Abstract
For giant pandas, the ecological impact of prophylactic postoperative antibiotics on their gut microbial communities and resistome is not well characterized. Here, we assessed the impact of intravenous cefotaxime administration by analyzing longitudinal fecal samples from five giant pandas via 16 S rRNA sequencing (n = 304 samples) and shotgun metagenomics (n = 22 samples). 16 S-based analysis revealed that antibiotic exposure significantly altered bacterial community structure, resulting in a pronounced increase in the abundance of Pseudomonadota (from 50% ± 24% to 60% ± 38%; P < 0.001) and a reduction in Shannon diversity (from 2.8 ± 0.4 to 2.4 ± 1.3; P < 0.05). In contrast, metagenomic analysis indicated that cefotaxime exposure did not significantly increase the overall diversity of antimicrobial resistance genes (ARGs) or virulence factor genes (VFGs). However, we observed a marked expansion in the diversity of the CTX-M β-lactamase family (blaCTX−M), which persisted into the recovery phase. We also recovered 10 metagenome-assembled genomes (MAGs) harboring both ARGs and VFGs, identifying them as potential antibiotic-resistant pathogens (ARPs). Their abundance, however, remained unchanged throughout treatment. These findings provide new insights into the effects of short-term antibiotic exposure in giant pandas, highlighting its transient effect on microbial community structure and a limited effect on resistome diversity.
Similar content being viewed by others
Introduction
The giant panda (Ailuropoda melanoleuca), a vulnerable and endemic species in China, has experienced gradual population recovery due to enhanced habitat protection and reduced human interference in nature reserves. Nonetheless, gastrointestinal diseases remain a major threat to survival and a leading cause of mortality in both wild and captive populations1,2,3. Common pathogenic and opportunistic bacteria implicated in these diseases include Escherichia coli, Klebsiella, Campylobacter jejuni, Pseudomonas aeruginosa, Enterococcus hirae, β-hemolytic streptococci, Yersinia enterocolitica, and Clostridium welchii4,5,6.
Antimicrobial therapy is widely used in captive giant pandas to treat gastrointestinal infections4,7. However, antibiotics indiscriminately target both pathogenic and commensal bacteria, disrupting gut microbial homeostasis and impairing host physiological functions including digestion, immunity, and metabolism in human studies8,9. Furthermore, antibiotic use accelerates the dissemination of antibiotic resistance genes (ARGs), facilitates the emergence of multidrug-resistant strains, and promotes the invasion of ARG-carrying pathogens (ARPs), thereby posing severe risks to host health and ecosystem safety10,11,12,13,14. Long-term antibiotic exposure may also lead to persistent enrichment of ARGs, which has been linked in humans to chronic diseases such as type 2 diabetes and cardiovascular disorders15,16,17.
Studies specific to giant pandas indicate that captive individuals exhibit enriched ARGs, virulence factor genes (VFGs), and heavy metal resistance genes compared to their wild counterparts18. Geographic variations in ARG profiles have also been observed, with the Qinling population showing higher abundances of Clostridium and vancomycin resistance genes19. Multidrug resistance is prevalent among gut bacteria in pandas: Zhang et al. (2009)20 reported widespread multidrug resistance in isolates from 30 individuals, and Yan et al. (2023)21 noted an increase in multi-class resistant E. coli between 2014 and 2020. Similarly, Fan et al. (2022)22 and Li et al. (2023)23 documented multidrug-resistant Klebsiella pneumoniae and Raoultella strains in captive pandas, highlighting serious health risks for both animals and their caretakers.
While existing research has well documented the presence of ARGs in the giant panda gut microbiome, the dynamic responses of the gut microbiota and resistome to short-term antibiotic exposure remain poorly understood. Elucidating these responses is critical for evaluating the health implications of antibiotic use in this vulnerable species. We hypothesize that short-term antibiotic challenge will induce significant—yet largely reversible—shifts in microbial community structure and resistome composition, with minimal long-term effects on overall diversity. To test this, we conducted a longitudinal study tracking temporal changes in the gut microbiome and resistome of giant pandas receiving cefotaxime prophylaxis during post-surgical recovery. This approach enables characterization of microbial resilience and ARG dynamics following antibiotic perturbation.
Methods
Sampling and fecal DNA extraction
Five giant pandas (Table S1) were included in this study and sampled from February 2018 to March 2020. Among them, three (Fufu, Kelin, Maotao) underwent surgical procedures and received intravenous injection of cefotaxime (10 mg/kg bodyweight, once every three days for 14 days) for postoperative pain management and wound healing, as well as to promote recovery and reduce surgical complications. The other two giant pandas (Shuxiang and Yingying) had comparable health status, age, and diet to the surgically treated individuals but did not undergo any surgical procedures. None of the five pandas had received antibiotic treatment for at least four months prior to sampling.
Fecal samples from the three surgically treated pandas were collected longitudinally over a period of approximately one month. Samples from the two non-surgical pandas were collected during the same period to serve as controls without antibiotic treatment. Additionally, fresh fecal samples from Kelin and Fufu were collected prior to surgery to establish baseline controls without antibiotic treatment. Within 15 min of defecation, a fresh portion of feces was collected using sterile gloves, placed in sterilized bags on dry ice, and promptly transported to the laboratory. Samples were stored at −80 ℃ until DNA extraction and microbial analyses. The entire sample collection process complied with the animal welfare guidelines of the Chengdu Research Base of Giant Panda Breeding (No. 201701).
Total DNA was extracted from the fecal samples using a bead-beating method following protocols described by Xue et al. (2015)24 and Zhang et al. (2018)25, which were adapted for bamboo-rich samples. The extracted DNA was then diluted to a concentration of 50 ng/µL using sterile water.
16 S rRNA gene sequencing
A total of 304 samples (Table S2) were used for 16 S rRNA gene sequencing and categorized into three groups: group B (n = 111) consisted of samples collected from individuals that had not received antibiotic treatment for at least four months prior to sampling, all taken at day 0; group C1 (n = 117) included samples obtained during the antibiotic treatment phase (days 1–14 after surgery), which involved intravenous administration of cefotaxime; and group C2 (n = 76) consisted of samples from the recovery phase, during which no antibiotics were administered (from day 15 up to approximately 30 days post-operation).
The V3–V4 hypervariable region of the bacterial 16 S rRNA gene was amplified using the universal primers 338 F (5′-barcode- ACTCCTACGGGAGGCAGCA − 3′) and 806 R (5′- GGACTACHVGGGTWTCTAAT − 3′)26, which provide high phylogenetic resolution and broad coverage of both Gram-positive and Gram-negative bacteria. Each sample was tagged with a unique 6-bp barcode. PCR reactions were performed in a 40 µl total volume using 2× High-Fidelity PCR Master Mix (MedChemExpress, China), with 15 pmol of each primer and 50 ng template DNA. The cycling conditions consisted of 35 cycles of denaturation at 94 °C for 45 s, annealing at 50 °C for 60 s, and extension at 72 °C for 90 s, followed by a final extension at 72 °C for 10 min. Amplified products were pooled in equimolar concentrations, verified by 2% agarose gel electrophoresis, and purified using the Qiagen Gel Extraction Kit (Qiagen, Germany). Sequencing libraries were constructed and sequenced on the Illumina MiSeq PE-250 platform following standard protocols provided by Novogene Biotech Co., Ltd. (Beijing, China).
Microbial raw reads of 16 S rRNA gene sequencing were processed in QIIME2 (version 2024.2)27. After demultiplexing and primer removal, the DADA2 plugin was employed for quality filtering, denoising, and generation of amplicon sequence variants (ASVs), producing a high-resolution feature table of ASV counts27. Taxonomic classification was performed using the SILVA reference database (version 138)28. Sequences classified as Eukaryota, chloroplasts, or mitochondria were removed. The resulting feature table and taxonomies of bacterial members were uploaded for Microbiome Analyst (https://www.microbiomeanalyst.ca)29 for Alpha- and Beta-diversity analyses after samples were rarefied at a threshold of minimum sequence frequency across all samples.
