Abstract
Quantifying structural brain changes is critical for diagnosing and monitoring neurodegenerative diseases. Although magnetic resonance imaging (MRI) is the silver standard, limited accessibility and cost hamper routine use. We developed a deep learning–based framework using the nnU-Net for brain segmentation using computed tomography (CT) to assess cerebrospinal fluid (CSF) volume changes, as an indirect marker of tissue loss, and evaluated its utility across Alzheimer’s disease (AD) stages and frontotemporal dementia (FTD) subtypes. We included 2357 participants: cognitively unimpaired (CU, n = 595), mild cognitive impairment (MCI, n = 954), dementia of Alzheimer’s type (DAT, n = 663), and FTD subtypes (FTD, n = 145, behavioral variant FTD (bvFTD, n = 66), nonfluent variant primary progressive aphasia (nfvPPA, n = 29), and semantic variant PPA (svPPA, n = 50). CT-based segmentation was trained and validated using 3D T1-weighted MRI as reference. We assessed (1) segmentation accuracy via Dice similarity coefficients (DSCs), (2) reliability and precision using correlation and Bland–Altman analyses, and (3) clinical utility by identifying stage- and region-specific changes in CSF volumes. Key regions, including anterior and posterior lateral ventricles, showed DSCs above 0.93 and correlations ranging from 0.822 to 0.996. CT-based measurements revealed increasing CSF volumes from CU to DAT and distinct patterns of CSF volume enlargement across FTD subtypes. This framework enables accurate, reliable assessment of CSF volume changes as an indirect marker of atrophy, and supports early detection and differential diagnosis.
Similar content being viewed by others
Introduction
Neurodegenerative diseases, such as Alzheimer’s disease (AD) and frontotemporal dementia (FTD), are characterized by progressive neuronal loss, which leads to gyral shrinkage and sulcal widening1,2,3,4. Depending on each disease’s specific mechanisms and the selective vulnerability of particular brain regions, a unique pattern of brain tissue loss and corresponding CSF volume enlargement emerges—shaping the distinct clinical phenotypes observed in these conditions5,6,7. Therefore, quantifying brain structural changes (including CSF volume enlargement as an indirect marker of atrophy) is crucial for differential diagnosis and for monitoring the progression of each neurodegenerative disease3. Conventionally, magnetic resonance imaging (MRI) with 3dimensional (D) T1‐weighted scans has been used to quantify brain atrophy through techniques such as voxel-based morphometry or cortical thickness measurements8,9,10. However, MRI can pose risks for patients with implantable devices like pacemakers or cardioverter defibrillators, and its cost, limited accessibility, and longer acquisition times often make routine clinical use impractical.
Computed tomography (CT) offers a more accessible, rapid, and cost-effective imaging option compared to MRI11,12; however, its lower spatial resolution, two-dimensional acquisition in many clinical settings, and insufficient contrast between white and gray matter have historically made it difficult to perform automated volumetric analyses. As a result, clinicians have largely relied on visually assessing sulcal widening and ventricular enlargements—an inherently subjective process prone to inter- and intra-rater variability, prompting the development of standardized visual rating scales that still struggle with detecting subtle changes12,13,14. To overcome these challenges, previous research has consistently sought to integrate the rich anatomical information from MRI to enhance CT analysis. For example, Rorden et al.15 developed integrated CT-MRI atlases to improve anatomical labeling in CT scans, and more recently, Srikrishna et al.16 demonstrated that using MRI-derived labels to train deep learning models on CT enables accurate tissue classification. Despite such progress, reliably delineating fine structural boundaries across the whole brain remains difficult16,17,18. Building on these foundational concepts, our approach focuses on pronounced changes in CSF volume at sulci and ventricle regions, approximates the precision of 3D T1 MRI measurements, and provides detailed, region-specific information—offering a practical alternative for routine clinical assessment.
A key concept underpinning our work is the distinction between stage‐specific and region‐specific atrophy patterns across different neurodegenerative diseases, which in our study are indirectly reflected through changes in CSF volumes (ventricular and sulcal enlargement). In AD, tau Braak staging describes a progression of atrophy beginning in the medial temporal region, then extending to lateral temporal, parietal, and finally frontal areas7,19. Meanwhile, FTD subtypes manifest distinct regional signatures: behavioral variant FTD (bvFTD) typically involves widespread atrophy in bilateral frontal and temporal regions20, nonfluent variant PPA (nfvPPA) shows predominant atrophy in the left frontal region, and semantic variant PPA (svPPA) presents marked atrophy in the left temporal region21,22. These heterogeneous patterns highlight the importance of tailored volumetric assessments that can capture both disease progression and subtype-specific vulnerabilities.
In this study, we developed a robust deep learning model for CT-based brain segmentation using nnU-Net framework which automatically configures U-Net architectures based on the characteristics of the input dataset, to evaluate its clinical utility. First, we compared the segmentation results with 3D T1 MRI—our silver standard—and then examined the reliability and precision of our CT-based measurements. Finally, we investigated whether this approach could capture stage-specific changes in CSF volumes in the AD continuum, as well as the distinct region-specific patterns of CSF volume enlargement that characterize FTD subtypes.
