Abstract
Supervised learning has excellent segmentation performance in bladder tumor segmentation, but it relies on a large amount of labeled data. To learn the features of bladder tumors from limited labeled data and obtain accurate segmentation results, we propose a semi-supervised segmentation method for bladder tumors, UDS-MT. The method consists of a Mean Teacher network and a guided branch, which respectively undertake the tasks of segmentation prediction and supervision of prediction results. Mean teacher network uses the exponential moving average (EMA) mechanism to update the teacher network parameters to achieve fine-grained capture of the target shape; the guided branch uses uncertainty estimation to filter out pixel blocks with high confidence to obtain more reliable masks, and it suppresses overfitting on some certain extent. In addition, we propose a defend loss term that only calculates the loss for pixels with high prediction confidence of the model, thereby improving the reliability of the pseudo-label. After evaluation on a bladder tumor clinical medical image dataset, when the labeled data is limited to 15%, the Dice coefficient of the network segmentation target shape can reach up to 80.04%, which is at least 2.81% higher than other methods.
Similar content being viewed by others
Introduction
The pathogenesis of bladder cancer may be closely related to factors such as advanced age, male sex, smoking, and family history of bladder cancer1. Since the mid-2000s, bladder cancer has become an important public health issue; the rate of decline in the incidence of bladder cancer has changed2. Its rising incidence and high medical costs have brought heavy burdens to society and patients3.
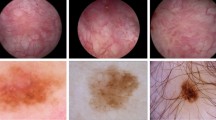
In order to achieve early detection and effective management of bladder cancer, cystoscopy and artificial intelligence are widely used in bladder cancer diagnosis4,5,6,7. However, bladder tumors, especially early-stage ones, are often small (≤ 5 mm) or hidden in bladder folds, diverticula, or near the ureteral orifice. Moreover, the class differences between lesion regions and normal regions are small8. Segmentation models need to accurately capture these subtle lesions to meet clinical requirements. Obtaining pixel-level annotations for cystoscopic images is not only costly but also time-consuming9. In recent years, 10 have shown that semi-supervised learning still has defects in the bladder cancer segmentation task. As shown in Fig. 1, the pixel ratio of tiny tumors in cystoscopic images is extremely small, and the features are not obvious, such as (a) and (b). There are also images with blurred boundaries and unclear features of the target area, such as (c) and (d). This situation depends on the judgment of professional doctors.
Some samples of the bladder tumor dataset.
This study aims to make full use of tiny unlabeled images, improve the accuracy of bladder tumor segmentation, and obtain more reliable results that better match the ground truth. The main contributions of this study are as follows:
(1) We propose a semi-supervised segmentation method for bladder tumors, UDS-MT, which integrates the Mean Teacher model and the guided branch, effectively solves the problems of limited labeled data, and significantly improves the performance of bladder tumor segmentation.
(2) The guided branch uses uncertainty estimation to generate masks, which helps to obtain high-quality pseudo labels andsuppresses overfitting.
(3) We propose a defend loss term that only calculates the loss for pixels with high prediction confidence of the model, which promotes the model to learn the reality of the target area, further improving the accuracy of segmentation prediction.
(4) Through comparative experiments on bladder tumor clinical image datasets, this method has demonstrated excellent segmentation results. Comparative experiments on the public colonoscopy dataset have verified that our method has generalization ability and robustness. Finally, ablation studies have been conducted to verify the role of each module.
Related work
Semi-supervised learning
Self-training architectures15 (such as local contrast loss16, uncertainty-guided ensemble self-training17,18 and co-training frameworks19,20 (such as deep co-training21, attention-generated adversarial networks22, integrating adversarial training and co-consistent learning23 are used to generate high-quality pseudo-labels for segmentation tasks, especially lesion segmentation in breast ultrasound images. These methods improve the reliability of pseudo-labels and segmentation performance through information exchange and uncertainty-aware mechanisms between models24. Consistency regularization25,26 (such as the π model27, time series combination model27, mean teacher model28 improves the robustness and consistency of the model through data augmentation and temporal prediction. Methods that combine pseudo-labels and consistency regularization (such as the cross-teaching consistency method29, MC-Net + network26, cross-consistency training30 further improve segmentation performance, but there is a challenge in synchronizing prediction results. Generative adversarial networks (GANs)31 improve segmentation performance by alternating updates of the discriminator and generator, but do not fully consider the preservation of multi-scale features and boundary detail information.
Semi-supervised learning has been developed in recent years10,11,12,13,14, such as mask learning (CGML)32, contrast agent pseudo-supervision task33, mean teacher model guided by triple uncertainty and contrastive learning34, shape-aware network35, Bayesian pseudo label36, etc., aiming to improve pseudo-label quality and segmentation performance. By introducing perturbations and consistency regularization to enhance the learning of unreliable regions, the Mismatch37 method incorporates differentiated morphological feature perturbations at the feature level, thereby improving the segmentation performance of the model when labeled data is limited. The method introduces a weak-to-strong perturbation strategy38 and the corresponding feature perturbation consistency loss to efficiently utilize unlabeled data and guide the model to learn reliable regions.
