Abstract
End-stage renal disease (ESRD) is associated with high morbidity and mortality. Identifying patients with stage 4 chronic kidney disease (CKD) at risk of short-term progression to ESRD remains challenging. Accurate prediction can improve advanced care planning and patient outcomes. This study aimed to develop and validate a machine learning (ML) model for predicting progression within 25 weeks (approximately six months) of ESRD in patients with stage 4 CKD. Electronic health records (EHRs) of patients with stage 4 CKD were analyzed. Nine ML models including Ridge regression (Ridge), random forest (RF), and eXtreme Gradient Boosting (XGBoost) were used to predict short-term progression to ESRD within 25 weeks. The models were trained and externally validated using the data of 346 and 105 patients. Of the 451 patients with stage 4 CKD, 219 developed ESRD. Among the evaluated models, XGBoost demonstrated the best overall performance. In the internal validation, it achieved an area under the curve (AUC) of 0.93, an accuracy of 0.90, and an F1 score of 0.89. In the external validation, XGBoost maintained the highest AUC (0.85), accuracy (0.79), and F1 score (0.79), along with the highest average precision (0.89) and a low log-loss (0.48), indicating strong discriminative ability and good generalizability. The top predictive features included high-density lipoprotein cholesterol, Alb, Cys C, ApoB, FGB, Bun, Neutrophil, and Total cholesterol. This study demonstrated the feasibility of ML for assessing ESRD prognosis based on easily accessible clinical features. XGBoost demonstrated superior performance in both internal and external validation, suggesting its potential for future patient screening.
Similar content being viewed by others
Introduction
Chronic kidney disease (CKD) is a major public health concern worldwide that contributes to end-stage renal disease (ESRD), cardiovascular disease (CVD), and even death. The Global Burden of Disease (GDB) predicts CKD becoming the fifth leading cause of mortality by 20401. Its estimated prevalence in China ranges from 8.2 to 10.8%2,3. CKD progression rates vary greatly among individuals. In advanced CKD, particularly in stage 4 (defined by an estimated glomerular filtration rate [eGFR] between 15 and 29 mL/kg/min), patients face a higher risk of progression to ESRD and a significantly greater likelihood of CVD compared to those in earlier stages4.
The Kidney disease: Improving Global Outcomes (KDIGO) guidelines recommend that the care target for stage 4 CKD should be to slow down the disease progression and prepare for renal replacement therapy, including options for kidney transplantation, dialysis, and vascular access5. Notably, improved survival and a higher probability of access to kidney transplantation are associated with early nephrological care in patients prior to dialysis6. However, a large proportion of patients feel ill-equipped for dialysis due to the short-term progression of ESRD. Therefore, accurate risk assessment is crucial to identify patients with stage 4 CKD at high short-term risk of ESRD progression. Early detection can enhance advanced care outcomes in patients with severe CKD by guiding decision-making and enabling more effective management of CKD. Furthermore, effective prediction tools can enhance the efficacy of health systems by tailoring care to individual risk profiles.
Advances in digitalization and the widespread use of electronic health records (EHRs) in clinical medicine have generated large amounts of real-world clinical data for the development of clinical decision-making systems. Machine learning (ML) approaches, with an ability to predict outcomes from multiple complex and nonlinear interactions, have demonstrated great potential in solving real-world problems of diagnosis and prognosis. However, few studies have applied ML to prognostic models for stage 4 CKD, with most focusing on diabetic kidney disease (DKD).
This study aimed to develop and externally validate a simple, accurate ML model for predicting short-term progression of stage 4 CKD to ESRD using data from two CKD cohorts. We hypothesized that using laboratory features alone in ML models would accurately identify patients at high risk of progressing to ESRD within 6 months.
Materials and methods
Study design
This retrospective cohort study was conducted to identify baseline clinical characteristics that predict short-term progression in patients with advanced CKD. We retrospectively reviewed EHRs and selected patients with confirmed stage 4 CKD, recording their subsequent progression to stage 5 CKD as the study outcome.
Study population
Development cohort
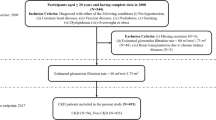
The development cohort was derived from the EHRs of the Second Affiliated Hospital of Harbin Medical University. We included adult patients (> 18 years) who were initially diagnosed with stage 4 CKD (eGFR 15–29 mL/min/1.73 m², according to KDIGO 2024 criteria) between January 2017 and December 2023.
Validation cohort
The validation cohort was derived from the EHRs of Harbin 242 Hospital of Genertec Medical. Using the same criteria, we included adult patients who were initially diagnosed with stage 4 CKD between January 2016 and July 2024.
