Abstract
Ischemic watershed strokes (IWS), also known as border zone infarcts hold significant importance in neurology, however, are often overlooked type of Ischemic Stroke. They are difficult to detect due to their subtle appearance and small sizes. Current diagnostic methods are limited in their ability to effectively identify IWS which represents only 5% of all Ischemic stroke cases and that is why most existing approaches for stroke segmentation and classification focus on other mostly occurring stroke types. This leaves a gap in the effective management of the disease. Additionally, machine learning models often struggle to differentiate between small and large strokes, limiting their ability to generalize across different stroke sub-types. This study aims to introduce a new method for identifying, segmenting, and classifying IWS using a fusion of CNN and Transformer models. The proposed method is evaluated using DWI MRI images. Additionally, we evaluate the proposed method on another type of stroke, hemorrhage stroke, using an open source dataset named Physio Net. The proposed CT-Transfusion is a CNN-Transformer fusion model and is embedded with a Feature Fusion Module (FFM) to integrate local and global features for enhanced stroke classification. The FFM aligns the spatial orientation of the features and unifies them at the patch level and finally, a CNN decoder reconstructs the final image. Experimental results demonstrate impressive scores with 94.79% accuracy, 93.0% precision, 95.0% recall, and 94.0% F1-Score, for the detection and classification of small ischemic watershed lesions. It outperforms benchmark models, including Modified Mobile-net-UNet, Efficient-net-Unet, DeepLabV3Plus, ResNet50, and InceptionV3. The proposed model is also assessed on haemorrhage stroke dateset characterized with relatively large size lesions. The model achieved an accuracy of 99.7%, precision of 99.8%, recall of 99.8%, and F1-Score of 99.8%. These metrics show that our proposed model outperformed the recently published models. It is concluded that the proposed dual-stream approach enhances diagnostic accuracy without needing to combine different types of medical images, making it a practical solution for stroke assessment in clinical settings. The model has multiple potential benefits: it can improve early stroke detection, assist doctors in making decisions, and potentially be integrated into automated image analysis workflows in hospitals.
Similar content being viewed by others
Introduction
According to C. Tsao, et al., 2023, and Hou, Shuai, et al., 2021, stroke is the second most frequent cause of death globally, characterized by its devastating impact on brain health. During a stroke, approximately 32,000 brain cells are destroyed every second, resulting in a 20% mortality rate. On the other hand, survivors have to face significant disabilities in many cases due to underestimating the severity of this disease. Strokes are broadly divided into three key categories: Ischemic Stroke, Hemorrhagic stroke and Transient Ischemic Attack (TIA). Throughout the past three decades, the incidence of stroke has risen by 70%, highlighting a growing public health concern1. According to WHO TIAs accounts for 15% of all cerebrovascular occurrences globally2.
In the United States only, a stroke occurrence is recorded every 40 s, highlighting its alarming frequency. Strokes occur when blood vessels in the brain are either blocked or ruptured, leading to reduced oxygen supply and subsequent death of brain cells3.
A study focusing on Saudi health context, in the Kingdom of Saudi Arabia (KSA), the incidence rate of all strokes ranged between 175.8 and 196.2 per 100,000 people. The rate for intracerebral hemorrhage was found between 39.7 and 48.6, whereas the incidence rate for Ischemic Stroke was between 131.0 and 151.54. A 2023 study predicted a general upward trend about stroke incidence in Saudi Arabia over the past three decades5. Ischemic Stroke is the most frequently occurring type among all stroke types. contributing to approximately 89% of cases. On the other hand, hemorrhagic strokes have a higher incidence rate of 65%6.
According to7, a study focusing on Ischemic Stroke in Pakistan states that there is high incidence of stroke in Pakistan with estimates around 250 per 100,000 people annually. Current studies report that the stroke rate in certain communities is upto 4.8%, mainly affecting people under the age of 457.
Researchers have categorized Ischemic Stroke in various ways, depending on factors such as the location of the blockage, severity of the stroke, and time of onset. For instance, James Jose et al.8 classified Ischemic Stroke into ten distinct categories, one of which is watershed stroke, accounting for approximately 5% of all stroke cases. Watershed Stroke Infarct (WSI) occurs because of blocking and bursting of blood vessels between the brain territories9,10,11. An infarct is classified as an occurring in a watershed region when the boundary between two primary arterial territories divides the infarct into two distinct sections. This specific criterion for classification explains the characteristically small size of watershed infarcts8. Detecting these infarcts using deep learning models is a challenging task.
During a stroke, brain cells are deprived of oxygen and become vulnerable to infarction, resulting in Ischemic Stroke. The evolution of an ischemic infarct occurs in distinct stages of increasing severity, which are categorized accordingly as: (1) Hyper-acute, (2) Acute, (3) Sub-acute, and (4) Chronic. In certain cases, the condition may further progress to complications such as hemorrhagic transformation or watershed infarction. Early diagnosis of stroke is critical to minimize brain damage and determine the most appropriate treatment based on the stage of the disease. Given the severe consequences of stroke, significant research efforts have been dedicated to advancing early diagnosis and improving treatment strategies. The shortage of relevant human experts across healthcare institutions is a pressing issue that can negatively impact patient care and outcomes.The shape of the watershed stroke is patchy and band-like, making it difficult to detect for less experienced radiologists.
Advancements in medical imaging, computational power and, artificial intelligence (AI) has played a key role in improving both the accuracy and efficiency of medical image analysis. Various medical imaging modalities are the main tools for extracting key insights about diseases using non-surgical methods. In this context, Magnetic Resonance Imaging (MRI) and its variants are the most powerful and versatile tools. Watershed Infarcts (WSI) can be prominently detected on diffusion-weighted MRI (DW-MRI) during the acute stage. Several imaging modalities such as Computed Tomography (CT), perfusion MRI, diffusion MRI (dMRI), MR angiography (MRA), and MR spectroscopy (MRS) are also used for detection and classification of the disease12.
Computer vision is a rapidly growing domain within artificial intelligence domain, focused on extracting crucial information from images. In the past decade, neural-network-based approaches have consistently outperformed other machine learning methods. Among various neural network approaches, convolutional neural networks are found better for image processing and classification. Using mathematical filters to distinguish complex image patterns, CNN-based approaches produce feature maps that capture essential information for further analysis. Finally, fully connected layers are employed in the network to obtain classification probability. Despite their strengths in object classification, CNN models suffer from different issues during training phase. For example, gradient vanishing, difficulty in recognizing objects present in different poses, and a limited understanding of spatial relationships within images. These limitations can bound their ability to generalize effectively to unseen cases. One notable limitation is the use of constrained receptive field which restricts CNN capacity to learn long-range dependencies within an input image.
To overcome this, approaches like enhancing kernel size, global feature extraction, local and global pooling, and deploying deeper networks are used13. Larger kernel sizes in deep learning models increase the network’s representational capacity and allow for more fine-grained adjustments, improving flexibility. Additionally, substantial filters with increased pooling strides enhance variance and prevent overfitting, ensuring that characteristics are generalized on unseen datasets. Conventional neural networks have limitations in natively supporting variable-sized inputs and are less efficient when scaling up to handle high-quality images.