Methods
Participants
A total of 2,357 participants who underwent both brain 3D T1 MRI and CT, were recruited from the Samsung Medical Center (SMC). The participants comprised individuals with cognitively unimpaired (CU), mild cognitive impairment (MCI), and dementia of Alzheimer’s type (DAT). CU individuals demonstrated no subjective cognitive complaints or functional impairments, with cognitive performance confirmed to be within normal limits through detailed neuropsychological assessments. MCI was diagnosed based on the National Institute on Aging-Alzheimer’s Association (NIA-AA) criteria23, characterized by measurable cognitive decline in one or more domains without significant interference in daily functional activities. Among the 954 individuals with MCI, 96 (10.1%) were classified as single-domain amnestic MCI, 850 (89.1%) as multiple-domain amnestic MCI, and 8 (0.8%) as non-amnestic MCI. DAT was diagnosed following the NIA-AA guidelines2, requiring evidence of significant cognitive decline, including memory impairment, that interfered with independence in daily life and was consistent with an Alzheimer’s disease etiology. Among the 663 individuals with DAT, 641 (96.7%) were classified as probable AD, while 22 (3.3%) were classified as possible AD, including logopenic PPA (lvPPA; N = 11), posterior cortical atrophy (PCA; N = 8), and frontal variant AD (fvAD; N = 3). Moreover, participants with FTD syndromes included those with a clinical diagnosis of bvFTD, nfvPPA, or svPPA. Probable bvFTD was clinically defined based on the criteria outlined by Rascovsky et al.20, whereas nfvPPA and svPPA were diagnosed based on the criteria provided by Gorno-Tempini et al.22 All FTD syndromes were diagnosed based on the patient’s clinical course, neurologic examination, neuropsychological testing, and brain imaging. Amyloid PET positivity, defined as a Klunk centiloid value > 20, was available for 2,346 participants and observed in 1,271 (54.9%) (see Table 1). Clinical diagnoses were made independently of amyloid biomarker results. We excluded participants who had any of the following conditions: (1) white matter hyperintensities due to radiation injury, multiple sclerosis, vasculitis, leukodystrophy or metabolic disorders; (2) traumatic brain injury; (3) territorial infarction; (4) brain tumor; and (5) rapidly progressive dementia(RPD). The RPD and monogenic dementia (e.g., MAPT, GRN, C9orf72 expansions) were excluded to focues on degenerative trajectories typical of sporadic AD/FTD. Exclusion of RPD relied on clinical judgment by neurologists, and no MMSE cut-off was used.
The study protocol received approval from the Institutional Review Board of SMC. Written informed consent was obtained from each participant and all procedures were conducted in accordance with the approved guidelines.
An additional set of 250 MRI scans from the Alzheimer’s Disease Neuroimaging Initiative (ADNI) dataset was used to further validate the robustness of the pipeline.
Neuropsychological tests
Most participants underwent the Seoul Neuropsychological Screening Battery (SNSB)24,25, a standardized tool widely used in Korea to assess cognitive function across five domains: attention, memory, language, visuospatial ability, and frontal/executive function. The SNSB has been validated and normed in Korea, with normative data derived from 1,067 healthy individuals. In this study, we administered the following core SNSB tests26: the Korean version of the Boston Naming Test (K-BNT) and the Controlled Oral Word Association Test (COWAT) for language; the Rey Complex Figure Test (RCFT, copy score) for visuospatial ability; the Seoul Verbal Learning Test (SVLT) and RCFT for verbal and visual memory; and attention/executive tasks including COWAT (animal, supermarket, phonemic), the Stroop color–reading test, the Korean Trail-Making Test–Part B (K-TMT-B), and Digit Span (forward and backward). Group-level performance measures are summarized in Table 1.
Functional impairment was assessed using the Korean Instrumental Activities of Daily Living (K-IADL) scale, a validated tool for elderly populations that evaluates daily functions across multiple domains (e.g., grooming, shopping, transportation, medication management). Caregiver input was incorporated to ensure accuracy of ratings27.
Acquisition of CT images and 3D T1 images
We acquired CT images from all subjects at Samsung Medical Center using a Discovery STe PET-CT scanner (GE Medical Systems, Milwaukee, WI, USA) in the three-dimensional scanning mode, which examines 47, 3.3-mm thick slices spanning the entire brain. CT images were also acquired using a 16-slice helical CT (140 keV, 80 mA, 3.75-mm section width) for attenuation correction were reconstructed in a 512 × 512 matrix. Voxel size of CT images acquired by PET-CT scanner are 0.5 mm × 0.5 mm × 3.27 mm. The signal-to-noise ratio was checked through Phantom study (3.75 mm slice thickness, 120 kVp, 190 mA), and it was conducted by GE Discovery STe PET-CT scanner.
To acquire 3D T1 turbo field-echo MRI scans from all participants at SMC, a 3.0 T MRI scanner (Philips 3.0 T Achieva; Philips Healthcare, Andover, MA, USA) was used with following parameters: sagittal slice thickness of 1.0 mm with 50% overlap, repletion time (TR) of 9.9 ms, echo time (TE) of 4.6 ms, flip angle of 8°, and matrix size of 240 × 240 pixels reconstructed to 480 × 480 over a field of view of 240 mm.
Preprocessing
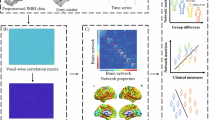
Preprocessing was performed on three different brain imaging modalities prior to applying the deep learning segmentation model (Fig. 1a). First, the 3D T1 images were resampled to 1 mm isotropic voxels and segmented using the SynthSeg function in FreeSurfer (v7.4.2; http://surfer.nmr.mgh.harvard.edu/)28,29. Subsequently, a stereotaxic atlas was employed to subdivide the cerebrospinal fluid (CSF) regions into 14 distinct areas. The CSF adjacent to the gray matter was partitioned into eight regions corresponding to the left and right frontal, occipital, parietal, and temporal lobes. Additionally, the lateral ventricle (LV) were divided into six regions, comprising the left and right anterior, posterior, and inferior regions30,31.
