Abstract
Curcumin (Cur), quercetin (Que), and piperine (Pip) are well-known plant-derived compounds with diverse pharmacological activities and health benefits. However, their clinical application is limited due to hydrophobicity and poor bioavailability. Polylactic-co-glycolic acid nanoparticles(PLGA NPs) provide efficient drug loading and sustained release, making them an ideal platform for controlled drug delivery. Cur, Que, and Pip co-loaded PLGA NPs (Cur-Que-Pip-PLGA NPs) were prepared using the emulsification-solvent evaporation method. Their physicochemical properties were characterized by transmission electron microscopy (TEM), scanning electron microscopy (SEM), dynamic light scattering (DLS),and high-performance liquid chromatography (HPLC). Biocompatibility was assessed through the CCK-8 cytotoxicity assay and Calcein AM/PI cell viability staining. Antimicrobial activity was investigated via the agar diffusion method. The average particle size of Cur-Que-Pip-PLGA NPs was 210.6 ± 0.22 nm, with a zeta potential of -8.57 ± 1.16 mV and a polydispersity index (PDI) of 0.186 ± 0.010. The encapsulation efficiencies of Cur, Que, and Pip were 97.1%, 95.5%, and 45.5%, respectively. The NPs were spherical with smooth surfaces. In vitro studies revealed that the cumulative release of Cur-Que-Pip-PLGA NPs in PBS after 96 h of sustained release was 26.9%, 57.5%, and 98%, respectively. In contrast, the cumulative release of free Cur, Que, and Pip after only 8 h of sustained release reached 92.1%, 94.8%, and 96.6%, respectively. This comparison highlights the significant slow-release effect of PLGA NPs. The NPs exhibited excellent stability at 25 ± 3°C for 14 days. The results of the CCK-8 cytotoxicity assay and Calcein AM/PI cell death staining indicated that Cur-Que-Pip-PLGA NPs exhibited good biocompatibility with RAW264.7, BMSC, and MC3T3 cells, with no significant cytotoxic effects. Agar diffusion method confirmed its good antibacterial activity against Staphylococcus aureus and Pseudomonas aeruginosa. Cur-Que-Pip-PLGA NPs demonstrate excellent biocompatibility, controlled release, and antimicrobial properties, suggesting potential for improving the therapeutic performance of hydrophobic compounds.
Similar content being viewed by others
Introduction
Traditional Chinese Medicine (TCM) has played a crucial role in healthcare for centuries, contributing significantly to the treatment of a wide range of human diseases. However, its broader development and international recognition are constrained by several factors, including the inherently low bioavailability of many active compounds, limited high-quality clinical evidence, and the lack of standardized formulations and dosing regimens1,2. These challenges not only restrict the reproducibility of therapeutic outcomes but also hinder the global integration of TCM into modern medical systems. To overcome these barriers, modern pharmaceutical technologies and scientific methodologies are increasingly being employed to isolate, modify, and optimize bioactive components from TCM, paving the way toward evidence-based and personalized therapeutic strategies3.
Curcumin (Cur), quercetin (Que), and piperine (Pip) are representative plant-derived compounds that exemplify this modern approach to TCM-inspired drug development. Cur, a polyphenol compound derived from turmeric rhizome, has been approved by the Food and Drug Administration (FDA) as a safe natural food additive. Que, a flavonoid antioxidant widely found in fruits, vegetables, and medicinal plants, and Pip, an alkaloid extracted from black and long pepper, both exhibit diverse pharmacological activities4,5,6. These compounds act through multiple molecular targets and signaling pathways, showing potent antioxidant, anti-inflammatory, antibacterial, anticancer, neuroprotective, and metabolic regulatory effects7,8,9,10. Clinical studies have confirmed that even at high doses, they demonstrate safety and good tolerability, supporting their potential for treating various diseases such as rheumatoid arthritis, Alzheimer’s disease, multiple sclerosis, atherosclerosis, and certain cancers11,12,13,14.
Despite these promising therapeutic effects, the clinical translation of Cur, Que, and Pip remains constrained by their inherent physicochemical drawbacks: poor water solubility, low bioavailability, and rapid metabolic clearance15,16. These characteristics significantly reduce their systemic absorption and therapeutic efficacy, underscoring the need for innovative delivery systems to enhance their stability, solubility, and application scope17.
In recent years, advances in nanotechnology have provided powerful solutions to long-standing challenges in drug delivery18. Nanoparticles (NPs), typically defined as particles ranging from 1 to 100 nm in diameter, have emerged as one of the most promising carriers for therapeutic agents19. For instance, Feng et al.20 successfully formulated Cur-loaded nanoparticles with diameters of 55.6 nm and 62.4 nm, significantly improving Cur solubility and extending its in vivo activity. Additionally, polymer NPs can range from 1 to 1000 nm depending on their type21. Compared with conventional formulations, NPs offer distinct advantages, such as enhanced solubility of poorly soluble drugs, excellent biodegradability and biocompatibility, prolonged circulation time, controlled drug release, targeted delivery, and reduced systemic toxicity22.
Among the various nanocarriers, polylactic-co-glycolic acid (PLGA) stands out as one of the most versatile and biocompatible polymers. Composed of polylactic acid (PLA) and polyglycolic acid (PGA), PLGA undergoes hydrolysis degradation into lactic acid and glycolic acid, which are further metabolized into water and carbon dioxide. This process ensures complete biocompatibility and safety23,24. It also demonstrates high encapsulation efficiency, non-immunogenicity, and excellent biosafety25. Owing to these characteristics, PLGA has been approved by both the U.S. FDA and the European Medicines Agency (EMA) for biomedical use26, and is now widely applied in drug delivery, diagnostic imaging, and tissue engineering27,28.
Building upon these advantages, PLGA nanoparticles can protect drugs from premature degradation, extend their therapeutic duration, and improve patient compliance29. For hydrophobic compounds such as Cur, Que, and Pip, polymeric NPs can substantially enhance pharmacokinetic behavior, solubility, and therapeutic efficacy30. Notably, Pip functions as a bioavailability enhancer and has been reported to potentiate the pharmacological activity of co-administered drugs31,32. Recent studies have also demonstrated that co-encapsulation of multiple bioactive agents within a single nanocarrier can achieve synergistic therapeutic effects and superior targeting efficiency compared to single-drug systems33.
Therefore, we hypothesize that co-loading Cur, Que, and Pip into a PLGA nanoparticle system will synergistically enhance their stability, sustain drug release, and improve antimicrobial efficacy while maintaining high biocompatibility. To validate this hypothesis, we designed and optimized Cur–Que–Pip co-loaded PLGA nanoparticles, characterized their physicochemical and release behavior, and systematically evaluated their cytocompatibility and antibacterial performance. This study aims to develop a multifunctional nanoplatform for the efficient delivery of multiple bioactive compounds, potentially improving therapeutic outcomes for hydrophobic agents.