Unlike CNNs, transformers are more robust as they can mine long-term associations within sequences. They can efficiently handle spatial information through their encoding and embedding mechanisms, making them more suitable for tasks that require understanding complex relationships across large contexts. For medical images, transformers partition the 2D images into a sequence of smaller patches14. Embedding mechanisms are then used to interpret the relationships between these patches. Furthermore, self-attention applied to these embedded patches effectively captures spatial relationships, facilitating the identification of long-range dependencies and efficiently extracting meaningful information. Despite the advantages of transformers in intricate pattern recognition, the significant demand for a large volume of data for training poses a notable constraint. This issue is particularly challenging when dealing with medical datasets where overlapping and similar visual representation of different information are common. Medical images are rich in semantic and pathological details. However, when these images are affected by noise, distortion, or occlusion, the patterns and relationships within the data can become disrupted, making it difficult for a model to interpret and extract meaningful information accurately. This highlights the presence or absence of inductive bias with limited data leading to distinct challenges. These challenges can be addressed by combining convolutional and transformer models to enhance generalization with limited data.
The watershed region, positioned between the Middle Cerebral Artery (MCA), Posterior Cerebral Artery (PCA), and Anterior Cerebral Artery (ACA) presents challenges due to its limited sample size and the small size of strokes within it. MRI data consists of volumetric samples whereas 2D and 3D Convolution Neural Networks have limitations in capturing contextual information across consecutive slices of this volumetric data. Analyzing data accurately while considering contextual information presents its own set of challenges.
This study introduces the CT Transfusion Neural Network to improve generalization on the limited Watershed stroke dataset, where each lesion is small. We aim to develop a neural network capable of extracting low-level features and learning long-range dependencies within the feature set. Based on the analysis and experimental outcomes, it is evident that Transformers and CNNs complement each other in image analysis tasks. Following is the list of contributions of this work:
Development of custom dataset
We collected a proprietary dataset of Watershed stroke which is annotated by expert radiologists. The details are given in the relevant section.
Development of segmentation and classification
The proposed method focuses on enhancing feature diversity by employing a fusion model to evaluate both small and large lesions within and between arterial regions of the brain. The developed model is trained and evaluated on a custom Watershed stroke dataset, along with the PhysioNet ICH benchmark dataset. The structure of the paper is outlined as follows: Sect. 2 covers the literature review employing CNN and fusion of CNN with transformer for binary classification of different types of the stroke. Section 3 depicts our proposed approach of transformers and CNN for binary classification of Watershed Stroke. Section 4 demonstrates experimental setup, results obtained and their analysis. Finally, the paper concludes in Sect. 5 with a discussion of the study.
Literature review
This section discusses the most prominent methods applied for the automatic detection of Ischemic Stroke. The review focuses on segmentation and classification methods based on CNN, Transformers, and their hybrid approaches.
Convolutional neural network based approaches
Several studies have employed convolutional neural networks for automated detection, classification, and segmentation of different types of Ischemic Strokes. These models capture the spatial hierarchies of features from medical image data. Long et al.15 used fully connected networks (FCN) for semantic segmentation. Building upon the success of FCN, subsequent advancements led to the development of deeper and wider networks such as VGG1, Res Net16, Dense Net17, and Google Net4. Ronneberger et al.16 introduced the U-shaped network called U-Net, leveraging an encoder-decoder structure, which yielded remarkable results in biomedical image segmentation. Following this breakthrough, various versions of the U-Net architecture are introduced, each contributing to improved performance for the above-mentioned tasks8. Various researchers have utilized CNN-based models for the binary and Multi-class classification of various types of Ischemic Strokes18,19,20,21,22. Soltan et al.23 proposed a novel approach for acute stroke prediction, demonstrating a multiple parallel 2D U-Net architecture with a pixel-level classifier. The model operates in two stages: initially, it gathers stroke lesion texture using parallel U-Net architectures, followed by differentiation between normal and abnormal images using a pixel-level classifier. Astonishingly, employing logistic regression as the pixel-level classifier produces remarkable performance i.e. Dice Similarity Coefficient (DSC) of 71.3%, Recall of 73.6%, and Volumetric Similarity (VS) of 82.1%. A novel RRCNN model was introduced by Luu-Ngoc Do. et al.24 to classify low and high DWI-ASPECTS groups. Augmented DWI slices are used as input to the model with five convolutional blocks to extract feature maps, along with one recurrent block for capturing sequential features. It demonstrated outstanding performance, attaining an AUC of 0.94 and an F1-score of 0.88. The Classifier-Segmenter network (CSNet), introduced by25 is an architectural innovation combining U-Net and fractal Net components. Comprising two integral sections, the classifier network exploits series of convolutional blocks to capture diverse global and local features, comprising network depth for analyzing disease image slices. A SoftMax layer is appended to yield the final classification between lesion and non-lesion images. The segmenter network part employs an encoder-decoder framework, with encoder blocks comprising one or two convolution layers along with max-pooling and a 3D spatial dropout layer for spatial information extraction and learning regularization. The decoder block utilizes transpose convolution as its along with regular convolution operation. The network demonstrates significant performance with an 83% accuracy (dice coefficient), 79% precision, and 89% recall score. In contrast, Yi-Chia Wei et al.26 used a proprietary dataset of 261 MRIs and applied a two-stage model for lesion segmentation and binary classification.They employed a two-stage model for lesion segmentation and binary classification and introduced SGD-Net Plus, a technique incorporating brain atlas images. In another study, Lo, Hung et al.27 introduced a DCNN architecture for detecting acute Ischemic Stroke, combining AlexNet, Inception-v3, and ResNet-101 with a Softmax layer. This approach achieved an impressive 97.12% accuracy based on NCCT images from 96 patients, all taken within six hours of stroke onset.
In 2022, Khezrpour et al.28 proposed a network based on a U-Net encoder-decoder structure that incorporates blocks consisting of five parallel layers. This model includes a preprocessing stage using CLAHE, which significantly impacts the results. de Vries L et al.29 proposed a PerfU-Net model that generates perfusion maps from source data for infarct core segmentation. This model employs encoder and decoder architecture, enhanced with two attention modules: one for the channel dimension and one for the temporal dimension, aiming to improve classification and segmentation performance. A compact asymmetric U-Net architecture was introduced by Kumar et al.30for segmenting acute Ischemic Stroke using the ISLES 2018 dataset. The model consists of 11 convolutional layers and 84,217 trainable parameters, with various loss functions applied for class balancing, showing improved results in terms of DSC, precision, recall, and AVD.
Sinha et al.31 proposed EnigmaNet, a model designed to segment Ischemic Stroke lesions in FLAIR and DWI images. It utilizes a modified Weighted Focal-Tversky-Dice (wFTD) Loss function to enhance the detection and segmentation of Ischemic Stroke lesions. The architecture comprises Genesis-k blocks in both the encoder and decoder stages, along with dual-headed attention gates. The system achieved a Dice score of 0.8965, sensitivity of 0.8776, and specificity of 0.9866 for FLAIR images, and for DWI images, it resulted in a Dice score of 0.8423, sensitivity of 0.8452, and specificity of 0.9754.
Another model for lesion detection was proposed by Ghosal, P. et al.32, utilizing a dual-channel CNN encoder-decoder architecture. One channel utilizes residual connections to mitigate the vanishing gradient problem and transmit fine details, while the other uses residual and spatial attention to capture long-range dependencies. By combining these strategies, the model effectively captures both global and local information, addressing the challenge of detecting variable MS lesion features.
