Abstract
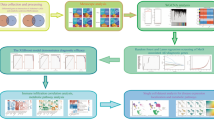
Type A aortic dissection (TAAD) is a vascular disease with high mortality; however, the role of gene methylation in its pathogenesis has received little attention. This study aimed to identify candidate markers of TAAD by integrating methylation and transcriptome sequencing analyses. Aortic tissue samples from five TAAD cases and five controls were sequenced on Illumina Hiseq sequencing platform and Infinium Methylation EPIC BeadChip microarray. A series of bioinformatics analyses and machine learning algorithms were used to identify key methylated genes from the differentially expressed genes and differentially methylated sites between TAAD and controls. An overexpression vector of ZC3H12A was constructed, and human vascular smooth muscle cells (HVSMCs) were transfected with the vector to explore the effects of key methylated genes on cell proliferation, migration, and phenotypic switch. Differential analysis and integration of gene expression and methylation levels between TAAD and control samples suggested 239 differentially methylated genes, mainly involved in nicotinamide nucleotide biosynthetic and metabolic processes. By applying protein-to-protein network and machine learning algorithms, we finally identified three methylated genes, including ZC3H12A, IRAK2, and CCL5, which could be used as potential markers of TAAD. Both correlation analysis and experimental validation results indicated that expression levels of these genes were significantly negatively regulated by their methylation levels. Among them, ZC3H12A was confirmed to significantly promote the proliferation and migration in HVSMCs in vitro, while inhibiting their phenotypic transformation. Three methylated genes were identified as potential diagnostic markers for TAAD. Among them, ZC3H12A might contribute to disease progression.
Similar content being viewed by others
Introduction
Aortic dissection (AD) is a life-threatening and aggressive vascular disease, with a high mortality rate owing to severe complications, such as aortic rupture, aortic regurgitation, lethal malperfusion syndrome, heart failure, and stroke1,2. The ascending aorta is involved in approximately 60–70% of the patients with AD, who are classified as Stanford type A AD (TAAD)3. Acute TAAD has a mortality rate approaching 40–50% within 48 h, and about 90% of the patients will die within three months if not promptly treated4,5. Over the past decade, emergency surgical intervention has significantly improved survival in patients with TAAD, yet postoperative hospitalization mortality remains as high as 22%6. Early clinical symptoms of TAAD may resemble those of other diseases, such as coronary syndromes, pulmonary embolism, and pneumothorax, often leading to the delayed diagnosis; therefore, early diagnosis, prompt treatment, and close follow-up are critical for patient survival7. Under the current situation, there is a lack of effective biomarkers for early and accurate diagnosis, as well as scientific stratification of high-risk populations, resulting in many patients experiencing delayed treatment and extremely poor prognosis. Although some TAAD risk genes and serum biomarkers have been identified in previous studies8,9, their value in clinical translational applications is limited, making it difficult to effectively guide disease prevention and treatment. Although current research has improved the diagnosis, treatment options, and clinical outcomes, the specific pathogenesis of TAAD is still unclear. Therefore, understanding the molecular mechanisms of TAAD may provide new therapeutic strategies to prevent the progression of aortic diseases.
Epigenetic regulation affect gene expression and produce inherited phenotypic changes without altering the DNA sequence10. Epigenetic modifications have been shown to contribute to the onset and progression of cardiovascular diseases11. DNA methylation is a major epigenetic mechanism that is maintained through the combined actions of methyltransferase and demethylase12. In cardiovascular diseases, including heart failure13, atherosclerosis14, and coronary heart disease15, external risk factors can influence downstream molecular events by altering DNA methylation levels. Recently, several studies have focused on DNA methylation patterns in AD. For instance, HOX gene regulation has been shown to be associated with changes in its methylation patterns in thoracic AD16. In addition, DNA methylation was reported to mediate the relationship between AD and inflammatory vascular remodeling17. However, altered gene methylation patterns in TAAD remain inconclusive, and only few studies have systematically examined the relationship between altered methylation levels and gene expression profiles.
The integration of methylation and transcriptome sequencing analyses provides a novel strategy for identifying diagnostic markers of diseases18; however, this approach has not yet been applied in TAAD. Therefore, in this study, we carried out methylation and transcriptome sequencing analyses using aortic vascular tissue samples, and the key methylated genes were identified as potential diagnostic markers of TAAD via integrative bioinformatics-based analyses. Subsequently, the effects of the key methylated genes on disease progression and phenotypic transformation in TAAD were experimentally explored. The epigenetic regulatory mechanisms revealed in this study can provide the basis for the development of new therapeutic strategies for patients with TAAD.
Methods
Study participants
Patients with TAAD enrolled in this study were diagnosed according to the guidelines for diagnosing thoracic aortic disease19. After emergency admission, preoperative aortic computed tomographic angiography (CTA) was carried out, and the involvement of the ascending aorta and aortic arch was intraoperatively confirmed. All subjects in the experimental group had their first attack and underwent surgical treatment within two weeks of symptom onset. No family history of death due to aneurysms was reported. In addition, ascending aortic vessel wall tissues from donor hearts for heart transplantation were included in the control group after normal pathology examination. Exclusion criteria included: a. Marfan syndrome, b. Ehlers-Danlos syndrome, c. Lloyd-Dietz syndrome, d. Turners syndrome, e. bicuspid aortic valve disease, f. aortic coarctation, arteritis, ascending aortic aneurysm, and other aorta-related diseases, g. Behcet’s disease, h. familial thoracoabdominal aortic aneurysm syndrome, i. trauma-induced aortic dissection, j. dissection during pregnancy, k. iatrogenic aortic dissection, l. concurrent heart failure and acute myocardial infarction, m. congenital heart disease and congenital vascular dysplasia, and n. cancer. The clinical characteristics of all participants were collected. After age and gender matching, this study finally included five TAAD patients and five control subjects, divided into the TAAD group (n = 5) and the control group (n = 5). This study complied with the Helsinki Declaration and was approved by the ethics committee of Tianjin Chest Hospital (2023KY-028–01). All participants signed informed consent.
Sample collection
Aortic vascular tissues were excised intraoperatively from the ascending aorta tissue samples in patients with TAAD. Tissue collection from the arterial wall tear site was avoided, as the aortic wall structure at this location is fully disrupted, and the excess components, including fat, were rinsed off with normal saline. One part of the tissue was immediately frozen in liquid nitrogen and stored in a − 80 °C ultra-low temperature freezer. The other part of the tissue was fixed with 4% neutral formaldehyde in a cool and ventilated place, embedded in paraffin, and then stored for further use.