Existing methods have made progress in generating high-quality pseudo-labels and improving segmentation performance, but there are still challenges in synchronizing model prediction, preserving boundary and detail information, and fusing multi-scale features.
Mean teacher model
Tarvainen et al.28 first proposed the mean teacher model in 2017, which has attracted widespread attention as an effective semi-supervised learning method. By performing exponential moving averaging (EMA) on the parameters of the teacher model and using consistency regularization to constrain the student model, the student model’s prediction results for labeled and unlabeled data are consistent.
Wang et al.39 introduced the mean teacher model into the area of medical image segmentation. To address the common problems of class imbalance and insufficient labeled data in medical images, this model can generate high-quality soft labels using little labeled data and a large amount of unlabeled data. Through experiments in medical image segmentation tasks, such as lungs and liver, the effectiveness and advantages of the mean teacher model in this field were verified. Based on the original mean teacher model, Liu et al.40 improved the structure of the teacher model and the student model and the calculation method of the consistency loss to improve the segmentation accuracy, especially when dealing with complex lesion areas and small target segmentation tasks. Li et al.41 proposed a multi-scale mean teacher model for extracting multi-scale image features. The model performs well in segmenting lesions of different sizes and shapes. Zhu et al.42 introduce 2D/3D hybrid dual-teacher and uncertainty weighted regularization to achieve approximate full supervision segmentation accuracy on very few labeled MRI data. Xiao et al.43 integrate local CNN details with Transformer global features on mean teacher, and significantly improve semi-supervised cardiac segmentation performance through cross-pseudo-labeling and uncertainty screening.
These studies demonstrate the potential of the mean teacher model in medical image segmentation, and our approach extends the Mean Teacher framework, introduces supervised branches, and integrates uncertainty estimation and consistency regularization to further improve the model’s performance.
Uncertainty estimation
Uncertainty estimation plays an increasingly important role in semi-supervised medical image segmentation work. Hua et al.44 proposed a method based on uncertainty-driven. The uncertainty estimation can effectively guide the model to focus on difficult-to-classify areas, significantly improving the performance of the segmentation model when labeled data is scarce. Zhu et al.42 incorporated uncertainty perception into the mean teacher model and found that the uncertainty perception mechanism can enable the model to better utilize unlabeled data. Wei et al.45 proposed an uncertainty-based multi-scale feature learning method. The model learns and filters multi-scale features through uncertainty estimation, which can better capture the characteristics of lesion areas of different sizes and shapes in the image. Wang et al.46 constructed a framework based on uncertainty-guided adversarial training. Uncertainty estimation is used to guide the training process of the generative adversarial network, and the images generated by the generator are adversarially learned from the real images based on the uncertainty features.
Uncertainty estimation shows strong potential in semi-supervised medical image segmentation. By combining it with semi-supervised learning methods, it can effectively utilize unlabeled data and improve the performance of segmentation models.
Method
Overview
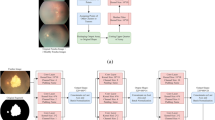
We propose a semi-supervised learning method, as shown in Fig. 2, which uses the guided branch to guide the generation of masks and jointly trains the supervised branch and the Mean Teacher model to fully utilize unlabeled data. To enhance the reliability of pseudo-labels, we calculate the loss by combining the masks with the pseudo-labels of the student model and the teacher model, respectively.
We propose the semi-supervised segmentation method : (1) For labeled data, the pseudo-labels generated by Model-G and Model-S are mutually constrained for consistency, and they are trained under the supervision of real labels to obtain the initial parameters of the model; (2) For unlabeled data, the pseudo-labels generated by Model-G generate masks under uncertain estimation, and the masksare used as supervisory signals to measure the reliability of the pseudo-labels of Model-S and Model-T.
We explain the work of the method as follows: Let \(\:D={D}_{l}+{D}_{u}\) represent the entire dataset, where \(\:{D}_{l}\) and \(\:{D}_{u}\) respectively represent the labeled dataset and the unlabeled dataset. We use \(\:\left({x}_{i},{y}_{i}\right)\in\:{D}_{l}\) to denote the labeled image pair, where \(\:{y}_{i}\) is the ground truth. The unlabeled image is represented as \(\:{u}_{j}\)∈\(\:{D}_{u}\). In our method, Model-T implements the segmentation task, and Model-G is used to participate in the supervision of the predicted probability of labeled and unlabeled images for each batch. The encoder and decoder of Model-G, Model-S, and Model-T have the same structure but with independent parameters. It is obvious that Pyramid Vision Transformer (PVT) encoder combines Transformer’s global modeling ability and CNN’s multi-scale feature advantages to have efficient feature extraction capabilities. As shown in Fig. 3, the model uses PVT encoder to extract four-layer feature information, and uses CFM to cross the feature information of the last three layers to reduce the differences between features and avoid redundant information. Prevent fuzzy feature information from being filtered, especially for tiny segmentation targets or segmentation boundaries of fuzzy pictures. We connect the features of the first layer with the crossed features to obtain the prediction probability of the model.
Network backbone flow work: We extract four-layer feature information through the PVT encoder, and use Cross Feature Module (CFM) to cross the feature information of the last three layers to reduce the differences between features and avoid redundant information. To prevent fuzzy feature information from being filtered, we connect the features of the first layer to obtain the prediction probability of the model.
