Abstract
Sequence data, such as nucleotides or amino acids, are crucial in advancing our understanding of biology. However, investigating and analyzing sequencing data and genotype–phenotype associations present several challenges, including noise components that arise from the sequencing, nonlinear genotype–phenotype associations, collinearity between input features, and high dimensionality of the input data. Machine learning (ML) algorithms have proven to be effective in detecting intricate and nonstructural patterns, making them a valuable tool for studies focused on genotype–phenotype associations. Yet, there needs to be more user-friendly ML implementations that leverage the unique features of high-volume DNA sequence data. Here, we introduce deepBreaks, a generic approach that detects important positions (genotypes) in sequence data that are associated with phenotypic traits. deepBreaks compares the performance of multiple ML algorithms and prioritizes positions based on the best-fit models. It is open-source software with online documentation and examples available at https://github.com/omicsEye/deepBreaks.
Similar content being viewed by others
Introduction
Advancements in sequencing and computational technologies have provided researchers with large-scale data, and the development of tools for analyzing such data is growing1. A major challenge in developing predictive DNA sequence-to-phenotype models is accounting for both the linear and non-linear effects of sequence variations on the phenotype while simultaneously considering the entire sequence which makes ML algorithms suitable to address these problems2. Decoding and interpreting results from predictive models are essential to make biological inferences3,4. Fitting machine learning (ML) and deep learning (DL) models on DNA sequence data to model traits has been studied in various frameworks, ranging from predicting drug resistance5,6,7 to cancer detection8,9. deepBreaks is developed to rank the performance of ML models that best fit the data and then, based on those models, prioritize and report the most discriminative positions of the sequence (genotypes) with respect to a given phenotype of interest. Early efforts in this field were implemented by a Bayesian method that used all marker data simultaneously to predict the phenotype10. ML approaches for genotype–phenotype associations have evolved, and some support more effective and reproducible use of multivariate genotype data for the prediction of quantitative traits11. Tools such as KOVER predict phenotypes based on reference-free genomes using k-mers (short DNA sequences) as the features, chi-square tests for filtering redundant features prior to modeling12, and for the features that have exact equal values, assign the same importance score12. Studies have attempted various approaches for predicting phenotypes based on the sequence, including using dense neural networks13, convolutional neural networks14,15, and ensemble learners14. Comparison of the outcomes of these different ML algorithms reveals no universally best predictive algorithm for the diversity of genotype–phenotype studies13. To find the best model suited for a given set of data, researchers compare several models and make inferences regarding feature importances only based on the best model16.
We developed a generic and computationally optimized tool, namely deepBreaks, to identify and prioritize important sequence positions in genotype–phenotype associations. Our approach is as follows: first, we prepare a training dataset based on the provided raw sequencing data. Second, we fit multiple ML algorithms and, based on their cross-validation score, select the best model. Then, we use this top model to find the most discriminative positions of the sequence. By doing this, we assess the phenotype’s predictability from sequences and use the most accurate models to identify and prioritize the most predictive sequence positions. This entails examining the variable components of the sequences to ascertain whether they are linked to the phenotype under investigation. It is essential to recognize that not all variable sites necessarily contribute to a phenotype17 and sometimes multiple mutations can contribute to the same phenotype18. Simply using the alignment or conservation score does not necessarily work; thus, our evaluation aims to discern the predictive potential of the phenotype, particularly in cases where the input sequence is a truncated segment or the phenotype, such as obesity or hair color, is influenced by factors beyond genetics16,19.
In this paper, we evaluate the performance of the deepBreaks approach on simulated data and assess its performance in finding the important variables in a dataset with ground truth. We have also applied deepBreaks to multiple datasets to show its wide applications and power to detect the most important positions in both nucleotide and amino acid sequence data. In the methods section, we elaborate on the steps that deepBreaks takes to prepare the data, fit models to the data, and interpret the results. deepBreaks is a generic software that can be applied in sequence-to-phenotype studies to show the feasibility of first predicting the phenotype based on a sequence and then determining what are the most discriminative parts of the sequence in predicting the phenotype.
Results
deepBreaks overview
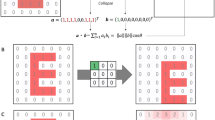
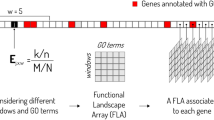
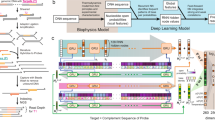
The input data of deepBreaks is a Multiple Sequence Alignment (MSA) file containing \({X}_{i}=({x}_{i1},{x}_{i2},...,{x}_{im})\), \(i\in \{\text{1,2},...,n\}\), n sequences of length \(m\) and a phenotypic metadata, with a vector of size n, and \({\pi }_{i}\) s, as phenotypes which are related to the \({i}^{th}\) sequence. \({x}_{ij}\) (the \({j}^{th}\) element of the sample \(i\)) can be a subset of \(\{A,T/U,C,G\}\), or any one-letter character representing an amino acid. Phenotypes (\({\pi }_{i}\) s) can be continuous measures such as height, BMI, categorical, or binary variables such as obese/healthy weight/underweight, antibacterial resistance/sensitivity, or mild/severe cases of a disease. deepBreaks has three phases described in Fig. 1a: i) preprocessing, ii) modeling, and iii) interpreting. In the data preprocessing phase, illustrated in Fig. 1b, we deal with imputing missing values, ambiguous reads, dropping zero-entropy columns, clustering correlated positions (features), and dropping redundant features which do not carry a significant amount of information in association with the phenotype under study. To keep track of positions before starting to drop the columns, all of them (by default) are named from \({p}_{1}\) to \({p}_{m}\). The names of the columns (positions) in the dataset are fixed, and dropping certain columns does not change the position names in a sequence. To identify the collinearity of features, we cluster features based on their pairwise distances. Then we cluster them using the density-based spatial clustering of applications with noise (DBSCAN) method20 algorithm and take the feature that is the closest to the center of the cluster as the representative of the cluster in the training data set21. We also normalize the training data using min–max normalization to have mean 0 and scale 1 before the training step. Two sets of models for continuous or categorical phenotypes are incorporated in our training model phase, and a complete list of these models with their default parameters is available in the Methods section. For model comparison, deepBreaks employs a default tenfold cross-validation method and ranks the models according to their average cross-validation score. Alternatively, users have the option to choose a train/test split. In either scenario, all preprocessing steps are exclusively applied to the training set. In a k-fold cross-validation study, this entails selecting the training folds and conducting preprocessing solely on those folds. The default performance metrics for regression and classification that deepBreaks uses are mean absolute error (MAE) and F-score, respectively. A complete list of other performance metrics is available in the Methods section. For interpreting the contribution of the positions in the sequence to the predictive models, we use the feature importance and coefficients. These importance values are then scaled to 0 and 1 (maximum importance). We also consider the same importance values for features that have been clustered together. We provide a detailed elaboration of the pipeline in the Methods section.