Transcriptome sequencing
In this study, the high-throughput sequencing of samples was performed using the double-end sequencing mode of the Illumina Hiseq sequencing platform. Skewer (v0.2.2) software was used to dynamically remove splice fragments and low-quality fragments from the 3’ end of the sequencing data20, and FastQC (v0.11.5) software was used for quality control of the preprocessed data21. Then, the preprocessed sequences were aligned with the reference genome of the sequenced species using STAR (2.5.3a) software22. After the transcripts were filtered and assembled using StringTie (v1.3.1c) software, Gffcompare (0.9.9) was applied to compare and annotate the assembled transcripts with the known gene reference genome position information23.
Methylation sequencing
The Illumina Infinium Methylation EPIC BeadChip microarray was used to examine the methylation status of CpG sites across the human genome. After obtaining the raw IDAT file, the methylation sites of the probes with detected P value > 0.01 were filtered out. The beta-mixture quantile dilation method was used to normalize the beta values24.
Bioinformatics analyses
Screening of differentially expressed genes (DEGs) and differential methylation sites (DMSs)
Based on the annotation file, the gene IDs were converted to symbols to obtain the expression matrix of the transcriptome. Analysis of differential gene expression levels between TAAD and normal tissue samples was performed using linear regression and empirical Bayesian methods provided in the limma package (Version 3.10.3)25 installed on R Studio. In differential gene screening, the setting of the log2fold change (FC) threshold is of vital importance, with commonly adopted thresholds being 1, 0.585, and 0.263. Since the number of DEGs was insufficient under the initial threshold, after taking into account experiences from relevant literature and making a comprehensive consideration, we reduced the log2FC threshold to below 0.263 to include more genes26. The statistically significant DEGs were selected at |log2FC|> 0.263 and a P value < 0.05. Similarly, IDs of methylated sites were converted to symbols, followed by screening for DMSs using the limma package. Statistical significance was set at a P value < 0.05.
Identifying methylation sites driving gene expression
Based on the identified DEGs and DMSs, the up- and downregulated DEGs were labeled as up and down, respectively, whereas the up- and downregulated DMSs were labeled as hyper and hypo, respectively. In order to find methylation sites that can drive gene expression, DEGs and DMSs were intersected to screen for hyper-down and hypo-up pairs of gene and methylation site relationships as differentially methylated genes (DMGs) for subsequent analysis.
Functional and pathway enrichment analyses
Gene ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG)-based pathway enrichment analyses were performed on DMGs using DAVID (version 6.8)27,28. In addition, the R package cluster Profiler was employed to perform Gene Set Enrichment Analysis (GSEA) on DEGs related to GO, aiming to identify the signaling pathways associated with DEGs. Terms with a P value < 0.05 were selected as significantly enriched functions and pathways.
Construction of a protein-to-protein interaction (PPI) network
The STRING (Version 11.5) database29 was used to construct a PPI network for DMGs involved in the identified key functions and pathways, and the results were visualized using Cytoscape (Version 3.8.2)30. After topological analysis of the closeness, degree, edge percolated component (EPC), maximal clique centrality (MCC), maximum neighborhood component (MNC), and radial stress for each node in the network, the top 20 genes in each metric were selected, and the intersection was subsequently used to identify the candidate genes.
Screening of key methylated genes
Based on the candidate genes, the support vector machine-recursive feature elimination (SVM-RFE)31 was used to obtain the optimal variables by cross-validation. Moreover, the least absolute shrinkage selection operator (LASSO) regression model was constructed using the R glmnet package (Version 4.1.3)32 to screen for significantly methylated genes. The overlapping methylated genes obtained from these two algorithms were identified as key methylated genes. Finally, Pearson’s correlation coefficients were calculated to assess the relationship between key methylation sites and their corresponding genes.
Quantitative real-time polymerase chain reaction (qRT-PCR)
The aortic wall tissue was ground and added to RNAiso Plus (Trizol, TaKaRa, 9109) for total RNA extraction. Then, the RNA purity was determined, followed by the estimation of sample concentration and quality using a microplate reader (BioTek, Epoch). After the reverse transcription reaction, the Power Up™ SYBR™ Green mix (Thermo, A25742) was added to the samples, and the PCR was carried out to detect the mRNA expression levels of the target genes using a PCR machine (ABI, 7900HT FAST). Primer sequences are shown in Table 1. The data was analyzed using the 2−ΔΔCt algorithm by normalizing to glyceraldehyde 3-phosphate dehydrogenase (GAPDH) expression.
Cell culture
Human vascular smooth muscle cells (HVSMCs) were purchased from BeNa Culture Collection (BNCC) and cultured in primary smooth muscle cell medium (PriMed-iCELL-004, BNCC, BNCC360626) containing 10% fetal bovine serum (FBS; Gibco, 10099141) and 1% penicillin–streptomycin (P/S; Gibco, 15140-122) under normal culture conditions. Furthermore, a methylase inhibitor, 5-aza-2'-deoxycytidine (Sigma, A3656-5MG) was added at 5 μM to the treatment group to inhibit cell methylation.
ZC3H12A overexpression vector construction
To construct a ZC3H12A overexpression model, the sequence of ZC3H12A was ligated into the linearized vector pCDH-CMV-MCS-EF1-copGFP-T2A-Puro. After electrophoretic recovery and ligand transformation, the digestion products were added to DH5α recipient cells and cultured overnight in plates containing ampicillin. Finally, the plasmids were extracted and verified by single enzyme digestion.
Cell transfection
The cultured HVSMCs were divided into three groups; mock, pCDH-ZC3H12A-NC (OE-NC), and pCDH-ZC3H12A (OE-ZC3H12A), in which the 293 T cells were transfected the plasmid vector after lentiviral packaging. During transfection, cells were incubated with fresh medium containing lentiviral suspension for 24 h at 37 °C. Subsequently, transfection was terminated by replacing the virus-containing medium with a fresh one.
Western blot analysis
Briefly, proteins were extracted, quantified, and separated on sodium dodecyl sulfate–polyacrylamide gel. Subsequently, equivalent quantities of proteins were transferred to a polyvinylidene fluoride membrane (Millipore, IPVH00010) on ice. After blocking the membranes with 5% non-fat milk powder, they were incubated overnight with an anti-ZC3H12A primary antibody (1:2000, ray biotech, 144-62425-50). The horseradish peroxidase (HRP)-conjugated Affinipure goat anti-rabbit IgG (H + L) secondary antibody (1:2000, proteintech, SA00001-2) was then added at room temperature for 2 h. Finally, the target proteins were visualized using a chemiluminescence imaging system (Tanon, 4600) and normalized to the expression level of GAPDH (1:3000, ShareBio, SB-AB0037).
