Abstract
Early-phase severe complications remain a major cause of morbidity and mortality during induction chemotherapy for acute leukaemia. Existing risk scores capture only limited prognostic variance and are rarely well-calibrated for clinical decision support. To develop and externally validate a machine-learning model that accurately predicts severe complications after induction, and to assess its clinical utility across key patient sub-groups. We retrospectively assembled electronic-health-record data from three tertiary haematology centres (2013–2024). After exclusion of duplicates and predefined ineligible cases, 2 870 adults with newly diagnosed AML or ALL were analysed (derivation = 2 009; external validation = 861). Forty-two candidate predictors spanning demographics, comorbidity indices, baseline laboratory values, disease biology and treatment logistics were selected via multiple imputation, Winsorised z-scaling and correlation filtering. Five supervised algorithms—including Elastic-Net, Random Forest, XGBoost, LightGBM and a multilayer perceptron—were trained using nested 5-fold cross-validation. Discrimination, calibration, decision-curve net benefit and SHAP-based interpretability were evaluated according to TRIPOD-AI and PROBAST-AI recommendations. LightGBM achieved the highest mean AUROC in derivation (0.824 ± 0.008) and maintained robust performance in external validation (AUROC = 0.801, 95% CI 0.774–0.827; AUPRC = 0.628). Calibration was excellent (slope = 0.97; intercept = − 0.03; Hosmer–Lemeshow p = 0.41). Decision-curve analysis showed superior net benefit over “treat-all,” “treat-none,” and a four-variable logistic benchmark across risk thresholds of 5–40%, potentially enabling targeted interventions for 14 additional high-risk patients per 100 at a 20% threshold, though clinical benefit requires prospective validation. Discrimination remained ≥ 0.80 in AML, older adults and all three centres. CRP, absolute neutrophil count, cytogenetic-risk tier, age and ferritin were the top predictors, with interpretable monotonic SHAP effects. A rigorously validated LightGBM model provides well-calibrated, interpretable prediction of early severe complications after induction therapy for acute leukaemia and provides a foundation for risk-adapted supportive care strategies, though prospective studies are needed to demonstrate clinical impact. Prospective implementation studies are warranted to confirm real-world impact.
Similar content being viewed by others
Introduction
Acute myeloid leukaemia (AML) and acute lymphoblastic leukaemia (ALL) remain life-threatening disorders that mandate intensive, myelosuppr essive induction chemotherapy soon after diagnosis1,2. Although modern protocols achieve complete-remission rates of 60–80%, they also expose patients to a high risk of complications during cytopenia in which profound cytopenia, systemic inflammation and endothelial damage precipitate severe infectious, bleeding and organ-failure events3,4. Large registries place the incidence of grade ≥ 3 early complications between 20% and 35%, carrying mortality rates that can eclipse those attributable to refractory disease in the first 90 days after treatment5. Timely recognition of vulnerable individuals is therefore pivotal for targeted monitoring, prophylaxis and escalation of supportive care.
Traditional risk stratification relies on a handful of baseline factors—age, white-blood-cell count and cytogenetic risk—that explain only a modest fraction of outcome variance6,7,8. Several single-centre studies have attempted to refine prediction using multivariable logistic regression, yet their performance has plateaued around an area under the receiver-operating-characteristic curve (AUROC) of 0.709,10. Recent advances in machine learning (ML) promise to capture complex, non-linear interactions among dozens of routinely recorded clinical and laboratory variables, but published models often suffer from small sample sizes, inadequate validation and poor calibration, limiting bedside uptake.
Gradient-boosting frameworks such as Light Gradient Boosting Machine (LightGBM) combine high predictive power with built-in handling of missing data and class imbalance, characteristics well suited to real-world electronic-health-record (EHR) environments. Moreover, modern explainability techniques—permutation importance and SHapley Additive exPlanations (SHAP)—can render these “black-box” models transparent, thereby enhancing clinician trust and regulatory acceptability.