Let \(\:\theta\:\) and\(\:{\theta\:}^{{\prime\:}}\) correspond to the weight parameters of Model-S and Model-T, respectively. Model-T weight parameters are updated through the EMA, that is, they are updated in combination with the previous network weight parameters \(\:\theta\:\) and the smoothing coefficient α, where t represents the training time.
When using labeled data, the labeled data is processed by Model-G and Model-S to extract the target features, and the output results are constrained by consistency loss. \(\:{l}_{sc}\) and supervision loss \(\:{l}_{s1\:\:}\)and \(\:{l}_{s2}\). When an unlabeled image is input, Model-G, Model-S, and Model-T are processed simultaneously to generate the prediction results, and the consistency loss and cross entropy loss constraints are imposed on each other, expressed by \(\:{l}_{c1}\)、\(\:{l}_{c2}\) and \(\:{l}_{c3}\). In order to reduce the coupling between models, we introduce uncertainty estimation in Model-G, generate pseudo labels and masks from the prediction results, and calculate the loss of the segmentation results of Model-S and Model-T under the constraint of the mask.
Progressive increase strategy of consistency weight
Consistency regularization is a method that calculates the loss by measuring the consistency between the prediction results of the student model and the teacher model in semi-supervised learning, thereby constraining the prediction behavior of the student model on labeled data and unlabeled data. During the training process, the consistency weight will gradually increase, so that the model mainly relies on labeled data for learning in the early stage of training, and gradually uses more information from unlabeled data in the later stage of training. When the labeled data is limited, this method can help the model fully explore the valuable information in unlabeled data and improve the generalization ability of the model.
Uncertainty estimation
In this method, so as to improve the quality of the target shape segmentation results, the method introduces an uncertainty estimation mechanism. By estimating the uncertainty of the prediction results of the unlabeled data in the supervised branch to guide the training and generate a mask, the mask guides the prediction results of other branches to perform pixel constraints. This strategy can fully mine reliable information in unlabeled data, thereby further improving model performance.
Based on the uncertainty estimation of information entropy, it can not only directly quantify ambiguity in probabilistic model outputs, but also captures both aleatoric and epistemic uncertainty, which works for fuzzy information caused by noise in cystoscopic images. It is computationally efficient and easy to integrate into training/workflows.
Loss function
We design a loss function to take into account both supervised and unsupervised learning scenarios, and enhance the performance of the semi-supervised segmentation work by combining consistency regularization, cross-entropy loss, and Dice loss. We denote the size of the input image by H×W, and the position of a pixel block in the image by \(\:\left(h,w\right)\). The loss function of the overall method is divided into two parts: one is the loss function for labeled data, and the other is the loss function for unlabeled data.
The loss function for labeled data \(\:{\varvec{L}}_{1}\)
The loss function for labeled data includes ① the supervision loss \(\:{l}_{s}\) between the prediction results of Model-G and Model-S and the true label, and ② the consistency loss \(\:{l}_{sc}\) between the prediction results of Model-G and the student model.
Dice loss is calculated under the constraint of the true label to measure the accuracy between the prediction result and the true label, while the consistency loss is evaluated by cross-entropy loss to measure the probability distribution difference between the two prediction results.
Unlabeled data loss function \(\:{\varvec{L}}_{2}\)
Due to the lack of true labels of manually annotated unsupervised data, the prediction results of Model-G provide pseudo-label supervision signals for the prediction results of Model-S and Model-T under the action of the softmax function. Unlabeled data loss includes ① consistency loss \(\:{l}_{c}\) between the prediction results of the Model-G, Model-S, and Model-T, and ② supervised information loss \(\:{l}_{p}\).
\(\:{l}_{c}\:\)is subjected to consistency loss to constrain the prediction results of the model under different conditions to the greatest extent possible.
We use the defend loss \(\:{l}_{p}\), combined with uncertainty estimation, to expand cross-entropy loss.
Pseudo-label supervision
Pseudo-labels \(\:{Y}_{g}\left({u}_{j}\right)\) are generated through the results of the Model-G \(\:{P}_{g}\left({u}_{j}\right)\) to guide the training of unlabeled images of Model-S and Model-T. The generation method is.
Uncertainty estimation
Since the pseudo-labels generated by the guide branch lack additional constraint information, their reliability may be low. In order to improve the reliability of pseudo-labels, we introduced an uncertainty estimation method based on information entropy. Specifically, when the probability distribution of the prediction results is more uniform, the information entropy is larger, indicating higher uncertainty; when the probability distribution is more concentrated, the information entropy is smaller, indicating lower uncertainty. We quantify uncertainty by calculating the information entropy of the pseudo-labels generated by the guide branch. The higher the entropy value, the more uncertain the model’s prediction for that area.
To filter out areas with high certainty of pseudo-labels, we set a dynamic threshold (\(\:{T}_{threshold}\)), which can flexibly filter out areas where the model predicts reliably according to different training stages.
Where \(\:T\) indicates the total number of training sessions, and \(\:t\) indicates the current number of iterations.