Materials and methods
Materials and chemicals
Curcumin, quercetin, and piperine (purity ≥95%, HPLC grade) were procured from Med Chem Express (Monmouth Junction, NJ, USA). Poly(lactic-co-glycolic acid) (PLGA; 50:50 ratio, molecular weight 55,000) was supplied by Jinan Daigang Bioengineering Co., Ltd. Polyvinyl alcohol (PVA;88% hydrolyzed) was purchased from Shanghai Aladdin Reagent Co., Ltd. Chromatographically pure methanol and ethyl acetate were acquired from Honeywell. Acetone and dichloromethane were provided by the laboratory, while Tween 80 was obtained from Sinopharm Chemical Reagent Co., Ltd. Fetal bovine serum (10%) was sourced from Gibco (USA). Dimethyl sulfoxide (DMSO), 0.25% trypsin, DMEM, and MEM-α medium were purchased from Thermo Fisher Scientific (Beijing, China).The Calcein AM/PI viability assay kit was sourced from Suzhou Youyilandi Biotechnology Co., Ltd., and 96-well cell culture plates were obtained from Corning (USA). The CCK-8 cell proliferation assay kit was procured from Shanghai Bai Sai Biotechnology Co., Ltd.Rabbit mesenchymal stem cell-specific culture medium (DMEM medium containing 5% fetal bovine serum, 1% penicillin, 1% streptomycin, and stem cell growth factors) was obtained from Shanghai Zhongqiao Xinzhou Biotechnology Co., Ltd. All other chemicals and solvents utilized in this study were of analytical grade.
Cell line and culture
Mouse monocyte macrophages (RAW264.7,CVCL-0493, Shanghai Cell Bank, CN), rabbit mesenchymal stem cells (RSC-RB,ZQ0682, Zhong Qiao Xin Zhou, CN), and mouse cranial parietal preosteoblasts subclone 14 (MC3T3-E1 Subclone 14,AW-CELLS-M0047, CVCL-5437, Anweisci, CN) were retrieved from liquid nitrogen storage, thawed in a 37 °C water bath, and resuscitated. After gentle agitation to facilitate thawing, all cell-handling procedures were conducted within a sterile laminar flow hood to minimize contamination risk. The cell suspension was pipetted into culture media comprising DMEM with 10% FBS, DMEM medium containing 5% FBS, 1% penicillin, 1% streptomycin, stem cell growth factors, or MEM-α medium supplemented with 10% FBS (Gibco, USA) and incubated at 37 °C in 5% CO₂. Cell morphology was observed using an inverted phase contrast microscope (IX2-SLP, OLYMPUS, Japan), and the culture medium was changed every other day. Upon reaching approximately 80% confluence, cells were passaged and subsequent experiments were conducted.
Formulation of nanoparticle drug preparation
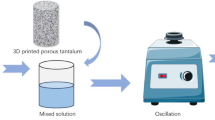
Cur-Que-Pip-PLGA NPs were prepared using the emulsification-solvent evaporation method (Fig. 1)34,35. Briefly, 200 mg of PLGA was dissolved in 3 mL of an organic solvent mixture (dichloromethane:acetone = 4:1) to form the organic phase. Subsequently, Cur, Que, and Pip powders were added at a mass ratio of 1:1:0.2, corresponding to 10 mg of Cur, 10 mg of Que, and 2 mg of Pip per batch, respectively. The total drug-to-polymer ratio was 1:10. The organic phase was slowly added dropwise into 5 mL of 2% polyvinyl alcohol (PVA) aqueous solution (used as a surfactant) under continuous stirring to form a coarse emulsion. The mixture was then sonicated at 40% amplitude for 5 min in an ice bath to generate a fine emulsion, which was subsequently transferred into 10 mL of 0.5% PVA solution. The suspension was magnetically stirred for 3 h at room temperature to evaporate the organic solvent.
Schematic representation of the preparation process for Cur-Que-Pip-PLGA NPs.
The nanoparticle suspension was centrifuged at 5000 rpm for 15 min to remove large aggregates, followed by centrifugation at 15000 rpm for 30 min at 4 °C to collect the nanoparticles. The obtained Cur–Que–Pip–PLGA NPs were washed three times with distilled water, re-suspended, and freeze-dried at −20°C for subsequent characterization. All formulations were prepared in triplicate.
Measurement of particle size, Zeta potential and polydispersion index (PDI)
The mean particle size, PDI, and zeta potential of the nanoparticles were measured using dynamic light scattering (DLS) on a Zetasizer Nano (Malvern Instruments, UK)36. The prepared Cur-Que-Pip-PLGA NPs were diluted with ultrapure water and analyzed using the instrument. Each measurement was performed in triplicate.
Transmission electron microscopy and scanning electron microscopy
Transmission electron microscopy (TEM, JEM-F200, JEOL) and scanning electron microscopy (SEM, JSM-IT700HR, JEOL) were employed to examine the morphology and surface characteristics of Cur-Que-Pip-PLGA NPs37,38. The nanoparticle samples were prepared by diluting Cur-Que-Pip-PLGA NPs with deionized water. TEM: Samples were deposited onto carbon-coated copper grids, allowed to dry for 2–3 min at room temperature, and stained with 1% phosphotungstic acid under light-protected conditions. After drying, the samples were imaged using a transmission electron microscope. SEM: Samples were dropped onto degradation-resistant slides, dried at room temperature, affixed to conductive carbon tape, and sputter-coated with gold for approximately 30 s. The coated samples were observed and imaged using a scanning electron microscope. All experiments were repeated three times.
Cur-Que-Pip-PLGA NPs encapsulation efficiency (EE%)
The encapsulation efficiency (EE%) of Cur, Que, and Pip in PLGA nanoparticles was determined using high-performance liquid chromatography (HPLC)39. Briefly, 5 mg of the freeze-dried Cur–Que–Pip–PLGA NPs were resuspended in 1 mL of distilled water and centrifuged at 15,000 rpm for 30 min at 4 °C to separate the nanoparticles from unencapsulated drugs. The supernatant was collected, and the concentrations of free Cur, Que, and Pip were quantified by HPLC. To determine the total drug content, another 1 mL of nanoparticle suspension was dissolved in 1 mL of methanol and sonicated at 40% amplitude for 20 min in an ice bath to fully release the encapsulated drugs. Methanol was selected as the extraction solvent due to the high solubility and chemical stability of Cur, Que, and Pip under these conditions, and no degradation was observed during the process. Chromatographic analysis was conducted using a mobile phase of acetonitrile: water (86:20, v/v) at a flow rate of 1.0 mL/min and a column temperature of 25°C. The injection volume was set to 20 µL. The encapsulation efficiency (EE%) was calculated using the following formula:
where initial drug represents the theoretical amount of each individual drug (Cur, Que, or Pip) initially added during nanoparticle preparation, and free drug represents the amount of unencapsulated drug in the supernatant.