Transformer based approaches
Vision transformers have become a promising alternative to CNN-based approaches in medical imaging applications such as segmentation and classification. Muhammad Ayoub et al.33 introduced an end-to-end vision transformer architecture that leverages self-attention to capture long-term dependencies between 16 × 16 patches. The model refines images by embedding the patches via a transformer encoder, and processes three distinct inputs: a class token, sequences of patches, and bounding box coordinates. The inclusion of an RNN enhances temporal pattern recognition, thereby boosting the accuracy of classification. Wang et al.34 presented METrans, a network that employees a multi-encoder architecture for multiscale feature extraction with CBAM and a transformer. The network culminates in a decoder that performs upsampling to generate the final output image. In a separate study, Luo et al.35 proposed UCATR, a transformer-based network for the segmentation of acute Ischemic Stroke. By replacing the CNN encoder with an enhanced multiheaded cross-attention mechanism. UCATR filters out irrelevant information.UCATR, when evaluated on clinical data from 11 patients, achieved a Dice similarity coefficient of 73.58% for lesion segmentation, surpassing both U-Net and Attention U-Net.
In 2024, Lo et al.36 proposed a method for feature extraction using a Vision Transformer (ViT) applied to carotid color Doppler (CCD) images, which included 513 stroke and 458 normal images. The extracted features were utilized in an SVM classifier to predict Ischemic Stroke. The system achieved 89% accuracy, 94% sensitivity, 84% specificity, and an AUC of 0.95.
CNN-Transformer fusion
Currently, the researchers are focusing on using CNNs to extract local features and then employing transformers to capture long-range dependencies within slice sequences. In a research, Zelin Wu et al.37 proposed MLiRA-Net, a multiscale long-range interactive and regional attention network. The initial patch partition block leverages CNNs for local feature extraction, and the STR block is used for scale conversion, a distinctive strength of the model. The SiTR, or STR subsampling interactive transformer, captures channel-level features using a dimensional attention mechanism. Lastly, the FIP restores the original image resolution through a cascaded upsampler and merges the encoded features with the interpolated ones. This model achieves better performance than the TransUNet model38. Zhixiang Xu et al.39 introduced a model that extracts local features from CNN layers using channel attention and spatial attention modules to capture more precise local feature maps. The output is then fed to a transformer-based encoder. To address the loss of positional information, positional embeddings are applied. The transformer’s output is combined with local information, and a decoder is used to accurately segment the images. Their model achieved 58.66% of DC for AIS infarct segmentation, surpassing other contemporary models. Hulin Kuang et al.40 proposed a hybrid CNN model for segmenting acute Ischemic Stroke lesions. This model integrates parallel CNN and transformer encoder components, with an interaction block facilitating circular feature exchange between the CNN and transformer. This combination effectively merges the features extracted by both architectures. To focus on bilateral differences in the brain, a CNN decoder calculates these differences and produces the final output image, leading to precise segmentation. This model achieved a dice score of 61.63 ± 20.07 on the AISD dataset and a private dataset. The two-stage segmentation framework proposed by Tingting Li et al.41 SrSNet employs a dual-model approach with coarse and refined segmentation models, utilizing a Symmetrical Attention Block (SAB) to capture both local and global features. The model detects Ischemic Stroke lesions by analyzing differences between symmetric regions’ feature maps, leading to a recall score of 83.80% on the ISLES’22 data set. Kousar et al.19 emphasis on a critical need to detect a broader spectrum of stroke sub-classes simultaneously, recommending the use of both publicly and privately available datasets to improve cerebral stroke detection accuracy. It was concluded that most of the prior research has focused on ischemic and hemorrhagic strokes, with limited attention given to the different sub-types of Ischemic Stroke. Our study is among initial efforts to detect lesions both between (Watershed a type of Ischemic Stroke) and outside the brain’s arteries.
Research gaps
In the literature review, various segmentation and classification models for Ischemic Stroke and its sub-types are extensively discussed(Table 1).However, there is a need for a more detailed examination to uncover the specific factors that make one model more effective than another in terms of accuracy, robustness, and scalability.(Meng F., 2022) et al.42 proposed a model for Hemorrhage detection with robust generalization, achieving an impressive predictive accuracy of 96.51%. This success is attributed to the use of a large dataset, though it comes at a significant computational cost. Our research aims to develop an enhanced feature learning approach to improve model accuracy with this Large dataset. Xiyue Wang et al.43, Soltan et al.23, de Vries L et al.29, and Hulin Kuang et al.40 have proposed models for acute Ischemic Stroke segmentation, but these models have predictive accuracies lower than 85%. These studies highlight that the limitations due to the use of small datasets.research. Through a critical review as listed in Table 1, it becomes clear that most previous research has concentrated on ischemic and hemorrhagic strokes, with limited attention to the various subtypes of Ischemic Stroke. Our study is one of the initial efforts to detect lesions both within (Watershed, a subtype) and outside the brain’s arteries.One reason for focusing on Watershed stroke is that it involves smaller strokes, typically diagnosed by experienced radiologists. An automated method would assist radiologists in diagnosing Ischemic Strokes more accurately. This study focuses on both watershed and hemorrhagic strokes, utilizing the same architecture. Although hemorrhagic strokes involve bleeding and watershed strokes result from ischemic events, both types exhibit common features, such as abnormal tissue density, brain region engagement, and disrupted blood flow. The proposed model that can focus on both detailed (micro) and larger (macro) image patterns enables it to detect these shared features across stroke types. With its ability to capture both local and global features, the architecture can distinguish and effectively analyze both hemorrhagic and watershed strokes within a single pipeline.
Methodology
Overview of the proposed model
Our proposed model, CT Transfusion, adopts a fusion strategy by integrating Convolutional Neural Networks (CNNs) to capture local attributes and applies the transformers to extract global features from volumetric inputs. This approach enables to extract stroke specific features by tokenizing patch extraction effectively. In CT Transfusion, first a CNN-based encoder is designed, drawing inspiration from13,38, to extract contextual feature maps of size 16 × 16 × 1 from image slices of 256 × 256 × 1. Next, a transformer generates tokens of equivalent dimensions. Patch embedding enables efficient processing, manages receptive fields and spatial hierarchies, and incorporates positional encoding to improve interoperability. Additionally, our model integrates a transformer-based encoder inspired by the Vision Transformer (ViT) and TransUNet approaches14,38. This encoder captures long-range dependencies and manages images of varying dimensions or resolutions. It combines local and global features using a Multilayer Perceptron (MLP), allowing the model to learn complex patterns through self-attention and MLP layers. Overall, the fusion of convolutional neural network, transformer encoder with an MLP enhances our model’s ability to interpret intricate medical images by efficiently capturing both localized global details.
The proposed approach effectively addresses over fitting on limited data by utilizing multiple strategies. Focused learning is initially applied to the disease area, ensuring the model prioritizes the relevant regions. Augmentation is performed specifically on the ischemic areas to diversify the data. Higher weights are assigned to the ischemic lesions while the rest of the image is given less importance, guiding the model’s attention toward critical areas. The use of CNNs and transformers, each featuring built-in regularization mechanisms like CNN pooling layers and the transformer’s attention mechanism, enhances the model’s robustness. Reducing the transformer encoder’s layers minimizes model complexity and further lowers the risk of overfitting, making the method ideal for limited data.