deepBreaks overall workflow. (a) deepBreaks begins with sequencing data organized in a multiple sequence alignment (MSA) format. Sequences can be nucleic acids or amino acids. The aligned sequences can be from a single region or concatenation of multiple regions. The phenotype of interest is also a required input parameter. Preprocessing steps are conducted to recode the sequencing data and phenotype into a format usable by the machine learning models. A modeling step follows in which various models are attempted and ranked. The best model is probed to identify the positions which best predict phenotype. These results are then merged and presented to the user as visualizations and interpretable tables. (b) The illustrated preprocessing steps implemented in the deepBreaks pipeline are essential for the approach efficiency by summarizing positions used in analyses and include dropping positions with no variation or data, imputing missing data, one-hot encoding, dropping insignificant positions, and finally clustering collinear features.
Simulation study
During data preprocessing, we drop redundant positions using p-value-based statistical tests and address feature collinearity by clustering correlated features and selecting a representative from each group. To evaluate the impact of these steps on estimating the effect of positions on the response variable (phenotype), we conducted a simulation study. We assessed the performance of three models (Adaboost, Decision Tree, and Random Forest) and their ability to estimate the true effect size of feature groups across various datasets and different preprocessing approaches. Additionally, we examined the effects of adjusting the threshold for both p-values and distance metrics.
Each data set is simulated based on this formula:
We first created a data matrix of size \({X}_{n\times m}\) with binary data (n samples and m features); then, we selected a subset of its m features as informative. The corresponding regression coefficients of these selected informative features were then sampled from a normal distribution, and the rest were considered zero. We call this data set the initial dataset which has no collinear features. Then, based on the initial dataset, we created datasets with collinear features. To add a collinear feature, we randomly selected one of the informative features and then replaced 40% of its values with random samples from a binary distribution. We repeat this process 5 times, and each time use the data set created in the last step as the initial data set. So, we start with data containing no collinear features and gradually increase their number, introducing different levels of collinearity that range up to moderate correlation. Figure 2a illustrates three steps of the simulation on a sample dataset with 10 features (4 informative, 6 redundant).
A simulation study design and results. (a) Creating training datasets with different levels of multicollinearity. The first dataset has independent features and only some of them are informative (1). Then, we start to replace the redundant features with samples from the informative features to add collinear features to the training dataset (2 to 5). For example, we took a sample from f1 and replaced it with f5 (2). (b–e) Results of simulation on a dataset with 1000 samples, 1000 features with 10 informative features. (f–i) Results of simulation on a dataset with 1000 samples, 2000 features with 20 informative features. Boxplots show the validation scores of each model with different levels of collinearity (x-axis) and line plots show the correlation between the true effect sizes and the predicted effect sizes by models. Collinearity (x-axis) zero means the starting dataset. (b, f) Using no clustering and no filtering. (c, g) Using no clustering but filtering redundant features with a p-value of 0.2. (d, h) Using clustering with a threshold of correlation 0.7 and no filtering. (e, i) Using clustering with a threshold of correlation of 0.7 and filtering redundant features with a p-value of 0.2. Machine learning algorithms used for evaluation include Adaboost (ada), Decision Tree (dt), and Random Forest (rf).
The correlations between sampled features in the dataset are a random number between 0.6 and 1 (exactly the same). With each data frame, we train models with different combinations of thresholds for correlation (no clustering, 0.7, and 0.9) and different thresholds for p-value (no filtering and 0.2) and then run 5 times repeated tenfold cross-validation on the data set. Finally, each data set with each of the combinations results in 50 performance metrics for the model and one value for the correlation between the estimated effect size and the true effect size. Here, we use correlation to evaluate how effectively each combination of analyses predicts the importance of a position, providing a quantitative measure of the model’s ability to identify true associations. We used two different datasets as the initial datasets. Dataset 1, has 1000 samples and 1000 features with 10 informative features, and dataset 2 has 1000 samples and 2000 features with 20 informative features.
We can see that in Fig. 2, the performance of the models stays in the same range, and preprocessing methods do not affect the predictive performances. On average, for dataset 1, Adaboost ~ 0.80, Decision Tree ~ 0.83, and Random Forest ~ 0.84, and for dataset 2 Adaboost ~ 0.74, Decision Tree ~ 0.59, and Random Forest ~ 0.69. However, suppose we do not cluster the collinear features together. In that case, we can see that the ability of the models to estimate the true effect sizes decreases substantially (Fig. 2b, c, f, g). The complete code for simulation and visualization of the results is available at https://github.com/omicsEye/deepbreaks/blob/master/simulation/simulation.ipynb. We extended our simulated data to benchmark performance against existing tools such as pyseer22 which applies linear models to evaluate the influence of genetic variation on a phenotype of interest, and demonstrated that deepBreaks outperforms pyseer in the presence of collinearity (Fig. S1), a common characteristic of sequence data positions.