Quality control for shotgun metagenomic data
To enable a cost-efficient longitudinal analysis of antibiotic resistance gene (ARG) dynamics, we conducted shotgun metagenomic sequencing on 11 fecal samples from the individual Fufu. These samples were collected at time points spanning from pre-treatment to antibiotic exposure (Group B: n = 2; Group C1: n = 5; Group C2: n = 4). Additionally, 11 control samples (Group B) obtained at comparable time points from three individuals without antibiotic treatment (Kelin, Shuxiang, and Yingying) were included for cross-sectional comparison (Table S2). Sequencing libraries were constructed and sequenced on the Illumina NovaSeq 6000 platform with a 2 × 150 bp configuration following standard protocols provided by Novogene Biotech Co., Ltd. (Beijing, China).
Raw reads were processed with Trimmomatic30 to remove low-quality sequences, including (i) all read less than 50 bp in length, (ii) reads including N bases, and (iii) all PCR duplicates, resulting in clean data. To eliminate potential host-derived contamination, the clean data were aligned against reference genomes of the giant panda (GCA_002007445.3_ASM200744v3_genomic) and human genome (hg37dec_v0.1), using bowtie231, and matching sequences were removed.
Taxonomic assignments for shotgun metagenomic data
Clean reads were assembled de novo using Megahit (v1.1.3)32 with the default k-mer set ‘–k-list 21, 29, 39, 59, 79, 99, 119, 141′. Open Reading Frames (ORFs) were then predicted from the assembled contigs for each sample using MetaGeneMark (v.2.10, http://topaz.gatech.edu/GeneMark/)33. The predicted ORFs were processed to remove redundancy using CD-HIT software34, with < 90% overlap and < 95% identity, to generate a non-redundant initial gene catalog.
The clean reads from each sample were mapped to this non-redundant gene catalog using Salmon35 to estimate abundance in TPM (transcripts per million reads). Genes with read counts ≤ 2 across all samples were filtered out. Putative functional annotations (COG/KEGG36/CAZy) of the predicted genes were performed by aligning them against the corresponding databases using DIAMOND37. Finally, taxonomic abundance profiling for the metagenomic samples was conducted with the HUMAnN3 pipeline38.
Antimicrobial resistance genes and virulence gene analysis
To characterize the resistome of the giant pandas, clean metagenomic sequencing reads were aligned against the MEGARes 3.0 database (https://megares.meglab.org) using the AmrPlusPlus pipeline (version 3.0)39. The MEGARes database integrates ARG sequences from multiple public resources, including ResFinder40, ARG-ANNOT41, the NCBI Lahey Clinic beta-lactamase archive, the Comprehensive Antibiotic Resistance Database (CARD)42, NCBI’s Bacterial Antimicrobial Resistance Reference Gene Database43, and BacMet44. Briefly, after quality control, adapter trimming, and removal of host and food (bamboo) derived reads, high-quality clean reads were aligned to MEGARes 3.0 using BWA. The resulting alignments were used to generate a count matrix of matches to AMR gene accessions. Duplicate sequences were filtered out, and rarefaction analysis was performed for all samples. Read counts were normalized for gene length and sequencing depth by calculating reads per kilobase per million bacterial reads (RPKM) using STAMP45. The deduplicated AMR abundance matrix was further processed by excluding entries categorized as “metals” or “multi-compound”. The relative abundance of ARGs in each sample was determined by dividing the RPKM value of each gene by the total RPKM of all ARGs in that sample.
A similar protocol was applied to identify virulence factor genes (VFGs). Unassembled reads were queried against the Virulence Factor Database (VFDB set A, containing experimentally verified virulence factors; http://www.mgc.ac.cn/VFs/download.htm; accessed on October 9, 2024)46 using the same alignment and normalization procedures.
Co-occurrence network analysis of ARGs and microbial taxa
A previous study suggested that strong and significant co-occurrence patterns between ARGs and microbial taxa may help infer potential hosts of ARGs47. In this study, a robust statistical correlation was defined as a Spearman’s correlation coefficient with an absolute value ≥ 0.8 and a significance level of P ≤ 0.0148. These criteria were used to construct microbial co-occurrence networks. Network analysis was performed in Gephi (version 0.9.2) using Spearman’s correlation coefficient obtained from relative abundance data of ARGs and microbial taxa derived from 16 S rRNA sequencing of the 22 samples. Only significant pairwise correlations were included in the network, with nodes representing individual ARGs and edges representing significant associations between them.
Metagenomic assembly and Binning
The assembled contigs were binned using BASALT (v1.0.1)49 (Qiu et al., 2024) with default parameters to obtain non-redundant metagenome-assembled genomes (MAGs). Taxonomic classification of the MAGs was performed with GTDB-Tk (v1.7.0)50 based on the GTDB r202 database under default settings. Relative abundances of the MAGs were estimated using the quant_bin module in MetaWRAP (v1.2) with default parameters and normalized according to the method described by Zhu et al. (2023)51. ARGs and VFGs within these non-redundant MAGs were identified by conducting a DIAMOND search against the CARD and VFDB databases, respectively, following the same approach as detailed previously. Phylogenetic analysis of the non-redundant MAGs was conducted using GTDB-Tk based on 120 bacterial universal marker genes, and the resulting phylogeny was visualized with iTOL52 and the itol.toolkit R package53.
The raw sequencing data generated in this study have been deposited in the CNGB Sequence Archive (CNSA) of the China National GeneBank DataBase (CNGBdb) under accession number CNP0007979.
Statistical analysis and data visualization
Differential enrichment of bacterial taxa was assessed using a pairwise Mann-Whitney test while differences across timepoints were evaluated with Wilcoxon matched-pairs signed rank test. Multiple hypothesis testing was corrected by controlling the false discovery rate (FDR) using Benjamini-Hochberg (BH) method54. Group differences in beta diversity were tested using permutation based multivariate analysis of variance (PERMANOVA).
Compositional shifts were quantified and compared across individuals or sample pairs using Bray–Curtis and Jaccard distance metrics for shotgun metagenomic data. Principal coordinates analysis (PCoA) and significance testing were performed using the vegan package of R. Heatmaps were generated with the Pheatmap R package (v.1.0.12). Analysis of variance (ANOVA) followed by Tukey’s HSD test was used to examine significant differences in the abundance of ARGs and VFGs between groups. P-values ≤ 0.05 were considered statistically significant and were further adjusted for multiple comparisons using the BH method where applicable. Data visualization was conducted using the online tool ImageGP (http://www.ehbio.com/ImageGP/)55.
Random forest analysis56 was employed to identify significant bacterial taxa, ARGs, and VFGs discriminating between groups, using the “randomForest” R package with 1,000 trees and default parameters.
All experimental procedures were conducted in accordance with relevant guidelines and regulations, including the Animal Research: Reporting of In Vivo Experiments (ARRIVE) guidelines. The study protocol was reviewed and approved by the Scientific Research Ethics Committee of Mianyang Normal University (Project License: MSL202402).
Bacterial nomenclature follows the International Code of Nomenclature of Prokaryotes (ICNP). Notably, the phyla formerly known as ‘Firmicutes’ and ‘Bacteroidetes’ are hereafter referred to as ‘Bacillota’ and ‘Bacteroidota’, respectively, in alignment with current taxonomic standards and consistently throughout this manuscript.
Results
Baseline characterization of the gut bacterial community
A total of 304 samples across all phases were used to analyze bacterial communities based on 16 S rRNA gene sequencing data (Table S2). Following paired-end read assembly and quality control, a total of 16,389,523 non-chimeric high-quality reads were retained, with read counts per sample ranging from 16,844 to 75,495 per sample (mean: 53912) (Table S2). These reads were grouped into 4,907 ASVs. A species accumulation boxplot (Figure S1) indicated that sufficient sampling depth was achieved to capture the majority of bacterial diversity present.