Cell counting kit-8 (CCK8)
HVSMCs suspension (100 μL/well) was inoculated into 96-well plates and cultured for 24, 48, or 72 h. Then, 10 μL of the CCK-8 solution (Beyotime, C0038) was added to each well, followed by incubation for another 4 h. The absorbance of each well was measured at 450 nm using amicroplate reader to estimate the cell viability.
Transwellassay
The cells were digested and adjusted to a density of 5 × 105 cells/mL. Afterwards, 200 μL of cell suspension was inoculated in the upper chamber, whereas 800 μL of complete medium containing 20% FBS was added to the lower chamber for routine incubation. After 48 h, the chambers were removed for cleaning and fixation, and stained using crystal violet staining. Finally, the cell migration ability was observed and recorded under a microscope.
Statistical analysis
Graphpad prism 9.0.5 (Graphpad Software, San Diego, CA) was used for statistical analysis of the data. The grayscale values of the western blot strips were quantified using Image J. All experiments were carried out in at least three replicates, and all results are presented as the mean ± standard deviation33. For normally distributed data with equal variances, comparisons between two groups were performed using the unpaired t-test, whereas comparisons among multiple groups were conducted using one-way analysis of variance (ANOVA). In cases of non-normality or heteroscedasticity, the Kruskal–Wallis test, followed by multiple comparisons analysis, was conducted to assess the differences among groups. A P value < 0.05 was considered statistically significant.
Results
Screening for DEGs and DMSs between TAAD and control groups
Comparison of the gene expression profiles between the TAAD and control groups revealed 2464 upregulated (up) and 3441 downregulated (down) DEGs (Fig. 1A,B) with a threshold of |log2FC|> 0.263 and P value < 0.05. Moreover, comparison of the methylation sites between the two groups showed 1407 upregulated (hyper) and 3570 downregulated (hypo) DMSs (Fig. 1C,D). To further screen for methylation sites that could drive gene expression, hyper-down and hypo-up pairs were selected from the overlap between DEGs and DMSs. Therefore, a total of 239 DMGs were selected for further analyses (Fig. 1E).
Screening for DEGs and DMSs in TAAD and control groups to select DMGs. A, B: Volcano plot (A) and heatmap (B) showing 2464 upregulated (up) and 3441 downregulated (down) DEGs. C, D: Volcano plot (C) and heatmap (D) showing 1407 upregulated (hyper) and 3570 downregulated (hypo) DMSs. (E) Venn diagram displaying the intersection of DEGs and DMSs.
Enrichment analysis of DMGs
To investigate the potential functions and pathways potentially associated with the identified DMGs, GO and KEGG enrichment analyses were carried out. GO results suggested that DMGs were mainly enriched in functions related tonicotinamide nucleotide biosynthetic processes, nicotinamide nucleotide metabolic processes, ligand-gated ion channel activity involved in regulation of presynaptic membrane potential, regulation of smooth muscle cell–matrix adhesion, and regulation of aortic smooth muscle cell differentiation (Fig. 2A). Among the genes involved in the biosynthetic and metabolic pathways of nicotinamide nucleotides, ENO1 (log2FC = 1.0427, P value = 0.0214), NAMPT (log2FC = 1.6212, P value = 0.0065), and HK2 (log2FC = 1.8229, P value = 0.0141) all exhibit high expression levels in the disease group (Supplementary Figure 1 A-C). Their regulatory relationship with TAAD is particularly crucial, with their underlying mechanisms encompassing energy metabolism, methylation regulation, and dysfunction of vascular smooth muscle cells (VSMCs).
GO function (A) and KEGG pathway (B) enrichment analyses of DMGs.
Among the eight enriched KEGG pathways (Fig. 2B), Hypoxia-inducible factor-1 (HIF-1) signaling pathway (including ENO1, PFKFB3, SERPINE1, and HK2 genes) was found to induce phenotype switch of VSMCs34. Therefore, DMGs involved in the aforementioned functions and HIF-1 signaling pathway genes were integrated for subsequent analysis.
Through GSEA analysis, we found that the nicotinamide nucleotide synthesis and metabolism pathway did not exhibit statistically significant differences (Supplementary Figure 1D). However, from a trend perspective, this pathway demonstrated a downregulated characteristic. Although this result did not reach statistical significance, it provides crucial clues for us to further understand the relevant biological processes.
Construction of the PPI network to screen for candidate genes
DMGs involved in key GO functions and KEGG pathways were used to construct a PPI network (Fig. 3A), and metrics, including closeness, degree, EPC, MCC, MNC, and radial stress for each node in the network were estimated. Based on the intersection of the top 20 nodes of each metric, a total of 13 node genes (including ZC3H12A, IRAK2, CCL5, TLR9, GBP1, SERPINE1, TNFRSF10B, IFNA1, HGF, JAK3, IL11, NOD1, and CXCL6) were identified as candidate genes (Fig. 3B).
Constructing a PPI network to screen for candidate genes. (A) DMGs involved in the key GO functions and KEGG pathways were used to construct a PPI network. (B) The intersection of the top 20 nodes of each metric.
Identification of key methylated genes using machine learning algorithms
By applying SVM-RFE, genes with optimal variables were obtained through cross-validation, and the top five genes (ZC3H12A, IRAK2, CCL5, TLR9, and GBP1) were selected. Furthermore, based on the variation features of variable coefficients, the model with excellent performance and with the least number of variables was screened by the cross-validation of LASSO regressions. Ultimately, the combination of five genes (ZC3H12A, IRAK2, CCL5, HGF, and IL11) was considered the optimal model (Fig. 4A). Based on the intersection of five genes from SVM-RFE and five genes from LASSO, ZC3H12A, IRAK2, and CCL5 were identified as key methylated genes (Fig. 4B). The correlation scatter plots suggested significant negative correlations between the methylation level and transcriptome expression level for these three methylated genes (Fig. 4C).
Identification of key methylated genes using SVM-RFE and LASSO algorithms. (A) LASSO coefficient distribution (top panel) and its likelihood deviation (bottom panel). (B) Venn diagram showing the intersection of genes selected from the SVM-RFE and LASSO algorithms. (C) The correlations between methylation levels and transcriptome expression levels of the three key methylated genes.