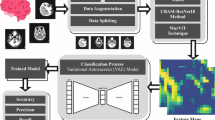
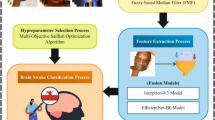
The proposed method provides multiple benefits like better image details extraction, focused analysis on specific parts and more efficient processing. 2) Combining local and global features allows us to look at both deep and coarse details simultaneously. This approach helps in finding micro changes with focus on important areas, understanding tissue details, reducing the impact of image noise, and observing large patterns and relationships, ensuring scalability and interoperability, making it suitable for various medical image analysis tasks. Figure 1 and Algorithm 1 show working of the proposed method.
The overall framework of the proposed approach.
The key features of this study are outlined below:
-
1.
This paper introduces a dual-path CNN architecture that combines local feature extraction with a global context, enhancing stroke detection, especially in challenging regions like the brain’s peripheral or border zones. The model’s ability to combine detailed local features with broader contextual information enables it to distinguish between small strokes and other potential artefacts better, minimizing false positives which makes it more suitable for accurate and robust diagnosis of small stroke detection.
-
2.
The model was assessed on a diverse dataset consisting of both small and large stroke cases. Compared to earlier studies, the model achieved higher accuracy with less false positive detection for large strokes. Although the model performed well with small strokes, its ability to detect large strokes with higher precision highlights its effectiveness in addressing more noticeable cases.
CT transfusion local and global features extractions algorithm.
Dataset description & preprocessing
The dataset, used in this research, was originally developed by Cetinoglu, Koska, Uluc, et al. (2021)18. The image format of this dataset is DICOM that is converted to NII (Neuroimaging Informatics) format to ensure consistent spatial orientation, enhancing its usability for automated analysis. Patient age and gender were recorded. The MRI scans were performed with a 1.5 Tesla MRI machine (MAGNETOM, Siemens Healthcare, Erlangen, Germany), and two distinct DWI protocols were employed.
The IWS datasets in this study comprises 365 disease-focused Trace DWI slices, and 365 Normal slices from 150 patients collected between January 2017 and April 2022. The images are divided into normal and watershed stroke classes.Two different DWI Image acquisition protocols were utilized, and Table 2 presents the technical details of each protocol, along with the distribution of the acquisition protocols for the DWIs included in the study.
For assessment the performance of proposed approach in terms of its effectiveness on standard Dataset of intracranial hemorrhage (ICH comprising 381 slice images from the positive hemorrhage stroke class and 683 slice images from the normal class). Each slice has a thickness of 5 mm. with spatial resolution of 512 × 51244. Each image is provided with relevant ground truth which provides manual segmentation of disease area.
Several methods for data augmentation and pre-processing are applied. This helped in obtaining better results and dealing with class-imbalance problem. During the pre-processing phase..Ischemic watershed Dataset is initially in.nrrd format and converted into.nii file for the purposes of Aligning the slices that ensures anatomical consistency, reduces artifacts. After cleaning the data, the resulting slices are stored separately as images, which is essential for the model’s ability to process each slice independently. These slices are stored as individual images to create a diverse training dataset, with binary classes representing stroke and normal brain tissue. Further, pre-processing, steps such as normalization, resizing are applied Normalizing pixel intensity values (such as scaling them to a fixed range like [0, 1]) ensures that features are on a similar scale, resulting in more stable gradient updates during backpropagation. This leads to smoother training and better generalization across different imaging types, ultimately improving segmentation accuracy. Meanwhile, resizing the images to the same size (256 × 256 × 1 pixels), fixed resolution while preserving critical anatomical features reduces memory and computational demands, balancing spatial resolution with processing efficiency to handle dataset more effectively. Data augmentation were implemented to address the limited amount of data and improve model performance. Data augmentation through image translation was also used, which involved shifting the images along the X and Y axes or one of them.Moreover, To address the class imbalance issue, equal number of disease and normal slices are selected as an input batch and fed to the designed algorithm. The visualization & details are provided in figure 2 (Algorithm 1).Following figure 2 shows the examples from resulting image-set.
Visualization of small watershed lesion through DWI modalities of custom dataset.
Network architecture
Local feature extraction in CT transfusion: CNN encoder
The architecture of the CTtransfusion model focuses on the classification of watershed infarcts. Overall CNN Encoder is represented in Fig. 3. The process begins by feeding 256 × 256 × 1 input image slices through a convolutional block, which extracts local features and aids in learning complex patterns. Each convolutional block employs batch normalization and ReLU activation to stabilize the learning process and enhance feature extraction. Max pooling layers are employed to decrease the spatial dimensions of the feature maps. Subsequently, bilinear interpolation transforms the feature maps into specific patches of size 16× 16 × 1, as depicted in Eq. 1. Here \(\:w\:\)and \(\:b\) represent the weights and biases. Bilinear interpolation (BI) is used for intensity-based image resizing to generate local feature set patches.
Global feature extraction in CT transfusion: transformer encoder
Extracted feature vectors are input for a transformer encoder, inspired by the Vision Transformer (ViT) model. The input images are divided into patches or tokens, which then undergo a sequential transformation. This process results in feature vectors of size (n + 1, d) as shown in Eq. (2) and figure 3.
Convolutional neural network encoder.
The input image slice is split into n patches of dimensions (p, p,c). Each patch is flattened into a vector of shape (1, p2c). These vectors are then projected into d dimensions using a trainable dense layer, creating n embedded patches of shape (1, d). Positional embeddings are created and added to these embedded patches to incorporate spatial information, serving as a simple alternative to complex sinusoidal encoding as shown in Eq. 2. The entire patch embedding process is illustrated in Fig. 4. This approach offers a balanced computational load, captures both local details and global context, improves generalization and robustness, aids in understanding object layouts, and accelerates the training process. Embedded patches are passed to the Sequential Transformer Encoder, which calculates long-range dependencies between patches. The purpose of transformer encoders is to use self-attention mechanisms to capture relationships between all patches simultaneously. This allows the model to understand intricate interactions and dependencies between different image parts beyond the local convolutional operations. Multiheaded attention mechanism uses number of attention heads to extract features. Each head processes the input differently, and then the results are concatenated. More heads allow for capturing more aspects of the data. Table 3 lists the parameters corresponding to various aspects of the model architecture and training settings. These parameters collectively define the structure and behavior of the model, influencing its capacity to learn, process input data, and generate accurate predictions. Compared to ViT, using fewer layers in the transformer encoder minimizes the risk of overfitting, reduces model complexity, and is advantageous for limited data. Each layer consists of a multi-head self-attention mechanism and a feed-forward neural network. To minimize the effect of vanishing gradients and small batch sizes, normalization techniques are incorporated into the transformer encoder. A multilayer perceptron enables the model to learn complex patterns. The Multilayer Perceptron (MLP) consists of two layers. The first dense layer, with a weight matrix ℎ of shape (d,) transforms the input y from dimension d to which effectively transforms the input into a higher (or lower) dimensional space depending on the value of. Dropout is then applied for regularization. Increasing the dropout rate from 0.1 to 0.2 improves the model’s ability to generalize and reduces its dependence on specific neurons, which is particularly helpful when training data is limited in size. Incorporating convolutional layers within a transformer captures local patterns, and maintains spatial consistency, thereby improving performance through enhanced local and global context learning33,41. The second dense layer, with a weight matrix 0 of shape (,d) transforms back to the output dimension d without an activation function. Equation (3) represents the multi-headed self attention -mechanism (MHSA), where layer normalization is applied to the output of the previous layer (LN(h−1)). Equation (4) describes a residual connection within a layer of a transformer encoder using a multilayer perceptron (MLP). The MLP is a feed-forward network in which input is passed through the network layer by layer, to reduce the overfitting issue on limited data. In our model we use small number of layers which also leads to minimized computational load. In MLP, the Gaussian Error Linear Unit (GELU) activation function helps minimize the vanishing gradient problem by allowing a smoother flow of information unlike other functions that kill neurons by adjusting all negative values to zero, GELU allows specific negative values to pass through. Hence, all this process supports for smooth gradient flow as mentioned in Eq. (5). In Eq. (5), z derives a feature from the input and 0.5. represents the amount of attention given to each pixel. The cubic term makes the pixel value constant that are close to zero. Let \(X \in \mathbb{R}^{{B \times H \times W \times C}}\) be the 4-D output tensor from the transformer encoder, where B represents the batch size, H and W denotes the height and width of attribute maps, and C denotes the number of channels. After applying Global Average Pooling (GAP), the resultant global feature vector is \(G \in \mathbb{R}^{{B \times C}}\) for each channel. This vector is then reshaped and tiled to match the spatial dimensions of the original local features. Let \(G_{{tiled}} \in \mathbb{R}^{{B \times H \times W \times C}}\) be the tiled global features, and \(X \in \mathbb{R}^{{B \times H \times W \times C}}\) denotes the local attributes from CNN encoder. Concatenate the local and global attributes along the channel dimension to obtain the resultant hybrid embedding \(H_{{hybrid}} \in \mathbb{R}^{{B \times H \times W \times 2C}}\). The schematic diagram of transformer encoder is depicted in Fig. 5. This approach ensures that the model captures both the global context and the local details, enhancing its robustness and effectiveness, particularly in classification tasks. Equation (6) (7), (8) and (9) describe the tiling and concatenation process. After obtaining the combined refined image through the transformer encoder and CNN, we further processed it using the CNN decoder. This decoder up-samples the feature maps from 16 × 16 to 256 × 256, aiding in reconstructing high resolution output while preserving spatial details.
