Abstract
Chemotherapy-induced neutropenia (CIN) and febrile neutropenia (FN) significantly increase the risk of invasive infections, which can progress to septic shock or sepsis and may ultimately result in death. our purpose of this study is to construct a predictive model based on peripheral blood inflammatory markers to enable timely detection of bloodstream infections (BSI) and evaluation of infection severity in this population. A total of 112 patients with chemotherapy-induced neutropenia (CIN) who developed BSI and 145 patients without BSI (NBSI) were included. Inflammatory marker levels were compared across subgroups. The Logistic regression was performed to identify risk factors for BSI within the training cohort. Model performance was assessed by receiver operating characteristic curves (ROC), decision curve analysis (DCA) and diagnostic calibration curves. Patients with BSI had longer disease duration, a higher proportion of acute lymphoblastic leukemia, and significantly lower levels of complete blood count and inflammatory markers compared to those without BSI. Univariate analysis identified prolonged disease duration, hematologic diagnosis category, and reduced levels of NLR, MLR, PLR, SIRI, and SII as risk factors for BSI. A nomogram model was constructed incorporating disease duration, hematologic diagnosis category, MLR, PLR, and SII. ROC and DCA indicated a broad threshold range for clinical applicability and a high net benefit of predictive model, supporting its clinical utility. Diagnostic calibration curves showed good agreement between predicted and observed outcomes, with a Hosmer–Lemeshow test P-value of 0.3289. A nomogram model incorporating disease duration, hematologic diagnosis category, MLR, PLR, and SII was constructed and shown to provide a practical and effective tool for the assessment and early identification of BSI in patients with CIN.
Similar content being viewed by others
Introduction
Neutropenia is the most common hematologic adverse event and a major dose-limiting toxicity induced by chemotherapy. Chemotherapy-induced neutropenia (CIN) and febrile neutropenia (FN) substantially increase the risk of invasive infections, which may progress to sepsis and septic shock, ultimately resulting in death1,2. Therefore, early identification and evaluation of infection risk in patients with CIN are of critical importance. Bloodstream infections (BSI), the leading cause of infection-related complications in this population, are a primary clinical concern during the diagnosis and management of FN, often progressing to sepsis or septic shock3. Blood culture remains the gold standard for the diagnosis of BSI4; however, its clinical utility is limited by a low positivity rate, long turnaround time, large blood volume required, and a high risk of contamination, posing significant challenges to early clinical decision-making5,6,7. Therefore, identifying appropriate biomarkers to screen for BSI in patients with CIN may offer a more effective alternative. Previous studies have shown that peripheral blood inflammatory markers can effectively detect BSI at an early stage and assess infection severity, prognosis, and clinical outcomes in non-CIN populations, including those in intensive care units and on hemodialysis8,9,10.
However, the aforementioned inflammatory indices are derived from calculations involving neutrophil (N), lymphocyte (L), monocyte (M), and platelet (PLT) counts. In patients with CIN, the marked reduction in neutrophil levels results in distinct peripheral blood leukocyte profiles compared to those in non-chemotherapy populations. Consequently, it remains unclear whether inflammatory marker trends observed during BSI in CIN patients mirror those seen in non-CIN individuals. Moreover, studies specifically examining the inflammatory marker patterns associated with BSI in CIN patients are currently limited. To address this gap, we conducted a retrospective analysis of clinical data from 112 CIN patients with confirmed BSI, with the aim of evaluating whether peripheral blood inflammatory markers could serve as rapid and effective tools for BSI screening and infection severity assessment in this population. We further developed a predictive model based on these markers and assessed its performance.