The bacterial compositions of the 304 samples were dominated by three main phyla: Pseudomonadota, Bacillota, and Campilobacterota (Figure S2). At the phylum level, Pseudomonadota (mean relative abundance = 53.18%) was significantly higher than Bacillota (46.34%) (one way ANOVA, P < 0.001). A total of 7 families exhibited a relative abundance exceeding 1%, among which Enterobacteriaceae was the most abundant (51.23%) (Figure S3). At the genus level, 11 taxa each accounted for more than 1% of the total sequences across all samples. The top four most abundant genera were Escherichia-Shigella (45.53%), Lactococcus (12.72%), Clostridium_sensu_stricto_1 (12.38%), and Streptococcus (11.86%) (Figure S4). In addition, about 6.82% of the total sequences could not be classified at the genus level.
Response of microbial diversity and composition to antibiotic treatment
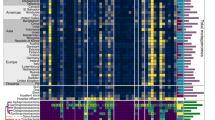
Across all samples in group B (n = 111), the dominant bacterial taxa (Fig. 1A), as well as the alpha (Fig. 1B) and beta (Fig. 1C) diversity, remained relatively stable throughout the two-year study period. Substantial shifts in microbial composition occurred following antibiotic exposure during the treatment phase (C1). Specifically, this phase was characterized by a significantly higher relative abundance of Pseudomonadota and a significantly lower relative abundance of Bacillota compared to both group B and the recovery phase (C2) (P < 0.001; Fig. 1A, Figures S5–S6). Antibiotic treatment markedly altered the bacterial community structure, with the first day of treatment (C1) showing the most pronounced changes: Pseudomonadota reached its highest relative abundance (92.63%), while Bacillota reached its lowest (7.35%). By day 2, the abundance of Pseudomonadota decreased to 90.34%, and further declined to 73.48% by day 3. It then stabilized at approximately 56% for the remainder of the treatment period (Fig. 1A). No significant differences in the relative abundances of Pseudomonadota and Bacillota were observed between group B and phase C2 (Figures S5–S6). At the individual level, all three animals exhibited a consistent decline in Pseudomonadota from phase C1 to C2 in (P < 0.001), reflecting the trend observed at the group level (Figure S5).
The bacterial diversity in this study based on 16 S rRNA gene sequencing. (A) the relative abundance of samples at phylum level. Samples on the x-axis are arranged in chronological order within each treatment group. (B) the Shannon index of samples. (C) unweighted principal coordinates analysis. (D) the 15 most important genera with random forest (RF) analysis. The mean decrease accuracy shows the ability of microbes to decrease node impurity in the RF model and a large value means that the genus is very important to determinize microbiota structure. The color bar to the right represents the relative abundance of each genus. B: the samples without antibiotic-treatment (day 0), C1: the samples were treated by intravenous injection of cefotaxime (day 1–14) and C2: the samples were for recovery (day 15–30).
Alpha diversity decreased significantly following antibiotic treatment: Group B showed significantly higher Shannon indices (T-test, P < 0.05; Fig. 1B and Figure S7; Table S2) and greater number of observed ASVs (T-test, P = 2.175E-5; Figure S7) than both C1 and C2. No significant differences in Shannon indices or observed ASVs were detected between C1 and C2 (Fig. 1B and Figure S7). Consistent with phylum-level changes, alpha diversity was lowest on the first day of treatment (mean Shannon index = 0.8), gradually increasing to 0.9 by day 2, 1.9 by day 3, and stabilizing around 2.7 thereafter (Fig. 1B). Individual variation was also evident: both Shannon and observed species indices increased significantly from C1 to C2 in Kelin; Shannon index increased significantly in Maotao; whereas no significant changes were observed in Fufu (Figure S7).
Principal coordinate analysis (PCoA) based on unweighted UniFrac analysis (sensitive to rarer taxa) revealed clear separation between B and C1, with 19.4% variance explained ([PERMANOVA] F-value: 21.573; R-squared: 0.12537; p-value: 0.001). In contrast, weighted UniFrac analysis (sensitive to abundances of taxa) showed weaker separation and greater overlap between B and C1 (Fig. 1C; Figure S8). Both weighted and unweighted UniFrac analysis indicated that group C2 could not be clearly distinguished from B or C1, a finding consistent with Random Forest classification, which yielded class error rates of 43.5% for C2, 1.4% for C1, and 0 for B. Among the top 15 genera identified by random forest as discriminative of gut microbiota structure, eight were overrepresented in group B: Clostridium_sensu_stricto_1, Turicibacter, Campylobacter, Lactobacillus, Anaerosporobacter, Cellulosilyticum, Helicobacter, and Epulopiscium (Fig. 1D). Three genera were overrepresented in C1, and four in C2 (Fig. 1D).
Differential abundance analysis using EdgerSuper2 in EasyMicrobiomeR57 identified several significantly altered ASVs. The largest differences involved members of Clostridium_sensu_stricto_1, including: ASV_3226 (enriched in B vs. C1 and C2), ASV_3458 (enriched in B vs. C1, depleted in C2 vs. C1), ASV_3510 (depleted in C2 vs. C1), and ASV_3293 (enriched in B vs. C1) (Fig. 2). Escherichia-Shigella was depleted in C2 compared to B (ASV_2387) and C1 (ASV_2402) (Fig. 2). Streptococcus was enriched in B compared to C1 (ASV_0911) and depleted in C2 compared to C1 (ASV_0911 and ASV_0912) (Fig. 2). Additionally, Lactobacillus was enriched in B compared to C1 (ASV_0348) and C2 (ASV_0349), while Rummeliibacillus (ASV_0281) and Pediococcus (ASV_0303) were also enriched in B compared to C2 (Fig. 2). These findings are consistent with the random forest results (Fig. 1D).
Multi-group difference analysis volcano plot. The figure displays only the top five ASVs between groups, as determined by EdgerSuper2.
PCoA based on Bray–Curtis distances revealed significant clustering by diet, which explained 44.7% of the variance between shoot and stem (PERMANOVA: F = 7.8462, R² = 0.051, P < 0.001) and between shoot and leaf (F = 103.09, R² = 0.258, P < 0.001) (Figure S9). Individual variation also significantly influenced microbial composition (F = 24.651, R² = 0.248, P < 0.001), although to a lesser extent than diet. Pairwise comparisons revealed the largest dissimilarity between Maotao and Shuxiang (R² = 0.333) and the smallest between Shuxiang and Yingying (R² = 0.051) (Figure S9).
Functional prediction of shotgun metagenomics
We next examined the effects of intravenous cefotaxime (a β-lactam antibiotic) on the composition of bacterial ARGs and VFGs over a one-month time-series experiment. The study included a pre-exposure phase (B; n = 13), a 14-day exposure phase (C1; n = 5), and a recovery phase spanning days 14 to about 30 (C2; n = 4). The use of cefotaxime directly accounted for the high abundance of β-lactam resistance genes observed in subsequent analyses. Sequencing yielded between 33 and 48 million paired-end reads per sample. Duplicate reads counted for 27% to 64% of the total (Figure S10), and overrepresented sequences constituted less than 1%. After removing low-quality reads and contaminants (including food and host DNA, representing 0.03–0.11% of all reads), a total of 599 Gbp of high-quality clean reads from the 22 samples were retained (Figure S10). De novo assembly produced 6.12 × 10⁶ contigs with an N50 of 1,676 bp.