For both cohorts, CKD staging was determined using ICD-10-CM discharge codes (N18.804-N18.907), which represent confirmed diagnoses in the hospital discharge summary system. Eligible patients were required to have at least two hospital admissions. Patients with a baseline diagnosis of stage 5 CKD (eGFR < 15 mL/min/1.73 m²) were excluded. There were no substantive differences between the development and validation cohorts in terms of healthcare setting, eligibility criteria, outcome definition, or candidate predictors.
Ethical approval and reporting standards
This retrospective cohort study was approved by the Clinical Ethics Review Committee of the Second Affiliated Hospital of Harbin Medical University (YSKY2023-212). All methods were performed in accordance with the relevant guidelines and regulations, including the ethical principles outlined in the Declaration of Helsinki. The requirement for informed consent was waived by the ethics committee due to the retrospective nature of the study. Reporting adhered to the STROBE guidelines (Strengthening the Reporting of Observational Studies in Epidemiology) and the TRIPOD + AI statement (Transparent Reporting of a Multivariable Prediction Model for Individual Prognosis or Diagnosis-AI Extension).
Laboratory and clinical characteristics
The candidate predictors, primarily laboratory markers, were identified through a two-step process: by literature review and the clinician’s and the patients’ engagement. The following clinical characteristics were collected at baseline: age, gender, 24-h urine urinary total protein (UTP), serum albumin (Alb), albumin/globulin ratio (AGR), blood urea nitrogen (BUN), Uric Acid (UA), Serum creatinine, blood urea nitrogen/creatinine ratio(BCR), total cholesterol, triglyceride (TG), high-density lipoprotein cholesterol (HDL-C), low-density lipoprotein cholesterol (LDL-C), hemoglobin (Hb), Cystatin-C (CysC), C-reactive protein (CRP), Cholinesterase (ChE), Gamma-glutamyltransferase (GGT), lactate dehydrogenase (LDH), serum sodium (Na+), serum potassium (K+), Serum chloride (Cl−), red blood cell (RBC), Neutrophil(Ne), Red blood cell distribution width, red blood cells per high-power field (RBC-HPF), percentage of lymphocytes in WBCs (%LYM), percentage of monocyte (Mon%) in WBCs, eosinophils in WBCs (EOS), prothrombin time (PT), Prothrombin time ratio (PTR), phosphorus (P), glucose (GLU), Urine specific gravity(SG), plasma fibrinogen (FIB), urine Epithelial cells (EPC), cardiac troponin I (cTnI), anion gap(AG), Apolipoprotein B (APOB), Direct bilirubin(Dbi).
Outcome definition
The primary outcome was defined as progression from baseline stage 4 CKD to stage 5 CKD within 25 weeks (approximately 6 months) of follow-up. Patients who did not progress to stage 5 within this 25-week window were classified as non-progressors. After the first mention, we refer to this outcome as “short-term progression”.
The observation window was selected based on the following considerations. First, 6-month horizons have been widely adopted in CKD research as clinically meaningful timeframes, for example in models developed from health insurance claim data7, aggregated longitudinal EHRs data in diabetic patients8, and longitudinal EHR-based models of diabetic kidney disease progression9. Second, in our participating hospitals, outpatient follow-up is typically scheduled every 3–6 months, which ensures sufficient data capture and aligns with routine clinical practice. Third, exploratory analysis demonstrated that a 25-week horizon provided an optimal balance between sufficient event numbers and model stability.
This definition is distinct from rapidly progressive glomerulonephritis (RPGN), which is defined as a ≥ 50% decline in eGFR within 3 months. For each patient, the time of progression was recorded as the date of the first confirmed diagnosis of stage 5 CKD (eGFR < 15 mL/min/1.73 m²) based on the ICD-10-CM discharge codes (N18.804–N18.907) in the EHRs.
Data pre-processing
Data on candidate variables, including laboratory values and ICD-10 billing codes, were obtained from EHR. The patient ID number was used as a unique identifier. Data were cleaned and formatted before establishing the model. Categorical variables were transformed into binary dummy variables. All variables were normalized to reduce the dimension-introduced bias using the Z-score standardization method10. The equation used was as follows
where xi is the ith data of the dataset, µ is the average of the feature, and σ stands for the standard deviation.
Model development and evaluation
As the study endpoint was operationalized as a binary outcome within a fixed 25-week window, the predictive task was formulated as a classification problem. The 25-week endpoint was pre-specified at the protocol stage and held fixed throughout model development to avoid post-hoc outcome redefinition (see “Outcome definition”). Nine ML algorithms were applied to estimate the short-term risk of CKD progression, including regression-based models (logistic regression [LR], Elastic Net, Ridge regression, and LASSO), tree-based models (random forest [RF] and eXtreme Gradient Boosting [XGBoost]), and additional non-linear classifiers (support vector machine [SVM], neural network [NN], and k-nearest neighbors [k-NN]).