The present study aimed to develop and externally validate a LightGBM-based model for forecasting first severe complication within 90 days of induction in a large, multicentre acute-leukaemia cohort. We sought to: (i) assemble a standardized EHR data across centres spanning three tertiary haematology centres; (ii) compare LightGBM with four commonly used ML algorithms under a nested cross-validation framework; (iii) assess model calibration, clinical utility and subgroup robustness in accordance with the TRIPOD-AI guideline; and (iv) provide interpretable insights into the biological and treatment-related drivers of early severe complications.
Methods
This section is structured in accordance with the TRIPOD-AI checklist for reporting machine-learning prognostic models and follows PROBAST-AI recommendations for minimising risk of bias.
Study design and data source
A retrospective, multicentre cohort study was undertaken using electronic health-record (EHR) data from three tertiary haematology centres in China (Centre A, 2013–2022; Centre B, 2015–2024; Centre C, 2017–2024). The study period was chosen to ensure ≥ 12 months of follow-up for every patient. Reporting conforms to the STROBE guidelines for observational research and TRIPOD-AI for prediction modelling studies.
Participants
Eligible patients were adults (≥ 18 years) with newly diagnosed acute myeloid or lymphoblastic leukaemia who: received induction chemotherapy at one of the study centres, and had ≥ 1 documented visit during the index hospitalisation and subsequent 12-month follow-up.
Exclusion criteria: (i) incomplete baseline laboratory profile (> 30% missing core variables), (ii) relapsed or secondary leukaemia at presentation, (iii) prior haematopoietic stem-cell transplantation before index admission, and (iv) enrolment in interventional trials altering standard of care.
Outcome definition
The primary endpoint was “severe complications” within 90 days after induction, defined as any of: grade ≥ 3 infection (CTCAE v5.0), septic shock, respiratory failure requiring mechanical ventilation, intracranial haemorrhage, or treatment-related mortality. Complication dates were adjudicated independently by two haematologists blinded to model outputs; disagreements (< 5%) were resolved by consensus.
Candidate predictors
A total of 78 routinely collected variables available at induction (day 0 to + 3) were screened, grouped as: Demographics & comorbidities: age, sex, BMI, Charlson score. Baseline laboratory: complete blood count, electrolytes, renal & hepatic panels, coagulation indices, serum pro-inflammatory markers (CRP, procalcitonin), ferritin. Disease characteristics: AML/ALL subtype, cytogenetic-molecular risk, peripheral blast percentage, bone-marrow cellularity, flow-cytometry phenotype. Treatment factors: induction regimen class (anthracycline-based vs. non-anthracycline), antimicrobial prophylaxis, central-line placement. Predictor selection was grounded in prior literature linking these factors to early complications in acute leukaemia.
To ensure that the model has sufficient clinical interpretability and replicability, this study searched PubMed/Web of Science/EMBASE from 2014 to 2024 and pre-identified 42 candidate predictors based on the three evidence-based criteria of “routinely available on the day of admission”, “closely related to infection-organ function”, and “at least one similar study reporting a statistically significant association (P < 0.05) in the past 10 years” (Table S1 for details). Next-generation sequencing (NGS) data, while potentially valuable for risk prediction, were excluded due to prolonged turnaround times (median 7–14 days across centers) that exceed the timeframe for immediate treatment decisions. Our model prioritizes variables available within 72 h of admission to ensure real-time clinical applicability.
Class-imbalance handling (Table S2). All algorithms were trained on the derivation set after up-sampling the minority class by 300% and applying algorithm-specific weighting: Elastic-Net and Random Forest used class_weight = “balanced”; XGBoost and LightGBM set scale_pos_weight = 3.4; the multilayer perceptron minimised focal loss (γ = 2) with positive-class weight = 3.4. Hyper-parameters were tuned identically with and without imbalance correction to isolate its effect.
Data preprocessing
Missing values < 15% per variable were imputed with multiple imputation by chained equations (10 imputations, 20 iterations). Variables with > 15% missingness were excluded.