A non-redundant microbial gene catalog was constructed, containing 3.9 × 10⁶ genes with an average length of 352.8 bp. Of these, 36.31% contained complete open reading frames (ORFs). Saturation curves indicated that the gene catalog achieved high coverage with the current sequencing depth (Figure S11). Functional annotation revealed that nearly half of the genes (1.49 × 10⁶) had significant matches in the COG database, suggesting that a substantial portion of the giant panda gut microbiome remains functionally uncharacterized. Among the annotated COG categories, 43.5% were associated with metabolic processes, particularly amino acid transport and metabolism, carbohydrate transport and metabolism, inorganic ion transport and metabolism, and energy production and conversion (Table S3). Additionally, 21.0% of genes were assigned to unknown functions (Table S3). KEGG annotation indicated that the most abundant functional categories included “Protein families: signaling and cellular processes” (16.9%), “Protein families: genetic information processing” (12.3%), and “carbohydrate metabolism” (Table S4). Principal coordinate analysis (PCoA) showed that cefotaxime administration did not significantly affect the functional profiles based on COG (Figure S12) or KEGG (Figure S13) annotations.
Taxonomic profiling using HUMAnN338 revealed that Pseudomonadota (mean ± SD: 84.69% ± 6.26%) and Bacillota (15.30% ± 6.26%) were the dominant phyla, together accounting for 99.99% of the total abundance. At the genus level, Escherichia (81.26% ± 6.67%) was most abundant, followed by Lactococcus (6.95% ± 2.25%) and Leuconostoc (6.05% ± 1.03%). A total of 79 genera were identified, with three—Escherichia, Leuconostoc, and Weissella—present in over 90% of the individuals.
Dynamics of ARGs across treatment phases
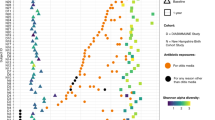
We next characterized the types and abundance of ARGs across the 22 samples from giant pandas using a read-based approach with AMRPlusPlus39. On average, 0.2% of the total reads were classified as ARGs, with per-sample values ranging from 0.08% to 0.4%. A total of 1,091 ARGs were identified, distributed across 20 antibiotic classes. Among these, beta-lactam resistance genes were the most abundant (32.48%), followed by MLS (macrolides–lincosamides–streptogramins; 12.6%), multi-drug resistance (12.1%), tetracyclines (9.3%), cationic antimicrobial peptides (8.5%), fluoroquinolones (6.4%), and aminoglycosides (6.2%); each of these classes accounted for more than 5% of the total ARG abundance (Fig. 3A). Antibiotic treatment significantly increased the relative abundance of MLS genes: group C1 showed a significantly higher abundance (16.0% ± 0.98%) compared to group B (10.7% ± 0.15%) (Fisher’s exact test, P < 0.01; Fig. 3A).
Comparisons of ARGs in the guts of giant pandas for three groups based on metagenome shotgun sequencing. (A) the relative abundance of ARGs for every sample at class level. The observed ARGs (B) and Simpson index (C) of antibiotic resistance genes across the three phases. The two indexes are presented with the median values indicated (central black horizontal lines); the 25th and 75th percentiles are indicated (boxes), and the whiskers extend from each end of the box to the most extreme values within 1.5 times the interquartile range from the respective end. The letter (a) for each group showed no significant difference. PCoA based on (D) Bray–Curtis distance (adonis R2: 0.15, adonis P − value: 0.0805) and (E) Jaccard distances (adonis R2: 0.15, adonis P − value: 0.043) showing the changes in ARG compositions in the three groups.
Alpha diversity of the resistome was assessed using the observed number of ARGs and the Simpson diversity index. Although antibiotic treatment increased the observed ARG count and decreased Simpson diversity between groups B and C1, these differences were not statistically significant (P > 0.05; Fig. 3B–C).
To evaluate the structural shifts in the resistome following antibiotic treatment, we examined beta diversity based on normalized ARG counts using both Bray–Curtis and Jaccard dissimilarity indices. Principal coordinate analysis revealed clear separation between groups B and C2 (Fig. 3D–E). PERMANOVA confirmed that resistome composition differed significantly between groups B and C2 (Jaccard: R² = 0.14, P < 0.05; Bray–Curtis: R² = 0.16, P < 0.05). In contrast, differences between groups B and C1 were not significant (Jaccard: R² = 0.09, P = 0.09; Bray–Curtis: R² = 0.08, P = 0.18). These results indicate that antibiotic treatment led to significant alterations in resistome composition by the recovery phase (C2), although considerable intra-group variation was observed. Random forest analysis did not identify any specific ARG subtype that reliably discriminated among the three groups.
Expansion of bla CTX−M gene following cefotaxime exposure
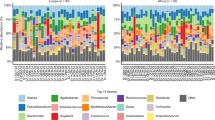
Of the ARGs identified in the gut microbiome of giant pandas, blaCTX−M exhibited the greatest diversity at the subtype level, with a total of 188 distinct genes accounting for 8.8% of the total ARG abundance. Among these, only 3 blaCTX−M genes (MEG_2435, MEG_2378, MEG_2430) were prevalent across all individuals, each representing more than 1% relative abundance (Fig. 4).
The heatmap of 188 blaCTX−M genes in the 22 samples. Here, the legend shows the relative abundance in total of ARGs. Each column represents a unique blaCTX−M gene accession from AMR + + v3.0, and each row corresponds to one sample. The heatmap depicts normalized abundance values after strict alignment filtering (≥ 80% coverage and identity).
Cefotaxime treatment resulted in two clearly separated clusters based on blaCTX−M composition. A greater number of blaCTX−M subtypes were detected in samples from the antibiotic-treatment groups (C1 and C2) compared to the pretreatment and untreated group (B). Specifically, 171 blaCTX−M genes were absent in group B, while 9 were present in more than 75% of samples and 37 were detected in over half of the samples for group C1 and C2 – none of which were observed in group B (Fig. 4). However, no significant difference in the relative abundance of blaCTX−M was observed between group C1 and C2 (P > 0.05; Fig. 4).
Overall, these findings indicate that cefotaxime exposure led to a diversification of the blaCTX−M gene composition, and that this effect persisted long after the cessation of antibiotic treatment.
Shifts in VFGs during and after antibiotic treatment
A total of 1690 VFGs were identified from the reads, representing 12 functional types in the giant pandas (Figure S13, Table S5). Among these, the most abundant VFG types included those associated with nutritional/metabolic factor (VFC0272) (34.3%–74.6%), adherence (VFC0001) (9.2%–32.7%), effector delivery system (VFC0086) (4.0%–20.5%), invasion (VFC0083) (5.0%–18.6%), motility (VFC0204) (0.3%−3.9%) and immune modulation (VFC0258) (0.07%–2.4%) were found to be more abundant VFG types (Figure S14).
Alpha diversity analysis of VFGs—assessed using the observed number of genes, Shannon index, and inverse Simpson index—revealed no significant effect of antibiotic treatment, with no notable differences among the three groups (B, C1, and C2; P > 0.05; Figure S15). Similarly, beta diversity analysis based on Bray–Curtis and Jaccard distances of normalized VFG counts showed no significant separation among the groups in PCoA (Figure S16).
At the bacterial species level, 26 species were associated with VFGs, with Escherichia coli containing the highest number (265 genes) (Table S5). These VFGs were distributed across six functional categories, among which effector delivery system (VFC0086) were the most represented (124 genes). No significant differences were observed in the relative abundances of VFGs among the three groups (Table S5). Random forest analysis did not identify any VFG subtype capable of distinguishing between the groups.
Co-occurrence networks between ARGs and bacterial taxa
Network analysis was conducted to examine co-occurrence patterns between ARG subtypes and bacterial taxa obtained from 16 S rRNA sequencing data. The resulting network contained 89 nodes and 159 edges, with a high modularity index of 0.90. Among these, Streptococcus and Acinetobacter exhibited the highest number of edges (more than 10 linkages each; Fig. 5). However, these correlations indicate potential ecological co-occurrence and do not necessarily imply that these taxa are direct hosts of the ARGs.
