and prognosis.
Materials and methods
ENCORI database
ENCORI16 is a comprehensive cancer database that integrates innovative algorithms and consolidates sequencing data from dozens of cancer types. Users can mine cancer-related molecules from this database based on their research interests and investigate their clinical significance in cancer. We utilized this tool to analyze the pan-cancer expression profile of LINC01354 and its association with patients’ OS.
GEPIA platform
GEPIA17 is a well-established cancer analysis server that employs specific algorithms to integrate RNA sequencing data from over 9700 tumor and 8500 normal samples. Researchers can perform differential expression analyses for specific cancer types using GEPIA’s customizable functionalities. In this study, we utilized this database to investigate the pan-cancer expression of LINC01354 in humans.
Receiver operating characteristic curve
Receiver operating characteristic (ROC) curves were employed to evaluate the diagnosis of LINC01354 in ESCA. First, the GSE53622 and GSE53624 datasets were downloaded from the Gene Expression Omnibus (GEO) database, followed by obtaining the expression data of LINC01354 in ESCA tissues and paired normal tissues. Finally, ROC analysis was performed based on these expression profiles, where a larger area under the curve (AUC) indicates superior diagnostic performance.
Enrichment analysis
Functional enrichment analysis was conducted to explore the biological roles of LINC01354 in ESCA. First, co-expressed genes of LINC01354 in ESCA samples were obtained from the GEPIA database, followed by analyzing the enriched pathways of these co-expressed genes using the online server WebGestalt18. Functional enrichment was based on Gene Ontology Biological Process (GO-BP) and the Kyoto Encyclopedia of Genes and Genomes (KEGG).
Cell culture and transfection
Four human ESCA cell lines (TE1, KYSE70, KYSE150, and KYSE450) and one esophageal epithelial cell line Het-1 A were cultured in RPMI 1640 medium supplemented with 10% fetal bovine serum (FBS). Het-1 A was obtained from Fuheng Biology (Shanghai, China). Human ESCA cell lines KYSE150 and TE1 were purchased from Procell (Wuhan, China). KYSE70 and KYSE450 were kindly provided by the Bioengineering and Transformation Laboratory of Zhengzhou University. The adherent cells were passaged following trypsin digestion and incubated at 37 °C with 5% CO₂. A recombinant plasmid for LINC01354 overexpression (OE-LINC01354) was purchased from GeneChem (Shanghai, China), and cell transfection was performed using Lipo3000 reagent.
Quantitative real-time polymerase chain reaction
Adherently cultured Het-1 A, TE1, KYSE70, KYSE150, and KYSE450 cells were detached using trypsin solution and collected into centrifuge tubes. Total RNA was extracted from the cells via the Trizol method, and RNA purity and concentration were measured. The RNA was reverse-transcribed into cDNA using a reverse transcription kit (Novoprotein, China). Quantitative real-time polymerase chain reaction (qRT-PCR) was performed using the SYBR kit (Novoprotein, China) with cDNA as the template. The housekeeping gene GAPDH served as the internal reference, and the relative expression of LINC01354 across cell lines was analyzed. Primer sequences are listed in Supplementary Table 1.
Fluorescence in situ hybridization (FISH)
The expression level of LINC01354 in a microarray including normal tissues and ESCA tissues (Servicebio, China) was detected by fluorescence in situ hybridization. Briefly, a paraffin tissue microarray was dewaxed in dewaxing transparent solution and gradient ethanol, and then soaked in DEPC water. Place the microarray in a citric acid repair box and incubate at 90℃ for 48 min. Add proteinase K solution and digest at 40℃ for 10 min. After pre-hybridization, hybridization solution containing fluorescent probes (Servicebio, China) was added to the microarray and incubated overnight at 40 °C. After DAPI staining, the microarray was placed under a fluorescence microscope to collect images. The probe sequences is shown in Supplementary Table 1.