Within the development cohort, the dataset was randomly split into training (80%) and independent testing (20%) sets. Model training and hyperparameter tuning were conducted exclusively within the training set using a 10-fold cross-validation (CV) framework to reduce overfitting and improve generalizability. In each iteration, nine folds were used for training and onefold for validation, and results were averaged across iterations. As each patient contributed only one baseline record, partitioning was inherently performed at the patient level, preventing data leakage across folds.
Hyperparameter tuning was performed using grid search with 10-fold CV in the training set. For each candidate parameter configuration, the mean area under the curve (AUC) across folds was calculated, and the configuration with the highest average AUC was selected. The final model with tuned parameters was then retrained on the full training set and subsequently evaluated on the independent test set to assess out-of-sample performance.
To enhance interpretability, Tree SHapley Additive exPlanations (SHAP) were applied to the optimal model. Local Shapley values were aggregated as mean absolute SHAP to obtain global feature importance. SHAP values quantify the contribution of each feature to the prediction, allowing identification of the variables with the greatest impact on the model’s predictions.
Model performance assessment
Model performance was assessed using AUC, sensitivity, specificity, accuracy, log loss, false positive (FP) rate, precision, F1, and average precision (AP), as recommended by clinical prediction model guidelines. An AUC of > 0.9, 0.7–0.9, ≤ 0.7 indicated good, moderate, and poor performance, respectively11,12. Paired t tests were used to compare AUC values in internal validation. Within internal CV, model AUCs were compared using paired t tests across folds. On the external validation cohort, differences in AUC were tested using DeLong’s nonparametric test for correlated receiver operating characteristic (ROC) curves (two-sided α = 0.05). Calibration was evaluated using the Brier score and the Expected Calibration Error (ECE), with lower values indicating better calibration. Calibration curves were plotted by grouping predictions into five equally spaced probability bins and comparing observed versus predicted risk against a 45° reference line.
Variable importance was assessed in each ML model. The relative importance was quantified by assigning a weight between 0 and 1 to each variable in the model. XGBoost and RF directly derived the variable importance during model training. The coefficients of the Elastic Net, LASSO, and Ridge models were used as measures of importance. The mean decrease accuracy method was used for the k-NN and SVM models by directly measuring the impact of each feature on model accuracy. Python (version 3.12.3) was used for model establishment and evaluation, respectively.
Statistical analysis
Categorical variables were expressed as percentages, whereas continuous measures were expressed as mean ± standard deviation for normally distributed variables or median (interquartile range; IQR) for non-normally distributed variables. Student’s t test was used for normally distributed continuous variables. Kruskal–Wallis test was used for non-normally distributed continuous variables, and χ2 for categorical variables. Statistical significance was set at two-sided P < 0.05.
Results
Baseline characteristics
The baseline demographic and clinical characteristics of the study population are summarized in Table 1. The development and validation cohorts consisted of 346 and 105 patients, respectively. Patients in both cohorts were on average middle-aged, with a broadly balanced distribution of males and females. At baseline, indicators of kidney function impairment were consistent with stage 4 CKD. The prevalence of short-term progression from stage 4 to stage 5 CKD was similar between the two cohorts.
Hyperparameter tuning
The average AUCs of the different ML models with tuned hyperparameters are as follows: Five models exhibited a single tuning parameter (LR, LASSO, Ridge, RF, and k-NN), whereas four exhibited two tuning parameters (ElasticNet, SVM, NN, and XGBoost) (Fig. 1). For LR, LASSO, and Ridge, the optimal regularization parameter alpha values were 1.0, 0.1, and 0.9, respectively. The RF model performed best with 70 randomly selected subtrees, whereas k-NN was optimal with k = 19. For models with two parameters, the ideal values were alpha = 0.1 and l1_ration = 0.4 for Elastic-Net, cost C = 3 and gamma = 0.001 for SVM, and a hidden layer size of 15 with alpha = 0.1 for the NN. XGBoost achieved the best results with a maximum depth of 6 and a minimum child weight of 4. The maximum ROC averages for each model with the corresponding parameters are listed (Table 2).
Tuning parameters for each model with re-sampling method.
Internal validation of ML models
The average ROC curves of the ML models on the test set using a 10-fold sample resampling procedure are presented in Fig. 2A and B. Most models showed moderate accuracy, with an average AUC of > 0.7. The optimized SVM model exhibited optimal performance, with an AUC of 0.86. Models are ranked in the box plot from top to bottom based on their mean AUC values.
Evaluations of the ML models on the test set. A Shows the average ROC curves of the ML models on the test set, using a 10-fold sample resampling procedure. B Shows the box plot of the mean AUC values with standard deviations of different prognostic models. The purple triangle in the box represents the mean values.
The AP was used to assess the tradeoff between precision and recall. A bar graph of the mean AP (mAP) for each model is shown in Fig. 3, with detailed values listed in Table 3. The mAP values of the LR, RF, and SVM models were > 0.8.