Experimental validation of the key genes at both expression and methylation levels
qRT-PCR of the collected tissue samples was used to verify the mRNA expression levels of three key methylated genes. The experimental results were consistent with the findings of transcriptome sequencing analysis, suggesting a significant increase in ZC3H12A (6.05 ± 3.50 vs. 1.03 ± 0.03, P = 0.033) and IRAK2 (5.69 ± 2.94 vs. 1.04 ± 0.03, P = 0.024), while decrease in CCL5 expression (0.49 ± 0.36 vs. 1.05 ± 0.05, P = 0.009) expression in the TAAD group, compared to that in the controls (Fig. 5A). To further investigate whether the methylation level of these genes might affect their expression, the methylase inhibitor, 5-aza-2'-deoxycytidine was added to HVSMCs. When cellular methylation was inhibited, the expression levels of ZC3H12A (2.18 ± 0.30 vs. 1.00 ± 0.06, P = 0.017), IRAK2 (3.01 ± 0.14 vs. 1.00 ± 0.07, P < 0.001), and CCL5 (2.81 ± 0.48 vs. 1.02 ± 0.14, P = 0.023) significantly increased (Fig. 5B), indicating that the methylation of these genes could significantly inhibit their expression.
Experimental validation of the key genes at expression and methylation levels. (A) Validation of the mRNA expression levels of ZC3H12A, IRAK2, and CCL5 in tissue samples using qRT-PCR (n = 5). (B) Altered expression levels of the three key genes after inhibition of methylation in HVSMCs (n = 3). *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Effect of ZC3H12A overexpression on cell proliferation, migration, and phenotypic transformation in vitro
In our preliminary literature review, many studies have demonstrated that ZC3H12A exerts a critical regulatory role in various cardiovascular diseases35,36,37,38,39. Therefore, we opted to validate ZC3H12A in vitro studies of TAAD to further explore its mechanistic role in TAAD. In order to explore the effects of the key genes on HVSMCs function, a ZC3H12A overexpression model was established. Results of the western blot analysis revealed that the expression of ZC3H12A was significantly higher in the OE-ZC3H12A group than that in the mock (1.98 ± 0.09 vs. 1.00 ± 0.12, P < 0.001) and OE-NC (1.98 ± 0.09 vs. 0.92 ± 0.18, P < 0.001) groups (Fig. 6A), validating our model. The results of the CCK8 analysis elucidated that the cell viability in the OE-ZC3H12A group was significantly higher than that in the mock (48 h: 1.36 ± 0.04 vs. 1.09 ± 0.05, P = 0.002; 72 h: 1.53 ± 0.05 vs. 1.33 ± 0.03, P < 0.001) and OE-NC (48 h: 1.36 ± 0.04 vs. 1.04 ± 0.05, P = 0.004; 72 h: 1.53 ± 0.05 vs. 1.31 ± 0.03, P = 0.002) groups after 48 and 72 h of culture (Fig. 6B), suggesting that ZC3H12A might promote the proliferation of HVSMCs. The transwell assay showed that ZC3H12A might also promote cell migration (Fig. 6C). In addition, the expression levels of phenotypic switch markers, including SM22α, SMA, and OPN, were found to be significantly downregulated in the OE-ZC3H12A group, compared to that in the mock (SM22α: 0.56 ± 0.01 vs. 1.00 ± 0.02, P < 0.001; SMA: 0.59 ± 0.07 vs. 1.00 ± 0.03, P = 0.003; OPN: 0.37 ± 0.11 vs. 1.02 ± 0.13, P = 0.019) and OE-NC (SM22α: 0.56 ± 0.01 vs. 0.94 ± 0.05, P < 0.001; SMA: 0.59 ± 0.07 vs. 0.95 ± 0.06, P = 0.006; OPN: 0.37 ± 0.11 vs. 0.96 ± 0.11, P = 0.028) group (Fig. 6D). The downregulation of SM22α and SMA indicated a weakening of the contractile phenotype characteristics in HVSMCs, while the decreased expression of OPN suggested that the synthetic phenotype characteristics of HVSMCs were also suppressed. Typically, phenotypic switching is accompanied by the downregulation of contractile markers and the upregulation of synthetic markers. However, in this experiment, overexpression of ZC3H12A resulted in the downregulation of both types of markers, suggesting that ZC3H12A may inhibit the complete phenotypic transition process of HVSMCs through an atypical mechanism, rather than merely suppressing either the contractile or synthetic characteristics40,41,42. These findings confirmed that ZC3H12A might promote the proliferation and migration of HVSMCs, whereas it inhibited their phenotypic switch in vitro.
In vitro experiments to study the effects of ZC3H12A overexpression on the proliferation, migration, and phenotypic transformation in HVSMCs. (A) Western blot analysis confirming the successful construction of the ZC3H12A overexpression model (n = 3). (B) The CCK8 assay to measure cell viability of HVSMCs after ZC3H12A overexpression (n = 6). (C) Transwell assays to detect changes in cell migration ability after overexpression of ZC3H12A. (D) qRT-PCR to detect changes in the mRNA expression levels of phenotypic switch markers after ZC3H12A overexpression (n = 3). *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Discussions
TAAD, a potentially life-threatening disease, requires greater attention in terms of disease manifestations, clinical course, and treatment urgency1. Moreover, an in-depth understanding of the molecular mechanism of TAAD is essential to improve the therapeutic approaches. In this study, transcriptome and methylation sequencing of vascular wall tissue samples was carried out to get a comprehensive understanding of the gene expression and methylation levels in TAAD. Through a series of bioinformatics analyses and machine learning algorithms, we finally identified three key methylated genes, including ZC3H12A, IRAK2, and CCL5. Tissue- and cell-based experiments confirmed the differential gene expression and methylation levels in patients with TAAD, as compared to the controls. Based on these findings, ZC3H12A, IRAK2, and CCL5 were proposed as diagnostic markers of TAAD. Among these candidates, ZC3H12A was confirmed to promote the proliferation and migration of HVSMCs in vitro while concurrently inhibiting their phenotypic switching, and its overall biological effect is inferred to facilitate the progression of TAAD.
Several previous studies have demonstrated that DNA methylation exerts significant influences on the pathogenesis of cardiovascular diseases, with a particular emphasis on TAAD9,16. Chen et al. utilized the Infinium Human Methylation 450 K BeadChip to conduct DNA methylation sequencing on aortic tissues obtained from four patients with TAAD and four control subjects9. Their analysis detected differential DNA methylation at CpG sites within five pivotal genes (Fas, ANGPT2, DUSP6, FARP1 and CARD6). Furthermore, they observed that the protein expression level of the gene Fas, which was associated with a hypomethylated position, increased by 1.78-fold, thereby suggesting a plausible role for DNA methylation in the regulation of gene expression. Liu P et al. employed whole-genome bisulfite sequencing to compare differential DNA methylation patterns in ascending aortic tissues between patients with TAAD and healthy control subjects16. Their investigation revealed a substantial number of differentially methylated regions, with the associated genes being notably enriched in pathways related to vasculature and heart development. Notably, Hox genes emerged as potential key players in the pathogenesis of TAAD. However, the integration of methylation and transcriptome sequencing analyses has not yet been applied in TAAD. In this study, we carried out methylation and transcriptome sequencing combined analyses using aortic vascular tissue samples, and the key methylated genes were identified as potential diagnostic markers of TAAD via integrative bioinformatics-based analyses.
