Abstract
Prostaglandin receptors are pharmacologically validated targets with implications in several medical indications, including glaucoma, cardiac cyanotic disease, pulmonary hypertension, oncology, and various rare diseases. In this study, we developed a ligand-based machine learning (ML) model to classify chemical compounds as either active or inactive against the prostaglandin receptor EP2. From an initial set of 1,826 descriptors, 20 were selected to train random forest algorithms, yielding an area under the curve score (AUC) of > 0.8 for compound classification in the test set. Our resulting ML classifier showed an overall accuracy of 88.9% towards newly experimentally tested EP2 ligands. This adaptable and tractable workflow can be extended to other EP receptors and possibly other similar targets.
Similar content being viewed by others
Introduction
Prostaglandins exhibit remarkable chemical diversity owing to their unique structural features and multiple isomeric forms. They are lipid compounds derived from arachidonic acid, a polyunsaturated fatty acid, through the enzymatic action of cyclooxygenases (COX)1. The classification of prostaglandins is based on their ring structure scaffold, with different series characterized by variations in side chain ring modifications. Additionally, within each series, specific prostaglandins can differ in the position and configuration of double bonds, hydroxyl groups, and functional substitutions, leading to a vast array of distinct molecules. The chemical diversity of prostaglandins contributes to their wide-ranging physiological role and underlines their importance as signaling molecules in various biological processes2.
The EP2 receptor is one of the four subtypes of prostaglandin E receptors, distinctly responsive to prostaglandin E2 (PGE2). Functionally, EP2 signals primarily via the activation of adenylate cyclase, leading to an increase in cyclic AMP levels within the cell. This intracellular signaling plays pivotal roles in mediating anti-inflammatory effects, vasodilation, and protection of the gastric mucosa3. Due to its central role in such a broad spectrum of physiological and pathological processes, the EP2 receptor presents as a promising target for therapeutic drug development and intervention in various pathologies, including cancer progression, neurodegenerative diseases, inflammatory conditions, and rare diseases4,5. While PGE2 is the natural endogenous ligand for EP2, synthetic agonists and antagonists have been developed for research and clinical purposes6.
Machine learning (ML) has emerged as a powerful tool in the field of ligand-based drug discovery, revolutionizing the process of identifying potential drug candidates7. Ligand-based drug discovery focuses on the interaction between small molecules (ligands) and their target receptors or proteins. With an abundance of available molecular data, machine learning algorithms can efficiently analyze and interpret these complex datasets to predict ligand-receptor interactions, identify bioactive compounds, and prioritize promising candidates for further experimental validation. Various machine learning approaches, such as support vector machines8, random forests9, and deep learning models through neural network architectures like Multi-Layer Perceptron10, have been employed to extract meaningful patterns from chemical structures. By leveraging these predictive ML models, researchers can significantly accelerate the drug discovery process, optimize lead compounds, and ultimately improve the chances of identifying novel, safe, and effective therapeutic agents for a wide range of diseases. However, challenges still exist, such as data imbalance and the need for robust validation strategies, which necessitate continuous refinement and development of machine learning methodologies in the pursuit of more efficient and reliable ligand-based drug discovery approaches.
In this study, following preliminary evaluations of various machine learning algorithms and architectures, we developed an efficient ML classifier to predict compounds that elicit EP2 receptor activity, agonists in particular. It exhibited robust predictive performance, effectively identifying active compounds, including those with experimentally validated activity. This tool aids in the design and prioritization of novel EP2-targeting compounds by integrating diverse data sources and optimizing ML model parameters. Our strategy of combining relatively simple ML architectures, trained on peer-reviewed data, to identify patent-free compounds amenable for repurposing and new chemical entities could be used to support other ligand-based drug discovery initiatives for other GPCRs targets.
Results and discussion
Compilation of datasets
To serve as the basis for the category classification (active or inactive), we curated a collection of compounds sourced from literature, patents, and experimental data, focusing on their activity toward the EP2 receptor (DATASET1); out of these, 757 compounds had experimentally verified activity as either agonists or antagonists.