The bar graph of mean AP for each ML model.
We calculated the confusion matrix for the nine ML models. The LR and k-NN models exhibited the highest number of mispredictions, including false negatives (FNs) and false positives (FPs), with 23 errors each (Table 4). Among these, k-NN produced the largest number of underdiagnoses (FNs = 19), whereas LR generated the most misdiagnoses (FP = 12). Although the other models exhibited fewer FNs and FPs, RF showed good performance with only one FN and one FP.
Table 5 summarizes the performance of various ML models in the internal validation cohort. Among the models, Random Forest (RF) demonstrated the highest AUC (0.98), followed by Neural Network (NN) with an AUC of 0.95 and XGBoost with an AUC of 0.93. These models also achieved high accuracy and F1 scores, indicating strong discriminative ability. Specifically, XGBoost yielded an accuracy of 0.90, a sensitivity of 0.85, a specificity of 0.94, and an F1 score of 0.89, along with a relatively low log-loss of 0.34, suggesting a favorable balance between performance and calibration. While RF achieved slightly higher AUC and F1, its false positive rate (0.24) and precision (0.97) were comparable to XGBoost. Traditional models such as LR (AUC = 0.69) and k-NN (AUC = 0.80) performed less favorably across most metrics. Overall, XGBoost, RF, and NN were the top-performing models in the internal validation phase.
The mean AUC values of each model were compared using t-tests (Fig. 4). k-NN showed significant differences from RF, SVM, and XGBoost. However, no statistically significant differences were observed in the predictive power among LR, Elastic Net, Lasso, and XGBoost, indicating that these models performed similarly.
Comparison between mean AUC values. The differences of average AUC values between models are reflected by the number and color (red is big and blue is small) of the cube, respectively. *p < 0.05, ***p < 0.001.
External validation of ML models
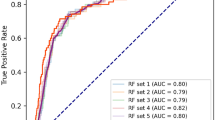
To validate the ML models developed in this study, clinical data from patients at Harbin 242 Hospital of Genertec Medical were collected for the external validation set. Table 6 presents the performance of the ML models in the external validation cohort. Compared to the internal validation, most models exhibited a decline in predictive performance, highlighting the challenge of model generalizability. In the external validation cohort, XGBoost achieved the best overall performance, with an AUC of 0.85 and the ROC curve closest to the ideal top-left corner (Fig. 5). It maintained 70% sensitivity at a 10% false positive rate, demonstrating strong discrimination and clinical applicability. Random Forest (AUC = 0.76) and Neural Network (AUC = 0.74) also performed well but showed notable declines from their internal validation results.
Linear models exhibited limited generalizability. LR achieved an AUC of 0.75, outperforming Lasso and Ridge, which both dropped to 0.56. SVM showed extreme imbalance, with high specificity (0.96) but very low sensitivity (0.21), while k-NN displayed poor specificity (0.27) despite high sensitivity.
Overall, the rightward shift of most ROC curves compared to internal validation highlights the challenge of domain shift. XGBoost remained the most robust and balanced model across metrics, supporting its selection as the optimal approach.
ROC curves of the ML models for identifying short-term progression in patients with stage 4 CKD on the external validation set.
Pairwise AUC comparisons (DeLong test)
Pairwise DeLong tests showed that XGBoost had significantly higher AUCs than k-NN, RF, LASSO, Ridge, and SVM (all p < 0.05), whereas differences versus LR, NN, and Elastic Net were not significant (p ≥ 0.05) (Table 6; Supplementary Table S1).
Calibration of predicted probabilities
In the external validation cohort, probability calibration varied across models (Fig. 6; Table 7). XGBoost showed the best overall calibration (Brier = 0.12; ECE = 0.03), with a reliability curve closely tracking the 45° reference line across risk bins. Random Forest achieved a low ECE (0.03) but a comparatively high Brier score (0.26), suggesting limited sharpness despite small average bin-wise deviations. Linear models (LR, Elastic Net, LASSO, Ridge) had higher Brier scores (0.24–0.27) and larger ECEs (0.08), with visible over-/underestimation at probability extremes. k-NN yielded a near-zero ECE (0.00) but a higher Brier score (0.25), with a flat reliability curve indicating shrinkage toward the cohort mean. The neural network showed intermediate calibration (Brier = 0.24; ECE = 0.06). Collectively, these findings indicate that XGBoost provided the most reliable risk estimates for clinical use in our setting.
External calibration of predicted risk across models.