Based on the differential and integrative analyses of the gene expression and methylation levels, this study identified 239 DMGs involved in nicotinamide nucleotide biosynthesis and metabolism. Nicotinamide riboside is essential for biological oxidation and reduction processes, and its glycosidicmoiety includes a positively charged ring, resulting in instability and cleavage43. Deficiency in nicotinamide adenine dinucleotide phosphate (NADPH) oxidase 1 (NOX1) can reduce the risk of angiotensin II-induced AD44. The involvement of nicotinamide in redox reactions is also implicated in the development of a range of cardiovascular diseases. Our study found multiple DEGs enriched in the biosynthesis and metabolic pathways of nicotinamide nucleotides, such as ENO1, NAMPT, and HK2, which are closely related to the redox process and exhibit complex cross-regulation. Enolase 1 (ENO1) is a key glycolytic enzyme. Increased glycolytic activity of ENO1 causes lactic acid buildup, lowering intracellular pH45. This indirectly inhibits NAD+-dependent deacetylases, affecting mitochondria and VSMCs’ stability. It may also reduce nicotinamide-to-NAD+ conversion, leading to genome-wide hypomethylation and contributing to TAAD vascular wall degeneration46,47. Nicotinamide phosphoribosyl transferase (NAMPT) functions as the rate-limiting enzyme in the biosynthesis of nicotinamide nucleotides, and alterations in its expression directly dictate the intracellular levels of NAD+48. In the context of TAAD, the overexpression of NAMPT results in an excessive utilization of NAD+ in mitochondrial oxidative phosphorylation49. Consequently, this depletes the amount of NAD+ in the cytoplasm that is required for the synthesis of the methylation donor, ultimately leading to genome-wide hypomethylation50. Additionally, the pro-inflammatory properties of NAMPT disrupt the redox balance by activating signaling pathways such as the NF-κB pathway, thereby aggravating damage to the vascular wall51. Hexokinase 2 (HK2) acts as the initial enzyme in glycolysis, and its activity influences the reduction state of NAD+. Overexpression of HK2 may interact with nicotinamide metabolism through metabolic reprogramming, leading to alterations in NAD+ levels52. This, in turn, affects the redox status of VSMCs, resulting in exacerbated cell death and fibrosis53. Alterations in redox balance may influence methylation regulation through multiple mechanisms. On one hand, reactive oxygen species (ROS) can directly oxidize and damage DNA methyltransferases (DNMTs), such as DNMT1, DNMT3A, and DNMT3B, thereby affecting their activity and stability. These DNMTs are crucial enzymes responsible for adding methyl groups to DNA, and any impairment in their function can disrupt the normal methylation pattern. On the other hand, changes in redox status can impact the synthesis and metabolism of the intracellular methyl donor, S-adenosylmethionine (SAM). The synthesis of SAM relies on the methionine cycle, and the activity of multiple enzymes involved in this cycle is regulated by the cellular redox state. Disruption of redox balance due to decreased synthesis and metabolism activity of nicotinamide nucleotides may interfere with the methionine cycle, leading to a reduction in SAM synthesis. Consequently, this limits the supply of methyl groups for DNA methylation, ultimately resulting in aberrant methylation. Therefore, in this study, these DMGs might affect the redox balance via altering the nicotinamide nucleotide biosynthesis and metabolism, resulting in aberrant methylation regulation and gene expression and ultimately contributing to TAAD progression. In addition, NOX1 might play an important role in this process; however, the specific upstream and downstream regulatory mechanisms still need to be experimentally elucidated.
In this study, ZC3H12A, IRAK2 and CCL5 were identified as potential diagnostic markers for TAAD. Our data confirmed that in tissue samples from patients with TAAD, CCL5 expression was significantly reduced, whereas ZC3H12A and IRAK2 expression levels were significantly elevated, compared to those in control samples. In addition, bioinformatics analysis and in vitro experimental results indicated that gene expression levels were significantly negatively regulated by their methylation levels. Reduced plasma levels of chemokine ligand 5 (CCL5), an inflammatory product, were shown to be useful in the diagnosis of TAAD54. This is consistent with our findings, suggesting that CCL5 methylation might be activated in TAAD to inhibit its expression, whereas the upstream inflammatory response might play a regulatory role. Similarly, IL-1R-associated kinase2 (IRAK2) is closely related to inflammation and involved in glucose metabolism and oxidative phosphorylation in the mitochondria55. Although the relationship between IRAK2 and TAAD has not been yet reported, IRAK2 was shown to be involved in immune cell infiltration in the cardiac tissue and could be used as a therapeutic target for stroke-heart syndrome56. Therefore, we speculated that aberrant IRAK2 methylation levels and gene expression in TAAD might affect the immune regulatory mechanisms in cardiac tissues. ZC3H12A, which encodes the MCP-1-induced protein (MCPIP1), acts as an innate immunomodulator and exerts anti-inflammatory effects through its ribonuclease and deubiquitinating enzyme activities57. ZC3H12A is involved in a range of autoimmune diseases, as well as cardiovascular diseases58, but its role in TAAD has not been reported. Herein, ZC3H12A methylation levels were suppressed in TAAD, resulting in upregulation of gene expression. In addition, ZC3H12A overexpression was found to promote the proliferation and migration of HVSMCs, while inhibit their phenotypic transformation. Overall, the role of ZC3H12A in TAAD is complex. On the one hand, it promotes HVSMCs proliferation and migration. CCK8 and transwell assays showed that ZC3H12A overexpression significantly boosts these capabilities in HVSMCs. Since aortic dissection is closely tied to HVSMCs proliferation, migration, and vascular wall remodeling, this effect may directly drive dissection progression. On the other hand, it inhibited phenotypic transformation. While this might protect blood vessels by preserving HVSMCs contractile function, it could also leave HVSMCs in a poorly differentiated state, making it hard to say if it’s protective or promotive40,42. A conflict emerges in this context. Phenotypic inhibition may stabilize the vascular wall by reducing inflammation, but proliferation and migration could offset this and worsen damage. Possible reasons include time-dependent effects, cell-type differences, and signaling pathway cross-talk. In conclusion, the phenotypes of HVSMCs observed in vitro are closely linked to the pathogenesis of TAAD. The role of ZC3H12A is complex and warrants further investigation.
