Abstract
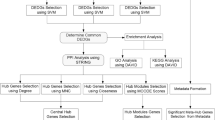
Plasticizers, such as dimethyl phthalate (DMP), diethyl phthalate (DEP), and dioctyl phthalate (DOP), are ubiquitously used in industrial products but pose potential hepatotoxic risks. This study employs network toxicology, molecular docking, and molecular dynamics (MD) simulations to investigate the molecular mechanisms underlying plasticizer-induced liver damage. The potential targets of DMP, DEP, and DOP were predicted using multiple databases (ChEMBL, STITCH, etc.), and liver damage-related genes were retrieved from GeneCards, OMIM, and DisGeNET. Overlapping targets were identified through Venn diagrams. A protein-protein interaction (PPI) network was constructed using STRING and Cytoscape, with hub genes prioritized by the Maximal Clique Centrality(MCC) algorithm. Functional enrichment analysis (GO/KEGG) was performed to identify and characterize key pathways. Molecular docking (AutoDock Vina) and MD simulations (AMBER18) were used to validate interactions between plasticizers and key targets. A total of 100 overlapping targets were identified, with KRAS, KIT, LCK, NR3C1, and FLT1 emerging as hub genes. Enrichment analysis highlighted pathways including PI3K-Akt, Ras signaling, and proteoglycans in cancer. Molecular docking revealed stable interactions between FLT1-DEP/DMP and KIT-DOP, supported by MD simulations showing low RMSD/RMSF values. This integrative approach identifies critical targets (e.g., FLT1, KIT) and pathways (e.g., PI3K-Akt/Ras) that mediate plasticizer hepatotoxicity, providing novel insights into mechanistic pathways and potential therapeutic targets.
Similar content being viewed by others
Introduction
Plasticizers, a class of polymeric chemical additives widely used in industrial manufacturing, are designed to enhance the processability and mechanical properties of plastics. By increasing material pliability and resilience, these compounds significantly broaden the application scope of plastic products across diverse sectors, from medical devices to food packaging1,2. Among them, Dimethyl phthalate (DMP), Diethyl phthalate (DEP), and dioctyl phthalate (DOP) are most commonly used (Table 1)3,4. In the era of advanced industrialization and modernization, plastic-derived products have become ubiquitous and indispensable in daily life. However, the lipophilic nature and environmental persistence of these compounds not only facilitate their widespread application but also contribute to their bioaccumulative potential in both biological systems and ecosystems5. A growing body of evidence indicates that certain plasticizers may elicit a spectrum of adverse health effects upon human exposure, particularly manifesting hepatotoxic consequences6,7. This study employs an integrative approach that combines network toxicology, molecular docking, and molecular dynamics (MD) simulations to systematically unravel the mechanistic underpinnings of plasticizer-induced liver damage.
Materials and methods
Underlying targets of plasticizers and liver damage
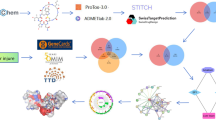
The standard structures and Simplified Molecular-Input Line-Entry System (SMILES) notations of DEP, DMP, and DOP were identified by querying PubChem8. Using the retrieved structures, potential molecular targets of DEP, DMP, and DOP were predicted through multiple databases, including ChEMBL 33, STITCH, SwissTargetPrediction, Similarity Ensemble Approach (SEA), and PharmMapper. The structural consistency of the search results was carefully examined and compared. Potential targets identified from the searches were consolidated, and duplicates, including redundant ChEMBL IDs and STITCH codes, were removed. The target names were standardized using the UniProt database.
Targets associated with Liver damage were collected by searching the GeneCards, OMIM, and DisGeNET databases using the keyword “Liver damage.” Additionally, Venn diagrams were employed to identify common potential targets between “Plasticizers” and “Liver damage,” with the overlapping targets considered as the key candidates for further analysis.
Construction of protein-protein interaction (PPI) network and target clustering analysis
The STRING (https://string-db.org)9 database was utilized to construct a protein-protein interaction (PPI) network of functionally related targets and to extract general data on candidate targets. The confidence score for protein interactions was set to a value greater than 0.4. Using Cytoscape 3.9.1 software (https://cytoscape.org/), the general data were employed to construct molecular pathway networks depicting the toxicity mechanisms of DEP, DMP, and DOP in Liver damage. The top ten hub genes were identified using the Maximal Clique Centrality (MCC) algorithm and degree ranking within the Cytoscape software.