Patch embedding.
Transformer encoder block.
CNN decoder
The preprocessed images are arranged as sequential patches and fed to the CNN decoder as input, producing high-resolution output without affecting spatial information. At the decoder level, employing UpSampling2D and Conv2DTranspose layers can generate high-resolution images from low-resolution feature maps. Using convolutional layers with Conv2DTranspose provides balanced control over feature extraction and spatial resolution adjustment. Intermediate convolutions with 32, 64, and 128 filters enhance feature learning before upsampling, which can lead to better predictions. To initialize the network weights, the random initializer is used, leading to fast convergence. To maintain consistency in feature map size, padding=’same’ is used. ReLU activation function is used in the intermediate layers, while sigmoid activation is used for the binary classification of Watershed Stroke.
In Eq. (10), \(\:F\) defines each transformation (convolution, transposed convolution, upsampling, or activation function), the symbol (\(^\circ\)) denotes function composition, and N represents the number of layers. The Model is trained by using weighted binary cross-entropy as a loss function, which is defined by Eq. (11) where \(\:x\:\:\)represents the actual label (0 or 1) as the abnormal class’s predicted probability.w1 and w0 represents higher weights to the focused Watershed regions (class 1) and lower weights to the healthy regions (class 0).
Experiments and results
All experiments were conducted on a Microsoft Windows system 11 pro powered by an 11th-generation Intel® Core™ i7-11700KF CPU running at 3.60 GHz, featuring 8 cores and 16 logical processors, 32 GB of RAM, and a 2.78 TB HDD. Graphical processing was handled by a CUDA-enabled NVIDIA GeForce RTX 3080 GPU with 32 GB of memory. The code was implemented using Keras with a TensorFlow backend.
For model evaluation, the validation split technique is employed, reserving 20% of the images from the combined dataset for validation. The loss function was optimized using the Adam optimizer with a low learning rate of 0.0007. Training, using the standard dataset of ICH, was performed over 140 epochs, using a mini-batch size of 4 and a dropout rate of 0.2.
An empirical approach was used to select the optimal learning rate for the model. Various learning rates, including 0.01, 0.0001, 0.001, 0.0005, and 0.0007, were tested and evaluated. SLR (adaptive learning rate methods) method is also used, The transformer model performed most effectively with a learning rate of 0.0007. A small batch size of 4 was used because it allows the model to generate patches during processing, improving space complexity. While multiple batch sizes were tried, the smaller batch size produced superior results. The model was trained for 50 epochs, and as the epochs increased, the loss stabilized, leading to the model halting once it stopped improving.
Comparison with state-of-the-art methods
Our proposed model demonstrated outstanding performance in detecting small-size watershed strokes, achieving an accuracy of 94.79%, a precision of 93%, a recall of 95%, an F1-score of 94%, and a Dice Similarity Coefficient (DSC) of 94%. In this study, we compare the performance of our proposed CT-transfusion model against different widely adopted architectures, including Modified Mobile-net-UNet45, Efficient Net-Unet46, DeepLabV3Plus47, ResNet5048, and InceptionV349—models commonly used for stroke prediction tasks.
All models chosen for comparison are evaluated on the same dateset using a consistent pre-processing pipeline, which involves feature scaling and image enhancement. To address class imbalance, an equal number of pathological and normal slices are selected and input into each network. The models were assessed using key performance metrics: accuracy, precision, recall, and F1-score. Detailed results for all models are presented in Table 3.
Table 3 demonstrates the CTtransfusion model’s performance, emphasizing its effective capture of sequential patterns for enhanced prediction accuracy. While ResNet50 and InceptionV3 maintain strong precision, indicating reliable positive instance identification, they exhibit limitations in capturing all true positives.
To evaluate the proposed method’s performance, we conducted experiments on multiple datasets, including the benchmark ICH hemorrhage dataset, which presents larger lesions compared to watershed strokes. Our model achieved accuracy of 99.7%, outperforming existing methods. A comparative analysis was performed against VGG-1942,50, CNN&RNN43, DenseNet-20151, and Ensemble 2D CNN52. Table 5 presents the detailed performance metrics, including accuracy, precision, recall, and F1-score, DSC and jaccard Index for these models. Our model’s performance, characterized by a dice Similarity coefficient(DSC), recall, precision, accuracy and Jaccard Index demonstrates its effectiveness in infarct segmentation and classification, surpassing the comparative methods. Figures 7 and 8 illustrate the training and validation accuracy graphs for the ICH hemorrhage and watershed datasets, respectively. Figures 6 and 9 present the training and validation loss during the training of the ICH hemorrhage dataset and watershed stroke dataset.
The selected research, for performance comparison, have used deep-learning models for haemorrhage stroke detection using physionet dataset. Fanhua Meng et al.42 proposed the AIMA-ICHDC model based on VGG-19, achieving an overall accuracy of 96.51% on the ICH dataset. Hai Ye et al.43 proposed a CNN-RNN-based model trained on 2836 ICH images, achieving an accuracy of over 80% for all stroke types. S. Santhoshkumar et al.51 introduced the DL-ELM model with Tsallis entropy and the Grasshopper Optimization Algorithm for segmentation and DenseNet201 for feature extraction, achieving 96.34% accuracy on the ICH dataset. Additionally, Xiyue Wang et al.52 presented a deep learning model combining 2D CNN with sequence models for ICH detection, achieving 94.3% accuracy.