Areas with higher uncertainty are considered unreliable. We filter out areas with higher reliability as masks based on the threshold. \(\:M\) denotes the number of elements in the set \(\:\phi\:\) of pixels selected by the mask. Using pseudo-labels as comparison results and masks as supervision signals, the supervised information loss of the prediction results of Model-S and Model-T is measured.
The weight of the consistency loss \(\:{\upgamma\:}\) gradually increases during the training process, which makes the model mainly use the labeled data for learning in the early stage of training, and gradually use more information from the unlabeled data in the later stage of training. This design can effectively play the role of unlabeled data, thereby improving the generalization ability and segmentation accuracy of the network.
The total loss function combines the loss function of the labeled data and the unlabeled data. By dynamically adjusting the consistency weight, the role of labeled data and unlabeled data is balanced.
Dataset
The experiment used the cystoscopy dataset provided by the Affiliated Hospital of Anhui Medical University and the open access polyp datasets Kvasir-SEG, CVC-ColonDB, and ETIS-LaribPolypDB for comparative experiments to verify the stability of the network. These datasets are authentic and validated by experienced experts, and sample images are shown in Fig. 4.
Sample images from each dataset.
(1) Bladder tumor dataset: This dataset includes 1948 images of different bladder areas with a resolution of 1920 × 1072. Cystoscopy images are pivotal for diagnosing bladder cancer, which has a high recurrence rate. Tumor margins often lack distinct boundaries, and blood/mucus interference degrades image quality, while rare tumors require careful sampling to avoid bias.
(2) Kvasir-SEG47: The dataset is a critical dataset for colorectal polyp segmentation and contains resolutions ranging from 332 × 487 to 1920 × 1072. It has 1000 images with different polyp regions. Polyps vary in size, shape, and texture, with flat lesions blending into the mucosal background. The dataset was expanded with a subset of 196 challenging flat polyps, enhancing its utility for real-world clinical scenarios.
(3) CVC - ColonDB dataset 48: The dataset has a resolution of 500 × 573 and contains 300 images extracted from 13 polyp video sequences. It is foundational for polyp detection research.
(4) ETIS - LaribPolpyDB dataset49: The dataset has a resolution of 1225 × 966 and contains 196 images. Mucosal folds and vascular patterns often obscure polyp boundaries, requiring models to learn subtle texture differences. With fewer images compared to Kvasir-Seg, overfitting is a risk.
When training different datasets, we use a 6:2:2 ratio to split images between training, validation, and testing. On the comparison dataset, we used the same evaluation metrics.
Evaluation metrics
We use 6 indicators to evaluate, including: Dice coefficient (Dice), sensitivity (sm), mean Intersection over Union (mIoU), specificity (em), Mean Absolute Error (MAE), and Accuracy (Acc).
Detailed demonstration
We use the PyTorch library to implement our method. We use the SGD optimizer, and initialize the learning rate to 0.001 and decay it by a factor of \(\:\frac{epoch}{({n}_{epoch}{)}^{0.9}}\) In 500 epochs of training networks. In our experimental setting, the images processed at each step contain two labeled data and two unlabeled data. All experiments are run on the experimental cloud platform.
Experiments and results
Our method is compared and evaluated with several state-of-the-art methods. These methods include: Mean Teacher framework (MT)28, as well as the advanced Uncertainty-Aware Mean teacher (UAMT)17, the Dual Task Consistency Strategy (DTC)50 that cleverly combines two complementary tasks, the Polyp-Aware Mixture and Dual-Level Consistency Regularization (PolypMix)51, the Twin Student Mean teacher (DSST)52,weak-to-strong perturbation consistency(W2sPC)38, and Expectation maximization pseudo labels(EMSSL)53. These network method comparisons are crucial for evaluating the performance of UDS-MT. In these experiments, 15% of the training data is randomly selected as labeled data.
We implemented these methods using published code and following the same settings. For data augmentation, we apply the same technique, performing simple resizing, random flipping, and rotation. Similarly, we use the same decoder and PVT encoder for all method backbones in the experiments.
Bladder tumor dataset segmentation results
Our method is compared with the fully supervised method. This comparison is important for evaluating the performance difference between our UDS-MT method and the fully supervised method. We train a fully supervised network using the same backbone network on 187(15%), 374(30%), and 1246(100%) labeled images, respectively, denoted as supervised. We use the same backbone network to demonstrate the effectiveness of the method.
As shown in Table 1, when the labeled data accounts for 30%, the results of fully supervised methods are similar to those of our method under the 15% limit. When the labeled data is only 15% of the bladder tumor training set, compared with other semi-supervised methods, Dice is improved by at least 2.81%, mIoU is improved by at least 3.48%, sm and em are also improved to varying degrees, MAE is reduced to 5.18%, and accuracy is also improved.