In vitro drug release experiment
The in vitro drug release behavior of Cur–Que–Pip–PLGA nanoparticles (NPs) was determined using dialysis bag diffusion method and compared with free Cur, Que, and Pip as controls. For each test, an equivalent of 10 mg Cur–Que–Pip–PLGA NPs (corresponding to approximately 0.5 mg Cur, 0.5 mg Que, and 0.1 mg Pip) was accurately weighed and placed in a dialysis bag (molecular weight cutoff 8,000–14,000 Da).
The dialysis bag was immersed in 30 mL of phosphate-buffered saline (PBS, pH 7.4) containing 1% Tween 80 to enhance the solubility of the poorly water-soluble drugs. The medium was maintained at 37 ± 0.5°C under gentle agitation at 100 rpm in an orbital shaker. The drug-to-medium ratio and the addition of Tween 80 ensured that sink conditions were maintained throughout the experiment, as the total drug concentration did not exceed 10% of the saturation solubility of Cur, Que, or Pip in the release medium.
At predetermined intervals (0.5, 1, 2, 4, 6, 8, 12, 24, 32, 48, 72, and 96 h), 2 mL of the release medium was withdrawn and replaced with an equal volume of fresh preheated medium to maintain constant volume and sink condition. The samples were filtered through a 0.45 μm membrane filter, and the concentrations of Cur, Que, and Pip were determined by HPLC at their respective wavelengths. The cumulative release percentages were calculated and plotted as a function of time to analyze the release kinetics40,41.
Storage stability research
Nanoparticle stability is a crucial parameter. The stability of Cur-Que-Pip-PLGA NPs was investigated by observing possible changes in the solution state, average particle size and PDI42. Free Cur-Que-Pip and freeze-dried Cur-Que-Pip-PLGA NPs were in PBS (pH 7.4) containing 1% Tween80, respectively, Vortexed mixer at 5 W for 1 min, and then stored in airtight glass vials. The samples were stored at 25±3°C for 2 weeks to assess their stability. Visual inspection for changes in appearance and color was performed at set time points (0, 2, 5, 8, 10, and 14 days). Samples were taken on days 1, 3, 5, 7, 9, 12, and 14, and the changes in mean particle size and PDI were analyzed using a DLS.
Biocompatibility test
The biocompatibility of Cur–Que–Pip–PLGA nanoparticles (NPs) was assessed using a CCK-8 assay and Calcein AM/PI live–dead staining43,44. RAW264.7 cells were seeded into 96-well plates at a density of 1×105 cells/well in DMEM supplemented with 10% FBS and cultured for 24 h at 37 °C in 5% CO₂. Untreated cells served as controls.
Cells were then exposed to different concentrations of free Cur–Que–Pip, blank PLGA NPs, and Cur–Que–Pip–PLGA NPs (0, 10, 30, 60, 80, 100, and 120 µg/mL), where the concentrations refer to the total nanoparticle mass per milliliter of medium. According to the drug feeding ratio (Cur:Que:Pip = 1:1:0.2), these NP concentrations correspond to approximate drug concentrations of Cur = 4.6 µg/mL, Que = 4.6 µg/mL, and Pip = 0.9 µg/mL for the 100 µg/mL NP sample, with proportional adjustments for the other groups.
After 24 h of incubation, the drug-containing medium was discarded and the cells were gently rinsed twice with PBS. Next, an equal volume of a mixture containing 100μL of fresh medium and 10μL of CCK-8 reagent was added to each well, and the plate was incubated for 1 hour. Absorbance (OD) at 450 nm was measured using a microplate reader to calculate cell viability. The cell survival rate(%)was determined using the formula: cell survival rate (%) = (OD of experimental group-OD of blank group)/(OD of control group-OD of blank group) ×100%.
For live/dead staining, RAW264.7, BMSC, and MC3T3 cells were seeded in 12-well plates (2×105 cells/well) and treated with free Cur–Que–Pip, blank PLGA NPs, or Cur–Que–Pip–PLGA NPs at 80 µg/mL total NP concentration (equivalent to Cur ≈ 3.6 µg/mL, Que ≈ 3.6 µg/mL, and Pip ≈ 0.7 µg/mL) for 48 h. After washing with PBS, cells were stained with Calcein AM/PI solution for 20 min in the dark, rinsed, and imaged under a fluorescence microscope (DMi8, Leica, Germany). Live cells appeared green, while dead cells appeared red. All experiments were performed in triplicate.
Antibacterial activity experiment
To evaluate the antibacterial activity using the agar well diffusion method45,46, the following procedure was adopted. Initially, Staphylococcus aureus and Pseudomonas aeruginosa were separately inoculated into nutrient broth and subjected to oscillatory culture at 37 °C for 24 h to reach the logarithmic growth phase. Subsequently, the cultured bacterial suspensions were diluted with sterile physiological saline, adjusting the density to 1.0×10⁷ CFU/mL. A sterile cotton swab was used to dip 100μL of the bacterial suspension and evenly spread it on a petri dish containing nutrient agar. Next, a sterile punch was used to create holes in the agar plate with a diameter of 6 mm, maintaining appropriate spacing between holes to prevent the overlap of inhibition zones. A micropipette was employed to add 50μL of nanoparticle solution (150μg/mL) into each hole. Concurrently, positive controls were set up with 50μL of penicillin (100ng/mL) and levofloxacin (16μg/mL), and negative controls with 50μL of distilled water. The inoculated petri dishes were placed in a constant temperature incubator at 37 °C for cultivation. At the predetermined time points, the petri dishes were removed, and the diameters of the inhibition zones surrounding each hole were measured with a vernier caliper. The antibacterial activity of the nanoparticles was assessed based on the size of the inhibition zone diameters. The experiment was repeated three times to ensure the reliability of the results.
Statistical analysis
All tests were performed at least three times, and the data are presented as mean± standard deviation. Statistical analyses were performed using Prism version 9 (GraphPad Software, San Diego, CA,USA). Comparisons between two groups with normally distributed data were conducted using Student’s t-tests; otherwise, the nonparametric Mann–Whitney test was used. For comparisons of multiple groups with normally distributed data, one-way ANOVA with Tukey’s post hoc test was applied. For non-normal distributed data, the Kruskal–Wallis test was used. A p-value of less than 0.05 was considered statistically.