Functional enrichment analysis
The key target genes were subjected to Gene Ontology (GO) biological process and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses using R version 4.0.5. The results were subsequently visualized for comprehensive interpretation10,11,12.
Molecular docking and MD simulation
The top five genes ranked by degree in the PPI network were selected for molecular docking with DEP, DMP, and DOP. We performed molecular docking of DEP, DMP, and DOP with the top five targets using AutoDock 1.5.7, PyMOL, and Chem3D 14.0. First, we downloaded the PDB crystal structures of the target proteins from RCSB (https://www.rcsb.org/). Then, we used PyMOL to remove the solvents/organic molecules (and extract one monomer if the protein was a multimer), saving the resulting structures as PDB files. Next, we imported ligand-receptor complexes into AutoDock 1.5.7 for hydrogenation (saving as PDBQT for docking), set the active pocket to cover most of the receptor (recording its coordinates), and ran docking 20 times via Vina (saving output as a txt file). The resulting PDBQT file was optimized in PyMOL to get the final docking conformation. Finally, we used R 4.0.5 to make a heatmap of binding energies and PyMOL to visualize docking structures13,14,15.
We used the AMBER18 software package to carry out the MD simulations. For the plasticizers, we set their parameters using the ANTECHAMBER module and GAUSSIAN09 software16,17. When processing the proteins, we used the FF14SB force field, while the GAFF2 force field was applied to the plasticizers. The addition of hydrogen atoms and sodium ions to protein-ingredient complexes was accomplished in the LEaP module to ensure overall charge neutrality. The proteins were solvated using a three-point water model in a dodecahedron box with dimensions of 1 nm in all directions. The energy minimization (EM) phase used the steepest descent integrator, terminating when the maximum force dropped below 10.0 kJ/mol. The system was equilibrated using Canonical (NVT) and Isobaric (NPT) for 100 picoseconds. During the MD simulation phase, we set the time step to 2 fs and simulated a total of 200 ns. Trajectories were recorded every 10 ps.
Result
Identification of the underlying targets of plasticizers and liver damage
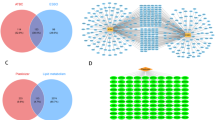
A total of 347 targets of plasticizers were initially identified through ChEMBL, STITCH, Swiss Target Prediction, the SEA, and PharmMapper. 2,513 liver damage-associated targets were curated from the GeneCards, OMIM, and DisGeNET databases. After integrating and deduplicating these datasets, 100 overlapping targets were identified as potential mediators of plasticizer-induced liver toxicity (Fig. 1A and B).
Targets of plasticizers and liver damage. (A) Potential targets of plasticizers (B) Venn diagram of the targets of plasticizers and Liver damage.
Construction of PPI network and identification of hub genes
The 100 potential targets associated with plasticizer-induced liver damage were analyzed using the STRING database to generate a PPI network. Subsequently, the PPI network was subjected to cluster analysis via the MCODE plugin in Cytoscape (v3.9.1), which identified the top five hub targets as KRAS, KIT, LCK, NR3C1, and FLT1 (Fig. 2).
Hub targets of plasticizers and liver damage.
Functional enrichment analysis
Functional enrichment analysis of the 100 potential targets was conducted through GO and Kyoto Encyclopedia of KEGG assessments. GO analysis revealed significant associations with biological processes (BP), cellular components (CC), and molecular functions (MF), including wound healing, positive regulation of kinase activity, vesicle lumen, protein tyrosine kinase activity, and endopeptidase activity. KEGG pathway enrichment demonstrated prominent involvement in the PI3K-Akt signaling pathway, Ras signaling pathway, proteoglycans in cancer, and checkpoint regulation in cancer (Fig. 3A,B).
Functional enrichment analysis of intersecting target genes. (A) GO enrichment analysis of public target genes. (B) KEGG enrichment analysis of intersecting target genes.