Overview of the proposed framework for CSF volume analysis and W-score calculation pipeline in this study. The figure illustrates the algorithmic modeling process for measuring volume and W-score of each ROI: (a) Preprocessed input images, (b) Ground truth segmentation labels for CSF regions (14 ROIs) used to train the model, (c) Example output segmentation from CT and MRI, (d) Multiple linear regression model, (e) W-score calculation. The pipeline consists of two main steps: Segmentation and regression. In the segmentation step, the model is trained using preprocessed multimodal images (MRI, CT, and synthetic), along with ground truth segmentation labels for multiple ROIs. The regression model is designed to calculate W-score using the segmented volume, demographics factors (age and sex), and imaging modality. The Total CSF volume is defined as the sum of the CSF volumes across all predefined ROIs. The W-score is computed for further analysis. Abbreviations: MRI = magnetic resonance imaging; CT = computed tomography; HU = housefield unit; ROI = region of interest; CSF = cerebrospinal fluid volume; ICV = intracranial volume; LV = lateral ventricle.
The CT images were first corrected for brain tissue Hounsfield units (HU) and then co-registered to the corresponding T1 MRI using Advanced Normalization Tools32. The details of the HU correction method are described in the original methodology paper15.
To generate synthetic images, we randomly selected one of the pre-segmented label maps with detailed brain region annotations and applied geometric augmentation through random spatial transformations. Then, we generated preliminary images by sampling from a randomly initialized Gaussian mixture model (GMM), conditioned on the transformed label map. The generated images undergo a series of sequential transformations, including random bias field augmentation, noise injection, intensity rescaling to a range between 0 and 1, and voxel-wise exponentiation. To simulate low-resolution and partial volume effects, we further apply Gaussian blurring followed by random low-resolution subsampling. Finally, training pairs are obtained by defining the deformed label map as ground truth and resampling the low-resolution images back to the 1 mm isotropic voxels. These synthetic images were incorporated at the training stage together with real CT/MRI data. They were designed to mimic rare anatomical variants, noise, and intensity differences that are underrepresented in clinical datasets, thereby improving the model’s robustness and generalizability.
Segmentation methods for regions of cerebrospinal fluid
The algorithm for computing CSF volume and W-score33 for each region of interest (ROI) volume follows a structured pipeline. The W-score is a statistical metric that adjusts for specific covariates, namely age, sex, and imaging modality (Fig. 1d). An overview of the proposed workflow is presented in Fig. 1. In the first step, to enable accurate segmentation of the CSF regions, we utilized a multi-modal training dataset comprising MRI, CT, and synthetic images (Fig. 1b). Further details are provided in Supplementary Table 1. In this step, the ROIs generated during the preprocessing stage were defined as the ground truth and shared with the co-registered CT, ensuring that each imaging modality had a corresponding label map.
We employed 3D nnU-Net model34, a self-configuration deep learning-based segmentation framework, which automatically adapts its architecture and training pipeline to the given dataset. We selected the 3D nnU-Net because its superior performance and flexibility have been consistently demonstrated across numerous publications and international segmentation challenges35. This self-configuring design minimizes manual tuning and reduces the risk of overfitting. Model performance was evaluated using fivefold cross-validation. Preprocessed images were further refined using the default nnUNet preprocessor and were trained based on the following implementation details: leaky rectified linear unit as the activation function, and loss function was a combination of Dice loss and cross-entropy loss. The stochastic gradient descent was adopted as the optimizer, with a learning rate of 1e−2, a weight decay of 3e−5, and trained for 1000 epochs. The batch size was set to 2. For model evaluation, we used the Dice similarity coefficient (DSC) to quantify the overlap between ground truth and the predicted segmentation. The DSC was computed as follows Eq. (1):
where TP denotes true positive, FP denotes false positives, and FN denotes false negatives. The volumes of each ROI were computed by summing the voxel values within the predicted ROIs, with each voxel representing the presence or absence of a target region.
Regression for W-score of each region of cerebrospinal fluid
To assess the deviation of CSF volume in each ROI from an expected normative value, we employed a W-score based on CSF volumes normalized by ICV (Fig. 1c). This approach enables the identification of region-specific changes in CSF volume while accounting for individual differences.
The expected CSF volume for each ROI was estimated using a multiple linear regression model trained on 1200 scans (600 MR and 600 CT scans) from CU subjects. The regression model was formulated as follows Eq. (2):
where age, sex, and modality were included as covariates to account for individual differences. The modality variable in Eq. (2) distinguishes between MRI and CT scans, allowing the model to adjust for imaging-related differences. The regression coefficients and model performance metrics are provided in Supplementary Table 2.
The W-score for each ROI was then computed as follows Eq. (3):
where \({\text{V}}_{{{\text{ROI}}}}\) represents the observed CSF volume of the ROI and \(\sigma_{{W_{ROI} }}\) denotes the standard deviation of the residuals from normative model in the CU group. Importantly, since an increase in CSF volume is indicative of brain tissue loss, we applied a negative sign to the W-score in Eq. (3) to reflect this biological interpretation (Fig. 1e). Therefore, lower W-scores indicate enlarged CSF volume or brain atrophy in the respective regions.
Statistical analysis
All statistical analyses were performed using R (R Studio version 2023.12.1 + 402). In the demographic analysis, categorical variables were compared using the chi-square tests, while continuous variables were analyzed using analysis of variance (ANOVA). Tukey’s post hoc tests were applied. Extreme values exceeding three standard deviations from the mean were classified as outliers and excluded from further statistical analysis. For the W-score analysis, group differences in continuous variables were also analyzed with an Analysis of variance (ANOVA). Tukey’s post hoc tests applied for cognitive stage (CU, MCI and DAT) following ANOVA when relevant. For FTD subtype analyses, the same procedure was applied. In both cases, all raw p-values were adjusted for multiple comparisons using the false discovery rate (FDR) method, and statistical significance was defined as adjusted p < 0.05. In FTD subtype comparisons, correction for multiple comparisons was performed using the false discovery rate (FDR). The distribution pattern of the W-scores for each ROI across the groups was visualized and analyzed using boxplots. Pearson’s correlation coefficients (r) were calculated to assess the relationship between W-score obtained from MR and CT methods. The agreement between W-scores from MR and CT methods was further evaluated using a Bland–Altman plot.