Results
Average particle size, Zeta potential, and PDI
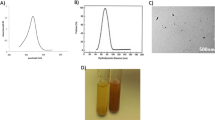
Dynamic light scattering (DLS) analysis revealed that the Cur–Que–Pip–PLGA NPs exhibited a uniform size distribution with an average diameter of 210.6 ± 0.22 nm (Fig. 2A), indicating homogeneous nanoparticle formation. The Zeta potential was –8.57 ± 1.16 mV (Fig. 2B; Table 1), suggesting moderate surface charge and colloidal stability. The PDI values ranged from 0.176 to 0.195, confirming the monodispersity of the nanoparticles (Table 1).
DLS Measurement of Cur-Que-Pip-PLGA NPs (A) particle size distribution, (B) zeta potential. The results were presented as mean± standard deviation based on three independent experiments.
These results demonstrate that the emulsification–solvent evaporation method produced nanoscale particles with consistent morphology and narrow distribution, suitable for drug encapsulation and in vivo delivery. The relatively small particle size and negative surface potential are expected to promote stability in aqueous media and favorable interactions with biological systems47.
Furthermore, the zeta potential of Cur–Que–Pip–PLGA NPs was measured as –8.57 ± 1.16 mV, indicating a moderately negative surface charge. This relatively low zeta potential may result from the adsorption of PVA on the nanoparticle surface, which masks part of the negatively charged carboxyl groups of PLGA and provides steric stabilization instead of purely electrostatic repulsion. In addition, weak interactions between the encapsulated drugs and the PLGA matrix may further reduce the surface charge density. Nevertheless, the nanoparticles exhibited good colloidal stability and monodispersity, as evidenced by their low PDI values.
Cur-Que-Pip-PLGA NPs drug entrapment efficiency (EE%)
Entrapment efficiency (EE%) of Cur, Que, and Pip in PLGA NPs were determined to be 97.1%, 95.5%, and 45.5%, respectively. These high EE% indicate that the majority of the drugs were effectively encapsulated within the PLGA NPs (Table 1).
Morphological evaluation
The structural characteristics of Cur-Que-Pip-PLGA NPs were meticulously examined using TEM and SEM. As shown in Fig. 3, TEM imaging revealed that the NPs possessed a predominantly spherical or spheroidal morphology with uniform dispersion, consistent particle size, and a smooth, rounded surface. SEM imaging was employed to further investigate the surface morphological features of Cur-Que-Pip-PLGA NPs, offering detailed insights into their microstructure and potential surface heterogeneity.
TEM and SEM observations of the morphology of Cur-Que-Pip-PLGA NPs (A-B) TEM representative image (multiple: x15000 scale: 200nm), (C) SEM representative image (multiple: x20000 scale: 1 μm). Based on three independent experiments.
In vitro release assay
The in vitro release behavior of Cur-Que-Pip-PLGA NPs and free Cur-Que-Pip were compared using PBS (pH 7.4) containing 1% Tween 80 as the release medium to improve drug solubility. The release behavior of the encapsulated drugs was monitored over 96 h (Fig. 4A). The cumulative release of Cur, Que, and Pip from Cur-Que-Pip-PLGA NPs reached 26.9%, 57.5%, and 98.0%, respectively, showing a sustained and gradual release pattern. In contrast, free Cur, Que, and Pip exhibited rapid release within 8 h, reaching 92.1%, 94.8%, and 96.9%, respectively (Fig. 4B). Notably, during the same period, Que and Pip in the nanoparticle formulation achieved cumulative release levels of 26.8% and 75.5%, while Cur displayed the slowest release rate (2.9%), indicating its stronger interaction with the PLGA matrix. These results demonstrate that PLGA encapsulation effectively slows drug diffusion and provides a controlled, sustained-release behavior, suggesting that Cur-Que-Pip-PLGA NPs can serve as an efficient platform to improve the solubility and utilization of hydrophobic drugs.
In vitro release behavior of Cur-Que-Pip-PLGA NPs (A) Drug release behavior of free Cur-Que-Pip, (B) Drug release behavior of Cur-Que-Pip-PLGA NPs. The results were presented as mean± standard deviation based on three independent experiments.
Particle stability
Nanoparticle stability is a critical factor influencing drug storage and transportation. The Cur-Que-Pip-PLGA NPs solutions maintained a pale yellow color throughout the storage period, displaying uniform dispersion in water without noticeable changes in appearance or color (Fig. 5A). In contrast, free Cur-Que-Pip solutions initially exhibited slight solubility, followed by drug aggregation, precipitation, and eventual stratification at the bottom of the container over time (Fig. 5A).Additionally, characterization parameters, including particle size and PDI of Cur-Que-Pip-PLGA NPs, showed no significant variation over two weeks (Fig. 5B-C).These findings suggest that the prepared nanoparticle drug formulations exhibit excellent stability.
Stability evaluation of Cur-Que-Pip-PLGA NPs. (A) Visual comparison of Cur-Que-Pip-PLGA NPs and free Cur-Que-Pip solutions over a two-week storage period. (B) Particle size stability of Cur-Que-Pip-PLGA NPs, (C) PDI stability of Cur-Que-Pip-PLGA NPs. The Results are presented as mean ± standard deviation based on three independent experiments.
Biocompatibility evaluation of nanoparticles in vitro
The effects of free Cur-Que-Pip, PLGA NPs, and Cur-Que-Pip-PLGA NPs on cell viability were evaluated using the CCK-8 cytotoxicity assay and Calcein AM/PI live/dead cell staining. As shown in Figure 6A, none of the tested agents, including free Cur-Que-Pip, PLGA NPs, and Cur-Que-Pip-PLGA NPs, exhibited detectable cytotoxicity in RAW264.7 cells, even at concentrations as high as 120 µg/mL. Consequently, 80 µg/mL was selected as the optimal drug concentration for subsequent experiments. During co-culture with RAW264.7, MBSC, and MC3T3 cells for 48 h, these agents did not adversely affect cell growth, and no dead cells (indicated in red) were observed (Fig. 6B-D).In summary, these findings demonstrate that Cur-Que-Pip-PLGA NPs exert less impact on cell viability compared to free Cur-Que-Pip at equivalent concentrations, underscoring the potential of PLGA NPs as an ideal delivery system for Cur-Que-Pip.
Biocompatibility assessment of Cur-Que-Pip-PLGA NPs in vitro. (A) Cytotoxicity of free Cur-Que-Pip, PLGA NPs, and Cur-Que-Pip-PLGA NPs on RAW264.7 cells was evaluated using the CCK-8 assay after 24 h of incubation. Results are expressed as mean ± standard deviation from three independent experiments. (B-D) Live/dead cell staining and viability assessment of RAW264.7, BMSC, and MC3T3 cells co-cultured with free Cur-Que-Pip, PLGA NPs, and Cur-Que-Pip-PLGA NPs (80 µg/mL) for 48 h. Live cells are stained green, and dead cells are stained red. Scale bar = 100 µm. Based on three independent experiments.