Molecular docking and MD simulation
Integration of KEGG pathway enrichment and network analysis results prioritized five hub targets (KRAS, KIT, LCK, NR3C1, FLT1) for molecular docking with core plasticizer components. Fifteen docking complexes were systematically evaluated and visualized via a binding energy heatmap (Fig. 4). Notably, FLT1-DEP, FLT1-DMP, and KIT-DOP exhibited the lowest binding energies, indicating the most stable molecular interactions. Both 2D and 3D views of the docking modes show how van der Waals, electrostatic, hydrogen bonding, and hydrophobic interactions contribute to the total binding energy (Table 2). FLT1-DEP: Total binding energy = -6.1 kcal/mol, with LYS861 and VAL909 as key residues (mainly contributing through van der Waals and electrostatic effects).FLT1-DMP: Total binding energy = -6.0 kcal/mol, with ASP1040 and VAL909 as key residues (mainly contributing through van der Waals and electrostatic effects).KIT-DOP: Total binding energy = -7.7 kcal/mol, with VAL654 and LYS623 as major contributors (mainly through van der Waals and electrostatic impact).
The binding free energy of molecular docking.
To further validate the precise binding mechanisms and interaction stability, MD simulations were performed on the three prioritized complexes. Root Mean Square Deviation (RMSD) analysis revealed minor fluctuations across all systems, with all complexes achieving equilibrium stability after 75 ns. Complementarily, Root Mean Square Fluctuation (RMSF) analysis was employed to assess residue-level flexibility and active site dynamics over a 200 ns time frame. The RMSF profiles demonstrated that FLT1-DEP, FLT1-DMP, and KIT-DOP complexes exhibited low flexibility stabilization, particularly at critical active site residues, suggesting strong ligand-induced structural rigidity (Fig. 5A,B).
RMSD and RMSF of MD simulation. (A) The RMSD of FLT1-DEP. (B) The RMSD of FLT1-DMP. (C) The RMSD of KIT-DOP. (D)The RMSF of FLT1 when conjugated with DEP. (E) The RMSF of FLT1 when conjugated with DMP. (F) The RMSF of KIT when conjugated with DOP.
Discussion
In this study, we employed a combination of computational methods to investigate how exposure to plasticizers may lead to liver damage at the molecular level. We identified 100 key targets that are linked to both plasticizers and liver damage. This finding not only demonstrates that plasticizers act on multiple targets (instead of just one) but also helps connect environmental chemical exposure to the actual biological changes that cause liver problems. Next, we’ll discuss how these findings fit into the broader context of toxicology research, examine the implications of our methods for future studies, and suggest specific steps that can be taken to explore this topic further.
Hub targets as drivers of liver damage
The prioritization of KRAS, KIT, LCK, NR3C1, and FLT1 as top hub targets aligns with their documented roles in liver injury cascades. KRAS, a linchpin of the PI3K-Akt and Ras pathways, is a known mediator of chemically induced hepatic fibrosis via TGF-β/Smad3 activation and extracellular matrix remodeling18,19. Its prominence in our network suggests that plasticizers may mimic pro-fibrotic stimuli, exacerbating parenchymal damage. FLT1 (VEGFR1), exhibiting the strongest binding affinities with DEP/DMP, emerged as a critical node in both docking and dynamics analyses. FLT1’s role in regulating sinusoidal endothelial permeability implies that plasticizer binding could disrupt hepatic vascular integrity, potentiating hypoxia-driven injury20,21,22,23. KIT’s interaction with DOP highlights stem cell factor (SCF)/KIT signaling in hepatic progenitor cell activation—a double-edged sword mediating both regenerative responses and fibrotic scarring24,25,26.
Pathway convergence: from toxicity to carcinogenesis
The enrichment of PI3K-Akt, Ras, and cancer-related pathways unveils a striking overlap between plasticizer toxicity and oncogenic signaling27,28,29,30. The PI3K-Akt pathway, a master regulator of cell survival and proliferation, is frequently hyperactivated in hepatocellular carcinoma (HCC)31,32,33. Our findings suggest that chronic exposure to plasticizers may mimic growth factor signaling, thereby creating a pro-oncogenic microenvironment. The prominence of wound healing and kinase activity regulation in GO terms may reflect compensatory hepatocyte repair mechanisms34,35. However, sustained activation of these processes under plasticizer burden could drive maladaptive remodeling36,37.
