Abstract
Endometrial cancer (EC) is the most common gynecologic malignancy, with a steadily increasing incidence worldwide. Abnormal vaginal bleeding, a hallmark symptom, enables early diagnosis, which is critical for improving clinical outcomes. Pelvic magnetic resonance imaging (MRI) serves as the primary imaging modality for EC evaluation, offering detailed visualization of endometrial and myometrial invasion.This study proposes DCS-Net, a multi-task deep learning framework for the automated detection and staging of early-stage EC in MRI. The framework incorporates an advanced object detection module to accurately localize and crop the uterine region, followed by a convolutional neural network for staging classification.Experimental results show that the detection module achieves high localization performance, and the classification network reaches an accuracy of 90.8%. The region-focused approach improves staging accuracy by 5% compared to direct classification using unprocessed images.These results underscore the potential of DCS-Net to improve diagnostic efficiency and accuracy in alignment with clinical workflows. Future work will explore the integration of multi-parametric MRI data to further enhance diagnostic performance and address broader clinical needs.
Similar content being viewed by others
Introduction
Endometrial cancer (EC) is the most common gynecological malignancy in developed countries, with its global incidence on the rise1. According to the latest data from the National Cancer Institute (NCI) in 2022, the mortality rate from uterine cancer among women of all races and ethnicities increased by an average of 1.8% annually between 2010 and 2017. Factors such as socioeconomic changes, rising obesity rates, declining fertility rates, and the use of exogenous estrogen contribute to the increasing incidence of EC among the younger generation worldwide2. Endometrial cancer originates from the glandular cells of the endometrium, initially developing in the uterine lining and subsequently invading the myometrium3. According to the surgical staging system for endometrial cancer established by the International Federation of Gynecology and Obstetrics (FIGO) in 20094, endometrial cancer is classified into four stages: Stage I, Stage II, Stage III, and Stage IV, with each stage further subdivided into specific subcategories. Abnormal uterine bleeding is a primary clinical manifestation of endometrial cancer, facilitating the early detection of EC. Notably, the five-year survival rate for early-stage EC exceeds 95%5. Therefore, early screening, accurate diagnosis, and timely treatment are crucial for preventing disease progression and improving patient survival rates. Accurate staging of endometrial cancer is essential for developing individualized treatment plans6. Precise staging not only helps standardize clinical treatment protocols but also significantly enhances patient prognosis.
Magnetic Resonance Imaging (MRI) is an advanced medical imaging technology that utilizes strong magnetic fields and harmless radio waves to generate high-resolution images of the human body’s internal structures7. It is highly favored in early screening due to its exceptional soft tissue contrast, particularly in detecting early-stage endometrial cancer and other pathological changes, thereby enabling precise visualization of subtle lesions and facilitating early diagnosis and effective treatment planning8. According to the 2022 edition of the Chinese National Health Commission’s guidelines for the diagnosis and treatment of endometrial cancer, pelvic MRI scanning is recommended as the primary imaging modality. It provides clear delineation of the endometrial and myometrial structures, facilitating detailed assessment of lesion size, location, depth of myometrial invasion, involvement of cervix and vagina, extracorporeal extension to the uterus, vagina, bladder, and rectum, as well as tumor spread within the pelvic, retroperitoneal, and inguinal lymph nodes9. However, manual interpretation of MRI scans suffers from subjectivity, variability in expertise, and high time and cost demands, leading to potential diagnostic errors. Despite the irreplaceability of human expertise in certain scenarios, these inherent limitations underscore the increasing recognition and adoption of automated and computer-assisted technologies in medical image analysis.
In recent years, significant advancements and importance of deep learning technology in the field of medical imaging have been demonstrated. Particularly, convolutional neural networks (CNNs) play a crucial role in medical image analysis10. CNNs leverage convolutional operations and hierarchical connections to effectively extract and optimize image features, thereby enhancing understanding of complex medical data and diagnostic accuracy11. Training effective medical image models necessitates large, precisely annotated datasets and rigorous data preprocessing and augmentation12. Deep learning technology has made significant progress in medical image classification13, detection14, and segmentation tasks15, not only improving diagnostic accuracy but also reducing reliance on manual intervention, thereby significantly enhancing clinical productivity for radiologists. Compared to traditional computer-aided diagnostic systems, deep learning models can directly extract useful features from data without the need for manual feature selection steps, thereby demonstrating clear advantages and potential applications in complex medical image analysis16. This study focuses on the application of deep learning technology in assisting the diagnosis of early-stage endometrial cancer using MRI images. The key to diagnosing early-stage endometrial cancer lies in accurately capturing subtle lesion features, and deep learning, with its outstanding image analysis capability, can effectively extract and identify crucial features from MRI images17.