deepBreaks identifies amino acids associated with color sensitivity
Opsins are genes involved in light sensitivity and vision, and when coupled with a light-reactive chromophore, the absorbance of the resulting photopigment influences physiological phenotypes like color sensitivity (Fig. 3a). We analyzed the amino acid sequence of rod opsins because previously published mutagenesis work established mechanistic connections between 12 specific amino acid sites and phenotypes23. Therefore, we hypothesized that ML approaches could predict known associations between amino acid sites and absorbance phenotypes. We identified opsins expressed in rod cells of vertebrates (mainly marine fishes) with absorption spectra measurements (λmax, the wavelength with the highest absorption). The dataset contains 175 samples of opsin sequences, including samples with experimental mutations. Amino acid sequences were aligned with multiple alignment using fast Fourier transform (MAFFT)24 and trimmed with GBlocks25 (a program used to trim multiple sequence alignments) (see Frazer et al.26 for details). We next applied deepBreaks on this dataset to find the most important sites contributing to the variations of λmax. We analyzed the data with a tenfold cross-validation, and the best-performing model on the validation sets was random forest (R2 = 0.54, MAE 4.4, MSE = 46, RMSE = 6.3, MAPE = 0.0087) followed by Lasso and Lasso Least Angle Regression (Lars). We then performed hyperparameter tuning using the default parameters provided by deepBreaks. This involved selecting the maximum features for the random forest using either the square root or log2 functions and adjusting the alpha parameter for both Lasso and Lasso Lars models. Specifically, for the Lasso and Lasso Lars models, we searched for the optimal alpha values in the range of 0.01 to 100, using five evenly spaced values within this range. For the random forest model, we explored using either the square root or log2 of the number of features as the maximum number of features. We found sites 37, 39, 50, 83, 124, 127, 137, 158, 165, 173, 225, 261, and 292 to be important in terms of affecting the λmax (Fig. 3b). Some of these sites are known from published mutagenesis experiments23 to strongly affect λmax. Frazer et al.26 investigated the associations and looked deeply at how some of these mutations affect the λmax such as A292S and F261Y. For a more detailed investigation of the positions and their association with λmax please see Frazer et al.26. Figure 3c illustrates the effects of mutations in positions 261 and 292 of the sequences.
Anatomy of the eye involved in light sensing. (a) The overall view from a vertebrate eye, structure of the retina, and position of opsins in rod and cone cells. The figure is created using BioRender at https://www.biorender.com/. (b) Important positions in opsin amino acid sequences. (c) Functional changes in positions 292 and 261 as exemplary with LambdaMax associated with each amino acid variant. Panels b and c are outputs of deepBreaks.
deepBreaks identifies HIV regions with potentially important functions
Subtypes of the human immunodeficiency virus type 1 (HIV-1) group M are different in the envelope (Env) glycoproteins of the virus. These parts of the virus are displayed on the surface of the virion and are targets for both neutralizing antibodies and cell-mediated immune responses27. The third hypervariable domain (V3) of HIV-1 gp120 is a cysteine-bounded loop structure usually composed of 105 nucleotides and labeled as the base (nu 1:26 and 75:105), stem (nu 27:44 and 54:74), and turn (nu 45:53) regions27. Among all of the hyper-variable regions in gp120 (V1-V5), V3 plays the main role in the virus infectivity28. Here, we use deepBreaks to identify regions in the V3 loop that are important in terms of associating the V3 sequences to subtypes B and C. We used the publicly available Los Alamos HIV Database29 (www.hiv.lanl.gov) to gather the aligned nucleotide sequences of the V3 loop of subtypes B and C. We then dropped the repeated samples from the same patients and the final dataset contained 35,424 sequences with a combination of 24,042 (67.87%) sequences of subtype B and 11,382 (32.13%) sequences of subtype C. The maximum length of the sequences was 105 nucleotides. Three distinct communities (clusters) with potentially different biological functions were detected using the omeClust30 (a zoom-out approach) based on V3 loop distances between samples, suggesting there are variations in the V3 loop with potential functions (Fig. 4a). We then implemented deepBreaks, a zoom-in approach to identifying important mutations (Fig. 4b–d). We split the dataset into 80% train (28340 samples) and 20% test (7084 samples) datasets and then performed a tenfold cross-validation on the train set to find the best model. Figure 5b shows the performance metrics of the top 2 models on test data as well as cross-validation scores. We then performed hyperparameter tuning using the default parameters provided by deepBreaks for the Extremely Randomized Trees (Extra Trees) model. Specifically, we searched for the optimal maximum depth values of 4, 6, and 8, and the number of estimators set to either 500 or 1000. We used a much larger dataset than the previous studies, and the important positions reported by deepBreaks cover all the previously detected positions31. Also, several additional positions are reported to be significantly different that were not mentioned previously. As previously investigated, the most important changes are in the stem and turn parts of the V3 loop27,31 (Fig. 4c). It has been shown that these sequence differences cause specific phenotypic traits and affect the role of V3 loop in both virus replication and virus-host interaction27,31,32. Among the important positions, p52,p53, and p38,p39 show higher importance (Fig. 4d). It has been investigated that these positions are responsible for the structure of turn in both subtypes B and C31.
Classification of HIV-1 subtypes B and C based on nucleotide sequences of the V3 loop. (a) Cluster analysis of the sequences with ground truth labels from the Los Alamos National Lab database. (b) Results of tenfold cross-validation (cv) and test data of top 2 classification models, XGBOOST (xgb) and lightGBM (lgbm), trained to predict the subtypes of the HIV-1 based on the V3 loop. (c) Important positions reported by deepBreaks based on the results of the top three models labeled with the sections of the sequence. (d) Stacked bar plots of the top 5 positions that contribute to the classification models for predicting HIV subtypes ‘B’ and ‘C’.
Mutations in Haemophilus parainfluenzae from human oral systems are associated with sampling sites. (a) Aggregated importance of positions across the top three models. (b) Stacked barplot of three top positions showing the frequency of nucleotides in each position for different niches.