Results
Participant characteristics
The demographics of participants are presented in Table 1. A total of 2,357 participants were included in the study, consisting of CU (595, 25.2%), MCI (954, 40.5%), DAT (663, 28.1%), bvFTD (66, 2.8%), nfvPPA (29, 1.2%), or svPPA (50, 2.1%). The mean age was 70 ± 9.6 years, with 57.5% female participants. The average years of education was 11.8 ± 4.7, and the mean Mini-Mental State Examination (MMSE) score was 23.1 ± 7.1. Detailed neuropsychological test results were presented in Table 1.
Segmentation performance of each ROI
The segmentation performance for 14 region of interests (ROIs) and total CSF was evaluated using the test dataset consisting of 983 cases (Fig. 2). The anterior and posterior LV achieved the highest DSC values: 0.937 for the left anterior LV, 0.933 for the right anterior LV, 0.935 for the left posterior LV, and 0.932 for the right posterior LV. The inferior LV also showed relatively high DSCs, with 0.753 for the left region and 0.756 for the right region. Among the extracerebral CSF (eCSF) regions, the DSC values were 0.728 for both the left and right frontal, 0.719 for the left parietal, 0.717 for the right parietal, 0.695 for the left temporal, and 0.703 for the right temporal. The occipital regions showed the lowest performance, with DSC values of 0.564 for the left occipital and 0.570 for the right occipital. The overall DSC for the total CSF region was 0.823.
Evaluation of segmentation performance for predicted CSF volumes. The figure illustrates the segmentation performance of the proposed model, evaluating using the dice similarity coefficient against the silver standard. Abbreviations: CSF = cerebrospinal fluid volume; Left Anterior LV = left anterior lateral ventricle; Right Anterior LV = right anterior lateral ventricle; Left Posterior LV = left posterior lateral ventricle; Right Posterior LV = right posterior lateral ventricle; Left Inf Lat Vent = left inferior lateral ventricle; Right Inf Lat Vent = right inferior lateral ventricle; Left Frontal = left frontal CSF; Right Frontal = right frontal CSF; Left Temporal = left temporal CSF; Right Temporal = right temporal CSF; Left Parietal = left parietal CSF; Right Parietal = right parietal CSF; Left Occipital = left occipital CSF; Right Occipital = right occipital CSF; TotalCSF = total CSF.
To further evaluate the generalizability of the proposed pipeline, we tested its performance on an independent external cohort of 250 MRI scans from the Alzheimer’s Disease Neuroimaging Initiative (ADNI). The model achieved consistently high Dice scores across brain regions (Supplementary Fig. 1), supporting its robustness beyond the original training data. In addition, stratified fivefold cross-validation within the primary dataset demonstrated consistent convergence of training and validation curves and stable pseudo-Dice coefficients across folds (Supplementary Fig. 2).
Correlation between CT-derived and MRI-derived CSF W-scores across different ROIs
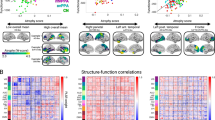
As shown in Fig. 3a, scatter plots show the relationship between the CT-based and MRI-derived W-scores, with a strong linear correlation observed in most ROIs. The correlation coefficients (r) ranged from 0.822 to 0.996, indicating high agreement between CT-based and MRI-derived W-scores. LV ROIs (Left and Right anterior LV, posterior LV, and inferior LV) exhibited the highest correlations (r > 0.98), suggesting that ventricular volume estimation is highly consistent across both modalities. In contrast, eCSF regions such as the parietal and occipital CSF demonstrated slightly lower correlations (r ≈ 0.82–0.90), potentially due to the lower contrast resolution of CT in distinguishing cortical structures compared to MRI.
(a) Scatter plots showing the correlation between CT‐derived and MRI‐derived W‐score measurements, and (b) Bland‐Altman plots comparing CT‐ and MRI‐based methods. The analysis includes the following ROIs for both the left and right hemispheres: anterior lateral ventricle (Anterior LV), posterior lateral ventricle (Posterior LV), inferior lateral ventricle (Inferior LV), frontal CSF, temporal CSF, parietal CSF, and occipital CSF: MRI = magnetic resonance imaging; CT = computed tomography; LV = lateral ventricle; CSF = cerebrospinal fluid; R = right; L = left.
The Bland–Altman plots presented in Fig. 3b illustrate the differences between CT-based and MRI-derived brain W-score measurements across various ROIs. Across most ROIs, the mean differences between CT-based and MRI-derived measurements were close to zero, indicating minimal bias. However, specific ROIs, particularly those in the occipital and temporal regions, displayed greater variability in the differences, as evidenced by wider limits of agreement. This suggests that CT-based measurements may slightly underestimate or overestimate brain volumes in these areas compared to MRI.
Regional variations in W-scores among CU, MCI, and DAT
Figure 4a shows the distribution and statistical significance of W-scores for each ROI across the CU, MCI, and DAT groups in the CT-based method (Fig. 4a). All ROIs showed significant differences in W-scores between CU and both MCI and DAT groups (all, p value < 0.05), indicating consistent patterns of differentiation across the two methods. Significant differences in W-scores between MCI and DAT groups were observed across all ROIs in two methods (all, p value < 0.05). To provide a group-level overview of these patterns, we reorganized the boxplots by group (Supplementary Fig. 3a).