Recent series of studies have underscored significant advancements of deep learning in MRI imaging. Similarly, early research contributions have laid a solid foundation for further development in MRI analysis of endometrial cancer. Mao et al. developed a deep learning-based automated segmentation model that achieved high accuracy in staging early endometrial cancer by analyzing the ratio of tumor area to uterine area in MRI images, with AUCs of 0.86, 0.85, and 0.94 for axial T2WI, axial DWI, and sagittal T2WI images, respectively18. Feng et al. proposed a multitask framework named ECMS-Net, employing Transformer algorithm for sequence classification and U2-Net model for precise automatic segmentation of early endometrial cancer. Results demonstrated sequence classification accuracy of 0.985 and maxF1 value of 0.970 for tumor segmentation, significantly enhancing diagnostic accuracy and efficiency, thereby alleviating clinical workload19. Aiko et al. compared convolutional neural network (CNN)-based deep learning models with radiologists in diagnosing endometrial cancer, showing comparable AUCs of 0.88–0.95 on single and combined image sets. Integration of different image types further improved CNN diagnostic performance on specific single image sets20. A study by Stanzione evaluated a machine learning model based on MRI radiomics for identifying deep myometrial invasion (DMI) in endometrial cancer patients, exploring its clinical applicability. The model achieved accuracies of 86% in cross-validation and 91% in final testing, with AUCs of 0.92 and 0.94, respectively, significantly enhancing diagnostic performance compared to radiologists with and without ML assistance21. Mainenti et al. explored the application of radiomics and machine learning in stratifying risk of endometrial cancer based on MRI. Using features extracted from T2-weighted images, the SVM algorithm-based model achieved accuracies of 0.71 and 0.72 on training and testing sets, demonstrating good performance and versatility in identifying low-risk endometrial cancer patients22. Chen et al. evaluated the diagnostic performance of deep learning models in determining myometrial invasion depth in MRI of endometrial cancer based on T2WI. They first trained a detection model using YOLOv3 algorithm to localize lesion areas and then inputted detected regions into a deep learning network-based classification model for automatic recognition of myometrial invasion depth23. Dong et al. employed U-Net with ResNet34, VGG16, and VGG11 encoders to establish a CNN-based AI model, comparing its experimental results on contrast-enhanced T1WI and T2WI slices with diagnoses by radiologists. The AI’s interpretation of contrast-enhanced T1WI was more accurate (79.2%) compared to radiologists’ (77.8%), while accuracy on T2WI was lower (70.8%). Despite no significant difference in diagnostic accuracy between AI and radiologists on T1WI and T2WI images, AI tended to provide more erroneous interpretations in patients with benign uterine leiomyomas or polypoid tumors24. Drawing insights from prior research, we systematically outline the methodology to guide our study. Inspired by previous work, this paper comprehensively details the entire process including data collection25, data preprocessing methods, model selection26, detailed model analysis27, visualization of experimental processes28, validation using external data, clinical validation by healthcare professionals, and interpretability analysis of the model. Inspired by these advancements, this study focuses on applying deep learning techniques to assist in diagnosing early-stage EC from MRI scans. The key challenge in diagnosing early-stage EC lies in accurately identifying subtle lesion features, and deep learning models have shown significant promise in extracting and analyzing these critical features. This paper aims to outline a comprehensive methodology for training and validating deep learning models for early EC diagnosis, including data collection, preprocessing, model selection, and evaluation.
Materials and methods
Figure 1 represents the architecture of this study.
The architecture of this study.
To meet the format requirements for input datasets in neural network models, we first preprocessed early endometrial cancer MRI images in DICOM format. DICOM is an international standard used for storing, transmitting, and processing medical imaging information, ensuring interoperability and data integrity across different devices and systems29. We have named this proposed data processing method “DRIL.” Firstly, we utilized RadiAntViewer to visualize the source data included in the experiment. This step facilitates close collaboration between physicians and software developers in non-hospital environments, enabling in-depth analysis of image data characteristics, which provides a foundation for subsequent annotation and processing. Next, experienced radiologists used ITK-SNAP to meticulously annotate the images included in the study and saved the processed images30. During this process, we placed particular emphasis on the anonymization of image data. This step ensures that while the overall image information is maximally preserved, no patient-related data is included in the processed images, thereby effectively safeguarding patient privacy. Lastly, the radiologist employed Labelimg software to annotate the images in a format suitable for detection datasets, forming a detection data set compatible with the neural network model31. These annotated data will be used for model training, ensuring that the model can accurately identify and analyze early endometrial cancer MRI images32. Through these steps, the DRIL method not only guarantees the scientific rigor and standardization of data processing but also provides high-quality data support for neural network model training, ensuring the final model’s accuracy and reliability.