Novel insights of niche associations in the oral microbiome
Bacterial communities evolve to adapt to novel environmental conditions33,34, and microbial species tend to adapt at the genome level to the niche in which they live. Here, we use microbial strain representatives from metagenomics data of healthy adults from the Human Microbiome Project35. Each microbial strain representative is a concatenation of marker genes using the StrainPhlAn tool36 which generates MSA files per species across samples (each row in the MSA file represents the species in the corresponding sample). The input for deepBreaks consists of 1) an MSA file of nucleotide sequences with 1006 rows, each a representative strain of a specific microbial species, here Haemophilus parainfluenzae, with 49839 lengths of only marker genes used by strainPhlAn; and 2) labels for deepBreaks prediction are body sites from which samples were collected: buccal mucosa, supragingival plaque, and tongue dorsum. We split the dataset into 80% train (805 samples) and 20% test (201 samples) datasets and then performed a 75%-25% train-validation split on the train set to find the best model. The gradient boosting classifier performed the best on the validation set (Accuracy = 0.89, AUC = 0.96, F1 = 0.88, Recall = 0.88, Precision = 0.88), and it also performed the best on the test set (Accuracy = 0.89, AUC = 0.96, F1 = 0.87, Recall = 0.87, Precision = 0.87). Figure 5a illustrates the important positions in the sequence, and there are some positions within a specific range that have similar importance values. With respect to location, buccal mucosa and supragingival plaque are closer37 and have similar mutation rates compared to tongue dorsum (Fig. 5b). The mutations block of position 2100–2350 are specifically in energy transducer TonB protein (Pfam Id: PF03544, Gen bank: QOR16081.1). In gram-negative bacteria, which have a distinctive two-membrane envelope structure, TonB plays a crucial role in the energy transduction system from cytoplasmic membrane (CM) to the outer membrane (OM)38,39. The OM helps gram-negative bacteria to adapt and thrive in diverse environments, and mutations in Tonb can affect the nutrient transfer to OM38. TonB-ExbB-ExbD complex transfers energy from protons in Gram-negative bacteria (e.g., Haemophilus parainfluenzae in different oral sites), helping outer membrane receptors absorb iron40. This suggests environmental conditions, such as pH and temperature in oral sites, potentially cause microbial species mutation to adapt to the niche in which they live. The mutations in a block of positions 20150–20200 are associated with mutations in outer membrane protein assembly factor BamC (NCBI protein ID: QOR23430.1). BamC is a member of a complex group of proteins known as β-barrel assembly machinery (BAM), which are responsible for folding and inserting OM proteins into the membrane41. The BAM complex functions specifically in the folding and insertion of nascent β-barrel outer membrane proteins, which highlights its essential role in protein transport across the membrane rather than in the processing of all outer membrane proteins41 and it is shown that mutation in bam complex in Escherichia coli helps the bacteria to survive under hostile environments and develop antibiotic resistance42. These findings show that mutations in proteins that are associated with the OM proteins play important roles in helping the bacteria live in different niches.
deepBreaks reveals important SARS-CoV-2 regions associated with Alpha and Omicron variants
Without a procedure for correcting the replication, RNA viruses are more prone to mutations than organisms with DNA-based genomes43. Most mutations in the SARS-CoV-2 genome do not affect the functioning of the virus. However, mutations in the spike protein of SARS-CoV-244, which binds to receptors on cells lining the inside of the human nose, may make the virus easier to spread or affect how well vaccines protect people. Other mutations may lead to SARS-CoV-2 being less responsive to treatments for COVID-1945,46. Variants of SARS-CoV-2 have been categorized into multiple variants, but based on their effect on public health, five of these—Alpha, Beta, Delta, Gamma, and Omicron—have been labeled as variants of concern and associated with enhanced transmissibility and increased virulence47,48. We used the publicly available data from GSAID49 and obtained 10,000 sequences of Spike protein region for SARS-CoV-2 samples of the Alpha variant—one of the first variants of concern identified by the World Health Organization (WHO)—and 10,000 sequences of the spike protein region for SARS-CoV-2 samples of the Omicron variant—one of the newest variants of concern identified by the WHO. Then, we used MAFFT24 with PAM 20050 to align these sequences with the reference sequence spike protein (NCBI accession: NC_045512.2). The final data set after dropping the replicates consisted of 9863 sequences of the Alpha variant and 9618 sequences of the Omicron variant (19481 total samples). Then, we used deepBreaks to analyze the data and find the most important (predictive) positions in these sequences in terms of classifying the variants (Fig. 6a). We split the dataset into 80% train (15585 samples) and 20% test (3897 samples) dataset and then performed a 70%-30% train-validation split on the train set to find the best model. The gradient boosting classifier performed the best on the validation set (Accuracy = 0.99, AUC = 0.99, F1 = 0.99, Recall = 0.99, Precision = 0.99), and it also performed the best on the test set (Accuracy = 0.99, AUC = 0.99, F1 = 0.99, Recall = 0.99, Precision = 0.99). We then performed hyperparameter tuning using the default parameters provided by deepBreaks for the Random Forest and Gradient Boosting Classifier (GBC) models. For the Random Forest model, we explored using either the square root or log2 of the number of features as the maximum number of features. For the Gradient Boosting Classifier, we tested maximum depth values ranging from 3 to 5, maximum features using either the square root or log2 of the number of features, 200, 500, and 800 estimators, and learning rates ranging from 0.001 to 0.1 with two evenly spaced values. The mutations in this part of the sequence were highly correlated and happened almost concurrently. This was also mentioned in an earlier study46. We have shown 6 of the positions with mutations in these sequences and their detailed changes in Fig. 6b. These differences between the Omicron and Alpha variants, observed mostly concurrently in the spike protein, have different effects on the virus (see Supplement).
Classifying the SARS-CoV-2 variants based on the spike protein sequences. (a) Important positions in the spike protein (S) of SARS-CoV-2 in terms of predicting Alpha and Omicron variants. (b) Details of how the mutations appear in variants. Insertions and deletions are marked as ‘GAP’, used internally to avoid parsing conflicts and ensure robust preprocessing across libraries.
Discussion
In this study, we provided an integrated generic approach to find the most discriminative changes in a sequence in association with a given phenotype of interest. Our approach is based on first training accurate ML models and then using their information to interpret the predictive power of each position in the sequence data. However, having an accurate predictive model for sequence-to-phenotype studies is a challenging task. One of the major challenges in training an accurate model is rooted in the high-dimensional MSA files with lengthy sequences and a limited number of samples, known as the curse of dimensionality51. The other tremendous challenge for training and interpreting the models is the collinearity between positions of an MSA file which has a negative effect on the performance of the models21. We showed that by implementing multiple filtering methods in the data preprocessing step, deepBreaks not only finds the most accurate model based on the given data but also allows for the interpretation of the trained models. To justify our approaches in terms of dealing with the redundancy of features and multicollinearity, we also conducted a simulation study on multiple datasets with different levels of collinear features. The results of these simulation studies show that our method not only reduces data dimensionality by clustering collinear features and removing redundancy, but also enables models to accurately estimate feature importance under varying levels of multicollinearity. In contrast, existing tools such as pyseer exhibit a decline in performance in the presence of feature collinearity (see Supplement).
We also evaluated the performance of deepBreaks on real data with 4 different datasets. For each study, deepBreaks pointed out the important positions in the sequences relative to phenotypes of interest. Some of the reported positions were targeted for individual experimental studies, and some of the important positions have not been mentioned in the literature before, opening new topics for further research. Finally, by applying deepBreaks in different scenarios ranging from predicting a continuous phenotype, such as light sensitivity with amino acid sequences of opsins, to categorical phenotypes, such as different niches of Haemophilus parainfluenzae based on its genome sequence, and finding significant results, we showed its wide applicability.