Materials and methods
Study population
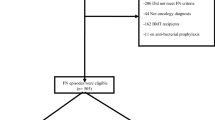
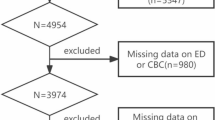
This retrospective study included patients with hematological malignancies and CIN who were diagnosed with BSI via blood culture between April 2020 and March 2025 at the Department of Hematology, Quanzhou First Hospital Affiliated to Fujian Medical University. A total of 112 patients were enrolled, comprising 57 males and 55 females, aged 18 to 85 years, with a median age of 47 years. As disease controls, 145 patients without BSI (NBSI)—matched by sex and age—were selected during the same period. These control patients were diagnosed by blood culture and final clinical evaluation as having non-bloodstream infections. The NBSI group included 81 males and 64 females, aged 16 to 72 years, with a median age of 54 years. For patients with multiple hospitalizations or repeated testing, only data from the initial clinical presentation were included. All protocols were approved by Ethics Committee of Quanzhou First Hospital (China), the protocol number is [2025] K218.
Inclusion and exclusion criteria
Inclusion criteria
Patients were eligible for inclusion if they met all of the following criteria: ① Presence of clinical signs suggestive of infection, including fever, nausea and vomiting, abdominal pain or diarrhea, chest tightness or dyspnea, urinary frequency, or other relevant symptoms; ② Underwent both blood culture and complete blood count testing during the same clinical episode; ③ Had a documented diagnosis of a primary hematological malignancy, such as acute leukemia, lymphoma, or myelodysplastic syndrome; ④ patients were undergoing chemotherapy at the time of blood culture collection, with CIN confirmed by an absolute neutrophil count (ANC) of < 2.0 × 109/L11.
Exclusion criteria
Patients were excluded if they met any of the following criteria: ① CIN resulting from chemotherapy for non-hematological malignancies; ② Secondary hematologic disorders caused by drugs or immune-related conditions; ③ ANC < 2.0 × 109/L without clear evidence that neutropenia was induced by chemotherapy; ④ Incomplete clinical or laboratory data; ⑤ Blood cultures in which only one of multiple bottles tested positive and the isolated organism was suspected to be a contaminant, making it impossible to distinguish between true BSI and contamination.
Sample preparation
Peripheral venous blood (2 mL) was collected from each patient into an EDTA-K₂ anticoagulant tube (EDTA3K 2 mL, Fuzhou Changgeng MEDICAL Devices Co., Ltd., China). After gentle inversion for mixing, samples were processed using an automated hematology analyzer (Sysmex XN9000, Sysmex Corporation, Japan) in accordance with the manufacturer’s instructions. The following parameters were recorded: white blood cell (WBC), N, L, M, and PLT counts. Inflammatory indices were calculated as follows: NLR, monocyte-to-lymphocyte ratio (MLR), PLR, systemic inflammatory response index (SIRI = N × M / L), and systemic immune inflammation index (SII = N × PLT / L).
Blood culture and microbial identification
Venous blood (8–10 mL) was aseptically collected from each patient by clinical staff and inoculated into blood culture bottles (Becton, Dickinson and Company [BD], USA) for microbiological testing. Cultures were processed using an automated blood culture system (BACTEC FX, BD, USA). Blood culture results were interpreted based on instrument reports as follows: ① Negative: No positive signal reported within 120 h; ② Positive: A positive signal reported within 120 h, after which the culture bottle was removed and subcultured onto blood agar and China blue agar plates (Guangzhou Dijing Microbial Technology Co., Ltd, China). Bacterial colonies were inspected the following day, and species identification along with antimicrobial susceptibility testing was performed using an automated system (Phoenix 100, BD, USA).
Clinical data collection
Clinical information was obtained from the electronic medical record system of Quanzhou first hospital. Data collected included sex, age, history of hypertension, history of diabetes, hematologic diagnosis, duration of illness, presence of infections other than BSI, presence of chronic internal medical conditions (e.g., hepatitis, coronary artery disease, chronic kidney disease, chronic obstructive pulmonary disease), and infection severity in patients with BSI.