The ZC3H12A, IRAK2, and CCL5 identified in this study not only serve as potential biomarkers for TAAD but also harbor significant potential for therapeutic development in their regulatory pathways. Currently, treatment options for TAAD are limited, and traditional methods struggle to fundamentally intervene in disease progression. Targeted epigenetic intervention of ZC3H12A can halt disease deterioration from the root of gene expression regulation. IRAK2 inhibitors hold promise for precise action at a key node in inflammatory signal transduction, making up for the deficiencies of existing anti-inflammatory treatments. The development of CCL5 neutralizing antibodies can effectively block immune cell infiltration, a crucial factor in aortic damage. Moreover, there may be a complex interaction network among these three genes that synergistically affects TAAD pathogenesis. Based on this, a comprehensive treatment strategy could be constructed in the future, such as combining targeted ZC3H12A intervention with IRAK2 inhibitors and CCL5 neutralizing antibodies to block disease progression through multiple targets and pathways, bringing new breakthroughs to TAAD treatment and improving patient prognosis. However, extensive experiments are still needed to verify their safety and efficacy from basic research to clinical application.
There are some limitations in this study. Firstly, the sample sizes we utilized were relatively modest, with only five cases compared to five controls. Within the same group, the presence of sample heterogeneity might have an impact on the experimental outcomes. Secondly, participants in this study were primarily recruited from the same region and shared a similar ethnic background, which may limit the generalizability of the research findings. Populations from different ethnic groups and geographical regions exhibit differences in genetic background, lifestyle, and environmental exposures, among other factors, which could influence gene expression and methylation patterns. Future studies should consider including participants from diverse ethnic groups and regions to enhance the external validity of the research results. Thirdly, although this study validated the effects of ZC3H12A on the proliferation, migration, and phenotypic switching of HVSMCs through in vitro experiments, these experimental results require further verification in more complex in vivo models. There are significant differences between the in vivo environment and in vitro culture conditions, which may affect the functional performance of genes. Future studies should utilize animal models to validate the biological functions identified in the current research and explore the relevant molecular mechanisms. In addition, due to limited research funding, we couldn’t conduct comprehensive validation experiments on all three genes simultaneously and instead chose ZC3H12A as the research focus for in vitro validation. In subsequent studies, pending research progress and sufficient funding availability, validation work on IRAK2 and CCL5 will be sequentially carried out to the extent possible. Finally, this study primarily focuses on the regulatory role of DNA methylation in gene expression, without considering other epigenetic regulatory mechanisms. However, epigenetic regulation is a complex network that also includes histone modifications, non-coding RNAs, and various other mechanisms. These mechanisms may interact with each other to collectively influence the occurrence and progression of TAAD. Future studies should adopt multi-omics integration analysis methods and comprehensively consider multiple epigenetic regulatory mechanisms to more thoroughly reveal the molecular mechanisms underlying TAAD.
Conclusions
By integrating transcriptome and methylation sequencing analyses, we identified three potential biomarkers of TAAD, including ZC3H12A, IRAK2, and CCL5. These gene expression levels were significantly negatively regulated by their methylation levels. Among them, in vitro experiments revealed that ZC3H12A promotes the proliferation and migration of HVSMCs, exerts a complex regulatory role in the process of cellular phenotypic switching, and the overall effect is to drive the TAAD progression.
Data availability
Sequence data that support the findings of this study have been deposited in GEO database with the primary accession code GSE267434, GSE269847, and GSE270377 (https://www.ncbi.nlm.nih.gov/geo/). Please contact corresponding author for further information.
References
Gudbjartsson, T. et al. Acute type A aortic dissection—A review. Scand. Cardiovasc. J. 54, 1–13. https://doi.org/10.1080/14017431.2019.1660401 (2020).
Jiang, T. & Si, L. Identification of the molecular mechanisms associated with acute type A aortic dissection through bioinformatics methods. Braz. J. Med. Biol. Res. 52, e8950. https://doi.org/10.1590/1414-431x20198950 (2019).
Pisano, C., Rita Balistreri, C., Fabio Triolo, O., Argano, V. & Ruvolo, G. Acute type A aortic dissection beyond the diameter. J. Heart Valve Dis. 25, 764–768 (2016).
Peng, X. et al. A bioinformatics analysis of the susceptibility genes in Stanford type A aortic dissection. J. Thorac. Dis. 15, 1694–1703. https://doi.org/10.21037/jtd-23-308 (2023).
Hines, G., Dracea, C. & Katz, D. S. Diagnosis and management of acute type A aortic dissection. Cardiol. Rev. 19, 226–232. https://doi.org/10.1097/CRD.0b013e3182203ed9 (2011).
Zhu, Y. et al. Type A aortic dissection-experience over 5 decades: JACC historical breakthroughs in perspective. J. Am. Coll Cardiol. 76, 1703–1713. https://doi.org/10.1016/j.jacc.2020.07.061 (2020).
Lian, R., Zhang, G., Yan, S., Sun, L. & Zhang, G. Identification of molecular regulatory features and markers for acute type A aortic dissection. Comput. Math. Methods Med. 2021, 6697848. https://doi.org/10.1155/2021/6697848 (2021).
Rylski, B., Schilling, O. & Czerny, M. Acute aortic dissection: Evidence, uncertainties, and future therapies. Eur. Heart J. 44, 813–821. https://doi.org/10.1093/eurheartj/ehac757 (2023).
Chen, Y. et al. DNA methylation alternation in Stanford- A acute aortic dissection. BMC Cardiovasc. Disord. 22, 455. https://doi.org/10.1186/s12872-022-02882-5 (2022).
Zhang, L., Lu, Q. & Chang, C. Epigenetics in Health and Disease. Adv. Exp. Med. Biol. 1253, 3–55. https://doi.org/10.1007/978-981-15-3449-2_1 (2020).
Boileau, A., Lindsay, M. E., Michel, J. B. & Devaux, Y. Epigenetics in ascending thoracic aortic aneurysm and dissection. Aorta (Stamford) 6, 1–12. https://doi.org/10.1055/s-0038-1639610 (2018).
Chen, C. et al. DNA methylation: From cancer biology to clinical perspectives. Front Biosci. (Landmark Ed.) 27, 326. https://doi.org/10.31083/j.fbl2712326 (2022).