Our fusion model achieved an accuracy of 94.79% in detecting watershed stroke using dataset developed by Cetinoglu, Koska, Uluc, et al. (2021)53 and 99.7% in detecting hemorrhage using physioNet data-set of ICH. The proposed model outperforms the selected recent research, with an accuracy of 99.7%, a precision of 99.8%, a recall of 99.8%, and an F1-score of 99.8%. Figure 10 demonstrates the proposed CTTransfusion model’s predictions on four watershed stroke testing images. Working with large datasets is crucial for improved image recognition and enhanced CNN performance. Additionally, extracting features that are more precise and capturing long-range dependencies between them can further improve model performance.
Training and validation loss of proposed model on ICH dataset.
Training and validation accuracy of proposed model on benchmark dataset (ICH).
Training and validation accuracy of proposed model on private Ischemic water-shed stroke dateset.
Training and validation loss of proposed model on prosperity watershed dataset.
To address class imbalance in the dataset, we focused on instances of interest (e.g., slices containing hemorrhage or watershed) while excluding irrelevant or non-informative cases (e.g., slices without hemorrhage or watershed). To further address this issue, we applied cost-sensitive learning to prioritize minority classes. This approach reduced computational load, enhanced accuracy, and improved the model’s generalization performance.
The time complexity of proposed model for one epoch is given below in Eq. (12)54,55,56,57,58,59,60:
And Time Complexity for all training epochs is:
Where:
-
\(\:E\)= number of epochs (50).
-
M = dataset size (number of training samples) (36,600).
-
B = batch size (4).
-
H, W = image size (256 × 256).
-
N=(H/P) (W/P) = number of patches (256).
-
d = hidden dim (768).
-
mlp_dim = 3072.
-
\(\:C\_in,\:C\_out\): Number of input and output channels.
-
\(\:k\): Kernel size.
Space complexity in deep learning models is mainly influenced by three key factors. Thee include model parameters, which include the weights and biases across all layers, such as convolutional, dense, and attention layers, which need memory to store the model’s learned parameters. Next, intermediate activations, generated during the forward pass, are stored for backpropagation and can occupy significant memory, particularly in large models with complex inputs. Finally, temporary memory is needed for operations such as reshaping tensors, matrix multiplications, and patch extraction, which require buffers that, while short-lived, still contribute to the model’s overall memory consumption54,55,56,57,58,59,60. The computed space complexity is given below:
Where
B = Batch size
P = Number of pateches
H = Hidden dimensions
Heads = Number of attention heads
M = mlp_dim
C = \(num_{{channels}}\)
I = image_size
F = Number of filters
The detailed computation of time and space complexity is given in appendix-A54,55,56,57,58,59,60.
Our proposed model handles the local and global features separately and then fusing them after independent processing, This model can achieve a more precise integration of the two types of information. This method allows the global features to be processed in a way that emphasizes their broader context, which is particularly important for tasks involving large-scale image understanding, such as medical imaging. In contrast, TransUNet uses a Transformer to capture global context from tokenized patches of CNN feature maps, and then the decoder upsamples and merges these encoded features with high-resolution CNN maps for accurate localization. In a benchmark dataset comparison, CT Transfusion achieves an impressive DSC of 99.9 for hemorrhage detection, while TransUNet has a DSC of 84.36. In Architectural design of UNETR, a 3D input volume (e.g., 4 channels for MRI images) is split into non-overlapping patches. These patches are then projected into an embedding space using a linear layer.The patch embedding, combined with position embedding, are processed by a Transformer encoder.The encoded representations from multiple layers of the Transformer are extracted and fused with the decoder via skip connections to generate the final segmentation output. Table 6. represents Comparison of number of parameters, Flops and averaged inference time for various models to proposed model.
Predictions generated through proposed CTtransfusion model on watershed stroke dataset.
This approach captures fine-grained details and reduces over-fitting issues, as smaller patches decrease the complexity of the data fed into the model at once. Additionally, this method improves the model’s flexibility by enabling multi-scale analysis.
The comprehensive analysis of our model’s performance on the standard dataset of ICH physio-net reveals its high precision (99.3%), Dice Similarity Coefficient (DSC) of 0.9987%, and a relatively low false negative rate (0.06%). These results demonstrate the model’s effectiveness in accurately detecting haemorrhage cases while minimizing erroneous predictions. Such performance is essential in clinical settings, where accurate diagnostics can significantly influence patient outcomes and the allocation of healthcare resources.
The performance of the hemorrhage segmentation model was assessed by clinical experts using a 5-point scale.The results were overwhelmingly positive, with 85% of images rated as clinically acceptable (score 5), demonstrating the model’s potential for high-quality segmentation.However, 15% of images required minimal edits (score 2) due to anatomical misidentifications, which could be corrected with further fine-tuning and enhanced data with some data augmentation approaches.To quantify the agreement between the expert ratings and the model’s predictions, calculated the Cohen’s Kappa score. The Cohen’s Kappa score, which measures inter-rater reliability, was found to be 0.75, indicating substantial agreement between the expert radiologists and the model.Despite these minor issues, the model shows great potential for broader clinical use after further refinement.The model identifies key features such as hyperdense regions in the brain indicating bleeding, the size and location of the hemorrhage, and critical associated features like brain compression, abnormal tissue changes, and signs of increased pressure or fluid buildup, all of which assist in clinical decision-making.The inter-rater agreement between Observer 1 (Expert Radiologist) and Observer 2(Pgr4) (Model) was evaluated As shown in Table 7.Although there was one instance of disagreement, the model’s overall agreement rate of 91.67% demonstrates its strong clinical reliability.This suggests that the model is suitable for use in routine hemorrhage detection, with only slight adjustments needed for more complicated cases.
The watershed stroke assessment was reviewed by two independent observers using a 5-point Likert scale. Agreement was observed in 80% of cases, predominantly rated as clinically acceptable (score 4) or unusable (score 1) as shown in Table 8. A single case revealed a major discrepancy, highlighting the role of subjective judgment. The Cohen’s Kappa score of 0.55 indicates moderate agreement, reflecting that while the approach shows promise, variability between observers remains. Importantly the reason of limited performance is, watershed stroke differs from focal infarcts, consisting instead of multiple patchy lesions that may shift positionally within the brain. This anatomical complexity likely contributes to the observed disagreement and model performance challenges. Overall, these results demonstrate the potential of the method in clinical application, though further refinement is necessary to ensure consistent interpretation.
Despite its good performance on limited datasets, the model has the following limitations:
-
1.
Fine-tuning hyper parameters for both the CNN, such as layer complexity and learning rate, and the Transformer encoder, including the number of heads, depth of attention layers, and dropout, are complex tasks because tuning one component affects the performance of the other. This process is also time intensive. Additionally, the model’s performance may be sub-optimal when applied to different disease types or varying imaging modalities. Balancing the features learned by CNN and the Transformer encoder can be challenging, as misalignment or redundancy in features might reduce the overall effectiveness of the hybrid model.
-
2.
Combined architecture is harder to interpret compared to using a single model type, complicating the understanding of how each component influences the final outcome. This may impact the model’s accuracy when applied to different brain illnesses.
-
3.
The developed method has only been evaluated on binary classification tasks, and further evaluation on multi-class detection tasks is required to ensure its resilience.
-
4.
Current method uses a fixed image size due to the hyper parameters of the transformer component. This limitation could be overcome by developing a hybrid approach that supports variable image sizes and multi-scale inputs.
-
5.