Feature importance interpretation (XGBoost SHAP Summary)
Figure 7 displays the SHAP summary plot for the XGBoost model. The most influential features included ApoAI, Alb, CysC, HDL, and ApoB. Lower levels of ApoAI, Alb, and HDL were associated with higher predicted risk, while elevated CysC and ApoB also contributed positively. Other key predictors such as TC, BUN, neutrophils, FGB, and AG reflected the combined effects of renal dysfunction, lipid abnormalities, and inflammation. These patterns support the clinical interpretability and biological plausibility of the model’s predictions.
SHAP summary plot of top 10 features in the XGBoost model. X-axis represented the impact of the feature value on the model output, with positive values increasing the stage 4 CKD short-term progression, and negative values decreasing the progress. Y-axis listed the top 10 contributing features. Color gradient showed the feature value, with blue indicating lower values and pink indicating higher values.
Discussion
In this multicenter retrospective study, we developed and externally validated nine ML models to predict the short-term risk of progression to ESRD in patients with stage 4 CKD using real-world data. Among them, the XGBoost model demonstrated strong performance, with AUC values of 0.93 and 0.85 for the training and external validation datasets, respectively, indicating its potential as a robust tool for early risk stratification. Key features driving short-term progression included decreased levels of ApoAI, albumin, and HDL, along with elevated Cystatin C, ApoB, and inflammatory markers.
Model performance
ML methods are powerful tools for predicting the occurrence and progression of CKD. A major strength of ML approaches lies in their proficiency in processing complex, multidimensional information and uncovering nonlinear connections between different variables. Previous studies comparing the fitness of ML methods in predicting ESRD in patients with stage 4 CKD have reported promising results. For example, Takkavatakarn et al. used ML algorithms to develop a prediction model for ESRD in patients with advanced CKD over a 3-year follow-up period13. Furthermore, Xiao et al. evaluated CKD progression using urine protein employing different ML models14. Our study further confirms that ML methods offer more accurate predictions. Notably, the XGBoost model demonstrated superior predictive performance, particularly in external validation, highlighting its strong generalizability and robustness. As a gradient boosting ensemble, XGBoost effectively models complex interactions, incorporates regularization to reduce overfitting, and is robust to noise and redundant features common in clinical data. These strengths align with prior findings on its medical predictive power. Our findings align with previous studies reporting the superiority of XGBoost in medical prediction tasks. Its excellent performance not only makes it well-suited for predicting short-term progression risk in CKD patients but also provides valuable insights for developing predictive models in other complex diseases.
In contrast, k-NN and LR performed poorly. k-NN’s reliance on distance calculations makes it sensitive to irrelevant or noisy features, reducing accuracy in high-dimensional settings. LR’s assumption of linearity limits its ability to capture complex relationships, and its performance declines with many features and multicollinearity. These limitations highlight the superiority of advanced nonlinear ML methods for predicting CKD progression.
Variables
Clinicians and researchers have emphasized identifying critical factors that may prevent the progression of CKD to ESRD. However, the risk factors for CKD deterioration are often extremely complex. Our models relied on laboratory markers to predict the risk of short-term ESRD progression in patients with stage 4 CKD. In this study, SHAP was used to visually interpret the XGBoost model.
Our SHAP-based interpretation of the XGBoost model revealed several key biological predictors of ESRD progression, reinforcing established pathogenic mechanisms in advanced CKD. Notably, decreased levels of apolipoprotein A-I (ApoA-I), albumin (Alb), and high-density lipoprotein (HDL) indicate poor nutritional and lipid profiles, consistent with the malnutrition–inflammation–atherosclerosis (MIA) syndrome. ApoA-I and HDL are central to reverse cholesterol transport and possess anti-inflammatory properties; their reduction suggests impaired lipid clearance and increased oxidative stress15,16. Serum Alb, a marker of protein-energy wasting and inflammation, has long been associated with poor prognosis in CKD17. Elevated Cystatin C (CysC) was another strong predictor, indicating glomerular filtration decline that may be underestimated by creatinine alone18.
In addition, increased apolipoprotein B (ApoB) and alpha-1-acid glycoprotein (AG) levels highlight the pro-atherogenic and inflammatory milieu in CKD. ApoB is a marker of atherogenic lipoproteins and has been independently linked to ESRD risk19,20, while AG, an acute-phase reactant, reflects systemic inflammation that may accelerate glomerulosclerosis.
Together, these findings underscore the intertwined mechanisms of malnutrition, dyslipidemia, and inflammation in the short-term progression to ESRD, and suggest that early interventions targeting these pathways may be clinically beneficial.