Antibacterial activity
The antibacterial activity of Cur-Que-Pip-PLGA NPs against Staphylococcus aureus (S. aureus) and Pseudomonas aeruginosa(P. aeruginosa) was determined using the agar diffusion method, as shown in Fig. 7. The control group (distilled water) exhibited no inhibitory effect, with robust bacterial growth. In contrast, the antibiotic group (penicillin at 100 ng/mL and levofloxacin at 16 µg/mL) demonstrated significant antibacterial effects, forming distinct zones of inhibition around both S. aureus and P. aeruginosa (Table 2). This indicates that these antibiotics possess strong inhibitory effects against these two bacterial strains. Similarly, Cur-Que-Pip-PLGA NPs exhibited significant antibacterial activity against S. aureus and P. aeruginosa, with their antibacterial effects increasing over time and resulting in clear zones of inhibition (Table 2). This suggests that the combination of Cur, Que, and Pip with the biocompatible PLGA NPs enables the effective and sustained release of antibacterial activity. Table 2. lists the diameters of the inhibition zones obtained from the microbial samples tested within 24 h.
Antibacterial effects of antibiotics and Cur-Que-Pip-PLGA NPs against Staphylococcus aureus (S. aureus) and Pseudomonas aeruginosa (P. aeruginosa). (A) Representative image of S. aureus. (B) Representative image of P. aeruginosa. Based on three independent experiments.
Discussion
Traditional herbal-derived compounds such as Cur, Que, and Pip have attracted increasing attention for their broad-spectrum pharmacological effects, including antioxidant, immunomodulatory, analgesic, antimicrobial, anti-inflammatory, and anticancer activities48,49,50,51. However, their clinical translation remains constrained by poor aqueous solubility, instability, and low utilization Rate. In this study, we mechanistically rationalized the co-encapsulation of Cur, Que, and Pip within poly(lactic-co-glycolic acid) (PLGA) nanoparticles (NPs) to overcome these limitations and achieve synergistic therapeutic effects. Cur and Que exhibit complementary antioxidant and antibacterial mechanisms, while the combined use of pip and cur enhances cur’s bioavailability52,53,54. The PLGA nanocarrier provides further benefits by protecting drugs from degradation and enabling sustained release, collectively contributing to improved pharmacokinetic profiles and therapeutic efficacy.
Referring to previous studies, the emulsification-solvent evaporation method has been demonstrated to be highly effective for encapsulating a diverse range of hydrophobic compounds55. Given the hydrophobic nature of Cur, Que, and Pip, reducing the size of oil-in-water emulsions provides an optimal approach for their encapsulation56. In this study, the prepared Cur-Que-Pip-PLGA NPs exhibited a uniform spherical morphology with a mean diameter of approximately 210.6 ± 0.22 nm and a narrow polydispersity index (PDI < 0.2), suggesting excellent homogeneity and dispersion stability. The slightly negative zeta potential (−8.57 ± 1.16 mV) may be attributed to the terminal carboxyl groups of PLGA and the adsorption of PVA residues on the particle surface. Although the magnitude of zeta potential was moderate, electrostatic repulsion combined with steric stabilization provided by PVA effectively prevented aggregation, conferring good colloidal stability57,58,59. Such nanoscale size and surface charge are optimal for cellular internalization via endocytosis, facilitating efficient intracellular drug delivery while maintaining minimal cytotoxicity60.
Additionally, the high encapsulation rates demonstrated in this study (Cur: 97.1%, Que: 95.5%, Pip: 45.5%) surpass those reported in similar single drug systems61. For instance, Feltrin et al.62 reported an EE of 72% for Cur-PLGA NPs, while Hussein et al.63 achieved an EE of 92% for Que-PLGA NPs.The relatively lower encapsulation of Pip may result from its physicochemical properties, particularly higher polarity and solubility in the external aqueous phase, which reduce its entrapment efficiency during emulsification64. Nonetheless, all three drugs remained stably encapsulated, and the nanoparticles demonstrated excellent storage stability in PBS containing 1% Tween 80 without aggregation, attributed to the interfacial stabilization provided by PVA.
The in vitro release study revealed a biphasic release pattern characterized by an initial burst phase followed by a sustained release over 96 h. The initial burst is likely due to desorption of surface-bound drugs, while the subsequent gradual release reflects diffusion through and degradation of the PLGA matrix65,66. Distinct release profiles were observed among the three drugs, consistent with their differential hydrophobicity and localization within the polymeric core. The slow and sustained release of Cur and Que, coupled with the faster diffusion of Pip, highlights the potential for time-staggered synergistic action, allowing Pip to enhance the utilization rate of the other two agents throughout the release process67,68. Such controlled release kinetics support the long-term therapeutic potential of the nanoparticle system.
In vitro cytotoxicity studies demonstrated that Cur-Que-Pip-PLGA NPs were non-toxic to RAW264.7, MC3T3, and BMSC cells, indicating excellent biocompatibility69,70. Furthermore, Cur-Que-Pip-PLGA NPs exhibited significant inhibitory effects against both Staphylococcus aureus and Pseudomonas aeruginosa. Interestingly, although the inhibition zones of NPs against S. aureus were slightly smaller than those of conventional antibiotics, the NPs displayed a comparable antibacterial effect against P. aeruginosa after 18–24 h of incubation. This observation suggests that the nanoparticle formulation achieved time-dependent enhancement in antibacterial efficacy, which may be attributed to its sustained drug release and improved penetration through the outer membrane of Gram-negative bacteria.
The distinct behavior between the two strains can be mechanistically explained by differences in cell envelope architecture. S. aureus, a Gram-positive bacterium, possesses a thick peptidoglycan layer that allows rapid antibiotic diffusion, resulting in stronger early inhibition by conventional drugs. In contrast, P. aeruginosa, a Gram-negative bacterium, has an additional outer membrane rich in lipopolysaccharides, which typically limits drug penetration. The smaller and more stable Cur-Que-Pip-PLGA NPs could gradually accumulate and permeate this barrier, leading to an antibacterial performance approaching that of standard antibiotics over extended exposure times71. Although the antimicrobial activity of individual Cur, Que, and Pip has been well-documented in previous studies, this work primarily focused on evaluating the co-loaded PLGA nanoparticle system (Cur-Que-Pip-PLGA NPs) as a synergistic delivery platform. Therefore, PLGA nanoparticles containing single drugs were not included as control groups. Future studies will compare the antibacterial efficacy among single-drug-loaded and co-loaded PLGA NPs to further elucidate potential synergistic effects.