The model has been trained on a relatively small dateset, which may limit its generalization ability and affect its overall performance. To mitigate these limitations, future research should explore using larger and more diverse datasets, evaluate the model across different sorts of imaging modalities and strokes, and introduce methods to treat image sizes and resolutions that vary in nature.
-
6.
unavailability of standard Dataset of watershed stroke.The lack of a standardized Dataset for watershed strokes creates a limitation in evaluating the model’s ability to generalize, as it prevents evaluating it on a new, unseen Dataset outside of the training and validation data. Due to the absence of a widely recognized Dataset for this stroke type, assessing the model’s performance on external data becomes challenging, thus restricting the demonstration of its generalization capabilities in real-world applications.
Conclusion and future work
Artificial intelligence is pivotal in healthcare, driving significant advancements in research. Accurate stroke detection remains challenging due to the need for expert-level skills and overlapping features in manual interpretation. Automatic extraction of relevant features is crucial for improving detection performance and accelerating recovery. Research on stroke detection in various brain regions is ongoing, with different studies employing standard datasets and neural networks for automatic detection.
Our research focuses on detecting a specific type of Ischemic Stroke, known as watershed stroke, which affects the broader zone areas of the brain. The proposed method, CTtransfusion, combines local and global feature extraction techniques, achieving 94.79% accuracy in watershed stroke detection. We tested our method on the ICH Benchmark Dateset and attained a 99.7% accuracy in haemorrhage detection. While our results are promising with the limited dataset, challenges remain. The complexity of integrating feature architecture exceeds that of standalone models, leading to more intricate hyperparameter tuning and increased training time. This architecture also requires additional finetuning when applied to other disease detections. Our model performs well on balanced class datasets. In future work, to address the class imbalance issue at the architectural level, this work can be extended by designing Hybrid Models with imbalance-specific heads. This approach will help the model perform better under imbalanced conditions by focusing on underrepresented classes during feature extraction. To achieve more generalized results, larger balanced datasets will be used. While the current model is designed for binary classification, future work will explore a multi-scale fusion model, which could improve performance for both binary and multi-class classification tasks.
Data availability
The datasets used or analyzed during the current study are available from the corresponding author upon reasonable request.
Change history
10 February 2026
The original online version of this Article was revised: The original version of this Article contained an error in the Funding section. Thecorrect Funding section now reads: “This work was supported by the Deanship of Scientific Research, Vice Presidency for GraduateStudies and Scientific Research, King Faisal University, Saudi Arabia Grant No. KFU254278.”
References
Tsao, C. et al. Heart disease and stroke statistics—2023 update: a report from the American heart association. Circulation 147 (8), e93–e621 (2023).
Feigin, V. L. et al. Global, regional, and National burden of stroke and its risk factors, 1990–2019: a systematic analysis for the global burden of disease study 2019. Lancet Neurol. 20 (10), 795–820 (2021).
Moawad, H. B.J.M., Everything you need to know about stroke. (2023).
Algahtani, H. & Shirah, B. Primordial and primary prevention of ischemic stroke in Saudi arabia: A combination approach and evolving concepts. Saudi J. Med. Med. Sci. 12 (1), 1–9 (2024).
Çınar, N., Kaya, B. & Kaya, M. Brain stroke detection from CT images using transfer learning method. In: 2023 13th International Conference on Advanced Computer Information Technologies (ACIT). (IEEE, 2023).
Krishnamurthi, R., Ikeda, T. & Feigin, V. Global, regional and countryspecific burden of ischaemic stroke, intracerebral haemorrhage and subarachnoid haemorrhage: a systematic analysis of the global burden of disease study 2017. Neuroepidemiology 54 (2), 171–179 (2020).
Farooq, A., Venketasubramanian, N. & Mohammad Wasay., and Stroke Care Pakistan Cerebrovasc. Dis. Extra 11.3 : 118–121. (2021).
Jose, J. & James, J. An mri based ischemic stroke classification–a mechanism oriented approach. Ann. Indian Acad. Neurol. 25 (6), 1019–1028 (2022).
Liu, L. et al. Deep convolutional neural network for automatically segmenting acute ischemic stroke lesion in multi-modality mri. Neural Comput. Appl. https://doi.org/10.1007/s00521-019-04096-x (2020).
Helwan, A. et al. Deep networks in identifying Ct brain hemorrhage. J. Intell. Fuzzy Syst. 35 (2), 2215–2228 (2018).
Torvik, A. The pathogenesis of watershed infarcts in the brain. Stroke 2, 221–223 (1984).
Bernal, J. et al. Deep convolutional neural networks for brain image analysis on magnetic resonance imaging: a review. Artif. Intell. Med. 95, 64–81 (2019).
X.Wang, R., Girshick, A. & Gupta, K. He,Non-local neural networks. In: Proceedings of the IEEE conference on computer vision and pattern recognition. 7794–7803 (2018).
Dosovitskiy, A. et al. An image is worth 16x16 words: Transformers for image recognition at scale. preprint at https://arXiv.org/abs/2010.11929 (2020).
Long, J., Shelhamer, E. & Darrell, T. Fully convolutional networks for semantic segmentation. In: Proceedings of the IEEE conference on computer vision and pattern recognition. (2015).
Ronneberger, O., Fischer, P. & Brox, T. U-net: Convolutional networks for biomedical image segmentation. In: Medical Image Computing and Computer-Assisted Intervention–MICCAI 2015: 18th International Conference Proceedings. Part III (Springer, 2015).
Simonyan, K. & Zisserman, A. Very deep convolutional networks for large-scale image recognition. Preprint at https://arXiv.org/abs/1409.1556 (2014).
Cetinoglu, Y. K. et al. Detection and vascular territorial classification of stroke on diffusion-weighted MRI by deep learning. Eur. J. Radiol. 145, 110050 (2021).
Kousar, T. et al. Applications of deep learning algorithms in ischemic stroke detection, segmentation, and classification. Artif. Intell. Rev. 58 (5), 1–48 (2025).
Suberi, A. M. et al. Deep transfer learning application for automated ischemic classification in posterior fossa CT images. Int. J. Adv. Comput. Sci. Appl. 10 (8), 459–465 (2019).
Kuang, H. et al. Hybrid CNN-Transformer Network with Circular Feature Interaction for Acute Ischemic Stroke Lesion Segmentation on Non-contrast CT Scans. IEEE Trans. Med. Imaging https://doi.org/10.1109/TMI.2024.3362879 (2024).
Ostmeier, S. et al. Random Expert Sampling for Deep Learning Segmentation of Acute Ischemic Stroke on non-contrast CT. J. NeuroIntervent. Surg. (2024).
Soltanpour, M. et al. Ischemic Stroke lesion prediction in ct perfusion scans using multiple parallel u-nets followed by a pixel-level classifier. In: 2019 IEEE 19th International Conference on Bioinformatics and Bioengineering (BIBE). (IEEE, 2019).
Do, L. et al. Automatic assessment of aspects using diffusion-weighted imaging in acute ischemic stroke using recurrent residual convolutional neural network. Diagnostics 10 (10), 803 (2020).
Kumar, A. et al. CSNet: A new deepnet framework for ischemic stroke lesion segmentation. Comput. Methods Programs Biomed. 193, 105524 (2020).
Y.-C.Wei, et al., Semantic segmentation guided detector for segmentation, classification, and lesion mapping of acute Ischemic Stroke in mri images, NeuroImage: Clinical 103044. (2022).