In the image processing tasks and model training, since early-stage endometrial cancer lesions are typically confined to the uterus, the first step is to focus on the uterine region within the entire pelvic scan image. To achieve this, we initially employed the YOLOv5 model to detect and crop the uterine region. This step enables accurate identification of the uterine area and its subsequent cropping for further analysis. Subsequently, we utilized the ResNet34 classification model to automate the staging of endometrial cancer.
Data set and processing
In the initial phase of our study, we provided a detailed description of the case screening procedures and dataset generation process, as shown in Fig. 2. To ensure the accuracy and consistency of the data, we outlined the specific processes for image annotation and data preservation, ensuring that the generated dataset seamlessly integrates with our computational model. Initially, we collected DICOM source data from 289 patients who underwent early-stage endometrial cancer MRI examinations at the Second Affiliated Hospital of Fujian Medical University between March 2018 and October 2022. Under the guidance of physicians and in strict accordance with inclusion and exclusion criteria, we ultimately selected 235 patients for this study. We applied a self-designed DRIL method for data preprocessing, generating 707 sagittal T2WI MRI images of early-stage endometrial cancer, which formed the preliminary experimental dataset. Subsequently, two senior physicians professionally annotated the images, clearly identifying the uterine region and differentiating cancer stages. Through this effort, we constructed a key region detection dataset consisting of 1,414 images, as well as a staging dataset containing 600 images, which was standardized according to the model training proportion, both Stage IA and Stage IB categories were balanced, each containing 300 T2-weighted MRI images. High-quality and standardized datasets are the foundation and critical guarantee for the success of model experiments.
Flowchart of data set selection.
The original imaging data consisted of T2-weighted MRI scans in DICOM format. All images were preprocessed using standard clinical software and exported to JPG format, during which pixel intensities were standardized to the 0–255 range and window width and level were fixed, resulting in consistent spatial resolution across the dataset. No additional intensity normalization, resampling, or windowing operations were performed. For the classification task, images were directly fed into the network without further augmentation. For the detection task, default augmentation strategies provided in the official implementation, including Mosaic, random flip, and color jitter, were employed without additional modifications. Early-stage endometrial cancer cases were annotated according to the FIGO staging system: Stage IA corresponds to tumors confined to the uterine body with < 50% myometrial invasion, while Stage IB corresponds to tumors confined to the uterine body with ≥ 50% myometrial invasion.To further validate the model’s generalization ability, we included data from 48 early-stage endometrial cancer patients from the First Affiliated Hospital of Quanzhou, Fujian Medical University, using the GE system for external validation, generating 96 sagittal T2WI MRI images. This step aimed to assess the model’s applicability and stability in different clinical settings. The study was approved by the Ethics Committee of the Medical College of Huaqiao University, and written consent was obtained, with the approval number M2024009. All participants provided written informed consent to participate in the study, and all methods were performed in accordance with relevant guidelines and regulations.
Uterine ROI detection
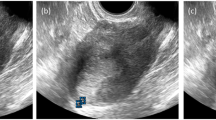
Based on the practical requirements of medical image diagnostics, we selected the YOLO series object detection model to precisely localize and crop the uterine region within pelvic scan images, laying the foundation for subsequent automatic staging tasks33. Given that early-stage endometrial cancer lesions are typically confined to the uterine cavity, subsequent staging detection will be performed based on the identified uterine ROI (region of interest), thereby enabling a more accurate and automated staging process. As depicted in Fig. 3, the schematic architecture of the renowned YOLOv5 detection model is intricately illustrated34. YOLOv5 is an advanced object detection model based on Convolutional Neural Networks (CNN). Developed upon the foundational architecture of the YOLO (You Only Look Once) series, it maintains the series’ hallmark efficiency and accuracy. YOLOv5 operates on the core principle of predicting multiple bounding boxes and class probabilities simultaneously within a single inference pass of the neural network. This model features several key enhancements, including a rapid inference speed, a modular design, an optimized loss function, sophisticated data augmentation techniques, and a streamlined training process. The Backbone of YOLO is examined starting from the network configuration file, analyzing the composition and propagation process of feature maps, and breaking down the components such as CSP, CBS, SPPF, and Bottleneck35. For the Bottleneck module, it is introduced with ResNet36, detailing the differences between ResNet and CSP. The Neck of YOLO employs the classic FPN + PAN design, which integrates top-down and bottom-up feature extraction methods.In object detection frameworks, the Feature Pyramid Network (FPN) and Path Aggregation Network (PAN) are essential modules for enhancing multi-scale feature representation. FPN employs a top-down pathway with lateral connections to progressively transmit high-level semantic information to shallow layers, thereby improving the recognition of small objects. Conversely, PAN introduces a bottom-up information flow that feeds rich spatial details from shallow layers back to deeper layers, further refining the localization of large objects. By integrating FPN and PAN, YOLOv5 constructs a bidirectional feature fusion network that enables efficient interaction between semantic and spatial information, leading to significantly improved detection performance across objects of varying scales.