In vivo tumor growth
A nude mouse xenograft experiment was conducted to evaluate the effects of LINC01354 overexpression on ESCA tumorigenesis and growth. In brief, KYSE150 cells were evenly seeded in culture dishes and transfected with plasmids (OE-NC and OE-LINC01354). Once sufficient cell numbers were achieved, the cells were detached using trypsin solution and collected into centrifuge tubes. Saline was added to resuspend the cells, and the cell concentration was determined. The ESCA cells were then subcutaneously injected into the right forelimbs of male BALB/c nude mice (purchased from Beijing HFK Company, China). After tumor formation, tumor size was measured using the formula: length × width² × 0.5. Finally, mice were anesthetized with isoflurane and euthanized via cervical dislocation, and tumors were excised for analysis.
Flow cytometry analysis
Flow cytometry analysis was employed to investigate the effects of LINC01354 overexpression on ESCA cell-cycle process based on a propidium iodide (PI) staining kit (7Sea, China) and a flow cytometer (Beckman, USA). In short, the transfected cells were collected in centrifuge tubes, and then 70% ethanol was added and the tubes were incubated at 4℃ for 3 h. Subsequently, the cells were stained with the PI solution at 37℃ for 30 min and then examined on a flow cytometer.
10× high-depth proteomics
The potential molecular mechanism by which LINC01354 overexpression suppresses ESCA tumorigenesis was investigated using high-depth proteomic technology (PTMBIO, China). Briefly, KYSE150 cells were uniformly seeded in culture dishes followed by plasmid transfection (OE-NC and OE-LINC01354). Upon reaching confluence, cells were detached using trypsin and harvested by centrifugation. Total cellular protein was extracted by adding lysis buffer containing 1% SDS and 1% protease inhibitor cocktail to the cell pellets. Total proteins were quantified and subsequently subjected to enzymatic digestion to generate peptides. Finally, the peptides were analyzed using 10× Proteomics technology for comprehensive detection and proteomic profiling.
Statistical analysis
The experimental results are presented as mean ± standard deviation. Statistical comparisons between any two groups were analyzed using t-test. Graphical illustrations and statistical analyses were performed using GraphPad Prism 9.0 and SPSS 19.0 softwares. A p-value less than 0.05 was considered statistically significant. We confirm that all methods were performed in accordance with the relevant guidelines and regulations.
Results
The expression landscape of LINC01354 across human pan-cancer
As a newly identified disease-associated lncRNA, LINC01354 has been rarely reported in cancer research. Current evidence from data mining and cellular model investigations has primarily elucidated its functional role in gastrointestinal cancers. To systematically characterize its expression landscape across human malignancies, we used the ENCORI database for comprehensive analysis. The results demonstrated that LINC01354 was commonly downregulated in human cancers. Tumor tissues exhibited significantly reduced LINC01354 expression levels compared with normal counterparts. This pan-cancer pattern was consistently observed in 12 cancer types, encompassing bladder urothelial carcinoma (BLCA, Fig. 1A), breast invasive carcinoma (BRCA, Fig. 1B), cholangiocarcinoma (CHOL, Fig. 1C), colon adenocarcinoma (COAD, Fig. 1D), ESCA (Fig. 1E), head and neck squamous cell carcinoma (HNSC, Fig. 1F), kidney renal papillary cell carcinoma (KIRP, Fig. 1G), lung adenocarcinoma (LUAD, Fig. 1H), lung squamous cell carcinoma (LUSC, Fig. 1I), stomach adenocarcinoma (STAD, Fig. 1J), thyroid carcinoma (THCA, Fig. 1K), and uterine corpus endometrial carcinoma (UCEC, Fig. 1L).
LINC01354 is down-regulated in pan-cancer. (A–L) By mining the ENCORI database, it was found that LINC01354 expression was reduced in 12 cancers, including BLCA (411T vs. 19 N), BRCA (1104T vs. 113 N), CHOL (36T vs. 9 N), COAD (471T vs. 41 N), ESCA (162T vs. 11 N), HNSC (502T vs. 44 N), KIRP (289T vs. 32 N), LUAD (526T vs. 59 N), LUSC (501T vs. 49 N), STAD (375T vs. 32 N), THCA (510T vs. 58 N), and UCEC (548T vs. 35 N), ****p < 0.0001. T represents tumor tissues and N represents normal tissues.