Lo, C., Hung, P. & Lin, D. Rapid assessment of acute ischemic stroke by computed tomography using deep convolutional neural networks. J. Digit. Imaging. 34 (3), 637–646 (2021).
Khezrpour, S. et al. Automatic segmentation of the brain stroke lesions from Mr flair scans using improved u-net framework. Biomed. Signal Process. Control. 78, 103978 (2022).
de Vries, L. et al. Perfu-net: baseline infarct Estimation from Ct perfusion source data for acute ischemic stroke. Med. Image. Anal. 85, 102749 (2023).
Kumar, A. A lightweight asymmetric U-Net framework for acute ischemic stroke lesion segmentation in CT and CTP images. Comput. Methods Programs Biomed. 226, 107157 (2022).
Sinha, S. et al. EnigmaNet: A novel Attention-Enhanced segmentation framework for ischemic stroke lesion detection in brain MRI. IEEE Access. 12, 91480–91498 (2024).
Ghosal, P. et al. Compound attention embedded dual channel encoder-decoder for ms lesion segmentation from brain MRI. Multimedia Tools Appl. 84, 1–33 (2024).
Ayoub, M. et al. End to end vision transformer architecture for brain stroke assessment based on multi-slice classification and localization using computed tomography. Comput. Med. Imaging Graph. 109, 102294 (2023).
Wang, J., Wang, S. & Liang, W. Metrans: Multi-encoder transformer for ischemic stroke segmentation. Electron. Lett. 58 (9), 340–342 (2022).
J., Zhang, Z., Xue, Q., He, T., Luo, C., Zhuo, K., … Xu, F., The robust UCATR algorithm enhances the specificity and sensitivity to detect the infarct of acute ischaemic stroke within 6 hours of onset via non-contrast computed tomography images. BMC neurology 22(1) 291 (2022).
Lo, C. M. & Hung, P. H. Predictive stroke risk model with vision transformer-based doppler features. Med. Phys. 51 (1), 126–138 (2024).
Wu, Z. et al. Multi-scale long-range interactive and regional attention network for stroke lesion segmentation. Comput. Electr. Eng. 103, 108345 (2022).
Chen, J. et al. Transunet: Transformers make strong encoders for medical image segmentation. preprint at https://arXiv.org/abs/2102.04306 (2021).
Xu, Z. & Ding, C. Combining convolutional attention mechanism and residual deformable transformer for infarct segmentation from ct scans of acute Ischemic Stroke patients. Front. Neurol. 14, 1178637. https://doi.org/10.3389/fneur.2023.1178637 (2023).
H.Kuang, et al., Hybrid cnn-transformer network with circular feature interaction for acute ischemic stroke lesion segmentation on Noncontrast Ct scans. IEEE Trans. Med. Imaging (2024).
Li, T. et al. Srsnet: Accurate segmentation of stroke lesions by a twostage segmentation framework with asymmetry information. Expert Syst. Appl. https://doi.org/10.1016/j.eswa.2024.124329 (2024).
FMENG F., W. J., ZHANG H., AND LI W., Artificial Intelligence-Enabled Medical Analysis for Intracranial Cerebral Hemorrhage Detection and Classification. J. Healthcare Eng.. 13 2017223 https://doi.org/10.1155/2022/2017223 (2022).
Ye, H. et al. Precise diagnosis of intracranial hemorrhage and subtypes using a three-dimensional joint convolutional and recurrent neural network. Eur. Radiol. 29, 6191–6201 (2019).
HSSAYENI, M. Computed tomography images for intracranial hemorrhage detection and segmentation (version 1.3.1). PhysioNet https://doi.org/10.13026/4nae-zg36 (2020). RRID:SCR_007345.
Jing, J., Wang, Z., Rätsch, M. & Zhang, H. Mobile-Unet: an efficient convolutional neural network for fabric defect detection. Text. Res. J. 92 (1–2), 30–42 (2022).
Mathews, M. R., Anzar, S. M., Krishnan, R. K. & Panthakkan, A. EfficientNet for retinal blood vessel segmentation. In 2020 3rd International Conference on Signal Processing and Information Security (ICSPIS). 1–4 (IEEE, 2020).
Liu, Y. et al. Image semantic segmentation approach based on DeepLabV3 plus network with an attention mechanism. Eng. Appl. Artif. Intell. 127, 107260 (2024).
Thorpe, M. & van Gennip, Y. Deep limits of residual neural networks. preprint at https://arXiv.org/abs/1810.11741 (2018).
Szegedy, C. V.V.e.a., Rethinking the Inception Architecture for Computer Vision. (2015).
Ma, J., Li, F. & Wang, B. U-mamba: enhancing long-range dependency for biomedical image segmentation. ArXiv Preprint. 2401 (04722), 1–10 (2024).
Santhoshkumar, S. et al. Machine learning model for intracranial hemorrhage diagnosis and classification. Electronics 10 (21), 2574 (2021).
Wang, X. et al. A deep learning algorithm for automatic detection and classification of acute intracranial hemorrhages in head Ct scans. NeuroImage: Clin. 32, 102785 (2021).
Cetinoglu, Y. et al. Detection and vascular territorial classification of stroke on diffusion-weighted mri by deep learning. Eur. J. Radiol. 145, 110050 (2021).
Goodfellow, I. et al. Large Language Models Lecture Notes on Transformer C omplexity. Deep learning. 1 2 (MIT Press, 2016).
Ba, J., Lei, J. R. & Kiros and Geoffrey E. Hinton Layer. Normalization. preprintat https://arXiv.org/abs/160706450 (2016).
Ioffe, S. & Szegedy, C. Batch normalization: Accelerating deep network training by reducing internal covariate shift. International conference on machine learning. (pmlr, 2015).
Nielsen, M. A. Neural Networks and Deep Learning. 25 (Determination press, 2015).
Mikolov, T. et al. Distributed representations of words and phrases and their compositionality. Adv. Neural. Inf. Process. Syst. https://doi.org/10.1016/j.yjbinx.2019.100047 (2013).
Goldberg, Y. Neural network methods for natural Language processing. Comput. Linguistics. 44 (1), 194–195 (2018).
Dumoulin, V. & Visin, F. A guide to convolution arithmetic for deep learning. Preprint at https://arXiv.org/abs/1603.07285 (2016).
Acknowledgements
The Researchers would like to thank the Deanship of Graduate Studies and Scientific Research at Qassim University for financial support (QU-APC-2025).
Funding
This work was supported by the Deanship of Scientific Research, Vice Presidency for Graduate Studies and Scientific Research, King Faisal University, Saudi Arabia Grant No. KFU254278.
Author information
Authors and Affiliations
Contributions
Tanzeela kousar (T.K.) * Analysis, Design the Methodology, Write the ManuscriptMuhammad shafry Mohammad Rahim (M.S.M.R.) Supervise and review the WorkMuhammad Alsuhaibani (M.A.)* Funding and reviewing the manuscript Sajid Iqbal (S.I.) & Muhammad Asif Raza (M.A.R.) prepared figures, the authors reviewed the manuscript, and Resource Funding.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Kousar, T., Rahim, M.S.M., Alsuhaibani, M. et al. Integrating convolutional and transformer networks for precise diagnosis of watershed and hemorrhagic stroke. Sci Rep 15, 41675 (2025). https://doi.org/10.1038/s41598-025-25539-7
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-25539-7