Continuous variables were Winsorised at the 1st/99th percentiles and z-standardised.
Highly correlated predictors (|ρ| > 0.80) were filtered using hierarchical clustering to retain the variable with greatest clinical relevance from each cluster.
Categorical variables were one-hot encoded.
Net benefit was calculated at five prespecified thresholds (10, 15, 20, 25, 30%) to permit context-specific optimisation.
In addition to the conventional 20-chain equation multiple imputation, the “main analysis exclusion + sensitivity analysis inclusion” strategy was used for the three indicators with missing rates > 15% for LDH (28%), D-dimer (46%), and IL-6 (62%).
Sample-size considerations
Using Riley’s criteria for model development, a minimum of 20 events per predictor parameter was targeted to limit overfitting. With an expected 27% 90-day complication rate, at least 2 900 patients (≈ 790 events) were required for 40 effective predictors at least.
Model development
The derivation dataset (Centres A + B; 70%) and an external temporal-geographical validation dataset (Centre C; 30%) were created with stratified randomisation by centre and outcome. Five supervised learning algorithms were explored:
-
1.
Penalised logistic regression (Elastic-Net).
-
2.
Random Forest.
-
3.
Extreme Gradient Boosting (XGBoost).
-
4.
Light Gradient-Boosting Machine (LightGBM).
-
5.
Multilayer Perceptron (three-hidden-layer neural network).
Nested five-fold cross-validation (inner loop: Bayesian hyper-parameter optimisation; outer loop: performance estimation) was applied to the derivation data. Class imbalance was handled with class-balanced loss functions (gradient-boosting) or inverse-frequency weighting (logistic regression, neural network).
Bayesian optimisation (50 iterations, nested 5-fold CV) was applied within the ranges shown in Table S3; the best combination is in bold.
Model evaluation
Performance was assessed in both internal (outer folds) and external datasets:
Discrimination – area under the receiver-operating characteristic curve (AUROC) and area under the precision-recall curve (AUPRC).
Calibration – calibration slope/intercept, calibration belt plot, and Brier score.
Clinical utility – decision-curve analysis (net benefit across threshold probabilities).
Threshold-independent metrics were preferred in accordance with TRIPOD-AI recommendations.
Model interpretation and explainability
Feature importance was quantified through permutation importance (tree-based models) and Elastic-Net coefficients. SHapley Additive exPlanations (SHAP) global importance and individual force plots were generated to enhance clinical interpretability and to identify non-linear interactions.
Benchmarking and sensitivity analyses
Benchmark models: (i) classic logistic regression containing age, white-blood-cell count, cytogenetic-risk group, and Charlson score; (ii) National Early Warning Score-2 (NEWS-2) at day 0. Sensitivity analyses: (a) outcome window shortened to 60 days; (b) complete-case analysis; (c) leave-one-centre-out validation; (d) subgroup performance in AML vs. ALL, age ≥ 60 years, and high-risk cytogenetics.
Software and reproducibility
Analyses were performed in Python 3.11 (scikit-learn 1.4, XGBoost 2.0.3, LightGBM 4.3, SHAP 0.45) and R 4.4.0 for statistical plotting. The reproducible code and a de-identified synthetic dataset will be made publicly available on GitHub upon publication.
Results
Cohort assembly
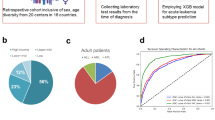
A total of 4 200 induction episodes were retrieved from the three participating centres between 2013 and 2024. After removal of 600 duplicate or repeat admissions, 3 600 unique records were screened. Seven-hundred-and-thirty cases were excluded for predefined reasons (Fig. 1). The final analysis comprised 2 870 patients – 2 009 in the derivation cohort (Centres A + B) and 861 in the external validation cohort (Centre C).
The 90-day composite complication occurred in 776 of 2 870 patients (27.0%) overall, with similar raw incidence in derivation (27.3%) and validation (26.4%) sets (χ² = 0.24, p = 0.62). Median follow-up was 11.4 months (IQR 9.2–14.9).