The expression heterogeneity of LINC01354 in human malignancies
Our integrative analysis of the GEPIA database revealed dysregulated expression of LINC01354 across 17 cancer types. The lncRNA demonstrated significant downregulation in 15 malignancies, including BLCA, BRCA, cervical squamous cell carcinoma and endocervical adenocarcinoma (CESC), COAD, ESCA, LUAD, LUSC, ovarian serous cystadenocarcinoma (OV), pancreatic adenocarcinoma (PAAD), prostate adenocarcinoma (PRAD), rectal adenocarcinoma (READ), STAD, THCA, UCEC, and uterine carcinosarcoma (UCS) (Fig. 2A and C). Conversely, elevated expression of LINC01354 was observed in 2 cancer subtypes, including glioblastoma multiforme (GBM) and low grade glioma (LGG) (Fig. 2D). These findings collectively demonstrate the heterogeneous regulatory patterns of LINC01354 expression in human malignancies.
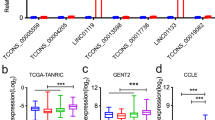
LINC01354 shows heterogeneous expression in human cancers. (A–D) Results from the GEPIA database showed that LINC01354 was deregulated in 17 cancer types, including BLCA (404T vs. 28 N), BRCA (1085T vs. 291 N), CESC (306T vs. 13 N), COAD (275T vs. 349 N), ESCA (182T vs. 286 N), LUAD (483T vs. 347 N), LUSC (486T vs. 338 N), OV (426T vs. 88 N), PAAD (179T vs. 171 N), PRAD (492T vs. 152 N), READ (92T vs. 318 N), STAD (408T vs. 211 N), THCA (512T vs. 337 N), UCEC (174T vs. 91 N), UCS (57T vs. 78 N), GBM (163T vs. 207 N), and LGG (518T vs. 207 N), ****p < 0.0001. (E) A comparative analysis of the results from ENCORI and GEPIA showed that LINC01354 was dysregulated in 9 cancers.
LINC01354 expression demonstrates significant prognostic associations across multiple cancer types
Cross-analysis of ENCORI and GEPIA results revealed consistent downregulation of LINC01354 in 9 malignancies, including BLCA, BRCA, COAD, ESCA, UCEC, LUAD, LUSC, STAD, and THCA (Fig. 2E). Through comprehensive survival analysis utilizing the ENCORI database, we systematically investigated the clinical relevance of LINC01354 expression in cancer prognosis. The data demonstrated that tumor elevated LINC01354 expression correlated with improved OS in BRCA, ESCA, KIRC, and uveal melanoma (UVM) patients (Fig. 3A and D), while paradoxically associating with poorer survival outcomes in COAD and STAD cohorts (Fig. 3E and F). These results indicate that LINC01354 is associated with patient prognosis.
LINC01354’s expression correlates with patient survival. (A–F) Data from ENCORI displayed that expression of LINC01354 was associated with overall survival of cancer patients. High expression of LINC01354 in tumors led to good prognosis in BRCA (HR = 0.68, p = 0.019), ESCA (HR = 0.58, p = 0.036), KIRC (HR = 0.53, p = 6.3E-5), and UVM (HR = 0.23, p = 0.0014) patients, but poor prognosis in COAD (HR = 1.81, p = 0.0035) and STAD (HR = 1.6, p = 0.0055) patients.
LINC01354 is downregulated in ESCA and shows promising diagnosis
Through integrated analyses of LINC01354 expression profiles (Fig. 2E) and clinical prognostic data (Fig. 3), we revealed that this lncRNA demonstrates reduced expression in both ESCA and BRCA, and correlates with shortened patient survival (Fig. 4A). These characteristics suggest LINC01354 may serve critical regulatory functions in these two cancers. Given our primary research focus on gastrointestinal malignancies, subsequent investigations were specifically designed to delineate the functional roles and mechanistic underpinnings of LINC01354 in ESCA pathogenesis. The biological significance of LINC01354 in BRCA warrants systematic elucidation in future investigations.