Methods
Approach
ML is a field of inquiry devoted to understanding and building methods that ‘learn’, that is, methods that leverage data to improve performance on some set of tasks52. Supervised learning, which is a branch of ML, aims to find a function \(f\) that maps input data \(X\), to output variable \(y\) through a training set of \(t = \{({X}_{1}, {\pi }_{1}), ({X}_{2}, {\pi }_{2}), ..., ({X}_{n}, {\pi }_{n})\}\). A supervised learning algorithm produces \(f({X}_{i})\) as an estimation for the \({\pi }_{i}\) (phenotype value for sample \(i\)). Supervised learning algorithms are designed to enhance their performance by minimizing the distance \(\| f({X}_{i})-{\pi }_{i}{\| }_{\mathcal{H}}\)53 for an appropriate choice of norm space \(\mathcal{H}\). Setting \(\mathcal{H}={\mathbb{L}}^{2}\) turns this optimization into a least square approach. In our case, \({X}_{i}=({x}_{i1},{x}_{i2},...,{x}_{im})\), \(i\in \{\text{1,2},...,n\}\) are sequences of length \(m\) (\(m\) nucleotides or amino acids) and \({p}_{i}\) s are phenotypes related to the \({i}^{th}\) sequence. For example \({x}_{ij}\) (the \({j}^{th}\) element of the sample \(i\)) can be a subset of \(\{A,T/U,C,G\}\), or any amino acid. Phenotypes (\({\pi }_{i}\) s) can also be a continuous measure such as light sensitivity, or categories such as virus subtype, niches of a bacterial species, or strains of a virus. Assuming that raw data (sequences and phenotypes) are provided, actions are still required to prepare the data for ML algorithms. deepBreaks has three phases: i) preprocessing the data, ii) fitting models to preprocessed data and comparing them, and iii) providing interpretable tables and visualizations for top predictor(s).
Preprocessing
Data preprocessing is a fundamental step for any ML algorithm. Sequence data may contain missing values, ambiguous reads, zero-entropy columns, correlated positions (features), and redundant features which do not carry a significant amount of information in association with the phenotype under study. The deepBreaks pipeline for preprocessing starts with dropping columns in the dataset that contain missing values over a certain threshold. The default threshold is 70% of the sample size. So, if we have 1000 samples, we drop all the positions that have over 700 missing values from the training set (this value can be changed based on user preference). Dropping the zero-entropy (constant) features from the dataset is the next step. It is worth mentioning that before starting to drop the columns, all of them (by default) are named from \({p}_{1}\) to \({p}_{m}\). The names of the columns (positions) in the dataset are fixed, and dropping certain columns does not change the position names in a sequence. The subsequent step involves handling missing values for the remaining positions. We address this by either labeling them as insertion/deletion (GAP) or imputing them with the mode (most frequent read) of the respective position. If the proportion of missing values for a position exceeds a specified threshold (default is 5%), we consider those missing values as GAP. Conversely, positions with missing values below the threshold are imputed using the mode of the corresponding position. Additionally, we handle rare values in a position by identifying characters that appear less frequently than a specified threshold (the default is 2%). If there is only one rare character, we replace these rare characters with the mode (most common character) in the column or combine multiple rare characters into a single “others” category (concatenating all the rare characters), ensuring more robust data for analysis or modeling. After this step, as we modified the reads, positions are again checked for entropy, and positions with zero entropy will be dropped from the training set which also saves us some computational cost. After handling missing values, the subsequent step involves reducing the number of positions in the training dataset by performing either chi-square tests (for categorical phenotypes) or Kruskal–Wallis tests (for continuous phenotypes). This helps in identifying and eliminating redundant positions. We apply one-hot encoding to the categorical features to prepare the data for these tests. To prevent multicollinearity, we employ the technique of dropping one category from each one-hot encoded set. Specifically, for a position \({p}_{i}\) with \(k\) levels, we generate \(k-1\) binary indicator variables, effectively excluding one level to ensure the encoded matrix remains full rank. We use these statistical tests to assess the significance of each position by running tests on all the positions against the phenotype one by one. Those features where the p-value of their test against the phenotype is less than a threshold54 (default p-value = 0.25) will be dropped. A list of all features and the corresponding p-values will be provided as a report to the user. As we consider each position in the sequences as a feature of our training dataset, we need to check for collinearity between our predictive variables, as it can cause issues for parameter estimation21. Then, we use a distance function that calculates the pairwise distances between features. The list of available metrics are Spearman correlation55, hamming56, Jaccard57, normalized mutual information58, adjusted mutual information59, and adjusted Rand score60. The result of this step is a symmetric distance matrix with values between 0 (being the same) to 1 (uncorrelated). We then use this symmetric matrix of distance values and feed it into the density-based spatial clustering of applications with noise (DBSCAN) method20 for clustering the features based on their pairwise distances. The reason behind this is to cluster the features that provide the same information21. After that, we select one feature from each cluster by keeping the one that is the closest to the center of the cluster as the representative of that cluster and drop the rest of the features in that cluster from the training set. Although we drop the rest of the features in each cluster except the representative, we keep the information of the members of the clusters for interpretation after the modeling step. The default parameters of the DBSCAN, epsilon (distance between centers), and the minimum points for a cluster are set to 0.2 and 2, respectively. The remaining variables are then standardized to have a mean of 0 and a variance of 1 to have a common scale, preventing features with larger variances from overshadowing smaller ones. This minimizes bias in features with higher numeric values, ensuring fair contributions to discriminating pattern classes and maintaining equal feature importance in prediction. This approach is especially valuable for statistical learning methods, where all features equally contribute to the learning process61.
Models
We use different sets of models for continuous or categorical phenotypes. For continuous phenotypes, we fit linear regression, Ridge Regression62, Lasso Regression63, Bayesian Regression, Lasso Least Angle Regression64, Huber Regressor65, Extremely Randomized Trees (Extra Trees)66, Extreme Gradient Boosting (XGBoost)67, Light Gradient Boosting Machine (LightGBM)68, Random Forest69, Decision Tree70, and AdaBoost71. For problems with a categorical phenotype, we use Extra Trees, XGBoost, LightGBM, Random Forest, Decision Tree, AdaBoost, Gradient Boosting, and Logistic Regression. For all the above-mentioned models, we use the default hyperparameters from the scikit-learn library in python72 and a grid search (expandable by user preference) parameter set that is provided in the documentation. For model comparison, deepBreaks, by default, uses a tenfold cross-validation approach and ranks the models based on their average cross-validation score. K-fold cross-validation is a resampling method that partitions the whole dataset into k separate equal-sized parts and then uses k-1 parts for training the model and 1 part for testing the performance. This process is repeated k times, and the average score of all k models is called the cross-validation score.