The sources reported results from two main types of experimental assays: (1) binding assays, which measure a compound’s ability to displace radiolabeled ligands (e.g., [3H]PGE2) from EP2 receptors, typically reported as \(K_i\) values; and (2) functional assays, which assess changes in intracellular cAMP levels in EP2-expressing cells, with \(EC_{50}\) values indicating agonist activity and \(IC_{50}\) values indicating antagonist activity based on inhibition of a known agonist. When multiple measurements existed for a single compound, we calculated the arithmetic mean of the log-transformed values (e.g., \(pEC_{50}\), \(pIC_{50}\), or \(pK_i\)) to obtain a unified pEP2 score (see Methods). Of note, as most compounds have one to two measurements, using the median instead of the mean for data aggregation would not have led to significant changes in the final pEP2 values and in model’s performance. This approach allowed us to assign agonist or antagonist labels based on experimentally reported functional activity.
One of the key aspects of prostaglandin ligand discovery is the choice of chemical scaffold: most EP2 natural agonists have a cyclopentane prostanoid scaffold (e.g., PGE2), whereas non-prostanoid clinically approved (Taprenepag isopropyl / PF-04217329, Pfizer11; JV-GL1, Jenivision12) or market-approved (Omidenepag isopropyl / OMLONTI®, Santen13) EP2 agonists include sulfonamide or amide scaffolds. Our DATASET1 showed a balanced structural distribution among these scaffolds (Fig. 1a, b, c and Table 1). Also, chemical scaffolds were found approximately equally distributed in both EP2 agonists and antagonists, indicating that the structural frameworks are compatible with either activating or blocking the receptor function; as expected given its natural agonist function, the cyclopentane scaffold was almost exclusively associated with agonist activity (Supplementary Fig. S1).
An important point to note is that our training and test DATASET1 contained both agonists and antagonists; this choice was motivated by the need for sufficiently large training and testing sets and by the substantial scaffold overlap between agonists and antagonists (see Supplementary Figures S1 and S2a plus S2b).
A total of four datasets were compiled in this study: (1) the aforementioned DATASET1, (2) a dataset of structurally related decoys (DATASET2), (2) a compilation of EP2 ligands, and referenced as active, from patent and public databases (DATASET3), and (4) a collection of patent-protection-free and novel chemical entities (NCE) EP2 ligands, for which experimental agonist activity was determined (DATASET4) (Fig. 2, Supplementary Fig. S3 and Table S1). These last three datasets are discussed in more detail further below.
Descriptors generation and selection
We computed a total of 1,826 molecular descriptors using Mordred14 that provided quantitative (both 2D and 3D) representations of the molecular structures in the compounds in DATASET1. Also, we computed their mutual information (MI) values, which quantify the amount of information shared between each descriptor and the target variable (EP2 activity in our case). This first set of descriptors was further narrowed down following the workflow illustrated in Fig. 3a . As shown in Fig. 3b , MI-ranked values varied depending on the activity cutoff. As expected, low (pEP2>3) and high (pEP2>8) cutoffs yielded poor MI values, as in these cases most of the compounds were mainly labeled either active or inactive. Activity cutoffs of pEP2> 6 and pEP2> 7 yielded the highest MI scores, as observed in the activity distribution in DATASET1. Finally, we chose pEP2 > 6 as a cutoff, as it aligned with the median value (See the Supplementary Fig. S2 a, b and c) and since it is recognized as the threshold where compounds with 1 \(\upmu\)M activity are considered bona fide hits.
The feature set was narrowed down to the most informative descriptors (Supplementary Table S2) to balance between ML classifier performance and interpretability. Descriptors with low variance and those exhibiting low correlation relative to higher-ranked features were excluded.
Machine learning algorithms generation and selection
We performed preliminary experiments with various ML algorithms (e.g., logistic regression, K-Nearest Neighbor, Gradient Boosting, Support Vector Machine) and some Deep Learning architectures (eg., Multi-Layer Perceptron), all with different molecular representations (morgan and rdkit fingerprints, Mordred descriptors14, ChemBERTa15 pretrained model’s output).
The results of these preliminary tests led to similar results in both ROC AUC and Average Precision (AP), above 0.9 and 0.7, respectively, as presented in Supplementary Fig. S4 and detailed in Supplementary Table S3. While the deep learning Multi-Layer Perceptron (MLP) model seems to underperform relative to the other ML algorithms, with a margin of up to 0.2 in both ROC AUC and AP (primarily when using Mordred descriptors and ChemBERTa as features), we reasoned that this is likely due to suboptimal MLP model architecture. With further optimization, it would achieve comparable performance as the others.