The architecture of detection networks and prediction of key uterine anatomical structures.
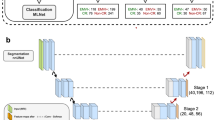
Cancer staging
After accurately detecting the uterine region, we used the Save-crop method to extract the uterine region from the complete pelvic scan images and applied the classic classification model ResNet34 for automated cancer staging36. The Save-crop method is an advanced image enhancement and dataset preprocessing technique, designed to facilitate the precise cropping and saving of target regions within input images during the training process. This method enables the accurate extraction of regions of interest (e.g., specific objects or target structures) and saves them as individual image files. Such preprocessing is crucial for subsequent data analysis, augmentation, and the independent training of object detection models, ensuring higher quality and more focused datasets for improved model performance. ResNet34, a classic architecture within the ResNet (Residual Network) series, is widely recognized as a benchmark model in deep convolutional neural networks. The model architecture is shown in Fig. 4. It addresses key challenges in training deep networks, such as vanishing gradients and performance degradation, which often arise as network depth increases. As a relatively shallow variant of the ResNet family, ResNet34 features a 34-layer structure, striking an optimal balance between model performance and computational efficiency. Its hallmark innovation is the introduction of a residual learning framework, leveraging skip connections within residual blocks. This mechanism effectively mitigates the vanishing gradient problem, enhances gradient flow, and substantially improves both the training efficiency and stability of deep neural networks. These characteristics make ResNet34 particularly well-suited for applications in medical image analysis, where robust model performance and computational efficiency are crucial for tasks such as disease classification and lesion detection.
The architecture of cancer staging network.
Evaluation metrics
To evaluate the effectiveness of the model, we selected precision, recall, mean average precision at IoU threshold 0.5 (mAP@0.5), and mean average precision across IoU thresholds from 0.5 to 0.95 (mAP@0.5:0.95) as the evaluation metrics for detecting uterine ROI in early endometrial cancer. Precision, recall, mAP@0.5, mAP@0.5:0.95, accuracy, and specificity were calculated using Eqs. (1)–(5), respectively. Precision measures the accuracy of the model’s positive predictions. Recall indicates the model’s ability to correctly identify positive instances from all actual positive instances. Average Precision (AP) is the area under the Precision-Recall (P-R) curve, and mean Average Precision (mAP) is the mean AP across all categories, with a maximum value of 1. Accuracy represents the ratio of the number of correct predictions to the total number of samples. Specifically, accuracy measures the proportion of correct predictions (including both positive and negative classes) out of all predictions made by the model. In medical image analysis, accuracy is commonly used to evaluate the overall classification performance of the model in distinguishing between lesion and non-lesion regions. These evaluation metrics collectively provide a comprehensive assessment of the model’s performance in the task of detecting uterine ROI in early endometrial cancer.
Experiments set
In our experiments, we utilizing pretrained weights significantly reduces the required training time and computational resources, while enhancing the initial performance and stability of the models. For tasks with limited data, pretrained weights effectively mitigate overfitting and improve the generalization capabilities of the models. The experiment is conducted using a 12th Gen Intel(R) Core (TM) i5- 12400F CPU and an NVIDIA GeForce RTX3050 GPU with 16 GB video memory and 32 GB RAM. The computer operates on a 64-bit Windows 10 system with PyCharm as the programming software and uses Python 3.7 as the programming language.For the object detection task (YOLOv5-n), we strictly followed the standard YOLOv5 training protocol. The model was trained using the SGD optimizer with an initial learning rate of 0.01, coupled with a cosine annealing learning rate scheduler (final learning rate 0.01) and a 3-epoch linear warmup. Training was performed for 100 epochs with a batch size of 8, and the model weights achieving the highest mean average precision (mAP) on the validation set were retained. For the image classification task (ResNet-34), the Adam optimizer was used (learning rate 0.0001, weight decay 0.01) with a batch size of 8, trained for 150 epochs. Early stopping was triggered if the validation accuracy did not improve for 15 consecutive epochs. This standardized experimental setup ensured the reproducibility of our results while maintaining consistency with prevailing benchmarks in the field.
Results
Detection performance
In the comparative model experiments, we tested classic deep learning detection models provided by official open-source codes, including YOLOv5, YOLOX37, YOLOv838, and the two-stage target detection network Faster R-CNN39. Our evaluation of the experimental results revealed that YOLOv5 exhibited the most outstanding performance in this experimental task, with specific data presented in Table 1. The YOLOv5n model achieved an mAP@0.5 score of 0.934, with a training time of only 27.07 s per epoch. After completing the model training, we utilized external data for prediction, accurately identifying the critical uterine regions in the images and providing confidence score evaluations, partial prediction results are shown in Fig. 3. Through comparative experiments, we identified YOLOv5 as the most effective model. This model demonstrated superior performance in accurately identifying the uterine ROI within the uterine region.