Molecular docking and MD simulation
Molecular docking and MD simulations helped us understand how plasticizers (DMP, DEP, DOP) interact with key targets, supporting the links we identified through network analysis. In docking tests with AutoDock Vina, FLT1 stuck strongly to DEP and DMP, while KIT formed stable pairs with DOP. These results matched the targets that were most important in the PPI network. The low binding energies for these pairs tell us these interactions aren’t random.
To check if these docking results hold up in real conditions, we did MD simulations with AMBER18 on the top three complexes. We set up the simulations to mimic the body’s environment, using FF14SB for proteins, GAFF2 for plasticizers, and TIP3P for water. The results showed all complexes settled into a stable state within 75 ns. Low RMSD values indicated that their structures didn’t change significantly over time. Additionally, the active parts of FLT1 and KIT exhibited low RMSF values, indicating that plasticizers made these areas less flexible. This stability and reduced flexibility support the docking findings: these plasticizer-target interactions aren’t short-lived. They may remain in the body, disrupting normal protein function and contributing to liver damage through pathways such as PI3K-Akt and Ras signaling. Together, docking and MD simulations suggest that FLT1, KIT, and their related pathways may play a crucial role in how plasticizers harm the liver.
Methodological innovations and limitations
Our framework advances toxicological screening by unifying network pharmacology with dynamics validation and prioritizing KEGG-driven targets. However, limitations necessitate caution, including database bias, metabolite oversight, and cell type specificity.
Translational implications and future directions
The FLT1-DEP/DMP and KIT-DOP complexes represent high-priority candidates for biomonitoring. Small-molecule inhibitors of KRAS or FLT1 could be repurposed to mitigate the toxicity of plasticizers. Future work should focus on multi-omics validation, dynamic exposure modeling, and interspecies extrapolation.
Conclusion
In conclusion, this research integrates network toxicology, molecular docking, and molecular dynamics simulations to systematically explore the hepatotoxic mechanisms of plasticizers (DMP, DEP, DOP). A total of 100 overlapping targets between plasticizers and liver damage were identified, with KRAS, KIT, LCK, NR3C1, and FLT1 emerging as key hub genes. Functional enrichment analysis emphasized the involvement of the PI3K-Akt and Ras signaling pathways, suggesting a potential association between plasticizer exposure and pro-fibrotic or oncogenic processes. Molecular docking and MD simulations further verified strong and stable interactions between FLT1-DEP/DMP and KIT-DOP, indicating that these complexes may play crucial roles in disrupting hepatic function. These findings lay a mechanistic foundation for understanding plasticizer-induced liver injury and highlight potential therapeutic targets for intervention. Future research should incorporate experimental validation and more comprehensive toxicological assessments to confirm these computational findings.
Data availability
All data supporting the findings of this study are available within the paper and its Supplementary Information.
References
Chung, B. Y. et al. Risk assessment of phthalates in pharmaceuticals. J. Toxicol. Environ Health A. 82, 351–360. https://doi.org/10.1080/15287394.2019.1598053 (2019).
Arrigo, F., Impellitteri, F., Piccione, G. & Faggio, C. Phthalates and their effects on human health: focus on erythrocytes and the reproductive system. Comp. Biochem. Physiol. Toxicol. Pharmacology: CBP. 270, 109645. https://doi.org/10.1016/j.cbpc.2023.109645 (2023).
Wang, Y. & Qian, H. Phthalates and their impacts on human health. Healthcare 9, 603 (2021).
Hahladakis, J. N., Velis, C. A., Weber, R., Iacovidou, E. & Purnell, P. An overview of chemical additives present in plastics: Migration, release, fate and environmental impact during their use, disposal and recycling. J. Hazard. Mater. 344, 179–199. https://doi.org/10.1016/j.jhazmat.2017.10.014 (2018).
Martínez-Ibarra, A. et al. Multisystemic alterations in humans induced by bisphenol A and phthalates: Experimental, epidemiological and clinical studies reveal the need to change health policies. Environ. Poll. (Barking, Essex: 1987) 271, 116380. https://doi.org/10.1016/j.envpol.2020.116380 (2021).