High tumor expression of LINC01354 holds diagnostic potential in ESCA. (A) A comparative analysis of LINC01354 expression and prognosis revealed that LINC01354 was decreased in ESCA and BRCA and led to poor prognosis in patients. (B, C) The high expression of LINC01354 in ESCA was validated in additional two cohorts. (D) Paired analysis showed that LINC01354 was down-regulated in 95% of ESCA patients in GSE53622 cohort. (E) Further analysis depicted that LINC01354 was decreased in 89.9% of ESCA patients in GSE53624 cohort. (F, G) ROC analysis was used to evaluate the diagnostic potential of LINC01354 in the GSE53622 (95% confidence interval: 0.8317 to 0.9544) and GSE53624 (95% confidence interval: 0.8374 to 0.9277) cohorts. (H) The expression of LINC01354 in Het-1 A, TE1, KYSE70, KYSE150, and KYSE450 cells was examined by qRT-PCR, **p < 0.01, ***p < 0.001, independent-samples T test. (I) FISH was employed to detect the expression of LINC01354 in ESCA TMA, scale bar = 800 μm.
Given that results from both ENCORI and GEPIA databases indicated low expression of LINC01354 in ESCA, we incorporated two additional ESCA datasets (GSE53622 containing 60 paired tissue samples; GSE53624 containing 119 paired tissue samples) for analysis. Unsurprisingly, LINC01354 exhibited significantly lower expression levels in ESCA compared with paired normal tissues (Fig. 4B and C). Further paired analysis revealed that in the GSE53622 cohort, LINC01354 showed reduced expression in 95% of ESCA cases (Fig. 4D). Similarly, in the GSE53624 cohort, LINC01354 was downregulated in 89.9% of ESCA samples (Fig. 4E). These findings suggest an essential role of LINC01354 in ESCA pathogenesis. We then investigated the diagnostic potential of LINC01354 underexpression in ESCA. ROC analysis demonstrated excellent diagnostic performance with AUC values all exceeding 0.88 (Fig. 4F and G). Subsequent qRT-PCR confirmed significantly lower transcription levels of LINC01354 in ESCA cell lines compared with normal esophageal epithelial cell line Het-1 A (Fig. 4H). We next used FISH to detect the expression of LINC01354 in the ESCA tissue microarray, and confirmed once again its low expression status in ESCA tumors (Fig. 4I). Collectively, these results indicate that LINC01354 underexpression in ESCA represents a promising diagnostic biomarker.
Overexpression of LINC01354 suppresses ESCA progression
Given the observed downregulation of LINC01354 in ESCA and its association with poor patient prognosis, we hypothesized that this lncRNA may function as a tumor suppressor. To investigate this, we obtained LINC01354 co-expressed gene sets from the GEPIA database and performed functional enrichment analysis. Both GO-BP and KEGG pathway analyses consistently demonstrated LINC01354’s involvement in cell cycle regulation, suggesting its role in modulating malignant proliferation in ESCA (Fig. 5A and B). Subsequently, we conducted xenograft tumor experiments in nude mice. ESCA cells overexpressing LINC01354 were subcutaneously inoculated into nude mice. Consistent with our hypothesis, LINC01354 overexpression significantly inhibited in vivo tumor growth, as evidenced by marked reductions in both tumor volume and weight (Fig. 5C and E). The inhibition of LINC01354 on ESCA cell-cycle progression was further demonstrated by flow cytometry analysis (Fig. 5F and G). These data collectively indicate that LINC01354 overexpression suppresses ESCA progression.
LINC01354 is involved in cell-cycle regulation and curbs tumorigenesis in ESCA. (A, B) Functional enrichment analysis of LINC01354 associated genes in ESCA revealed the biological function of LINC01354. (C–E) The effect of overexpression of LINC01354 on ESCA tumor growth was evaluated in a nude mouse model, **p < 0.01. (F, G) The inhibition of LINC01354 on KYSE150 cell cycle process was validated by flow cytometry analysis, ****p < 0.0001.