Network analysis for the co-occurrence patterns between ARG subtypes and bacterial taxa at the genus level. Only strong (Spearman’s correlation coefficient > 0.8) and significant (P < 0.01) correlations are shown. Nodes represent ARGs and bacterial taxa, the size of each node is proportional to its number of connections. The edge weight represents the absolute Spearman’s correlation coefficient, which ranges from 0.8 to 1.0. A thicker edge indicates a stronger correlation. The color legend showed the nodes represent genes to indicate antimicrobial classes of ARGs.
Beta-lactam resistance genes were the most abundant class, followed by multidrug resistance genes. At the genus level, 24 genera were associated with beta-lactam resistance genes and 26 genera were linked to multidrug resistance genes. The most frequently detected ARG subtypes included POXT, TUFAB, CMY, and FEXB. Three genera—Agromyces, Rhodococcus, and Variovorax—were correlated with blaCTX−M resistance genes (Fig. 5). It is important to note that these correlations may arise from shared environmental constraints or community-level interactions and do not constitute definitive evidence of ARG carriage.
MAGs reveal potential antibiotic-resistant pathogens
After binning, the quality of metagenome-assembled genomes (MAGs) was assessed using CheckM58. MAGs with completeness greater than 50% and contamination less than 10% were retained, resulting in 46 non-redundant MAGs for downstream analyses. Among these, 16 MAGs had completeness > 90% and contamination < 5%, and three MAGs exhibited completeness > 97% with no detectable contamination (Table S6). Taxonomic classification against the GTDB database (release 214) assigned the 46 MAGs to 5 phyla, 6 classes, 10 orders, 15 families, 27 genera, and 25 species (Fig. 6 and Table S6). All MAGs were classified at least to the family level, 45 (97.8%) to the genus level, and 31 (67.4%) to the species level (Table S6).
the information of 46 non-redundant MAGs with completeness > 50% and contamination < 10%. Phylogenomic tree was shown in the inner circle and the color indicated the MAG belonging to bacteria taxa at family level. The middle circle shows the relative abundance in every sample. The bar shows ARG and VFG numbers for every MAG and the details can be found in Table S6. The figure was visualized using the toolkit package v.1.1.953 in R and the online tool iTOL52.
To evaluate the distribution of ARGs within the fecal metagenome of giant pandas, we annotated ARGs in each MAG. A total of 55 ARGs were identified across the 46 MAGs. The most prevalent ARG was vanY, detected in 19 MAGs, followed by vanT, arnT, and vanG, all of which are associated with antibiotic target alteration (Table S6).
At the family level, ARGs were most abundant in MAGs assigned to Enterobacteriaceae. The most frequently observed ARG subtypes within this family were arnT (88.9%; 8 out of 9 MAGs), kpnH (77.8%), kpnF (77.8%), and vanG (55.6%) (Fig. 6 and Table S6). Similarly, all MAGs classified as Enterococcaceae carried at least two ARGs, with vanY (80%) and AAC(6′)-Iid (40%) being the most prevalent.
At the species level, Escherichia coli (sur-bin-20) contained the highest number of ARGs (30), followed by Buttiauxella sp002321695 (sur-bin-38) with 14 ARGs and Cedecea davisae (sur-bin-37) with 13 ARGs. These species showed intermediate relative abundance among the 46 MAGs (Fig. 6 and Table S6). In contrast, Basfia_A sp014843355 (sur-bin-39), which had the highest relative abundance, carried only one ARG (Fig. 6 and Table S6). No significant differences in the relative abundance of these 46 MAGs were observed among groups B, C1, and C2 following antibiotic exposure (Fig. 6 and Table S6).
To identify potential ARG-carrying pathogens (ARPs) that may pose health risks to giant pandas, we performed functional annotation on MAGs that contained both ARGs and VFGs. Among the 46 MAGs carrying VFGs, a total of 78 VFGs were identified. Notably, sur-bin-22 (classified as Pseudomonas helleri) and sur-bin-1 (classified as Aeromonas caviae) each carried 34 VFGs. Based on the co-occurrence of ARGs and VFGs, 10 MAGs were predicted as potential ARPs. These 10 MAGs span 3 phyla, 3 classes, 4 orders, 5 families, and 8 genera, with 7 assigned to known species (Table S6). Most of these putative ARPs belonged to the Enterobacteriaceae family. However, it should be noted that the identification of these MAGs as ARPs is based on in silico annotation and does not imply confirmed pathogenic function or host infection capability. Furthermore, antibiotic exposure did not significantly alter the relative abundance of these 10 putative ARPs across groups B, C1, and C2 (Fig. 6 and Table S6).
Discussion
In our longitudinal experiment, we advance the ecological framework for understanding the dynamics of host-associated microbiomes by monitoring bacterial recovery in giant pandas that received antibiotic-treatment during post-surgical recovery. Our results demonstrate that a short-course cefotaxime intervention induced a transient yet significant increase in the dissimilarity of bacterial community composition between baseline and antibiotic-treated samples (Figs. 1 and 2). Furthermore, our work reveals that cefotaxime exposure had minimal to non-significant effects on resistome diversity (Fig. 3). However, the treatment led to marked increase in the abundance of blaCTX−M genes in the antibiotic-treated group compared to the untreated controls—an effect that persisted well into the recovery phase (Fig. 4).
In this study, giant pandas treated with antibiotics exhibited a sharp decline in the alpha diversity of their gut bacterial communities, followed by a period of adaptation leading to a stabilized state (Fig. 1, S5-S7). A diverse gut microbiota is widely recognized as crucial for animal health59, and reductions in microbial diversity can reduce the abundance of beneficial taxa while facilitating the proliferation of potential pathogens60,61. Consistent with this, our analyses revealed a pronounced decline in beneficial microbes such as Lactobacillus—which plays a key role in maintaining gut homeostasis and inhibiting pathogens62—as well as hydrolytic and fermentative genera including Clostridium_sensu_stricto_1 and Cellulosilyticum, which are critical for fiber digestion and short-chain fatty acid production. Concurrently, we observed an increase in Escherichia/Shigella, a group that encompasses opportunistic pathogens known to be associated with intestinal inflammation63. These shifts -- consistent across PERMANOVA, Wilcoxon, and Random Forest analyses (Figs. 1 and 2) -- suggest that cefotaxime treatment may compromise gut functional integrity and promote the expansion of potentially pathogenic bacteria, even during the recovery phase (C2).
Previous studies have indicated that Enterobacteracea, particularly Escherichia, dominate the gut microbiome of giant panda25,64,65, which is consistent with our detection of Escherichia as a predominant genus. In support of this, a MAG (sur-bin-20) was annotated with Escherichia coli and found to carry 30 ARGs (Table S6), aligning with earlier reports of multidrug-resistant E. coli and a high prevalence of ARGs in Escherichia within giant pandas4,7,66. Notably, Yang et al. (2024)67 identified Escherichia as a major ARG carrier in this host species, while Yan et al. (2023)68 reported that dominant ARGs were primarily derived from Pseudomonadota. In contrast, our findings indicate that Streptococcus and Acinetobacter were identified as the primary potential carriers of ARGs (Fig. 5), a conclusion further supported by MAG analysis. Among the 46 MAGs meeting quality thresholds (completeness > 50%, contamination < 10%), seven were classified as Streptococcus, exhibited high abundance, and were found to harbor ARGs (Fig. 6; Table S6). These results are consistent with those reported by Mustafa et al. (2021)69, who also observed a positive correlation between Streptococcus and prevalent ARGs in the giant panda gut microbiome. These discrepancies may stem from variations in antibiotic type and treatment duration across studies. It is well established that the class of antibiotic administered profoundly shapes the selection and emergence of specific ARGs within the gut microbiota70,71,72,73. Likewise, treatment duration significantly modulates microbial community structure and resistance dynamics74,75,76. Therefore, the distinct antibiotic regime used in our study may account for the observed prominence of Acinetobacter and Streptococcus in harboring ARGs.