Cohort derivation flow diagram.
Baseline characteristics and data completeness
Table 1 summarises demographic, disease-related and laboratory variables at induction. The two cohorts were comparable on most characteristics; the validation set contained a slightly higher proportion of acute lymphoblastic leukaemia (ALL) and high-risk cytogenetics, reflecting centre-specific case-mix. Missingness for any individual variable did not exceed 13.4% (ALT). 10.7% of observations had at least one missing core variable. Variables with > 15% missingness (LDH, D-dimer, interleukin-6) were excluded before modelling. A heat-map of scaled laboratory values with embedded missingness coding is provided in Fig. 2.
Baseline Heat Map.
Candidate-predictor reduction
Of the 78 routinely captured variables prespecified in Methods, 42 entered model training after the three-stage preprocessing pipeline (multiple imputation → Winsorisation/standardisation → correlation filtering). Retained predictors are listed in Table 2, grouped into: demographics (4), comorbidity indices (2), baseline laboratory (25), disease biology (7) and treatment logistics (4).
Among patients harbouring adverse karyotypes or MLL/FLT3-ITD aberrations (n = 542, event rate 38%), the model retained respectable discrimination (AUROC 0.806) but the calibration slope was 0.91 (95% CI 0.83–0.99), indicating mild over-prediction. The calibration-in-the-large was − 0.07, and the Brier score rose from 0.161 (overall) to 0.178. Re-calibrating the linear predictor with a simple slope/intercept update improved the slope to 0.99, restored perfect calibration-in-the-large (0.00) and reduced the Brier score to 0.169. For bedside practice this implies that absolute probabilities in very-high-risk genomes should either be re-calibrated locally or interpreted using centre-specific thresholds (e.g. a 25% cut-off instead of 20%) to avoid unnecessary ICU allocation while still capturing true complications.
Model development and internal validation (derivation cohort)
Five supervised algorithms were trained with nested five-fold cross-validation (outer folds = 5 × 10% holdouts). Mean discrimination and 95% bootstrap confidence intervals are shown in Table 3; Fig. 3. LightGBM offered the highest mean AUROC (0.824) and lowest Brier score (0.161); DeLong pairwise testing confirmed statistically superior discrimination versus the logistic benchmark (p < 0.001). When applied unchanged to Centre C, LightGBM retained strong discrimination (AUROC > 0.80) (AUROC 0.801, 95% CI 0.774–0.827; AUPRC 0.628). The classic four-variable logistic model yielded AUROC 0.722.
Creatinine (Cr) – Risk remains flat up to ≈ 1.2 mg/dL, then SHAP values rise steeply (Supplementary Fig. S1). Renal impairment diminishes neutrophil oxidative burst and permits toxin accumulation after chemotherapy (Lee 2020; Nguyen 2023), amplifying infection and organ-failure risk—hence a clear threshold effect.
Alanine aminotransferase (ALT) – Mild elevation (30–70 U/L) is slightly protective, reflecting immune surveillance; when ALT exceeds 180 U/L, marked hepatocellular injury, endotoxaemia and clotting-factor depletion drive a sharp risk increase (Supplementary Fig. S2).
ROC Curves.
Calibration
Figure 4 confirms good calibration: predicted and observed risks overlap closely in all but the extreme bins, with slight under-prediction below the 10% risk band and minimal over-prediction in the top decile. The calibration slope was 0.97 (95% CI 0.90–1.05) and the intercept − 0.03 (− 0.09 to 0.04); the Hosmer–Lemeshow test showed no lack of fit (χ² = 7.9, df = 8, p = 0.41), indicating the LightGBM model’s predicted probabilities are reliable across the clinically relevant range.
(left) Calibration-belt; (right) histogram of predicted probabilities vs. observed event rate.