High-depth proteomics reveals the mechanism by which LINC01354 inhibits ESCA
Given that LINC01354 is downregulated in ESCA and its overexpression significantly suppresses ESCA growth, we sought to investigate its tumor-suppressive mechanisms. To this end, we employed 10× high-depth proteomics technology to profile proteomic alterations in ESCA cells overexpressing LINC01354 (Fig. 6A and B). High-throughput screening revealed that LINC01354 upregulation increased the expression of 46 proteins and decreased 81 proteins (Fig. 6C and E). Functional enrichment analysis of the 81 downregulated proteins demonstrated their predominant involvement in proliferation-associated pathways, including cell division and cell cycle regulation (Fig. 6F and H). Notably, LINC01354 overexpression reduced the levels of 24 cell division-related proteins (including AMBRA1, AURKA, AURKB, BRCA2, BUB1, CCND1, CDC20, CDCA2, CDCA8, CENPJ, CENPL, EVI2B, HJURP, HUS1, KLHL42, MIS18BP1, NSD2, PCLAF, PLK1, PRR11, RAD51D, RRM2, SOX9, and UBE2S) (Fig. 6I and J), molecularly validating its tumor-suppressive function. These high-depth proteomic findings collectively indicate that LINC01354 suppresses ESCA progression via inhibiting cell division signaling pathways.
Overexpression of LINC01354 inhibits cell division signaling in ESCA. (A–E) High-depth proteomics was conducted to explore the changes of protein levels in ESCA cells following LINC01354 overexpression. The differential expression threshold is defined as a fold change greater than 1.5 and a p-value less than 0.05. (F–H) GO-BP, Reactome, and KEGG analyses were carried out to investigate LINC01354-related signaling of 81 reduced proteins. (I, J) Overexpression of LINC01354 in ESCA cells resulted in decreased expression of 24 proteins associated with cell division signaling pathways, *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
Discussion
Both aberrant cell division signaling and dysregulated lncRNA expression contribute to cancer initiation and progression19,20. However, the interplay between them and their roles in malignant proliferation remain poorly understood. This study delineates LINC01354 as a pan-cancer tumor suppressor, with pronounced roles in ESCA. Its underexpression correlates with aggressive disease and poor survival, while restoration inhibits tumor growth via cell division blockade.
The rapid advancement of high-throughput sequencing technology has significantly enhanced our understanding of complex cellular and molecular events within tumors and accelerated the translation of molecular oncology from bench to bedside21,22,23. Although aberrantly expressed lncRNAs in ESCA have been extensively documented, most studies have focused on upregulated oncogenic lncRNAs24,25,26, while the functional characterization of downregulated lncRNAs remains limited. In this study, we systematically constructed the pan-cancer expression profile of LINC01354 by integrating multiple tumor databases. Our analysis revealed its consistent downregulation across nine cancer types, including ESCA, suggesting its potential role as a conserved tumor suppressor across multiple cancer types.
The functional diversity and tissue specificity of lncRNAs endow them with substantial clinical potential. For instance, lnc191 is upregulated in ESCA, and its overexpression correlates with poor patient prognosis27. Similarly, NAMPT-AS exhibits elevated expression in colorectal cancer tissues and circulating levels, suggesting its utility as a diagnostic biomarker28. In this study, we identified that LINC01354 is downregulated in ESCA, and its reduced expression serves as an independent risk factor for shortened patient survival. Furthermore, low LINC01354 levels demonstrate promising diagnostic potential for ESCA. Although its clinical diagnostic and prognostic value requires further validation, our findings provide first-line evidence to support its future translation into clinical applications. We noted that the expression of LINC01354 was relatively low in COAD and STAD, and its high expression indicated a poorer prognosis for patients with these two types of cancers, which contradicted the general understanding. We hypothesized that LINC01354 might play a contradictory role in the complex processes of the occurrence and progression of COAD/STAD. Therefore, the biological function and prognostic value of LINC01354 in these two cancers are worthy of in-depth study.