Subgroup classification within the BSI cohort
Based on clinical data, the 112 patients with BSI were stratified into subgroups according to the following criteria:
① Co-infection status: Patients were categorized into the simple BSI (si-BSI, n = 44), who had BSIs without other concurrent infections, and the BSI with co-infections (co-BSI, n = 68), who had concurrent infections such as pulmonary infections, urinary tract infections, or peritonitis. ② Infection severity: Patients were classified as having general BSI (ge-BSI, n = 80) if they showed clinical improvement and were discharged following blood culture collection and treatment, or serious BSI (se-BSI, n = 32) if they experienced shock, cardiopulmonary arrest, multiorgan failure, or required emergency resuscitation. ③ Microbial classification: According to pathogen type, patients were grouped into Gram-negative BSI (G-BSI, n = 81) and Gram-positive BSI (G + BSI, n = 20). Patients with fungal (n = 4) or mixed infections (n = 7) were not included in subgroup comparisons due to their small numbers.
Cohort division for model development and validation
The BSI and NBSI cohorts were randomly divided into training and validation sets in an 8:2 ratio. The training cohort included 87 patients with BSI and 115 patients with NBSI, while the validation cohort included 25 patients with BSI and 30 with NBSI. There was no statistically significant difference between the positive groups of the training and validation sets, nor between the negative groups of the two sets. Data from the training cohort were used to develop the predictive model, and data from the validation cohort were used to evaluate its performance.
Statistical analysis
Statistical analyses were performed using SPSS version 21.0 and R version 4.2.1. Mean ± standard deviation (X ± SD) represents continuous variables with normally distributed, such as age, were expressed as and compared between groups using independent samples t-tests. Disease duration, WBC, N, L, M, PLT, NLR, MLR, PLR, SIRI, and SII were non-normally distributed variables, presented as median (interquartile range, IQR) and compared using the Mann–Whitney U test. Categorical variables were reported as percentages: sex was expressed as the proportion of male patients; history of hypertension, diabetes, presence of co-infections, and chronic internal diseases were reported as positive rates; hematologic diagnoses were expressed as the distribution of disease types. Group comparisons for categorical variables were conducted using the chi-square (χ2) test.
Binary logistic regression was used to identify risk factors for BSI in the training cohort. Spearman correlation analysis was applied to assess correlations among inflammatory indices. Variance inflation factor (VIF) analysis was performed to diagnose collinearity and select the most informative predictors for construction of a nomogram predictive model. The diagnostic performance of individual predictors and the predictive model was assessed using receiver operating characteristic curves (ROC). Decision curve analysis (DCA) and diagnostic calibration curves were used to validate and evaluate the predictive model. A two-tailed P value < 0.05 was considered statistically significant.
Results
Pathogen distribution in the BSI cohort
Among the 112 patients with BSI, a total of 119 pathogenic strains were isolated, including 7 cases of mixed infections involving two pathogens (6.25%). Of the isolates, 81 (72.32%) were Gram-negative (G −) bacteria, 20 (17.86%) were Gram-positive (G +) bacteria, and 4 (3.57%) were fungi. The three most frequently isolated pathogens—all Gram-negative—were Escherichia coli (25.00%), Klebsiella pneumoniae subsp. pneumoniae (23.21%), and Pseudomonas aeruginosa (16.96%). Notably, among the seven cases of mixed infection, five involved K. pneumoniae subsp. pneumoniae combined with other pathogens (Table 1).
Comparison of inflammatory marker levels among BSI subgroups
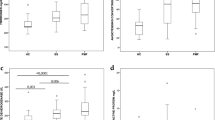
Comparison of inflammatory index levels between different subgroups within the BSI group. The results showed that SII and NLR can be used to distinguish whether there is co infection and evaluate the severity of infection (Fig. 1). Among patients with BSI, those in the co-BSI group had significantly higher SII levels compared to those in the si-BSI group, with a statistically significant difference (P = 0.033). The NLR level in the se-BSI group was significantly higher than that in the ge-BSI group, and the difference was statistically significant (P = 0.006). There was no statistically significant difference in the levels of all inflammatory indicators between the G-BSI group and the G + BSI group.
(A) Comparison of inflammatory marker levels between the co-BSI group and the si-BSI group. (B) Comparison of inflammatory marker levels between the ge-BSI group and the se-BSI group. (C) Comparison of inflammatory marker levels between the G-BSI group and the G + BSI group.