Since all the tested ML algorithms demonstrated similar performance, we chose the Random Forest Classifier (RFC) due to its efficacy and ease of interpretability; RFC is also the only one among those tested that facilitates a feature selection process that we considered key for our general approach. In short, RFC builds an ensemble of Decision Trees, each trained on a random subset of the data. Each tree votes for the class (inactive or active), and the mean vote determines the probability of a compound being classified as active. If this probability exceeds a threshold, the compound is labeled as “active.” Decision Trees split data by selecting features that maximize class purity at each node9.
Once the RFC algorithm was selected, the DATASET1 was split into training and validation sets, with 80% of the data used for training and 20% for validation. Multiple RFC models were generated using various numbers of input features (descriptors), and hyperparameter tuning was carried out using 5-fold cross-validation within the training set to minimize overfitting and optimize model’s performance. This process iteratively adjusted their parameters to ensure the best configuration for predictive accuracy (Fig. 2, Table 2).
Assessment of model predictive performance
The predictive performance of the machine learning classifiers was evaluated using the Area Under the Receiver Operating Characteristic Curve (ROC AUC) and Average Precision (AP) as primary metrics. These measures provide insights into the ML classifiers’ ability to discriminate between active and inactive compounds. Each of them was trained using a various number of features (descriptors) as input to evaluate their impact on the ML classifiers’ prediction power.
In both metrics, RFC models utilizing relatively few descriptors performed well (Supplementary Table S2), reaching an ROC AUC and AP of 0.9 with only 3 descriptors (Fig. 3c and d; Table 3). To rule out the possibility of an overfitting, DATASET2 was generated as a decoy dataset of compounds not active against EP2; decoys were confirmed to be structurally related to other datasets using a PCA analysis that showed that most compounds (for all datasets) map to the same PCA clusters (Supplementary Fig. S5). As expected, the introduction of DATASET2 reduced the ROC AUC and significantly dropped the AP (Fig. 3c and d). The decoy set was integrated as part of the validation set to select the ML classifiers that distinguish between active and inactive compounds.
As presented in Fig. 3c and d, with the introduction of decoys (validation + DATASET2), the addition of new descriptors in the ML classifier’s input does increase its performance by an important amount up to 20 descriptors, where it starts to reach a plateau. After that the gain of performance slows down, by gaining only marginal information in comparison to the addition of complexity (gain of 0.07 in AP between 10 and 20 descriptors against a gain of 0.014 between 20 and 30 descriptors). Hence why we decided to continue with the RFC model using the 20 best descriptors for inference and experimental analysis, as it was considered a good balance between predictability and simplicity. Details of the performance of this final model can be found in Supplementary Table S3.
Model inference on public datasets
We created a dataset (DATASET3) with approximately 85,000 compounds that gathered EP2-ligands reported from public patents (WIPO, Patentscope, SureChEMBL16) and pharmacological databases (IUPHAR/BPS17, GoStar18, and ChEMBL19). The overarching goal of this DATASET3 was to identify compounds amenable for redevelopment (with an expired patent protection). To prepare those compounds for inference, their 20 descriptors used by the selected RFC model have been generated from their canonical SMILES. The compounds for which the descriptors computation failed have been removed, leading to a final number of 84,863 molecules to perform inference on.
As the patent dataset does not contain experimental activity data, we compared the number of compounds identified as active between DATASET3 and the decoys DATASET2, using the final RFC model obtained (utilizing the top 20 descriptors) and three activity thresholds: pEP2 \(\ge\)0.5, \(\ge\)0.7 and \(\ge\)0.85. A threshold of 0.5 can be seen as the default assumption to distinguish positives and negatives. It may however, not lead to the optimal performance depending on the ML model. A threshold of \(\ge\)0.7 corresponds to that in which most of the active compounds tested in-house are recovered while avoiding at maximum the false positives (DATASET4, see below), and \(\ge\)0.815 is the highest score obtained by a compound from DATASET2 (decoys). The results are compiled in the Supplementary Table S4.
At a probability score threshold of \(\ge\)0.5, a significant 29.89% of DATASET3 compounds were identified as active, compared to 13.82% from DATASET2. Progressively refining the threshold to \(\ge\)0.7, and \(\ge\)0.815 markedly decreased the incidence of false positives, highlighting its precision, with only 0.45% of DATASET2 (decoys) compounds meeting this most stringent threshold. These results indicate a capacity of the RFC model to minimize false positives while effectively identifying the most viable candidates from DATASET3.