The default performance metrics for regression and classification that deepBreaks uses are Mean Absolute Error (MAE) and F-score.
The default list of metrics that deepBreaks reports is provided in the documentation, and the user can provide predefined custom metrics or a set of metrics from the scikit-learn library in Python72.
Interpretation
For interpreting the contribution of sequence positions to the predictive models, we use the feature importance, coefficients, and weights, as different algorithms have different kinds of output. For xgboost73 and LightGBM74 the reported feature importance represents the number of times a feature appears in a tree. For AdaBoost, random forest, decision tree, extra tree, and gradient boosting, the importance of a feature is its Gini importance, which is computed as the normalized total reduction of the criterion brought by that feature72.
If we have \(N\) data samples (rows in the train set) that reache the node \(j\) of a tree and \(G\) be the impurity of the node \(j\), the importance of node \(j\), \({ion}_{j}\) is calculated as follows:
Based on this, the feature importance value of the \({i}^{th}\) feature is:
And then, we can normalize the feature importance value for the \({i}^{th}\) feature by dividing it by the sum of the importance of all the features:
These calculations are just based on one tree and in other tree-based algorithms such as random forest, AdaBoost, and extra tree, the final importance of a feature is its average over all of the fitted trees. For the linear models, the regression coefficients or weight is considered as the feature importance72.
In the preprocessing phase, we one-hot encoded the positions into training features, so, each position (based on the number of its unique characters), is transformed into one or more than one feature. For example, if position \(pi\) consists of \(\left\{A,T,C\right\}\), its features are \(pi\_A\) and \(pi\_T\) in the form of one-hot encoded features (we drop \(pi\_C\) to avoid collinearity). These features have separate importances, and we average the absolute value of their importance and the importance of position \(pi\) is the average of importances of the feature \(pi\_A\) and \(pi\_T\). After that, to normalize the importances, all the importances will be divided by their maximum value. We consider zero importance for all the positions that have been dropped during the preprocessing steps. For all those positions that were in the same group and dropped based on their distance values and DBSCAN clustering, we consider the same feature importance value. deepBreaks, by default, considers the top three best-fitted models, but the user can change this number.
deepBreaks output
deepBreaks creates reports of 1) p-values from statistical tests for each position against a phenotype, 2) the related distance matrix, 3) clusters of correlated positions, 4) a table of fitted models with their performance metrics, 5) feature importance values for each of the top models and their merged results, 6) plots for importance values based on individual models and merged results 7) box plot (continuous phenotype) or stacked bar plot (categorical phenotype) for most discriminative positions 8) top fitted and tuned models as .pkl files, 9) all the visualizations as .pkl files, 10) a text file with all the important positions information along with the highlighted position, and 11) a report on the performance of the models on the test dataset. In the importance report file, we detail the significance of each binary feature derived from positions. To enhance visualization, especially in cases where positions encompass multiple training features (e.g., those with more than two nucleotide types), we adopt the maximum value (though users have the flexibility to opt for alternative computations like mean or median) as the default indicator of importance for a given position. Consequently, when generating importance plots, the bar heights reflect the maximum importance value among all training features associated with that position.
deepBreaks implementation
In the data preprocessing, we use NumPy75, Pandas76, and SciPy77 Python libraries. For model comparisons and cross-validation pipeline, we use scikit-learn72, XGBoost67, and lightgbm68. Visualizations generated by deepBreaks make use of the seaborn78 and matplotlib79 libraries in Python. We used deepBreaks 1.1.4 for all the applications and evaluations in this manuscript.
Data availability
To facilitate the usage of this tool, we have provided it as an open-source Python library with various sample codes, tutorials for installation and usage, data used in this study and the Jupyter notebooks that were used to produce the results at https://github.com/omicsEye/deepbreaks. A Docker image of *deepBreaks* with instructions on how to run the tool is available at https://hub.docker.com/r/omicseye/deepbreaks-dc.
References
Ritchie, M. D., Holzinger, E. R., Li, R., Pendergrass, S. A. & Kim, D. Methods of integrating data to uncover genotype–phenotype interactions. Nat. Rev. Genet. 16, 85–97 (2015).
Moore, J. H., Asselbergs, F. W. & Williams, S. M. Bioinformatics challenges for genome-wide association studies. Bioinformatics 26, 445–455 (2010).
Doshi-Velez, F. & Kim, B. Towards A Rigorous Science of Interpretable Machine Learning. arXiv [stat.ML] (2017).
Leung, M. K. K., Delong, A., Alipanahi, B. & Frey, B. J. Machine learning in genomic medicine: A review of computational problems and data sets. Proc. IEEE 104, 176–197 (2016).
Yang, Y. et al. Machine learning for classifying tuberculosis drug-resistance from DNA sequencing data. Bioinformatics 34, 1666–1671 (2018).
Hadikurniawati, W., Anwar, M. T., Marlina, D. & Kusumo, H. Predicting tuberculosis drug resistance using machine learning based on DNA sequencing data. J. Phys. Conf. Ser. 1869, 012093 (2021).
Adam, G. et al. Machine learning approaches to drug response prediction: challenges and recent progress. NPJ Precis Oncol 4, 19 (2020).
Wan, N. et al. Machine learning enables detection of early-stage colorectal cancer by whole-genome sequencing of plasma cell-free DNA. BMC Cancer 19, 832 (2019).
Kurian, B. & Jyothi, V. L. Breast cancer prediction using an optimal machine learning technique for next generation sequences. Concurrent Eng. Res. Appl. 29, 49–57 (2021).
Lee, S. H., van der Werf, J. H. J., Hayes, B. J., Goddard, M. E. & Visscher, P. M. Predicting unobserved phenotypes for complex traits from whole-genome SNP data. PLoS Genet. 4, e1000231 (2008).
Guzzetta, G., Jurman, G. & Furlanello, C. A machine learning pipeline for quantitative phenotype prediction from genotype data. BMC Bioinf. 11(Suppl 8), S3 (2010).