Targeting and knocking down oncogenic lncRNAs represents a promising strategy against cancer, yet achieving systemic suppression of their expression remains challenging29,30. In contrast, overexpression of tumor-suppressive lncRNAs may emerge as a novel anticancer paradigm. This study demonstrates that LINC01354 overexpression effectively inhibits ESCA growth in vivo and retards cell-cycle progression in vitro. It is therefore imperative to validate the therapeutic potential of LINC01354 supplementation for ESCA using diverse animal models. Continuous cell division is required for the uncontrolled malignant proliferation of tumors. Our findings reveal that LINC01354 overexpression alters the expression status of multiple cell division-related proteins. To our knowledge, this is the first report demonstrating suppression of cell division signaling by LINC01354 in ESCA.
In summary, multi-database analyses identified LINC01354 as a downregulated molecule across various malignancies that correlates significantly with patient prognosis, suggesting its potential as a clinical biomarker for ESCA and other cancers. The tumor-suppressive effects of LINC01354 overexpression in xenograft models indicate its therapeutic candidacy for ESCA. High-depth proteomics technology revealed significant suppression of cell division signaling following LINC01354 overexpression. These findings address the knowledge gap regarding ESCA pathogenesis associated with under-expression of tumor-suppressive lncRNAs, while highlighting the therapeutic potential of the LINC01354-cell division signaling axis in ESCA treatment.
Data availability
Data are available from the corresponding author upon reasonable request.
References
Bray, F. et al. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 74 (3), 229–263 (2024).
Sung, H. et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71 (3), 209–249 (2021).
Chen, W. et al. Cancer statistics in China, 2015. CA Cancer J. Clin. 66 (2), 115–132 (2016).
Sabatelle, R. C., Colson, Y. L., Sachdeva, U. & Grinstaff, M. W. Drug delivery opportunities in esophageal cancer: current treatments and future prospects. Mol. Pharm. 21 (7), 3103–3120 (2024).
Qu, H. T. et al. Esophageal cancer screening, early detection and treatment: current insights and future directions. World J. Gastrointest. Oncol. 16 (4), 1180–1191 (2024).
Chunhui, G., Yanqiu, Y., Jibing, C., Ning, L. & Fujun, L. Exosomes and non-coding rnas: bridging the gap in Alzheimer’s pathogenesis and therapeutics. Metab. Brain Dis. 40 (1), 84 (2025).
Zhang, Y. & Wen, Y. The role of LncRNA NEAT1 in gastric cancer: tumor development and potential therapeutic target. Curr. Pharm. Des. 29 (28), 2213–2222 (2023).
Wang, Z., Wang, P. S. & Yang, C. Dysregulation of long non-coding RNAs-the novel Lnc in metal toxicity and carcinogenesis. Curr. Environ. Health Rep. 12 (1), 3 (2024).
Liang, Y. et al. LncRNA CASC9 promotes esophageal squamous cell carcinoma metastasis through upregulating LAMC2 expression by interacting with the CREB-binding protein. Cell. Death Differ. 25 (11), 1980–1995 (2018).
Cui, Y. et al. RNA m6A demethylase FTO-mediated epigenetic up-regulation of LINC00022 promotes tumorigenesis in esophageal squamous cell carcinoma. J. Exp. Clin. Cancer Res. 40 (1), 294 (2021).
Lin, C. et al. Functional role of a novel long noncoding RNA TTN-AS1 in esophageal squamous cell carcinoma progression and metastasis. Clin. Cancer Res. 24 (2), 486–498 (2018).
Zhang, C., Cui, Y. & Targeting TSPEAR-AS2 suppresses tumor growth and interferon signaling in esophageal cancer. Sci. Rep. 14 (1), 28768 (2024).
Wang, X. et al. A 15-lncRNA signature predicts survival and functions as a CeRNA in patients with colorectal cancer. Cancer Manag Res. 10, 5799–5806 (2018).
Li, J., He, M., Xu, W. & Huang, S. LINC01354 interacting with hnRNP-D contributes to the proliferation and metastasis in colorectal cancer through activating Wnt/β-catenin signaling pathway. J. Exp. Clin. Cancer Res. 38 (1), 161 (2019).
Fang, F. et al. LINC01354 enhances the metastatic ability of gastric cancer cells by adjusting miR-153-5p/CADM2 expression. Am. J. Cancer Res. 13 (6), 2630–2643 (2023).