Model inference on experimental data
To evaluate our RFC model against experimentally validated in-house data, we created DATASET4. DATASET4 consisted of 27 compounds selected from three complementary sources:(i) 6 representative molecules from DATASET1, included as experimental benchmarks; (ii) a subset of 8 compounds from DATASET3, prioritized for their patent-free status (repurposing potential), high EP2 selectivity relative to other EP receptors, and predicted potency; and (iii) 13 new chemical entities (NCEs) designed by our medicinal chemistry team.
The corresponding chemical entities of DATASET4 were either aggregated from different vendors or synthesized de novo (structures not shown). The EP2-agonist activity was then assessed using Homogeneous Time-Resolved Fluorescence (HTRF) by measuring the increase of the cytosolic pool of cAMP in CHO cells overexpressing the human EP2 receptor20. Briefly, the cells expressing EP2 were plated and incubated with test compounds, then lysed for fluorescence measurements of cAMP levels. Fluorescence resonance energy transfer was used to determine cAMP concentration by the ratio of emissions at 665 nm to 620 nm, with results expressed as a percentage of the control response to PGE2. The EC50 for PGE2 was calculated from a dose-response curve in each experiment.
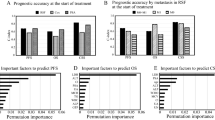
The compounds in DATASET4 exhibited variable EP2-agonist activity levels (Fig. 4a), with most of the active compounds including either sulfonamide- or amide scaffolds (Fig. 4c). By plotting experimental agonist activity vs. the predicted score value (Fig. 4b), and fixing the threshold at 0.70, our final RFC model was capable of detecting active compounds with a sensitivity of 83.3%, a specificity of 90.5% and an overall accuracy of 88.9%, as shown in the confusion matrix in Supplementary Fig. S6.
Final remarks and conclusions
A limitation of the present model is that it does not distinguish between agonists and antagonists; however, our objective was to predict EP2 binding capacity irrespective of functional outcome, and therefore both classes were combined during model development. This decision was driven by the need for sufficiently large training and testing sets, as well as by the significant scaffold overlap between agonists and antagonists. The resulting ligand-binding model was subsequently applied to libraries enriched in nominal EP2 agonists, thereby bypassing the need for an exclusively agonist-focused classifier.
One recent study reported predicting compound activity towards EP2 using machine learning21, which employed a regression-based model, focusing on agonists with small number of molecular descriptors as input features. In our work, we focused on global binding ability by aggregating agonist and antagonist activity data, given its shared chemical scaffold and potential pharmacology applications. Additionally, we focused on developing a ML classification rather than a regression model, proven to be efficient in virtual screening and reducing potential noise due to data aggregation from multiple sources.
Furthermore, the RFC model’s ability to predict novel EP2 binders, as well as its generalization to new compounds not included in the training set, demonstrates its versatility and reliability. This predictive RFC model provides a valuable tool for guiding further experimental validation, offering insights into the design and optimization of compounds targeting the EP2 receptor. It can be extended to explore additional receptor interactions or new chemical scaffolds, enhancing the scope of future drug development strategies.
In this study, we present a robust and adaptable machine learning model designed to predict the activity toward the EP2 receptor to contribute to ligand prioritization and drug discovery. By combining multiple datasets, optimizing model hyperparameters, feature curation, and various validation procedures, our RFC model achieved good predictive accuracy when applied to experimentally validate EP2 ligands. Furthermore, it provides a practical tool for prioritizing EP2 ligands with favorable freedom-to-operate (FTO) status, including both patent-free compounds and novel chemical entities for second-medical use or composition patent protection, respectively.
Methods
Datasets
DATASET1 was compiled from patents and non-patent literature (e.g., Markovič et al. 20166); the activity reported varied depending on the source and experimental methods; generally, this included binding assays (Ki) or the cellular response, either as an agonist activity (e.g. \(EC_{50}\)) or antagonist activity (e.g., \(IC_{50}\)). DATASET2 was compiled by generating 16,300 decoys based on DATASET1, using DUD\(\cdot\)E22 (Irwin and Shoichet Laboratories, UCSF. DUD\(\cdot\)E). DATASET3 (with approximately 85k compounds) was compiled from several databases:(1) A list of patent families was obtained by querying patent public databases (e.g., WIPO, Patentscope) using “EP2” OR “EP4” as keywords in either titles, abstracts, or claims; the molecules associated with these families were retrieved by querying the database SureChEMBL16. Of note, enumeration of Markush structures from patents was not included in the DATASET3 (2) Compounds annotated as ligands of (EP1 to EP4) were covered from IUPHAR/BPS17, GoStar18 and ChEMBL19. DATASET4 was compiled from compounds identified as bona fide repurposing molecules, or NCE, designed by Medetia Pharmaceuticals.