Figure 5 provides a comprehensive visualization of the model’s performance evolution throughout the training process, capturing the fluctuations of evaluation metrics and loss across successive training epochs. The Precision-Recall (PR) curve elucidates the trade-off between precision and recall at varying thresholds, with higher recall typically correlating with lower precision, and vice versa. Furthermore, the visualized prediction results highlight the model’s accuracy in detecting the uterine region. Collectively, these visualizations offer an insightful overview of the model’s training trajectory, facilitating a thorough assessment of the classification model’s effectiveness and robustness.
Performance of the model in detecting the uterine region.
In this section, we utilized the YOLOv5 family of models for lesion detection in early-stage endometrial cancer MRI scans. The YOLOv5-n, s, m, and l variants differ mainly in network depth and width, with parameter count and computational complexity increasing from n to l. Although larger models generally offer stronger feature representation, our dataset is relatively small and the lesion features are localized. As a result, the lightweight YOLOv5-n effectively reduces overfitting while providing faster inference and more stable training. Across multiple experiments, YOLOv5-n outperformed the larger variants in both detection accuracy and efficiency, demonstrating its practical suitability for small-sample medical imaging applications.The primary objective of this study is to propose a novel framework for early-stage endometrial cancer MRI analysis. The comparative experiments are intended for validation rather than exhaustive benchmarking. To ensure representativeness, we selected widely used models such as YOLOvX and YOLOv8 as reference baselines, thereby enabling a fair and reasonable evaluation of the proposed method.
Cancer staging performance
In the detection and classification of early-stage endometrial cancer lesions, the YOLO model was employed to detect and crop the lesion regions, effectively reducing redundant background information and providing more precise input for the subsequent classification model. In the cancer staging experiment, the performance of the ResNet34 model was compared with that of the Swin-Transformer40 and EfficientNet41 models. The detailed evaluation metrics are presented in Table 2. The ResNet34 model achieved the highest accuracy of 0.908. Through comparative analysis, we identified ResNet34 as the most effective model, showcasing exceptional performance in the accurate staging of cancer.
To provide a comprehensive evaluation of the proposed ResNet34 model’s performance, we employed a confusion matrix to present classification outcomes and utilized prediction visualization to offer an intuitive representation of the model’s performance on test set samples. As shown in Fig. 6, the confusion matrices for the ResNet34 model and the baseline models on the test set are displayed, where the horizontal axis represents predicted classes and the vertical axis denotes actual classes. Diagonal elements correspond to correctly classified samples, while off-diagonal elements indicate misclassifications. The prediction visualizations further highlight the ResNe34t model’s high accuracy and robustness in the classification task. Nonetheless, to improve the model’s ability to distinguish challenging boundary cases, increasing the diversity of the training dataset is essential.
Confusion matrix and prediction visualization for cancer staging.
Ablation experiments
To comprehensively evaluate the contributions of the components within the proposed method and analyze their impact on overall performance, we conducted an ablation study, with results summarized in Table 3. Initially, we assessed the baseline ResNet34 model, which did not integrate the YOLO-based uterine region cropping step. This configuration exhibited a relatively low accuracy of 0.858, highlighting the negative impact of redundant background information on the classification process. Upon integrating the YOLO-based cropping method, we observed a significant improvement in accuracy, reaching 0.908. Comparative experiments with other models further validated the efficacy of the cropping method in providing more precise input for the classification model. These findings from the ablation study emphasize the crucial role of uterine region cropping in enhancing model performance, thereby providing a robust framework for the automated staging of early-stage endometrial cancer.
Discussion
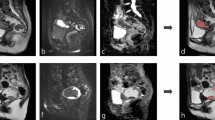
Deep learning models have demonstrated outstanding performance in medical image analysis, but their opacity remains a significant drawback. Due to the complexity and non-linear nature of these models, their decision-making processes are often incomprehensible to humans. To address this issue, this study employs Class Activation Map (CAM) technology to visualize the decision-making process of the model in medical image detection28. As shown in Fig. 7, we present the original image, the uterus region detection image, the CAM visualization, the cropped uterus region image, and the prediction map. CAM highlight the regions of the image that the model focuses on, thus elucidating the rationale behind its decisions and enhancing the interpretability of the model. By visualizing the areas of the image that the model considers important, clinicians can gain intuitive insights into the features the model references during diagnosis. If the regions highlighted by the model align with the clinically significant features identified by the physicians, it indicates a reasonable decision-making process. Conversely, if the model focuses on irrelevant or incorrect areas, it suggests potential biases that need further optimization. Additionally, interpretable models are more likely to gain the trust of clinicians, facilitating their application in actual clinical settings. Therefore, utilizing CAM to visualize the decision-making process of deep learning models not only significantly enhances their interpretability but also improves their credibility in medical image detection, ultimately providing more reliable support for clinical diagnosis and treatment.