Furthermore, the co-delivery of Cur, Que, and Pip may contribute to this outcome through multiple synergistic mechanisms: Cur disrupts bacterial membranes; Que inhibits DNA gyrase and cell wall synthesis; and Pip interferes with efflux pumps and biofilm formation. Collectively, these effects support a multi-target antibacterial mechanism that compensates for the intrinsic resistance of P. aeruginosa and enhances the long-term inhibitory effect of the nanoformulation72,73,74,75.
Although only two representative bacterial strains (S. aureus and P. aeruginosa) were tested in this study due to experimental constraints, they serve as models for Gram-positive and Gram-negative bacteria. Further studies involving a broader range of microbial species, including fungi and drug-resistant strains, as well as mechanistic analyses, will be conducted in the future to fully assess the broad-spectrum antimicrobial potential of the nanoparticles.
In summary, our findings suggest that the co-encapsulation of Cur, Que, and Pip within PLGA nanoparticles provides a feasible and synergistic strategy to improve the delivery and therapeutic performance of natural compounds. This nanoformulation helps mitigate the inherent drawbacks of plant-derived drugs, such as poor solubility, instability, and limited utilization rate, while enabling sustained release and enhanced antimicrobial activity. Future studies will be directed toward in vivo pharmacokinetic assessment, mechanistic elucidation of cellular uptake and trafficking, and evaluation against a broader spectrum of microbial and drug-resistant strains. Overall, this work offers a promising foundation for the further development of nanoparticle-based delivery systems for infectious and inflammatory diseases.
Conclusion
In this study, Cur, Que, and Pip were successfully co-loaded into PLGA NPs using the emulsification-solvent evaporation method. The results demonstrated that the PLGA NPs exhibited high drug encapsulation efficiency. In vitro release studies revealed a slow, sustained release of Cur, Que, and Pip from the PLGA NPs, with no significant changes observed throughout the storage period, indicating the physical stability of the formulated nanoparticles. The PLGA NPs enhanced the bioavailability of the drugs, thereby improving their biopharmaceutical and pharmacodynamic properties. This strategy also helped overcome some of the inherent limitations of these hydrophobic compounds, while maintaining excellent biocompatibility. Therefore, PLGA NPs hold significant potential as nanocarriers for the co-delivery of Cur, Que, and Pip. Our findings offer valuable insights for advancing nanoparticle technology, enhancing the efficacy and safety of combination therapies, and providing new perspectives in the field of nanomedicine. The antibacterial schematic diagram of PLGA NPs is shown in Fig. 8.
Schematic diagram of the antibacterial mechanism of PLGA NPs.
Future outlook
There are certain limitations in the present study. The in vitro drug release was conducted under a simplified condition using PBS (pH 7.4) containing 1% Tween 80 to enhance drug solubility. In physiological or pathological environments with different pH values, ionic strengths, or enzymatic activities, the release behavior of the nanoparticles may differ. In addition, due to current equipment constraints, advanced spectroscopic characterizations such as FTIR, DSC, and PXRD were not performed. These analyses will be incorporated in future work to elucidate drug–polymer interactions and confirm the physical state of encapsulated compounds.
Moving forward, a 3D printing–based centrifugal flow reactor will be employed to systematically optimize formulation parameters, including PVA concentration, ultrasonication time, and drug ratios, in order to achieve the best encapsulation and release performance. Furthermore, in vivo pharmacokinetic and pharmacodynamic studies will be conducted using infection or inflammation models to evaluate the therapeutic efficacy and clarify the underlying molecular mechanisms.
Ultimately, the co-loading strategy will be extended to additional hydrophobic and synergistic natural drugs, aiming to develop multifunctional PLGA-based nanoplatforms for controlled delivery and improved treatment outcomes in regenerative medicine and infectious diseases.
Data availability
The datasets used and/or analysed during the current study available from the corresponding author on reasonable request.
References
Liu, B. et al. Thinking and practice of accelerating transformation of traditional Chinese medicine from experience medicine to evidence-based med -icine. Front. Med. 5, 163–170 (2011).
Huang, K. et al. Traditional Chinese Medicine (TCM) in the treatment of COVID-19 and other viral infections: efficacies and mechanis- ms. Pharmacol. Ther. 225, 107843 (2021).
Li, F. S. & Weng, J. K. Demystifying traditional herbal medicine with modern approach. Nat. Plants 3, 17109 (2017).
Hatab, H. M. et al. A combined treatment of curcumin, piperine, and taurine alters the circulating levels of IL-10 and miR-21 in hepatocellular carcinoma patients: a pilot study. J. Gastrointest. Oncol. 10, 766–776 (2019).
Jagota, S. & Rajadas, J. Effect of phenolic compounds against A β aggregation and A β-induced toxicity in transgenic C. elegans. Neurochem. Res. 37, 40–48 (2012).
Rather, R. A. & Bhagat, M. Quercetin as an innovative therapeutic tool for cancer chemoprevention: molecular mechanisms and implications in human health. Cancer Med. 9, 9181–9192 (2020).
Dos Santos, P. D. F. et al. The nanoencapsulation of curcuminoids extracted from Curcuma longa L. and an evaluation of their cytotoxic, enzymatic, antioxidant and anti-inflammatory activities. Food Funct. 10, 573–582 (2019).
Memarzia, A. et al. Experimental and clinical reports on anti-inflammatory, antioxidant, and immunomodulatory effects of curcuma longa and curcumin, an updated and comprehensive review. Biofactors 47, 311–350 (2021).
Yadav, N. et al. PLGA-quercetin nano-formulation inhibits cancer progression via mitochondrial dependent caspase-3,7 and independent FoxO1 activation with concomitant PI3K/AKT suppression. Pharmaceutics 14, 1326 (2022).
Duse, L. et al. Photodynamic therapy of ovarian carcinoma cells with curcumin-loaded biodegradable polymeric nanopar- ticles. Pharmaceutics 11, 282 (2019).
Panknin, T. M. et al. Curcumin supplementation and human disease: a scoping review of clinical trials. Int. J. Mol. Sci. 24, 4476 (2023).
Hong, Y. et al. Quercetin induces anticancer activity by upregulating pro-NAG-1/GDF15 in differentiated thyroid cancer cells. Cancers (Basel). 13, 3022 (2021).
Heidari, H. et al. Curcumin-piperine co-supplementation and human health: a comprehensive review of preclinical and clinical studi- es. Phytother. Res. 37, 1462–1487 (2023).
Corrêa Carvalho, G. et al. Curcuma longa: nutraceutical use and association with nanotechnology. Adv. Healthc. Mater. 13, e2400506 (2024).