Li, J. H., Liu, S., Zhou, H., Qu, L. H. & Yang, J. H. StarBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res. 42 (Database issue), D92–D97 (2014).
Tang, Z., Kang, B., Li, C., Chen, T. & Zhang, Z. GEPIA2: an enhanced web server for large-scale expression profiling and interactive analysis. Nucleic Acids Res. 47 (W1), W556–W560 (2019).
Elizarraras, J. M. et al. WebGestalt 2024: faster gene set analysis and new support for metabolomics and multi-omics. Nucleic Acids Res. 52 (W1), W415–W421 (2024).
Gugnoni, M., Kashyap, M. K., Wary, K. K. & Ciarrocchi, A. LncRNAs: the unexpected link between protein synthesis and cancer adaptation. Mol. Cancer. 24 (1), 38 (2025).
Siddique, R. et al. LncRNAs and cyclin-dependent kinases: unveiling their critical roles in cancer progression. Pathol. Res. Pract. 258, 155333 (2024).
Tao, S. et al. Long noncoding RNA study: genome-wide approaches. Genes Dis. 10 (6), 2491–2510 (2022).
Khawar, M. B. et al. Diagnostic, prognostic and therapeutic potential of long noncoding RNAs in cancer. Mol. Biol. Rep. 49 (3), 2311–2319 (2022).
Grillone, K. et al. A systematic review of non-coding RNA therapeutics in early clinical trials: a new perspective against cancer. J. Transl Med. 22 (1), 731 (2024).
Cui, Y. et al. ESCCAL-1 promotes cell-cycle progression by interacting with and stabilizing galectin-1 in esophageal squamous cell carcinoma. NPJ Precis Oncol. 6 (1), 12 (2022).
Wang, M. et al. LncRNA U731166 increases the accumulation of TGFBR1 by sponging miR-3607-3p in esophageal squamous-cell carcinomas (ESCC) to promote tumor metastasis. Iran. J. Biotechnol. 22 (3), e3391 (2024).
Zhang, C., Lian, H., Xie, L., Yin, N. & Cui, Y. LncRNA ELFN1-AS1 promotes esophageal cancer progression by up-regulating GFPT1 via sponging miR-183-3p. Biol. Chem. 401 (9), 1053–1061 (2020).
Wei, S. et al. Hypoxia induced Lnc191 upregulation dictates the progression of esophageal squamous cell carcinoma by activating GRP78/ERK pathway. Adv. Sci. (Weinh). 12 (4), e2406674 (2025).
Rizk, N. I. et al. Revealing the role of serum exosomal novel long non-coding RNA NAMPT-AS as a promising diagnostic/prognostic biomarker in colorectal cancer patients. Life Sci. 352, 122850 (2024).
Subaiea, G. M. et al. Non-coding RNAs (ncRNAs) and multidrug resistance in glioblastoma: therapeutic challenges and opportunities. Pathol. Res. Pract. 253, 155022 (2024).
Ammad, M. et al. Advancements in long non-coding RNA-based therapies for cancer: targeting, delivery, and clinical implications. Med. Oncol. 41 (11), 292 (2024).
Funding
This study was supported by the Medical Science and Technology Project of Henan Province (SBGJ202403054), the Scientific Research Project of Zhengzhou Central Hospital (KYQDJJ2023001).
Author information
Authors and Affiliations
Contributions
YBC conceived this study, performed bioinformatics analysis, and wrote the manuscript. XYW, FFG and MHS participated in experiments and manuscript revision.
Corresponding author
Ethics declarations
Consent for publication
All authors approved the manuscript for publication.
Ethic statement
The animal experiment was approved by the Ethical Review Committee of Zhengzhou Central Hospital. The study was reported in accordance with ARRIVE guidelines.
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Cui, Y., Wu, X., Gong, F. et al. Overexpression of LINC01354 predicts well prognosis and suppresses esophageal cancer progression. Sci Rep 15, 36366 (2025). https://doi.org/10.1038/s41598-025-20455-2
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-20455-2