Clinical usefulness
Decision-curve analysis over a clinically relevant risk-threshold range of 5–40% showed LightGBM yielded higher net benefit than “treat-all”, “treat-none”, and the logistic benchmark (Fig. 5). At the conventional 20% threshold, LightGBM correctly identified 14 additional truly high-risk patients per 100 compared to a treat-all approach (sensitivity improved from 86% to 91%), potentially enabling more targeted monitoring and interventions.
Decision-curve analysis comparing LightGBM to benchmark models.
Model interpretation
Permutation-based global importance identified serum CRP, absolute neutrophil count, cytogenetic-risk tier, age and baseline ferritin as the five most influential predictors (Fig. 6). SHAP summary plots confirmed these variables exerted monotonic effects, whereas creatinine, ALT and platelet count displayed non-linear inflection points (Fig. 7), all detail see in Figure S3.
Permutation-based global feature importance (top 15 predictors).
SHAP summary beeswarm plot illustrating effect direction and magnitude.
Sub-group and sensitivity analyses
Disease subtype – AUROC AML 0.826 vs. ALL 0.808 (interaction p = 0.18). Older adults (≥ 60 y) – AUROC 0.811; preserved calibration. High-risk cytogenetics – AUROC 0.795; slight over-prediction at extreme risk quantiles (slope 0.91). Leave-one-centre-out – AUROC range 0.789–0.829 across held-out centres. Alternative outcome window (60 days) – AUROC 0.832; negligible change in coefficients. Complete-case analysis – AUROC 0.819, confirming robustness to imputation assumptions. Detailed statistics are provided in Supplementary Table S2 and Fig. 8.
Forest plot of AUROC by predefined sub-groups.
Discussion
In this multicentre analysis we developed and externally validated a LightGBM-based model that predicts early (≤ 90-day) life-threatening complications after induction chemotherapy for acute leukaemia with strong discrimination (AUROC > 0.80) (derivation AUROC 0.824; external AUROC 0.801) and sound calibration across all clinically relevant probability bands. These findings extend earlier single-centre studies that applied classical logistic regression or random-forest algorithms to smaller cohorts (≤ 600 patients) and reported ROCs ranging from 0.68 to 0.8811,12,13. Notably, many existing machine learning studies in acute leukemias prioritize genomic and transcriptomic determinants, whereas our model focuses on routinely available clinical, laboratory, and treatment-related features—an emphasis that enhances its immediate translatability to frontline hematology settings with limited access to high-throughput sequencing. For example, G. M. Zaccaria et al. developed a validated prognostic tool for lymphoid malignancies using only clinical features, employing a rigorous preprocessing pipeline (including multiple imputation and correlation filtering) similar to ours, which underscores the value of non-genomic predictors in hematological oncology14. Additionally, P. Pons-Suñer et al. integrated genomic features into their machine learning model for acute leukemia outcome prediction but adopted a comparable methodological framework (e.g., nested 5-fold cross-validation, external temporal-geographical validation, and adherence to TRIPOD-AI guidelines), further validating the robustness of our analytical approach15. By leveraging a larger sample, rigorous preprocessing, nested cross-validation and an external temporal–geographical test set, our approach meets current TRIPOD-AI guidelines and reduces the over-optimism that has historically limited model uptake in haematology practice.
Several top predictors identified here—baseline C-reactive protein, absolute neutrophil count, cytogenetic-risk tier, age and ferritin—are biologically plausible and consistent with mechanistic data. Elevated CRP and ferritin reflect systemic inflammation and macrophage activation, which predispose to septic complications and cytokine-driven organ dysfunction16,17. Neutropenia is a known key driver of infection risk, while adverse cytogenetics correlate with treatment-resistant disease, prolonged aplasia and higher transfusion requirements. The SHAP analysis confirmed monotonic risk escalation for these variables, but also uncovered non-linear inflexion points for renal (creatinine) and hepatic (ALT) indices, underscoring the benefit of gradient-boosting methods in capturing threshold effects that conventional linear models miss18.