VFGs may facilitate the colonization and infection of hosts by potentially enhancing bacterial adhesion, motility, growth, and survival, thereby aiding putative pathogens in evading or modulating host defenses77,78,79. Of particular concern are bacteria co-harboring multidrug resistance genes and virulence factors, as the convergence of these traits may amplify pathogenic potential and the capacity to cause infection80. Thus, microorganisms carrying both ARGs and VFGs could pose a health risk to giant pandas.
In this study, we identified 10 MAGs carrying both ARGs and VFGs (Fig. 6 and Table S6). It should be noted that the classification of these MAGs as potential ARPs is based on genomic annotations and does not confirm functional pathogenicity or infectivity in giant pandas. Among these, Aeromonas caviae (sur-bin-1), which has been associated with clinical symptoms including diarrhea and soft tissue infections in humans81, was annotated with 34 VFG subtypes across 6 functional categories and 2 ARG subtypes. Enterococcus hirae (sur-bin-13 and sur-bin-25), linked to human infections such as bacteremia and urinary tract infections82, carried 2 ARG subtypes and 1 VFG subtype. Additionally, Morganella morganii (sur-bin-44), an emerging public health concern implicated in bloodstream infections83,84, was annotated with 2 VFG subtypes and 3 ARG subtypes. These MAGs exhibited low to medium relative abundance across all samples, with no significant differences observed among treatment groups (Fig. 6 and Table S6), suggesting that the antibiotic intervention in this study did not markedly alter their abundance. Importantly, although these taxa include known pathogens in humans and other animals, their potential pathogenicity in giant pandas remains to be empirically validated.
In order to control these ARPs, long-term sampling and monitoring of these pathogenic bacteria using metagenomics‐based sequencing technologies are essential for antimicrobial resistance analysis. Furthermore, to limit the dissemination of ARPs in giant pandas, alternative strategies should be developed. These may include antibiotic combinations therapies, bacteriophage therapy, antibiotic adjuvants, probiotics, monoclonal antibodies, and vaccines85,86,87. There is increasing interest in modulating intestinal microbiota resistance through probiotic supplementation88. Thus, identifying functional probiotics specific to giant pandas, as explored by Yu et al. (2024)89 and Zhang et al. (2024)65, is of considerable importance. Additionally, compared with oral administration, injection of antibiotics may reduce the amplification and emergence of gut ARGs under equivalent conditions90. In this study, intravenous cefotaxime administration likely contributed to the limited shifts in ARGs and VFGs profiles across the three groups (Fig. 3 and S14-S15). This suggests that when antibiotic therapy is required, intravenous injection may represent a preferable route to minimize impact on the gut resistome and virulence factor pool, thereby supporting host health without substantially promoting the expansion of resistance or virulence genes.
Moreover, we observed that multidrug resistance represents one of the predominant types of ARGs in the giant panda gut microbiome (Fig. 3A). This finding aligns with previous reports indicating that multidrug resistance genes are highly abundant in both wild and captive giant pandas18,19,64,69,91. Multidrug resistance, which confers tolerance to multiple antibiotic classes, complicates clinical treatment and poses a potential threat to giant panda health. Therefore, increased attention should be directed toward this category of ARGs to develop effective strategies that limit their dissemination and mitigate the risks associated with antimicrobial resistance.
Study limitations and strengths
This study provides new insights into the effects of short-term antibiotic exposure in giant pandas, highlighting its transient influence on microbial community structure and a limited influence on resistome diversity. Nevertheless, several limitations should be acknowledged. First, the extremely small sample size—only three pandas received antibiotic treatment, with two serving as controls—considerably limits the statistical power and generalizability of the findings. Moreover, as an endangered species, giant pandas are challenging subjects for systematic intervention-based research. Only one individual (Fufu) provided longitudinal shotgun metagenomic data across all three time points (B, C1, C2). Therefore, the dynamics of ARGs and VFGs reported here should be interpreted with caution. Second, potential confounders including diet and inter-individual variation may have shaped the microbial diversity and composition in giant pandas (Figure S9)24,25. These observations align with human studies reporting considerable interpersonal variation in gut microbiota responses to antibiotic exposure92. Such factors could also modulate the response to antibiotic intervention. In addition, the dramatic microbial shift observed on day 1 may be partly attributable to physiological stress associated with the surgical procedure itself, complicating the interpretation of early time-point data. To address these limitations, future studies should aim to include larger sample sizes and more comprehensive temporal sampling to improve the robustness and generalizability of functional insights. Higher-resolution tracking of MAGs aligned to individual hosts is also recommended. Furthermore, the origins and mechanisms driving the persistence or decline of specific ARGs remain unclear. Thus, wet-lab validations—such as quantitative PCR (qPCR) of key ARGs (e.g., blaCTX‑M variants) to confirm temporal and individual dynamics, culture-based antibiotic susceptibility testing of dominant genera (e.g., Acinetobacter and Streptococcus) to link genotypes to phenotypes, and plasmid conjugation assays to evaluate ARG transferability—would greatly enhance the mechanistic understanding of resistome dynamics.
Conclusions
Our results demonstrate that prophylactic post-surgical intravenous cefotaxime administration significantly disrupted the short-term structure of the gut bacterial community in giant pandas, although it did not substantially affect the overall diversity of the resistome or VFGs. Nevertheless, the limited sample size of the treatment group constrains the generalizability of our findings. Despite this limitation, this study provides valuable baseline information for the management and conservation of captive giant panda populations. Further research incorporating longitudinal sampling and deeper metagenomic sequencing is warranted to better evaluate antibiotic efficacy and monitor subsequent health outcomes, including infection susceptibility, in this species.
Data availability
We have shared the link to our data in our manuscript.
References
Qiu, X. & Mainka, S. A. Review of mortality of the giant panda (Ailuropoda melanoleuca). J. Zoo Wildl. Med. 24, 425–429 (1993).
Edwards, M. S., Zhang, G., Wei, R. & Liu, X. Nutrition and dietary husbandry. In Giant Pandas: Biology, Veterinary Medicine and Management (eds Zhang, A. et al.) 101–158 (Cambridge University Press, 2006).
Zhao, S., Li, C., Zhu, T., Jin, L. & Zou, L. Diversity and composition of gut bacterial community in giant panda with anorexia. Curr. Microbiol. 78, 1358–1366 (2021).
Zou, W. et al. Frequency of antimicrobial resistance and integron gene cassettes in Escherichia coli isolated from giant pandas (Ailuropoda melanoleuca) in China. Microb. Pathog. 116, 173–179 (2018).
Chen, D. et al. Serotype and antimicrobial resistance of Escherichia coli isolated from feces of wild giant pandas (Ailuropoda melanoleuca) in Sichuan Province, China. J. Wildl. Dis. 54, 691–699 (2018).
Gan, B. et al. Dynamic monitoring of changes in fecal flora of giant pandas in mice: Co-Occurrence network reconstruction. Microbiol. Spectr. 11 (2), e0199122 (2023).
Guo, L. et al. Antimicrobial and disinfectant resistance of Escherichia coli isolated from giant pandas. J. Appl. Microbiol. 119, 55–64 (2015).
Maurice, C. F., Haiser, H. J. & Turnbaugh, P. J. Xenobiotics shape the physiology and gene expression of the active human gut Microbiome. Cell 152, 39–50 (2013).
Sommer, F. & Backhed, F. The gut microbiota — masters of host development and physiology. Nat. Rev. Microbiol. 11, 227–238 (2013).
Ventola, C. L. The antibiotic resistance crisis: part 1: causes and threats. Pharm. Ther. 40, 277 (2015).