CAM maps and the process of staging prediction (a–e).
This study has some acknowledged limitations and constraints. First, the limited size of the data set inevitably introduces variability in the experimental outcomes, a common challenge in the field of deep learning. The restricted data set size may result in insufficient generalization capability of the model during training, thereby impacting its performance in real-world applications. Additionally, our analysis was conducted using MRI images from a single location and single sequence, which restricts the generalizability and applicability of the findings. Medical imaging analysis typically involves multi-location and multi-sequence MRI images, which possess diverse characteristics and information densities, providing a more comprehensive description of lesions and a stronger diagnostic basis. To enhance the comprehensiveness and applicability of our research, future studies should integrate MRI images from multiple locations and sequences. Such a multidimensional and diversified data set would not only better utilize existing data resources but also more closely mirror the diagnostic processes and procedural norms of radiologists in clinical practice. By encompassing a broader range of image data, we can validate the model’s robustness and generalizability across various clinical scenarios, thereby ensuring its reliability and accuracy in practical applications.
Moreover, future research should focus on increasing the diversity and scale of the data set to further optimize model performance. This includes incorporating data from patients with different pathological types, age groups, genders, and ethnicities to ensure the model’s applicability and fairness across diverse populations. Additionally, further studies could explore the combination of various deep learning models and algorithms to improve detection accuracy and diagnostic capability.
The FIGO staging system is the internationally recognized standard for classifying uterine cancer, widely employed for assessing disease progression, guiding treatment strategies, and predicting patient prognosis. The most recent update, the FIGO 2023 staging system, introduces significant refinements, including more granular classification criteria for pathological and imaging characteristics. These improvements enhance the ability to stratify prognostic groups with greater precision. Notably, the 2023 update incorporates tumor molecular features, integrating biological behavior alongside anatomical attributes, thereby providing a more comprehensive and individualized basis for therapeutic decision-making. This study, being retrospective in nature, utilized data based on earlier FIGO staging criteria standards. However, the methodologies and insights presented herein remain broadly applicable and provide a foundation for future research that leverages the updated FIGO 2023 staging system.
In conclusion, although this study has made notable progress, there remains a need to enhance the robustness and generalizability of the experimental results through expanding the data set size, increasing data diversity, and optimizing algorithms. Only through continuous improvement and refinement can deep learning technology in medical image analysis truly achieve clinical application, thereby providing more reliable support for clinical diagnosis and treatment.
Conclusion
This study introduces a deep learning-based approach to effectively detect the uterine region and perform automatic staging of early endometrial cancer using MRI images. By seamlessly integrating cancer staging tasks into the clinical workflow of medical imaging departments, the proposed method improves staging efficiency and reduces inference time, establishing a fast, efficient, and accurate computer-aided diagnostic model for endometrial cancer MRI sequences. Moreover, the image processing methodology minimizes the influence of subjective factors in radiologists’ diagnostic processes, addressing the growing demand for imaging-based diagnosis and treatment. This study primarily utilizes sagittal T2-weighted MRI images of endometrial cancer. Future research will focus on a comprehensive evaluation of multi-positional and multi-sequence imaging guided by the FIGO 2023 staging criteria to better align with clinical diagnostic needs. The proposed artificial intelligence solution provides a robust and scalable approach to alleviating the increasing workload challenges in clinical imaging.
Data availability
Due to hospital policies and individual privacy, these datasets are not fully publicly available; we have produced a partial example dataset published at “https://github.com/LxiangFeng/EC-MRI-.git”, and the full dataset is available from the corresponding author upon reasonable request.
References
Crosbie, E. J. et al. Endometrial cancer. Lancet 399(10333), 1412–1428 (2022).
Sung, H. et al. Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71, 209–249. https://doi.org/10.3322/caac.21660 (2021).
Lortet-Tieulent, J., Ferlay, J., Bray, F. & Jemal, A. International patterns and trends in endometrial cancer incidence, 1978–2013. JNCI J. Natl. Cancer Inst. 110, 354–361. https://doi.org/10.1093/jnci/djx214 (2017).
Lewin, S. N. Revised FIGO staging system for endometrial cancer. Clin. Obstet. Gynecol. 54(2), 215–218. https://doi.org/10.1097/GRF.0b013e3182185baa (2011).