Box plots of W‐scores for CSF in ROI, showing regional variations in (a) the CU, MCI, and DAT and (b) FTD subtypes (bvFTD, nfvPPA, svPPA). All W-scores are derived from CT-based segmentation. An asterisk (*) indicates a significant difference from CU, and a dagger (†) indicates a significant difference from DAT, based on adjusted p < 0.05 after false discovery rate (FDR) correction. Abbreviations: CT = computed tomography; LV = lateral ventricle; CSF = cerebrospinal fluid; ROI = region of interest; CU = cognitively unimpaired; MCI = mild cognitive impairment; DAT = dementia of Alzheimer’s type; bvFTD = behavioural variant frontotemporal dementia; nfvPPA = nonfluent variant primary progressive aphasia; svPPA = semantic variant primary progressive aphasia.
Regional variations in W-scores among FTD subtypes
In Fig. 4b, we compared boxplots of various ROIs among CU, DAT, and FTD subtypes. In the left hemisphere, FTD subtypes showed significant differences compared to CU across all ROIs. When comparing FTD subtypes with DAT, bvFTD exhibited reduced volumes in the anterior LV, frontal, and temporal regions; svPPA demonstrated significant volume increases in the temporal region and inferior lateral ventricle; and nfvPPA showed lower volumes in the anterior LV, with a trend in the frontal region. In contrast, in the right hemisphere, bvFTD showed a similar trend to that observed in the left hemisphere, whereas the language variants (svPPA and nfvPPA) did not show significant differences from DAT. These complementary group-level patterns are also illustrated in Supplementary Fig. 3b. A representative stereotaxic atlas illustrating these 14 subdivisions is provided in Supplementary Fig. 4.
W-scores across CDR stages
W-scores tracked disease severity within diagnostic groups, showing progressive decreases across higher CDR stages in both AD and FTD cohorts (Supplementary Fig. 5). These resultsThese indicate that W-scores are sensitive to within-group variation in disease severity, complementing their utility in cross-sectional group differentiation.
Discussion
In this study, we developed a robust deep learning framework for CT‐based brain segmentation, and applied it to differentiate cognitive stages in AD as well as to distinguish distinct FTD subtypes. Our major findings were as follows. First, segmentation performance, evaluated using DSC, showed excellent agreement with the silver standard (3D T1 MRI segmentation), confirming the reliability of our CT‐based method. Second, there was a strong correlation between CT‐based measurements and 3D T1‐based measurements, with Bland–Altman analysis revealing minimal bias and narrow limits of agreement. Third, CT‐based measurements effectively distinguish between CU, MCI, and DAT, reflecting the algorithm’s ability to detect stage-specific changes in CSF volumes. Finally, FTD subtypes exhibited distinct, region-specific patterns of CSF volume enlargement and a relative hemispheric asymmetry in these changes, which is consistent with clinical phenotypes. Taken together, our findings demonstrate that CT‐based volumetric analysis not only replicates 3D T1 MRI measurements with high fidelity but also provides detailed, region-specific information that aligns with clinical phenotypes. These results underscore the superior clinical readiness of CT‐based assessment as a practical tool for early diagnosis and monitoring of neurodegenerative disorders, a capability that is critical for differential diagnosis.
Our first major finding was that segmentation performance exhibited excellent concordance with the silver standard (3D T1 MRI segmentation), affirming the robustness of our CT‐based method. Our detailed analysis revealed that the anterior and posterior lateral ventricles achieved DSCs above 0.93, while the inferior lateral ventricles and eCSF regions in the frontal, parietal, and temporal lobes also performed well. Notably, the occipital regions displayed the lowest DSC values; however, since occipital involvement is typically observed only in advanced stages of neurodegenerative diseases36 (except in posterior cortical atrophy), this limitation is less critical for early diagnosis. Taken together, our findings suggest that our CT‐based approach reliably approximates 3D T1-based measurements, offering significant clinical utility due to CT’s superior accessibility and readiness for routine neurodegenerative assessment.
Segmentation performance was slightly lower in the inferior LV and occipital CSF regions, likely reflecting both anatomical complexity (e.g., smaller structures, partial volume effects) and reduced image contrast. Furthermore, the use of a silver-standard reference may have introduced variability, as automated labels are not fully accurate across all participants. These factors together likely contributed to the consistent lowering of Dice scores across regions. Importantly, similar regional patterns of reduced segmentation performance have been reported in MRI-based studies, suggesting that these challenges are not modality-specific but rather reflect intrinsic anatomical and methodological factors.
Our second major finding was that CT‐based measurements and 3D T1‐based measurements demonstrated a very strong linear correlation across multiple ROIs. As shown in Fig. 3, scatter plots revealed correlation coefficients ranging from 0.822 to 0.996, with lateral ventricular ROIs—specifically, the left and right anterior, posterior, and inferior lateral ventricles—exhibiting the highest correlations (r > 0.98). Bland–Altman analysis also confirmed this strong agreement by showing minimal bias and narrow limits of agreement between the two modalities. These results are consistent with previous studies such as Srikrishna et al.16 Moreover, while previous studies primarily focused on whole-brain segmentation by distinguishing CSF, gray matter, and white matter16,37, our approach measures areas with pronounced changes in CSF volume—such as ventricular enlargement and sulcal widening that are carefully evaluated by radiologists—and analyzes the resulting region-specific CSF volume enlargement patterns. Additionally, our method provides detailed lobe-specific and hippocampal CSF volume assessments, which not only facilitate early AD diagnosis—given that the hippocampus is one of the earliest affected regions38—but also enable differentiation of FTD subtypes based on distinct lobe-specific CSF volume change patterns.