There are two main reasons for the improvement in the evaluation index effect. First, the deep learning methods54,55 rely heavily on the large number and high quality of the training set, and the limited labeled dataset limits the performance of the fully supervised. Our method utilizes a large amount of unlabeled images, and through techniques such as pseudo-label generation and consistency regularization, the model can learn richer feature representations under limited labeled data. This strategy effectively alleviates the problem of insufficient labeled data and improves the efficiency of the model in utilizing unlabeled data, thereby achieving significant improvements in segmentation accuracy. However, in the absence of labeled data, overfitting may occur. By introducing uncertainty estimation and consistency loss, we not only improve the accuracy of segmentation but also enhance the robustness of the model to noise and outliers. This robustness enables the model to output high-quality segmentation results more stably when facing complex bladder tumor images, further improving the performance of key indicators such as Dice and mIoU. At the same time, the improvement of sm and Em indicators shows that the model also performs well in structure preservation and edge detection, while the reduction of MAE shows that the model has improved to a certain extent in prediction accuracy.
As shown in Fig. 5, we provide visualization results of different methods under a limited dataset of 15%. Test results of (a) and (b) indicate that our method works in the early diagnosis of tiny bladder tumors. The visualization results in the figure show that our method can provide relatively accurate results under limited labeled data. Whether it is a small tumor, a blurry low-quality image, or an image with highlights, it shows good segmentation results.
Test results of different methods under 15% limited bladder tumor dataset training.
Comparative dataset segmentation results
In order to verify the effectiveness of our method in the bladder tumor segmentation task, we conducted extensive comparative experiments on the publicly accessible polyp dataset. We followed the same settings as the bladder tumor dataset and used 15% of the training set for experiments.
The Kvasir-SEG dataset shows a variety of polyp morphologies and imaging conditions, making it a very challenging test platform. According to the data partition ratio, we randomly selected 90 (15%) and 180 (30%) images as labeled data. We simulated the actual situation with limited labeled data to evaluate the effectiveness of our method. The results are shown in Table 2.
The ETIS_LaribPolypDB dataset contains 196 images. According to the same training set division as the bladder tumor dataset, the data involved in the training consists of 118 images, of which 18 are used as labeled data, and the other 100 images are used as test sets for testing. Test results are shown in Table 3.
The CVC-conlonDB dataset contains 300 images. According to the same training set division as the bladder tumor dataset, there are 180 images involved in the training data, of which 27 are used as labeled data, and the other 120 images are used as test sets for testing. The test results are shown in Table 4.
Table 2 Tables 3 and 4 show the test results on the comparison dataset, respectively. Judging from the results, although our method cannot guarantee the optimality of each indicator, the Dice coefficient of the overlap between the predicted results and the real label has been improved, which shows that our method has generalization ability and robustness.
We visualize the Dice results of each dataset under different methods, visually demonstrating the advantages of our method. As shown in Fig. 6.
Visualization of Dice results with different datasets using different methods on a limited dataset of 15%.
Combined with Figs. 5 and 6, we can see that the method works. Uncertainty estimation based on information entropy makes the pseudo-labels generated by the segmentation model clearer in the target area and also filters some of the redundant information. During the iteration process, the accurate feature information is extracted and added to the next iteration training, and the accuracy of the segmentation result is further improved. Reliable feature information plays an important role in segmenting image blur or small tumors. Moreover, the consistency constraints between pseudo-labels suppress the occurrence of overfitting to a certain extent, making the method robust.
In order to see the segmentation effect more intuitively, we select two pictures from each comparison dataset and display the segmentation results, as shown in Fig. 7.
We select six of the three data segmentation results of Kvasir-SEG, CVC-ColonDB, and ETIS-LaribPolypDB for display.
Test results of 30% limited dataset
So as to improve the effectiveness of our method verification, we tested different semi-supervised methods on the bladder tumor dataset and Kvasir-SEG dataset with 30% limited annotations. The result percentages are shown in Tables 5 and 6. The results show that with 30% limited data, our method still has better segmentation ability over other methods. On a limited dataset of 30%, the visualization of each method is shown in Fig. 8, which verifies that our methods have more advantages than other methods on limited datasets.
Visualization of Dice results on bladder dataset and Kvasir-SEG dataset using various methods on a limited dataset of 30%.
With 30% limited annotations, the testing results of our method on the bladder dataset and Kvasir-SEG dataset still have certain advantages. The ability to segment the target area is the best compared to other methods. In Fig. 8, we can see it intuitively.
Computational costs
In this section, we study the computational cost of UDS-MT in comparison with other semi-supervised methods. We use two key metrics to evaluate the computational cost: the single image time (ITime) for the test set to pass the network for image segmentation and the total training time (TTime) required to train the network on the training set.
Table 7 shows the comparison results of computational cost. Because additional inference is introduced on the mean teacher, our method takes more time to train than the mean teacher. But it shows similar inference cost, which indicates that we can make improvements in module lightweighting. It also shows that our method can be a viable candidate for clinical applications as the inference cost is less than the clinically required 30ms.
Ablation analysis
To explore the impact of small modules on the overall network performance, we conducted experiments by training the model and removing or modifying some parts. The experiments followed the settings described in the network and used the bladder tumor dataset for ablation studies. The network used the mean teacher framework as a baseline, where Model-S and the Model-T use the same backbone network structure but do not share it. As shown in Table 8, adding only the Model-G to the mean teacher base network improves Dice by 2.56% and mIoU by 2.35%; introducing only uncertainty estimation(As shown in Fig. 9) improves Dice by 0.28% and slightly reduces mIoU. In addition, our network improves performance by combining uncertainty estimation with consistency regularization and the introduction of pseudo-labels, resulting in an increase of 3.27% and 3.28% in dice and mIoU.