Drouin, A. et al. Predictive computational phenotyping and biomarker discovery using reference-free genome comparisons. BMC Genomics 17, 754 (2016).
Montesinos-López, A., Montesinos-López, O. A., Gianola, D., Crossa, J. & Hernández-Suárez, C. M. Multi-environment genomic prediction of plant traits using deep learners with dense architecture. G3 8, 3813–3828 (2018).
Ma, W. et al. A deep convolutional neural network approach for predicting phenotypes from genotypes. Planta 248, 1307–1318 (2018).
Liu, Y. et al. Phenotype prediction and genome-wide association study using deep convolutional neural network of soybean. Front. Genet. 10, 1091 (2019).
Lee, Y.-C. et al. Using machine learning to predict obesity based on genome-wide and epigenome-wide gene-gene and gene-diet interactions. Front. Genet. 12, 783845 (2021).
Wang, L., Shen, H., Liu, H. & Guo, G. Mixture SNPs effect on phenotype in genome-wide association studies. BMC Genomics 16, 3 (2015).
GTEx Consortium. The GTEx Consortium atlas of genetic regulatory effects across human tissues. Science 369, 1318–1330 (2020).
Orgogozo, V., Morizot, B. & Martin, A. The differential view of genotype–phenotype relationships. Front. Genet. 6, 179 (2015).
A Density-Based Algorithm for Discovering Clusters in Large. https://www.aaai.org › KDD › 1996 › KDD96-037https://www.aaai.org › KDD › 1996 › KDD96-037.
Dormann, C. F. et al. Collinearity: a review of methods to deal with it and a simulation study evaluating their performance. Ecography 36, 27–46 (2013).
Lees, J. A., Galardini, M., Bentley, S. D., Weiser, J. N. & Corander, J. Pyseer: A comprehensive tool for microbial pangenome-wide association studies. Bioinformatics 34, 4310–4312 (2018).
Yokoyama, S., Tada, T., Zhang, H. & Britt, L. Elucidation of phenotypic adaptations: Molecular analyses of dim-light vision proteins in vertebrates. Proc. Natl. Acad. Sci. U. S. A. 105, 13480–13485 (2008).
Katoh, K., Rozewicki, J. & Yamada, K. D. MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Brief. Bioinform. 20, 1160–1166 (2017).
Talavera, G., Castresana, J., Kjer, K., Page, R. & Sullivan, J. Gblocks. http://molevol.cmima.csic.es/castresana/Gblocks.html.
Frazer, S. A., Baghbanzadeh, M., Rahnavard, A., Crandall, K. A. & Oakley, T. H. Discovering genotype–phenotype relationships with machine learning and the visual physiology opsin database (VPOD). GigaScience 13, giae073 (2024).
Lynch, R. M., Shen, T., Gnanakaran, S. & Derdeyn, C. A. Appreciating HIV type 1 diversity: subtype differences in Env. AIDS Res. Hum. Retroviruses 25, 237–248 (2009).
Felsövályi, K., Nádas, A., Zolla-Pazner, S. & Cardozo, T. Distinct sequence patterns characterize the V3 region of HIV type 1 gp120 from subtypes A and C. AIDS Res. Hum. Retroviruses 22, 703–708 (2006).
Compendium, H. Foley B, LT, Apetrei C, Hahn B, Mizrachi I, Mullins J, Rambaut A, Wolinsky S & Korber B, Eds. Biophysics Group, Los Alamos National Laboratory
Rahnavard, A. et al. Omics community detection using multi-resolution clustering. Bioinformatics 37, 3588–3594 (2021).
Patel, M. B., Hoffman, N. G. & Swanstrom, R. Subtype-specific conformational differences within the V3 region of subtype B and subtype C human immunodeficiency virus type 1 Env proteins. J. Virol. 82, 903–916 (2008).
Fouchier, R. A. et al. Phenotype-associated sequence variation in the third variable domain of the human immunodeficiency virus type 1 gp120 molecule. J. Virol. 66, 3183–3187 (1992).
Culyba, M. J. & Van Tyne, D. Bacterial evolution during human infection: Adapt and live or adapt and die. PLoS Pathog. 17, e1009872 (2021).
McDonald, M. J. Microbial Experimental Evolution - a proving ground for evolutionary theory and a tool for discovery. EMBO Rep. 20, e46992 (2019).
Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature 486, 207–214 (2012).
Truong, D. T., Tett, A., Pasolli, E., Huttenhower, C. & Segata, N. Microbial strain-level population structure and genetic diversity from metagenomes. Genome Res. 27, 626–638 (2017).
Lloyd-Price, J. et al. Strains, functions and dynamics in the expanded human microbiome project. Nature 550, 61–66 (2017).
Brinkman, K. K. & Larsen, R. A. Interactions of the energy transducer TonB with noncognate energy-harvesting complexes. J. Bacteriol. 190, 421–427 (2008).
Samantarrai, D., Lakshman Sagar, A., Gudla, R. & Siddavattam, D. TonB-dependent transporters in sphingomonads: unraveling their distribution and function in environmental adaptation. Microorganisms 8, 359 (2020).
Biou, V. et al. Structural and molecular determinants for the interaction of ExbB from Serratia marcescens and HasB, a TonB paralog. Commun. Biol. 5, 355 (2022).
Knowles, T. J., Scott-Tucker, A., Overduin, M. & Henderson, I. R. Membrane protein architects: The role of the BAM complex in outer membrane protein assembly. Nat. Rev. Microbiol. 7, 206–214 (2009).
Georgieva, M. et al. Mutations in the essential outer membrane protein BamA contribute to Escherichia coli resistance to the antimicrobial peptide TAT-RasGAP317-326. J. Biol. Chem. 301, 108018 (2025).
Holmes, E. C. What does virus evolution tell us about virus origins?. J. Virol. 85, 5247–5251 (2011).
Rahnavard, A. et al. Epidemiological associations with genomic variation in SARS-CoV-2. Sci. Rep. 11, 23023 (2021).
Lauring, A. S. & Malani, P. N. Variants of SARS-CoV-2. JAMA https://doi.org/10.1001/jama.2021.14181 (2021).
Ilmjärv, S. et al. Concurrent mutations in RNA-dependent RNA polymerase and spike protein emerged as the epidemiologically most successful SARS-CoV-2 variant. Sci. Rep. 11, 13705 (2021).