Our third major finding was that CT‐based measurements effectively distinguish between CU, MCI, and DAT, reflecting stage-specific changes in CSF volumes. Although we initially had concerns that emphasizing pronounced changes in CSF volume might compromise sensitivity to subtle structural changes, our findings demonstrated significant differences in W-scores across all ROIs at the MCI status relative to CU, with further distinctions observed between DAT and MCI. W-scores progressively decreased from CU to DAT, suggesting increased CSF volume associated with advanced tissue loss. These results underscore the sensitivity of our CT‐based method for tracking structural brain changes across the disease continuum.
Our final major finding was that FTD subtypes exhibited distinct, region-specific patterns of CSF volume enlargement that indirectly reflect tissue loss consistent with clinical phenotypes. In particular, bvFTD demonstrated prominent CSF volume increases in bilateral frontal (enlargements in the anterior ventricles and frontal eCSF) and temporal (enlargements in the inferior lateral ventricle and temporal eCSF) regions. In contrast, nfvPPA was characterized primarily by increased CSF volume in the left frontal region, while svPPA showed marked volume enlargement in the left temporal region. Notably, for the language variants, several ROIs in the right hemisphere—especially in posterior regions (including the posterior ventricle, parietal, and occipital eCSF areas)—did not differ significantly from CU, highlighting selective vulnerability. These findings underscore that our CT-based regional volumetric analysis could effectively differentiate between DAT and FTD subtypes—a capability that is critical for differential diagnosis. Moreover, our CDR stage-wise analyses further demonstrated that W-scores captured disease severity within diagnostic groups, with progressive declines observed across higher stages. Importantly, regional trajectories differed between AD and FTD, consistent with their distinct underlying pathologies. These findings underscore the potential of CT-based W-scores as not only cross-sectional biomarkers but also as practical indices of clinical staging.
Our study leverages CSF volume differences in CT images across multiple ROIs to achieve robust, reproducible segmentation and quantification over diverse imaging conditions, as well as across cognitive stages and FTD subtypes. However, there were several limitations. First, the inherent lower soft-tissue contrast of CT compared to MRI limits the accurate delineation of fine cortical structures, leading to reduced performance in regions such as the occipital lobe. Second, although our CT-based method provides detailed region-specific CSF volume assessments, it may be less sensitive to subtle early-stage changes compared to advanced MRI techniques. Also, the limited sample size for certain FTD subtypes, particularly nfvPPA, may constrain the generalizability and statistical power of our findings. Third, while robustness was supported by stratified fivefold cross-validation within the primary dataset and further confirmed using an independent cohort of MRI scans from the ADNI study, true external validation with independent CT datasets was not performed. Establishing generalizability in such CT cohorts will be an essential step to confirm the clinical applicability of our framework. Fourth, our cohort was recruited from a tertiary referral center, leading to potential sampling bias such as an overrepresentation of cognitively impaired individuals and an FTD group with overlapping age distribution with AD. Moreover, MMSE may underestimate disease severity in some FTD subtypes. While this heterogeneity limits strict representativeness, it underscores the pipeline’s robustness in real-world clinical settings where such variability is common. Finally, although the W score-based pipeline shows promise for clinical translation, it has not yet been directly compared with clinician visual ratings and should be considered a complementary tool requiring further validation and workflow optimization for routine use in dementia clinics. In the future, it will also be important to extend this approach beyond group-level validation to evaluate its potential utility for single-subject classification, for example, distinguishing AD from FTD in individual patients using CT scans. Nonetheless, our findings provide valuable insights into the potential of CT‐based segmentation for capturing clinically relevant patterns of CSF volume enlargement and differentiating between cognitive stages and FTD subtypes, supporting its utility as a practical tool for early diagnosis, monitoring disease progression, and aiding differential diagnosis of dementia.
Conclusion
In summary, our CT‐based volumetric analysis not only replicates 3D T1 MRI measurements with high fidelity but also provides detailed insights by capturing stage‐specific changes in CSF volumes that effectively differentiate cognitive stages in AD, as well as region‐specific CSF volume enlargement patterns that enable the distinction of FTD subtypes. These findings underscore the clinical utility of CT‐based assessments as a practical, accessible tool for early diagnosis, monitoring, and differential diagnosis of neurodegenerative disorders such as AD and FTD subtypes.
Data availability
The dataset(s) supporting the conclusions of this article is(are) included within the article (and its additional file(s)).
References
Chetelat, G. et al. Relationship between atrophy and beta-amyloid deposition in Alzheimer disease. Ann. Neurol. 67, 317–324. https://doi.org/10.1002/ana.21955 (2010).
McKhann, G. M. et al. The diagnosis of dementia due to Alzheimer’s disease: recommendations from the National Institute on aging-Alzheimer’s association workgroups on diagnostic guidelines for Alzheimer’s disease. Alzheimers Dement 7, 263–269. https://doi.org/10.1016/j.jalz.2011.03.005 (2011).
Pini, L. et al. Brain atrophy in Alzheimer’s disease and aging. Ageing Res. Rev. 30, 25–48. https://doi.org/10.1016/j.arr.2016.01.002 (2016).
Wardlaw, J. M. et al. Neuroimaging standards for research into small vessel disease and its contribution to ageing and neurodegeneration. Lancet Neurol. 12, 822–838. https://doi.org/10.1016/S1474-4422(13)70124-8 (2013).
Fathy, Y. Y. et al. Differential insular cortex sub-regional atrophy in neurodegenerative diseases: A systematic review and meta-analysis. Brain Imaging Behav. 14, 2799–2816. https://doi.org/10.1007/s11682-019-00099-3 (2020).