Francino, M. P. Antibiotics and the human gut microbiome: dysbioses and accumulation of resistances. Front. Microbiol. 6, 1–11 (2016).
Langdon, A., Crook, N. & Dantas, G. The effects of antibiotics on the Microbiome throughout development and alternative approaches for therapeutic modulation. Genome Med. 8, 39 (2016).
Casals-Pascual, C., Vergara, A. & Vila, J. Intestinal microbiota and antibiotic resistance: perspectives and solutions. Hum. Microbiome J. 9, 11–15 (2018).
Abd El-Aziz, N. K. et al. Extensive drug-resistant Salmonella enterica isolated from poultry and humans: prevalence and molecular determinants behind the co-resistance to Ciprofloxacin and Tigecycline. Front. Microbiol. 12, 738784 (2021).
Yang, Z. et al. Preliminary analysis showed country-specific gut resistome based on 1,267 feces samples. Gene 581(2), 178–82 (2016).
Gibson, M. K., Crofts, T. S. & Dantas, G. Antibiotics and the developing infant gut microbiota and resistome. Curr. Opin. Microbiol. 27, 51–56 (2015).
Shuai, M. et al. Human gut antibiotic resistome and progression of diabetes. Adv. Sci. (Weinh). 9 (11), e2104965 (2022).
Guo, W. et al. Comparative study of gut microbiota in wild and captive giant pandas (Ailuropoda melanoleuca). Genes 10, 827 (2019).
Hu, T. et al. Geographic pattern of antibiotic resistance genes in the metagenomes of the giant panda. Microb. Biotechnol. 14 (1), 186–197 (2021).
Zhang, A. Y. et al. Phenotypic and genotypic characterisation of antimicrobial resistance in faecal bacteria from 30 giant pandas. Int. J. Antimicrob. Agents. 33, 456–460 (2009).
Yan, X. et al. Research progress in the antibiotic resistance of giant panda-derived bacteria. Acta Microbiol. Sinica. 63 (3), 977–992 (2023). (In Chinese).
Fan, S. et al. Antibiotic-resistant Escherichia coli strains isolated from captive giant pandas: a reservoir of antibiotic resistance genes and virulence-associated genes. Vet. Sci. 9 (12), 705 (2022).
Li, Y. et al. Prevalence of antimicrobial resistance and virulence genes in Klebsiella pneumoniae and Congenetic Raoultella isolates from captive giant pandas. PLoS One 18(3), e0283738 (2023).
Xue, Z. et al. The bamboo-eating giant panda harbors a carnivore-like gut microbiota, with excessive seasonal variations. mBio 6, e00022–e00015 (2015).
Zhang, W. et al. Age associated Microbiome shows the giant panda lives on hemicelluloses, not on cellulose. ISME J. 12, 1319–1328 (2018).
Caporaso, J. G. et al. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. PNAS 108, 4516–4522 (2011).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible Microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857 (2019).
Pruesse, E. et al. SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res. 35, 7188–7196 (2007).
Dhariwal, A. et al. MicrobiomeAnalyst - a web-based tool for comprehensive statistical, visual and meta-analysis of Microbiome data. Nucleic Acids Res. 45, W180–188 (2017).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with bowtie 2. Nat. Methods. 9, 357–359 (2012).
Li, D., Liu, C. M., Luo, R., Sadakane, K. & Lam, T. W. MEGAHIT: an ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph. Bioinformatics 31, 1674–1676 (2015).
Mende, D. R. et al. Assessment of metagenomic assembly using simulated next generation sequencing data. PLoS One. 7, e31386 (2012).
Fu, L., Niu, B., Zhu, Z., Wu, S. & Li, W. CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28, 3150–3152 (2012).
Patro, R., Duggal, G. & Kingsford, C. Salmon: accurate, versatile and ultrafast quantification from RNA-seq data using lightweight-alignment. Biorxiv 021592 (2015).
Kanehisa, M., Furumichi, M., Sato, Y., Matsuura, Y. & Ishiguro-Watanabe M. KEGG: biological systems database as a model of the real world. Nucleic Acids Res. 53, D672–D677 (2025).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nat. Methods. 12, 59–60 (2015).
Beghini, F. et al. Integrating taxonomic, functional, and strain-level profiling of diverse microbial communities with biobakery 3. Elife 10, e65088 (2021).
Bonin, N. et al. MEGARes and AMR++, v3.0: an updated comprehensive database of antimicrobial resistance determinants and an improved software pipeline for classification using high-throughput sequencing. Nucleic Acids Res. 51 (D1), D744–D752 (2023).
Florensa, A. F., Kaas, R. S., Clausen, P. T. L. C., Aytan-Aktug, D. & Aarestrup, F. M. ResFinder – an open online resource for identification of antimicrobial resistance genes in next-generation sequencing data and prediction of phenotypes from genotypes. Microb. Genomics. 8, 000748 (2022).
Gupta, S. K. et al. ARG-ANNOT, a new bioinformatic tool to discover antibiotic resistance genes in bacterial genomes. Antimicrob. Agents Chemother. 58, 212–220 (2014).
Alcock, B. P. et al. CARD 2020: antibiotic resistome surveillance with the comprehensive antibiotic resistance database. Nucleic Acids Res. 48, D517–D525 (2020).
Feldgarden, M. et al. Validating the amrfinder tool and resistance gene database by using antimicrobial resistance genotype-phenotype correlations in a collection of isolates. Antimicrob. Agents Chemother. 63, e00483119 (2019).
Pal, C., Bengtsson-Palme, J., Rensing, C., Kristiansson, E. & Larsson, D. G. J. BacMet: antibacterial biocide and metal resistance genes database. Nucleic Acids Res. 42, D737–D743 (2014).
Parks, D. H., Tyson, G. W., Hugenholtz, P. & Beiko, R. G. STAMP: statistical analysis of taxonomic and functional profiles. Bioinformatics 30, 3123–3124 (2014).
Liu, B., Zheng, D. D., Zhou, S. Y., Chen, L. H. & Yang, J. VFDB 2022: a general classification scheme for bacterial virulence factors. Nucleic Acids Res. 50(D1), D912–D917 (2022).
Xiang, Q. et al. Adsorbed sulfamethoxazole exacerbates the effects of polystyrene (~ 2 µm) on gut microbiota and the antibiotic resistome of a soil Collembolan. Environ. Sci. Technol. 53 (21), 12823–12834 (2019).
Quintela-Baluja, M. et al. Spatial ecology of a wastewater network defines the antibiotic resistance genes in downstream receiving waters. Water Res. 162, 347–357 (2019).
Qiu, Z. et al. BASALT refines Binning from metagenomic data and increases resolution of genome-resolved metagenomic analysis. Nat. Commun. 15, 2179 (2024).
Chaumeil, P. A., Mussig, A. J., Hugenholtz, P. & Parks, D. H. GTDB-Tk: a toolkit to classify genomes with the genome taxonomy database. Bioinformatics 36, 1925–1927 (2019).
Zhu, L. et al. Deciphering basic and key traits of antibiotic resistome in influent and effluent of hospital wastewater treatment systems. Water Res. 231, 119614 (2023).
Letunic, I. & Bork, P. Interactive tree of life (iTOL) v5: an online tool for phylogenetic tree display and an-notation. Nucleic Acids Res. 49, W293–W296 (2021).
Zhou, T. et al. itol.toolkit accelerates working with iTOL (Interactive tree of Life) by an automated generation of annotation files. Bioinformatics 39 (6), btad339 (2023).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R Stat. Soc. Ser. B Methodol. 57 (1), 289–300 (1995).
Tong, C., Liu, Y. X. & Huang, L. ImageGP: An easy-to-use data visualization web server for scientific researchers. iMeta 1, e5 (2022).
Breiman, L. Random forests. Mach. Learn. 45, 5–32 (2001).