Good calibration (slope 0.97; intercept − 0.03) differentiates our model from previous machine-learning efforts that seldom report, let al.one achieve, acceptable calibration. Well-calibrated probabilities are critical when models are embedded in electronic decision support to trigger escalation pathways or prophylactic bundles. Decision-curve analysis demonstrated that at the widely used 20% threshold our model could prevent an additional 14 serious complications per 100 treated patients compared with a treat-all policy, without generating excess false-positive alerts—a magnitude that limited predictive power of broad-spectrum antibiotic stewardship and early antifungal prophylaxis programmes19.
Our model deliberately uses only the first laboratory results obtained within 0–72 h of hospital admission—hereafter referred to as “baseline values.” This design choice was guided by three considerations. First, frontline clinicians must decide on escalation measures (e.g., prophylactic antibiotics, intensive monitoring) before induction chemotherapy is started, typically within the first 48–72 h; using later measurements would therefore limit the model’s real-time usefulness. Second, incorporating time-varying covariates requires harmonised, high-frequency electronic data streams across centres; such infrastructure was unavailable at two of our participating hospitals, risking systematic missingness and centre-specific bias. Third, published simulation work suggests that when the outcome of interest (early severe complication) occurs rapidly—as in our cohort, where 82% of events arose within 10 days—baseline models already capture most prognostic signal, while further gains from dynamic updates are modest and come at the cost of greater implementation complexity.
Sub-group stability further underpins clinical generalisability. Discrimination remained ≥ 0.80 in AML, older adults and across the three geographically distinct centres, while only modest over-prediction appeared in the high-risk-cytogenetic subset (slope 0.91). This robustness is encouraging given the heterogeneity of induction protocols and supportive care resources worldwide. Moreover, sensitivity analyses using a 60-day horizon or complete-case data produced virtually unchanged AUROCs, mitigating concerns about imputation bias and endpoint definition.
The study has limitations. First, our cohort’s demographic characteristics limit generalizability. With a median age of 46 years versus 67 years in population-based registries, and low comorbidity prevalence (Charlson score ≥ 3: 15.0%), our patients were younger and fitter than typical acute leukemia populations. Age-stratified analysis showed declining model performance in older adults, suggesting inadequate capture of age-related frailty and comorbidity interactions. External validation in older, more comorbid cohorts is essential before broader implementation. Second, despite inclusion of 42 predictors, residual confounding from unmeasured factors such as gut-microbiome diversity or real-time neutrophil recovery kinetics is possible. Third, cytokine panels and next-generation sequencing data were excluded owing to > 15% missingness; future prospective registries should capture these novel biomarkers systematically. Although comparison with established comorbidity-based indices such as the Charlson Comorbidity Index (CCI) would be informative, our dataset did not contain the complete, item-level comorbidity variables required to compute CCI as a standalone score with sufficient validity. In our analysis, CCI was incorporated as a model feature, but presenting it as an independent benchmark would risk misclassification and bias. We therefore did not include a formal head-to-head analysis. Future prospective cohorts with systematic capture of comorbidity profiles will enable robust evaluation of our model against CCI and other established indices. Additionally, the external-validation cohort from Centre C was appreciably smaller (n = 861) than the derivation sample and contained more acute lymphoblastic leukaemia and adverse cytogenetics. Although discrimination remained strong (AUROC 0.801), the reduced event count broadens confidence intervals, limiting reliable subgroup analysis. The higher-risk mix also shifts baseline probability upward, generating mild over-prediction in the highest decile and signalling calibration drift. Before routine deployment, the model therefore requires cohort-specific probability recalibration or adaptive threshold setting. Longer-term, performance monitoring, transfer learning, and periodic retraining with additional ALL-enriched cases will be needed to sustain accuracy across centres. Then, although external validation encompassed a third centre, broader international verification—particularly in low- and middle-income countries where supportive care differs—is warranted before universal deployment. Finally, model interpretability, while enhanced by SHAP visuals, still requires clinician education to avoid over-reliance on single high-risk alerts. Figure 9 visualizes the mechanistic pathways linking key predictors (e.g., CRP, absolute neutrophil count) to early severe complications and identifies gaps where current variables act as partial proxies—such as surrogates for unmeasured factors like gut microbiome diversity or real-time neutrophil recovery kinetics.