Ubeyitogullari, A. & Ciftci, O. N. A novel and green nanoparticle formation approach to forming low-crystallinity curcumin nanoparticles to improve curcumin’s bioac-cessibility. Sci. Rep. 9, 19112 (2019).
Esfandiarpour-Boroujeni, S. et al. Fabrication and study of curcumin loaded nanoparticles based on folate-chitosan for breast cancer therapy application. Carbohydr. Polym. 168, 14–21 (2017).
Sarker, M. R. & Franks, S. F. Efficacy of curcumin for age-associated cognitive decline: a narrative review of preclinical and clinical studies. Geroscience 40, 73–95 (2018).
Xin, L. et al. Evaluation of METase-pemetrexed-loaded PEG-PLGA nanoparticles modified with anti-CD133-scFV for treatment of gastric carcinoma. Biosci. Rep. 38, BSR20171001 (2018).
Joudeh, N. & Linke, D. Nanoparticle classification, physicochemical properties, characterization, and applications: a comprehensive review for biologists. J. Nanobiotechnol. 20, 262 (2022).
Feng, R., Song, Z. & Zhai, G. Preparation and in vivo pharmacokinetics of curcumin-loaded PCL-PEG-PCL triblock copolymeric nanoparticles. Int. J. Nanomed. 7, 4089–4098 (2012).
Zielińska, A. et al. Polymeric nanoparticles: production, characterization. Toxicol. Ecotoxicol. Mol. 25, 3731 (2020).
Liu, W. et al. Multiple nano-drug delivery systems for intervertebral disc degeneration: current status and future perspectives. Bioact. Mater. 23, 274–299 (2022).
Chereddy, K. K., Payen, V. L. & Préat, V. PLGA: From a classic drug carrier to a novel therapeutic activity contributor. J. Control Release 289, 10–13 (2018).
Su, Y. et al. PLGA-based biodegradable microspheres in drug delivery: recent advances in research and application. Drug Deliv. 28, 1397–1418 (2021).
Rocha, C. V. et al. PLGA-based composites for various biomedical applications. Int. J. Mol. Sci. 23, 2034 (2022).
Patel, J. et al. Targeted delivery of monoclonal antibody conjugated docetaxel loaded PLGA nanoparticles into EGFR overexpressed lung tumour cells. J. Microencapsul. 35, 204–217 (2018).
Li, M. et al. Osteogenesis effects of magnetic nanoparticles modified-porous scaffolds for the reconstruction of bone defect after bone tumor resection. Regen. Biomater. 6, 373–381 (2019).
Parveen, S., Misra, R. & Sahoo, S. K. Nanoparticles: a boon to drug delivery, therapeutics, diagnostics and imaging. Nanomedicine 8, 147–166 (2012).
Hassan, R. H. et al. Chitosan nanoparticles for intranasal delivery of olmesartan medoxomil: pharmacokinetic and pharmacodynamic perspectives. Int. J. Pharm. 628, 122278 (2022).
Arcaro, C. A. et al. Piperine, a natural bioenhancer, nullifies the antidiabetic and antioxidant activities of curcumin in streptozotocin-diabetic rats. PLoS One 9, e113993 (2014).
Saini, N., Chopra, B. & Dhingra, A. K. Synergistic effect of piperine and its derivatives: a comprehensive review. Curr. Drug Res. Rev. 15, 105-121. (2023).
Kesarwani, K., Gupta, R. & Mukerjee, A. Bioavailability enhancers of herbal origin: an overview. Asian. Pac. J. Trop. Biomed. 3, 253–266 (2013).
Liu, K. et al. Co-encapsulation systems for delivery of bioactive ingredients. Food Res. Int. 155, 111073 (2022).
Paswan, S. K. & Saini, T. R. Purification of drug loaded PLGA nanoparticles prepared by emulsification solvent evaporation using stirred cell ultrafiltration technique. Pharm. Res. 34, 2779–2786 (2017).
Pulingam, T. et al. Exploring various techniques for the chemical and biological synthesis of polymeric nanoparticles. Nanomaterials (Basel) 12, 576 (2022).
Alkholief, M. et al. Employing a PLGA-TPGS based nanoparticle to improve the ocular delivery of Acyclovir. Saudi Pharm. J. 27, 293–302 (2019).
Yadav, B. et al. RGD-decorated PLGA nanoparticles improved effectiveness and safety of cisplatin for lung cancer therapy. Int. J. Pharm. 633, 122587 (2023).
Ismail, J. et al. PEG-Lipid-PLGA hybrid particles for targeted delivery of anti-inflammatory drugs. Pharmaceutics. 16(2), 187 (2024).
Farid, A., Michael, V. & Safwat, G. Melatonin loaded poly(lactic-co-glycolic acid) (PLGA) nanoparticles reduce inflammation, inhibit apoptosis and protect rat’s liver from the hazardous effects of CCL4. Sci. Rep. 13, 16424 (2023).
Sun, Y. et al. Cell permeable NBD peptide-modified liposomes by hyaluronic acid coating for the synergistic targeted therapy of metastatic inflammatory breast cancer. Mol. Pharm. 16, 1140–1155 (2019).
Hussein, R. M. et al. Effect of quercetin-loaded poly (lactic-co-glycolic) acid nanoparticles on lipopolysaccharide-induced memory decline, oxidative stress, amyloidogenesis, neurotransmission, and Nrf2/HO-1 expression. Heliyon 10, e23527 (2023).
Wan, S. et al. Resveratrol-loaded PLGA nanoparticles: enhanced stability, solubility and bioactivity of resveratrol for non-alcoholic fatty liver disease therapy. R. Soc. Open Sci. 6, 182173 (2019).
Zhang, H. et al. Development of hierarchical porous bioceramic scaffolds with controlled micro/nano surface topography for accelerating bone regeneration. Mater. Sci. Eng. C Mater. Biol. Appl. 130, 112437 (2021).
Yu, H. et al. Isoginkgetin-loaded reactive oxygen species scavenging nanoparticles ameliorate intervertebral disc degeneration via enhancing autophagy in nucleus pulposus cells. J. Nanobiotechnol. 21, 99 (2023).
Sugitha, S. K. J. et al. Green synthesis of Al-ZnO nanoparticles using Cucumis maderaspatanus plant extracts: analysis of structural, antioxidant, and antibacterial activities. Nanomaterials (Basel) 14, 1851 (2024).
García-Vela, S. et al. Antimicrobial peptides from porcine blood cruor hydrolysates as a promising source of antifungal activity. Foods 14, 8 (2024).
Yang, L. et al. Dual-dynamic-bond cross-linked injectable hydrogel of multifunction for intervertebral disc degeneration therapy. J. Nanobiotechnol. 20, 433 (2022).