Clinical implications
The proposed model has important implications for clinical practice, research, and public health policy. To the best of our knowledge, most previous ML prediction models for CKD progression have been established in patients with moderate CKD21, DKD22, or those with ESRD23,24. This study developed and externally validated an ML model in patients with CKD stage 4, a critical group concerned with slowing CKD progression and preparing for renal replacement therapy. Previously, Grams et al.25 developed a CKD G4 risk calculator to predict the probability of KRT, death, and CVD events at 2 and 4 years. Moreover, Kullaya et al.13 focused on ESRD prediction in CKD stage 4 over 3 years, whereas the The European QUALity (EQUAL) study involved the follow-up of older patients with stage 4 CKD for 4 years26. However, the progression rate varies widely in CKD stage 4 due to patient heterogeneity and complications, with many patients progressing rapidly to renal failure in < 1 year or even 6 months27,28,29. Therefore, suitable prediction models are required for high-risk patients experiencing short-term progression.
Our model addresses this gap by identifying patients at high risk of ESRD progression within 6 months, enabling nurses to promptly collaborate with physicians in adjusting follow-up frequencies, optimizing medication guidance, and initiating pre-dialysis preparations30, thereby mitigating delays in care. Moreover, the prediction tool facilitates information sharing among interprofessional teams, including nurses, physicians, dietitians, and rehabilitation therapists, thus promoting cohesive multidisciplinary collaboration. In terms of patients, this proactive approach may also aid in managing patients’ physical health while enhancing their psychological well-being by reducing anxiety and confusion. On the other hand, with no extra clinical tests or treatments, medical costs for patients and the burden on the healthcare system can be alleviated.
Both the KDIGO and NICE emphasize the necessity of risk assessment in patients with CKD, and powerful tools such as the KFRE31, KPNW32, and Z6 score33 models have been developed. Nevertheless, most studies have focused on patients with stage 3 ~ 5 CKD, leaving the prognosis of CKD stage 4 less understood.
A global meta-analysis of 28 cohorts reported a higher risk for kidney failure, CVD events, hospitalizations, and death among the patients with CKD stage 4 undergoing kidney replacement therapy34. Consequently, clinical decisions are often based on clinicians’ experience, which can lead to delays in treatment for those progressing to kidney failure or unnecessary interventions for those who do not. Early identification of patients at a higher risk of kidney failure can promote primary care-based treatment and improve the prognosis of advanced CKD.
Strengths and limitations
One notable strength of our study is the establishment and external validation of a highly precise prediction tool. This model is user-friendly and utilizes features that are routinely collected during clinical situations. It can also be easily connected to hospital information systems, making it a valuable tool for early identification and stratified interventions.
Nevertheless, several limitations should be acknowledged. First, the retrospective design and modest sample may introduce bias and limit generalizability (e.g., via overfitting); the COVID-19 pandemic-related attrition caused loss to follow-up. Therefore, future prospective cohort studies with larger sample sizes are warranted. Nevertheless, our analysis comprehensively incorporated all available patient data from both participating medical centers to ensure maximal representativeness under these challenging conditions. Second, although we applied regularization and 10-fold CV, external AUCs were lower and confidence intervals wider than internal estimates, which is expected under distributional shift and given the smaller external sample (n = 105); cautious deployment with local recalibration and larger prospective validation is warranted. Moreover, we used classification rather than survival models because the endpoint was protocol-defined as a binary 25-week outcome and long-term follow-up was incomplete. This design choice (appropriate for the present study) precluded explicit modeling of censoring and unequal follow-up. In line with the pre-specified endpoint, we did not assess alternative windows (e.g., 26 weeks); such choices could modestly shift performance and warrant prospective evaluation. Regarding interpretation, SHAP was used for global explanations by aggregating local values. Standard SHAP does not capture higher-order feature interactions, so interaction effects may be underrepresented, and its attributions are associational rather than causal. Third, the predictor scope was restricted by available data. Comorbidities, medications, lifestyle factors, and CKD etiology were not included. Etiology is a well-recognized determinant of CKD progression, with diabetic nephropathy generally progressing faster than IgA nephropathy or other glomerular diseases. However, reliable etiological information was often unavailable in our cohorts, as renal biopsy for IgA nephropathy was rarely performed and ICD-based coding was incomplete. To avoid misclassification bias, etiology was not incorporated. Future studies with standardized and comprehensive etiological data should include this factor to improve risk stratification and clinical applicability.
Conclusion
We developed and externally validated an ML-based risk prediction model for 25 weeks disease progression in a hospital-based cohort of adults with stage 4 CKD. We also demonstrated the feasibility of ML in assessing the prognosis of ESRD based on easily accessible clinical features. The XGBoost model demonstrated adequate performance, highlighting its potential use for future patient screening. Future studies should include the development of a web-based risk prediction tool for patients with stage 4 CKD to optimize risk identification in clinical practice.
Data sharing statement
The data that support the findings of this study are not publicly available due to containing information that could compromise the privacy of research participants but are available from the corresponding author [Y.F.] upon reasonable request.
References
Foreman, K. J. et al. Forecasting life expectancy, years of life lost, and all-cause and cause-specific mortality for 250 causes of death: reference and alternative scenarios for 2016–40 for 195 countries and territories. Lancet 392, 2052–2090. https://doi.org/10.1016/S0140-6736(18)31694-5 (2018).