Activity threshold definition
A common comparable activity variable score is described by Eq. (1) below:
Here, the variable Activity refers to Ki, \(EC_{50}\), \(IC_{50}\), each expressed in M/L (Molar), or their arithmetic mean when the compound has multiple data points available. This formula implies that a compound with an activity around 1 \(\upmu\)M has a pEP2 value of 6, implying that the higher the pEP2 value, the more active the compound. This Activity Threshold refers to the cutoff value for pEP2, above which the molecules are considered active and below which they are labeled as inactive.
Data computations
All data manipulation, analysis, and modeling were primarily conducted using Python (Python Software Foundation, version 3.9.15). The NumPy package23, developed by the consensus of the NumPy and wider scientific Python community (NumPy project, version 1.23.5), was used for numerical computations. For data handling, we employed pandas, maintained by NumFOCUS, Inc. (pandas, version 1.5.2). Machine learning tasks were performed with scikit-learn24, provided by the scikit-learn developers (version 1.3.0). Figures were created using the Python package Plotly, developed by Plotly Technologies Inc. (Collaborative Data Science, Montréal, QC, 2015, version 5.9.0). The random seed was fixed at the value 42 for all computational processes that introduce randomness (random split, Mutual Information computation, random forest initializations, etc.) in order to enable reproducibility.
Chemical space
A representation of DATASET1 and DATASET4 chemical space was performed for visualization purposes. The Morgan fingerprints, also known as Extended-Connectivity fingerprints25, of its compounds were generated with a radius of 2 and were next passed to a t-SNE algorithm26 to generate a 2-dimensional embedding.
Molecular descriptors
Molecular descriptors were computed using the Mordred Python package (version 1.2.0)14. The canonical SMILES of the molecules were converted into their graph representations using the RDKit package (RDKit: Open-source cheminformatics, version 2023.03.1), which were subsequently used as input for descriptor computation. Descriptors with computational errors for at least one compound were excluded from the analysis.
The descriptors were normalized for subsequent steps using min-max scaling: based on the training set, each descriptor’s range was rescaled between 0 and 1 according to Eq. (3) below, where X represents the descriptor vector, \(X_{i}\) is the descriptor value for the i-th compound, and \(X'\) is the normalized vector:
Descriptors with zero variance were removed based on the training set. Additionally, from each group of highly correlated descriptors (Pearson correlation coefficient27, \(corr > 0.95\)), only the most informative descriptor with respect to EP2 activity (pEP2) was kept.
We selected the top 1 to 100 most informative descriptors based on Mutual Information (MI) as defined in Eq. (4); this approach has been previously used and validated in QSAR modeling28,29.
The MI was computed for each descriptor (X in Eq. (4)) relative to EP2 activity (Y in Eq. (4)). Descriptors were ranked in descending order, from the most to the least informative. This supervised selection approach was performed using various activity cutoffs ranging from pEP2> 3 to pEP2 > 8.
Modeling
Data splitting
DATASET1 was randomly split into two sets before modeling: a training set and a validation set. The training set included 80% of our dataset, and it was used for the ML model optimization process and training; the validation set (with 20% of our original dataset) was used to assess the performance of the optimized and trained ML models. The training set is split into 5 random folds used for cross-validation during the optimization process.
Optimization process
Several ML models were created using the training set; they were built with varying numbers of molecular descriptors, ranging from 10 to 100 in increments of 10. The goal was to identify the most effective combination of approach, cutoff, and features by assessing performance on the test set to predict on the DATASET4. The optimization process for each ML classifier was as follows: the hyperparameters of the algorithm are optimized using a 5-fold random cross-validation, stratified by the EP2 activity class on the training set. The optimization metric used is the Area Under the Receiver Operating Characteristic Curve (ROC AUC) and Precision-Recall curve (see the Performance Evaluation section below). The tuned hyperparameters are shown in Table 2. All other hyperparameters are set to their default values. Finally, the entire training set is used to train the final ML classifier, ensuring the best possible performance.
Performance evaluation
Receiving operating characteristic curve
The Receiving Operating Characteristic (ROC) curve is drawn as follows: the True Positive Rate (TPR, see Eq. (5)), also called Sensitivity or Recall) is computed along with the False Positive Rate (FPR, see Eq. (6)) at various probability thresholds p (Supplementary Fig. S7). Typically, each threshold p corresponds to each unique probability returned by the ML model for the tested set. At a given threshold p, positives (here active compounds) that are rightly predicted will be labeled as True Positives (TP) while wrongly predicted ones are False Negatives (FN). For Negatives (here inactive compounds), rightly predicted ones are labeled as true negatives, while wrongly predicted ones are False Positives (FP).