Wen, T. et al. The best practice for Microbiome analysis using R. Protein Cell. 14 (10), 713–725 (2023).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 25, 1043–1055 (2015).
Chen, C. et al. Expanded catalog of microbial genes and metagenome-assembled genomes from the pig gut Microbiome. Nat. Commun. 12 (1), 1106 (2021).
Sassone-Corsi, M. & Raffatellu, M. No vacancy: how beneficial microbes cooperate with immunity to provide colonization resistance to pathogens. J. Immunol. 194 (9), 4081–4087 (2015).
Litvak, Y., Byndloss, M. X., Tsolis, R. M. & Baumler, A. J. Dysbiotic Proteobacteria expansion: a microbial signature of epithelial dysfunction. Curr. Opin. Microbiol. 39, 1–6 (2017).
Zhou, Y. et al. The panda-derived Lactobacillus plantarum G201683 alleviates the inflammatory response in DSS-induced panda microbiota-associated mice. Front. Immunol. 12, 747045 (2021).
Pakbin, B., Brück, W. M. & Rossen, J. W. A. Virulence factors of enteric pathogenic Escherichia coli: a review. Int. J. Mol. Sci. 22, 9922 (2021).
Zhu, Z. et al. High prevalence of multi-drug resistances and diversity of mobile genetic elements in Escherichia coli isolates from captive giant pandas. Ecotoxicol. Environ. Saf. 198, 110681 (2020).
Zhang, W. et al. The giant panda gut harbors a high diversity of lactic acid bacteria revealed by a novel culturomics pipeline. mSystems 9 (7), e0052024 (2024).
Wang, X. et al. Antimicrobial resistance of Escherichia coli, Enterobacter spp., Klebsiella pneumoniae and Enterococcus spp. isolated from the feces of giant panda. BMC Microbiol. 22, 102 (2022).
Yang, S. et al. Reference gene catalog and metagenome-assembled genomes from the gut Microbiome reveal the microbial composition, antibiotic resistome, and adaptability of a lignocellulose diet in the giant panda. Environ. Res. 245, 118090 (2024).
Yan, Z. et al. The impact of bamboo consumption on the spread of antibiotic resistance genes in giant pandas. Veterinary Sci. 10 (11), 630 (2023).
Mustafa, G. R. et al. Metagenomic analysis revealed a wide distribution of antibiotic resistance genes and biosynthesis of antibiotics in the gut of giant pandas. BMC Microbiol. 21 (1), 15 (2021).
Looft, T. et al. In-feed antibiotic effects on the swine intestinal Microbiome. PNAS 109 (5), 1691–1696 (2012).
Ajami, N. J., Cope, J. L., Wong, M. C., Petrosino, J. F. & Chesnel, L. Impact of oral Fidaxomicin administration on the intestinal microbiota and susceptibility to clostridium difficile colonization in mice. Antimicrob. Agents Chemother. 62 (5), e02112–e02117 (2018).
Liu, P. et al. Antibiotic-induced dysbiosis of the gut microbiota impairs gene expression in gut-liver axis of mice. Genes 14 (7), 1423 (2023).
Laconi, A. et al. Amoxicillin and thiamphenicol treatments May influence the co-selection of resistance genes in the chicken gut microbiota. Sci. Rep. 12, 20413 (2022).
Jernberg, C., Löfmark, S., Edlund, C. & Jansson, J. K. Long-term ecological impacts of antibiotic administration on the human intestinal microbiota. ISME J. 1 (1), 56–66 (2007).
Ladirat, S. E. et al. Exploring the effects of galacto-oligosaccharides on the gut microbiota of healthy adults receiving amoxicillin treatment. Br. J. Nutr. 112 (4), 536–546 (2014).
Dhariwal, A. et al. Differential response to prolonged amoxicillin treatment: long-term resilience of the Microbiome versus long-lasting perturbations in the gut resistome. Gut Microbes. 15 (1), 2157200 (2023).
Yang, S. C., Hung, C. F., Aljuffali, I. A. & Fang, J. Y. The roles of the virulence factor IpaB in Shigella spp. in the escape from immune cells and invasion of epithelial cells. Microbiol. Res. 181, 43–51 (2015).
Sicard, J. F., Le Bihan, G., Vogeleer, P., Jacques, M. & Harel, J. Interactions of intestinal bacteria with components of the intestinal mucus. Front Cell. Infect Microbiol. 7, 387 (2017).
Jurado-Martín, I., Sainz-Mejías, M. & McClean, S. Pseudomonas aeruginosa: an audacious pathogen with an adaptable arsenal of virulence factors. Int. J. Mol. Sci. 22, 3128 (2021).
Sarowska, J. et al. Virulence factors, prevalence and potential transmission of extraintestinal pathogenic Escherichia coli isolated from different sources: recent reports. Gut Pathog. 11 (1), 10 (2019).
Dai, Y., Gao, J. & Jiang, M. Case report: a rare infection of multidrug-resistant Aeromonas caviae in a pediatric case with acute lymphoblastic leukemia and review of the literature. Front. Pediatr. 12, 1233600 (2024).
Jorge, R., Teixeira, S., Marques, M., Pereira, J. & Paiva, J. A. Enterococcus Hirae bacteremia associated with traumatic soft tissue infection: A case report. Cureus 16 (11), e74092 (2024).
Karlbom, O., Sunnerhagen, T. & Ljungquist, O. Morganella Morganii bloodstream infection affects the elderly in close contact with health care. IJID Reg. 13, 100480 (2024).
Alsaadi, A. et al. Epidemiology and clinical characteristics of Morganella Morganii infections: a multicenter retrospective study. J. Infect. Public. Health. 17 (3), 430–434 (2024).
Cordaillat-Simmons, M., Rouanet, A. & Pot, B. Live biotherapeutic products: the importance of a defined regulatory framework. Exp. Mol. Med. 52, 1397–1406 (2020).
Jit, M., Anderson, M. & Cooper, B. Quantifying the benefits of vaccines in combating antimicrobial resistance. Eurohealth 26, 16–19 (2020).
Maugeri, A., Barchitta, M. & Agodi, A. Vaccination coverage in Italian children and antimicrobial resistance: an ecological analysis. Antimicrob. Resist. Infect. Control. 11, 136 (2022).
Fu, Y. et al. Gut microbiota research nexus: one health relationship between human, animal, and environmental resistomes. mLife 2 (4), 350–364 (2023).
Yu, S. et al. Clostridium Butyricum isolated from giant panda can attenuate dextran sodium sulfate-induced colitis in mice. Front. Microbiol. 15, 1361945 (2024).
Zhang, L., Huang, Y., Zhou, Y., Buckley, T. & Wang, H. H. Antibiotic administration routes significantly influence the levels of antibiotic resistance in gut microbiota. Antimicrob. Agents Chemother. 57, 3659–3666 (2013).
Zhu, D. et al. Insights into the roles of fungi and protist in the giant panda gut Microbiome and antibiotic resistome. Enviro Int. 155, 106703 (2021).
Koo, H. et al. Individualized recovery of gut microbial strains post antibiotics. NPJ Biofilms Microbiomes. 5 (1), 30 (2019).
Acknowledgements
We thank Lijun Zheng, Fuyao Han and other staff at the Chengdu Research Base of Giant Panda Breeding for assistance with sampling.
Author information
Authors and Affiliations
Contributions
W.Z. and S.S.E. wrote the main manuscript text. W.Z. and M.Z. prepared the main figures. J.X., H.H., and W.L. collected the samples. W.Z., H.L., and D.L. supervised the project.All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Zhang, W., Zhang, M., Xie, J. et al. Dynamics of the gut microbiome and resistome in response to prophylactic antibiotic treatment in post-surgical giant pandas. Sci Rep 15, 43425 (2025). https://doi.org/10.1038/s41598-025-25645-6
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-25645-6