Development of a nomogram predictive model for BSI prediction
Comparison of clinical characteristics between the BSI and NBSI groups
Compared with the NBSI group, patients in the BSI group had a significantly longer disease duration and a higher proportion of acute lymphoblastic leukemia (P values are 0.002 and < 0.001, respectively). In addition, complete blood count parameters and inflammatory marker levels were significantly lower in the BSI group, with all differences reaching statistical significance (P < 0.001, except for PLR which had a P value of 0.009) (Table 2).
Logistic regression analysis of BSI risk factors
Univariate logistic regression analysis identified prolonged disease duration, hematologic diagnosis category (primarily acute leukemia), and decreased levels of inflammatory markers as individual risk factors for the occurrence of BSI (P values are all < 0.05). However, multivariate analysis revealed that only prolonged disease duration was an independent predictor of BSI (P = 0.004) (Tables 3).
ROC analysis of individual inflammatory markers
ROC analysis was performed to evaluate the diagnostic value of NLR, MLR, PLR, SIRI, and SII for identifying BSI. All markers demonstrated diagnostic utility, with areas under the curve (AUC) ranging from 0.607 to 0.775. Among them, SII exhibited the highest diagnostic performance, with an AUC of 0.775. The corresponding diagnostic metrics for each marker are summarized in Table 4.
Correlation analysis of inflammatory markers
As all inflammatory indices were derived from N, L, M, and PLT counts, potential collinearity was expected. Spearman correlation analysis was conducted to assess the relationships among these markers. Strong correlations (r > 0.7) were observed for the following pairs: NLR vs. SIRI (r = 0.715), NLR vs. SII (r = 0.868), MLR vs. SIRI (r = 0.856), and SIRI vs. SII (r = 0.713). Based on both correlation patterns and AUC performance, MLR, PLR, and SII were selected as candidate variables for nomogram predictive model construction.
Collinearity diagnostics
VIF were calculated for the candidate variables included in the predictive model: disease duration (VIF = 1.052), hematologic diagnosis (VIF = 1.084), MLR (VIF = 1.215), PLR (VIF = 1.622), and SII (VIF = 1.713). All VIF values were below 2, indicating weak collinearity among variables.
Nomogram predictive model construction
Disease duration, hematologic diagnosis, MLR, PLR, and SII—were incorporated into the construction of a nomogram for predicting BSI (Fig. 2).
Nomogram predictive model for predicting BSI. Evaluate the probability of developing BSI in patients with CIN hematological disorders by assigning scores to different indicators and calculating the total score. Note: ‘Disease duration (m)’ refers to the actual duration of illness in months. ‘Diagnosis’ is numerically coded as follows: 1 = acute myeloid leukemia, 2 = acute lymphoblastic leukemia, 3 = mature B-cell lymphoma, 4 = myelodysplastic syndrome, 5 = other hematologic diseases. ‘MLR’, ‘PLR’, and ‘log (SII + 1)’ values represent the actual calculated indices. The SII variable was transformed using log (SII + 1) to mitigate issues of complete or quasi-complete separation during model construction, as direct inclusion of raw SII values led to excessive predicted probabilities of 0 or 1.
Model validation
ROC analysis
ROC analysis was conducted to assess the discriminative ability of the predictive model for identifying BSI in both the training and validation cohorts. In the training cohort, the model demonstrated an AUC of 0.817 (95% CI: 0.758–0.876), with a sensitivity of 81.62%, specificity of 76.52%, positive predictive value (PPV) of 72.45%, and negative predictive value (NPV) of 84.62% (Fig. 3A). In the validation cohort, the model yielded an AUC of 0.781 (95% CI: 0.658–0.905), with a sensitivity of 92.00%, specificity of 63.33%, PPV of 67.65%, and NPV of 90.48% (Fig. 3B).
Validation of predictive model. (A) The ROC of the predictive model screening training cohort BSI. (B) ROC of the predictive model screening validation cohort BSI. (C) DCA of the predictive model for BSI in the validation cohort. (D) Diagnostic calibration curve of the predictive model for BSI in the validation cohort.