Campanale, C., Massarelli, C., Savino, I., Locaputo, V. & Uricchio, V. F. A detailed review study on potential effects of microplastics and additives of concern on human health. Int. J. Environ. Res. Public Health 17 https://doi.org/10.3390/ijerph17041212 (2020).
Zuri, G., Karanasiou, A. & Lacorte, S. Human biomonitoring of microplastics and health implications: A review. Environ. Res. 237, 116966. https://doi.org/10.1016/j.envres.2023.116966 (2023).
Kim, S. et al. PubChem 2023 update. Nucleic Acids Res. 51, D1373–D1380. https://doi.org/10.1093/nar/gkac956 (2022).
Szklarczyk, D. et al. STRING v11: protein–protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 47, D607–D613. https://doi.org/10.1093/nar/gky1131 (2018).
Kanehisa, M., Furumichi, M., Sato, Y., Matsuura, Y. & Ishiguro-Watanabe, M. KEGG: biological systems database as a model of the real world. Nucleic Acids Res. 53, D672–d677. https://doi.org/10.1093/nar/gkae909 (2025).
Kanehisa, M. Toward Understanding the origin and evolution of cellular organisms. Protein Science: Publication Protein Soc. 28, 1947–1951. https://doi.org/10.1002/pro.3715 (2019).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30. https://doi.org/10.1093/nar/28.1.27 (2000).
Rigsby, R. E. & Parker, A. B. Using the PyMOL application to reinforce visual Understanding of protein structure. Biochem. Mol. Biology Education: Bimon. Publication Int. Union Biochem. Mol. Biology. 44, 433–437. https://doi.org/10.1002/bmb.20966 (2016).
Seeliger, D. & de Groot, B. L. Ligand Docking and binding site analysis with PyMOL and Autodock/Vina. J. Comput. Aided Mol. Des. 24, 417–422. https://doi.org/10.1007/s10822-010-9352-6 (2010).
Tang, S. et al. Accelerating AutoDock Vina with GPUs. Molecules (Basel Switzerland) 27 https://doi.org/10.3390/molecules27093041 (2022).
Lee, T. S. et al. Alchemical binding free energy calculations in AMBER20: advances and best practices for drug discovery. J. Chem. Inf. Model. 60, 5595–5623. https://doi.org/10.1021/acs.jcim.0c00613 (2020).
Zhang, H. et al. CHARMM-GUI free energy calculator for practical ligand binding free energy simulations with AMBER. J. Chem. Inf. Model. 61, 4145–4151. https://doi.org/10.1021/acs.jcim.1c00747 (2021).
Kemeny, N. E. et al. KRAS mutation influences recurrence patterns in patients undergoing hepatic resection of colorectal metastases. Cancer 120, 3965–3971. https://doi.org/10.1002/cncr.28954 (2014).
Chung, S. I. et al. Comparison of liver oncogenic potential among human RAS isoforms. Oncotarget 7, 7354–7366. https://doi.org/10.18632/oncotarget.6931 (2016).
Kremer, V. et al. Neuro-oncological ventral antigen 2 regulates splicing of vascular endothelial growth factor receptor 1 and is required for endothelial function. Reproductive Sci. (Thousand Oaks Calif). 30, 678–689. https://doi.org/10.1007/s43032-022-01044-4 (2023).
Ciribilli, Y. et al. The coordinated p53 and Estrogen receptor cis-regulation at an FLT1 promoter SNP is specific to genotoxic stress and Estrogenic compound. PloS One. 5, e10236. https://doi.org/10.1371/journal.pone.0010236 (2010).
Raikwar, N. S., Liu, K. Z. & Thomas, C. P. N-terminal cleavage and release of the ectodomain of Flt1 is mediated via ADAM10 and ADAM 17 and regulated by VEGFR2 and the Flt1 intracellular domain. PloS One. 9, e112794. https://doi.org/10.1371/journal.pone.0112794 (2014).
Das, S. et al. ETV2 and VEZF1 interaction and regulation of the hematoendothelial lineage during embryogenesis. Front. cell. Dev. Biology. 11, 1109648. https://doi.org/10.3389/fcell.2023.1109648 (2023).