Aleem, A., Akbar Samad, A. B. & Slenker, A. K. Emerging Variants of SARS-CoV-2 And Novel Therapeutics Against Coronavirus (COVID-19). in StatPearls (StatPearls Publishing, Treasure Island (FL), 2022).
Tracking SARS-CoV-2 variants. https://www.who.int/activities/tracking-SARS-CoV-2-variants.
Khare, S. et al. GISAID’s role in pandemic response. China CDC Wkly 3, 1049–1051 (2021).
Dayhoff, M., Schwartz, R. & Orcutt, B. 22 a model of evolutionary change in proteins. Atlas Protein Sequence Struct. 5, 345–352 (1978).
Clarke, R. et al. The properties of high-dimensional data spaces: implications for exploring gene and protein expression data. Nat. Rev. Cancer 8, 37–49 (2008).
Hierons, R. Machine learning. Tom M. Mitchell. Published by McGraw-Hill, Maidenhead, U.K., International Student Edition, 1997. ISBN: 0-07-115467-1, 414 pages. Price: U.K. £22.99, soft cover. Software Testing, Verification and Reliability vol. 9 191–193 Preprint at https://doi.org/10.1002/(sici)1099-1689(199909)9:3<191::aid-stvr184>3.0.co;2-e (1999).
Nordhausen, K. The Elements of Statistical Learning: Data Mining, Inference, and Prediction, Second Edition by Trevor Hastie, Robert Tibshirani, Jerome Friedman. International Statistical Review vol. 77 482–482 Preprint at https://doi.org/10.1111/j.1751-5823.2009.00095_18.x (2009).
Hosmer, D. W. Jr., Lemeshow, S. & Sturdivant, R. X. Applied Logistic Regression (John Wiley & Sons, 2013).
Coefficient, S. R. C. In The Concise Encyclopedia of Statistics. Preprint at (2008).
Hamming Distance. in Encyclopedia of Biometrics (eds. Li, S. Z. & Jain, A.) 668–668 (Springer US, Boston, MA, 2009).
Hancock, J. M. Jaccard Distance (Jaccard Index, Jaccard Similarity Coefficient). in Dictionary of Bioinformatics and Computational Biology (2014).
Kvalseth, T. O. Entropy and correlation: some comments. IEEE Trans. Syst. Man Cybern. 17, 517–519 (1987).
Vinh, N. X., Epps, J. & Bailey, J. Information theoretic measures for clusterings comparison. Proceedings of the 26th Annual International Conference on Machine Learning—ICML ’09 Preprint at https://doi.org/10.1145/1553374.1553511 (2009).
Morey, L. C. & Agresti, A. The measurement of classification agreement: An adjustment to the rand statistic for chance agreement. Educ. Psychol. Meas. 44, 33–37 (1984).
Singh, D. & Singh, B. Investigating the impact of data normalization on classification performance. Appl. Soft Comput. 97, 105524 (2020).
Hoerl, A. E. & Kennard, R. W. Ridge regression: biased estimation for nonorthogonal problems. Technometrics 12, 55–67 (1970).
Santosa, F. & Symes, W. W. Linear inversion of band-limited reflection seismograms. SIAM J. Sci. and Stat. Comput. 7, 1307–1330 (1986).
Efron, B., Hastie, T., Johnstone, I. & Tibshirani, R. Least angle regression. aos 32, 407–499 (2004).
Huber, P. J. & Ronchetti, E. M. Robust Statistics. Wiley Series in Probability and Statistics Preprint at https://doi.org/10.1002/9780470434697 (2009).
Geurts, P., Ernst, D. & Wehenkel, L. Extremely randomized trees. Mach. Learn. 63, 3–42 (2006).
Chen, T. & Guestrin, C. XGBoost. Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining Preprint at https://doi.org/10.1145/2939672.2939785 (2016).
Ke, Meng, Finley & Wang. Lightgbm: A highly efficient gradient boosting decision tree. Adv. Neural Inf. Process. Syst.
Breiman, L. Random forests. Mach. Learn. 45, 5–32 (2001).
Breiman, L., Friedman, J. H., Olshen, R. A. & Stone, C. J. Classification and Regression Trees (Chapman & Hall/CRC, 2017).
Freund, Y. & Schapire, R. E. A decision-theoretic generalization of on-line learning and an application to boosting. J. Comput. System Sci. 55, 119–139 (1997).
Pedregosa, F. et al. Scikit-learn: Machine learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Xgboost: Scalable, Portable and Distributed Gradient Boosting (GBDT, GBRT or GBM) Library, for Python, R, Java, Scala, C++ and More. Runs on Single Machine, Hadoop, Spark, Dask, Flink and DataFlow. (Github).
LightGBM: A Fast, Distributed, High Performance Gradient Boosting (GBT, GBDT, GBRT, GBM or MART) Framework Based on Decision Tree Algorithms, Used for Ranking, Classification and Many Other Machine Learning Tasks. (Github).
Harris, C. R. et al. Array programming with NumPy. Nature 585, 357–362 (2020).
McKinney, W. Data Structures for Statistical Computing in Python. in Proceedings of the 9th Python in Science Conference (SciPy, 2010). https://doi.org/10.25080/majora-92bf1922-00a.
Virtanen, P. et al. SciPy 1.0: fundamental algorithms for scientific computing in Python. Nat. Methods 17, 261–272 (2020).
Waskom, M. seaborn: Statistical data visualization. J. Open Sour. Softw. 6, 3021 (2021).
Hunter, J. D. Matplotlib: A 2D graphics environment. Comput. Sci. Eng. 9, 90–95 (2007).
Acknowledgements
This work was supported by the National Science Foundation grants DEB-2028280 and DEB-2109688 to AR and KAC and IOS-1754770 to THO.
Author information
Authors and Affiliations
Contributions
A.R. and K.A.C. conceived the method; M.B. implemented the software; M.B., B.S., and S.A.F. tested and packaged the software and evaluated the performance; M.B. and A.R. provided online documents and software. M.B., A.R., B.S., T.D., and T.H.O. analyzed application datasets. M.B., A.R., B.S., T.D., T.H.O., and K.A.C. drafted the manuscript. All authors discussed the results and commented on the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Baghbanzadeh, M., Dawson, T., Sayoldin, B. et al. deepBreaks identifies and prioritizes genotype–phenotype associations using machine learning. Sci Rep 15, 39095 (2025). https://doi.org/10.1038/s41598-025-25580-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-25580-6