TPR, also known as Sensitivity (SEN) or Recall, corresponds to the capacity of the ML classifier to distinguish positives in the dataset while FPR, also known as Fall-out, corresponds to the struggling of the ML classifier to distinguish negatives. TPR is the opposite of the True Negative Rate (TNR, see 7), also known as Specificity (SPE), as \(FPR = 1 - SPE\), and SPE corresponds to the capacity of the ML classifier to distinguish negatives. This means that the ROC Curve gives the balance between SEN and SPE at each threshold, giving at which cost (FPR) the ML classifier is able to find positives (TPR).
From a ROC curve drawn this way, the Area Under the Curve (AUC) gives then an indication of the ML classifier performance: the higher, the better. A higher ROC AUC score suggests that it is able to find positives better (high SEN), for a lesser cost (low FPR, high SPE).
The computations to draw the ROC curve of each ML classifier and get their ROC AUC were made through the functions sklearn.metrics.roc_curve and sklearn.metrics.auc from the scikit-learn package.
Precision-recall curve and average precision
The Precision-Recall curve is drawn as follows: the Recall (or TPR, see Eq. (5)) is computed at various probability thresholds p in the same manner as the ROC curve, but along with the Precision (also called Positive Predictive Value, PPV) detailed in Eq. (8).
The Average precision (AP) is then computed from the Precision-Recall curve as described in Eq. (9) below:
Medchem, scaffold selection and pharmacology
Compounds were provided by Medetia Pharmaceuticals. In short, selected compounds were EP2 agonists with either N,N-alkyl alkoxy-sulfonamide or N,N-alkyl alkoxy-amide scaffolds. Solubility of the compounds was verified visually in assay conditions. Compounds were stored in DMSO at -20\(\phantom{0}^\circ\).
Representation of the scaffolds of interest in DATASET1 (a) Molecular structures of the three scaffolds of interest. (b) Low-dimensional visualization of the t-SNE similarity landscape based on Morgan fingerprints, with colors representing distinct scaffolds and (c) with colors indicating compounds’ affinity type toward EP2 (agonist/antagonist).
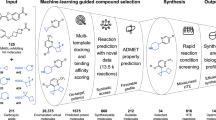
Workflow of the data gathering, ML model optimization, feature selection and experimental validation processes.
Descriptor computation and feature selection informativeness ranking. (a) The workflow outlines the steps taken from initial SMILES data through Mordred to compute 1826 descriptors, followed by the exclusion criteria leading to the selection of the top 100 descriptors based on variance and correlation thresholds. (b) The plot displays the mutual information values of descriptors for varying activity score cutoffs (pEP2> 3 to > 8), with descriptors ordered by informativeness at each cutoff. This ranking illustrates that descriptor relevance can change with different activity thresholds. (c) and (d): Evolution of the RFC models’ performance by area under the Receiving Operating Characteristic curve (ROC AUC, (c)) or Average Precision (from Precision Recall curve (d)) as a function of the number of features (descriptors) in input, toward the validation set alone (blue) or with the addition of the DATASET2 (orange). Random prediction on validation only gives a ROC AUC value of 0.5 and an Average Precision value of 0.47; Random prediction on validation+DATASET2 give a ROC AUC value of 0.5 and an Average Precision of 0.004 (active ratio of 0.4% in this configuration).
Experimental validation of the selected RFC model on DATASET4 colored by the compound’s scaffold: (a) Barplot of the experimental cellular response toward drugs in percentage compared to EP2 natural agonist PGE2; (b) scatter plot of the biological response (%) at \(10^{-6}M\) against the predicted probability of activity; (c) t-SNE representation of the DATASET4 with the DATASET1 representation from Figure 1 in the background.