Harper, L. et al. Patterns of atrophy in pathologically confirmed dementias: A voxelwise analysis. J. Neurol. Neurosurg. Psychiatry 88, 908–916. https://doi.org/10.1136/jnnp-2016-314978 (2017).
Scheltens, P. et al. Atrophy of medial temporal lobes on MRI in “probable” Alzheimer’s disease and normal ageing: Diagnostic value and neuropsychological correlates. J. Neurol. Neurosurg. Psychiatry 55, 967–972. https://doi.org/10.1136/jnnp.55.10.967 (1992).
Jack, C. R. Jr., Petersen, R. C., O’Brien, P. C. & Tangalos, E. G. MR-based hippocampal volumetry in the diagnosis of Alzheimer’s disease. Neurology 42, 183–188. https://doi.org/10.1212/wnl.42.1.183 (1992).
Orellana, C. et al. Measuring global brain atrophy with the brain volume/cerebrospinal fluid index: Normative values cut-offs and clinical associations. Neurodegener. Dis. 16, 77–86. https://doi.org/10.1159/000442443 (2016).
Wei, M. et al. A new age-related cutoff of medial temporal atrophy scale on MRI improving the diagnostic accuracy of neurodegeneration due to Alzheimer’s disease in a Chinese population. BMC Geriatr. 19, 59. https://doi.org/10.1186/s12877-019-1072-8 (2019).
de Leon, M. J., George, A. E., Stylopoulos, L. A., Smith, G. & Miller, D. C. Early marker for Alzheimer’s disease: The atrophic hippocampus. Lancet 2, 672–673. https://doi.org/10.1016/s0140-6736(89)90911-2 (1989).
Wattjes, M. P. et al. Diagnostic imaging of patients in a memory clinic: Comparison of MR imaging and 64-detector row CT. Radiology 253, 174–183. https://doi.org/10.1148/radiol.2531082262 (2009).
Rossi, R., Joachim, C., Smith, A. D. & Frisoni, G. B. The CT-based radial width of the temporal horn: Pathological validation in AD without cerebrovascular disease. Int. J. Geriatr. Psychiatry 19, 570–574. https://doi.org/10.1002/gps.1132 (2004).
Wahlund, L. O. et al. Imaging biomarkers of dementia: Recommended visual rating scales with teaching cases. Insights Imaging 8, 79–90. https://doi.org/10.1007/s13244-016-0521-6 (2017).
Rorden, C., Bonilha, L., Fridriksson, J., Bender, B. & Karnath, H. O. Age-specific CT and MRI templates for spatial normalization. Neuroimage 61, 957–965. https://doi.org/10.1016/j.neuroimage.2012.03.020 (2012).
Srikrishna, M. et al. Deep learning from MRI-derived labels enables automatic brain tissue classification on human brain CT. Neuroimage 244, 118606. https://doi.org/10.1016/j.neuroimage.2021.118606 (2021).
Aguilar, C. et al. Automated CT-based segmentation and quantification of total intracranial volume. Eur. Radiol. 25, 3151–3160. https://doi.org/10.1007/s00330-015-3747-7 (2015).
Srikrishna, M. et al. CT-based volumetric measures obtained through deep learning: Association with biomarkers of neurodegeneration. Alzheimers Dement 20, 629–640. https://doi.org/10.1002/alz.13445 (2024).
Braak, H. & Braak, E. Staging of Alzheimer’s disease-related neurofibrillary changes. Neurobiol. Aging 16(271), 278. https://doi.org/10.1016/0197-4580(95)00021-6 (1995) (discussion 278-284).
Rascovsky, K. et al. Sensitivity of revised diagnostic criteria for the behavioural variant of frontotemporal dementia. Brain J. Neurol. 134, 2456–2477. https://doi.org/10.1093/brain/awr179 (2011).
Botha, H. et al. Classification and clinicoradiologic features of primary progressive aphasia (PPA) and apraxia of speech. Cortex 69, 220–236. https://doi.org/10.1016/j.cortex.2015.05.013 (2015).
Gorno-Tempini, M. L. et al. Classification of primary progressive aphasia and its variants. Neurology 76, 1006–1014. https://doi.org/10.1212/WNL.0b013e31821103e6 (2011).
Albert, M. S. et al. The diagnosis of mild cognitive impairment due to Alzheimer’s disease: Recommendations from the National Institute on aging-Alzheimer’s association workgroups on diagnostic guidelines for Alzheimer’s disease. Alzheimers Dement 7, 270–279. https://doi.org/10.1016/j.jalz.2011.03.008 (2011).
Ahn, H. J. et al. Seoul neuropsychological screening battery-dementia version (SNSB-D): A useful tool for assessing and monitoring cognitive impairments in dementia patients. J. Korean Med. Sci. 25, 1071–1076. https://doi.org/10.3346/jkms.2010.25.7.1071 (2010).
Kang, Y., Na, D. & Hahn, S. Seoul neuropsychological screening battery. Incheon: Human brain research & consulting co (2003).
Hahn, A. et al. The preclinical amyloid sensitive composite to determine subtle cognitive differences in preclinical Alzheimer’s disease. Sci. Rep. 10, 13583. https://doi.org/10.1038/s41598-020-70386-3 (2020).
Won, C. W., Rho, Y. G., SunWoo, D. & Lee, Y. S. The validity and reliability of korean instrumental activities of daily living(K-IADL) scale. Ann. Geriatr. Med. Res. 6, 273–280 (2002).
Billot, B. et al. SynthSeg: Segmentation of brain MRI scans of any contrast and resolution without retraining. Med. Image Anal. 86, 102789. https://doi.org/10.1016/j.media.2023.102789 (2023).