The uncertainty estimation generates a mask to supervise the Mean Teacher’s segmentation prediction: Model-G, Model-T, and Model-S generate prediction probabilities. The prediction probabilities of multiple forward propagations on the Model-G are calculated to get uncertainty, and the high-confidence area is selected as the mask. The pixel block loss of Model-T and Model-S’s pseudo-labels is calculated under the supervision of the mask.
Limits and future work
While UDS-MT achieved relatively good results for bladder tumor segmentation, there are certain limits. Compared with the truth labels, as shown in Fig. 4 the segmentation results of our method still have some differences at the edge. The pixel-level annotation itself has 1–2 pixel system errors, and it may also be that the maximum down-sampling of UDS-MT causes the thinnest edge features to be lower than the average thickness of the bladder wall, resulting in structural defects. Additional inference takes more time to train, which indicates that we can make improvements in module lightweighting. Ghost modules can be tried. Replacing standard convolution with Ghost modules, the channel will first reduce the dimensionality and then expand linearly to speed up the inference speed. Multi-scale deep supervision can also be introduced, and the decoding layer synchronously outputs edge maps without introducing additional up-sampling. Prior knowledge is also a method that accelerates the reasoning process. Focusing on edge micro information will be the main content of the future work.
Conclusion
We designed a method for semi-supervised tasks, and our experimental verification on the bladder tumor dataset, Kvasir-SEG, ETIS_LaribPolypDB and CVC-conlonDB datasets shows that our method outperforms other methods. Additionally, we also conducted ablation experiments to further verify the effectiveness of each module and highlight the superiority of this method. However, further efforts are still needed in terms of inference time and segmentation accuracy. By rationally utilizing unlabeled data, the segmentation performance is significantly improved on the bladder tumor dataset, which is expected to improve the accuracy and efficiency of bladder cancer diagnosis and benefit patients.
Data availability
The datasets generated and/or analysed during the current study are not publicly available due to the data being owned by a third party and authors do not have permission to share the data but are available from the corresponding author on reasonable request.
References
Dobruch, J. et al. Gender and bladder cancer: A collaborative review of Etiology, Biology, and outcomes. Eur. Urol. 69, 300–310. https://doi.org/10.1016/j.eururo.2015.08.037 (2016).
Siegel, R. L., Kratzer, T. B., Giaquinto, A. N., Sung, H. & Jemal, A. Cancer statistics, 2025. Ca-a Cancer J. Clin. 75, 10–45. https://doi.org/10.3322/caac.21871 (2025).
Jubber, I. et al. Epidemiology of bladder cancer in 2023: A systematic review of risk factors. Eur. Urol. 84, 176–190. https://doi.org/10.1016/j.eururo.2023.03.029 (2023).
Nicolas, J. et al. Deep learning algorithms for multi-region bladder cancer segmentation. J. Urol. 209, E112–E112 (2023).
Wu, E. et al. Deep learning approach for assessment of bladder cancer treatment response. Tomography 5, 201–208. https://doi.org/10.18383/j.tom.2018.00036 (2019).
Lee, M. C. et al. Development of deep learning with RDA U-Net network for bladder cancer segmentation. Cancers 15 https://doi.org/10.3390/cancers15041343(2023).
Zheng, Z. et al. Pathology-based deep learning features for predicting basal and luminal subtypes in bladder cancer. Bmc Cancer. 25 https://doi.org/10.1186/s12885-025-13688-x (2025).
Wenger, K. Semi-Supervised Learning Approach for Bladder Cancer Diagnosis (Toronto Metropolitan University, 2021).
Wenger, K. et al. A semi-supervised learning approach for bladder cancer grading. Mach. Learn. Appl. 9, 100347 (2022).
Xia, Q. et al. A comprehensive review of deep learning for medical image segmentation. Neurocomputing, 128740 (2024).
Mittal, S., Tatarchenko, M. & Brox, T. J. I. t. o. p. a. & intelligence, m. Semi-supervised semantic segmentation with high-and low-level consistency. IEEE Trans. Pattern Anal. Mach. Intell. 43, 1369–1379 (2019).
Han, K. et al. Deep semi-supervised learning for medical image segmentation: A review. Expert Syst. Appl. 245 https://doi.org/10.1016/j.eswa.2023.123052 (2024).
Yu, X., Ma, Q., Ling, T., Zhu, J. & Shi, Y. Enhancing semi-supervised medical image segmentation with bidirectional copy-paste and masked image reconstruction. Int. J. Mach. Learn. Cybernet. 16, 2603–2613. https://doi.org/10.1007/s13042-024-02410-1 (2025).
Peng, J. & Wang, Y. Medical image segmentation with limited supervision: A review of deep network models. Ieee Access. 9, 36827–36851. https://doi.org/10.1109/access.2021.3062380 (2021).
Amini, M. R. et al. Self-training: A survey. Neurocomputing 616 https://doi.org/10.1016/j.neucom.2024.128904 (2025).