Amant, F. et al. Endometrial cancer. Lancet 366(9484), 491–505. https://doi.org/10.1016/S0140-6736(05)67063-8 (2005).
Hamilton, C. A. et al. Endometrial cancer: A society of gynecologic oncology evidence-based review and recommendations. Gynecol. Oncol. 160(3), 817–826. https://doi.org/10.1016/j.ygyno.2020.12.021 (2021).
Lefebvre, T. L. et al. Development and validation of multiparametric MRI–based radiomics models for preoperative risk stratification of endometrial cancer. Radiology 305(2), 375–386. https://doi.org/10.1148/radiol.212873 (2022).
Li, T. et al. Multi-parametric MRI for radiotherapy simulation. Med. Phys. https://doi.org/10.1002/mp.16256 (2023).
Cui, T. et al. Peritumoral enhancement for the evaluation of myometrial invasion in low-risk endometrial carcinoma on dynamic contrast-enhanced MRI. Front. Oncol. 11, 793709. https://doi.org/10.3389/fonc.2021.793709 (2022).
Adegun, A. A., Viriri, S. & Ogundokun, R. O. Deep learning approach for medical image analysis. Comput. Intell. Neurosci. 2021, 6215281. https://doi.org/10.1155/2021/6215281 (2021).
Salehi, A. W. et al. A study of CNN and transfer learning in medical imaging: advantages, challenges, future scope. Sustainability 15, 5930. https://doi.org/10.3390/su15075930 (2023).
Chinn, E., Arora, R., Arnaout, R. & Arnaout, R. ENRICHing medical imaging training sets enables more efficient machine learning. J. Am. Med. Inform. Assoc. 30(6), 1079–1090. https://doi.org/10.1093/jamia/ocad055 (2023).
Zhang, J., Xie, Y., Wu, Q. & Xia, Y. Medical image classification using synergic deep learning. Med. Image Anal. 54, 10–19. https://doi.org/10.1016/j.media.2019.02.010 (2019).
Zhao, Y. et al. Deep learning solution for medical image localization and orientation detection. Med. Image Anal. 81, 102529. https://doi.org/10.1016/j.media.2022.102529 (2022).
Wang, S. et al. Annotation-efficient deep learning for automatic medical image segmentation. Nat. Commun. 12(1), 5915. https://doi.org/10.1038/s41467-021-26216-9 (2021).
Lakhani, P., Gray, D. L., Pett, C. R., Nagy, P. & Shih, G. Hello world deep learning in medical imaging. J. Digit. Imaging 31(3), 283–289. https://doi.org/10.1007/s10278-018-0079-6 (2018).
Castiglioni, I. et al. AI applications to medical images: From machine learning to deep learning. Physica Med. 83, 9–24. https://doi.org/10.1016/j.ejmp.2021.02.006 (2021).
Mao, W., Chen, C., Gao, H., Xiong, L. & Lin, Y. A deep learning-based automatic staging method for early endometrial cancer on MRI images. Front. Physiol. 13, 974245. https://doi.org/10.3389/fphys.2022.974245 (2022).
Feng, L. et al. ECMS-NET: A multi-task model for early endometrial cancer MRI sequences classification and segmentation of key tumor structures. Biomed. Signal Process. Control 93, 106223. https://doi.org/10.1016/j.bspc.2024.106223 (2024).
Urushibara, A. et al. The efficacy of deep learning models in the diagnosis of endometrial cancer using MRI: A comparison with radiologists. BMC Med. Imaging 22(1), 80. https://doi.org/10.1186/s12880-022-00808-3 (2022).
Stanzione, A. et al. Deep myometrial infiltration of endometrial cancer on MRI: A radiomics-powered machine learning pilot study. Acad. Radiol. 28(5), 737–744. https://doi.org/10.1016/j.acra.2020.02.028 (2021).
Mainenti, P. P. et al. MRI radiomics: A machine learning approach for the risk stratification of endometrial cancer patients. Eur. J. Radiol. 149, 110226. https://doi.org/10.1016/j.ejrad.2022.110226 (2022).
Chen, X. et al. Deep learning for the determination of myometrial invasion depth and automatic lesion identification in endometrial cancer MR imaging: A preliminary study in a single institution. Eur. Radiol. 30(9), 4985–4994. https://doi.org/10.1007/s00330-020-06870-1 (2020).
Dong, H.-C., Dong, H.-K., Yu, M.-H., Lin, Y.-H. & Chang, C.-C. Using deep learning with convolutional neural network approach to identify the invasion depth of endometrial cancer in myometrium using MR images: A pilot study. Int. J. Environ. Res. Public Health 17, 5993. https://doi.org/10.3390/ijerph17165993 (2020).