Duan, J. L. et al. Age-related liver endothelial zonation triggers steatohepatitis by inactivating pericentral endothelium-derived C-kit. Nat. Aging. 3, 258–274. https://doi.org/10.1038/s43587-022-00348-z (2023).
Wang, W., Shui, L., Liu, Y. & Zheng, M. C. -Kit, a Double-Edged sword in liver regeneration and diseases. Front. Genet. 12, 598855. https://doi.org/10.3389/fgene.2021.598855 (2021).
Duan, J. L. et al. Notch-Regulated c-Kit-Positive liver sinusoidal endothelial cells contribute to liver zonation and regeneration. Cell. Mol. Gastroenterol. Hepatol. 13, 1741–1756. https://doi.org/10.1016/j.jcmgh.2022.01.019 (2022).
Glaviano, A. et al. PI3K/AKT/mTOR signaling transduction pathway and targeted therapies in cancer. Mol. Cancer. 22, 138. https://doi.org/10.1186/s12943-023-01827-6 (2023).
He, Y. et al. Targeting PI3K/Akt signal transduction for cancer therapy. Signal. Transduct. Target. Therapy 6 https://doi.org/10.1038/s41392-021-00828-5 (2021).
Holderfield, M. et al. Concurrent Inhibition of oncogenic and wild-type RAS-GTP for cancer therapy. Nature 629, 919–926. https://doi.org/10.1038/s41586-024-07205-6 (2024).
Chen, K., Zhang, Y., Qian, L. & Wang, P. Emerging strategies to target RAS signaling in human cancer therapy. J. Hematol. Oncol. 14 https://doi.org/10.1186/s13045-021-01127-w (2021).
Tian, L. Y., Smit, D. J. & Jücker, M. The role of PI3K/AKT/mTOR signaling in hepatocellular carcinoma metabolism. Int. J. Mol. Sci. 24 https://doi.org/10.3390/ijms24032652 (2023).
Sun, B. et al. FDX1 downregulation activates mitophagy and the PI3K/AKT signaling pathway to promote hepatocellular carcinoma progression by inducing ROS production. Redox Biol. 75, 103302. https://doi.org/10.1016/j.redox.2024.103302 (2024).
Sun, E. J., Wankell, M., Palamuthusingam, P., McFarlane, C. & Hebbard, L. Targeting the PI3K/Akt/mTOR pathway in hepatocellular carcinoma. Biomedicines 9 https://doi.org/10.3390/biomedicines9111639 (2021).
Kiourtis, C. et al. Hepatocellular senescence induces multi-organ senescence and dysfunction via TGFβ. Nat. Cell Biol. 26, 2075–2083. https://doi.org/10.1038/s41556-024-01543-3 (2024).
Cheng, N., Kim, K. H. & Lau, L. F. Senescent hepatic stellate cells promote liver regeneration through IL-6 and ligands of CXCR2. JCI Insight 7 https://doi.org/10.1172/jci.insight.158207 (2022).
Boran, T. et al. An evaluation of a hepatotoxicity risk induced by the microplastic polymethyl methacrylate (PMMA) using HepG2/THP-1 co-culture model. Environ. Sci. Pollut. Res. Int. 31, 28890–28904. https://doi.org/10.1007/s11356-024-33086-3 (2024).
Rosellini, M. et al. Impact of plastic-related compounds on the gene expression signature of HepG2 cells transfected with CYP3A4. Arch. Toxicol. 98, 525–536. https://doi.org/10.1007/s00204-023-03648-4 (2024).
Acknowledgements
We would like to express our sincere gratitude to the editors for their professional guidance and efficient handling of this manuscript throughout the submission and review process. We are also deeply appreciative of the anonymous reviewers for their insightful and constructive comments, which have significantly helped improve the quality and rigor of this work.
Author information
Authors and Affiliations
Contributions
All authors actively contributed to the conceptual development of this article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Feng, C., Chun, L., KeLei, S. et al. Exploring the mechanism of plasticizers action in liver damage based on network toxicology, molecular docking, and molecular dynamic simulation. Sci Rep 15, 42333 (2025). https://doi.org/10.1038/s41598-025-26361-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-26361-x