DCA
The decision curve of the predictive model was markedly distant from the “None” and “All” extremes (Fig. 3C), indicating a wide threshold range and a relatively high net clinical benefit. These results suggest that the model may offer substantial value in guiding clinical decision-making.
Diagnostic calibration curve
Diagnostic calibration curve of the predictive model in the validation cohorts, Hosmer-Lemeshow test result P = 0.3289, indicating good model fit (Fig. 3D).
Discussion
This study first investigated the distribution of pathogens isolated from 112 patients with BSI, providing preliminary insight into the predominant causative organisms of BSI in CIN patients within the local region. Subsequent comparisons of inflammatory marker levels across BSI subgroups revealed that the SII and NLR may be useful in evaluating the presence of co-infections and infection severity, respectively. A nomogram predictive model based on routine hematological inflammatory markers was then developed using training and validation cohorts to predict the risk of BSI in CIN patients. The predictive efficacy of this model was confirmed by ROC analysis, DCA, and diagnostic calibration curve assessment, demonstrating its potential to serve as a valuable tool in guiding clinical decision-making.
Among the 112 patients with BSI, a total of 119 pathogenic strains were isolated, including 7 cases involving mixed infections with two pathogens. The three most frequently isolated organisms were all G − bacteria: Escherichia coli, Klebsiella pneumoniae subsp. pneumoniae, and Pseudomonas aeruginosa. These findings indicate that BSI in hematologic patients with CIN at our institution are predominantly caused by G − bacteria and are characterized by a diverse spectrum of pathogens. Notably, in 5 of the 7 cases of mixed infection, K. pneumoniae subsp. pneumoniae was co-isolated with other pathogens, suggesting a potential association between K. pneumoniae subsp. pneumoniae infection and increased susceptibility to mixed infection. However, in this study, there were only 7 cases of mixed infection, which is too small to support this viewpoint. Therefore, this viewpoint still needs to be further validated by expanding the sample size or conducting multicenter studies. This pattern is consistent with previously reported pathogen profiles among hematologic patients with BSI12,13,14. However, compared with similar studies on hematologic patients with CIN, researchers such as Piukovics, Liu, and Guo have reported that Staphylococcus aureus and coagulase-negative staphylococci—both G + organisms—were among the top three pathogens isolated in cases of BSI15,16,17. This discrepancy may be attributable to differences in inclusion criteria: the aforementioned studies focused on patients with severe neutropenia (ANC < 0.5 × 109/L), whereas the present study included patients with neutropenia defined as ANC < 2.0 × 109/L. Another possible explanation lies in the study design. Our retrospective analysis was limited to CIN patients who underwent both blood culture and complete blood count testing simultaneously, and was conducted at a single center. These factors introduce notable limitations and may not fully capture the comprehensive pathogen spectrum of BSI in hematologic patients with CIN. In the future studies, a large number of specimens and multicenter participation are warranted to further characterize the microbiological profiles in this population.
To further explore the clinical relevance of inflammatory markers, the BSI group was stratified by infection characteristics, and intergroup comparisons were conducted. The results indicate that SII levels were significantly higher in the co-BSI group compared to the si-BSI group, while NLR levels were significantly elevated in the se-BSI group relative to the ge-BSI group. These results indicate that inflammatory markers may reflect different infection profiles in patients with BSI, especially SII and NLR. Specifically, elevated SII levels may indicate the presence of additional infections beyond BSI, whereas an increase in NLR may signal a more severe infection burden. The role of NLR observed in this study aligns with findings by Roldgaard in patients with sepsis not related to CIN10. However, no prior studies have been identified that specifically examine the relationship between SII and infection characteristics in BSI, making direct comparisons unavailable. Additionally, comparisons between the G − BSI and G + BSI groups revealed no significant differences in any of the inflammatory marker levels. This suggests that such markers may not be suitable for assessing the type of infecting organisms in CIN patients with BSI. Nonetheless, the findings regarding NLR and SII may support clinical assessment of infection severity and inform timely adjustments in therapeutic strategies.