Papait, R., Serio, S. & Condorelli, G. Role of the epigenome in heart failure. Physiol. Rev. 100, 1753–1777. https://doi.org/10.1152/physrev.00037.2019 (2020).
Dai, Y., Chen, D. & Xu, T. DNA methylation aberrant in atherosclerosis. Front. Pharmacol. 13, 815977. https://doi.org/10.3389/fphar.2022.815977 (2022).
Xia, Y., Brewer, A. & Bell, J. T. DNA methylation signatures of incident coronary heart disease: Findings from epigenome-wide association studies. Clin. Epigenet. 13, 186. https://doi.org/10.1186/s13148-021-01175-6 (2021).
Liu, P. et al. Altered DNA methylation pattern reveals epigenetic regulation of Hox genes in thoracic aortic dissection and serves as a biomarker in disease diagnosis. Clin. Epigenet. 13, 124. https://doi.org/10.1186/s13148-021-01110-9 (2021).
Pan, S. et al. DNA methylome analysis reveals distinct epigenetic patterns of ascending aortic dissection and bicuspid aortic valve. Cardiovasc. Res. 113, 692–704. https://doi.org/10.1093/cvr/cvx050 (2017).
Liu, Q. et al. Epigenome-wide DNA methylation and transcriptome profiling of localized and locally advanced prostate cancer: Uncovering new molecular markers. Genomics 114, 110474. https://doi.org/10.1016/j.ygeno.2022.110474 (2022).
Hiratzka, L. F. et al. 2010 ACCF/AHA/AATS/ACR/ASA/SCA/SCAI/SIR/STS/SVM guidelines for the diagnosis and management of patients with thoracic aortic disease: executive summary. A report of the American College of Cardiology Foundation/American Heart Association Task Force on Practice Guidelines, American Association for Thoracic Surgery, American College of Radiology, American Stroke Association, Society of Cardiovascular Anesthesiologists, Society for Cardiovascular Angiography and Interventions, Society of Interventional Radiology, Society of Thoracic Surgeons, and Society for Vascular Medicine. Catheter. Cardiovasc. Interv. 76, E43–86. https://doi.org/10.1002/ccd.22537 (2010).
Jiang, H., Lei, R., Ding, S. W. & Zhu, S. Skewer: A fast and accurate adapter trimmer for next-generation sequencing paired-end reads. BMC Bioinform. 15, 182. https://doi.org/10.1186/1471-2105-15-182 (2014).
de Sena Brandine, G. & Smith, A. D. Falco: High-speed FastQC emulation for quality control of sequencing data. Research 8, 1874. https://doi.org/10.12688/f1000research.21142.2 (2019).
Tamura, T. et al. Virological characteristics of the SARS-CoV-2 XBB variant derived from recombination of two Omicron subvariants. Nat. Commun. 14, 2800. https://doi.org/10.1038/s41467-023-38435-3 (2023).
Shumate, A., Wong, B., Pertea, G. & Pertea, M. Improved transcriptome assembly using a hybrid of long and short reads with StringTie. PLoS Comput. Biol. 18, e1009730. https://doi.org/10.1371/journal.pcbi.1009730 (2022).
Teschendorff, A. E. et al. A beta-mixture quantile normalization method for correcting probe design bias in Illumina Infinium 450 k DNA methylation data. Bioinformatics 29, 189–196. https://doi.org/10.1093/bioinformatics/bts680 (2013).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47. https://doi.org/10.1093/nar/gkv007 (2015).
Zhang, L. et al. Revealing the pathogenic changes of PAH based on multiomics characteristics. J. Transl. Med. 17, 231. https://doi.org/10.1186/s12967-019-1981-5 (2019).
Kanehisa, M., Furumichi, M., Sato, Y., Matsuura, Y. & Ishiguro-Watanabe, M. KEGG: Biological systems database as a model of the real world. Nucleic Acids Res. 53, D672–D677. https://doi.org/10.1093/nar/gkae909 (2025).
da Huang, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57. https://doi.org/10.1038/nprot.2008.211 (2009).
Szklarczyk, D. et al. The STRING database in 2017: Quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Res. 45, D362-d368. https://doi.org/10.1093/nar/gkw937 (2017).
Shannon, P. et al. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504. https://doi.org/10.1101/gr.1239303 (2003).
Sanz, H., Valim, C., Vegas, E., Oller, J. M. & Reverter, F. SVM-RFE: Selection and visualization of the most relevant features through non-linear kernels. BMC Bioinform. 19, 432. https://doi.org/10.1186/s12859-018-2451-4 (2018).
Gao, J., Kwan, P. W. & Shi, D. Sparse kernel learning with LASSO and Bayesian inference algorithm. Neural Netw. 23, 257–264. https://doi.org/10.1016/j.neunet.2009.07.001 (2010).
Qi, H. et al. Ammonium tetrathiomolybdate relieves oxidative stress in cisplatin-induced acute kidney injury via NRF2 signaling pathway. Cell Death Discovery 9, 259. https://doi.org/10.1038/s41420-023-01564-1 (2023).
Xiao, W. et al. Branched-chain α-ketoacids aerobically activate HIF1α signalling in vascular cells. Nat. Metab. 6, 2138–2156. https://doi.org/10.1038/s42255-024-01150-4 (2024).
Niu, J., Jin, Z., Kim, H. & Kolattukudy, P. E. MCP-1-induced protein attenuates post-infarct cardiac remodeling and dysfunction through mitigating NF-κB activation and suppressing inflammation-associated microRNA expression. Basic Res. Cardiol. 110, 26. https://doi.org/10.1007/s00395-015-0483-8 (2015).
Ribeiro, A. et al. Macrophage-specific MCPIP1/regnase-1 attenuates kidney ischemia-reperfusion injury by shaping the local inflammatory response and tissue regeneration. Cells https://doi.org/10.3390/cells11030397 (2022).
Zou, X. et al. MCPIP-1 knockdown enhances endothelial colony-forming cell angiogenesis via the TFRC/AKT/mTOR signaling pathway in the ischemic penumbra of MCAO mice. Exp. Neurol. 369, 114532. https://doi.org/10.1016/j.expneurol.2023.114532 (2023).
Jin, Z. et al. Essential role of endothelial MCPIP in vascular integrity and post-ischemic remodeling. Int. J. Mol. Sci. https://doi.org/10.3390/ijms20010172 (2019).
Xue, M. et al. Up-regulated MCPIP1 in abdominal aortic aneurysm is associated with vascular smooth muscle cell apoptosis and MMPs production. Biosci. Rep. https://doi.org/10.1042/bsr20191252 (2019).