Zhang, L. et al. Prevalence of chronic kidney disease in china: a cross-sectional survey. Lancet 379, 815–822. https://doi.org/10.1016/s0140-6736(12)60033-6 (2012).
Wang, L. et al. Prevalence of chronic kidney disease in china: results from the sixth China chronic disease and risk factor surveillance. JAMA Intern. Med. 183, 298–310. https://doi.org/10.1001/jamainternmed.2022.6817 (2023).
Jankowski, J., Floege, J., Fliser, D., Böhm, M. & Marx, N. Cardiovascular disease in chronic kidney disease: pathophysiological insights and therapeutic options. Circulation 143, 1157–1172. https://doi.org/10.1161/circulationaha.120.050686 (2021).
KCWG. KDIGO 2024 clinical practice guideline for the evaluation and management of chronic kidney disease. Kidney Int. 2024;105:S117–S314. https://doi.org/10.1016/j.kint.2023.10.018 (2024).
Piveteau, J. et al. Pre-dialysis care trajectory and post-dialysis survival and transplantation access in patients with end-stage kidney disease. J. Nephrol. 36, 2057–2070. https://doi.org/10.1007/s40620-023-01711-y (2023).
Krishnamurthy, S. et al. Machine learning prediction models for chronic kidney disease using National Health Insurance Claim Data in Taiwan. Healthcare (Basel) 9 (5), 546. https://doi.org/10.3390/healthcare9050546 (2021).
Aminnejad, N., Greiver, M. & Huang, H. Predicting the onset of chronic kidney disease (CKD) for diabetic patients with aggregated longitudinal EMR data. PLOS Digit. Health. 4 (1), e0000700. https://doi.org/10.1371/journal.pdig.0000700 (2025). PMID: 39841679; PMCID: PMC11753645.
Makino, M. et al. Artificial intelligence predicts the progression of diabetic kidney disease using big data machine learning. Sci. Rep. 9 (1), 11862. https://doi.org/10.1038/s41598-019-48263-5 (2019). PMID: 31413285; PMCID: PMC6694113.
Everitt, B. S. & Hothorn, T. An Introduction to Applied Multivariate Analysis with R (2011).
Steyerberg, E. W. & Vergouwe, Y. Towards better clinical prediction models: seven steps for development and an ABCD for validation. Eur. Heart J. 35, 1925–1931. https://doi.org/10.1093/eurheartj/ehu207 (2014).
Collins, G. S. et al. TRIPOD + AI statement: updated guidance for reporting clinical prediction models that use regression or machine learning methods. Bmj 385, e078378. https://doi.org/10.1136/bmj-2023-078378 (2024).
Takkavatakarn, K., Oh, W., Cheng, E., Nadkarni, G. N. & Chan, L. Machine learning models to predict end-stage kidney disease in chronic kidney disease stage 4. BMC Nephrol. 24, 376. https://doi.org/10.1186/s12882-023-03424-7 (2023).
Xiao, J. et al. Comparison and development of machine learning tools in the prediction of chronic kidney disease progression. J. Transl. Med. 17, 119. https://doi.org/10.1186/s12967-019-1860-0 (2019).
Vlad, C. et al. Apolipoproteins A and B and PCSK9: nontraditional cardiovascular risk factors in chronic kidney disease and in End-Stage renal disease. J. Diabetes Res. 2019 https://doi.org/10.1155/2019/6906278 (2019).
Kimak, E. & Solski, J. ApoA- and apoB-containing lipoproteins and Lp(a) concentration in non-dialyzed patients with chronic renal failure. Ren. Fail. 24 (4), 485–492. https://doi.org/10.1081/jdi-120006775 (2002).
Kaysen, G. A. et al. Relationships among inflammation nutrition and physiologic mechanisms establishing albumin levels in hemodialysis patients. Kidney Int. 61, 2240–2249. https://doi.org/10.1046/j.1523-1755.2002.00076.x (2002).
Koye, D. N. et al. Risk of progression of nonalbuminuric CKD to end-stage kidney disease in people with diabetes: the CRIC (Chronic renal insufficiency Cohort) study. Am. J. Kidney Dis. 72, 653–661. https://doi.org/10.1053/j.ajkd.2018.02.364 (2018).
Kwon, S. et al. Apolipoprotein B is a risk factor for end-stage renal disease. Clin. Kidney J. 14, 617–623. https://doi.org/10.1093/ckj/sfz186 (2020).
Zhao, W. B. & Alberto, P. S. M. Serum apolipoprotein B/apolipoprotein A1 ratio is associated with the progression of diabetic kidney disease to renal replacement therapy. Int. Urol. Nephrol. 52 (10), 1923–1928. https://doi.org/10.1007/s11255-020-02550-7 (2020). Epub 2020 Jul 13. PMID: 32661625.