In this study, random sampling was used to divide both the BSI and NBSI groups in an 8:2 ratio. Data from the training cohort were used to construct the predictive model, while the validation cohort was used to evaluate its performance. Comparative analysis of clinical characteristics and inflammatory markers between the BSI and NBSI groups in the training cohort revealed that BSI patients had significantly longer disease durations and a higher proportion of acute lymphoblastic leukemia. Furthermore, routine hematologic and inflammatory marker levels were markedly lower in the BSI group. These findings are consistent with results obtained in our preliminary studies18. These results indicate that prolonged disease duration and increased chemotherapy exposure elevate the risk of developing BSI in patients with hematologic malignancies, specifically acute myeloid leukemia and acute lymphoblastic leukemia—being the most susceptible. Prolonged disease duration predisposes patients to BSI, which are closely associated with the frequency and length of hospitalizations. Repeated and extended admissions entail greater exposure to chemotherapy and invasive diagnostic or therapeutic procedures, resulting in progressive immunosuppression and an increased risk of BSI. In addition, the lower the WBC and differential counts during episodes of CIN, the higher the likelihood of BSI occurrence. This conclusion was supported by the subsequent logistic regression analysis and is consistent with current expert consensus, which recognizes CIN as a significant risk factor for invasive infections. On the other hand, inflammatory marker levels in the BSI group were significantly decreased, in stark contrast to their typically elevated levels observed in BSI patients without CIN8,9,19. The suggests that in hematologic patients with CIN, a lower level of systemic immune response is related to a higher risk of developing BSI. Therefore, in clinical practice, declining inflammatory marker levels in suspected cases of BSI among CIN patients should be closely monitored and not overlooked.
Given the narrow numerical range of WBC and its differential counts, particularly N, which typically fall within 0 to 2 × 109/L in CIN patients. This study opted to use composite inflammatory markers instead of absolute WBC or subtype counts for constructing the nomogram. These markers, which represent ratios of different leukocyte subtypes, provide more representative and scalable metrics. Considering the overlapping formulas used to derive these indices, we first conducted a correlation analysis. Strong correlations were observed among NLR, SIRI, and SII, as well as between MLR and SIRI. Based on their respective AUCs for BSI prediction and the distribution characteristics shown in Table 2, SII and MLR were selected as model variables. After collinearity diagnosis using VIFs, five variables (disease duration, hematologic diagnosis, MLR, PLR, and SII) were ultimately included in the final nomogram. Notably, direct inclusion of SII led to a high frequency of predicted probabilities clustering at 0 or 1, suggesting the presence of complete or quasi-complete separation. However, given the superior diagnostic performance of SII as a single predictor (AUC = 0.775, 95% CI: 0.711–0.839), it was retained in the model after logarithmic transformation as log (SII + 1), thereby preserving its predictive utility. Additionally, PLR exhibited a pattern in the predictive model whereby higher values corresponded to higher scores, which contradicts the earlier findings of this study. This inconsistency may be attributed to underlying correlations among the inflammatory markers included in the model—namely, MLR, PLR, and log (SII + 1).
The nomogram predictive model developed in this study was validated using ROC analysis, DCA, and diagnostic calibration curve assessment. ROC analysis demonstrated an AUC of 0.817 (95% CI: 0.758–0.876) in the training cohort and 0.781 (95% CI: 0.658–0.905) in the validation cohort, indicating favorable diagnostic performance for BSI screening in both cohorts. Notably, the model outperformed any individual inflammatory marker in the training cohort, suggesting that incorporating disease duration and hematologic diagnosis significantly enhanced predictive accuracy. In the DCA, the model’s decision curve was well-separated from the “None” and “All” extremes, indicating a wide threshold range and high net clinical benefit. This highlights the model’s potential utility in guiding clinical decision-making. The diagnostic calibration curve showed a Hosmer–Lemeshow P-value of 0.3289, suggesting good agreement between predicted and observed risks. However, the calibration curve slightly deviated above the ideal line within a certain probability range, indicating that the model may underestimate the actual risk of BSI in CIN patients to some extent. Consequently, for CIN patients with suspected BSI, clinicians should consider empirical prophylactic antibiotic therapy based on individual clinical assessment.