Zhang, N. et al. Assessing serum levels of SM22α as a new biomarker for patients with aortic aneurysm/dissection. PLoS ONE 17, e0264942. https://doi.org/10.1371/journal.pone.0264942 (2022).
Xia, X. et al. The relationship between estrogen-induced phenotypic transformation and proliferation of vascular smooth muscle and hypertensive intracerebral hemorrhage. Ann. Transl. Med. 8, 762. https://doi.org/10.21037/atm-20-4567 (2020).
Zhu, S. B. et al. TGF-β1 induces human aortic vascular smooth muscle cell phenotype switch through PI3K/AKT/ID2 signaling. Am. J. Transl. Res. 7, 2764–2774 (2015).
Kim, H. J. & Benner, S. A. A direct prebiotic synthesis of nicotinamide nucleotide. Chemistry 24, 581–584. https://doi.org/10.1002/chem.201705394 (2018).
Hu, X., Jiang, W., Wang, Z., Li, L. & Hu, Z. NOX1 negatively modulates fibulin-5 in vascular smooth muscle cells to affect aortic dissection. Biol. Pharm. Bull. 42, 1464–1470. https://doi.org/10.1248/bpb.b18-01012 (2019).
Yang, T. et al. Enolase 1 regulates stem cell-like properties in gastric cancer cells by stimulating glycolysis. Cell Death Dis. 11, 870. https://doi.org/10.1038/s41419-020-03087-4 (2020).
Duarte-Pereira, S., Matos, S., Oliveira, J. L. & Silva, R. M. Study of NAD-interacting proteins highlights the extent of NAD regulatory roles in the cell and its potential as a therapeutic target. J. Integr. Bioinform. https://doi.org/10.1515/jib-2022-0049 (2023).
Besselink, N. et al. The genome-wide mutational consequences of DNA hypomethylation. Sci. Rep. 13, 6874. https://doi.org/10.1038/s41598-023-33932-3 (2023).
Revollo, J. R., Grimm, A. A. & Imai, S. The regulation of nicotinamide adenine dinucleotide biosynthesis by Nampt/PBEF/visfatin in mammals. Curr. Opin. Gastroenterol. 23, 164–170. https://doi.org/10.1097/MOG.0b013e32801b3c8f (2007).
Wang, P., Li, W. L., Liu, J. M. & Miao, C. Y. NAMPT and NAMPT-controlled NAD metabolism in vascular repair. J. Cardiovasc. Pharmacol. 67, 474–481. https://doi.org/10.1097/fjc.0000000000000332 (2016).
Hocaoglu, H., Wang, L., Yang, M., Yue, S. & Sieber, M. Heritable shifts in redox metabolites during mitochondrial quiescence reprogramme progeny metabolism. Nat. Metab. 3, 1259–1274. https://doi.org/10.1038/s42255-021-00450-3 (2021).
Li, D. et al. NAMPT mediates PDGF-induced pulmonary arterial smooth muscle cell proliferation by TLR4/NF-κB/PLK4 signaling pathway. Eur. J. Pharmacol. 961, 176151. https://doi.org/10.1016/j.ejphar.2023.176151 (2023).
He, K. et al. The role of HK2 in tumorigenesis and development: Potential for targeted therapy with natural products. Int. J. Med. Sci. 22, 790–805. https://doi.org/10.7150/ijms.105553 (2025).
Zhang, W. et al. Nicotinamide phosphoribosyltransferase in NAD(+) metabolism: Physiological and pathophysiological implications. Cell Death Discov. 11, 371. https://doi.org/10.1038/s41420-025-02672-w (2025).
Liu, X., Zhang, A., Dong, N. & Wang, Z. Expression of TLR4 Is upregulated in patients with sporadic acute Stanford type A aortic dissection. Cardiol. Res. Pract. 2022, 3806462. https://doi.org/10.1155/2022/3806462 (2022).
Zhou, H. et al. IL-1 induces mitochondrial translocation of IRAK2 to suppress oxidative metabolism in adipocytes. Nat. Immunol. 21, 1219–1231. https://doi.org/10.1038/s41590-020-0750-1 (2020).
Zheng, J., Ma, Y., Guo, X. & Wu, J. Immunological characterization of stroke-heart syndrome and identification of inflammatory therapeutic targets. Front. Immunol. 14, 1227104. https://doi.org/10.3389/fimmu.2023.1227104 (2023).
Cifuentes, R. A., Cruz-Tapias, P., Rojas-Villarraga, A. & Anaya, J. M. ZC3H12A (MCPIP1): Molecular characteristics and clinical implications. Clin. Chim. Acta 411, 1862–1868. https://doi.org/10.1016/j.cca.2010.08.033 (2010).
Yan, B., Guo, Y., Gui, Y., Jiang, Z. S. & Zheng, X. L. Multifunctional RNase MCPIP1 and its role in cardiovascular diseases. Curr. Med. Chem. 28, 3385–3405. https://doi.org/10.2174/0929867327999201113100918 (2021).
Acknowledgements
We acknowledge Jingneng Biotechnology (Shanghai) Co., Ltd. for providing sequencing services.
Funding
This work was supported by Tianjin Science and Technology Plan Project (24JCYBJC01480); Tianjin Natural Science Foundation Key Project (21JCZDJC00610); Tianjin Science and Technology Plan Project (22JCZDJC00580); Tianjin Education Commission Research Program Project (2022YGYB09); Tianjin Key Medical Discipline Construction Project (TJYXZDXK-3-030C).
Author information
Authors and Affiliations
Contributions
Qingliang Chen and Jie Geng conceived the project; Chao Chang, Meng Wang and Yunpeng Bai designed experiments; Chao Chang, Meng Wang, Yunpeng Bai and Kai Zhang performed the majority of experiments with the support from Qingliang Chen and Jie Geng; Chao Chang and Meng Wang collected data; Yunpeng Bai and Kai Zhang analyzed data; Chao Chang wrote the manuscript and all authors contributed edits.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
This study complied with the Helsinki Declaration and was approved by the ethics committee of Tianjin Chest Hospital (2023KY-028-01). All participants signed informed consent.
Consent for publication
Informed consent was obtained from all individual participants included in the study.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Chang, C., Wang, M., Bai, Y. et al. Genome wide DNA methylation and transcriptome integration analysis reveals potential markers in type A aortic dissection pathogenesis. Sci Rep 15, 42452 (2025). https://doi.org/10.1038/s41598-025-26684-9
Received:
Accepted:
Published:
Version of record:
DOI: https://doi.org/10.1038/s41598-025-26684-9