Bai, Q., Su, C., Tang, W. & Li, Y. Machine learning to predict end stage kidney disease in chronic kidney disease. Sci. Rep. 12, 8377. https://doi.org/10.1038/s41598-022-12316-z (2022).
Zou, Y. et al. Development and internal validation of machine learning algorithms for end-stage renal disease risk prediction model of people with type 2 diabetes mellitus and diabetic kidney disease. Ren. Fail. 44, 562–570. https://doi.org/10.1080/0886022x.2022.2056053 (2022).
Garcia-Montemayor, V. et al. Predicting mortality in hemodialysis patients using machine learning analysis. Clin. Kidney J. 14, 1388–1395. https://doi.org/10.1093/ckj/sfaa126 (2021).
Rankin, S. et al. A machine learning model for predicting mortality within 90 days of Dialysis initiation. Kidney 360 (3), 1556–1565. https://doi.org/10.34067/kid.0007012021 (2022).
Grams, M. E. et al. Predicting timing of clinical outcomes in patients with chronic kidney disease and severely decreased glomerular filtration rate. Kidney Int. 93, 1442–1451. https://doi.org/10.1016/j.kint.2018.01.009 (2018).
Jager, K. J. et al. The EQUAL study: a European study in chronic kidney disease stage 4 patients. Nephrol. Dial Transpl. 27 (Suppl 3), iii27–31. https://doi.org/10.1093/ndt/gfs277 (2012).
Rao, A. et al. Using datasets to ascertain the generalizability of clinical cohorts: the example of the European quality study on the treatment of advanced chronic kidney disease. Nephrol. Dial Transpl. 37, 540–547. https://doi.org/10.1093/ndt/gfab002 (2022).
Lee, M. J. et al. Can we predict when to start renal replacement therapy in patients with chronic kidney disease using 6 months of clinical data? PLoS One. 13, e0204586. https://doi.org/10.1371/journal.pone.0204586 (2018).
Cheng, L. C., Hu, Y. H. & Chiou, S. H. Applying the temporal abstraction technique to the prediction of chronic kidney disease progression. J. Med. Syst. 41, 85. https://doi.org/10.1007/s10916-017-0732-5 (2017).
Eckardt, K. U. et al. Improving the prognosis of patients with severely decreased glomerular filtration rate (CKD G4+): conclusions from a Kidney Disease: Improving Global Outcomes (KDIGO) Controversies Conference. Kidney Int. 93, 1281–1292. https://doi.org/10.1016/j.kint.2018.02.006 (2018).
Tangri, N. et al. A predictive model for progression of chronic kidney disease to kidney failure. Jama 305, 1553–1559. https://doi.org/10.1001/jama.2011.451 (2011).
Schroeder, E. B. et al. Predicting 5-Year risk of RRT in stage 3 or 4 CKD: development and external validation. Clin. J. Am. Soc. Nephrol. 12, 87–94. https://doi.org/10.2215/cjn.01290216 (2017).
Zacharias, H. U. et al. A predictive model for progression of CKD to kidney failure based on routine laboratory tests. Am. J. Kidney Dis. 79, 217–230. https://doi.org/10.1053/j.ajkd.2021.05.018 (2022).
Evans, M. et al. Risk factors for prognosis in patients with severely decreased GFR. Kidney Int. Rep. 3, 625–637. https://doi.org/10.1016/j.ekir.2018.01.002 (2018).
Acknowledgements
We would like to thank Editage’s (www.editage.cn) English language editing.
Funding
The author(s) declare financial support was received for the research, authorship, and/or publication of this article. This study received support from the Basic Scientific Research business expenses of provincial colleges and universities in Heilongjiang Province [grant number not yet assigned] and the Key Research and Development Program of Heilongjiang Province [grant number 2024ZX12C30].
Author information
Authors and Affiliations
Contributions
JL: Writing—review & editing, Writing—original draft, Visualization, Validation, Supervision, Methodology, Investigation, Data curation. Funding acquisition. XD: Project administration, Supervision, Methodology. RZ: Project administration, Supervision, Investigation. XL: Writing—review & editing, Validation, Software. JX: Validation, Statistics, Software, Visualization. XS: Writing—review & editing, Data Resources, Data curation. YZ: Writing—review & editing, Data curation. LL: Writing—review & editing, Data Resources, Data curation. GX: Writing—review & editing, Data Resources, Data curation. YF: Writing—review & editing, Supervision, Project administration, Investigation.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, J., Du, X., Zhang, R. et al. Machine learning models for predicting short-term progression in patients with stage 4 chronic kidney disease: a multi-center validation study. Sci Rep 15, 39285 (2025). https://doi.org/10.1038/s41598-025-23037-4
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-23037-4