Finally, this study was a retrospective analysis rather than a prospective investigation. Therefore, when comparing the BSI and NBSI groups, CIN patients matched by age and sex were selected to reduce baseline differences. However, due to substantial missing data on body mass index (BMI) among study participants, comparative analysis involving age, sex, and BMI could not be conducted. However, previous studies have suggested that increasing age may be associated with a higher risk of infection in patients with CIN20,Moreover, a lower baseline BMI has been associated with poorer infection-related outcomes21. Therefore, future studies will adopt a prospective design and incorporate additional variables such as age, sex, and BMI to further refine and improve the predictive model.
This study identified several characteristic features of BSI in hematologic patients with CIN, including prolonged disease duration, predominance of acute leukemia, and significantly reduced levels of routine hematologic inflammatory markers. Among these markers, the SII and NLR demonstrated some utility in reflecting infection severity. A nomogram incorporating disease duration, hematologic diagnosis, MLR, PLR, and SII was developed and validated, offering a straightforward and effective tool for BSI risk assessment and screening in clinical settings. However, this was a single-center retrospective study with inherent limitations. Future prospective multicenter studies and external validation are warranted to further refine and validate the predictive model.
Data availability
All data generated or analyzed during this study are included in this published article.
Abbreviations
- CIN:
-
Chemotherapy-induced neutropenia
- BSI:
-
Bloodstream infections
- NLR:
-
Neutrophil-to-lymphocyte ratio
- MLR:
-
Monocyte-to-lymphocyte ratio
- PLR:
-
Platelet-to-lymphocyte ratio
- SIRI:
-
Systemic inflammatory response index
- SII:
-
Systemic immune inflammatory index
References
Ba, Y. et al. Current management of chemotherapy-induced neutropenia in adults: key points and new challenges: Committee of neoplastic supportive-care (CONS), China anti-cancer association committee of clinical chemotherapy. China Anti-Cancer Association. Cancer Biol. Med. 17(4), 896–909 (2020).
China Anti-Cancer Association Tumor Clinical Chemotherapy Professional Committee, China Anti-Cancer Association Tumor Support Therapy Professional Committee. Consensus on clinical diagnosis, treatment, and prevention of chemotherapy-induced neutropenia in China (2023 edition). Chin. J. Oncol. 45 (07): 575–583 (2023)
Hansen, B. A. et al. Febrile Neutropenia in Acute Leukemia. Epidemiology, Etiology, Pathophysiology and Treatment. Mediterr J. Hematol. Infect. Dis. 12(1), e2020009 (2020).
Society of Clinical Microbiology and Infection of China International Exchange and Promotion Association for Medical and Healthcare, Clinical Microbiology Group of the Laboratory Medicine Society of the Chinese Medical Association, Clinical Microbiology Group of the Microbiology and Immunology Society of the Chinese Medical Association. Chinese expert consensus on the clinical practice of blood culture in the diagnosis of bloodstream infection. Chin. J. Lab. Med.. 45 (02): 105–121 (2022)
Zboromyrska, Y. et al. Evaluation of the Magicplex™ sepsis real-time test for the rapid diagnosis of bloodstream infections in adults. Front. Cell Infect. Microbiol. 9, 56 (2019).
Peri, A. M., Harris, P. & Paterson, D. L. Culture-independent detection systems for bloodstream infection. Clin. Microbiol. Infect. 28(2), 195–201 (2022).
Ignak, S., Unay-Demirel, O. & Yuksel, M. Early prediction of bloodstream infection with complete blood count parameters: an ex-vivo human whole blood model. Clin. Lab. 70(4), 725 (2024).
Li, Y. et al. Pathogenic bacteria features of central line-associated bloodstream infections in ICU patients: focus on the early predictive value of neutrophil-to-lymphocyte and platelet-to-lymphocyte ratios. Front. Cell Infect. Microbiol. 15, 1525758 (2025).