EP2 in vitro pharmacology
The agonist activity of compounds at the human EP2 receptor expressed in transfected CHO cells was evaluated by measuring their effects on cAMP production using HTRF. In short, cells were suspended in HBSS buffer supplemented with 20 mM HEPES (pH 7.4) and 500 \(\upmu\)M IBMX, then plated at a density of 104 cells/well in microplates and incubated for 30 minutes at \(37^\circ\)C with either the test compound at 1 \(\upmu\)M final concentration, the reference agonist, or no compound (control). For stimulated control measurements, wells contained 10 \(\upmu\)M PGE2. Following incubation, the cells were lysed, and fluorescence acceptor (D2-labeled cAMP) and donor (anti-cAMP antibody labeled with europium cryptate) were added. After a 60-minute incubation at room temperature, fluorescence transfer was measured using a microplate reader at excitation 337 nm and emission wavelengths of 620 and 665 nm. The cAMP concentration was calculated by dividing the signal at 665 nm by that at 620 nm (ratio). Results were expressed as a percentage of the control response to 10 \(\upmu\)M PGE2. The \(EC_{50}\) value of the standard reference agonist, PGE2, was determined from a concentration-response curve generated in each experiment30. CHO cell lines were obtained from the Department of In-vitro Pharmacology, GlaxoSmithKline Medicines Research Center (Gunnels Wood Road, Stevenage); the experiments were performed by Eurofins Discovery (Poitiers, France).
Data Availability
Due to intellectual property and strategic considerations, datasets have not been uploaded in public data warehouses ; however, the datasets used and/or analyzed during the current study are available from the corresponding author.
References
Cooper, P. The prostaglandins. Midwife. Health Visit. Community Nurse 19, 321. https://doi.org/10.2307/1294030 (1983).
Funk, C. D. Prostaglandins and leukotrienes: Advances in eicosanoid biology. Science 294, 1871–1875. https://doi.org/10.1126/science.294.5548.1871 (2001).
Jiang, J. & Dingledine, R. Prostaglandin receptor EP2 in the crosshairs of anti-inflammation, anti-cancer, and neuroprotection. Trends Pharmacol. Sci. 34, 413–423. https://doi.org/10.1016/j.tips.2013.05.003 (2013).
Garcia, H. et al. Agonists of prostaglandin E2 receptors as potential first in class treatment for nephronophthisis and related ciliopathies. Proc. Natl. Acad. Sci. U. S. A. 119, 1–11. https://doi.org/10.1073/pnas.2115960119 (2022).
Ganesh, T., Ganesh, Thota, Ganesh, T. & Ganesh, Thota. Prostanoid receptor EP2 as a therapeutic target. J. Med. Chem. 57, 4454–4465. https://doi.org/10.1021/jm401431x (2024).
Markovič, T., Jakopin, Ž, Dolenc, M. S. & Mlinarič-Raščan, I. Structural features of subtype-selective EP receptor modulators. Drug Discov. Today 22, 57–71. https://doi.org/10.1016/j.drudis.2016.08.003 (2017).
Catacutan, D. B., Alexander, J., Arnold, A. & Stokes, J. M. Machine learning in preclinical drug discovery. Nat. Chem. Biol. https://doi.org/10.1038/s41589-024-01679-1 (2024).
Cortes, C. & Vapnik, V. Support-vector networks. Mach. Learn. 20, 273–297. https://doi.org/10.1007/BF00994018 (1995).
Breiman, L. Random forests. Mach. Learn. 45, 5–32. https://doi.org/10.1023/A:1010933404324 (2001).
Rosenblatt, F. The perceptron: A probabilistic model for information storage and organization in the brain. Psychol. Rev. 65, 386–408. https://doi.org/10.1037/h0042519 (1958).
Prasanna, G. et al. Effect of PF-04217329 a prodrug of a selective prostaglandin EP 2 agonist on intraocular pressure in preclinical models of glaucoma. Experiment. Eye Res. 93, 256–264. https://doi.org/10.1016/j.exer.2011.02.015 (2011).
Coleman, R. A. et al. The affinity, intrinsic activity and selectivity of a structurally novel EP 2 receptor agonist at human prostanoid receptors. British Pharmacol. 176, 687–698. https://doi.org/10.1111/bph.14525 (2019).
Iwamura, R. et al. Identification of a Selective, Non-Prostanoid EP2 Receptor Agonist for the Treatment of Glaucoma: Omidenepag and its Prodrug Omidenepag Isopropyl. J. Med. Chem. 61, 6869–6891. https://doi.org/10.1021/acs.jmedchem.8b00808 (2018).
Moriwaki, H., Tian, Y.-S., Kawashita, N. & Takagi, T. Mordred: A molecular descriptor calculator. J. Cheminform. 10, 4. https://doi.org/10.1186/s13321-018-0258-y (2018).