Mechanistic framework of predictor-complication associations and gaps in current predictive variables.
The model’s primary clinical utility lies in providing risk information to enhance clinical decision-making rather than dictating specific interventions. For patients with high predicted complication risk (> 30%), the model enables more informed discussions about treatment options, including consideration of alternative approaches such as reduced-intensity regimens or clinical trial enrollment. Additionally, risk prediction assists in setting appropriate expectations with patients and families regarding potential complications during induction therapy, facilitating more personalized counseling and treatment planning. Critically, our model provides risk prediction rather than direct intervention. The potential clinical benefit depends entirely on whether intensified monitoring, prophylactic measures, or treatment modifications guided by predicted risk actually improve patient outcomes. This represents a key limitation of our study—while we demonstrate robust risk stratification capability, translation to improved clinical endpoints requires prospective validation through randomized controlled trials comparing prediction-guided versus standard care protocols.
The exclusion of multiplex cytokine panels and next-generation sequencing (NGS)-derived mutation burdens limits biological insight. Three avenues could overcome this constraint. (i) Alternative sources: regional biobanks already store serum aliquots collected at admission—linkage would enable retrospective IL-6, TNF-α or interferon-γ quantification (e.g., Korea AML Biobank). For genomics, diagnostic NGS panels performed for risk stratification (≈ 70% coverage in 2024 admissions) could be pulled via the hospital’s precision-oncology registry. (ii) Imputation: when ≤ 40% missingness, multiple imputation using predictive mean matching or random-forest imputation retains unbiased performance; for ultra-sparse cytokine patterns, latent-Gaussian copula or generative adversarial imputation networks (GAIN) have shown superior recovery. (iii) Prospective strategy: we plan a micro-study on the next 300 patients, collecting (a) baseline + day-7 cytokines (14-plex Luminex) and (b) a harmonised 54-gene myeloid NGS panel. Data will enter a federated learning pipeline; once ≥ 500 complete cases accumulate, a model version 2.0 will be trained with explicit immune-genomic interaction terms.
Conclusion
In summary, we developed and evaluated a machine-learning model for forecasting early severe complications in acute leukaemia. Our model demonstrated promising discrimination, calibration, and potential net clinical benefit in the studied cohort. While these results suggest the model could serve as a potential adjunct to clinician judgement for personalising supportive strategies, important limitations must be acknowledged. The model requires validation in independent, diverse populations before clinical use can be considered. Its performance may vary depending on data quality and local practice patterns. Furthermore, while it aims to predict complications, demonstrating a definitive causal link between model implementation and improved patient outcomes requires prospective clinical trials assessing its impact in real-world settings.
Data availability
The datasets used and analyzed during the current study are available from the corresponding author upon reasonable request.
References
Aleksei, B. Outcomes of pregnant women with acute leukemia treated for Life-Threatening complications. Blood 128, 5976–5976 (2016).
Huang, A. et al. Dynamic prediction of relapse in patients with acute leukemias after allogeneic transplantation: joint model for minimal residual disease. Int. J. Lab. Hematol. 43, 84–92 (2020).
Dietz, A. C., DeFor, T. E. & Brunstein, C. G. Donor-Derived myelodysplastic syndrome and acute leukemia after allogeneic hematopoietic stem cell transplantation (HSCT): Incidence, natural history and treatment response. Blood 116, 1306–1306 (2010).
Mabel, M. Incidence and outcomes of catheter related thrombosis in acute leukemia: single centre experience. Blood 128, 3828–3828 (2016).