Zhang, J. et al. Ultra-attention: Automatic recognition of liver ultrasound standard sections based on visual attention perception structures. Ultrasound Med. Biol. 49(4), 1007–1017. https://doi.org/10.1016/j.ultrasmedbio.2022.12.016 (2023).
Alzubaidi, L. et al. Review of deep learning: Concepts, CNN architectures, challenges, applications, future directions. J. Big Data 8(1), 53. https://doi.org/10.1186/s40537-021-00444-8 (2021).
Liu, P. et al. Benchmarking supervised and self-supervised learning methods in a large ultrasound muti-task images dataset. IEEE J. Biomed. Health Inform. https://doi.org/10.1109/JBHI.2024.3382604 (2024).
Zhou, B., Khosla, A., Lapedriza, A., Oliva, A. & Torralba, A. Learning Deep Features for Discriminative Localization (2015).
Bennett, W., Smith, K., Jarosz, Q., Nolan, T. & Bosch, W. Reengineering workflow for curation of DICOM datasets. J. Digit. Imaging 31(6), 783–791. https://doi.org/10.1007/s10278-018-0097-4 (2018).
Yushkevich, P. A., Gao, Y. & Gerig, G. ITK-SNAP: An interactive tool for semi-automatic segmentation of multi-modality biomedical images. In 2016 38th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC) 3342–3345. https://doi.org/10.1109/EMBC.2016.7591443 (2016).
Aljabri, M., AlAmir, M., AlGhamdi, M., Abdel-Mottaleb, M. & Collado-Mesa, F. Towards a better understanding of annotation tools for medical imaging: A survey. Multimed. Tools Appl. 81(18), 25877–25911. https://doi.org/10.1007/s11042-022-12100-1 (2022).
Rutherford, M. et al. A DICOM dataset for evaluation of medical image de-identification. Sci. Data 8(1), 183. https://doi.org/10.1038/s41597-021-00967-y (2021).
Terven, J., Córdova-Esparza, D.-M. & Romero-González, J.-A. A comprehensive review of YOLO architectures in computer vision: From YOLOv1 to YOLOv8 and YOLO-NAS. Mach. Learn. Knowl. Extract. 5(4), 1680–1716. https://doi.org/10.3390/make5040083 (2023).
Zhu, X., Lyu, S., Wang, X. & Zhao, Q. TPH-YOLOv5: Improved YOLOv5 Based on Transformer Prediction Head for Object Detection on Drone-captured Scenarios (2021).
Koh, P. W. et al. Concept Bottleneck Models (2020).
He, K., Zhang, X., Ren, S. & Sun, J. Deep residual learning for image recognition. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR) (2016).
Ge, Z., Liu, S., Wang, F., Li, Z. & Sun, J. YOLOX: Exceeding YOLO Series in 2021 (2021).
Sohan, M., Sai Ram, T. & Rami Reddy, C. V. A review on YOLOv8 and its advancements. In Data Intelligence and Cognitive Informatics (eds Jacob, I. J., Piramuthu, S. & Falkowski-Gilski, P.) 529–545 (Springer, 2024).
Girshick, R. Fast R-CNN. In: Proceedings of the IEEE International Conference on Computer Vision (ICCV) (2015).
Liu, Z. et al. Swin Transformer: Hierarchical Vision Transformer using Shifted Windows (2021).
Tan M. & Le Q. EfficientNet: Rethinking model scaling for convolutional neural networks. In: Proceedings of the 36th International Conference on Machine Learning (eds Chaudhuri, K. & Salakhutdinov, R.) Vol. 97, 6105–6114 (PMLR, 2019).
Funding
This work was supported by the grants from National Natural Science Foundation of Fujian (2021J011394, 2021J011404), the Quanzhou Scientific and Technological Planning Projects (2022NS057, 2022C006R), the Scientific Research Funds of Huaqiao University(605-50Y23038), the Joint Funds for the innovation of science and Technology, Fujian province(Grant number:2024Y9376).
Author information
Authors and Affiliations
Contributions
L.W. and L.X.F. conceptualized the study, designed the experiments, and provided the overall direction for the research. L.X.F. and J.S.Z. developed the methodology, analyzed the data, and wrote the main manuscript text. Y.P.L. contributed to the data collection and interpretation of the clinical images. Y.L.F. and P.Z.L. provided funding support and contributed to the design and validation of the deep learning models. Q.H.C. and Y.T.H. provided critical insights on the clinical aspects of the study and assisted with the interpretation of results. F.H. supervised the study, revised the manuscript as the corresponding author. All authors reviewed and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Wang, L., Feng, L., Zhang, J. et al. DCS-NET: a multi-task model for uterine ROI detection and automatic staging of early endometrial cancer in MRI. Sci Rep 15, 41024 (2025). https://doi.org/10.1038/s41598-025-25004-5
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-25004-5