Yang, J. et al. Diagnostic value of systemic inflammatory response index for catheter-related bloodstream infection in patients undergoing haemodialysis. J. Immunol. Res. 2022, 7453354 (2022).
Roldgaard, M., Benfield, T. & Tingsgård, S. Blood neutrophil to lymphocyte ratio is associated with 90-day mortality and 60-day readmission in Gram negative bacteremia: a multi-center cohort study. BMC Infect. Dis. 24(1), 255 (2024).
Freites-Martinez, A. et al. Using the common terminology criteria for adverse events (CTCAE - Version 5.0) to evaluate the severity of adverse events of anticancer therapies. Actas Dermosifiliogr (Engl Ed). 112(1), 90–92 (2021).
Li, Q. et al. Machine learning-based prognostic model for bloodstream infections in hematological malignancies using Th1/Th2 cytokines. BMC Infect. Dis. 25(1), 415 (2025).
Wu, H. et al. Pathogenic spectrum and drug resistance of bloodstream infection in patients with acute myeloid leukaemia: a single centre retrospective study. Front. Cell Infect. Microbiol. 14, 1390053 (2024).
Che, M. T. et al. A single-center analysis of pathogenic bacteria distribution and drug resistance in bacterial bloodstream infections among patients with hematological diseases. Zhonghua Xue Ye Xue Za Zhi 45(10), 937–943 (2024).
Piukovics, K. et al. Evaluation of bloodstream infections during chemotherapy-induced febrile neutropenia in patients with malignant hematological diseases: single center experience. Eur. J. Microbiol. Immunol. (Bp). 5(3), 199–204 (2015).
Liu, C. G. & Liao, A. J. Feature analysis of different neutrophil levels on the distribution of pathogens in bloodstream infection in patients with hematologic malignancy. Zhonghua Xue Ye Xue Za Zhi 44(10), 857–860 (2023).
Guo, Q. et al. Analysis of pathogenic bacterial spectrum, drug resistance and risk factors for mortality of bloodstream infection in patients with hematologic diseases. Zhongguo Shi Yan Xue Ye Xue Za Zhi 31(5), 1556–1562 (2023).
Zhang, D. et al. The correlation between inflammatory indicators related to blood routine and bloodstream infections in blood disease patients with neutropenia. J. Clin. Hematol. (China) 38(06), 472–476 (2025).
Xie, W. et al. Levels of PCT, NLR, SAA and PLR of newborns with bloodstream infection and their values in early diagnosis. Chin. J. Nosocomiol. 12, 1841–1845 (2025).
Ossenkoppele, G. & Löwenberg, B. How I treat the older patient with acute myeloid leukemia. Blood 125(5), 767–774 (2015).
Baumgartner, A. et al. Association of nutritional parameters with clinical outcomes in patients with acute myeloid leukemia undergoing haematopoietic stem cell transplantation. Ann. Nutr. Metab. 69(2), 89–98 (2016).
Acknowledgements
NO.
Funding
NO.
Author information
Authors and Affiliations
Contributions
Yabin Chen wrote the manuscript and collected data; Jing Jing wrote the mian manuscript and revised the paper; Dayan Zhang collected clinical data and performed the experiments; Tingjin Zheng and Chengyong Huang analysed data; Zhishan Zhang designed the experiments; Huidan Chen revised the paper. Yabin Chen, Jing Jing and Dayan Zhang contributed equally to this work and share first authorship. Zhishan Zhang and Huidan Chen shared co corresponding authors. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
The study was approved by Ethics Committee of Quanzhou First Hospital (China), the protocol number is [2025] K218. All participants provided written informed consent. And all methods were performed in accordance with Declaration of Helsinki.
Consent for publication
Not applicable.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chen, Y., Jing, J., Zhang, D. et al. Inflammatory markers predict bloodstream infections risk in hematological malignancy patients with chemotherapy induced neutropenia. Sci Rep 15, 40013 (2025). https://doi.org/10.1038/s41598-025-23770-w
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-23770-w