Billot, B. et al. Robust machine learning segmentation for large-scale analysis of heterogeneous clinical brain MRI datasets. Proc. Natl. Acad. Sci. U S A 120, e2216399120. https://doi.org/10.1073/pnas.2216399120 (2023).
Collins, D. L., Holmes, C. J., Peters, T. M. & Evans, A. C. Automatic 3-D model-based neuroanatomical segmentation. Hum. Brain Mapp. https://doi.org/10.1002/hbm.460030304 (1995).
Collins DL, Z. A., Baaré WFC, Evans AC. ANIMAL+INSECT: Improved Cortical Structure Segmentation. Information Processing in Medical Imaging, 210-223 (1999). https://doi.org/10.1007/3-540-48714-X_16
Avants, B. B., Epstein, C. L., Grossman, M. & Gee, J. C. Symmetric diffeomorphic image registration with cross-correlation: Evaluating automated labeling of elderly and neurodegenerative brain. Med. Image Anal. 12, 26–41. https://doi.org/10.1016/j.media.2007.06.004 (2008).
La Joie, R. et al. Region-specific hierarchy between atrophy, hypometabolism, and beta-amyloid (Abeta) load in Alzheimer’s disease dementia. J. Neurosci. 32, 16265–16273. https://doi.org/10.1523/JNEUROSCI.2170-12.2012 (2012).
Isensee, F., Jaeger, P. F., Kohl, S. A. A., Petersen, J. & Maier-Hein, K. H. nnU-Net: A self-configuring method for deep learning-based biomedical image segmentation. Nat. Methods 18, 203–211. https://doi.org/10.1038/s41592-020-01008-z (2021).
Antonelli, M. et al. The Medical Segmentation Decathlon. Nat. Commun. 13, 4128. https://doi.org/10.1038/s41467-022-30695-9 (2022).
Katsumi, Y. et al. Dissociable spatial topography of cortical atrophy in early-onset and late-onset Alzheimer’s disease: A head-to-head comparison of the LEADS and ADNI cohorts. Alzheimers Dement 21, e14489. https://doi.org/10.1002/alz.14489 (2025).
Leung, K. K. et al. Robust atrophy rate measurement in Alzheimer’s disease using multi-site serial MRI: tissue-specific intensity normalization and parameter selection. Neuroimage 50, 516–523. https://doi.org/10.1016/j.neuroimage.2009.12.059 (2010).
Lehmann, M. et al. Cortical thickness and voxel-based morphometry in posterior cortical atrophy and typical Alzheimer’s disease. Neurobiol. Aging 32, 1466–1476. https://doi.org/10.1016/j.neurobiolaging.2009.08.017 (2011).
Acknowledgements
We gratefully acknowledge the dedication of all participants and the partnership team members whose efforts continue to make the K‐ROAD study possible. A complete list of the members of the K-ROAD study groups appears in Supplementary data 2. Data used in preparation of this article were obtained from the Alzheimer’s Disease Neuroimaging Initiative (ADNI) database (adni.loni.usc.edu). The investigators within the ADNI contributed to the design and implementation of ADNI and/or provided data but did not participate in the analysis or writing of this report. A complete listing of ADNI investigators can be found at https://adni.loni.usc.edu/wp-content/uploads/how_to_apply/ADNI_Acknowledgement_List.pdf
Funding
This research was supported by a grant of the Korea Dementia Research Project through the Korea Dementia Research Center (KDRC), funded by the Ministry of Health & Welfare and the Ministry of Science and ICT, Republic of Korea (RS-2020-KH106434); by the Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI), funded by the Ministry of Health & Welfare, Republic of Korea (RS-2025-02223212); by the National Research Foundation of Korea (NRF) grant funded by the Korea government (MSIT) (RS-2019-NR040057); by the Institute of Information & Communications Technology Planning & Evaluation (IITP) grant funded by the Korea government (MSIT) (RS-2021-II212068, Artificial Intelligence Innovation Hub); by the Future Medicine 20*30 Project of Samsung Medical Center (SMX1250081); and by the Korea National Institute of Health research project (2024-ER1003-01).
Author information
Authors and Affiliations
Contributions
**Seongbeom Park:** conceptualization, methodology, writing—original draft. **Kyoungmin Kim:** conceptualization, writing—original draft. **Kyoungyoon Lim:** methodology, visualization. **Duk L. Na:** data curation, supervision. **Hee Jin Kim:** data curation, review. **Hyemin Jang:** data curation. **Jun Pyo Kim:** review, validation. **Sung Hoon Kang:** data curation, investigation. **Jihwan Yun:** data curation. **Min Young Chun:** data curation. **Sang Won Seo:** conceptualization, writing—review and editing, funding acquisition **. Kichang Kwak:** conceptualization, methodology, writing—review and editing.
Corresponding authors
Ethics declarations
Competing interests
Sang Won Seo is co-founder of BeauBrain Healthcare Inc. Seongbom Park, Kyoung Yoon Lim, and Kichang Kwak were employed by BeauBrain Healthcare Inc. The other authors declare that they have no competing interests.
Ethics approval
The institutional review board of Samsung Medical Center (No. 2021-02-135) approved this study.
Informed consent
All participants provided informed consent to participate in the study, and the data were collected in accordance with the Declaration of Helsinki.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Park, S., Kim, K., Lim, K.Y. et al. Computed tomography-based nnU-Net for region-specific brain structural changes across the alzheimer’s continuum and frontotemporal dementia subtypes. Sci Rep 15, 42597 (2025). https://doi.org/10.1038/s41598-025-26604-x
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-26604-x