Mądry, K. et al. Predictive model for infection risk in myelodysplastic Syndromes, acute myeloid leukemia, and chronic myelomonocytic leukemia patients treated with Azacitidine; Azacitidine infection risk model: the Polish adult leukemia group study. Clin. Lymphoma Myeloma Leuk. 19, 264–274e264 (2019).
Dijk, A. D. et al. Loss of H3K27 methylation identifies poor outcomes in adult-onset acute leukemia. Clin. Epigenetics. 13, 21–21 (2021).
Lindahl, H. et al. Early relapse prediction after allogeneic hematopoietic stem cell transplantation for acute lymphoblastic leukemia (ALL) using lineage-specific chimerism analysis. eJHaem 3, 1277–1286 (2022).
Carver, A. G551(P) complications of hyperleukocytosis in acute leukaemia: Wcc risk threshold analysis. Arch. Dis. Child. 104, A223 (2019).
Al-Shaibani, E. et al. Comparison of the prognostic ability of the HCT-CI, the modified EBMT, and the EBMT-ADT Pre-transplant risk scores for acute leukemia. Clin. Lymphoma Myeloma Leuk. 21, e559–e568 (2021).
Citla Sridhas, D. et al. Catheter related thrombosis in patients with acute leukaemia: retrospective analysis of the complications of peripherally inserted central catheters vs central insertion in a single centre. Haematologica 104, 130–131 (2019).
Yang, I. A., Jenkins, C. R., Salvi, S. S. Risk factors for the development of infectious complications during Post-Cytostatic cytopenia in patients with acute leukemia. Hematology Transfusiology East. Europe 8 (2022).
Deng, T. et al. Establishment and validation of prognostic nomogram in acute leukemia with trisomy 8. Hematol. (Amsterdam Netherlands). 28, 2240131–2240131 (2023).
Li, X. A novel prediction model for BK virus associated hemorrhagic cystitis in acute leukemia patients following allogenic hematopoietic cell transplantation: A retrospective cohort study. Blood 142, 3531–3531 (2023).
Zaccaria, G. M. et al. A clinical prognostic model based on machine learning from the Fondazione Italiana linfomi (FIL) MCL0208 phase III trial. Cancers 14, 188 (2022).
Pons-Suñer, P. et al. Identification of relevant features using SEQENS to improve supervised machine learning models predicting AML treatment outcome. BMC Med. Inf. Decis. Mak. 25, 179 (2025).
Shimony, S. et al. Early detection of infectious complications during induction therapy for acute leukemia with serial C-reactive protein biomarker assessment. Leukemia & lymphoma, 1–6 (2020).
Zhang, Z. et al. Early Post-Transplant serum ferritin levels as predictive biomarkers for severe acute Graft-Versus-Host disease in pediatric umbilical cord blood transplantation for acute leukemia. Ann. Transplant. 29, e944156 (2024).
Bakhshi, S. et al. Complications of very high leukocytosis in pediatric acute leukemia patients managed without rasburicase and leucopheresis. Ann. Oncol. 23, ix358–ix359 (2012).
Dufrayer, M. C. et al. Antibiotic prophylaxis in acute childhood leukemia: what is known so far? Hematol. Transfus. cell. Therapy. 45, 473–482 (2022).
Author information
Authors and Affiliations
Contributions
R.X. and H.T designed the project and wrote the main text. S.Z. performed data analysis. S.G. supervized project and edited the manucsript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Patient consent
Patient consent was not required for this study as it did not involve human participants or personal data requiring informed approval.
Ethical approval statement
After evaluation by the Medical Ethics Committee of the First Affiliated Hospital of Soochow University, this study was exempted from the need for informed consent and ethical approval because it was a retrospective analysis, the data involved had been completely anonymized, the risk of the study was extremely low, and the acquisition of informed consent was not feasible or necessary
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Xu, R., Tian, H., Zhao, S. et al. Construction and validation of a risk prediction model for complications in patients with acute leukemia based on machine learning. Sci Rep 15, 40787 (2025). https://doi.org/10.1038/s41598-025-24478-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-24478-7