Chaitanya, K., Erdil, E., Karani, N. & Konukoglu, E. Local contrastive loss with pseudo-label based self-training for semi-supervised medical image segmentation. Med. Image. Anal. 87 https://doi.org/10.1016/j.media.2023.102792 (2023).
Yu, L., Wang, S., Li, X., Fu, C. W. & Pheng-Ann, H. Uncertainty-Aware Self-ensembling Model for Semi-supervised 3D Left Atrium Segmentation in 10th International Workshop on Machine Learning in Medical Imaging (MLMI) / 22nd International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI). 605–613 (2019).
Zhang, Y., Gong, Z., Zhao, X. & Yao, W. Uncertainty guided ensemble self-training for semi-supervised global field reconstruction. Complex. Intell. Syst. 10, 469–483. https://doi.org/10.1007/s40747-023-01167-4 (2024).
Chen, B. et al. Debiased Self-Training for Semi-Supervised Learning in 36th Conference on Neural Information Processing Systems (NeurIPS). (2022).
Liu, X. et al. ACT: Semi-supervised Domain-Adaptive Medical Image Segmentation with Asymmetric Co-training in 25th International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI). 66–76 (2022).
Peng, J., Estrada, G., Pedersoli, M. & Desrosiers, C. Deep co-training for semi-supervised image segmentation. Pattern Recogn. 107 https://doi.org/10.1016/j.patcog.2020.107269 (2020).
Han, L. et al. Semi-supervised segmentation of lesion from breast ultrasound images with attentional generative adversarial network. Comput. Methods Programs Biomed. 189 https://doi.org/10.1016/j.cmpb.2019.105275 (2020).
Tang, Y., Wang, S., Qu, Y., Cui, Z. & Zhang, W. Consistency and adversarial semi-supervised learning for medical image segmentation. Comput. Biol. Med. 161, 107018. https://doi.org/10.1016/j.compbiomed.2023.107018 (2023).
Lu, L., Yin, M., Fu, L. & Yang, F. Uncertainty-aware pseudo-label and consistency for semi-supervised medical image segmentation. Biomed. Signal Process. Control. 79 https://doi.org/10.1016/j.bspc.2022.104203 (2023).
Tack, J. et al. Consistency Regularization for Adversarial Robustness in 36th AAAI Conference on Artificial Intelligence / 34th Conference on Innovative Applications of Artificial Intelligence / 12th Symposium on Educational Advances in Artificial Intelligence. 8414–8422 (2022).
Wu, Y. et al. Mutual consistency learning for semi-supervised medical image segmentation. Med. Image. Anal. 81 https://doi.org/10.1016/j.media.2022.102530 (2022).
Laine, S. & Aila, T. J. a. p. a. Temporal ensembling for semi-supervised learning. arXiv preprint arXiv:1610.02242. (2016).
Tarvainen, A. & Valpola, H. Mean teachers are better role models: Weight-averaged consistency targets improve semi-supervised deep learning results in 31st Annual Conference on Neural Information Processing Systems (NIPS). (2017).
Luo, X., Hu, M., Song, T., Wang, G. & Zhang, S. Semi-Supervised Medical Image Segmentation via Cross Teaching between CNN and Transformer in 5th International Conference on Medical Imaging with Deep Learning (MIDL). 820–833 (2022).
Ouali, Y., Hudelot, C. & Tami, M. & Ieee. Semi-Supervised Semantic Segmentation with Cross-Consistency Training in IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR). 12671–12681 (2020).
Zhu, R. Generative adversarial network and score-based generative model comparison in 2023 IEEE International Conference on Image Processing and Computer Applications (ICIPCA). 1–5 (IEEE).
Li, W., Lu, W., Chu, J., Tian, Q. & Fan, F. Confidence-guided mask learning for semi-supervised medical image segmentation. Comput. Biol. Med. 165 https://doi.org/10.1016/j.compbiomed.2023.107398 (2023).
Zhao, X. et al. Rectified contrastive Pseudo supervision for semi-supervised medical image segmentation. IEEE J. Biomedical Health Inf. 28, 251–261 (2023). Rcps.
Wang, K. et al. Tripled-Uncertainty Guided Mean Teacher Model for Semi-supervised Medical Image Segmentation in International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI). 450–460 (2021).
Liu, J., Desrosiers, C., Yu, D. & Zhou, Y. Semi-Supervised medical image segmentation using Cross-Style consistency with Shape-Aware and local context constraints. IEEE Trans. Med. Imaging. 43, 1449–1461. https://doi.org/10.1109/tmi.2023.3338269 (2024).
Xu, M. C. et al. Bayesian Pseudo Labels: Expectation Maximization for Robust and Efficient Semi-supervised Segmentation in 25th International Conference on Medical Image Computing and Computer Assisted Intervention (MICCAI). 580–590 (2022).
Xu, M. C. et al. MisMatch: calibrated segmentation via consistency on differential morphological feature perturbations with limited labels. IEEE Trans. Med. Imaging. 42, 2988–2999. https://doi.org/10.1109/tmi.2023.3273158 (2023).
Yang, Y., Sun, G., Zhang, T., Wang, R. & Su, J. Semi-supervised medical image segmentation via weak-to-strong perturbation consistency and edge-aware contrastive representation. Med. Image. Anal. 101 https://doi.org/10.1016/j.media.2024.103450 (2025).