Chithrananda, S., Grand, G. & Ramsundar, B. Chemberta: Large-scale self-supervised pretraining for molecular property prediction (2020). 2010.09885.
Papadatos, G. et al. SureChEMBL: a large-scale, chemically annotated patent document database. Nucleic Acids Res. 44, D1220–D1228. https://doi.org/10.1093/nar/gkv1253 (2016).
Harding, S. D. et al. The IUPHAR/BPS Guide to PHARMACOLOGY in 2024. Nucleic Acids Res. 52, D1438–D1449. https://doi.org/10.1093/nar/gkad944 (2024).
Jagarlapudi, S. A. R. P. & Kishan, K. V. R. Database Syst. Knowl. Discovery 159–172 (publisherHumana Press, addressTotowa, NJ, 2009).
Mendez, D. et al. ChEMBL: towards direct deposition of bioassay data. Nucleic Acids Res. 47, D930–D940. https://doi.org/10.1093/nar/gky1075 (2019).
Goedken, E. R. et al. HTRF-based assay for microsomal prostaglandin E2 synthase-1 activity. J. Biomol. Screen. 13, 619–625. https://doi.org/10.1177/1087057108321145 (2008).
Jawarkar, R. D. et al. QSAR and docking based lead optimization of nitrogen heterocycles for enhanced prostaglandin EP2 receptor agonistic potency. Chem. Phys. Impact 8, 100484. https://doi.org/10.1016/j.chphi.2024.100484 (2024).
Mysinger, M. M., Carchia, M., Irwin, J. J. & Shoichet, B. K. Directory of useful decoys, enhanced (DUD-E): Better ligands and decoys for better benchmarking. J. Med. Chem. 55, 6582–6594. https://doi.org/10.1021/jm300687e (2012).
Harris, C. R. et al. Array programming with NumPy. Nature 585, 357–362. https://doi.org/10.1038/s41586-020-2649-2 (2020).
Pedregosa, F. et al. Scikit-learn: Machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Rogers, D. & Hahn, M. Extended-Connectivity Fingerprints. J. Chem. Inform. Model. 50, 742–754. https://doi.org/10.1021/ci100050t (2010).
van der Maaten, L. & Hinton, G. E. Visualizing Data using t-SNE. J. Mach. Learn. Res. 9, 2579–2605 (2008).
Benesty, J., Chen, J., Huang, Y. & Cohen, I. Pearson Correlation Coefficient. 1–4 (publisherSpringer Berlin Heidelberg, addressBerlin, Heidelberg, 2009).
Venkatraman, V., Dalby, A. R. & Yang, Z. R. Evaluation of mutual information and genetic programming for feature selection in QSAR. J. Chem. Inform. Comput. Sci. 44, 1686–1692. https://doi.org/10.1021/ci049933v (2004).
Danishuddin & Khan, A. U. Descriptors and their selection methods in QSAR analysis: paradigm for drug design. https://doi.org/10.1016/j.drudis.2016.06.013 (2016).
Wilson, R. J. et al. Functional pharmacology of human prostanoid EP2 and EP4 receptors. European Pharmacol. 501, 49–58. https://doi.org/10.1016/j.ejphar.2004.08.025 (2004).
Acknowledgements
We are grateful to Pierrick Craveur for his help on the compilation of ChEMBL data subset. We are grateful to Maud Jusot and Brice Hoffman (Iktos) for providing the decoy libraries. We are also grateful to Maud Fleury and Celine Gros (Eurofins) for the discussion on EP2 binding assays and biology.
Author information
Authors and Affiliations
Contributions
P.D., V.O., S.W., P.N., J-P.A. L.B-R. conceived the experiment(s); P.D. C.L. compiled the datasets; P.D., A.R., J-P.A, L.B-R, designed molecules; P.D. conducted the coding and ML experiments; P.D., V.O., S.W., P.N., J-P.A. L.B-A. analysed the results. P.N. P.D. and L.B-R drafted the manuscripts and addressed reviewers’ comments. All authors reviewed the manuscript
Corresponding author
Ethics declarations
Competing interests
LB-R and J-PA are equity holders of Medetia Pharmaceuticals and coinventors in the patent application WO2019075369 “Methods for Treating Diseases Associated With Ciliopathies.” All the other authors declared no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Dupuyds, P., Rayar, A., Opuu, V. et al. Ligand-based machine learning models to classify active compounds for prostaglandin EP2 receptor. Sci Rep 16, 2844 (2026). https://doi.org/10.1038/s41598-025-23764-8
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-23764-8