Sobska, J. et al. Counterion-insulated near-infrared dyes in biodegradable polymer nanoparticles for in vivo imaging. Nanoscale Adv. 4, 39–48 (2021).
Mohseni, M. et al. The clinical use of curcumin on neurological disorders: An updated systematic review of clinical trials. Phytother. Res. 35, 6862–6882 (2021).
Kobori, M. et al. Quercetin suppresses immune cell accumulation and improves mitochondrial gene expression in adipose tissue of diet-induced obese mice. Mol. Nutr. Food Res. 60, 300–312 (2016).
Smilkov, K. et al. Piperine: old spice and new nutraceutical?. Curr. Pharm. Des. 25, 1729–1739 (2019).
Anand, P. et al. Design of curcumin-loaded PLGA nanoparticles formulation with enhanced cellular uptake, and increased bioactivity in vitro and superior bioavailability in vivo. Biochem. Pharmacol. 15, 102–143 (2016).
Chen, Z. et al. Multifaceted role of phyto-derived polyphenols in nanodrug delivery systems. Adv. Drug Deliv. Rev. 176, 113870 (2021).
Prasad, S. et al. Curcumin, a component of golden spice: from bedside to bench and back. Biotechnol. Adv. 32, 1053–1064 (2014).
Beconcini, D. et al. Cherry extract from prunus avium l. to improve the resistance of endothelial cells to oxidative stress: mucoadhesive chitosan vs. poly(lactic-co-glycolic acid) nanoparticles. Int. J. Mol. Sci. 20, 1759 (2019).
Xie, X. et al. PLGA nanoparticles improve the oral bioavailability of curcumin in rats: characterizations and mechanisms. J. Agric. Food Chem. 59, 9280–9289 (2011).
Seke, F. et al. Effect of freeze drying and simulated gastrointestinal digestion on phenolic metabolites and antioxidant property of the natal plum (Carissa macrocarpa). Foods 10, 1420 (2021).
Baier, G. et al. BSA adsorption on differently charged polystyrene nanoparticles using isothermal titration calorimetry and the influence on cellular uptake. Macromol. Biosci. 11, 628–638 (2011).
Merhi, M. et al. Study of serum interaction with a cationic nanoparticle: Implications for in vitro endocytosis, cytotoxicity and genotox- icity. Int. J. Pharm. 423, 37–44 (2012).
Oh, N. & Park, J. H. Endocytosis and exocytosis of nanoparticles in mammalian cells. Int. J. Nanomed. 9, 51–63 (2014).
Chen, X. et al. Curcumin-loaded mPEG-PLGA nanoparticles attenuates the apoptosis and corticosteroid resistance induced by cigarette smoke extract. Front. Pharmacol. 13, 824652 (2022).
Feltrin, F. D. S. et al. Selection and control of process conditions enable the preparation of curcumin-loaded poly(lactic-co-glycolic acid) nanoparticles of superior performance. ACS Appl. Mater. Interfaces 15, 26496–26509 (2023).
Hussein, R. M. et al. Effect of quercetin-loaded poly (lactic-co-glycolic) acid nanoparticles on lipopolysaccharide-induced memory decline, oxidative stress, amyloidogenesis, neurotransmission, and Nrf2/HO-1 expression. Heliyon 10, e23527 (2023).
Guineo-Alvarado, J. et al. Degree of crosslinking in β-cyclodextrin-based nanosponges and their effect on piperine encapsulation. Food Chem. 340, 128132 (2021).
Ramazani, F. et al. Strategies for encapsulation of small hydrophilic and amphiphilic drugs in PLGA microspheres: state-of-the-art and challenges. Int. J. Pharm. 499, 358–367 (2016).
Sokol, M. B. et al. Pharmaceutical approach to develop novel photosensitizer nanoformulation: an example of design and characterization rationale of chlorophyll α derivative. Pharmaceutics 16, 126 (2024).
Bhattacharjee, S. Understanding the burst release phenomenon: toward designing effective nanoparticulate drug-delivery systems. Ther. Deliv. 12, 21–36 (2021).
Wang, B. et al. Paclitaxel and etoposide co-loaded polymeric nanoparticles for the effective combination therapy against human osteosarcoma. J. Nanobiotechnol. 13, 22 (2015).
Gupta, S. C., Patchva, S. & Aggarwal, B. B. Therapeutic roles of curcumin: lessons learned from clinical trials. AAPS J. 15, 195–218 (2013).
Lu, X. et al. Development of L-carnosine functionalized iron oxide nanoparticles loaded with dexamethasone for simultaneous therapeutic potential of blood brain barrier crossing and ischemic stroke treatment. Drug Deli. 28, 380–389 (2021).
Kho, W. et al. Strain-specific anti-biofilm and antibiotic-potentiating activity of 3’,4’-difluoroquercetin. Sci. Rep. 10, 14162 (2020).
Chittasupho, C. et al. Effects of quercetin and curcumin combination on antibacterial, antioxidant, in vitro wound healing and migration of human dermal fibroblast cells. Int. J. Mol. Sci. 23, 142 (2021).
Yuan, G. et al. Antibacterial activity and mechanism of plant flavonoids to gram-positive bacteria predicted from their lipophilicities. Sci. Rep. 11, 10471 (2021).
Cushnie, T. P. & Lamb, A. J. Antimicrobial activity of flavonoids. Int. J. Antimicrob. Agents 27, 181 (2006).
Adamczak, A., Ożarowski, M. & Karpiński, T. M. Curcumin, a natural antimicrobial agent with strain-specific activity. Pharmaceuticals (Basel) 13, 153 (2020).
Acknowledgements
The experiments in this study were conducted at the Ningxia Institute of Bone and Joint Diseases. The authors would like to express their gratitude to the laboratory for providing the experimental facilities and equipment.
Funding
This study was supported by the Key Research and Development Programme of Ningxia Hui Autonomous Region, China (Project No. 2021BEG02037), the Natural Science Foundation of Ningxia (Project No. 2021AAC03396) and the Natural Science Foundation of Ningxia (Project No. 2025AAC030863).
Author information
Authors and Affiliations
Contributions
Conceptualization, Zhihai Zhang and Hui Ma; Funding acquisition, Zhidong Lu; Methodology, Zhihai Zhang and Hui Ma; Project administration, Zhidong Lu; Supervision, Zhidong Lu; Writing–original draft, Zhihai Zhang; Writing–review & editing, Mengxiao Tantai, Sihao Yu, Yi Zhang, Junliang Song, Gangning Feng and Fei Liu.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, Z., Tantai, M., Ma, H. et al. Synthesis and performance optimization of multi-bioactive compound-loaded nanocarriers for antimicrobial applications. Sci Rep 15, 41324 (2025). https://doi.org/10.1038/s41598-025-25043-y
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-25043-y