Wang, Q., Li, X., Chen, M., Chen, L. & Chen, J. A regularization-driven mean teacher model based on semi-supervised learning for medical image segmentation. Phys. Med. Biol. 67 https://doi.org/10.1088/1361-6560/ac89c8 (2022).
Liu, L., Tian, J., Shi, Z. & Fan, J. Semi-supervised Medical Image Segmentation with Semantic Distance Distribution Consistency Learning in 5th Chinese Conference on Pattern Recognition and Computer Vision (PRCV). 323–335 (2022).
Li, H., Wang, Y. & Qiang, Y. A semi-supervised domain adaptive medical image segmentation method based on dual-level multi-scale alignment. Sci. Rep. 15, 8784. https://doi.org/10.1038/s41598-025-93824-6 (2025).
Zhu, J., Bolsterlee, B., Chow, B. V. Y., Song, Y. & Meijering, E. Hybrid dual mean-teacher network with double-uncertainty guidance for semi-supervised segmentation of magnetic resonance images. Comput. Med. Imaging Graph. 115 https://doi.org/10.1016/j.compmedimag.2024.102383 (2024).
Xiao, Z., Su, Y., Deng, Z. & Zhang, W. Efficient combination of CNN and transformer for Dual-Teacher Uncertainty-guided Semi-supervised medical image segmentation. Comput. Methods Programs Biomed. 226, 107099. https://doi.org/10.1016/j.cmpb.2022.107099 (2022).
Hua, Y., Shu, X., Wang, Z. & Zhang, L. Uncertainty-Guided Voxel-Level supervised contrastive learning for Semi-Supervised medical image segmentation. Int. J. Neural Syst. 32 https://doi.org/10.1142/s0129065722500162 (2022).
Wei, J. et al. A Semi-Supervised Multi-Region segmentation framework of bladder wall and tumor with wall-Enhanced Self-Supervised Pre-Training. Bioengineering 11, 1225 (2024).
Wang, Z. et al. Uncertainty estimation- and attention-based semi-supervised models for automatically delineate clinical target volume in CBCT images of breast cancer. Radiat. Oncol. 19, 66. https://doi.org/10.1186/s13014-024-02455-0 (2024).
Jha, D. et al. Kvasir-SEG: A Segmented Polyp Dataset in 26th International Conference on MultiMedia Modeling (MMM). 451–462 (2020).
Bernal, J. et al. WM-DOVA maps for accurate polyp highlighting in colonoscopy: validation vs. saliency maps from physicians. Comput. Med. Imaging Graph. 43, 99–111. https://doi.org/10.1016/j.compmedimag.2015.02.007 (2015).
Silva, J., Histace, A., Romain, O., Dray, X. & Granado, B. Toward embedded detection of polyps in WCE images for early diagnosis of colorectal cancer. Int. J. Comput. Assist. Radiol. Surg. 9, 283–293. https://doi.org/10.1007/s11548-013-0926-3 (2014).
Luo, X., Chen, J., Song, T. & Wang, G. & Assoc Advancement Artificial, I. Semi-supervised Medical Image Segmentation through Dual-task Consistency in 35th AAAI Conference on Artificial Intelligence / 33rd Conference on Innovative Applications of Artificial Intelligence / 11th Symposium on Educational Advances in Artificial Intelligence. 8801–8809 (2021).
Jia, X. et al. Enhancing semi-supervised polyp segmentation with polyp-aware augmentation. Comput. Biol. Med. 170 https://doi.org/10.1016/j.compbiomed.2024.108006 (2024). PolypMixNet.
Li, B., Wang, Y., Xu, Y. & Wu, C. D. S. S. T. A dual student model guided student-teacher framework for semi-supervised medical image segmentation. Biomed. Signal Process. Control. 90 https://doi.org/10.1016/j.bspc.2023.105890 (2024).
Xu, M. et al. Expectation maximisation Pseudo labels. Med. Image. Anal. 94 https://doi.org/10.1016/j.media.2024.103125 (2024).
Litjens, G. et al. A survey on deep learning in medical image analysis. Med. Image. Anal. 42, 60–88. https://doi.org/10.1016/j.media.2017.07.005 (2017).
Altaf, F., Islam, S. M. S., Akhtar, N. & Janjua, N. K. Going deep in medical image analysis: Concepts, Methods, Challenges, and future directions. Ieee Access. 7, 99540–99572. https://doi.org/10.1109/access.2019.2929365 (2019).
Author information
Authors and Affiliations
Contributions
M.L. conducted relevant investigation and data analysis, developed the model architecture and verified it through experimentaldesign and data processing, completed most of the experimental data analysis, and finally wrote and revised the manuscript. C.X. and Z.L. provided guidance on the research to ensure the quality of academic content. M.L. provides resources and helpsrefine study design and interpretation of results. Y.C.: Data Interpretation and Review.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, M., Li, Z., Xu, C. et al. Semi-supervised medical image segmentation of bladder tumors based on supervised branches and uncertainty estimation. Sci Rep 15, 41517 (2025). https://doi.org/10.1038/s41598-025-25248-1
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-25248-1